Abstract
Acute leukaemia is the major subtype of paediatric cancer with a cumulative risk of 1 in 2000 for children up to the age of 15 years. Childhood acute lymphoblastic leukaemia (ALL) is a biologically and clinically diverse disease with distinctive subtypes; multiple chromosomal translocations exist within the subtypes and each carries its own prognostic relevance. The most common chromosome translocation observed is the t(12;21) that results in an in-frame fusion between the first five exons of ETV6 (TEL) and almost the entire coding region of RUNX1 (AML1).
The natural history of childhood ALL is almost entirely clinically silent and is well advanced at the point of diagnosis. It has, however, been possible to backtrack this process through molecular analysis of appropriate clinical samples: (i) leukaemic clones in monozygotic twins that are either concordant or discordant for ALL; (ii) archived neonatal blood spots or Guthrie cards from individuals who later developed leukaemia; and (iii) stored, viable cord blood cells.
Here, we outline our studies on the aetiology and pathology of childhood ALL that provide molecular evidence for a monoclonal, prenatal origin of ETV6-RUNX1+ leukaemia in monozygotic identical twins. We provide mechanistic support for the concept that altered patterns of infection during early childhood can deliver the necessary promotional drive for the progression of ETV6-RUNX1+ pre-leukaemic cells into a postnatal overt leukaemia.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Acute Lymphoblastic Leukaemia (ALL), a disease of the bone marrow, accounts for about 30 % of cancer diagnosed in children under the age of 15 years (Dickinson 2005). The disease is biologically and clinically diverse with distinctive subtypes, each characterized by an association between age at presentation of overt leukaemia and various recurrent genetic alterations. Multiple chromosomal translocations exist within the subtypes and each carries its own prognostic relevance (reviewed in (Rowley et al. 2015)).
The most common chromosome translocation observed in ALL is the t(12;21) (Golub et al. 1995; Romana et al. 1995). The translocation results in an in-frame fusion between the first five exons of ETV6 and almost the entire coding region of RUNX1; bringing together the PTD and repression domains of ETV6 and the DNA binding (RHD), repression and transactivation domains of RUNX1 (Golub et al. 1995; Romana et al. 1995), Fig. 14.1. Both RUNX1 and ETV6 are important transcription factors required for normal haematopoiesis (Okuda et al. 1996; Wang et al. 1996).
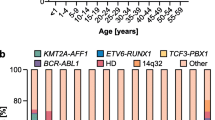
Although cryptic at the level of karyotype, both FISH and RT-PCR studies have shown the ETV6-RUNX1 fusion to be present in around 25 % of cases of B-cell precursor ALL (BCP-ALL), with an age related distribution peak of 2–5 years that matches the peak of incidence of the leukaemia (Shurtleff et al. 1995).
To further our understanding of the aetiology and natural history of childhood ALL, two key questions have been addressed; (1) precisely when and how is the ETV6-RUNX1 fusion gene generated in the development and clonal evolution of overt leukaemia and (2) whether occurrence of the fusion gene is a leukaemia initiating event sufficient for overt leukaemia.
2 Identical Twins with Concordant Leukaemia
The notion that genetic changes necessary for overt leukaemia might occur before birth was raised over 50 years ago and based on studies of concordant leukaemia in identical (monozygotic) twins (Clarkson and Boyse 1971). Clarkson and Boyse suggested that a demonstration of shared, non-constitutive, cytogenetic abnormalities in leukaemic cells isolated from such twin pairs might provide a prenatal, monoclonal explanation for the concordant leukaemia.
Monozygotic identical twins occur when a single egg is fertilized by a single sperm to form one zygote. Subsequently, the zygote will divide into two separate embryos, the timing of which is critical to the formation of the placenta(s) and amniotic sac(s), Fig. 14.2. If the zygote splits within the first 3 days, two separate placentas and amniotic sacs are formed (dichorionic and diamniotic). If the split occurs between days four and nine after fertilization then the twins will share one placenta with separate sacs (monochorionic and diamniotic) and notably will share their supply of blood. 60 % of monozygotic twins are in this category. If the split occurs after 9 days then the twins will share a single placenta and sac (monochorionic and monoamniotic). Non-identical or fraternal twins result from the fertilization of two separate eggs by two separate sperm (dizygotic) and consequently do not share their blood supplies. Monozygotic twins are genetically identical unless there has been a mutation in development.
