Abstract
ABA, a sesquiterpene, is a small molecule that has a non-planar shape and contains various functional groups. The C15 ABA-skeleton is common in biosynthetic precursors (xanthoxin, abscisic alcohol, and abscisic aldehyde) and oxidised catabolites (8′-hydroxy-ABA, phaseic acid, and dihydrophaseic acid). Thus, the conformation of the molecule and its functional groups are the key factors governing whether these ABA derivatives function as ABA mimics that can activate ABA receptors. Recent reports of the crystal structures of the ABA receptor proteins, PYR/PYL/RCAR (PYL), give significant clues regarding the structural requirements for eliciting ABA responses. The findings from structural studies of PYL are generally consistent with those from structure–activity studies based on bioassays using ABA analogues. The structural requirements for biosynthetic and catabolic enzymes and for ABA transporters remain unknown, as these structures have not yet been solved. However, ABA 8′-hydroxylases have been well investigated using in vitro enzyme assays. These studies show that different ABA-binding proteins have somewhat different structural requirements. Based on this knowledge, we can design a selective ligand that targets a specific ABA-binding protein.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1.1 Structures, Physicochemical Properties, and Physiological Functions
1.1.1 ABA
ABA(1) has a chiral centre (C-1′). Consequently, two enantiomers, S-(+)-ABA and R-(−)-ABA (Fig. 1.1), are formed during chemical synthesis. All naturally occurring ABAs have the S-configuration, and its R counterpart has never been detected in plants. The physicochemical properties of both isomers of ABA are identical except for their optical characteristics, such as optical rotation and circular dichroism. ABA is sufficiently heat-stable to be dissolved in boiling water without decomposition occurring. Although ABA is relatively stable towards a broad range of pH values, it is converted to γ-lactone (2) under strong acidic conditions such as formic acid–hydrochloric acid (Mallaby and Ryback 1972) (Fig. 1.2). Under alkaline conditions, the hydrogens at C-3′, -5′, -7′, -2, and -6 exchange with deuterium when dissolved in deuterated water (Milborrow 1984; Willows et al. 1991; Gray et al. 1974) (Fig. 1.2). The six hydrogens at C-3′, -5′, and -7′ are particularly easily exchanged with deuterium, making ABA-d 6 (3) a useful internal standard for LC–MS and GC–MS analysis.
ABA is moderately photosensitive due to the dienoic acid side chain and the ring enone. Irradiation with UV at 365 nm causes photoisomerization at the C-2 double bond to give an equimolar mixture of ABA and 2E-ABA (2-trans-ABA, 5) (Fig. 1.2), which is biologically inactive (Todoroki et al. 2001). This isomerization occurs even upon irradiation using fluorescent and tungsten lamps. UV light at wavelengths shorter than 305 nm decomposes the ABA molecule into unidentified compounds, in addition to causing photoisomerization (Cornelussen et al. 1995). This photolabile property of ABA limits its applications in agriculture. Consequently, several ABA analogues have been designed and synthesised to increase the photostability of ABA. For example, compounds 6–8 have a rigid side chain with an aromatic ring (Chen and Mctaggart 1986; Kim et al. 1992; Asami and Yoshida 1999) (Fig. 1.3). Recently, Wenjian et al. (2013) reported that 2,3-cyclopropanated ABA (9) is more photostable than ABA (Fig. 1.3).
ABA behaves as a weak acid due to the side chain carboxy group. Thus, the lipophilicity of ABA depends greatly on pH and is more lipophilic at lower pH. Because the undissociated form of ABA is cell-membrane-permeable, the in vivo distribution of ABA changes depending on the pH conditions (Wilkinson et al. 1998; Jiang and Hartung 2008). This means that the movement of ABA within plants does not always require specific transport proteins; nevertheless, some ABA transporters seem to be involved in physiological processes mediated by ABA (Kang et al. 2010; Kuromori et al. 2010; Seo and Koshiba 2011; Kanno et al. 2012).
The shape of the ABA molecule depends largely on the conformation of the cyclohexenone ring. The crystal structure of ABA shows that the ring adopts a slightly distorted sofa conformation with the pseudoaxial side chain (Ueda and Tanaka 1977; Schmalle et al. 1977) (Fig. 1.4). The preferred form in solution, revealed by NMR and CD analyses, is a half-chair with the pseudoaxial side chain (Milborrow Milborrow 1984; Willows and Milborrow 1993; Koreeda et al. 1973; Harada 1973). Theoretical conformational analyses revealed that the minimum-energy conformer is a half-chair with the pseudoaxial side chain (Todoroki et al. 1996) (Fig. 1.4). Because the cyclohexenone ring can invert into another half-chair with the pseudoequatorial side chain by paying only a small energy penalty (∼1.4 kcal mol−1) (Todoroki and Hirai 2000a, b) (Fig. 1.4), the shape of the ABA molecule when bound to proteins such as receptors, transporters, and catabolic enzymes cannot be estimated based on the preferred form of the uncomplexed ABA molecule. Todoroki et al. (1996) demonstrated that the biologically active conformation of ABA is a half-chair with the pseudoaxial side chain, based on the biological activity of cyclopropane analogues of ABA. Furthermore, the crystal structures of the ABA receptor proteins PYR/PYL/RCAR (PYL) bound to ABA revealed that ABA adopts a half-chair with the pseudoaxial side chain in the binding pocket (Melcher et al. 2009; Miyazono et al. 2009; Nishimura et al. 2009; Santiago et al. 2009; Yin et al. 2009) (Fig. 1.4). However, this does not guarantee that the conformation of ABA in other binding proteins, including transporters and catabolic enzymes, is also a half-chair with the pseudoaxial side chain.
