Abstract
The stem cell epigenome reflects a sensitive balance of chromatin (de-)modification processes. Here, we review our recent achievements towards a mechanistic understanding of this balance.
We introduce a computational model of stem cell populations, where each cell contains an artificial genome. Transcription of the genes encoded by this genome is controlled by DNA methylation, histone modification and the action of a cis-regulatory network. Model dynamics are determined by molecular crosstalk between these different mechanisms.
The epigenetic states of the genes are subject to different types of fluctuations. We demonstrate that the timescales of these fluctuations control whether the state associated with a particular gene will undergo drifts during ongoing cell replication. In particular, our model suggests that changes in DNA methylation states are determined by histone modification dynamics. Herewith, our model provides a mechanistic understanding of the origin of tissue, age and cancer-specific DNA methylation profiles.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
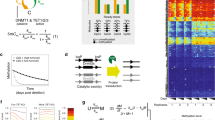
Among the plethora of known chromatin modifications, DNA methylation is probably the one that has been analyzed in most detail. Changes in DNA methylation are observed during stem cell differentiation and ageing and also in the course of many diseases (Bergman and Cedar 2013). A particular well-documented phenomenon is hyper-methylation of CpG-rich promoters during cancer development (Berdasco and Esteller 2010). This local increase of CpG methylation is often associated with a down-regulation of expression of the affected genes and thus can induce cancer phenotypes. In fact, DNA methylation patterns have been used in several tissues to classify cancer subtypes with different clinical outcomes (Hinoue et al. 2012; Sturm et al. 2012) (see Fig. 7.1).
Hierarchical clustering of DNA methylation pattern of human colorectal cancer (CRC) samples. Shown are methylation levels (red: low, white: high) that have been calculated based on 27k Illumina methylation arrays from 37 healthy and 53 tumour samples (columns). They have been clustered using the 200 most variant CpGs (rows). Data were taken from the TCGA data repository (Network 2012). Most of the CpGs become hyper-methylated in CRC, while only a few become hypo-methylated. The patterns allow to distinguish two or three different CRC methylation patterns. The origin of this epigenetic reorganization remains largely unknown
Regardless of the enormous amount of molecular data collected so far, a mechanistic understanding of how the observed changes in DNA methylation are induced and how they impact gene expression is still largely missing. Within the last years, many experimental groups observed correlations between DNA methylation and other chromatin marks. One of the first examples here was an observation made for a certain group of genes whose promoters become hyper-methylated during ageing and cancer development. It was found that the nucleosomes associated with the promoter of these genes are often tri-methylated at lysine 4 and 27 of histone 3 (Teschendorff et al. 2010; Rakyan et al. 2010). Such findings suggest that there exists a complex molecular crosstalk between the machinery of DNA methylation and that of histone modification. Some specific molecular interactions have been identified in the meanwhile (Rose and Klose 2014).
Here, we will demonstrate that mathematical modelling of this kind of molecular crosstalk can provide new insights into regulatory principles of the epigenome and can help to establish a mechanistic understanding of epigenetic reorganization, e.g. following loss of tissue homeostasis. For this purpose, in the following, we provide a brief introduction into a multi-scale model of epigenetic regulation of transcription. First, we introduce its molecular components enabling to describe DNA methylation, histone modification and cis-regulatory networks. Afterwards, we explain its extension to the cell population level. Finally, we provide some first simulation results on epigenetic changes during stem cell ageing and tissue transformation.
2 Modelling DNA Methylation Dynamics
2.1 Background
The role of DNA methylation (here: 5-mC methylation) in cancer development has been recognized already more than 40 years ago (Magee 1971); yet, the molecular details of the enzymatic machinery leading to establishment and maintenance of DNA methylation in normal tissue remained unknown for a long time. Despite missing knowledge about the enzymes involved, first mathematical models for DNA-methylation dynamics were proposed already in the 1990s (Otto and Walbot 1990; Pfeifer et al. 1990).
These models tried to explain the conservation of methylation states given the fact that all CpGs on the de novo synthesized DNA daughter strands are initially unmethylated. They proposed that this passive de-methylation is compensated by simultaneous action of maintenance and de novo methylation, leading to a genome-wide methylation equilibrium after a finite number of replication cycles. DNA methyltransferases (Dnmts) involved in these processes were identified experimentally some years later. Three types of Dnmts were identified in mammals, namely, Dnmt3a, Dnmt3b and Dnmt1. De novo methylation has been attributed to the action of the isoforms Dnmt3a and Dnmt3b (Okano et al. 1999), while Dnmt1 was found to be mainly responsible for maintaining the parental methylation pattern in daughter cells (Pradhan et al. 1999).
Improved models of DNA methylation, which considered these experimental findings, were introduced by Sontag et al. (2006). They introduced a linear model for independent action of Dnmt1 and Dnmt3a/b and a nonlinear model assuming cooperative dynamics between them.
2.2 Basic Model of DNA Methylation
Since experimental support for cooperation between Dnmts is still limited, we designed a first layer of our multi-scale model adopting a simple version of the linear model by Sontag et al. (see Fig. 7.2a). We assume a single-stranded DNA molecule. Accordingly, all CpG sites of a finite DNA region (N CG CpG sites) are either methylated or not. CpG methylations existing in the mother cell (n CG) are restored by maintenance methylation with probability D main in their daughters, while CpG sites that are unmethylated in the mother (N CG − n CG) might become de novo methylated in the daughters with probability D novo. In our model, de novo methylation acts only on CpG sites that were unmethylated in the mother and not on those where methylation has not been restored in the daughters.
