Abstract
Acquired resistance to fungicides in fungal plant pathogens is a challenge in modern crop protection. Fungi are indeed very able to adapt to changing environmental conditions, such as the introduction of a new fungicide in the agricultural practice. Several genetic mechanisms may underlay fungicide resistance and influence the chance and time of its appearance and spreading in fungal populations. Resistance may be caused by mutations in major genes (monogenic or oligogenic resistance) or in minor genes (polygenic resistance) which may occur in nuclear genes as well as in cytoplasmic genes. They are immediately expressed in haploid fungi, while they may be dominant or recessive in diploid fungi. Allelic variants may cause different levels of resistance and/or different negative pleiotropic effects on the fitness of resistant mutants. The sexual process, where occurring, plays an important role in releasing new recombinant genotypes in fungal populations. Heterokaryosis provides multinucleate fungi with a further mechanism of adaptation. Resistant mutants can be obtained from samples representative of field population of a pathogen or under laboratory conditions through selection of spontaneous mutations or following chemical or physical mutagenesis. Nowadays, molecular tools, such as gene cloning, sequencing, site-directed mutagenesis and gene replacement, make genetic studies on fungicide resistance amenable even in asexual fungi for which classical genetic analysis of meiotic progeny is not feasible.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Resistance to chemicals in microorganisms is a very common phenomenon occurring whenever antimicrobial compounds are used against pathogens of plants, animals or humans. Natural or innate resistance refers to intrinsic features (e.g. the lack of a specific molecular target and/or a metabolic pathway) protecting the organism from the effects of antimicrobials. For example, strobilurin-producing organisms, including wood-degrading Basidiomycetes, such as Strobilurus tenacellus, have innate resistance to their own strobilurins that show, instead, activity against a very broad spectrum of fungi and Oomycetes. Acquired resistance refers to organisms that in their wild-type form are sensitive and may develop resistance after their exposure to an antimicrobial compound. Acquired resistance is due to genetic modifications transmissible to the progeny so that a chemical that was once effective against the organism is no longer effective.
Resistance to fungicides used in agriculture as well as in animal or human health care is a more recent phenomenon than resistance to antibiotics (Coplin 1989; Cookson 2005) and insecticides (Brown 1977). Until the late 1960s, fungicides used in crop protection (e.g. sulphur, copper derivatives, dithiocarbamates) were indeed essentially multisite inhibitors, affecting multiple target sites and hence interfering with many metabolic processes of the pathogen. Despite their protracted and widespread use, acquired resistance to multisite fungicides is still a rare event. This is because there is a low probability that a number of mutations at different loci, needed for the onset of the resistance, simultaneously occur in fungal cells and, if this happens, the mutated isolates remain viable. Afterwards, with the introduction of single-site fungicides and as a consequence of their frequent and repeated use, fungicide resistance has become a major concern in modern crop protection seriously threatening effectiveness of several fungicides (Brent and Hollomon 2007a, b).
Fungicide resistance is hence a result of adaptation of a fungus to a fungicide due to a stable and inheritable genetic change, leading to the appearance and spread of mutants with reduced fungicide sensitivity (Delp and Dekker 1985).
2 Genetic Bases of Fungicide Resistance
Genetics of fungicide resistance have been previously reviewed by Grindle (1987), Grindle and Faretra (1993), Steffens et al. (1996) and Ma and Michailides (2005), and deeper information is available on the website of the Fungicide Resistance Action Committee (www.frac.info).
Fungal genetic backgrounds and genetic bases of resistance are key factors in the intrinsic risk of resistance and influence its evolution in the pathogen populations. For example, the occurrence of genetic recombination through the sexual process, where it regularly occurs in nature, or parasexuality, in essentially asexual fungi, may greatly influence the dynamics of resistant subpopulations producing new combinations of resistance and fitness traits originally occurring in separate individuals.
Most genetic studies on fungicide resistance have been carried out on ‘model’ saprophytic Ascomycetes, such as Aspergillus nidulans, Neurospora crassa and Saccharomyces cerevisiae. Nevertheless, the genetics of fungicide resistance has been investigated in several pathogenic fungi (Table 2.1).
Key factors in the genetic bases of fungicide resistance are (1) the number of loci involved, (2) the number of allelic variants at each locus, (3) the existence and relevance of dominant or recessive relationship between resistant and wild-type alleles (Borck and Braymer 1974) and (4) the additive or synergistic interactions between resistance genes.
