Abstract
It is clear that RNA is more than just a messenger between gene and protein. The mammalian genome is pervasively transcribed, giving rise to tens of thousands of noncoding transcripts, especially long noncoding RNAs (lncRNAs). Whether all of these large transcripts are functional remains to be elucidated, but it is evident that there are many lncRNAs that seem not to be the “noise” of the transcriptome. Recent studies have set out to decode the regulatory role and functional diversity of lncRNAs in human physiological and pathological processes, and accumulating evidence suggests that most of the functional lncRNAs achieve their biological functions by controlling gene expression. In this chapter, we will organize these studies to provide a detailed description of the involvement of lncRNAs in the major steps of gene expression that include epigenetic regulation, RNA transcription, posttranscriptional RNA processing, protein translation, and posttranslational protein modification and highlight the molecular mechanisms through which lncRNAs function, involving the interactions between lncRNAs and other biological macromolecules.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
2.1 Introduction
A large range of biological processes involved in cancer progression, such as cell differentiation, proliferation, apoptosis, and metastasis, are widely reported to be associated with long noncoding RNAs (lncRNAs), which are thought to work in cis on neighboring genes or in trans to regulate distantly located genes or molecular targets in the nucleus and cytoplasm. It has been clear that lncRNAs can function through quite diverse mechanisms and that their interaction with DNA, RNA, or protein is a well-established action mode [1]. On the basis of the intermolecular interactions, most of the characterized lncRNAs are shown to function in gene expression control by acting as decoys, guides, or scaffolds [2, 3]. The “guiding” lncRNAs, like Kcnq1ot1 [4] and lincRNA-p21 [5], are associated with chromatin regulatory protein complexes or transcriptional co-regulators and recruit them to specific genomic DNA regions to regulate transcription; “decoying” lncRNAs, such as GAS5 [6], Lethe [7], and PANDA [8], mimic and compete with their consensus DNA-binding motifs for binding nuclear receptors or transcriptional factors in the nuclei; “scaffolding” lncRNAs, including HOTAIR [9], XIST [10], and NRON [11], bring specific regulatory proteins into proximity with each other to function as a unique complex. Furthermore, many lncRNAs are exclusively expressed in specific stages of tissue differentiation and development or present apparent cell-type-specific expression patterns and distinct subcellular localization [12, 13]. Although some of such lncRNAs are merely by-products of transcription that don’t possess any regulatory function, they can faithfully reflect the action of gene expression or activation of signaling pathway and therefore act as “signaling” molecules [3].
Altogether, the current studies suggest that lncRNAs could be involved in almost each step of gene expression, such as epigenetic regulation, RNA transcription, posttranscriptional RNA processing, protein translation, and posttranslational protein modification, through in cis or in trans manner, and the dysregulation of lncRNAs could cause broad changes in cell signaling pathways.
2.2 Epigenetic Regulation
2.2.1 lncRNAs Involved in Histone Modification
Epigenetic regulation of gene expression, which is characterized as the altered transcription without any change in gene sequence, is generally reported to play roles in organism development as well as in tumorigenesis. Recently, the involvement of lncRNAs in epigenetic regulation has been widely documented. Kcnq1ot1, a nuclear-retained lncRNA with the length of 91.5 kb, is transcribed by RNA polymerase II (RNAP II) from the intron 10 of Kcnq1 gene in an antisense orientation to Kcnq1 [14]. Kcnq1ot1 was first found to be associated with the lineage-specific silencing of dozens of genes within the Kcnq1 locus [14, 15], and the subsequent studies further indicated that its silencing effect was achieved by the interactions with chromatin and with the H3K9- and H3K27-specific histone methyltransferases G9a and polycomb repressive complex 2 (PRC2) [4]. HOTAIR and Air are the other two well-characterized lncRNAs involved in chromatin remodeling, which exert function through a similar fashion, including accumulation at the chromatin regions of silenced genes and the subsequent mediation of repressive histone modification through recruiting specific histone modifiers such as G9a, PRC1, and PRC2 [9, 16–18].
2.2.2 lncRNAs Involved in DNA Methylation
Although interaction with histone modifiers is a major mechanism through which lncRNAs function in epigenetic regulation, some lncRNAs are also implicated in establishment and maintenance of DNA methylation patterns. DNA methylation is mediated by the members of the DNA methyltransferase (DNMT) family, conventionally classified as de novo (DNMT3a and DNMT3b) and maintenance (DNMT1) DNMTs. The ecCEBPA represents one of the lncRNAs that modulate DNA methylation by interacting with DNMT1, which functions as a decoying transcript to sequestrate DNMT1, resulting in prevention of CEBPA gene locus methylation [19]. The best characterized DNMT3a- and DNMT3b-binding lncRNA is Dum, which recruits both de novo methylation and maintenance DNMTs to silence its neighboring gene in cis [20].
Interestingly, some lncRNAs, especially the antisense lncRNAs, have been identified to simultaneously mediate DNA and histone modifications at the loci of silenced genes [21]. TMS1/ASC is a tumor suppressor gene that encodes a pro-apoptotic signaling factor operating in the intrinsic and extrinsic cell death pathways [22]. TMS1/ASC was originally identified as a downstream target of DNA methyltransferase-1 (DNMT1), and subsequent studies further showed that it was subjected to the hypermethylation-mediated epigenetic silencing in a wide range of human tumors [23–27]. In addition to DNA methylation, the inactivation of TMS1/ASC is also controlled by certain other epigenetic events such as the G9a-mediated histone H3K9 methylation [28], which is commonly linked to the methylation of nearby CpG sites [29, 30]. Biochemical interactions between DNA and histone methyltransferases were thought to provide a molecular explanation, at least in part, for the combinatorial pattern of DNA and histone modifications in chromatin [29–32]. In a recent study, an antisense lncRNA of TMS1/ASC, termed TMS1AS, was further revealed to regulate outputs of its sense counterpart by interacting with the DNMT1/G9a complex and aiding in recruitment of the complex to the sense promoter (see Fig. 2.1). This interesting finding not only highlighted the significant involvement of lncRNAs in epigenetic regulation of gene expression but also revealed a potential crosstalk between DNA methylation and histone modification established by lncRNA. Kcnq1ot1 is another example of such antisense lncRNAs. In addition to its role in regulating histone modification, Kcnq1ot1 was also reported to be required for the silencing of ubiquitously imprinted genes (UIGs) through guiding and maintaining the CpG methylation at methylated regions flanking the UIGs [33].