Model of DNA methylation. (a) Sketch of the processes considered in our model. Maintenance methylation by Dnmt1 and de novo methylation by Dnmt3a,b are either active with rate D main and D novo, or inactive with rate 1-D main and 1-D novo, respectively. (b) Different pairs (D novo, D main) can result in the same methylation level as seen for solutions of the deterministic system applying (0.05/τ, 0.95/τ), red curve, and (0.3/τ, 0.7/τ), yellow curve. Stochastic simulations for 30 CpGs show large fluctuations (magenta, cyan, blue lines) around the solution of the deterministic system (black line). (c) Transition probabilities for the stochastic system and the basic set of equations for the deterministic system
We assume that DNA methylation can occur only immediately after cell replication, in a short time frame compared to the cell cycle time τ. This effectively makes DNA methylation levels a direct function of the number of successive replication events. Therefore, following Sontag et al., we implemented a probabilistic model with discrete time steps and discrete updates of CpG-methylation states. Formally, this defines a discrete Markov chain model with transition probabilities D novo and D main. The transition scheme for our model is shown in Fig. 7.2c (left part). The thin-lined curves in Fig. 7.2b show the results of three different simulations of the stochastic model. Shown is the fraction m (given by n CG/N CG) of methylated CpGs that has been observed by analyzing the methylation dynamics of 30 CpGs.
Alternatively, changes of m during ongoing replication can be analyzed using a differential equation approach (see Fig. 7.2c, right part). Such a time continuous approach is helpful to estimate methylation equilibria and convergence times. The solution of this equation yields an exponential increase or decrease to an equilibrium methylation level, depending on the initial methylation level m 0. The equilibrium methylation is given by D novo/r. Examples of the temporal dynamics are shown in Fig. 7.2b. Note that D novo and D main represent modification rates per cell cycle time τ. Different combinations of them can lead to convergence to the same methylation level; yet, the time needed for convergence differs.
Our stochastic DNA methylation model is limited in some regards. For instance, the model alone cannot explain the coexistence of hyper- and hypo-methylated states after a large number of replications, as observed in aged tissues and during cancer development (Bergman and Cedar 2013). In their nonlinear model, Sontag et al. explained these phenomena suggesting that the efficiency of de novo methylation depends on the density of hemi-methylated sites observed after DNA replication but before maintenance and/or de novo methylation. A similar model was recently proposed by Haerter et al. (2014). Here, we use a different approach.
It is well known that recruitment of de novo Dnmts strongly depends on local histone modification states (Rose and Klose 2014). This suggests that coexisting DNA methylation states are controlled by local histone modification states. In the following, we introduce a model of cooperative histone modification dynamics which enables us to describe such regulation.
3 Modelling Histone Modification Dynamics
3.1 Background
Today, a huge number of chemical modifications on histone tails are known, including methylation, acetylation, phosphorylation, sumoylation and ubiquitination. These modifications can contribute to activation or repression of gene expression. Their combinatorial complexity is further increased by the possibility of different modification levels, e.g. mono-, di- or tri-methylation (me1, me2 or me3). Thereby, different levels of modification might have different effects on chromatin structure and transcription (Hoffman et al. 2013).
Several theoretical models of histone methylation and acetylation dynamics have been proposed so far (Dodd et al. 2007; Sedighi and Sengupta 2007). For a review of these models, see Rohlf et al. (2012). A common feature of these models is that they are based on cooperative modification dynamics. In our model of histone methylation, we enable cooperative behaviour by assuming that modified nucleosomes cooperatively recruit their own histone methyltransferase (HMT).
In the following, we first introduce a model of histone methylation for a finite number of cooperatively acting nucleosomes (Binder et al. 2013). Afterwards, we outline our strategies to integrate crosstalk between histone and DNA methylation into the model.
3.2 Basic Model of Histone Methylation
In our model, we only consider modification complexes that can write and read a specific modification. This is motivated by properties of polycomb group (PcG) and trithorax group (trxG) proteins (Kundu and Peterson 2009). The basic assumptions of our model regarding a reader–writer complex catalyzing histone modifications, in the following called ‘interaction complex’ (IC), are summarized in Fig. 7.3a. The regulatory processes are explained for an activating modification.
Model of histone methylation. (a) Sketch of the regulatory interactions between ICs and chromatin. For an explanation, see text. (b) Fluctuations of the methylation level within a CU containing 48 nucleosomes, as derived from a stochastic model realization (blue curve). Appreciate the sudden switch from high to low modification level after about 5,000 simulation time steps and back about 3,500 steps later. The two stable solutions of the deterministic system are shown for comparison (red and green curves). (c) Basic equations of the model. In S1–S4, the X labels the lysine, e.g. 4 or 9
We assume that each IC binds to one DNA-response element (RE) which contains a variable number n BS of binding sites (BS). Specifically, we identify BS with CpGs. Binding to CpGs depends on their methylation state. Adjacent REs form cooperative units (CUs) of length L CU, given in units of the number of base pairs (bp) involved. Formation of CUs might occur via chromatin looping as proposed by Tiwari et al. (2008). Each CU is associated with N H = L CU/200 nucleosomes, where n HM of them are in a modified (HM) histone state and the remaining N H − n HM are in an unmodified (H0) histone state. We call a nucleosome modified if one of its histones is modified. In addition to the DNA BS, also the n HM-modified nucleosomes within a CU facilitate IC binding. Bound ICs catalyze histone modifications, giving rise to a positive feedback loop between IC binding and histone modification. Gene transcription is activated after IC binding and repressed after IC release.