Placental status in twin embryos. The schematic shows the placenta as a red oval, the amnion a grey oval and the chorion in blue (Frequency data is taken from Strong and Corney 1967)
Over seventy pairs of monozygotic twins with concordant acute leukaemia have been recorded in the literature (Greaves et al. 2003) and such cases usually share the same morphological and immunological subtype of leukaemia and development of their clinical symptoms usually occurs within a short time of each other (Greaves et al. 2003). The concordance rate for leukaemia in infant twins (<1 year) is almost 100 %, while that for older identical twins, including those with ETV6-RUNX1+ ALL, is less at 10–15 % (Greaves et al. 2003) suggesting the occurrence of additional, postnatal, genetic events.
3 Molecular Evidence for a Monoclonal, Prenatal Origin of ETV6-RUNX1+ Leukaemia in Identical Twins
Chimeric fusion genes are formed by normal, error-prone repair of DNA double-strand breaks (DSBs) (Wiemels and Greaves 1999). Gene fusions between ETV6 and RUNX1 involve the noncoding introns of each gene and the breaks are both scattered and diverse within the respective breakpoint cluster regions (Golub et al. 1995). The breakpoints on chromosome 12 cluster within a single 12 kb intron of ETV6 whereas those on chromosome 21 occur mainly within the large (~150 kb) first intron of RUNX1 (Wiemels and Greaves 1999) and Fig. 14.3). As a consequence, each break and subsequent fusion junction is both clonotypic and patient specific at the DNA level and therefore the genomic fusion sequence provides a unique marker of clonal identity and a stable imprint of single cell origin. We reasoned that cloning and sequencing of the ETV6-RUNX1 fusion region in twin pairs with concordant t(12;21) childhood ALL should provide unambiguous evidence for any clonal relationship as well as provide clues to the mechanism of recombination. We first cloned the ETV6-RUNX1 fusion gene from a pair of monozygotic twins who were diagnosed at ages 3 years 6 months and 4 years 10 months respectively (Ford et al. 1998). Sequence analysis of twin 1 identified nucleotides within intron 5 of the ETV6 gene and intron 1 of RUNX1. An identical fusion sequence in twin 2 confirmed that the twin leukaemias were derivatives of the same single cell or clone in which the unique and non-constitutive ETV6-RUNX1 fusion had first arisen. Clonal identity was further supported by the finding that the leukaemic cells in the two twins shared an identical rearranged immunoglobulin heavy chain gene (IGH) allele (Ford et al. 1998). The most reasonable explanation for this finding was a single cell origin of the ETV6-RUNX1 fusion in one foetus in utero , followed by an intraplacental metastasis of clonal progeny to the other twin via the shared vascular anastomoses.
Further unequivocal evidence to support the pre-natal origin of childhood ETV6-RUNX1+ leukaemia was provided by the scrutiny of neo-natal blood spots, or Guthrie cards, taken at birth from a second pair of identical twins with concordant leukaemia. Guthrie cards are prepared by heel prick in the first days of life and are usually used for detection of inherited mutations and in screening for inborn errors of metabolism such as phenylketonuria. Given the natural history of childhood leukaemia, the assumption was that concordant identical twins with ETV6-RUNX1+ ALL might have cells with fusion gene sequences already present in their blood at birth. A simple way of testing this idea was through a backtracking analysis of the Guthrie cards of such patients. We studied a pair of identical twins diagnosed with concordant ETV6-RUNX1+ ALL at age 4 and for whom Guthrie cards were still available (Wiemels et al. 1999a). Diagnostic DNA was first used to establish that the sequence of the ETV6-RUNX1 fusion was identical between the twin pairs and then individual segments of Guthrie card were used to confirm the presence of the fusion gene in the blood at birth and consequently the in utero clonal origin of the leukaemia.
A third twin pair provided new and unexpected insight into the time frame necessary for critical sequential events to occur. Unusually, these twins were diagnosed with ETV6-RUNX1+ ALL over 8 years apart; at ages five and fourteen (Wiemels et al. 1999b). Cloning and sequencing of the ETV6-RUNX1 fusion present in each twin showed perfect identity, again indicative of a single cell origin. However at the time when the first twin was diagnosed, the bone marrow of the second twin was haematologically normal and remained so for 8 years. Retrospective analysis by PCR of an archived bone marrow smear from the then ‘unaffected’ twin showed the presumptive ETV6-RUNX1+ pre-leukaemic clone to be present 8 years before clinical diagnosis of ALL. These data suggest that subsequent to initiation of a prenatal, pre-leukaemic clone, almost certainly as a result of ETV6-RUNX1 fusion alone, the period required for appearance of overt leukaemia can be both extremely variable and protracted, with latency of up to 14 years (Wiemels et al. 1999b).
