Abstract
The definition of the automatic protein function means designating the function with the automation by utilizing the data that already revealed unknown protein function. The demand for analysis on the sequencing technology such as the next generation genome analysis (NGS) and the subsequent genome are on the rise; thus, the need for the method of predicting the protein function automatically has been more and more highlighted. As for the existing methods, the studies on the definition of function between the similar species based on the similarities of sequence have been primarily conducted. However, this paper aims to designate by automatically predicting the function of genome by utilizing InterPro (IPR) that can represent the properties of the protein family, which similarly groups the protein function. Moreover, the gene ontology (GO), which is the controlled vocabulary to describe the protein function comprehensively, is to be used. As for the data used in the experiment, the analysis on properties was conducted in the sparse state that is deflected to one side. Thus, this paper aims to analyze the prediction method for protein function automatically through selecting the features, assigning the data processing and weights and applying a variety of classification methods to overcome that property.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
88.1 Introduction
With the rapid development of the next genome sequencing technology (Next Generation Sequencing), the proteins whose function has not been defined are on the rise and it takes a lot of time and economic costs to assign the function manually. Therefore, the requirements for the automatic annotation processing (functional gene annotation) that can designate the protein function automatically have been gradually increased. As for the traditional method, the biologists defined the function by conducting the experiment manually in order to define the protein function. But there are some limitations in terms of time and efforts due to the explosive increase in data with the development of sequencing equipment in order to define the protein function by a manual method. As for new proteins that have not been revealed experimentally, the protein functions could be automatically annotated by the automatic annotation processing if the model was created by using the known protein function; therefore, it would be possible to predict the function through the much more reduced time than the existing method based on experimentation. Many of the existing studies have conducted the researches by utilizing the DB (protein functional site, gene expression, etc.) in which the pattern of base sequence, similarities and protein function, in order to define the protein function. Of those, InterPro (IPR) [9] defines the protein group (family), domain, etc. as the database to be used to define those unknown new protein functions. In the case of newly revealed proteins through InterProScan [15], IPR can be easily obtained. Thus, it would be appropriate to use it as a feature to define a function. In addition, Gene ontology [2] can be regarded as a language to represent the properties of genes, and it consists of the three independent hierarchical structures that include molecular function, biological process and cellular component. In the consortium, the database is still in progress at present in order to describe the comprehensive feature of genome. And many of the search tools and automation classification tools, which are based on this, have been researched. This paper aims to analyze the methods that allow for the automatic annotation processing through comparing the relationship between IPR and Gene ontology by a variety of methods as utilizing the known data.
88.2 Related Studies
In the case of the functional annotation that identifies the function of genome automatically, various studies related to the database and automatic annotation are in progress in order to define the function of genome of various species from human beings to small microorganisms. The studies are in progress to allow for the automatic prediction of the protein function by an easy access through web or automatic installation, and mostly the database to manage this systematically is also in progress as it has continued to be updated. However, the method of defining the protein function automatically is still at the initial phase; thus, the accuracy is not very high. In the case of using the interpro2go that was manually mapped as in GOA [3], mapping is relied for defining the function; thus, the accuracy is not high. Most studies are conducted by small species to increase the accuracy, and the prediction methods for the protein function based on the calculation that has been researched makes a judgment mostly by utilizing the similarities of sequence.
As for the most frequently used tools, they include Gotcha [13], OntoBLAST [16], Blast2GO [4], AutoFACT [11], etc. Gotcha [13] can have the similarity of sequence and the Directed Acyclic Graph (DAG) of Gene ontology, in other words, the parent node can have several offspring nodes. Thus, it is the method of utilizing the property in which the parent node means the functions of more comprehensive meaning. It is the method of automatically naming GO by assigning a score to GO owned by the genome that is determined to be similar by judging the similarity of sequence. Blast2GO [4] is the method of annotating new protein functions that cannot be known by Gene ontology that is owned by a similar sequence after judging the similarity of sequence by utilizing BLAST [1]. It is the prediction model for the accuracy by assigning weights in accordance with the evidence code that is the annotation code of GO at this point. The evidence code means the code of GO to indicate whether it is automatically named (IEA) and it is determined by the similarity (ISS). OntoBLAST [16] is the method of finding possible protein functions from GO, which are obtained also from BLAST search. AutoFACT [11] proposed a fast annotation method by utilizing BLAST with the relevant database.