In our multi-scale model, we implemented a stochastic version of this model at the single nucleosome level. The transition probabilities for the nucleosome states per simulation time step Δt are given in Fig. 7.3c (left part). We assume that de novo modification of a histone can occur only if an IC is bound to a nearby RE. The probability of this state is quantified by the RE-occupancy Θ, which can take values between 0 and 1. Accordingly, the probability of de novo modification is given by k M Θ, where k M is a constant. De-modification events are assumed to occur permanently with probability k D.
Using arguments from mass action kinetics, the binding process of an IC can be formalized (Binder et al. 2013). The resulting equation for the RE-occupancy Θ is given in Fig. 7.3c (Eq. D1). It is governed by the free enthalpy change Δg of IC binding, which can be decomposed into a basic repulsive term ε 0 >0, and two attractive terms n BS ε BS <0 and n HM ε HM <0 representing the enthalpy changes according to binding of the IC to n BS DNA-binding sites and to n HM nucleosomes of the CU which already carry the IC-specific modification, respectively.
In Fig. 7.3b, simulation results are given for a realization of the stochastic modification process within a single CU of 48 nucleosomes. For the chosen parameter set, the system shows bistable behaviour. Stochastic fluctuations lead to switches between the two attractor states.
The temporal dynamics of the fraction θ HM (given by n HM/N H) of modified histones in a CU can be described by a differential equation (Eq. D2 in Fig. 7.3c), similar to the DNA methylation process. Here, the terms k D, k MΘ describe rates per simulation time step Δt. Again this approach allows estimating equilibrium states of the system. It can be shown that, for relatively wide parameter ranges, the system exhibits bistable behaviour (Rohlf et al. 2012; Binder et al. 2013). Thereby, the solutions strongly depend on the number N H of nucleosomes contained in the CU. In Fig. 7.3b, the solutions of the deterministic system (red and green lines) are compared with those of the stochastic process.
The above model can be extended easily to combinations of histone modifications. For this purpose, one has to take into account sets of transition probabilities and model parameters for every modification. Thereby, different modifications might influence each other, either directly or indirectly. In the following, we study tri-methylation of lysine 4 at histone 3 (H3K4me3) in parallel with tri-methylation of lysine 9 at histone 3 (H3K9me3). We assume that these two modifications affect each other only indirectly via their effects on DNA methylation. This kind of crosstalk is described in the following.
3.3 Crosstalk Between DNA and Histone Methylation
It is in general accepted that there is a complex crosstalk between histone modifications and DNA methylation (D’Alessio and Szyf 2006). For example, several HMTs have been demonstrated to include binding motifs either for unmethlyated (CXXC, e.g. HMTs writing H3K4me3 (Thomson et al. 2010; Fujita et al. 2003)) or for methylated (MDB, e.g. HMTs writing H3K9me3 (Fujita et al. 2003)) CpGs. So, on the one hand, local DNA methylation status impacts the recruitment of HMTs. On the other hand, the recruitment of Dnmts is affected by histone modifications. For example, H3K4me3 has been demonstrated to repel Dnmt3a (Ooi et al. 2007), whereas H3K9me3 recruits it (Feldman et al. 2006).
We model this kind of crosstalk by accounting for effects of DNA methylation on IC (i.e. HMT) recruitment. We assume that unmethylated and methylated CpGs throughout the CUs act as binding sites for the ICs catalyzing H3K4me3 and H3K9me3, respectively. In addition, we account also for effects of the histone modifications on the recruitment of Dnmts. This is achieved by modifying the de novo Dnmt probability by a factor that depends on the actual histone methylation level of the associated nucleosomes. For details, see Przybilla et al. (2013, 2014).
Simulation of the impact of DNA methylation and histone modifications on gene transcription requires model representations of the genes controlled by these epigenetic marks. For this purpose, we adopt an artificial genome model, which defines a transcription factor (TF) network that exhibits realistic gene expression properties.
4 Modelling TF Networks: The Artificial Genome Approach
Artificial genomes (AGs) were originally introduced to generate gene regulatory networks that cover important biological features of real TF networks. The first idea of building an AG was published by Reil (1999). In extension, we introduced mechanisms to analyze structural evolution in AGs (Rohlf and Winkler 2009). Moreover, we added a thermodynamic model of transcriptional regulation (Binder et al. 2010), which was adapted from Bintu et al. (2005). In its present form, the model allows calculations of gene expression based on the DNA binding and regulatory action of two types of TFs, namely, repressors and activators. Moreover, it enables straightforward integration of our DNA and histone methylation model. Thus, it represents an ideally suited backbone of our multi-scale model of transcriptional regulation.
4.1 Construction of an AG
According to the suggestions by Reil (1999), we generate an AG of length L genome by calculating a random string composed of four different digits [0,1,2,3], where each digit denotes one DNA base [A,T,C,G] (Fig. 7.4a). We consider a single strand DNA only, neglecting all effects caused by the second DNA strand. We are using L genome = 400.000 ‘bases’. Motivated by the frequent association of promoters with TATA-boxes, we assume all short sequences [010100] that are found on the AG to represent base promoters of genes. The next L cod = 8 bases downstream of the promoters are assumed to represent their coding regions. All bases upstream of a gene up to the end of the coding region of the preceding gene define the regulatory region L reg of the gene. Its length can be different for every gene. According to these assumptions, each gene of the AG is divided into three regions, a regulatory, a base promoter and a coding region. All genes potentially encode TFs.
