Abstract
Research on detecting, recognising and interpreting cardiovascular magnetic resonance images (CMRIs) has started since the 1980s. Time consuming and the need of expert evaluation are the key problems in the manual tracing efforts of CMRIs in a routine investigation. CMRIs manual tracing is also dependent on image quality, and there is no one-size-fits-all MRI setting for an optimum image result. In this paper, we present an approach using 2-Standard Division (2-SD) correlation along with the Sum of Absolute Difference technique and Otsu Watershed to automatically detect the left ventricle (LV) wall and blood pool in the effort to automatically assist the assessment of cardiac function. We test the approach using the Sunnybrook Cardiac Data, a standard benchmark dataset. The results shown that the proposed method had improved the automatic detection of the epicardium and endocardium.
Access provided by CONRICYT-eBooks. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Cardiovascular disease (CVD) is a term for describing disease related to the heart or blood vessels and had been one of the major reasons of death all over the world [1]. For the past two decades, cardiovascular magnetic resonance (CMR) has emerged as an alternative noninvasive modality to assess and detect CVD. During a CMR scan, series of images are produced, and known as cardiovascular magnetic resonance images (CMRIs). These CMRIs depicts the valve anatomy that can be used for quantitative evaluation of stenosis and regurgitation.
The World Health Organisation (WHO) has listed CVD as one of the non-communicable diseases (NCDs) and currently part of the Global Action Plan for the Prevention and Control of NCDs 2013–2020 [1]. In conjunction with the action plan, various semi-automatic and fully automatic techniques were developed to assess the cardiac function and estimate cardiac parameters for clinical use. Common cardiac parameters that are researched are ejection fraction (EF), end-systolic volume (ESV), end-diastolic volume (EDV) and myocardial mass. CMRI has become a reference examination for cardiac morphology, function and perfusion in humans. CMRI automatic analysis is an open research due to the characteristics of CMR and variability of the images among patients; the problem of cardiac cavity; weak edges information; papillary muscles identification; and signal intensity.
Our research focuses on the mapping between CMRI and ontology to guide medical decision making as it can be used to characterise the state and behavior of a patient’s disease both in terms of natural course and as the result of therapeutic interventions. This paper presents part of our work in Phase 3, which is the automatic detection of spatial regions in CRMI, described in Awang Iskandar et al. [2]. It also contributes as phenomenon-centric data to support health care research. Besides being part of the big data science research, transforming the data into knowledge requires involvement of computer science techniques to aggregate the data in hierarchically organised knowledge that will be more understood by non-experts, such as medical students and researchers from other domains.
CMRI captures the space of liver, lungs, stomach and other surrounding organs. The heart only comprises of approximately a 20 % of the CMRI and left ventricle (LV) takes comparatively less space. The LV is our region of interest in this research. Semi-automatic methods use manual cardiac expert involvement to localise LV detection and its walls. In this paper, we present a proposed approach to automatically detect the left ventricle (LV) walls and compared the results against a reputable research by Wijnhout et al. [3]. In particular we have improved the automatic detection of the epicardium and endocardium. One of the main challenges of automatic segmentation of LV walls is the accuracy in segmenting the LV epicardium contour due to the ballooning problem of the epicardium contours at the region between myocardium and lung parenchyma.
The remainder of this paper is organised as follows. In Sect. 2, we present the background on existing related work. In Sect. 3, we describe the proposed method. In Sect. 4, we explain the benchmark dataset and experiment. In Sect. 5, we analyse and evaluate the findings. We conclude in Sect. 6 along with our suggestions for future works.
2 Related Work
Florentine et al. [4] recognized the problem with manual tracing efforts hampering the adoption of cardiac MRI as routine investigation. Manual tracing is also dependent on image quality, and there is no one-size-fits-all MRI setting for the optimum image result. They compared results from a graph-based searching method with the manual tracings, though they acknowledged that this in itself can be considered a bias, as only decent images can be used. Attili et al. [5] recently reviewed the various advances in CMRI image acquisition and processing. They remarked on the lack of an integrated image analysis package for cardiac MRI, as the various solutions only cater to selected elements within a cardiac MRI examination.
Research conducted to segment the LV walls automatically without expert intervention and assessment of cardiac function involves several techniques including fuzzy clustering [6]; dynamic programming [7]; active appearance model [8]; active shape model [9]; deformable model [10]; graph cuts [11] image-driven approaches [12] image-based methods(include region growing and thresholding [13]; pixel classification methods using classification and clustering [14] and atlas guided method [15]. As the study of this segmentation method prove that the LV still need to delineate accurate and fast automatic segmentation.
One of the most referred work for LV segmentation is done by Wijnhout et al. [3] using a hybrid approach of active appearance model and active shape model. Recently, Marino et al. [16] proposed a fully automated technique to detected points inside the LV cavity; calculate the minimal biases and narrow limits of agreement for LV volumes, ejection fraction and mass. On the other hand, Ringenberg et al. [17] proposed a combination novel window-constrained accumulator thresholding technique to drive the segmentation of the right ventricle.