88.3 Methods
88.3.1 Features of Data
The data to be used is Saccharomyces Cerevisiae; it is one of yeast fungus; thus, it belongs to the fungus class and it is the most well-known data by the experiments. Since it forms a relatively small dataset as compared with the other species and it already comes out its related function; thus, it would be an appropriate data for establishing a model for the automatic annotation processing. For the extraction of this data, 4,370 proteins could be obtained as a result of searching and extracting Saccharomyces Cerevisiae only from SWISS-PROT.
The property to be used as a feature to create a model of data is IPR. IPR has the appropriate features for the reference data that include the protein family binding the protein functions in a similar way and the functions of Prosite, Prints, Pfam, Prodom, SMART, TIGRFams and PIR SuperFamily that play the central role to refer to the functional domain database. GO is utilized as the reference data for defining the function automatically. GO forms a hierarchical structure and divided into the three big classes—cellular component, molecular function and biological process.
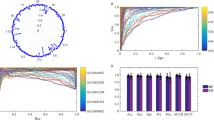
When counting the total number of IPR and GO term possessed by the 4,370 extracted proteins of Saccharomyces Cerevisiae, it was found to have 2,624 IPRs and 2,438 GO terms. When this data had one of the properties of IPR or GO term for each protein, it was represented in a binary form. It is represented in a large matrix (4,370 × 2,438) of GO in a binary form by representing “1” when the proteins have one term of particular GO terms and “0” when they do not have one as parsing Gene ontology at Ontology in the data section of SWISS-PROT. Also as for IPR, the IPR data was configured in a matrix form of 4,370 × 2,624 by a matrix of binary form as representing whether each protein has it through listing IPRs possessed by Saccharomyces Cerevisiae proteins after extracting InterPro in the family and domain database section with the same method as described above. A diagram for representing GO of IPR for each protein in a matrix and lining up the quantity of GOs that can be represented by “1”, in other words, the quantity owned by the proteins would be the same as shown in Fig. 88.1. As shown in Fig. 88.1, it has the problem that it does not have a sufficient quantity for each GO to conduct the learning. When viewed from the perspective of one single GO, the number of case in which it has only one single GO is 414. This means that only one protein owns the relevant GO; thus, it would not appropriate to utilize it as the learning data. In addition, the validity was tested through the 10-fold cross validation; thus, GO that has fewer quantity than a certain level would not be appropriate for the use as the learning data.
However, the biggest problem of the data is that the data exists sparse even though it has a relatively sufficient quantity to be utilized as the learning data. And the fact that the protein data not having the relevant IPR or GO are inclined to one side than the data having the relevant IPR or GO when viewed from a particular IPR or GO term is also a problem. For instance, only 50 proteins have a particular GO out of 4,370 proteins when viewed from the perspective of a particular single GO; therefore, they are represented by “1” and the remaining 4,320 proteins are represented by “0” since they do not have it. In the case of conducting the learning and experiment with such data, it is quite often predicted that most do not have it since the learning is conducted as being excessively inclined to “0” that is not owned by the learning result; thus, it cannot become an effective model for the automatic function prediction and command processing that has to assign new functions. There are many cases represented by “0” representing “not having” in the case of IPR in addition to GO. This cannot be utilized as an effective feature. Due to such properties of these two features, this paper aims to apply the method as to the feature selection and balanced dataset. Moreover, it aims to analyze the results by converting the binary form of the data into a non-binary form (weighted IPR) by utilizing the correlation coefficients since the data to be processed is not a binary form.