The artificial genome (AG). (a) All sequences [010100] are considered to represent base promoters. They divide the genome into genes of different length. The 8 digits downstream this sequence ([01201312] for the most right gene) define a transcript, which is translated into a protein with a specific DNA binding motif (here [12312023]). The protein can bind to DNA wherever this motif occurs. Bound proteins act as TF and regulate the nearest downstream gene (green arrow). (b) Distribution of the transcription values of AG genes for a fraction of repressors p a = 0.74. Shown is an average over 100 AGs (black line) applying the data set described in the text (green line: smoothed version). The result is compared with a distribution measured for colon tissue by RNA-seq (Network 2012) (red line). (c) Basic equations enabling calculation of expression T i of gene i. The product in D2 runs over all TFs j regulating gene i
All TFs together form a TF network. This network is constructed by the following rules: Each coding region of a gene defines a transcript. The transcript is translated into a protein with a specific binding motif. This motif is calculated by applying a simple transition rule: each digit of the coding region is updated by adding 1, if the sum is 4 it is replaced by 0. Accordingly, a coding sequence (01201312) will be translated to binding motif (12312023). The protein can bind to identical DNA motifs in the regulatory regions of all genes and act as a TF. Bound TFs regulate the next downstream gene.
According to these building rules, each AG has an intrinsic TF-network structure, which is completely defined by the promoter length, the length of the coding region, and the length of the genome. Using the parameters given above, each TF can regulate on average 6 different genes and each gene can be regulated on average by 6 TFs. For statistical properties of such kind of TF networks, we refer to Binder et al. (2010).
Whether a TF binds to DNA or not depends on the binding energies ε TF and on the concentration of the TF which is set to be equal to the expression level T of its transcript. Bound TFs regulate the occupancy θ Pro of the nearest downstream promoter by RNA-polymerase II by changing the polymerase binding energy. Activators increase it and repressors decrease it. Whether a TF is an activating or a repressing one is chosen with probability p a and (1-p a), respectively. The promoter occupancy θ Pro,i of the base promoter of gene i is assumed to be proportional to the transcription of the gene. A typical distribution of the expression values of an AG is shown in Fig. 7.4b. It agrees very well with experimentally measured distributions.
The equations describing how the transcription T i of an individual gene i is calculated for our AG are given in Fig. 7.4c. Here, δ is the degradation constant of the transcript and P max is the maximum promoter activity. Both parameters are assumed to be identical for all genes of the AG.
4.2 Crosstalk Between Chromatin Modifiers and Polymerase II Binding
Our AG model describes regulation of gene expression by a TF network. However, experimental results indicate that there is in addition a complex interplay between the expression of genes and their epigenetic status (Cui et al. 2009). In the following, we summarize our assumptions regarding this kind of crosstalk. A sketch describing the interactions considered in the model is given in Fig. 7.5.
First of all, implementing such crosstalk requires linking the AG model to our model of epigenetic regulation of transcription. For that one has to specify the nucleosomes whose status can contribute to the regulation of a particular gene. Given the structure of our AG, we assume that the modification of all nucleosomes associated with the regulatory region of gene i affect the transcription of this gene. This means that we identify the regulatory regions of the genes of our AG with the CUs of our histone model. The modification state of a particular gene is thus the modification state of the nucleosomes associated with the regulatory region of this gene.
As already pointed out in Sect. 7.3, histone modifications can activate as well as repress gene expression. The H3K4me3 mark has been suggested to contribute to gene activation via an improved recruitment of polymerase II in presence of this modification (Vermeulen et al. 2007). In our model we assume that the transcription level of gene i is proportional to the occupancy Θ i H3K4me3 of the REs of this gene by H3K4me3-specific ICs; i.e. by H3K4-HMTs. Accordingly, we replace P max in Eq. D1 in Fig. 7.4 by P max Θ i H3K4me3 . H3K9me3 is frequently associated with silenced genes. However, we assume that it affects transcription only indirectly via its activity in recruiting de novo DNA methylation.
Recently, it has been shown that not only transcription depends on the histone modification state of the gene but that, vice versa, also the stability of the histone modification states is affected by the transcriptional activity of the associated genes. Experimental findings suggest that the C-terminal domain of the RNA polymerase II subunit Rpb1 undergoes dynamic phosphorylation and that this process helps recruiting the H3K4-HMT complex during early elongation (Buratowski and Kim 2010). In our model, we assume that the recruitment of H3K4me3-specific ICs is enforced at transcribed promoters. Actually, we assume that the basic repulsive term ε 0 of the free enthalpy of binding of H3K4me3-specific ICs is equal to ε 1-ln(Tδ/P max), where ε 1 is a constant. This leads to a positive feedback, stabilizing H3K4me3 at transcribed regions. Binding of H3K9me3-specific ICs is assumed to be not affected by transcription. Accordingly, we assume ε 0 = ε 2 to be constant for this modification.
The assumed crosstalk determines the dynamics of our multi-scale model. In particular, it controls the stability of its regulatory states. In the next section, we describe how these states can be inherited through iterative replication cycles and how regulatory states evolve on the population scale.