2.2.3 Correlation Between the Sense and Antisense Transcripts
The antisense lncRNAs described above represent guiding transcripts that modulate their sense counterparts negatively. However, sense-antisense pairs are frequently revealed to express in a concordant manner [34, 35]. TARID is one of the antisense lncRNAs that positively correlate with their sense counterparts. The underlying mechanism implicates its simultaneous association with sense promoter and the regulator of DNA demethylation GADD45A, which in turn recruits thymine-DNA glycosylase along with members of TET family to induce sense promoter demethylation [36]. Thus, it can be speculated that although many antisense lncRNAs regulate their sense counterparts through a general mechanism serving as a genomic address label for specific epigenetic modification enzymes, they may exert completely opposite effect on gene expression on the basis of property of the associated enzyme partner.
Certainly, certain lncRNAs involved in epigenetic regulation also possess the ability to control multiple targets in addition to their sense counterparts. It has been speculated that lncRNAs recruit epigenetic modification complexes by binding to target sites through three mechanisms: tethering to its nascent transcription locus, directly hybridizing to genomic targets, or interacting with a DNA-binding protein [37].
2.3 RNA Transcription
Transcription is a tightly regulated process in eukaryotes. In addition to the well-known protein factors such as RNAP, general transcriptional factors, and gene-specific transcriptional factors, it is also suggested that ncRNAs, which include small ncRNAs and lncRNAs, exert regulatory functions in the complicated network to make gene expression more symphonic. Thus, a current central issue is to obtain a full understanding of the potential role of lncRNAs in regulated gene transcription programs, possibly through diverse mechanisms.
2.3.1 Role of NcRNAs in Controlling Initiation and Elongation of Transcription
The expression of protein-coding genes in mammalian genomes begins with the assembly of the preinitiation complex (PIC) that brings RNAP II to gene promoters. Following this step, U1 snRNA can induce transcriptional initiation by specifically binding to and stimulating TFIIH to phosphorylate the C-terminal domain of RNAP II [38]. However, after transcriptional initiation and promoter clearance, RNAP II is frequently paused near the transcription start site on numerous genes, and the regulation of RNAP II pause release has been recognized as a critical step in activation of gene transcription [39].
The transition of RNAP II to productive elongation requires active recruitment of P-TEFb, a cyclin-dependent kinase responsible for phosphorylation of the C-terminal domain of RNAP II and other key transcription elongation factors. The 7SK RNA is able to repress transcription elongation by, in combination with HEXIM1/2, forming an inactive complex that sequesters P-TEFb and then prevents its active recruitment [40, 41]. The SR-splicing factor SRSF2 is a newly identified component of 7SK complex assembled at gene promoters. Upon its binding to promoter-associated nascent RNA, SRSF2 can mediate the switch of P-TEFb from the 7SK complex to RNAP II, making the paused transcription elongation reactive [42].
2.3.2 lncRNAs Regulate Transcription of Specific Genes
Through recruiting and modulating the activities of co-regulators, lncRNAs may act as selective ligands to prevent the transcription of target genes. DNA damage has been reported to induce the production of several lncRNAs from the 5’ regulatory region of cyclin D1 (CCND1). These induced lncRNAs specifically bind and allosterically modify TLS, a regulatory sensor of DNA damage, leading to the interaction of the modified TLS with CREB-binding protein (CBP) that thereby inhibits the transcription of CCND1 [43]. Martianov et al. have reported another example of inhibitory lncRNA that is induced by serum starvation and functions as a promoter-specific transcriptional repressor. Upon being transcribed from the upstream minor promoter of DHFR gene, this lncRNA not only bonds transcriptional factor II B (TFIIB) to prevent its association with the major promoter but also formed a stable complex with the major promoter that interfered with the promoter-directed recruitment of TFIIB [44].
In addition to the inhibitory effects on transcription, lncRNAs can also act as activators or coactivators to upregulate gene expression. For instance, steroid receptor RNA activator (SRA) is an lncRNA that acts as a eukaryotic transcriptional coactivator for steroid hormone receptors [45]. Another lncRNA, called Evf-2, which is transcribed from one of the two Dlx-5/6 conserved intergenic regions, activates transcriptional activity of homeodomain proteins by directly influencing Dlx-2 activity [46]. The functional importance of gene enhancers in regulated gene expression has been well established, and the subsequent identification of bidirectional lncRNAs transcribed from enhancers, termed as enhancer RNAs (eRNAs), adds another functional layer to the transcriptional regulatory elements [47–49]. A subtype of eRNAs, which are derived from enhancers adjacent to E2-upregulated coding genes, has been observed to contribute to E2-dependent gene activation by stabilizing E2-/ERα-/eRNA-induced enhancer-promoter looping, suggesting that eRNAs are not merely a reflection of enhancer activation but a functional player [50].
Based on the ability to repress proto-oncogenes, PSF functions as a tumor suppressor protein, whose oncogenesis suppression activity, however, is impaired by the enhancement of its RNA-binding ability [51, 52]. The subsequent studies identify several PSF-binding lncRNAs from mice and humans, which include retroposon-derived lncRNAs such as VL30 as well as frame-disrupted and tumorigenesis-related noncoding transcripts such as MALAT1. All the lncRNAs promote tumorigenesis in mice and humans through a mechanism of reversible regulation on proto-oncogene transcription, including the protein PSF that binds to the regulatory region of a proto-oncogene and represses transcription and PSF-binding lncRNA that binds to PSF, forming a PSF-lncRNA complex that dissociates from a proto-oncogene, activating transcription [53–57] (see Fig. 2.2).
Regulation of gene transcription by PSF protein and PSF-binding lncRNAs. The first diagram on the left shows the PSF binding to the promoter (P) of a gene via the DNA-binding domain (DBD), which represses transcription of the coding region (C). The second diagram in the center shows the binding of an lncRNA to the RNA-binding domain (RBD) of PSF. The third diagram on the right shows the release of PSF from the promoter and initiation of transcription
2.3.3 lncRNAs Join P53 Network by Regulating Transcription
P53 is considered to be one of the most common denominators in human cancer that plays a central role in regulatory networks responsible for cancer-related stress [58–60]. It seems controversial to explain p53 pathway only from the perspective of protein-coding genes [61–63]. As expected, a large number of miRNAs and conserved miRNA families are found to be direct transcriptional targets of p53 that mediate the p53 function or serve as the upstream regulators of p53 [64]. In addition to the small ncRNAs, several studies further illustrate the linkage between lncRNAs and p53 pathway from the perspective of molecular biology. Among these lncRNAs, p53-activated lincRNA-p21 serves as a repressor, through interacting with heterogeneous nuclear ribonucleoprotein K (hnRNP-K), in p53-dependent transcriptional response. Inhibition of lincRNA-p21 results in a global change in expression of hundreds of gene targets, a majority of which are downstream targets repressed by p53 and responsible for p53-mediated apoptosis [5]. PANDA RNA is another downstream target of p53. Upon being induced by p53, PANDA RNA acts as a decoy through an interaction with the transcriptional factor NF-YA that prevents NF-YA from activating expression of pro-apoptosis genes, leading to cell cycle arrest [8].