Since ETV6-RUNX1+ leukaemia in twins is no different, biologically or clinically from that seen in single children, at least some singletons are also likely to have prenatal initiation of leukaemia. We used nine sets of diagnostic samples with paired blood spots to backtrack the fusion gene to birth in non-twinned children with ETV6-RUNX1+ ALL and provided more direct evidence that this disease can at least initiate in utero (Wiemels et al. 1999a).
4 Is an ETV6-RUNX1 Fusion Gene Sufficient for Overt Leukaemia?
Taken together, these studies provide strong evidence, in most cases, for a prenatal origin of ETV6-RUNX1+ leukaemia. However, it is now clear that not all individuals with a ETV6-RUNX1 fusion gene go on to develop overt disease. In a retrospective study of over 600 normal new born cord bloods, we showed the frequency of fusion gene positive cord bloods to be 1 %; approximately 100 times the collective frequency of overt, clinically diagnosed leukaemia with ETV6-RUNX1 fusion (Mori et al. 2002). The data, along with the modest rate of twin concordance (5–10 %), supports the view that detectable ETV6-RUNX1+ cells in healthy children represent expanded clones of pre-leukaemic cells that can remain pathologically and clinically silent or covert in the absence of additional, postnatal genetic hits, perhaps for up to 14 years. However, a postnatal fusion of ETV6 and RUNX1 in some cases, cannot be ruled out.
5 ETV6-RUNX1 as an Initiating or ‘Founder’ Event in ALL
A number of studies on singletons and pairs of monozygotic twins with ETV6-RUNX1+ leukaemia have now been described that shed light on the important genetic events ‘secondary’ to gene fusion (Ford et al. 1998; Wiemels et al. 1999b; Broadfield et al. 2004; Teuffel et al. 2004; Maia et al. 2004; Bateman et al. 2010; Bungaro et al. 2008; Alpar et al. 2015). FISH analyses at diagnosis of ETV6-RUNX1+ ALL show the fusion gene to be present in every leukaemic cell (Anderson et al. 2011) and the majority of cases also show some sub clonal deletion of the non-translocated ‘normal’ ETV6 allele (Raynaud et al. 1996; Kempski and Sturt 2000). The deletions vary in size between patients and both FISH and loss of heterozygosity (LOH) studies show that 73 % of ETV6-RUNX1+ cases have a partially or fully deleted second ETV6 allele (Patel et al. 2003). Although the second ETV6 allele was identified in the remaining patients, no ETV6 expression was detected. Taken together, these findings support the hypothesis that loss of ETV6 expression may be a critical secondary event for leukaemogenesis in ETV6-RUNX1+ ALL and the assumption that ETV6 can act as a tumor suppressor gene .
Recurrent copy number alterations (CNAs) are the likely “driver” events that contribute critically to clonal diversification and selection. In ETV6-RUNX1+ ALL they typically include deletions of genes involved in B-cell development and differentiation such as PAX5, BTG1, the RAG family and the wild-type copy of ETV6 (Mullighan 2012). If deletions of the normal copy of ETV6 and indeed all other recurrent ‘driver’ CNAs are consistently secondary to ETV6-RUNX1 fusion and therefore postnatal, then a testable prediction would be that these deletions should be distinct or different within monozygotic twin pairs. To address this idea we first used paired interphase FISH and SNP array information to identify recurrent CNAs in 5 pairs of twins with ETV6-RUNX1+ ALL. Sporadic CNAs classified as non-functional “passengers” were either identical (4/19) in the twin pairs and thought to precede the ETV6-RUNX1 fusion event, or were distinct (15/19) (Bateman et al. 2010). Significantly, all 32 CNAs identified between the twin pairs that were regarded as being ‘drivers’ of leukaemia were discordant (Fig. 14.4). As expected, this discordance was further reflected by singletons and twin pairs that shared the same ETV6 deletion but harboured different deletion boundaries (Ford et al. 2001; Maia et al. 2001; Bateman et al. 2010).
Genomics of ETV6-RUNX1 ALL in monozygotic twins. Combined data from 5 sets of twins with concordant ETV6-RUNX1+ ALL (Data taken from Bateman et al. 2015)
In a second study on a single set of monozygotic twins with ETV6-RUNX1+ ALL we used a whole genome sequencing approach to better determine the developmental timing of these events. We identified the ETV6-RUNX1 translocation to be the only recognised fusion product shared by the twins and, despite the presence of LOH in 27 and 41 cytoband regions respectively, we found the only other mutation in common to be an inactivating germline mutation of neurofibromatosis type 1 (Ma et al. 2013). Despite a paucity of single base and indel ‘driver’ mutations within the leukaemia clones, none of the mutations identified were found to be shared, further supporting the concept that these genetic changes are both secondary to ETV6-RUNX1 fusion and post-natal.