Many reports have demonstrated that both the S- and R-isomers of ABA show similar hormonal activities in several assay systems (Lin et al. 2005), suggesting that plants have a mechanism which permits the R-isomer, which is not naturally occurring, to mimic the endogenous hormone. The ABA molecule, although a chiral molecule, is relatively symmetric about the plane constructed by C-5, C-1′, and C-4′ (Fig. 1.5). When the structural formula of R-ABA is drawn on paper, the orientation of the side chain and the hydroxy group at C-1′ of S-ABA is usually exchanged. However, this method of drawing does not accurately represent the pseudo-symmetric property of the ABA molecule. Instead, the molecule should be flipped to have the side chain and hydroxy group in the same orientation, as in the case of S-ABA. By doing so, a large structural difference between the enantiomers becomes evident at the α-oriented, axial methyl group (C-8′). This means that the enantiomeric recognition of ABA by ABA-binding proteins depends on their sensitivity to the location of this methyl group (Fig. 1.5). The crystal structure of the PYL3-R-ABA complex shows that R-ABA is oriented in the pocket in a manner similar to S-ABA (Zhang et al. 2013), indicating that the ability to accommodate C-8′ provides PYL with a relatively high affinity for R-ABA. However, although some PYL proteins have measurable affinities for R-ABA, generally this affinity is weaker than that for S-ABA (Park et al. 2009; Zhang et al. 2013). Of the Arabidopsis PYL proteins, PYL5 shows the strongest binding affinity for R-ABA, whereas PYL9 does not function in the presence of R-ABA (Zhang et al. 2013). On the other hand, recombinant ABA 8′-hydroxylases (CYP707A enzymes), which oxidise the C-8′ of ABA, do not bind R-ABA (Kushiro et al. 2004). This is as expected, since CYP707A must recognise C-8′ in order to conduct oxidation, so stereospecific ligand binding is critical for this enzyme. Interestingly, C-8′ recognition by CYP707A seems to depend on the ring enone of ABA (Ueno et al. 2007). The difference in sensitivity to the orientation of C-8′ between CYP707A and PYL proteins may explain the comparable biological activity of R-ABA and S-ABA in some bioassays. R-ABA is metabolized to (−)-7′-hydroxy-ABA (21) (Cowan and Railton 1987; Boyer and Zeevaart 1986; Balsevich et al. 1994a) and (+)-PA (23) (Gillard and Walton 1976; Balsevich et al. 1994b; Dashek et al. 1979; Okamoto and Nakazawa 1993) (Fig. 1.6). Since the recombinant CYP707A enzymes do not bind R-ABA (Kushiro et al. 2004), these metabolites may be produced by the nonspecific oxidation of R-ABA.
1.1.2 Biosynthetic Precursors
The biosynthetic precursors that have an ABA-skeleton are xanthoxin (10), abscisic alcohol (ABAlc, 16), and abscisic aldehyde (ABAld, 11) (Fig. 1.6). The side chain of xanthoxin cannot be axial owing to the 1′,2′-epoxide, so the conformation of xanthoxin must be quite different from that of ABAlc, ABAld, and ABA. Crystal structures of the PYL–ABA complex suggest that these precursors, which have no carboxylic acid moiety, generally have poor affinity to PYL proteins because the salt bridge between the C-1 carboxylate of ABA and the ε-ammonium group of Lys of PYL is critical for forming a stable PYL–ABA complex. In fact, these precursors induce a slight reduction in PP2C activity in the presence of PYL proteins (Kepka et al. 2011).