2.3.4 Circular Intronic RNA (ciRNA)
During the process of pre-mRNA splicing, the 5’ splice site within intron sequence undergoes a nucleophilic attack by the downstream branch point to form a circular structure called the lariat, and the resultant free 5’ exon then attacks the 3’ splice site within intron, leading to the joint of two exons and the release of the intron lariat. In general, the intron lariats are debranched and degraded rapidly. However, a debranching failure, which seems to be determined by the presence of specific elements near the 5’ splice site and the branch point site, can result in the formation of the circular intronic RNAs, referred to as ciRNAs [65–67].
ciRNAs are prominently found in the nucleus and are often more stable than their parent linear mRNAs [67]. Once generated, ciRNAs seem to function in cis to regulate the transcription of their parent genes. For example, a relatively abundant ciRNA called ci-ankrd52 was found to interact with elongating RNAP II and facilitate transcription. Many ciRNAs remain at their “sites of synthesis” in the nucleus. However, a portion of ciRNAs do localize to additional sites in the nucleus, suggesting that they may have certain transacting effects other than regulating the transcription of their parent genes [67].
2.4 Posttranscriptional RNA Processing
2.4.1 Alternative Splicing
For all eukaryotes, one of the most versatile processing steps in the life of an mRNA molecule is pre-mRNA splicing, through which the noncoding introns are removed and the neighboring exons are ligated together before protein translation. Although the spliceosome functions in splicing catalysis, some additional transacting protein factors, such as the serine-/arginine-rich (SR) nuclear proteins (SR proteins), the SR-related proteins, the small nuclear ribonucleoproteins (snRNPs), and the heterogeneous nuclear ribonucleoproteins (hnRNPs), are well characterized to be required for the choice of splice sites, where spliceosome assembles and the following splicing occurs [68–70]. More than 90 % of human multi-exon-containing genes are speculated to be alternatively spliced in tissue- and cell-specific manners [71, 72], and in some spectacular examples, thousands of distinct gene products (protein isoforms) can be generated from a single gene [73], pointing to alternative pre-mRNA splicing as a common mechanism in regulation and diversification of gene function. Interestingly, several recent studies have highlighted the significant implication of lncRNAs in the regulation of pre-mRNA splicing.
2.4.1.1 MALAT1
Metastasis-associated lung adenocarcinoma transcript 1 (MALAT1), also known as nuclear-enriched autosomal transcript 2 (NEAT2), was originally identified as a prognostic marker in non-small cell lung cancer [74], and the subsequent studies showed that it was also overexpressed in many other human cancers [75–79].
This nuclear-enriched noncoding transcript predominantly localizes to nuclear speckles, the highly dynamic nuclear subdomains that are enriched with pre-mRNA splicing factors and are thought to serve as sites for the assembly, modification, and/or storage of the pre-mRNA processing machinery [80, 81]. Another nuclear-enriched autosomal noncoding transcript, referred to as NEAT1, has been shown to be significantly implicated in the structural maintenance of paraspeckles, the nuclear subdomains that control mRNA nuclear export [82–85]. MALAT1, however, does not possess a similar capacity to establish or maintain the functional nuclear subdomains since its depletion does not disrupt the architecture of nuclear speckles [86]. Nevertheless, MALAT1 is required for proper localization of several SR proteins to nuclear speckles [86], and its depletion has been reported to compromise the recruitment of SR proteins from nuclear speckles to the sites of transcription, where splicing occurs [87]. Furthermore, it has been demonstrated that MALAT1 can act as a “molecular sponge” by interacting with SR proteins, especially SRSF1, and thereby modulate the concentration of “splicing-competent” SR proteins in cells [86]. Altogether, these studies suggest that MALAT1 regulates pre-mRNA splicing by influencing the localization and/or activity of SR proteins. This “sponge” mechanism is also applicable to the untranslated region (UTR) of certain protein-coding genes. For instance, the repeat-containing RNA from the CTG expansion in 3’UTR of DMPK gene accumulates in the nucleus and affects the activity of splicing factors; similarly, CGG repeats occurred in FMR1 gene can recruit a set of splicing regulators into nuclear inclusions [88].
It has been suggested that the MALAT1enrichment in nuclear speckles occurs only when RNAP II-dependent transcription is active [87]. Since nuclear speckles do not represent the major sites of transcription or splicing, this finding seems contrary to the recent studies showing that MALAT1 binds many nascent pre-mRNAs derived from actively transcribed gene loci [89, 90]. Although the subcellular location of MALAT1 remains puzzling, these studies highlight the significant involvement of lncRNAs in the control of pre-mRNA splicing and the potential contribution of lncRNAs to the links between gene transcription and pre-mRNA splicing.
MALAT1 has a strong influence on pre-mRNA splicing patterns in human HeLa and fibroblast cells [86, 91, 92]. However, significant splicing changes were not observed when MALAT1 gene was knocked out in human lung cancer cells or in MALAT1 knockout mice [93, 94]. Although some other aspects of gene expression such as transcription have been shown to be affected in these cases, these studies still implicate that human and mouse genomes may encode certain functionally redundant products that possess the ability to regulate pre-mRNA splicing and compensate the loss of MALAT1. Certainly, we cannot exclude the possibility that MALAT1 only exerts the regulatory function in pre-mRNA splicing under a particular cell state or in specific cellular contexts.
2.4.1.2 sno-lncRNAs
The sno-lncRNAs (snoRNA-related long noncoding RNAs), a class of newly identified nuclear-enriched lncRNAs, are derived from the intron sequences and terminate in either box C/D or box H/ACA small nucleolar RNA (snoRNA) structures at their 5’ and 3’ ends [95]. At least 19 endogenous sno-lncRNAs have been identified in human, rhesus monkey, and mouse [96]. Most of them have been demonstrated to present apparent tissue- and species-specific expression patterns. Furthermore, their terminal snoRNA structures, but not the internal sequences, are highly conserved across species evolution.