88.3.2 Prediction Method of Protein Function
This paper aims to compare and analyze the prediction method of protein function by utilizing the data having a sufficient quantity of data as the data of learning and experiment of this paper among the data described in 3.1. Before the analysis, it would be essentially required to have a process of reconfiguring it as a balanced dataset due to the feature of not being balanced with the sparseness of data. It aims to compare the case of applying the feature selection by the three mutually different learning methods and the case of conducting the weighted IPR that adds weights to the data respectively.
This paper shows to comparison and analysis of the prediction methods by the methods presented in Table 88.1. First as for the learning methods, adaboosting [7] is the method of creating an optimal classification through several times of learning by assigning weights as to the instances wrongly classified by the method of weak learner. SVM is the method of seeking a boundary that makes the error of margin that can differentiate the class to be classified at the hyperplane; thus, it is one of the learning methods of machine learning. SMO [10] is the most well-known tool of libsvm [5]; thus, it can be regarded as the method that has simplified the complexity of SVM by the sequential minimal optimization. As for the methods to be presented in Sect. 88.3.2.2., the case of using the feature selection method and the case of not using it were compared and analyzed as W/O in Table 88.1 meant Without and W meant With. Furthermore, it compared the case of using the method called weighted IPR to be stated in Sect. 88.3.2.3 with the case of using the original data as it was.
88.3.2.1 Dataset Reconfiguration to Adjust Balance
As shown in Fig. 88.2, there are more proteins not having the relevant GO than those having it when viewed based on a particular GO. However, there are more proteins not having a particular GO, in other words, negative proteins, when learning with such data; therefore, there would be a high degree of probability for the modeling that most of learning results turned out to be not having it. However, it is only possible to find it out by creating a model having the relevant GO rather than a model not having GO. When experimenting with proteins that are not able to perform the function, it is impossible to obtain the desired result. Thus, balanced sampling approach is employed to overcome this handicapped data property.
There are the under-sampling method and the over-sampling method in terms of reconfiguring the data that consists of balanced proteins; the over-sampling [6] is the method of making the number equal by generating the data that become the major in terms of quantity as many as the quantity at which the relatively fewer data becomes the major in a random way. The under-sampling [12] is the method of meeting the ratio by selecting more data randomly based on the data whose quantity is few. In this experiment, the data that is relatively few in quantity is more important information; therefore, this paper reduces that quantity by utilizing the under-sampling. As shown in Fig. 88.2, the data indicated by “1” is to be named as positive protein, whereas the data indicated by “0” is to be named as negative protein. And it is supposed to learn with proteins that are fewer than 4,370 in terms of the quantity of protein by reconfiguring the data for the learning model at each GO through selecting the negative proteins just as many as the quantity of positive proteins.
88.3.2.2 Feature Selection
As shown above Fig. 88.1, IPR has the matrix of many binary features of 2,624 when viewed based on one GO. It is the well-known fact that learning and experimenting by selecting only meaningful features would reduce the time to be taken and have a better result as compared with learning and experimenting the method presented above by these many matrices [8, 14]. When representing the case in which “1” representing that each IPR has protein by positive data and the case of not having it by “0”, the positive negative data is to be counted for each protein. The positive data is represented as “IPos”, “GPos” and the negative data is represented as “INeg”, “GNeg” at IPR and GO in respectively and it is possible to classify the state of IPR and GO for each protein. They can become 4 states as shown in Table 88.2.