Genes responsible for fungicide resistance may be located on chromosomes inside the nucleus or on extrachromosomal genetic determinants. Nuclear and cytoplasmic genes can be distinguished by their inheritance patterns. Nuclear genes typically show classical biparental (disomic) inheritance in sexual crosses, i.e. the zygote receives one allele of each gene from each of its parents. In contrast, genetic material in the cytoplasm has a non-Mendelian inheritance and is characterized by uniparental (usually maternal) transmission (Griffiths 1996). In addition, cytoplasmic genes differ from nuclear genes in showing vegetative segregation and intracellular selection potentially affecting resistance stability (Birky 2001; Ziogas et al. 2002).
Most fungicide-resistance genes are located on nuclear chromosomes. In most cases, there is only one copy of resistance gene in the genome and mutations are usually located in gene sequences encoding enzymatic or structural proteins. However, multidrug resistance (MDR) in B. cinerea and other fungi is caused by overexpression of membrane efflux transporter genes resulting in an increased efflux of toxicants that reduces fungal sensitivity to several unrelated fungicides as well as plant defence chemicals (reviewed by Kretschmer 2012). In MDR1 strains of B. cinerea, resistance is conferred by mutations in the regulator mrr1 gene encoding a transcription factor controlling the ABC transporter AtrB gene, whereas in MDR2 strains resistance is caused by an insertion of a retrotransposon-derived sequence in the promoter region of the facilitator superfamily (MFS) transporter gene mfsM2 (Kretschmer et al. 2009).
Fungicide resistance may result from mutations in single major genes (Georgopoulos 1988) or from additive (Kalamarakis et al. 1991; Lasseron-de Farandre et al. 1991) or synergistic interactions (Molnar et al. 1985) between several mutant genes.
Monogenic and oligogenic resistance are caused, respectively, by one or few major genes. Major genes have an appreciable influence on the phenotype, and resistance mutations cause a qualitative change in the response to a fungicide with the appearance in the field of new fungicide-resistant subpopulation(s) well distinguishable from the wild-type sensitive one (Fig. 2.1). Most cases of fungicide resistance are due to mutations in major genes (Table 2.1). Mutations in major genes conferring resistance to fungicides having different modes of action may also occur in a same isolate, causing multiple resistance. In oligogenic resistance, several different major genes are involved, any one of which can mutate to cause an increase in resistance to a same fungicide. For instance, kasugamycin resistance in Pyricularia oryzae as well as resistance to the two fungicides ethirimol and triadimenol in Blumeria graminis f.sp. hordei may be controlled by three different loci where a resistance allele at any one locus confers resistance (Taga et al. 1979; Brown et al. 1992). Furthermore, differently from what is usually observed in most fungi where a single multiallelic gene is responsible for resistance to benzimidazole fungicides, the resistance of Fusarium oxysporum to benzimidazoles is caused by mutations in two major genes which interact synergistically conferring high degrees of fungicide resistance (Molnar et al. 1985).
Population dynamics of fungicide resistance in monogenic resistance (upper) and polygenic resistance (bottom). Disruptive or directional selection is caused by the usage of fungicides having the same mode of action at risk of resistance. Stabilizing selection is due to possible reduction of fitness of fungicide-resistant mutants
Different mutations in a same gene may cause different levels of resistance to a particular fungicide; this is known as multiallelic resistance. In the past, multiallelic resistance could be assessed only on the ground of phenotypic differences in the level of resistance and/or pleiotropic effects of mutations. With the availability of molecular and sequencing tools, nowadays it is clear that multiallelic resistance is quite common (Table 2.1).
Each mutant allele can be partially/completely dominant or partially/completely recessive to its wild-type allele. That is, when mutant and wild-type alleles of the same gene are combined in the same fungal cells or hyphae, the phenotype may be fungicide resistant (mutant) or fungicide sensitive (wild type).