The sno-lncRNAs encoded by the q11-q13 region of human chromosome 15 have been well characterized [95]. During exonucleolytic trimming, the terminal snoRNA structures protect the internal sequences between them from degradation, leading to the accumulation of these sno-lncRNAs to high levels (expression similar to that of some histone mRNAs). Although sno-lncRNAs and snoRNAs are processed by the same machinery and both of them are generally originated from the introns of protein-coding genes, sno-lncRNAs mainly accumulate near their sites of synthesis but do not co-localize with nucleoli or Cajal bodies, where snoRNAs reside [95, 97]. The difference in their cellular distribution strongly indicates that these sno-lncRNAs do not have a similar function as snoRNAs, which play roles in the modification of other noncoding RNAs including rRNAs and snRNAs. Indeed, these sno-lncRNAs strongly associate with RbFox2, a member of Fox family splicing regulators that is known to regulate many posttranscriptional events including alternative splicing. Moreover, these sno-lncRNAs can significantly influence the RbFox2-regulated splicing events through acting, at least in part, as molecular sinks that prevent RbFox2 from targeting its mRNA targets [95].
2.4.2 lncRNAs Function as miRNA Sponges
On the basis of direct base pairing to target sites within untranslated regions of mRNAs, the small regulatory ncRNAs such as miRNAs and endo-siRNAs can function as important posttranscriptional regulators that trigger mRNA degradation or translational inhibition. Since lncRNAs achieve their functions through employing quite diverse strategies, they are generally considered to be a class of regulatory RNAs fundamentally distinct from the small RNAs. Nevertheless, current studies have revealed that lncRNAs can serve as the precursors of certain small ncRNAs, such as miR-675 derived from the H19 lncRNA [98, 99]. Furthermore, the link between lncRNA and miRNA has been presented by the finding that several lncRNAs possess the ability to affect the miRNA activity, which is attributed to the internal miRNA-matching sequence. For instance, in human hepatocellular carcinoma metastases, the lncRNA activated by TGF-β, termed as lncRNA-ATB, was reported to act as a sponge for miR-200 family. The sponge effect was mediated by the selectively conserved miRNA target sites contained in lncRNA-ATB, which strongly bond miR-200 s to suppress their activities, resulting in increased levels of ZEB1 and ZEB2, two miR-200 s targets, and the ultimate induction of epithelial-mesenchymal transition (EMT) and invasion [100]. Moreover, identification of other sponge lncRNAs, such as tumor-suppressive PTEN competitive endogenous RNAs (ceRNAs) and muscle-specific linc-MD1 that targets miR-133 and miR-135, suggests that the lncRNA-mediated miRNA sponge effects can be a general phenomenon in diseases as well as in normal physiology [101–106].
In addition to the canonical linear mRNAs, it is now clear that thousands of protein-coding genes can also be processed to generate circular RNAs (circRNAs), which are resistant to exonuclease-mediated degradation due to the fact that they do not have 5’ and 3’ ends and the two ends have been joined together [107–113]. Distinct from ciRNAs that are derived from introns and reside in the nuclei, circRNAs are almost exclusively encoded by exons and predominantly localize in the cytoplasm. circRNAs are produced by noncanonical splicing events called “backsplicing,” in which a splice donor site is jointed to a splice acceptor site further upstream in the primary transcript [111]. The strategy employed by circRNAs to exit the nucleus remains to be elucidated. Since the nuclear envelope breaks down during mitosis, it seems reasonable to hypothesize that circRNAs exit the nucleus during this phase of the cell cycle. However, expression of certain circRNAs such as ciRS-7 is also detected in neuronal tissues, where mitotic division occurs at a low frequency [110, 114].
CiRS-7 and Sry are the two well-characterized circRNAs [110, 114]. These particular circRNAs contain many binding sites for specific miRNAs (miR-7 and miR-138, respectively). Following their accumulation in the cytoplasm, they can act as sponges that titrate the miRNAs from their other RNA targets, thereby modulating the miRNA activities. Although current attention has been focused on circRNA’s sponge effects, several lines of evidence also suggest their other functional possibilities. For instance, expression of ciRS-7 but not miR-7 has been detected in some areas of the mouse adult hippocampus, suggesting that ciRS-7 may have certain roles other than interacting with the miRNA [110]. Furthermore, most other circRNAs have been identified to contain few miRNA binding sites, indicating that they may achieve their functions through other strategies such as binding to RNA-binding proteins to form RNA-protein complexes [108, 113].
2.5 Protein Translation
lincRNA-RoR is one of the few lncRNAs whose detailed function in the regulation of p53 pathway has been characterized. Distinct from the p53-regulated lincRNA-p21 and PANDA, lincRNA-RoR acts as an upstream regulator of p53 in response to DNA damage. MDM2 is a well-known upstream negative p53 regulator, which causes p53 degradation through the ubiquitin-proteasome pathway. LincRNA-RoR, however, has been demonstrated to function via a different mechanism, involving its direct interaction with hnRNP-I that suppresses p53 translation and inhibits p53-mediated cell cycle arrest and apoptosis [115].
As described above, lincRNA-p21 was initially revealed to be induced by p53 during DNA damage and recruit hnRNP-K via physical interaction to facilitate p53-mediated repression of gene transcription [5]. Interestingly, the subsequent study showed that it possessed an additional role in translational control, the output of which was determined by the presence or absence of RNA-binding protein HuR [116]. In the presence of HuR, the association of HuR with lincRNA-p21 facilitates the recruitment of let-7/Ago2 to lincRNA-p21, leading to lower lincRNA-p21 stability. On the contrary, lincRNA-p21 is stable and accumulates when HuR is absent. It then interacts with the mRNAs CTNNB1 and JUNB and translational repressor Rck, repressing the translation of the targeted mRNAs.