1.1.3 Catabolites
Common catabolites of ABA in many plants include phaseic acid (PA, 13), dihydrophaseic acid (DPA, 14), epi-dihydrophaseic acid (epi-DPA, 15), 7′-hydroxy-ABA (17), and ABA glucose ester (ABA-GE, 18) (Ref) (Fig. 1.6). 9′-Hydroxy-ABA (19) and neoPA (20) are found in some plants, including Arabidopsis (Zhou et al. 2004; Okamoto et al. 2011). 8′-Hydroxy-ABA (12), which is produced by the hydroxylation of C-8′ by CYP707A enzymes, is thermodynamically unstable and spontaneously isomerizes to the more stable tautomer, PA. Thus, 8′-hydroxy-ABA is not stably maintained in the absence of 8′-O-protection following isolation. Zou et al. (1995) isolated 8′-hydroxy-ABA as a borate complex by heating PA and boric acid in glacial acetic acid. The isomerization of 8′-hydroxy-ABA to PA is an intramolecular Michael-type reaction which is accelerated under basic conditions. At 25 °C, the half-life of 8′-hydroxy-ABA is 30 h at pH 3, 4 h at pH 7, and shorter than 1 min at pH 10 (Todoroki and Hirai 2000a, b). 8′-Hydroxy-ABA is observed upon HPLC analysis of enzyme assay mixtures using recombinant CYP707A (Saito et al. 2004). This suggests that CYP707A catalyses only the hydroxylation reaction, and not the isomerization reaction to PA.
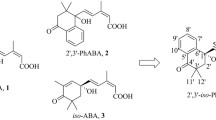
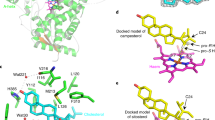
Is 8′-hydroxy-ABA biologically active? It is difficult to answer this question because its thermodynamic instability prevents us from determining its true activity. Zou et al. (1995) reported that a borate complex of 8′-hydroxy-ABA is active in lipid and oleosin biosynthesis in microspore-derived embryos of B. napus. Kepka et al. (2011) extrapolated the activity of 8′-hydroxy-ABA from that of the tetralone analogue of 8′-hydroxy-ABA (24). This analogue cannot isomerize to a PA-type derivative because the C-2′ is in an aromatic ring (Fig. 1.7). Tetralone-ABA is as effective as ABA in some bioassays using Arabidopsis, whereas the 8′-hydroxylated analogue is much less effective than tetralone-ABA. In PP2C assays using the Arabidopsis ABA receptors, PYL9 and PYR1, 8′-hydroxylated tetralone-ABA is less effective than tetralone-ABA and ABA by a factor of 100–1000. This suggests that 8′-hydroxy-ABA is essentially inactive, at least in Arabidopsis. Because some space is evident around the C-8′ of ABA in both the PYR1-ABA and PYL9-ABA complexes (PDB code: 3K90 and 3OQU, respectively), the 8′-hydroxy group is accommodated sterically. Furthermore, since ABA stabilizes the gate-closed conformation of PYL proteins by producing a hydrophobic network, a protic polar moiety around C-8′ may impair the critical function of the proteins. In any case, currently there is no experimental evidence to help resolve the discrepancy in ABA activity between Brassica napus and Arabidopsis.
PA is a more rigid molecule than ABA. The conformation of the six-membered ring of PA is a rigid chair with an axial side chain. This conformation is similar to that of ABA bound to PYL proteins; however, PA has much less affinity with PYL compared to ABA (Kepka et al. 2011). PA has an α-axial substituent at C-2′, like the substituent at C-6′ of R-ABA. However, the C-2′ substituent of PA is not a methyl group, but an ether oxygen. This is a significant disadvantage for interacting with PYL because the ether oxygen reduces the affinity of PA with PYL due to steric hindrance and hydrophilicity. In fact, PA is inactive in most bioassays. Although rarely mentioned, solutions of PA inevitably contain a trace amount of 8′-hydroxy-ABA; the equilibrium ratio of 8′-hydroxy-ABA/PA is 2:98 at pH 3–10 (Todoroki and Hirai 2000a, b). Consequently, a peak and signals due to 8′-hydroxy-ABA are evident in the HPLC and 1H NMR analyses of PA, respectively.
1.1.4 ABA Mimics Lacking an ABA-skeleton
Although ABA is produced by fermentation using ABA-producing fungi at relatively low cost, ABA mimics with a simpler structure than ABA would be practical and advantageous if their production costs were lower than that of ABA (Fig. 1.8). In early studies, lunularic acid (25) was considered to play an ABA-like physiological role in liverworts (Pryce 1972), although currently this hypothesis is in question because ABA has been detected in two species of liverworts (Nakayama et al. 1993). Some synthetic ABA mimics containing an aromatic ring instead of the cyclohexenone ring of ABA (26–28) have been reported, although their activities are lower than that of ABA (Bittner et al. 1977; Ladyman et al. 1988; Yoshikawa et al. 1992). The Lewis formulae of these compounds somewhat resemble ABA. However, these molecules are relatively flat, so they bear little resemblance to ABA sterically.