5 Modelling Cell Population Behaviour
5.1 Model of Cell Replication
In our model, gene expression depends on the histone modification states of the genes. These states are coupled to the local DNA methylation status, which changes during cell division. These changes are different between the daughter cells. Consequently, long-term drifts of transcription, DNA and histone methylation states of the cells can be captured by explicit simulation of cell replication only, and their analysis requires the simulation of large cell populations. We model cell replication assuming that each cell undergoes stochastic growth steps with rate R and divides after N R successful growth steps.
The changes in DNA methylation during cell division (after initial equilibration) due to limited maintenance and de novo methylation are rather moderate perturbations of the regulatory state of the cells. Much stronger changes can result in parallel from processes of nucleosome re-assembly. During cell division, the core nucleosomes of the mother cell are distributed onto the daughter cells and there become complemented by de novo synthesized, unmodified nucleosomes. This leads to a strong dilution of the modified nucleosomes in the daughter cells. These changes can be different in each daughter due to an unequal distribution of modified nucleosomes of the mother cell onto its daughters (Margueron and Reinberg 2010). In accordance with experimental results (Xu et al. 2010), we assume a random distribution (see Fig. 7.6). In parallel, we assume that de novo synthesis of unmodified nucleosomes guarantees that in the daughter cells the same number of nucleosomes is established as in the mother cell. Thus, we neglect any variance in this property.
Epigenetic changes during cell replication. During cell replication, the mother nucleosomes are randomly distributed onto the daughter strands and there become complemented with de novo synthesized, unmodified nucleosomes. Thus, the number of modified nucleosomes on each daughter strand becomes diluted. In parallel, DNA methylation state becomes updated separately on each strand. Both processes can induce an asymmetric phenotype in the daughter cells
The histone modification states immediately after cell division are non-equilibrium states and drift until stable states are reached. If the modification state of a particular gene is bistable, the strong dilution of modified histones after cell division can lead to spontaneous de-modification of the histones. As a result, an asymmetry of the daughters with respect to the modification state of these genes will become manifest. These changes might induce subsequent transcriptional changes and those even further regulatory changes. The regulatory states approached after cell division can be different across daughter cells and thus can induce a strong heterogeneity in an expanding cell clone on all levels of regulation. In order to cover this heterogeneity, we model cell populations, where in each cell the same regulatory network is active but undergoes independent development of its regulatory states.
5.2 Regulatory States at the Population Level
Transcriptional states (Wu et al. 2014) and in part DNA methylation states (Guo et al. 2013) of individual cells can be measured experimentally. In contrast, histone modification states are currently accessible on the population level only. In our model, all these states are calculated on the single cell level. Thus, to compare model results with available experimental data, we have to average the regulatory states of the individual genes over a population of cells.
An example of such a calculation is given in Fig. 7.7a showing simulated epigenetic drifts in a proliferative population. The parameter set used in this simulation is given in Table 7.1. In this example, two histone modification states, the H3K4me3 and H3K9me3 state, have been considered. Initially all nucleosomes were marked by both modifications. Due to the dynamics described above, H3K4me3 as well as H3K9me3 is lost within the regulatory region of many genes over time. Loss of H3K4me3 induces decreasing transcription and enables, as de novo DNA methylation is no longer blocked, stabilization of the repressed state by DNA methylation. Thereby, the stability of the modification depends on the number of cooperative nucleosomes. In order to visualize this phenomenon, we have sorted the genes from bottom to top by the increasing number N H of cooperatively acting nucleosomes associated with the gene. For low numbers, loss of H3K4me3 occurs fast, while for higher numbers it slows down and modified states become stable. Similar effects are observed for H3K9me3. Here, conservation of the initial modification state is associated with DNA methylation.
Simulated drifts on the population level. Shown is the development of regulatory states for a proliferative active (a) and a quiescent (b) cell population. Regulatory states of individual genes are characterized by their transcription level (Log10(T)), their DNA methylation level and by the level of H3K4me3 and H3K9me3 modification of the associated nucleosomes. In their initial state, the expression of all genes was set to the equilibrium state of the isolated AG; DNA methylation level was set to 0 (red) and the histone modification levels to 1 (white). The genes have been sorted from bottom to top by the increasing number N H of nucleosomes associated with them. Drifts of the regulatory states are seen for both compartments. In the quiescent population, no changes of the DNA methylation status can occur
The effect of cell division on the regulatory states can be analyzed comparing the dynamics of the regulatory states of proliferative active (Fig. 7.7a) with that of quiescent cells (Fig. 7.7b). In a quiescent cell, the number of cooperating nucleosomes that is required to ensure stable modification decreases and more genes remain stably modified. This is due to inactive DNA methylation, allowing de-modified nucleosomes to become modified again following fluctuations in their histone modification status.
Recently, we have shown that, according to this mechanism, age-dependent drifts in histone modification states are partly reversible if the cells become located in a quiescent niche (Przybilla et al. 2014). In this study, we simulated hematopoietic stem cells in their niche. In order to cope with experimental data (Dykstra et al. 2011; Verovskaya et al. 2013), we enabled the cells to switch between a compartment where they are proliferative active and one where they are quiescent. We assumed the transition rates per simulation time step between the compartments to depend on the number of cells in the compartment they leave. This enables a stabilization of the number of cells in each compartment. Alternative assumptions are described by Glauche et al. (2009).
6 Application of the Model: DNA Methylation Profiles in Tumours
Simulation of stem cell ageing and tissue transformation were major objectives guiding the development of our multi-scale model. In such simulations, we derive hypotheses about the mechanisms underlying the associated changes of the epigenome. So far, we have linked ageing to the limited inheritance of histone modification states (Przybilla et al. 2014) and suggested that epigenetic drifts during tissue transformation originate in an accelerated ageing process, which is often paralleled by drifts induced by mutation of epigenetic pathways (Przybilla et al. 2013).