6 A Candidate Pre-leukaemic Stem Cell Population with an Early B Lineage Phenotype
Childhood ALL is associated with a rare population of CD34+, CD38−/ low, CD19+ cells not usually detectable in normal bone marrow (Hotfilder et al. 2002; Castor et al. 2005; Hong et al. 2008; le Viseur et al. 2008) and is accompanied by clonal rearrangement of the IGH genes, indicative of a pre-B cell phenotype. In addition, ALLs characterised by ETV6-RUNX1 fusion maintain a phenotype of CD10+, CD19+, along with recombinase gene activity (RAG) and expression of TdT. The pre-B cell however, is not necessarily the cell in which the functional impact of the ETV6-RUNX1 fusion gene is first observed. In two early studies, distinct and specific B-cell receptor gene rearrangements were identified in one set of twins (Teuffel et al. 2004) suggesting that separate pre-leukaemic clones were already present at birth. However, Bungaro and colleagues (Bungaro et al. 2008) identified both shared and distinct rearrangements at diagnosis in a set of monozygotic twins with dichorionic placentas. Not only was this suggestive of a common clonal origin in utero but, in this case, is also indicative of the passage of cells from one foetus to the other via the blood system of the Mother. In a more detailed screen of IG/TCR rearrangements in 5 pairs of twins with concordant ETV6-RUNX1+ ALL, we revealed the pre-leukaemic initiating function of the ETV6-RUNX1 fusion to be associated with clonal expansion of an early foetal B-cell (Alpar et al. 2015). In all pairs of twins studied, the cells carried identical incomplete or complete IGH variable-diversity-joining (VDJ) regions together with substantial, sub-clonal and divergent rearrangements. In addition, most descendent cells with stem cell (self-renewal) activity were shared and maintained in both twins after birth and provided an opportunity for necessary postnatal, secondary genetic hits to occur (Fig. 14.5).
Evolutionary sequence of clonal immunogenotypic markers identified in a pair of monozygotic twins with concordant ETV6-RUNX1+ ALL. Rearrangements are depicted both in the context of an in utero origin of pre-leukaemia, perhaps occurring in a progenitor cell already committed to the B cell lineage and the diverging postnatal evolution of the overt leukaemia (Alpar et al. 2015) (CNA data was adopted from Bateman et al. (2015). HSC hematopoietic stem cell )
The notion that ETV6-RUNX1 fusion is an initiating event, insufficient itself for overt leukaemia, was also confirmed in a seminal study of identical twins with discordant ETV6-RUNX1+ leukaemia (Hong et al. 2008). One twin was diagnosed with ETV6-RUNX1+ pre-B cell ALL at age 2, while the other twin has remained healthy for over 10 years. The bone marrow of the leukaemic child contained the CD34+, CD38−/low, CD19+ cancer-propagating population while the blasts, as well as presenting the ETV6-RUNX1 fusion, showed additional loss of the nontranslocated ETV6 allele, loss of one copy of PAX5 and gain of 10p (Bateman et al. 2010), Fig. 14.6). Immuno-FISH for CD19 protein and the ETV6-RUNX1 fusion in the peripheral blood pre-B cells of the healthy child detected the fusion gene at a frequency of ~0.1 %, but these cells did not show additional chromosome aberrations. Cloning and sequencing of the respective ETV6-RUNX1 fusion junctions revealed their complete identity and added further support to the in utero origin of the pre-leukaemic clone (AF unpublished).
Interphase FISH to confirm CNA status in cells from monozygotic twins clinically discordant for ETV6-RUNX1+ ALL. (a) A normal bone marrow cell from the healthy twin (6b) shows 2 red (RUNX1 gene), 2 green (ETV6 gene), 2 pink (PAX5 gene), and 2 brown (chromosome 10p) signals, respectively. (b) A leukaemic cell from twin 6a shows the ETV6-RUNX1 gene fusion, 1 normal RUNX1 and the remnant from the rearranged RUNX1 gene, 1 copy of PAX5 and 3 signals for 10p. (c) A ETV6-RUNX1 fusion gene positive cell in the unaffected twin (6b) shows 1 ETV6-RUNX1 gene fusion, the normal RUNX1 and the remnant from rearranged RUNX1, 1 copy of normal ETV6, 2 copies of PAX5 and 2 signals for chromose 10p to show that loss of ETV6 and PAX5 and gain of 10p are not observed in the preleukaemic, ETV6-RUNX1+ cells of the unaffected twin. Five cells with the ETV6-RUNX1 fusion were identified in the unaffected twin (twin 6b), of a total of 4251 scored (Bateman et al. 2015)