Pyrabactin (29), a key compound for identifying ABA receptors amongst PYL proteins, exhibits weak ABA-like activity (Park et al. 2009). Although this compound does not superficially resemble ABA, the bent conformation functions as an effective gate-closing promoter in PYL1 (Melcher et al. 2010) (Fig. 1.8). Interestingly, pyrabactin binds to PYL2 in a manner different from its binding to PYL1 and does not induce closure of the gate. The ABA agonistic effect of pyrabactin is observed in in vitro experiments only for PYR1 and PYL1 (Okamoto et al. 2013). Quinabactin (30), which contains aromatic rings and a sulfonamide similar to pyrabactin, is a stronger ABA mimic than is pyrabactin (Okamoto et al. 2013). In vitro analysis revealed that quinabactin activates mainly dimeric receptors to elicit stomatal closure, ABA-regulated gene expression, and drought tolerance. Quinabactin also adopts a bent conformation in PYL2 (Fig. 1.8). The dimeric receptor-selective binding of quinabactin may depend on its slightly larger molecular size compared to ABA.
1.2 Structural Requirements
1.2.1 For Exhibiting Biological Activity
The biological activities of many ABA analogues have been examined using various plants and tissues under various conditions (Fig. 1.9). Although there are some exceptions, the absence of an existing functional group, or the addition of a new functional group, generally reduces the ABA activity. The most critical functional group is the C-1 carboxy group, although ABA analogues that have a C-1 hydroxy (ABAlc), aldehyde (ABAld), or methyl ester (Me ABA, 31) are relatively active under some assay conditions. For example, the aba3 mutant, which is deficient in molybdenum cofactor (MoCo) sulfurase necessary for ABAld oxidase (AAO3), is insensitive to ABAlc and ABAld (Kepka et al. 2011). The relatively high activity of these ABA precursors results from ABA being generated from them in plants. Since plant cells generally exhibit high esterase activity and rapidly hydrolyse esters (Kusaka et al. 2009), the activity of Me ABA must also result from its conversion to ABA by enzymatic hydrolysis.
The other two polar groups in ABA are also important for ABA activity, although they are less crucial than the C-1 carboxy group: The reported activities of 1′-deoxy-ABA (32), 1′-fluoro-ABA (33) (Todoroki et al. 1995a, b, c), 1′-O-methyl-ABA (34) (Rose et al. 1996), 4′-deoxo-ABA (35) (Oritani and Yamashita 1974), 1′,4′-cis/trans-diol-ABAs (36 and 37) (Walton and Sondheimer 1972), and trans-4′-alcoxy-ABA (38–40) (Asami et al. 2000; 2002) are weaker than that of ABA by a factor of 10–100. The cis isomer of 4′-alcoxy-ABA is much less potent than the trans isomer. Interestingly, some ABA analogues containing a larger functional group than ABA’s functional group at C-4′ (41–44) (Fig. 1.10) retain ABA activity, albeit weaker than ABA’s by a factor of 10–100 (Kohler et al. 1997; Asami et al. 1997; Kitahata et al. 2005). Because these compounds are substituted using a hydrozone linker at C-4′, it is possible that ABA is released by hydrolysis of the analogue in plant cells. On the other hand, the activity of 4′-octoxy-ABA (39) and 4′-benzyl-ABA (40) (Asami et al. 2002) cannot be derived from released ABA and must be due to the analogue, since an ether bond is more resistant to hydrolysis than a hydrazone. The significance of this will be discussed later.
The two methyl groups at C-3 in the side chain and at C-2′ in the ring (C-6 and C-7′, respectively) are more important for activity than the geminal methyl groups at C-6′ in the ring (C-8′ and C-9′). The C-6 and C-7′ methyl groups may contribute to specific recognition rather than to nonspecific hydrophobic interactions. The modification of C-8′ or C-9′ sometimes results in an increase in activity. In particular, ABA analogues, in which C-8′ is replaced by a trifluoromethyl (45) or an acetylenyl (46), and where C-9′ is replaced by a propagyl (47), are the most potent analogues tested to date in bioassays (Todoroki et al. 1995a, b, c; Cutler et al. 2000) (Fig. 1.11). 5′α,8′-Cyclo-ABA (48), which contains a direct linkage between C-5′ and C-8′, is also remarkably effective in some bioassays (Todoroki et al. 1996). The amplified activity of these analogues may depend partially on their long lifetime in plant cells, since modification of C-8′ or its neighbouring groups makes these compounds resistant to CYP707A enzymes (Cutler et al. 2000). Nevertheless, since recombinant Arabidopsis ABA 8′-hydroxylase CYP707A3 is not inhibited by the 8′-acetylenic analogue of ABA (Ueno et al. 2005), another factor may be involved. 2E-ABA (5) is also inactive, probably because the 2E-configuration greatly affects the orientation of the C-1 carboxy group. The C-2′ double bond in the ring can be changed to a single bond with only a slight loss of activity if C-7′ is cis to the side chain (49), that is, when the ring prefers a half-chair conformation with the pseudoaxial side chain (Fig. 1.11). This agrees with the finding that the ring conformation is a significant factor for activity. As described above, the active conformation of ABA is a half-chair with the pseudoaxial side chain.