Figure 7.8 summarizes some of our simulation results. Shown are results of a hierarchical clustering of DNA methylation pattern. These results have been obtained analyzing the consequences of changing activity and of mutations of chromatin modifiers. It can be see that changes of the modifiers can induce both DNA hyper- and hypo-methylation phenotypes. As expected, hypo-methylation is seen for inefficient DNA maintenance methylation (D main = 0.5). However, similar patterns are induced also by a knock-out of the H3K4 histone demethylase (HDM) activity (90 % reduction). Hyper-methylation is induced by a knock-out of the H3K9 HDM activity (90 % reduction) and also in case of decelerated histone modification dynamics (90 % reduction). The latter two patterns are distinguished by the methylation level of only a few genes (compare: cluster X). This suggests that even small groups of CpGs could be very important markers for specific kinds of deregulation.
Hierarchical clustering of simulated DNA methylation pattern. Shown are results for five different simulation scenarios, each of them in three replicates. The colour code quantifies the average methylation of CpGs within the regulatory region of genes at a defined time point (red: low, white: high). We have selected the 40 most variant genes. The characters denote different regulatory conditions, A normal ageing, B slow histone modification velocity, C inefficient maintenance DNA methylation, D knock-out of the H3K4 HDM activity, E knock-out of the H3K9 HDM activity. Replicates cluster together on the first level. At the second level, conditions C and D and F and B cluster, due to their similar hypo- and hyper-methylation pattern, respectively. Hyper-methylation pattern F and B can be distinguished by the methylation of the CpG subset X
Overall these results demonstrate that our model is capable of explaining complex changes in DNA methylation pattern by changes in individual chromatin modification pathways. The model suggests that changes in DNA methylation pattern are governed by histone modification dynamics.
7 Discussion
Transcriptional changes during stem cell differentiation and also during tissue transformation are commonly thought to be induced by changes in cis-regulatory networks. Chromatin modifications appear to function in stabilization of these changes (Wutz 2013). However, chromatin reorganization can neither establish completely stable nor perfectly inheritable transcriptional states because cell replication induces strong perturbations of the regulatory states. As a consequence, continuous replication results in epigenetic drifts that contribute in controlling the emergence of age-related phenotypes.
We here have introduced a multi-scale model of transcriptional regulation that combines models of DNA and histone methylation with a model of cis-regulatory networks. The combined model enables to analyze the temporal changes of global regulatory states and their dependence on the activity of the individual regulatory layers. Moreover, it allows to generate substantial hypotheses about the interrelations between the different layers of transcriptional regulation and about the potential changes following loss or gain of function in chromatin modification. We have shown how this model can be extended to simulate regulatory phenomena in proliferative active cell populations and that proliferation does strongly feedback on the states of the epigenome.
Our multi-scale model clearly contains various simplifications. For instance: (1) our cis-regulatory model is based on a single strand AG that does neglect evolutionary developed non-random structures, (2) our model of DNA methylation does not consider active DNA de-methylation, (3) the model of histone modification describes only a specific kind of potentially inheritable modifications, namely those set by reader-writer complexes, and focuses on modifications of the cis-regulatory regions only.
Regardless of these simplifications our model provides new insights into transcriptional regulation, e.g. by pointing to the importance of the timescale ratio between proliferation and histone modification for the stability of regulatory states (Przybilla et al. 2013, 2014). Moreover, our model adds new arguments to the histone code debate, suggesting that chromatin computation acts on a very restricted state space, because only a few of the possible combinatorial states are stable.
Although covering several time and length scales of transcriptional regulation, our model still might lack some important regulatory processes. As an example we like to highlight 3D chromatin organization. Changes in the 3D organization potentially affect the cooperative behaviour of the histone modification process, and thus might substantially impact the regulatory states. Actually, we and others observed a dramatic change in the length distribution of specifically modified chromatin during stem cell differentiation processes (Steiner et al. 2012).
In the model, simulations presented here we largely neglected extrinsic regulation of the epigenome. In fact, environmental effects have been considered only by assuming compartment-specific signals that support or block proliferation. However, there are many more environmental signals affecting DNA and histone methylation (Burgess et al. 2014). These signals often depend on the spatial position of the cells in the tissue. For example, stem cells in spatially structured niches, e.g. intestinal stem cells, have been shown to receive local signals that trigger their phenotype. The associated regulatory changes involve also epigenetic changes (Sheaffer et al. 2014). As a first example of a spatially structured model, we plan simulating an intestinal crypt where an artificial genome is transcribed in each of the cells.
Models of the transcriptional regulation by epigenetic processes, as mathematical models in general, will never be comprehensive. However, even at the current state of the art, they allow to generate experimentally testable hypotheses about the mechanisms driving global re-organization of the stem cell epigenome. Thus, computational model approaches, as that presented here, are well on the way to support a better understanding of epigenetic dynamics during differentiation, ageing and tissue transformation.
References
Berdasco M, Esteller M. Aberrant epigenetic landscape in cancer: how cellular identity goes awry. Dev Cell. 2010;19:698–711.
Bergman Y, Cedar H. DNA methylation dynamics in health and disease. Nat Struct Mol Biol. 2013;20:274–81.