7 An Infectious Aetiology for Childhood ALL?
Given the age related peak incidence of 2–5 years for BCP-ALL, it has long been proposed that infection (s) in childhood might accelerate the transformation of ETV6-RUNX1+ pre-leukaemic cells to overt leukaemia (Kinlen 1988; Greaves 1988). Although we and others (MacKenzie et al. 1999, 2001; McNally and Eden 2004) have not identified any specific exogenous viral sequences, epidemiological studies support the notion that a relationship exists between improved social conditions and childhood ALL (Dockerty et al. 2001). In a highly protective unchallenged or ‘hygienic’ environment a delayed exposure of infants to an otherwise common infection may trigger a rare abnormal immune response by selection and expansion of pre-leukaemic (ETV6-RUNX1+) clones (Greaves 2006, 1988). In this context, we have shown that altered cytokine environments in the context of inflammation , such as TGFβ, can eliminate normal pre-B cell clones from the repertoire and support the selective outgrowth of pre-B cell clones that already harbour ETV6-RUNX1 (Ford et al. 2009).
Somatic recombination and mutation of IGH genes to create antibody diversity in B cells requires the proteins encoded by the RAG1 and RAG2 genes that introduce DNA double-strand breaks and the subsequent recombination of V(D)J gene segments (Oettinger et al. 1990). The enzyme activation-induced cytidine deaminase (AICDA) subsequently then enables somatic hypermutation of V region genes followed by class switching (Li et al. 2004). During normal B cell ontogeny the activities of these enzymes are kept strictly segregated (Hardy and Hayakawa 2001).
8 Genetic Changes that Complement ETV6-RUNX1 Fusion
The presence of V(D)J recombination signal sequences (RSS) close to CNAs commonly deleted in ETV6-RUNX1+ ALL has suggested a role for aberrant RAG endonuclease targeting at these loci (Zhang and Swanson 2008; Mullighan et al. 2008). To obtain a more detailed picture of these secondary genetic events we carried out genome analysis of diagnostic samples from 57 cases of ETV6-RUNX1+ ALL paired with matched remission (constitutive) DNA. We performed low-depth whole-genome sequencing and structural variation analysis on the leukaemic samples of 51 cases and used exome sequencing of 56 cases to search for recurrent somatic variants (Papaemmanuil et al. 2014).
We observed a paucity of recurrent coding-region mutations but resolved 354 of 523 structural variations at breakpoint sites to base-pair resolution. We searched for conserved RSS, along with proposed recognition motifs for the APOBEC family of enzymes that deaminate cytosine to uracil and for the presence of CpGs (Tsai et al. 2008). Although we did not find conserved RSS motifs near the breakpoints of the founding ETV6-RUNX1 rearrangement, consistent with this rearrangement arising in a very early B-lineage progenitor cell, enrichment for RSS was particularly prominent at gene deletions targeting known B-cell ALL genes such as ETV6 and BTG1. We did not observe specific enrichment of CpGs or any of the proposed AICDA recognition motifs at breakpoint junctions relative to other cancers; however another comprehensive analysis of translocation breakpoints did reveal a breakage mechanism that involved the RAG complex acting at AICDA-deaminated methyl-CpGs (Tsai et al. 2008). Nonetheless, 43 % of the 354 deletion breakpoints in our ETV6-RUNX1 study showed conserved RAG recognition motifs: CACACTG – spacer – ACAAAAACC; compared to a complete absence of RSS at over 12,000 breakpoints examined in breast, pancreatic and prostate cancer. These observations signify the existence of a lymphoid-specific endogenous mutagenesis programme (Papaemmanuil et al. 2014; Swaminathan et al. 2015).
To test the hypothesis that, in the context of inflammation , RAGs and AICDA can cooperate to induce secondary genetic lesions and accelerate transformation of a ETV6-RUNX1 pre-leukaemic clone to overt leukaemia, we subjected ETV6-RUNX1 expressing pre-B cells to consecutive rounds of stimulation with the inflammatory mimic lipopolysaccharide (LPS) in the presence or absence of IL7. Signalling through the IL7-R has been shown to safeguard human pre-B cells from premature activation of AICDA (Swaminathan et al. 2015). We noted both upregulation of RAG1/RAG2 mRNA levels in the presence of ETV6-RUNX1 alone and the subsequent upregulation of AICDA by over 20 fold in the presence of LPS and absence of IL7 (Swaminathan et al. 2015). Intravenous injection of these vulnerable cells into NOD/SCID recipients triggered development of leukaemia within 3 weeks. In contrast, pre-B cells isolated from Aicda −/− or Rag1−/− animals respectively delayed or abrogated leukaemia development (Swaminathan et al. 2015). These data provide sound genetic evidence that, in the context of inflammatory/repetitive infectious stimulation, clonal evolution of pre-leukaemic ETV6-RUNX1+ pre-B cells requires both RAG and AICDA activities.