1.2.2 For Binding to PYL Proteins
The affinity of ABA analogues to PYL proteins has rarely been measured directly or indirectly. We can only speculate on their affinity on the basis of the crystal structure of the PYL–ABA complex, where ABA is almost entirely surrounded by the PYL protein. This binding mode suggests that the addition of a large substituent anywhere in the structure reduces the affinity because of interference with gate-closing or PP2C binding. This is generally consistent with the structural requirements for ABA activity in bioassays, discussed above. Nevertheless, some exceptions exist. The most significant inconsistency is observed in C-4′. As described above, 4′-octyl-ABA and 4′-benzyl-ABA (39 and 40) retain ABA activity. In the crystal structure of the PYL–ABA–PP2C complex, the C-4′ ketone of ABA forms an indirect hydrogen bond via a water molecule with the indole moiety of Trp in PP2C. In addition, the gate Pro in PYL is close to the C-4′ ketone. Thus, the introduction of a large substituent at C-4′ must impair the function of PYL as an inhibitor of PP2C. n-Octoxy and benzyloxy groups, with lengths of ∼10 and 5 Å, respectively, seem to be too large to fit into the space around C-4′ of ABA in the PYL–ABA–PP2C complex. In this complex, the distance between the 4′-carbonyl oxygen of ABA and the indole nitrogen of Trp in PP2C is calculated to be approximately 4.5–5 Å, based on the crystal structures (Fig. 1.12).
Recently, Takeuchi et al. (2014) reported an ABA analogue, AS6, that acts as an inhibitor of PYL proteins. This molecule has an S-hexyl group at C-3′ and is an effective gate-closing promoter of PYL. Crystallographic data of PYR1 bound to AS6 demonstrate that the long S-hexyl chain of AS6 protrudes through a solvent-exposed tunnel to prevent PYL–PP2C interactions.
1.2.3 For Inhibition of CYP707A Enzymes
ABA 8′-hydroxylases, which are CYP707A enzymes, are not inhibited by R-ABA (Kushiro et al. 2004). This strict stereoselective recognition is in marked contrast to the loose recognition requirements of PYL proteins. Ueno et al. (2007) revealed that the C-1 carboxy and C-4′ carbonyl groups play a significant role in stereoselective binding. Based on the hypothesis that ABA is anchored via these two polar groups on the helix I residues in the substrate-binding pocket of CYP707A, the C-2′ side of the ring must be close to helix I (SRS 4) to keep C-8′ in close proximity to the heme iron. If R-ABA binds in the same manner as S-ABA, C-8′ should contact helix I (Fig. 1.13). Determination of the exact mechanism requires the crystal structure of CYP707A, which has yet to be obtained. Another large difference is that the side chain methyl (C-6) and C-1′ hydroxy groups which are required for the high ABA activity is dispensable for binding to CYP707A enzymes (Ueno et al. 2005) (Fig. 1.14). Focusing on this difference, Ueno et al. (2005) developed AHI1 (50), an ABA analogue which functions as a competitive inhibitor of CYP707A without causing an ABA response.
References
Asami T, Yoshida S. Abscisic acid analogues possessing hetero five-membered ring: design, synthesis and activity. RIKEN Rev. 1999;21:17–9.
Asami T, Lin S, Yamamoto M, Robertson M, Min YK, Murofushi N, Yoshida S. Fluorescence-labeled abscisic acid possessing abscisic acid-like activity in barley aleurone protoplasts. Biosci Biotechnol Biochem. 1997;61:1198–9.
Asami T, Min YK, Han SY, Kitahata N, Oh K, Murofushi N, Yoshida S. 4′-Alkoxy derivatives of abscisic acid: a promising new probe for mechanisms of abscisic acid perceptions. Plant Growth Regul. 2002;38:237–41.
Asami T, Min YK, Matsuyama T, Murofushi N, Yamaguchi I, Yoshida S. Synthesis and biological activity of 4′-methoxy derivatives of abscisic acid. Bioorg Med Chem Lett. 2000;10:1571–4.
Balsevich J, Abrams SR, Lamb N, Konig W. Identification of unnatural phaseic acid as a metabolite derived from exogenouslyadded (−)-abscisic acid in a maize cell suspension culture. Phytochemistry. 1994a;36:647–50.
Balsevich J, Cutler A, Lamb N, Friesen L, Kurz E, Perras M, Abrams SR. Response of cultured maize cells to (+)-abscisic acid, (−)-abscisic acid, and their metabolites. PlantPhysiol. 1994b;106:135–42.
Bittner S, Gorodetsky M, Har-Paz I, MIzrahi Y, Richmond AE (1977). Synthesis and biological effect of aromatic analogues of abscisic acid. Phytochemistry 16: 1143–51.
Boyer GL, Zeevaart JAD. 7′-Hydroxy (−)-R-abscisic acid: ametabolite of feeding (−)-R-abscisic acid to Xanthium strumarium. Phytochemistry. 1986;25:1103–5.