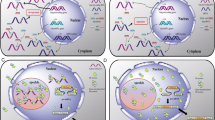
The lncRNA-mRNA gene pairs have been found to be prevalent in mammalian genomes, and the cluster of natural antisense transcripts (NATs) is revealed to constitute a surprisingly large fraction of lncRNAs [34, 35]. Unlike the nuclear-retained NATs such as Tsix, Airn, and HOTAIR that guide the epigenetic modification complexes to the target loci, some NATs can form RNA duplexes with the mRNA of paired gene, leading to the change in mRNA translation. Zeb2/Sip1 is a transcriptional repressor of E-cadherin. After the Snail1-induced EMT, expression of the Zeb2/Sip1 NAT leads to an increase in Zeb2/Sip1 protein level without any change at the mRNA level. The Zeb2/Sip1 NAT is complementary to the 5’ splice site of an intron in the 5’ UTR of Zeb2 mRNA. Expression of the Zeb2/Sip1 NAT upon EMT can mask the spice site, preventing deletion of the intron. As a result, the translation machinery can then recognize and bind to an internal ribosome entry site (IRES) in the retained intron, resulting in more efficient Zeb2/Sip1 translation [117]. The Uchl1 NAT is another antisense lncRNA that promotes translation of the paired mRNA. The Uchl1 contains a region that overlaps with the first 73 nucleotides of Uchl1 mRNA. Under stress conditions in which cap-dependent translation is inhibited, the Uchl1 NAT moves from the nucleus to the cytoplasm and hybridizes with Uchl1 mRNA to switch on its cap-independent translation; that is, the Uchl1 NAT acts as an internal ribosomal entry element to promote selective translation [118]. Although the above RNA duplexes work in promoting mRNA translation, the RNA-RNA pairing between TMS1AS and TMS1/ASC mRNA has been demonstrated to suppress TMS1/ASC translation by interfering with the ribosome assembly on TMS1/ASC mRNA. As described above, TMS1AS also regulates TMS1/ASC expression at the epigenetic level. Thus, this NAT links different effector mechanisms to simultaneously operate in the different aspects of TMS1/ASC regulation, contributing to establishing a more strict fashion of gene expression and enhancing the efficacy of gene expression control (see Fig. 2.3).
Proposed model of TMS1AS-mediated regulation of TMS1/ASC expression. TMS1AS is transcribed from the opposite strand of TMS1/ASC gene. Following transcription, a fraction of TMS1AS transcripts act in cis by recruiting the chromatin repressor proteins DNMT1 and G9a to TMS1/ASC promoter, and other transcripts interact with TMS1/ASC mRNA through direct base pairing, forming intermolecular duplexes that suppress TMS1/ASC translation by interfering with the ribosome assembly on TMS1/ASC mRNA. This novel mechanism results in a target-specific regulation of TMS1/ASC at both epigenetic and translational levels by its antisense counterpart
2.6 Posttranslational Protein Modification
Upon being synthesized by ribosomes, the nascent polypeptide chains undergo posttranslational modification (PTM) to form the mature protein product. In addition, PTM is also an important strategy employed to control protein activity. The initial evidence for a function of lncRNAs in PTM comes from the research on the lncRNA-mediated epigenetic gene silencing. As described above, lncRNAs have been associated with gene silencing through guiding enzymes involved in chromatin remodeling, such as PRC2 and G9a, to specific genomic DNA regions. In other words, lncRNAs can function in the recruitment of histone posttranslational modification machinery.
The function of most lncRNAs depends on their ability to interact with proteins, implying that lncRNAs may also directly interact with functional domains of signaling proteins and thus regulate signal transduction. NF-κB is a critical link between inflammation and cancer, and its aberrant activation has been observed in many tumors. Despite the classical negative regulators that include IκB, an lncRNA termed NKILA has been recently reported to play a significant role in regulating NF-κB signaling and repressing cancer-associated inflammation. NKILA binds to NF-κB/ IκB complex and inhibits NF-κB signaling by masking the phosphorylation sites of IκB and stabilizing the complex. Importantly, NKILA expression is significantly decreased in many breast cancers and is associated with cancer metastasis and poor patient prognosis [119]. This action mode is also applicable to lnc-DC. This transcript has been reported to regulate STAT3 signal transduction by interacting with STAT3 in the cytoplasm of dendritic cells and modulating its phosphorylation [120].
2.7 Summary and Perspectives
The discovery that lncRNAs are central to numerous pivotal biological processes may reflect ancient connections between lncRNAs and the regulation of developmental and physiological decisions, whose disruption can lead to physiological disorders as well as many types of diseases. Comparison of gene expression profiles of tumor and normal cells has revealed a linkage of lncRNAs with tumorigenesis. Most of the oncogenic and tumor suppressor lncRNAs are characterized to function in gene expression control, and their interaction with DNA, RNA, or protein is a well-established action mode [1]. However, there are only limited studies decoding the detailed molecular mechanism of these cancer-related lncRNAs. It has long been acknowledged that lncRNAs contain functionally redundant sequences. Nevertheless, their core functionality relies heavily on the cooperative action of their dispersed functional domains [121]. Similar to miRNAs, lncRNAs can also serve as the potential molecular targets for diagnosis and treatment of cancer. Thus, it is important to decode the molecular features of lncRNAs, which include the consensus motif and structural element that determine the intermolecular interactions, to help understand the involvement and significance of lncRNAs in tumor biology.
References
Guttman M, Rinn JL. Modular regulatory principles of large non-coding RNAs. Nature. 2012;482:339–46.
Ulitsky I, Bartel DP. LincRNAs: genomics, evolution, and mechanisms. Cell. 2013;154:26–46.
Wang KC, Chang HY. Molecular mechanisms of long non-coding RNAs. Mol Cell. 2011;43:904–14.
Pandey RR, Mondal T, Katayama F, et al. Kcnq1ot1 antisense non-coding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol Cell. 2008;32:232–46.
Huarte M, Guttman M, Feldser D, et al. A large intergenic non-coding RNA induced by p53 mediates global gene repression in the p53 response. Cell. 2010;142:409–19.
Kino T, Hurt DE, Ichijo T, et al. Non-coding RNA gas5 is a growth arrest- and starvation-associated repressor of the glucocorticoid receptor. Sci Signal. 2010;3:ra8.
Rapicavoli NA, Qu K, Zhang J, et al. A mammalian pseudogene lncRNA at the interface of inflammation and anti-inflammatory therapeutics. eLife. 2013;2:e00762.
Hung T, Wang Y, Lin MF, et al. Extensive and coordinated transcription of non-coding RNAs within cell-cycle promoters. Nat Genet. 2011;43:621–9.
Tsai MC, Manor O, Wan Y, et al. Long non-coding RNA as modular scaffold of histone modification complexes. Science. 2010;329:689–93.
Jeon Y, Lee JT. YY1 tethers Xist RNA to the inactive X nucleation center. Cell. 2011;146:119–33.
Sharma S, Findlay GM, Bandukwala HS, et al. Dephosphorylation of the nuclear factor of activated T cells (NFAT) transcription factor is regulated by an RNA-protein scaffold complex. Proc Natl Acad Sci U S A. 2011;108:11381–6.
Guttman M, Amit I, Garber M, et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature. 2009;458:223–7.
Ulitsky I, Shkumatava A, Jan CH, et al. Conserved function of lincRNAs in vertebrate embryonic development despite rapid sequence evolution. Cell. 2011;147:1537–50.