It is possible to calculate the four probabilities (NGPos_IPos/NPos, NGNeg_IPos/NPos, NGPos_INeg/NNeg, NGNeg_INeg/NNeg) by utilizing the 4 data, where NPos stands for the total number of positive proteins and NNeg means the total number of negative proteins. These probabilities represent a conditional probability, which the Gene ontology term may possess depending on the conditions of each IPR. When viewed the property by adding these conditional probabilities as an example of GO:0000329, the diagram as shown below could be viewed. The x-axis means several IPRs that are being experimented and the y-axis is the value of adding the conditional probabilities. It is possible to see the phenomenon of which most are concentrated in Area 1. On that account, 99 % of them are those IPRs having negative IPR term and also negative Gene ontology (Fig. 88.3). This paper selected the features based on the IPRs that are concentrated in Area 2 as excluding these IPRs. In other words, this is the learning method of utilizing only the selected index as a feature by selecting only the index of IPRs in Area 2 among the 2,624 IPRs by calculating the conditional probabilities above for each GO.
88.3.2.3 Weighted IPR
IPR that is utilized as the feature is the binary data that consists of 0 and 1. When converting this data into a continuous form rather than binary form by utilizing a correlation coefficient, IPR feature data would be expected to select a feature without partiality. This paper aims to analyze the performance between the two methods by the differences between the feature extraction using the binary data that consists of 0 and 1 and the weighted IPR of a continuous form as naming this data as the weighted IPR.
For instance, as shown in Table 88.3, the table that is composed of 0 and 1 would be modified into a table that utilizes a correlation coefficient (Table 88.4). A correlation coefficient becomes a value closer to 1 with a higher degree of correlation, whereas it is represented by a value close to 0 when there is no correlation. In addition, it becomes a negative value when there is a mutually contradicting correlation.
This paper aims to change to weight coefficients as proposed by Formula 1. Based on this correlation coefficient. First, each protein p possesses IPR from 1 to n. All the proteins to be experimented are represented by IPR of n units. A particular protein having IPR would be represented by 1, whereas those not having IPR would be represented by 0.
For instance, Protein 1 in Table 88.3 is represented as not having IPR 1, IPR 5 and IPR 6, which are 0, whereas IPR 2, IPR 3 and IPR 4 are represented by IPR possessed by the relevant protein. At this point, there is a relationship between IPR 5 and IPR 6 since IPR 1 is not a property that is not owned when viewed by each IPR of Protein 1. In reference with Table 88.4, the weight (IPR1) value of IPR 1 of corr(IPR1, IPR5) = 0.6124 and corr(IRP1, IPR6) = 0.1667 Protein 1 is 0.6124 + 0.1667 = 0. 7791. Moreover, the value of weight sum (IPR 1) is represented by IPR 1 = 0; therefore, the value of adding all the correlation coefficients of IPR 5 and IPR 6 becomes 0.7418. Essentially the value was the binary form of 0 and 1 in order to calculate the weighted sum (IPR 1) as to IPR 1 of Protein 1 of the calculated value; therefore, there finally generates the value of 0.5 × 0.7791/0.7418 = 0.5251 by giving the weighted value 0.5. A new data defined in the new weighted IPR would be generated by such method. Table 88.5 can be regarded as one of such cases.
Formula 1. Converting weighted IPR by the correlations and weights
Figure 88.4 is the diagramed result by a graph of the error rate of applying the 12 methods presented in Table 88.1 to GO:0003700. The 4 mutually different methods presented in Table 88.1 were represented by different colors. And a better performance would be shown with fewer error; thus, selecting the weighted method and features suggested by SVM or SMO resulted in a similar or less error rate. As to the 4 methods utilizing the feature selection and weighted method among the proposed 12 methods, the results of comparing and experimenting each GO term are as shown in Table 88.6. As shown in Table, they are found to have a high prediction rate of more than a range of 97–99.
88.4 Conclusion
This paper compared and evaluated the performance that could define the protein function by applying the classification algorithm by utilizing the feature selection and data transformation. As for the data to be processed, the data having GO term has been composed in much less quantity than the protein not having GO term when viewed by individual GO term. In addition, IPR that is set as the feature point is sparsely distributed; thus, it becomes difficult to learn all the protein data through the general classification algorithm. Due to such limitations, the performance as to the automatic annotation was compared by various classification methods through extracting only the GO term having the standard level or more as the learning subject. Moreover, the performance with the original data was also analyzed by the method of using the binarized data as the correlation coefficient through converting it into a newly weighted coefficient.