Binder H, Wirth H, Galle J. Gene expression density profiles characterize modes of genomic regulation: theory and experiment. J Biotechnol. 2010;149:98–114.
Binder H, Steiner L, Przybilla J, Rohlf T, Prohaska S, Galle J. Transcriptional regulation by histone modifications: towards a theory of chromatin re-organization during stem cell differentiation. Phys Biol. 2013;10:026006.
Bintu L, Buchler NE, Garcia HG, Gerland U, Hwa T, Kondev J, Kuhlman T, Phillips R. Transcriptional regulation by the numbers: applications. Curr Opin Genet Dev. 2005;15:125–35.
Buratowski S, Kim T. The role of cotranscriptional histone methylations. Cold Spring Harb Symp Quant Biol. 2010;75:95–102.
Burgess RJ, Agathocleous M, Morrison SJ. Metabolic regulation of stem cell function. J Intern Med. 2014;276:12–24.
Cui K, Zang C, Roh T-Y, Schones DE, Childs RW, Peng W, Zhao K. Chromatin signatures in multipotent human hematopoietic stem cells indicate the fate of bivalent genes during differentiation. Cell Stem Cell. 2009;4:80–93.
D’Alessio AC, Szyf M. Epigenetic tête-à-tête: the bilateral relationship between chromatin modifications and DNA methylation. Biochem Cell Biol. 2006;84:463–76.
Dodd IB, Micheelsen MA, Sneppen K, Thon G. Theoretical analysis of epigenetic cell memory by nucleosome modification. Cell. 2007;129:813–22.
Dykstra B, Olthof S, Schreuder J, Ritsema M, DE Haan G. Clonal analysis reveals multiple functional defects of aged murine hematopoietic stem cells. J Exp Med. 2011;208:2691–703.
Feldman N, Gerson A, Fang J, Li E, Zhang Y, Shinkai Y, Cedar H, Bergman Y. G9a-mediated irreversible epigenetic inactivation of Oct-3/4 during early embryogenesis. Nat Cell Biol. 2006;8:188–94.
Fujita N, Watanabe S, Ichimura T, Tsuruzoe S, Shinkai Y, Tachibana M, Chiba T, Nakao M. Methyl-CpG binding domain 1 (MBD1) interacts with the Suv39h1-HP1 heterochromatic complex for DNA methylation-based transcriptional repression. J Biol Chem. 2003;278:24132–8.
Glauche I, Moore K, Thielecke L, Horn K, Loeffler M, Roeder I. Stem cell proliferation and quiescence–two sides of the same coin. PLoS Comput Biol. 2009;5:e1000447.
Guo H, Zhu P, Wu X, Li X, Wen L, Tang F. Single-cell methylome landscapes of mouse embryonic stem cells and early embryos analyzed using reduced representation bisulfite sequencing. Genome Res. 2013;23:2126–35.
Haerter JO, Loevkvist C, Dodd IB, Sneppen K. Collaboration between CpG sites is needed for stable somatic inheritance of DNA methylation states. Nucleic Acids Res. 2014;42:2235–44.
Hinoue T, Weisenberger DJ, Lange CP, Shen H, Byun HM, van den Berg D, Malik S, Pan F, Noushmehr H, van Dijk CM, Tollenaar RA, Laird PW. Genome-scale analysis of aberrant DNA methylation in colorectal cancer. Genome Res. 2012;22:271–82.
Hoffman MM, Ernst J, Wilder SP, Kundaje A, Harris RS, Libbrecht M, Giardine B, Ellenbogen PM, Bilmes JA, Birney E, Hardison RC, Dunham I, Kellis M, Noble WS. Integrative annotation of chromatin elements from ENCODE data. Nucleic Acids Res. 2013;41:827–41.
Kundu S, Peterson CL. Role of chromatin states in transcriptional memory. Biochim Biophys Acta. 2009;1790:445–55.
Magee PN. The possible role of nucleic acid methylases in the induction of cancer. Cancer Res. 1971;31:599–604.
Margueron R, Reinberg D. Chromatin structure and the inheritance of epigenetic information. Nat Rev Genet. 2010;11:285–96.
Network CGA. Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012;487:330–7.
Okano M, Bell DW, Haber DA, Li E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell. 1999;99(3):247–57.
Ooi SKT, Qiu C, Bernstein E, Li K, Jia D, Yang Z, Erdjument-Bromage H, Tempst P, Lin S-P, Allis CD, Cheng X, Bestor TH. DNMT3L connects unmethylated lysine 4 of histone H3 to de novo methylation of DNA. Nature. 2007;448:714–7.
Otto SP, Walbot V. DNA methylation in eukaryotes: kinetics of demethylation and de novo methylation during the life cycle. Genetics. 1990;124:429–37.
Pfeifer GP, Steigerwald SD, Hansen RS, Gartler SM, Riggs AD. Polymerase chain reaction-aided genomic sequencing of an X chromosome-linked CpG island: methylation patterns suggest clonal inheritance, CpG site autonomy, and an explanation of activity state stability. Proc Natl Acad Sci U S A. 1990;87:8252–6.
Pradhan S, Bacolla A, Wells RD, Roberts RJ. Recombinant human DNA (cytosine-5) methyltransferase. I. Expression, purification, and comparison of de novo and maintenance methylation. J Biol Chem. 1999;274:33002–10.
Przybilla J, Buske P, Binder H, Galle J. Histone modifications control DNA methylation profiles during ageing and tumour expansion. Front Life Sci. 2013;7:31–43.