9 Summary
Our studies on the aetiology and pathology of childhood ALL have provided the molecular evidence for a monoclonal, prenatal origin of ETV6-RUNX1+ leukaemia in monozygotic identical twins and provide mechanistic support for the concept that altered patterns of infection during early childhood can deliver the necessary promotional drive for the progression of ETV6-RUNX1+ pre-leukaemic cells into a postnatal overt leukaemia.
References
Alpar, D., Wren, D., Ermini, L., Mansur, M. B., van Delft, F. W., Bateman, C. M., et al. (2015). Clonal origins of ETV6-RUNX1(+) acute lymphoblastic leukemia: Studies in monozygotic twins. Leukemia, 29, 839–846.
Anderson, K., Lutz, C., van Delft, F. W., Bateman, C. M., Guo, Y., Colman, S. M., et al. (2011). Genetic variegation of clonal architecture and propagating cells in leukaemia. Nature, 469, 356–361.
Bateman, C. M., Colman, S. M., Chaplin, T., Young, B. D., Eden, T. O., Bhakta, M., et al. (2010). Acquisition of genome-wide copy number alterations in monozygotic twins with acute lymphoblastic leukemia. Blood, 115, 3553–3558.
Bateman, C. M., Alpar, D., Ford, A. M., Colman, S. M., Wren, D., Morgan, M., et al. (2015). Evolutionary trajectories of hyperdiploid ALL in monozygotic twins. Leukemia, 29, 58–65.
Broadfield, Z. J., Hain, R. D., Harrison, C. J., Reza Jalali, G., McKinley, M., Michalova, K., et al. (2004). Complex chromosomal abnormalities in utero, 5 years before leukaemia. British Journal of Haematology, 126, 307–312.
Bungaro, S., Irving, J., Tussiwand, R., Mura, R., Minto, L., Molteni, C., et al. (2008). Genomic analysis of different clonal evolution in a twin pair with t(12;21) positive acute lymphoblastic leukemia sharing the same prenatal clone. Leukemia, 22, 208–211.
Castor, A., Nilsson, L., Astrand-Grundstrom, I., Buitenhuis, M., Ramirez, C., Anderson, K., et al. (2005). Distinct patterns of hematopoietic stem cell involvement in acute lymphoblastic leukemia. Nature Medicine, 11, 630–637.
Clarkson, B. D., & Boyse, E. A. (1971). Possible explanation of the high concoddance for acute leukaemia in monozygotic twins. Lancet, 1, 699–701.
Dickinson, H. O. (2005). The causes of childhood leukaemia. BMJ, 330, 1279–1280.
Dockerty, J. D., Draper, G., Vincent, T., Rowan, S. D., & Bunch, K. J. (2001). Case-control study of parental age, parity and socioeconomic level in relation to childhood cancers. International Journal of Epidemiology, 30, 1428–1437.
Ford, A. M., Bennett, C. A., Price, C. M., Bruin, M. C., Van Wering, E. R., & Greaves, M. (1998). Fetal origins of the TEL-AML1 fusion gene in identical twins with leukemia. Proceedings of the National Academy of Sciences of the United States of America, 95, 4584–4588.
Ford, A. M., Fasching, K., Panzer-Grumayer, E. R., Koenig, M., Haas, O. A., & Greaves, M. F. (2001). Origins of “late” relapse in childhood acute lymphoblastic leukemia with TEL-AML1 fusion genes. Blood, 98, 558–564.
Ford, A. M., Palmi, C., Bueno, C., Hong, D., Cardus, P., Knight, D., et al. (2009). The TEL-AML1 leukemia fusion gene dysregulates the TGF-beta pathway in early B lineage progenitor cells. The Journal of Clinical Investigation, 119, 826–836.
Golub, T. R., Barker, G. F., Bohlander, S. K., Hiebert, S. W., Ward, D. C., Bray-Ward, P., et al. (1995). Fusion of the TEL gene on 12p13 to the AML1 gene on 21q22 in acute lymphoblastic leukemia. Proceedings of the National Academy of Sciences of the United States of America, 92, 4917–4921.
Greaves, M. F. (1988). Speculations on the cause of childhood acute lymphoblastic leukemia. Leukemia, 2, 120–125.