Chen SC, Mctaggart JM. Abscisic acid analogueues with a geometrically rigid conjugated acid side-chain. Agric Biol Chem. 1986;50:1097–100.
Cornelussen MHM, Karssen CM, van Loon LC. UV-induced cross-linking of abscisic acid to binding proteins. Phytochemistry. 1995;39:959–68.
Cowan AK, Railton ID. The catabolism of (±)-abscisic acid by excised leaves of Hordeum vulgare L. cv Dyan and its modification by chemical and environmental factors. Plant Physiol. 1987;84:157–63.
Cutler AJ, Rose PA, Squires TM, Loewen MK, Shaw AC, Quail JW, Krochko JE, Abrams SR. Inhibitors of abscisic acid 8′-hydroxylase. Biochemistry. 2000;39:13614–24.
Dashek WV, Singh BN, Walton DC. Abscisic acid localization and metabolism in barley aleurone layers. Plant Physiol. 1979;64:43–8.
Gillard DF, Walton DC. Abscisic acid metabolism by a cell-free preparation from Echinocystis lobata liquid endoserum. Plant Physiol. 1976;58:790–5.
Gray RT, Mallaby R, Ryback G, Williams VP. Mass spectra of methyl abscisate and isotopically labelled analogueues. J Chem Soc Perkin Trans II. 1974;1974:919–24.
Harada N. Absolute configuration of (+)-trans-abscisic acid as determined by a quantitative application of the exciton chirality method. J Am Chem Soc. 1973;95:240–2.
Jiang F, Hartung W. Long-distance signalling of abscisic acid (ABA): the factors regulating the intensity of the ABA signal. J Exp Bot. 2008;59:37–43.
Kang J, Hwang J, Lee M, Kim Y, Assmann SM, Martinoia E, Lee Y. PDR-type ABC transporter mediates cellular uptake of the phytohormone abscisic acid. Proc Natl Acad Sci USA. 2010;107:2355–60.
Kanno Y, Hanada A, Chiba Y, Ichikawa T, Nakazawa M, Matsui M, Koshiba T, Kamiya Y, Seo M (2012). Identification of an abscisic acid transporter by functional screening using the receptor complex as a sensor. Proc Natl Acad Sci USA 109: 9653–8.
Kepka M, Benson CL, Gonugunta VK, Nelson KM, Christmann A, Grill E, Abrams SR. Action of natural abscisic acid precursors and catabolites on abscisic acid receptor complexes. Plant Physiol. 2011;157:2108–19.
Kim BT, Asami T, Morita K, Soh CH, Murofushi N, Yoshida S. Synthesis of new abscisic acid (ABA) analogues possessing a geometrically rigid cyclized side chain. Biosci Biotechnol Biochem. 1992;56:624–9.
Kitahata N, Nakano T, Kuchitsu K, Yoshida S, Asami T. Biotin-labeled abscisic acid as a probe for investigating abscisic acid binding sites on plasma membranes of barley aleurone protoplasts. Bioorg Med Chem. 2005;13:3351–8.
Kohler AD, Beale MH, Rollason R, Barratt DHP, Lewis MJ, Van der Meulen RM, Wang M. A radio-iodinated abscisic acid photoaffinity probe. J Chem Soc Perkin Trans. 1997;1:1543–7.
Koreeda M, Weiss G, Nakanishi K. The absolute configuration of natural (+)-abscisic acid. J Am Chem Soc. 1973;95:239–40.
Kuromori T, Miyaji T, Yabuuchi H, Sugimoto E, Kamiya A, Moriyama Y, Shinozaki K. ABC transporter AtABCG25 is involved in abscisic acid transport and responses. Proc Natl Acad Sci USA. 2010;107:2361–6.
Kusaka N, Maisch J, Nick P, Hayashi KI, Nozaki H. Manipulation of intercellular auxin in a single cell by light with esterase-resistant caged auxins. ChemBioChem. 2009;10:2195–202.
Kushiro T, Okamoto M, Nakabayashi K, Yamagishi K, Kitamura S, Asami T, Hirai N, Koshiba T, Kamiya Y, Nambara E. The Arabidopsis cytochrome P450 CYP707A encodes ABA 8′-hydroxylases: key enzymes in ABA catabolism. EMBO J. 2004;23:1647–56.
Ladyman JA, Sanborn JR, Eelkema EE. Biological activity of an aromatic analogueue of abscisic acid. Phytochemistry. 1988;27:3751–5.
Lin BL, Wang HJ, Wang JS, Zaharia LI, Abrams SR. Abscisic acid regulation of heterophylly in Marsilea quadrifolia L.: effects of R-(−) and S-(+) isomers. Plant Physiol. 2005;106:135–42.
Mallaby R, Ryback G. Chemistry of a color test for abscisic acid. J Chem Soc. 1972;8:919–21.