Mancini-Dinardo D, Steele SJ, Levorse JM, et al. Elongation of the Kcnq1ot1 transcript is required for genomic imprinting of neighboring genes. Genes Dev. 2006;20(10):1268–82.
Fitzpatrick GV, Soloway PD, Higgins MJ. Regional loss of imprinting and growth deficiency in mice with a targeted deletion of KvDMR1. Nat Genet. 2002;32(3):426–31.
Gupta RA, Shah N, Wang KC, et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature. 2010;464(7291):1071–6.
Nagano T, Mitchell JA, Sanz LA, et al. The Air non-coding RNA epigenetically silences transcription by targeting G9a to chromatin. Science. 2008;322(5908):1717–20.
Rinn JL, Kertesz M, Wang JK, et al. Functional demarcation of active and silent chromatin domains in human HOX loci by non-coding RNAs. Cell. 2007;129(7):1311–23.
Di Ruscio A, Ebralidze AK, Benoukraf T, et al. DNMT1-interacting RNAs block gene-specific DNA methylation. Nature. 2013;503(7476):371–6.
Wang L, Zhao Y, Bao X, et al. LncRNA Dum interacts with Dnmts to regulate Dppa2 expression during myogenic differentiation and muscle regeneration. Cell Res. 2015;25(3):335–50.
Wutz A. RNA-mediated silencing mechanisms in mammalian cells. Prog Mol Biol Transl Sci. 2011;101:351–76.
McConnell BB, Vertino PM. TMS1/ASC: the cancer connection. Apoptosis. 2004;9:5–18.
Conway KE, McConnell BB, Bowring CE, et al. TMS1, a novel proapoptotic caspase recruitment domain protein, is a target of methylation-induced gene silencing in human breast cancers. Cancer Res. 2000;60(22):6236–42.
Moriai R, Tsuji N, Kobayashi D, et al. A proapoptotic caspase recruitment domain protein gene, TMS1, is hypermethylated in human breast and gastric cancers. Anticancer Res. 2002;22(6C):4163–8.
Stone AR, Bobo W, Brat DJ, et al. Aberrant methylation and down-regulation of TMS1/ASC in human glioblastoma. Am J Pathol. 2004;165(4):1151–61.
Virmani A, Rathi A, Sugio K, et al. Aberrant methylation of TMS1 in small cell, non small cell lung cancer and breast cancer. Int J Cancer. 2003;106(2):198–204.
Yokoyama T, Sagara J, Guan X, et al. Methylation of ASC/TMS1, a proapoptotic gene responsible for activating procaspase-1, in human colorectal cancer. Cancer Lett. 2003;202(1):101–8.
Kapoor-Vazirani P, Kagey JD, Powell DR, Vertino PM. Role of hMOF-dependent histone H4 lysine 16 acetylation in the maintenance of TMS1/ASC gene activity. Cancer Res. 2008;68(16):6810–21.
Chang Y, Sun L, Kokura K, et al. MPP8 mediates the interactions between DNA methyltransferase Dnmt3a and H3K9 methyltransferase GLP/G9a. Nat Commun. 2011;2:533.
Hashimoto H, Vertino PM, Cheng X. Molecular coupling of DNA methylation and histone methylation. Epigenomics. 2010;2(5):657–69.
Cedar H, Bergman Y. Linking DNA methylation and histone modification: patterns and paradigms. Nat Rev Genet. 2009;10(5):295–304.
Estève PO, Chin HG, Smallwood A, et al. Direct interaction between DNMT1 and G9a coordinates DNA and histone methylation during replication. Genes Dev. 2006;20(22):3089–103.
Mohammad F, Pandey GK, Mondal T, et al. Long non-coding RNA-mediated maintenance of DNA methylation and transcriptional gene silencing. Development. 2012;139(15):2792–803.
Katayama S, Tomaru Y, Kasukawa T, et al. Antisense transcription in the mammalian transcriptome. Science. 2005;309(5740):1564–6.
Sigova AA, Mullen AC, Molinie B, et al. Divergent transcription of long non-coding RNA/mRNA gene pairs in embryonic stem cells. Proc Nat Acad Sci USA. 2013;110(8):2876–81.
Arab K, Park YJ, Lindroth AM, et al. Long non-coding RNA TARID directs demethylation and activation of the tumor suppressor TCF21 via GADD45A. Mol Cell. 2014;55(4):604–14.
Quinodoz S, Guttman M. Long non-coding RNAs: an emerging link between gene regulation and nuclear organization. Trends Cell Biol. 2014;24(11):651–63.
Kwek KY, Murphy S, Furger A, et al. U1 snRNA associates with TFIIH and regulates transcriptional initiation. Nat Struct Biol. 2002;9(11):800–5.
Adelman K, Lis JT. Promoter-proximal pausing of RNA polymerase II: emerging roles in metazoans. Nat Rev Genet. 2012;13(10):720–31.
Peterlin BM, Price DH. Controlling the elongation phase of transcription with P-TEFb. Mol Cell. 2006;23(3):297–305.
Yik JH, Chen R, Nishimura R, et al. Inhibition of P-TEFb (CDK9/Cyclin T) kinase and RNA polymerase II transcription by the coordinated actions of HEXIM1 and 7SK snRNA. Mol Cell. 2003;12(4):971–82.
Ji X, Zhou Y, Pandit S, et al. SR proteins collaborate with 7SK and promoter-associated nascent RNA to release paused polymerase. Cell. 2013;153(4):855–68.
Wang ET, Sandberg R, Luo S, et al. Alternative isoform regulation in human tissue transcriptomes. Nature. 2008;456(7221):470–6.
Martianov I, Ramadass A, Barros S, et al. Repression of the human dihydrofolate reductase gene by a non-coding interfering transcript. Nature. 2007;445:666–70.
Lanz RB, Razani B, Goldberg AD, O’Malley BW. Distinct RNA motifs are important for coactivation of steroid hormone receptors by steroid receptor RNA activator (SRA). Proc Natl Acad Sci U S A. 2002;99:16081–6.
Feng J, Bi C, Clark BS, et al. The Evf-2 non-coding RNA is transcribed from the Dlx-5/6 ultraconserved region and functions as a Dlx-2 transcriptional coactivator. Genes Dev. 2006;20:1470–84.
Hah N, Danko CG, Core L, et al. A rapid, extensive, and transient transcriptional response to estrogen signaling in breast cancer cells. Cell. 2011;145:622–34.
Kim TK, Hemberg M, Gray JM, et al. Widespread transcription at neuronal activity-regulated enhancers. Nature. 2010;465:182–7.