3 Automatic Segmentation of LV Walls
A CMRI slice is a series of images (in the form of video) that covers the whole cardiac cycle, starting from systole to the end of systole; then from the start of diastole to the end diastole. Existing research only manages to segment the LV walls from one image of a slice [3], in our approach, we segmented all images in a slice and render the motion along with the detected segmented walls. By accumulating the information from the motion, segments and wall contours, we derive the LV function.
Before being able to estimate the LV function, an accurate delineation of the LV walls is the primary task. In detecting the LV walls automatically—epicardium and endocardium, the first step is to perform LV localisation to detect the LV from a CMRI slice. We perform automatic region of interest (ROI) to detect the motion areas of the block from CMR short axis (SAX) images. The roundness matrix was used to draw the target circle around the LV ROI, we obtained the center point and calculate the distance from the center point to the endocardium wall.
The second step is to segment and extract the epicardium and endocardium based on the localised images. Besides segmenting the walls, this step also accurately remove the Papillary Muscle and Trabeculation from the contours of detected walls. Otsu watershed was used to solve the heterogeneity problem in the blood pool. The third step is to detect the wall contours in the CMRI slices. Here, we developed an algorithm to locate the beating heart from the CRMIs and detect the motions in a slice using the 2-Standrad Division (2SD) correlation along with the Sum of Absolute Differences technique. The 2SD is used to measure the similarity between blocks of image by calculating the difference between each pixel of original image to its corresponding pixel of the next image. Then absolutes of all differences are summed up. It results in a similarity block of simple metric form:
Here D is value of absolute differences and d is the difference of all pixels from I image and T the template image. If x − y = 0 then no difference is measured if x − y > 0 then it means pixels have changed.
4 Experiment
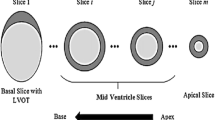
We conducted an experiment to evaluate the effectiveness of the proposed approach. We used a standard benchmark dataset from the previous 2009 Cardiac MR Left Ventricle Segmentation Challenge [18]. It is also known as the Sunnybrook Cardiac Data, containing 45 cases in total. Among the 45 cases, 12 cases were heart failure with ischemia (HF-I), 12 cases of heart failure without ischemia (HF-NI), 12 cases of hypertrophy (HYP) and nine normal (N) cases. The image data were obtained during 10–15 s breath-holds with a temporal resolution of 20 cardiac phases over the heart cycle. Six to twelve SAX images were obtained from the Atrioventricular ring to the apex (with the specification of: thickness 58\(\,\times \,\)10 mm, FOV 5320mm\(\,\times \,\)320 mm, matrix 5256\(\,\times \,\)256). In particular, the cardiac cine MRI LV short axis slices were used for the experiment.
The proposed automatic method of segmenting the LV was tested on different patients each by using three different slices with 20 phases for each slice. Figure 1 depicts a sample image of detected moving epicardium and endocardium contours that are highlighted in blue. The final endocardium border and contour detection (in Fig. 1III), uses the starting and ending points of the wall motion which provide accurate measures of LV.
5 Result Analysis and Evaluation
To analyse the effectiveness of the proposed algorithm, we evaluated the results against the measures available in Randau et al. [18]. Table 1 shows the result comparison between Wijnhout et al. [3] and the proposed approach—the good percentage of endocardium and epicardium contours that was successfully detected for the listed 12 cases. A detail comparison with other known research will be presented in future publication. In this result, the patient ID SC-HF-I-08, SC-HF-NI-07 and SC-HF-NI-11 are missing because the proposed algorithm was designed for diastolic measurement and in these cases the automatic drawn contours are not able to detect the diastolic process due to the nature of the image collection in the dataset. On average the proposed approach detected 90.24 % and 99.35 % of the endocardium and epicardium contours, respectively were detected within an average distance of 2.4 mm of the referenced manual contours. In most patient cases, the proposed approach managed to detect the endocardium better than Wijnhout’s.
6 Conclusion and Future Work
To summarise, we have described a proposed approach for detecting LV walls. This approach is a pipeline of operations to perform LV localisation, LV segmentation and LV wall contour detection. With the aim to accurately perform an automatic segmentation without human intervention using cardiac MRI images, the proposed approach shows a promising result of good contour detection of endocardium and epicardium contours for the assessment of cardiac function. Our future works include identification of similarity for clinical scoring, integrating data of temporal variations and knowledge representation model of semantics behind the cardiac functions.