Melcher K, Ng LM, Zhou XE, Soon F-F, Xu Y, Suino-Powell KM, Park S-Y, Weiner JJ, Fujii H, Chinnusamy V, Kovach A, Li J, Wang Y, Li J, Peterson FC, Jensen DR, Yong E-L, Volkman BF, Cutler SR, Zhu J-K, Xu HE. A gate–latch-lock mechanism for hormone signalling by abscisic acid receptors. Nature. 2009;462:602–8.
Melcher K, Xu Y, Ng LM, Zhou XE, Soon FF, Chinnusamy V, Suino-Powell KM, Kovach A, Tham FS, Cutler SR, Li J, Yong EL, Zhu JK, Xu HE. Identification and mechanism of ABA receptor antagonism. Nat Struct Mol Biol. 2010;17:1102–8.
Milborrow BV. The conformation of abscisic acid by N.M.R. and a revision of the proposed mechanism for cyclization during its biosynthesis. Biochem J. 1984;220:325–32.
Miyazono K, Miyakawa T, Sawano Y, Kubota K, Kang HJ, Asano A, Miyauchi Y, Takahashi M, Zhi Y, Fujita Y, Yoshida T, Kodaira KS, Yamaguchi-Shinozaki K, Tanokura M. Structural basis of abscisic acid signalling. Nature. 2009;462:609–14.
Nakayama M, Takase T, Yokota T. Abstract of the XV international botanical congress. Yokohama: Japan; 1993. p. 388.
Nishimura N, Hitomi K, Arvai AS, Rambo RP, Hitomi C, Cutler SR, Schroeder JI, Getzoff ED. Structural mechanism of abscisic acid binding and signaling by dimeric PYR1. Science. 2009;326:1373–9.
Okamoto M, Nakazawa H. Stereoselectivity of abscisic acid oxygenase in avocado fruits. Biosci Biotechnol Biochem. 1993;57:1768–9.
Okamoto M, Kushiro T, Jikumaru Y, Abrams SR, Kamiya Y, Seki M, Nambara E. ABA 9′-hydroxylation is catalyzed by CYP707A in Arabidopsis. Phytochemistry. 2011;72:717–22.
Okamoto M, Peterson FC, Defries A, Park SY, Endo A, Nambara E, Volkman BF, Cutler SR. Activation of dimeric ABA receptors elicits guard cell closure, ABA-regulated gene expression, and drought tolerance. Proc Natl Acad Sci USA. 2013;110:12132–7.
Oritani T, Yamashita K. Syntheses and biological activities of analogues of abscisic acid. Agric Bioi Chem. 1974;38:801–8.
Park SY, Fung P, Nishimura N, Jensen DR, Fujii H, Zhao Y, Lumba S, Santiago J, Rodrigues A, Chow TF, Alfred SE, Bonetta D, Finkelstein R, Provart NJ, Desveaux D, Rodriguez PL, McCourt P, Zhu JK, Schroeder JI, Volkman BF, Cutler SR. Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science. 2009;324:1068–71.
Pryce RJ. The occurrence of lunularic and abscisic acids in plants. Phytochemistry. 1972;11:1759–61.
Rose PA, Lei B, Shaw AC, Barton DL, Walker-Simmons MK, Abrams SR. Probing the role of the hydroxyl group of ABA: analogues with methyl ether at C-1′. Phytochemistry. 1996;41:1251–8.
Saito S, Hirai N, Matsumoto C, Ohigashi H, Ohta D, Sakata K, Mizutani M. Arabidopsis CYP707As encode (+)-abscisic acid 8′-hydroxylase, a key enzyme in the oxidative catabolism of abscisic acid. Plant Physiol. 2004;134:1439–49.
Santiago J, Dupeux F, Round A, Antoni R, Park SY, Jamin M, Cutler SR, Rodriguez PL, Marquez JA. The abscisic acid receptor PYR1 in complex with abscisic acid. Nature. 2009;462:665–8.
Schmalle VHW, Klaska KH, Jarchow O. Die Kristall- und Molekülstruktur von (R, S)-cis, trans-Abscisinsäure:5-(1-Hydroxy-2,6,6-trimethyl-4-oxo-2-cyclohexen-1-yl)-3-methyl-2,4-pentadiensäure. Acta Cryst. 1977;B33:2218–24.
Seo M, Koshiba T. Transport of ABA from the site of biosynthesis to the site of action. J Plant Res. 2011;124:501–7.
Takeuchi J, Okamoto M, Akiyama T, Muto T, Yajima S, Sue M, Seo M, Kanno Y, Kamo T, Endo A, Nambara E, Hirai N, Ohnishi T, Cutler SR, Todoroki Y. Designed abscisic acid analogues as antagonists of PYL-PP2C receptor interactions. Nat Chem Biol. 2014;10:477–82.
Todoroki Y, Hirai N. Analysis of isomerization process of 8′-hydroxyabscisic acid and its 3′-fluorinated analogue in aqueous solutions. Tetrahedron. 2000a;56:1649–53.