Wang D, Garcia-Bassets I, Benner C, et al. Reprogramming transcription by distinct classes of enhancers functionally defined by eRNA. Nature. 2011;474:390–4.
Li W, Notani D, Ma Q, et al. Functional roles of enhancer RNAs for oestrogen-dependent transcriptional activation. Nature. 2013;498:516–20.
Figueroa A, Fujita Y, Gorospe M. Hacking RNA: Hakai promotes tumorigenesis by enhancing the RNA-binding function of PSF. Cell Cycle. 2009;8:3648–51.
Galietta A, Gunby RH, Redaelli S, et al. NPM/ALK binds and phosphorylates the RNA/DNA-binding protein PSF in anaplastic large-cell lymphoma. Blood. 2007;100:2600–9.
Song X, Wang B, Bromberg M, et al. Retroviral-mediated transmission of a mouse VL30 RNA to human melanoma cells promotes metastasis in an immunodeficient mouse model. Proc Natl Acad Sci U S A. 2002;99:6269–73.
Song X, Sui A, Garen A. Binding of mouse VL30 retrotransposon RNA to PSF protein induces genes repressed by PSF: effects on steroidogenesis and oncogenesis. Proc Natl Acad Sci U S A. 2004;101:621–6.
Song X, Sun Y, Garen A. Roles of PSF protein and VL30 RNA in reversible gene regulation. Proc Natl Acad Sci U S A. 2005;102:12189–93.
Li L, Feng T, Lian Y. Role of human non-coding RNAs in the control of tumorigenesis. Proc Natl Acad Sci U S A. 2009;106:16794–8.
Wang G, Cui Y, Zhang G, et al. Regulation of proto-oncogene transcription, cell proliferation, and tumorigenesis in mice by PSF protein and a VL30 non-coding RNA. Proc Natl Acad Sci U S A. 2009;106:16794–8.
Lane DP, Fischer PM. Turning the key on p53. Nature. 2004;427:789–90.
Lowe SW, Cepero E, Evan G. Intrinsic tumour suppression. Nature. 2004;432:307–15.
Vogelstein BB, Lane DD, Levine AA. Surfing the p53 network. Nature. 2000;408:307–10.
Brugarolas J, Chandrasekaran C, Gordon JI, et al. Radiation-induced cell cycle arrest compromised by p21 deficiency. Nature. 1995;337:552–7.
Yu J, Zhang L, Hwang PM, et al. Identification and classification of p53-regulated genes. Proc Natl Acad Sci U S A. 1996;96:14517–22.
Zhao R, Gish K, Mruphy M, et al. Analysis of p53-regualted gene expression patterns using oligonucleotide arrays. Genes Dev. 2000;14:981–93.
Suzuki HI, Miyazono K. Dynamics of microRNA biogenesis: crosstalk between p53 network and microRNA processing pathway. J Mol Med (Berl). 2010;88:1085–94.
Gardner EJ, Nizami ZF, Talbot Jr CC, Gall JG. Stable intronic sequence RNA (sisRNA), a new class of non-coding RNA from the oocyte nucleus of Xenopus tropicalis. Genes Dev. 2012;26:2550–9.
Qian L, Vu MN, Carter M, Wilkinson MF. A spliced intron accumulates as a lariat in the nucleus of T cells. Nucleic Acids Res. 1992;20:5345–50.
Zhang Y, Zhang XO, Chen T, et al. Circular intronic long non-coding RNAs. Mol Cell. 2013;51:792–806.
Blencowe BJ. Alternative splicing: new insights from global analyses. Cell. 2006;126(1):37–47.
Lin S, Fu XD. SR proteins and related factors in alternative splicing. Adv Exp Med Biol. 2007;623:107–22.
Long JC, Caceres JF. The SR protein family of splicing factors: master regulators of gene expression. Biochem J. 2009;417(1):15–27.
Pan Q, Shai O, Lee LJ, et al. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat Genet. 2008;40(12):1413–5.
Wang X, Arai S, Song X, et al. Induced ncRNAs allosterically modify RNA-binding proteins in cis to inhibit transcription. Nature. 2008;454:126–30.
Schmucker D, Clemens JC, Shu H, et al. Drosophila Dscam is an axon guidance receptor exhibiting extraordinary molecular diversity. Cell. 2000;101(6):671–84.
Ji P, Diederichs S, Wang W, et al. MALAT-1, a novel non-coding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene. 2003;22:8031–41.
Guffanti A, Iacono M, Pelucchi P, et al. A transcriptional sketch of a primary human breast cancer by 454 deep sequencing. BMC Genomics. 2009;10:163.
Lai MC, Yang Z, Zhou L, et al. Long non-coding RNA MALAT-1 overexpression predicts tumor recurrence of hepatocellular carcinoma after liver transplantation. Med Oncol. 2011;29:1810–6.
Lin R, Maeda S, Liu C, et al. A large non-coding RNA is a marker for murine hepatocellular carcinomas and a spectrum of human carcinomas. Oncogene. 2007;26:851–8.
Luo JH, Ren B, Keryanov S, et al. Transcriptomic and genomic analysis of human hepatocellular carcinomas and hepatoblastomas. Hepatology. 2006;44:1012–24.
Schmidt LH, Spieker T, Koschmieder S, et al. The long non-coding MALAT-1 RNA indicates a poor prognosis in non-small cell lung cancer and induces migration and tumor growth. J Thorac Oncol. 2011;6:1984–92.
Hall LL, Smith KP, Byron M, Lawrence JB. Molecular anatomy of a speckle. Anat Rec A: Discov Mol Cell Evol Biol. 2006;288:664–75.
Lamond AI, Spector DL. Nuclear speckles: a model for nuclear organelles. Nat Rev Mol Cell Biol. 2003;4:605–12.
Chen LL, Carmichael GG. Altered nuclear retention of mRNAs containing inverted repeats in human embryonic stem cells: functional role of a nuclear non-coding RNA. Mol Cell. 2009;35:467–78.
Clemson CM, Hutchinson JN, Sara SA, et al. An architectural role for a nuclear non-coding RNA: NEAT1 RNA is essential for the structure of paraspeckles. Mol Cell. 2009;33:717–26.
Sasaki YT, Ideue T, Sano M, et al. MENepsilon/beta non-coding RNAs are essential for structural integrity of nuclear paraspeckles. Proc Natl Acad Sci U S A. 2009;106:2525–30.
Sunwoo H, Dinger ME, Wilusz JE, et al. MEN varepsilon/beta nuclear-retained non-coding RNAs are up-regulated upon muscle differentiation and are essential components of paraspeckles. Genome Res. 2009;19:347–59.