Greaves, M. (2006). Infection, immune responses and the aetiology of childhood leukaemia. Nature Reviews. Cancer, 6, 193–203.
Greaves, M. F., Maia, A. T., Wiemels, J. L., & Ford, A. M. (2003). Leukemia in twins: Lessons in natural history. Blood, 102, 2321–2333.
Hardy, R. R., & Hayakawa, K. (2001). B cell development pathways. Annual Review of Immunology, 19, 595–621.
Hong, D., Gupta, R., Ancliff, P., Atzberger, A., Brown, J., Soneji, S., et al. (2008). Initiating and cancer-propagating cells in TEL-AML1-associated childhood leukemia. Science, 319, 336–339.
Hotfilder, M., Rottgers, S., Rosemann, A., Jurgens, H., Harbott, J., & Vormoor, J. (2002). Immature CD34+CD19- progenitor/stem cells in TEL/AML1-positive acute lymphoblastic leukemia are genetically and functionally normal. Blood, 100, 640–646.
Kempski, H. M., & Sturt, N. T. (2000). The TEL-AML1 fusion accompanied by loss of the untranslocated TEL allele in B-precursor acute lymphoblastic leukaemia of childhood. Leukemia & Lymphoma, 40, 39–47.
Kinlen, L. (1988). Evidence for an infective cause of childhood leukaemia: Comparison of a Scottish new town with nuclear reprocessing sites in Britain. Lancet, 2, 1323–1327.
le Viseur, C., Hotfilder, M., Bomken, S., Wilson, K., Rottgers, S., Schrauder, A., et al. (2008). In childhood acute lymphoblastic leukemia, blasts at different stages of immunophenotypic maturation have stem cell properties. Cancer Cell, 14, 47–58.
Li, Z., Woo, C. J., Iglesias-Ussel, M. D., Ronai, D., & Scharff, M. D. (2004). The generation of antibody diversity through somatic hypermutation and class switch recombination. Genes & Development, 18, 1–11.
Ma, Y., Dobbins, S. E., Sherborne, A. L., Chubb, D., Galbiati, M., Cazzaniga, G., et al. (2013). Developmental timing of mutations revealed by whole-genome sequencing of twins with acute lymphoblastic leukemia. Proceedings of the National Academy of Sciences of the United States of America, 110, 7429–7433.
MacKenzie, J., Perry, J., Ford, A. M., Jarrett, R. F., & Greaves, M. (1999). JC and BK virus sequences are not detectable in leukaemic samples from children with common acute lymphoblastic leukaemia. British Journal of Cancer, 81, 898–899.
MacKenzie, J., Gallagher, A., Clayton, R. A., Perry, J., Eden, O. B., Ford, A. M., et al. (2001). Screening for herpesvirus genomes in common acute lymphoblastic leukemia. Leukemia, 15, 415–421.
Maia, A. T., Ford, A. M., Jalali, G. R., Harrison, C. J., Taylor, G. M., Eden, O. B., & Greaves, M. F. (2001). Molecular tracking of leukemogenesis in a triplet pregnancy. Blood, 98, 478–482.
Maia, A. T., Koechling, J., Corbett, R., Metzler, M., Wiemels, J. L., & Greaves, M. (2004). Protracted postnatal natural histories in childhood leukemia. Genes, Chromosomes & Cancer, 39, 335–340.
McNally, R. J., & Eden, T. O. (2004). An infectious aetiology for childhood acute leukaemia: A review of the evidence. British Journal of Haematology, 127, 243–263.
Mori, H., Colman, S. M., Xiao, Z., Ford, A. M., Healy, L. E., Donaldson, C., et al. (2002). Chromosome translocations and covert leukemic clones are generated during normal fetal development. Proceedings of the National Academy of Sciences of the United States of America, 99, 8242–8247.
Mullighan, C. G. (2012). Molecular genetics of B-precursor acute lymphoblastic leukemia. The Journal of Clinical Investigation, 122, 3407–3415.
Mullighan, C. G., Phillips, L. A., Su, X., Ma, J., Miller, C. B., Shurtleff, S. A., & Downing, J. R. (2008). Genomic analysis of the clonal origins of relapsed acute lymphoblastic leukemia. Science, 322, 1377–1380.
Oettinger, M. A., Schatz, D. G., Gorka, C., & Baltimore, D. (1990). RAG-1 and RAG-2, adjacent genes that synergistically activate V(D)J recombination. Science, 248, 1517–1523.