Todoroki Y, Hirai N. Conformational analysis of the cyclohexenone ring in abscisic acid and its analogues with a fused cyclopropyl ring. Tetrahedron. 2000b;56:8095–100.
Todoroki Y, Hirai N, Koshimizu K. 8′,8′,8′-Trifluoroabscisic acids as highly potent, long-lasting analogueues of abscisic acid. Phytochemistry. 1995a;38:561–8.
Todoroki Y, Hirai N, Koshimizu K. Synthesis and biological activity of 1′-deoxy-1′-fluoro- and 8′-fluoroabscisic acids. Phytochemistry. 1995b;40:633–41.
Todoroki Y, Nakano S, Hirai N, Ohigashi H. Synthesis, biological activity and metabolism of (S)-(+)-3′-Fluoroabscisic acid. Tetrahedron. 1995c;51:6911–26.
Todoroki Y, Nakano S, Hirai N, Ohigashi H. Ring conformational requirement for biological activity of abscisic acid probed by the cyclopropane analogueues. Tetrahedron. 1996;52:8081–98.
Todoroki Y, Tanaka T, Kisamori M, Hirai N. 3′-Azidoabscisic acid as a photoaffinity reagent for abscisic acid binding proteins. Bioorg Med Chem Lett. 2001;11:2381–4.
Ueda H, Tanaka J. The crystal and molecular structures of dl-2-cis-4-trans-abscisic acid. Bull Chem Soc Jpn. 1977;50:1506–9.
Ueno K, Araki Y, Hirai N, Saito S, Mizutani M, Sakata K, Todoroki Y. Differences between the structural requirements for ABA 8′-hydroxylase inhibition and for ABA activity. Bioorg Med Chem. 2007;13:3359–70.
Ueno K, Araki Y, Hirai N, Saito S, Mizutani M, Sakata K, Todoroki Y. Differences between the structural requirements for ABA 8′-hydroxylase inhibition and for ABA activity. Bioorg Med Chem. 2005;13:3359–70.
Walton DC, Sondheimer E. Activity and metabolism of C-(±)-abscisic acid derivatives. Plant Physiol. 1972;49:290–2.
Wenjian L, Xiaoqiang H, Yumei X Jinlong F, Yuanzhi Z, Huizhe L, Mingan W, Zhaohai Q (2013). Synthesis, photostability and bioactivity of 2,3-cyclopropanated abscisic acid. Phytochemistry. 2013;96:72–80.
Wilkinson S, Corlett JE, Oger L, Davies WJ. Effects of xylem pH on transpiration from wild-type and flacca tomato leaves. Plant Physiol. 1998;117:703–9.
Willows RD, Milborrow BV. Configurations and conformations of abscisic acid. Phytochemistry. 1993;34:233–7.
Willows RD, Netting AG, Milborrow BV. Synthesis of stably deuteriated abscisic acid, phaseic acid and related compounds. Phytochemistry. 1991;30:1483–5.
Yin P, Fan H, Hao Q, Yuan X, Wu D, Pang Y, Yan C, Li W, Wang J, Yan N. Structural insights into the mechanism of abscisic acid signaling by PYL proteins. Nat Struct Mol Biol. 2009;16:1230–6.
Yoshikawa H, Fujimoto E, Doi K (1992). Synthesis and biological activity of benzaldehyde O-alkyloximes as abscisic acid mimics (Part 1). Biosci Biotechnol Biochem 56: 256–60.
Zhang X, Jiang L, Wang G, Yu L, Zhang Q, Xin Q, Wu W, Gong Z, Chen Z. Structural insights into the abscisic acid stereospecificity by the ABA receptors PYR/PYL/RCAR. PLoS ONE. 2013;8:e67477.
Zhou R, Cutler AJ, Ambrose SJ, Galka MM, Nelson KM. A new abscisic acid catabolic pathway. Plant Physiol. 2004;134:361–9.
Zou J, Abrams GD, Barton DL, Taylor DC, Pomeroy MK, Abrams SR. Induction of lipid and oleosin biosynthesis by (+)–abscisic acid and its metabolites in microspore-derived embryos of Brassica napus L. cv. Reston. Biological responses in the presence of 8′-hydroxy abscisic acid. Plant Physiol. 1995;108:563–71.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media Dordrecht
About this chapter
Cite this chapter
Todoroki, Y. (2014). ABA and Its Derivatives: Chemistry and Physiological Functions. In: Zhang, DP. (eds) Abscisic Acid: Metabolism, Transport and Signaling. Springer, Dordrecht. https://doi.org/10.1007/978-94-017-9424-4_1
Download citation
DOI: https://doi.org/10.1007/978-94-017-9424-4_1
Published:
Publisher Name: Springer, Dordrecht
Print ISBN: 978-94-017-9423-7
Online ISBN: 978-94-017-9424-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)