Combinations of major genes may interact when they are present in the same fungal cells, so that the phenotype of a double mutant may be different from either single-gene mutants (Molnar et al. 1985). Usually, however, one mutant gene is epistatic to another mutant gene, which means that the double mutant has the same level of resistance of the single-gene mutants (Kappas and Georgopoulos 1970; Van Tuyl 1977). The presence of modifier genes affecting phenotypic response of resistant mutants has been suggested to influence the expression of response to phenylamides in Oomycete pathogens (Crute and Harrison 1988) or to mediate fitness of resistant mutants as found in mutants of N. crassa resistant to dicarboximides (Grindle and Dolderson 1986) and A. nidulans resistant to imazalil (van Tuyl 1977). The consequent increase in fitness will result in better survival and possible selection of resistant subpopulations in the field.
Polygenic resistance is due to mutations in minor genes. Those have individually a little effect on the phenotype and cause hence a negligible reduction in the sensitivity to a fungicide. However, numerous mutated minor genes may contribute, with an additive effect, to produce an appreciable increase of the level of resistance. In the field, the result is a quantitative decrease of the sensitivity to a fungicide with a slow, continuous and gradual shift of the fungal population towards increasing resistance levels (Fig. 2.1). Polygenic resistance is much more difficult to be detected and ascertained in the field. Polygenic resistance was demonstrated in B. graminis f.sp. hordei to ethirimol (Hollomon 1981) and triadimenol (Hollomon et al. 1984). Resistance to dodine is polygenic in Nectria haematococca var. cucurbitae (Kappas and Georgopoulos 1970). Ultraviolet-induced mutants of N. haematococca var. cucurbitae also show polygenic inheritance for resistance to fenarimol (Kalamarakis et al. 1991), fenpropimorph and terbinafine (Lasseron-deFalandre et al. 1991).
Cytoplasmic genes are present in mitochondria, plasmids and viruses. Mitochondrial genome, which contains mitochondrial rRNA genes and some of the proteins of the respiratory chain, is the most relevant among fungal extrachromosomal genetic elements affecting resistance to chemicals. However, antibiotic-resistance genes have been located on fungal episomes, plasmids or viruses (Guerineau et al. 1974).
Natural or induced resistance to QoI fungicides, inhibitors of mitochondrial respiration at the Qo site of the cytochrome bc1 complex (complex III), is usually conferred by point mutations in the mitochondrial cytb gene causing amino acid substitutions in the target protein. In particular, at least three possible codon changes have been associated to a moderate (F129L or G137R) or, more frequently, high (G143A) level of resistance to QoIs in several fungal species (Grasso et al. 2006; Fernández-Ortuño et al. 2008). The presence of a G143-associated group I-like intron in the cytb gene in some fungal species (i.e. Puccinia spp., Uromyces appendiculatus, Alternaria solani) or isolates (i.e. B. cinerea) prevents the occurrence of the G143A mutation and QoI resistance, since it would be lethal because it would be affecting the correct intron splicing process (Grasso et al. 2006).
Analysis of meiotic progenies of appropriate crosses between sensitive and resistant strains confirmed cytoplasmic (maternal) inheritance of QoI resistance in B. graminis (Robinson et al. 2002), Venturia inaequalis (Steinfeld et al. 2002) and B. cinerea (De Miccolis Angelini et al. 2012a). The segregation pattern in randomly collected progenies is expected to be in a phenotypic 1:0 ratio in most fungal species showing a uniparental, anisogamous inheritance of mitochondrial genome or 1:1 ratio in species, such as A. nidulans and B. graminis f.sp. tritici, showing an hermaphroditic, isogamous mitochondrial inheritance (Robinson et al. 2002).
Wild-type and mutated mitochondrial DNA carrying the G143A mutation in the cytb gene may coexist in heteroplasmic state within a single isolate, as demonstrated in several species, including V. inaequalis (Zheng et al. 2000), B. cinerea (Ishii et al. 2009) and other fungal pathogens (Ishii et al. 2007). Equilibrium between resistant and sensitive mitochondria depends on the strength of selective pressure (Ishii 2010). In Podosphaera leucotricha, the relative proportion of mutated and wild-type mitochondria is associated with differences in QoI sensitivity levels of the isolates (Lesemann et al. 2006). An instability of QoI resistance in heteroplasmic isolates grown in absence of selective pressure has been frequently reported (Ishii 2012a) suggesting a fitness cost associated to the resistance (Markoglou et al. 2006).
3 Ploidy Level
Differences in ploidy level, affecting the number of alleles at each locus, constitute a major genomic trait influencing the onset and subsequent evolution of fungicide resistance. Firstly, frequency of mutations that may arise in single individuals is directly related to the ploidy level as a result of the different numbers of mutational targets (Otto and Gerstein 2008).