References
World Health Organisation (2013) Global action plan for the prevention and control of noncommunicable diseases 2013–2020
Iskandar DNFA, Hamimah U (2015) Spatio-temporal semantic representation of cardiac MRI in heart attack patients. In: Proceedings of the 9th international conference on IT in Asia (CITA 2015), pp. 1–5
Wijnhout J, Hendriksen D, Assen HV, der Geest RV (2009) LV challenge LKEB contribution: fully automated myocardial contour detection. MIDAS J 43:2
Florentine MS, Grosskreutz CL, Chang W, Hartnett JA, Dunn VD, Ehrhardt JC, Fleagle SR, Collins SM, Marcus ML, Skorton DJ (1986) Measurement of left ventricular mass in vivo using gated nuclear magnetic resonance imaging. J Am Coll Cardiol 8(1):107–112
Attili AK, Schuster A, Nagel E, Reiber JH, van der Geest RJ (2010) Int J Cardiovasc Imaging 26(Suppl 1):27–40
Lorenzo-Valdés M, Sanchez-Ortiz GI, Elkington AG, Mohiaddin RH, Rueckert D (2004) Segmentation of 4D cardiac MR images using a probabilistic atlas and the EM algorithm. Med Image Anal 8(3):255–265
Rezaee MR, Van der Zwet PM, Lelieveldt BP, Van Der Geest RJ, Reiber JH (2000) A multiresolution image segmentation technique based on pyramidal segmentation and fuzzy clustering. IEEE Trans Image Process 9(7):1238–1248
Kurkure U, Pednekar A, Muthupillai R, Flamm SD, Kakadiaris I (2009) Localization and segmentation of left ventricle in cardiac cine-MR images. IEEE Trans Biomed Eng 56(5):1360–1370
Mitchell SC, Lelieveldt BP, Van der Geest RJ, Bosch HG, Reiver JHC, Sonka M (2001) Multistage hybrid active appearance model matching: segmentation of left and right ventricles in cardiac MR images. IEEE Trans Med Imaging 20(5):415–423
Cordero-Grande L, Vegas-Sánchez-Ferrero G, Casaseca-de-la-Higuera P, San-Román-Calvar JA, Revilla-Orodea A, Martn-Fernndez M, Alberola-Lpez C (2011) Unsupervised 4D myocardium segmentation with a markov random field based deformable model. Med Image Anal 15(3):283–301
Ayed IB, Lu Y, Li S, Ross I (2008) Left ventricle tracking using overlap priors. In: Medical image computing and computer-assisted intervention MICCAI 2008. Springer, Berlin, Heidelberg, pp. 1025–1033
Pednekar A, Kurkure U, Muthupillai R, Flamm S, Kakadiaris I (2006) Automated left ventricular segmentation in cardiac MRI. IEEE Trans Biomed Eng 53(7):1425–1428
Stalidis G, Maglaveras N, Efstratiadis SN, Dimitriadis AS, Pappas C (2002) Model-based processing scheme for quantitative 4-D cardiac MRI analysis. IEEE Trans Inf Technol Biomed 6(1):59–72
Lorenzo-Valdes M, Sanchez-Ortiz GI, Mohiaddin R, Rueckert D (2002) Atlas-based segmentation and tracking of 3D cardiac MR Images using non-rigid registration. In: Medical image computing and computer-assisted intervention MICCAI 2002. Springer, Berlin, Heidelberg, pp. 642–650
Zhuang X, Hawkes D.J, Crum WR, Boubertakh R, Uribe S, Atkinson D, Hill DLG (2008) Robust registration between cardiac MRI images and atlas for segmentation propagation. Med Imaging 691408. International Society for Optics and Photonics
Marino M, Veronesi F, Corsi C (2014) Fully automated assessment of left ventricular volumes and mass from cardiac magnetic resonance images. In: 36th Annual international conference of the IEEE in engineering and medicine and biology society (EMBC), pp. 1079–1082
Ringenberg J, Deo M, Devabhaktuni V, Berenfeld O, Boyers P, Gold J (2014) Fast, accurate, and fully automatic segmentation of the right ventricle in short-axis cardiac MRI. Comput Med Imaging Graph 38(3):190–201
Radau P, Lu Y, Connelly K, Paul G, Dick AJ, Wright GA (2009) Evaluation framework for algorithms segmenting short axis cardiac MRI. MIDAS J Cardiac MR Left Ventricle Segm Chall http://hdl.handle.net/10380/3070
Acknowledgments
This research was supported by Ministry of Education Malaysia through the Exploratory Research Grant (ERGS/ICT07(02)/1019 /2013(16)). The authors would also like to thank Universiti Malaysia Sarawak for providing the resources used in the conduct of this study.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media Singapore
About this paper
Cite this paper
Khan, A., Awang Iskandar, D.N.F., Ujir, H., Chai, W.Y. (2017). Automatic Segmentation of CMRIs for LV Contour Detection. In: Ibrahim, H., Iqbal, S., Teoh, S., Mustaffa, M. (eds) 9th International Conference on Robotic, Vision, Signal Processing and Power Applications. Lecture Notes in Electrical Engineering, vol 398. Springer, Singapore. https://doi.org/10.1007/978-981-10-1721-6_34
Download citation
DOI: https://doi.org/10.1007/978-981-10-1721-6_34
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-1719-3
Online ISBN: 978-981-10-1721-6
eBook Packages: EngineeringEngineering (R0)