However, as for the data sampling and feature selection processed in this paper, the GO term learned primarily the data of protein having a certain amount or more for the learning at Saccharomyces Cerevisiae; thus, there is the limitation that the quantity of learned data of GO term is small. If it is to learn by utilizing the data that includes a variety of species such as SWISS PROT in order to overcome this limitation, it will be possible to expect to utilize the automatic function prediction by learning more GO terms with the use of large quantity of data. Thus, this paper aims to study a learning method that is appropriate for this. In addition, it aims to prepare a base to allow for the automatic annotation by seeking for different features that can be utilized as a keyword in addition to IPR when trying to find out unknown protein functions by identifying the correlation with GO.
References
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25(17):3389–3402
Ashburner M, Ball CA, Blake JA et al (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25:25–29
Camon E, Magrane M, Barrell D, Lee V, Dimmer E, Maslen J, Binns D, Harte N, Lopez R, Apweiler R (2004) The gene ontology annotation (GOA) database: sharing knowledge in Uniprot with gene ontology. Nucleic Acids Res 32: D262–D266
Chang CC, Lin CJ (2011). LIBSVM: a library for support vector machines. ACM Trans Intell Syst Technol 2(3):27
Chawla N, Bowyer K, Hall L, Kegelmeyer P (2002) SMOTE: synthetic minority over-sampling technique. JAIR 16:321–357
Conesa A, Götz S, García-Gómez JM, Terol J, Talón M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21(18):3674–3676 Epub, Aug 4 2005
Freund Y, Schapire R (1996) A short introduction to boosting. J Japan Soc Artif Intell 14(5):771–780
Guyon I, Elisseeff A (2003) An introduction to variable and feature selection. J Mach Learn Res 3:1157–1182
Hunter S, Jones P, Mitchell A et al (2011) InterPro in 2011: new developments in the family and domain prediction database. Nucleic Acids Res 40:D306–D312
John CP Sequential minimal optimization: a fast algorithm for training support vector machines
Koski LB, Gray MW, Lang BF, Burger G (2005) AutoFACT: an automatic functional annotation and classification tool. BMC Bioinf 6:151
Kubat M, Matwin S (1997) Addressing the curse of imbalanced training sets: one-sided selection. In: Proceedings of the fourteenth international conference on machine learning, pp 179–186
Martin DM, Berriman M, Barton GJ (2004) GOtcha: a new method for prediction of protein function assessed by the annotation of seven genomes. BMC Bioinf 5:178
Quevillon E, Silventoinen V, Pillai S et al (2005) InterProScan: protein domains identifier. Nucleic Acids Res 33:W116–W120
Shahib A Al, Breitling R, Gilbert D (2005) Feature selection and the class imbalance problem in predicting protein function from sequence. Appl Bioinf 4(3):195–203
Zehetner G (2003) OntoBlast function: from sequence similarities directly to potential functional annotations by ontology terms. Nucleic Acids Res 31(13):803–3799
Acknowledgments
This research was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (2013R1A1A2063006).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media Dordrecht
About this paper
Cite this paper
Jung, J., Lee, H.K., Yi, G. (2014). Functional Annotation of Proteins by a Novel Method Using Weight and Feature Selection. In: Park, J., Zomaya, A., Jeong, HY., Obaidat, M. (eds) Frontier and Innovation in Future Computing and Communications. Lecture Notes in Electrical Engineering, vol 301. Springer, Dordrecht. https://doi.org/10.1007/978-94-017-8798-7_88
Download citation
DOI: https://doi.org/10.1007/978-94-017-8798-7_88
Published:
Publisher Name: Springer, Dordrecht
Print ISBN: 978-94-017-8797-0
Online ISBN: 978-94-017-8798-7
eBook Packages: EngineeringEngineering (R0)