Most phytopathogenic fungi are in haploid state for the major part of their life cycle. In contrast, Oomycetes typically show a diploid life cycle and the haploid phase is restricted to the gametes (Fincham et al. 1979). Furthermore, polyploids have been frequently identified among Oomycetes, such as Plasmopara viticola and Phytophthora spp. (Rumbou and Gessler 2006; Bertier et al. 2013).
In haploid fungi, mutations conferring resistance are immediately expressed and then directly exposed to selection, while in diploids or polyploids, mutations first appear in heterozygotic state and their phenotypic effects can be masked by dominant wild-type alleles on the homologous chromosome. For this reason, resistance mutations spread more rapidly in haploid than in diploid or polyploid populations. Fixation time may be reduced and selection against deleterious pleiotropic effects of mutations is more effective in haploids than in diploids (Anderson et al. 2004; Otto and Gerstein 2008).
CAA (carboxylic acid amide) fungicides, inhibitors of cellulose biosynthesis in Oomycete phytopathogens, are considered at low to medium resistance risk depending on the fungal species. Resistance to CAAs in P. viticola is controlled by one or two recessive nuclear genes, as demonstrated through sexual crosses between CAA-sensitive and CAA-resistant isolates and analysis of segregation patterns of sensitive and resistant phenotypes in F1 and F2 progenies (Gisi et al. 2007; Blum and Gisi 2008) and by sequence analysis of putative resistance genes (Blum et al. 2010). Classic genetic analysis also showed that resistance to all CAA fungicides co-segregates and has thus the same genetic basis (Young et al. 2005; Gisi et al. 2007). However, no cross resistance exists between CAA and other fungicides currently available against Oomycetes, such as phenylamides and QoI fungicides, where the intrinsic risk of resistance is estimated to be significantly higher than CAA due to their genetic differences. Resistance to phenylamides is indeed a monogenic trait, conferred by a semidominant chromosomal gene (Gisi and Cohen 1996; Knapova et al. 2002), while QoI resistance is due to mutations in the mitochondrial cytb gene (Gisi and Sierotzki 2008).
Similar to CAA, resistance to the new benzamide zoxamide in isolates of Phytophthora capsici is recessive and is conferred by two nontarget nuclear genes (Bi et al. 2014). This implies that resistance phenotype is expressed only in homozygous mutants, thus limiting resistance spreading and risk.
Nevertheless, the risk of resistance is significantly increased by the occurrence of gene recombination, even if several cycles of sexual process may be required for making resistance fixed and fully expressed in phenotypically aggressive and well-adapted isolates of the pathogen. Sexual recombination naturally occurring under field conditions has been proposed, for instance, as a possible explanation of the higher risk of CAA resistance assessed in field populations of Pseudoperonospora cubensis as compared to in vitro estimations (Zhu et al. 2007). Moreover, CAA resistance has been experienced in P. viticola field populations since shortly after their introduction, while no reduced sensitivity to CAA has been detected in other Oomycetes, such as the late blight pathogen, Phytophthora infestans, despite their intensive usage against these pathogens and extensive monitoring. It has been suggested that the lower risk of CAA resistance in P. infestans may be due to the lower frequency of sexual recombination under field conditions, as well as to polyploidy, heterokaryosis (Catal et al. 2010) and chromosomal aberrancies (Gisi 2012).
4 Heterokaryosis and Nuclear Number
The presence of two or more genetically different haploid nuclei, coexisting in a common hyphal compartment, occurs frequently in some fungal taxa and is a potential source of genetic variation. In multinucleate Ascomycetes, this condition, known as heterokaryosis, often permits changes in the proportions of different nuclei in response to selection and is a prerequisite to parasexual recombination (Davis 1966). In heterothallic Basidiomycetes, two distinct parental haploid nuclei coexist without fusion in each cell establishing a stable dikaryotic state. The dikaryon is genetically equivalent to a diploid, as two haploid genomes of different origins exist in each cell even if they remain separated in different nuclei.