Okuda, T., van Deursen, J., Hiebert, S. W., Grosveld, G., & Downing, J. R. (1996). AML1, the target of multiple chromosomal translocations in human leukemia, is essential for normal fetal liver hematopoiesis. Cell, 84, 321–330.
Papaemmanuil, E., Rapado, I., Li, Y., Potter, N. E., Wedge, D. C., Tubio, J., et al. (2014). RAG-mediated recombination is the predominant driver of oncogenic rearrangement in ETV6-RUNX1 acute lymphoblastic leukemia. Nature Genetics, 46, 116–125.
Patel, N., Goff, L. K., Clark, T., Ford, A. M., Foot, N., Lillington, D., et al. (2003). Expression profile of wild-type ETV6 in childhood acute leukaemia. British Journal of Haematology, 122, 94–98.
Raynaud, S., Cave, H., Baens, M., Bastard, C., Cacheux, V., Grosgeorge, J., et al. (1996). The 12;21 translocation involving TEL and deletion of the other TEL allele: Two frequently associated alterations found in childhood acute lymphoblastic leukemia. Blood, 87, 2891–2899.
Romana, S. P., Mauchauffe, M., Le Coniat, M., Chumakov, I., Le Paslier, D., Berger, R., & Bernard, O. A. (1995). The t(12;21) of acute lymphoblastic leukemia results in a tel-AML1 gene fusion. Blood, 85, 3662–3670.
Rowley, J. D., Le Beau, M. M., & Rabbitts, T. H. (2015). Chromosomal translocations and genome rearrangements in cancer. Cham: Springer.
Shurtleff, S. A., Buijs, A., Behm, F. G., Rubnitz, J. E., Raimondi, S. C., Hancock, M. L., et al. (1995). TEL/AML1 fusion resulting from a cryptic t(12;21) is the most common genetic lesion in pediatric ALL and defines a subgroup of patients with an excellent prognosis. Leukemia, 9, 1985–1989.
Strong, S. J., & Corney, G. (1967). The placenta in twin pregnancy. Oxford: Pergamon Press.
Swaminathan, S., Klemm, L., Park, E., Papaemmanuil, E., Ford, A., Kweon, S. M., et al. (2015). Mechanisms of clonal evolution in childhood acute lymphoblastic leukemia. Nature Immunology, 16, 766–774.
Teuffel, O., Betts, D. R., Dettling, M., Schaub, R., Schafer, B. W., & Niggli, F. K. (2004). Prenatal origin of separate evolution of leukemia in identical twins. Leukemia, 18, 1624–1629.
Tsai, A. G., Lu, H., Raghavan, S. C., Muschen, M., Hsieh, C. L., & Lieber, M. R. (2008). Human chromosomal translocations at CpG sites and a theoretical basis for their lineage and stage specificity. Cell, 135, 1130–1142.
Wang, Q., Stacy, T., Binder, M., Marin-Padilla, M., Sharpe, A. H., & Speck, N. A. (1996). Disruption of the Cbfa2 gene causes necrosis and hemorrhaging in the central nervous system and blocks definitive hematopoiesis. Proceedings of the National Academy of Sciences of the United States of America, 93, 3444–3449.
Wiemels, J. L., Cazzaniga, G., Daniotti, M., Eden, O. B., Addison, G. M., Masera, G., et al. (1999a). Prenatal origin of acute lymphoblastic leukaemia in children. Lancet, 354, 1499–1503.
Wiemels, J. L., Ford, A. M., Van Wering, E. R., Postma, A., & Greaves, M. (1999b). Protracted and variable latency of acute lymphoblastic leukemia after TEL-AML1 gene fusion in utero. Blood, 94, 1057–1062.
Wiemels, J. L., & Greaves, M. (1999). Structure and possible mechanisms of TEL-AML1 gene fusions in childhood acute lymphoblastic leukemia. Cancer Research, 59, 4075–4082.
Zhang, M., & Swanson, P. C. (2008). V(D)J recombinase binding and cleavage of cryptic recombination signal sequences identified from lymphoid malignancies. The Journal of Biological Chemistry, 283, 6717–6727.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Ford, A.M., Greaves, M. (2017). ETV6-RUNX1 + Acute Lymphoblastic Leukaemia in Identical Twins. In: Groner, Y., Ito, Y., Liu, P., Neil, J., Speck, N., van Wijnen, A. (eds) RUNX Proteins in Development and Cancer. Advances in Experimental Medicine and Biology, vol 962. Springer, Singapore. https://doi.org/10.1007/978-981-10-3233-2_14
Download citation
DOI: https://doi.org/10.1007/978-981-10-3233-2_14
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-3231-8
Online ISBN: 978-981-10-3233-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)