Tripathi V, Ellis JD, Shen Z, et al. The nuclear-retained non-coding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol Cell. 2010;39:925–38.
Bernard D, Prasanth KV, Tripathi V, et al. A long nuclear-retained non-coding RNA regulates synaptogenesis by modulating gene expression. EMBO J. 2010;29:3082–93.
Sellier C, Rau F, Liu Y, et al. Sam68 sequestration and partial loss of function are associated with splicing alterations in FXTAS patients. EMBO J. 2010;29:1248–61.
Engreitz JM, Sirokman K, McDonel P, et al. RNA-RNA interactions enable specific targeting of non-coding RNAs to nascent pre-mRNAs and chromatin sites. Cell. 2014;159:188–99.
West JA, Davis CP, Sunwoo H, et al. The long non-coding RNAs NEAT1 and MALAT1 bind active chromatin sites. Mol Cell. 2014;55:791–802.
Lin R, Roychowdhury-Saha M, Black C, et al. Control of RNA processing by a large non-coding RNA over-expressed in carcinomas. FEBS Lett. 2011;585:671–6.
Tripathi V, Shen Z, Chakraborty A, et al. Long non-coding RNA MALAT1 controls cell cycle progression by regulating the expression of oncogenic transcription factor B-MYB. PLoS Genet. 2013;9:e1003368.
Gutschner T, Hämmerle M, Eissmann M, et al. The non-coding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res. 2013;73:1180–9.
Zhang B, Arun G, Mao YS, et al. The lncRNA Malat1 is dispensable for mouse development but its transcription plays a cis-regulatory role in the adult. Cell Rep. 2012;2:111–23.
Yin QF, Yang L, Zhang Y, et al. Long non-coding RNAs with snoRNA ends. Mol Cell. 2012;48:219–30.
Zhang XO, Yin QF, Wang HB, et al. Species-specific alternative splicing leads to unique expression of sno-lncRNAs. BMC Genomics. 2014;15:287.
Matera AG, Terns RM, Terns MP. Non-coding RNAs: lessons from the small nuclear and small nucleolar RNAs. Nat Rev Mol Cell Biol. 2007;8:209–20.
Kapranov P, Cheng J, Dike S, et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science. 2007;316:1484–8.
Keniry A, Oxley D, Monnier P, et al. The H19 lincRNA is a developmental reservoir of miR-675 that suppresses growth and lgf1r. Nat Cell Biol. 2012;14:659–65.
Yuan JH, Yang F, Wang F, et al. A long non-coding RNA activated by TGF-β promotes the invasion-metastasis cascade in hepatocellular carcinoma. Cancer Cell. 2014;25:666–81.
Cesana M, Cacchiarelli D, Legnini I, et al. A long non-coding RNA controls muscle differentiation by functioning as a competing endogenous RNA. Cell. 2011;147:358–69.
Franco-Zorrilla JM, Valli A, Todesco M, et al. Target mimicry provides a new mechanism for regulation of microRNA activity. Nat Genet. 2007;39:1033–7.
Karreth FA, Tay Y, Perna D, et al. In vivo identification of tumor-suppressive PTEN ceRNAs in an oncogenic BRAF-induced mouse model of melanoma. Cell. 2011;147:382–95.
Poliseno L, Salmena L, Zhang J, et al. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature. 2010;465:1033–8.
Sumazin P, Yang X, Chiu HS, et al. An extensive microRNA-mediated network of RNA-RNA interactions regulates established oncogenic pathways in glioblastoma. Cell. 2011;147:370–81.
Tay Y, Kats L, Salmena L, et al. Coding-independent regulation of the tumor suppressor PTEN by competing endogenous mRNAs. Cell. 2011;147:344–57.
Conn SJ, Pillman KA, Toubia J, et al. The RNA binding protein quaking regulates formation of circRNAs. Cell. 2015;160:1125–34.
Guo JU, Agarwal V, Guo H, Bartel DP. Expanded identification and characterization of mammalian circular RNAs. Genome Biol. 2014;15:409.
Jeck WR, Sorrentino JA, Wang K, et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013;19:141–57.
Memczak S, Jens M, Elefsinioti A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495:333–8.
Salzman J, Gawad C, Wang PL, et al. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS ONE. 2012;7:e30733.
Salzman J, Chen RE, Olsen MN, et al. Cell-type specific features of circular RNA expression. PLoS Genet. 2013;9:e1003777.
Wilusz JE, Sharp PA. Molecular biology. A circuitous route to non-coding RNA. Science. 2013;340:440–1.
Hansen TB, Jensen TI, Clausen BH, et al. Natural RNA circles function as efficient microRNA sponges. Nature. 2013;495(7441):384–8.
Zhang A, Zhou N, Huang J, et al. The human long non-coding RNA-RoR is a p53 repressor in response to DNA damage. Cell Res. 2013;23:340–50.
Yoon JH, Abdelmohsen K, Srikantan S, et al. LincRNA-p21 suppresses target mRNA translation. Mol Cell. 2012;47(4):648–55.
Beltran M, Puig I, Peña C, et al. A natural antisense transcript regulates Zeb2/Sip1 gene expression during Snail1-induced epithelial-mesenchymal transition. Genes Dev. 2008;22(6):756–69.
Carrieri C, Cimatti L, Biagioli M, et al. Long non-coding antisense RNA controls Uchl1 translation through an embedded SINEB2 repeat. Nature. 2012;491(7424):454–7.
Liu B, Sun L, Liu Q, et al. A cytoplasmic NF-κB interacting long non-coding RNA blocks IκB phosphorylation and suppresses breast cancer metastasis. Cancer Cell. 2015;27(3):370–81.
Wang P, Xue Y, Han Y, et al. The STAT3-binding long non-coding RNA lncc-DC controls human dendritic cell differentiation. Science. 2014;344(6181):310–3.
Wutz A, Rasmussen TP, Jaenisch R. Chromosomal silencing and localization are mediated by different domains of Xist RNA. Nat Genet. 2002;30:167–74.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media Singapore
About this chapter
Cite this chapter
Li, L., Song, X. (2016). The Working Modules of Long Noncoding RNAs in Cancer Cells. In: Song, E. (eds) The Long and Short Non-coding RNAs in Cancer Biology. Advances in Experimental Medicine and Biology, vol 927. Springer, Singapore. https://doi.org/10.1007/978-981-10-1498-7_2
Download citation
DOI: https://doi.org/10.1007/978-981-10-1498-7_2
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-1496-3
Online ISBN: 978-981-10-1498-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)