Heterokaryons and dikaryons, harbouring several nuclei, offer the opportunity of genes to complement each other (genetic complementation). Heterokaryons harbouring both fungicide-resistant and fungicide-sensitive nuclei may be able to grow in the presence or absence of fungicides (Grindle 1987). They can adapt to fluctuations in fungicide exposure as a result of changes in the proportions and distribution of resistant-sensitive nuclei within cells (Meyer and Parmeter 1968; Ogden and Grindle 1983). Nucleotypic competition and selection after exposure to fungicides and the ability of heterokaryons to adapt to modified environmental conditions have been demonstrated, for instance, in B. cinerea strains resistant to dicarboximides (Summers et al. 1984) or to anilinopyrimidines (Santomauro et al. 2000).
Dominance or recessivity can be tested by inducing hyphal anastomosis between one strain carrying the wild-type allele and the other the mutant allele of a resistance gene to form heterokaryotic mycelium. Incompatibility impeding heterokaryon establishment can be overcome by fusion of protoplasts. The mutant allele is completely (or partially) dominant if the heterokaryon is phenotypically identical (or similar) to the mutant parent; it is completely (partially) recessive if the heterokaryon is phenotypically similar to the wild-type parent (Grindle 1987; Grindle and Faretra 1993).
5 Level of Resistance and Pleiotropic Effects of Resistance Mutations
Levels of resistance are usually quantified by determining from dose–response curves the concentration of fungicide needed to reduce ‘life’ parameters, such as colony or mycelium growth or spore germination by 50 % (effective concentration 50; EC50) and the minimal inhibitory concentration (MIC). A mutant can be designated resistant to a fungicide if its EC50 value is at least twice the EC50 value of sensitive wild-type isolates (Delp and Dekker 1985). However, small differences in EC50 values among resistant mutants and sensitive isolates may not be detected unless environmental variables are rigorously controlled and/or data from dose–response experiments are subjected to statistical analysis. Moreover, with small differences it may be difficult to establish whether resistance is due to a single major gene or to polygenes (Grindle and Faretra 1993).
A mutant gene conferring resistance to a particular fungicide often confers positive cross resistance to other fungicides having the same or related mode of action. On the contrary, mutant genes causing resistance to one fungicide may increase sensitivity to other chemicals (negative cross resistance) (Brent and Hollomon 2007a, b).
Resistance mutations may have deleterious pleiotropic effects on unrelated phenotypic characters, such as competitiveness, virulence, survival and reproductive success. Physiological mechanisms underlying resistance to fungicides may also be associated with a metabolic cost. Hence, resistant isolates may have lower fitness than wild-type sensitive isolates. Differences in fitness can be experimentally measured as reduction in mycelial growth rate, sporulation and conidial germination, pathogenicity, survival under stressing conditions, etc., in a fungicide-free environment. For instance, fitness penalty was observed in (1) DMI resistance in powdery mildews (Gisi et al. 2002); (2) resistance to dicarboximides and phenylpyrroles in several fungi, such as B. cinerea (Pollastro et al. 1996; Ochiai et al. 2001), N. crassa (Hollomon et al. 1997) and Monilinia laxa (Katan and Shabi 1982); (3) resistance to QoIs (G143A replacement) in field populations of P. viticola (Fernández-Ortuño et al. 2008) and Pyricularia grisea (Avila-Adame and Köller 2003) and in laboratory mutants of B. cinerea (Markoglou et al. 2006), C. beticola (Malandrakis et al. 2006) and Ustilago maydis (Ziogas et al. 2002), but not in B. graminis f.sp. tritici (Heaney et al. 2000; Chin et al. 2001), Mycosphaerella graminicola (Miguez et al. 2004) and Magnaporthe grisea (Avila-Adame and Köller 2003); (4) most of the mutations in the SdhB gene conferring resistance to SDHIs in B. cinerea except for SdhBH272Y (Lalève et al. 2014a; Veloukas et al. 2014); and (5) Penicillium expansum resistant to tebuconazole, fludioxonil and iprodione, but not to cyprodinil, coupled with reduction in patulin production (Karaoglanidis et al. 2011).