Przybilla J, Rohlf T, Loeffler M, Galle J. Understanding epigenetic changes in aging stem cells–a computational model approach. Aging Cell. 2014;13:320–8.
Rakyan VK, Down TA, Maslau S, Andrew T, Yang T-P, Beyan H, Whittaker P, Mccann OT, Finer S, Valdes AM, Leslie RD, Deloukas P, Spector TD. Human aging-associated DNA hypermethylation occurs preferentially at bivalent chromatin domains. Genome Res. 2010;20:434–9.
Reil T. Dynamics of gene expression in an artificial genome – implications for biological and artificial ontogeny. Adv Artif Life Proc. 1999;1674:457–66.
Rohlf T, Winkler CR. Emergent network structure, evolvable robustness, and nonlinear effects of point mutations in an artificial genome model. Adv Complex Syst. 2009;12:293–310.
Rohlf T, Steiner L, Przybilla J, Prohaska S, Binder H, Galle J. Modeling the dynamic epigenome: from histone modifications towards self-organizing chromatin. Epigenomics. 2012;4:205–19.
Rose NR, Klose RJ. Understanding the relationship between DNA methylation and histone lysine methylation. Biochim Biophys Acta. 2014;1839:1362–72.
Sedighi M, Sengupta AM. Epigenetic chromatin silencing: bistability and front propagation. Phys Biol. 2007;4:246.
Sheaffer KL, Kim R, Aoki R, Elliott EN, Schug J, Burger L, Schubeler D, Kaestner KH. DNA methylation is required for the control of stem cell differentiation in the small intestine. Genes Dev. 2014;28:652–64.
Sontag LB, Lorincz MC, Georg Luebeck E. Dynamics, stability and inheritance of somatic DNA methylation imprints. J Theor Biol. 2006;242:890–9.
Steiner L, Hopp L, Wirth H, Galle J, Binder H, Prohaska SJ, Rohlf T. A global genome segmentation method for exploration of epigenetic patterns. PLoS One. 2012;7:e46811.
Sturm D, Witt H, Hovestadt V, Al E. Hotspot mutations in H3F3A and IDH1 define distinct epigenetic and biological subgroups of glioblastoma. Cancer Cell. 2012;22:425–37.
Teschendorff AE, Menon U, Gentry-Maharaj A, Ramus SJ, Weisenberger DJ, Shen H, Campan M, Noushmehr H, Bell CG, Maxwell AP, Savage DA, Mueller-Holzner E, Marth C, Kocjan G, Gayther SA, Jones A, Beck S, Wagner W, Laird PW, Jacobs IJ, Widschwendter M. Age-dependent DNA methylation of genes that are suppressed in stem cells is a hallmark of cancer. Genome Res. 2010;20:440–6.
Thozmson JP, Skene PJ, Selfridge J, Clouaire T, Guy J, Webb S, Kerr AR, Deaton A, Andrews R, James KD, Turner DJ, Illingworth R, Bird A. CpG islands influence chromatin structure via the CpG-binding protein Cfp1. Nature. 2010;464:1082–6.
Tiwari VK, Mcgarvey KM, Licchesi JD, Ohm JE, Herman JG, Schubeler D, BAYLIN SB. PcG proteins, DNA methylation, and gene repression by chromatin looping. PLoS Biol. 2008;6:2911–27.
Vermeulen M, Mulder KW, Denissov S, Pijnappel WW, van Schaik FM, Varier RA, Baltissen MP, Stunnenberg HG, Mann M, Timmers HT. Selective anchoring of TFIID to nucleosomes by trimethylation of histone H3 lysine 4. Cell. 2007;131:58–69.
Verovskaya E, Broekhuis MJ, Zwart E, Ritsema M, van Os R, de Haan G, Bystrykh LV. Heterogeneity of young and aged murine hematopoietic stem cells revealed by quantitative clonal analysis using cellular barcoding. Blood. 2013;211:487–97.
Wu AR, Neff NF, Kalisky T, Dalerba P, Treutlein B, Rothenberg ME, Mburu FM, Mantalas GL, Sim S, Clarke MF, Quake SR. Quantitative assessment of single-cell RNA-sequencing methods. Nat Methods. 2014;11:41–6.
Wutz A. Epigenetic regulation of stem cells : the role of chromatin in cell differentiation. Adv Exp Med Biol. 2013;786:307–28.
Xu M, Long C, Chen X, Huang C, Chen S, Zhu B. Partitioning of histone H3-H4 tetramers during DNA replication-dependent chromatin assembly. Science. 2010;328:94–8.
Acknowledgements
This work was supported by the Federal Ministry of Education and Research (BMBF): HNPCC-Sys (grant No. 031 6065), MAGE (grant No. 50500541) and INDRA (grant No. 031A312). We thank Peter Buske for computing data sets used in Fig. 7.4b.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media Dordrecht
About this chapter
Cite this chapter
Przybilla, J., Rohlf, T., Galle, J. (2015). Towards a Mechanistic Understanding of Epigenetic Dynamics. In: Teschendorff, A. (eds) Computational and Statistical Epigenomics. Translational Bioinformatics, vol 7. Springer, Dordrecht. https://doi.org/10.1007/978-94-017-9927-0_7
Download citation
DOI: https://doi.org/10.1007/978-94-017-9927-0_7
Publisher Name: Springer, Dordrecht
Print ISBN: 978-94-017-9926-3
Online ISBN: 978-94-017-9927-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)