Mutations responsible for fungicide resistance may influence mycotoxin production. For instance, the production of 3-acetyl deoxynivalenol (3-ADON) was altered in isolates of Fusarium culmorum resistant to the DMI fungicide difenoconazole (D’Mello et al. 1997). A higher production of T-2 toxin, 4,15-diacetoxyscirpenol and neosolaniol was found in a carbendazim-resistant strain of Fusarium sporotrichioides (D’Mello et al. 1998, 2000). More recently, Zhang et al. (2009) found that benzimidazole resistance increased trichothecene production in F. graminearum. Laboratory mutants of Aspergillus parasiticus resistant to phenylpyrroles and dicarboximides produced more aflatoxins than the parental wild-type strain (Markoglou et al. 2008a). Similarly, laboratory mutant strains of A. parasiticus, A. ochraceus and F. verticillioides resistant to triazoles (epoxiconazole and flusilazole) and mutant strains of A. carbonarius and P. expansum resistant to fludioxonil produced significantly higher levels of mycotoxins (ochratoxins, patulin and fumonisins) compared to the parental sensitive strains (Doukas et al. 2008; Markoglou et al. 2008b, 2009).
It is generally assumed that fitness costs of resistance are invariable. However, Chin et al. (2001) showed that the cost of resistance to QoI fungicides in B. graminis varies with environmental conditions, such as temperature, being more costly under suboptimal conditions for the fungus.
6 Population Genetics
To develop effective resistance management strategies, it is crucial to know all the factors influencing relationship between sensitive and resistant strains.
The prevailing model explaining the selection of fungicide-resistant fungal populations considers random and rare mutations as the cause for pre-existing but infrequent resistant phenotypes prior to the introduction of a new fungicide (Torriani et al. 2009; Camps et al. 2012). Nevertheless, the evolutionary question on how populations adapt to novel environments, such as new antimicrobials, through de novo mutations or through selection from standing genetic variation, which affect the probability and speed of emergence of resistant alleles, is still debated (Hermisson and Pennings 2005; Hawkins et al. 2014).
Anyway, rare resistant mutants gain in competitiveness under the selection force of fungicide sprays and are selected to frequencies at which disease control becomes unsatisfactory (Milgroom et al. 1989; Skylakakis 1987; Wolfe 1982; Hobbelen et al. 2014). The shift towards resistance occurs at different rates depending on the number of genes conferring resistance. In monogenic resistance, a rapid shift towards resistance may occur, leading to discrete resistant subpopulation(s), while in polygenic resistance, the shift towards resistance progresses slowly, leading to a reduced sensitivity of the entire population. Resistant and wild-type subpopulations are in a dynamic equilibrium due to two selective pressures: i) the disruptive selection (directional selection in polygenic resistance), favouring resistant subpopulation(s), is due to repeated sprays with fungicides having the same mode of action at risk of resistance, and ii) stabilizing selection, favouring the wild-type sensitive populations, is caused by possible negative pleiotropic effect of resistance mutations leading to a reduced fitness (Fig. 2.1). Unfit mutants compete well only under the selection pressure of fungicide sprays, and, hence, resistance is at least partially reversible when the selection pressure is removed or minimized by applying resistance management strategies.
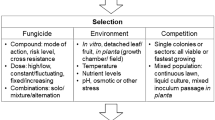
7 Obtainment of Resistant Mutants
Field isolates collected from diseased plants, plant debris, soil or air may include fungicide-resistant mutants, particularly if crops have been sprayed intensively with single-site fungicides. Resistant field isolates may be selected on appropriate agar media amended with a fungicide at a concentration inhibiting germination of conidia and/or mycelium growth of wild-type sensitive isolates. In choosing agar medium the mode of action of the fungicide must be complained. In the case of obligate biotrophic pathogens, plants or parts of plants must replace agar media. A number of monitoring methods are available (www.frac.info). Field isolates may display a broad variation making their genetic analysis more difficult than laboratory mutants (Grindle and Faretra 1993). It is advisable to obtain ‘monoconidial’ or ‘single hyphal tip’ isolates rather than ‘mass-conidial’ or ‘mass-hyphal’ isolates since they are likely to be genetically more homogeneous and stable. These traits are improved by repeated subculturing monoconidial isolates selecting the ‘most typical’ progeny.
Experiments under laboratory conditions are useful because it is possible to replicate them, to control the strength of selection and to use defined reference strains (Cowen et al. 2002). Resistant laboratory mutants can be generated in vitro from wild-type strains of known phenotype, and all mutants deriving from a same strain are near isogenic since their genomes are virtually identical, except for mutant gene(s) conferring resistance.
Selection of spontaneous mutations may be achieved by growing fungal colonies on media added with sublethal fungicide concentration; resistant hyphae grow better than the sensitive ones and produce vigorous sectors from slow-growing colonies. Alternatively, a high number of conidia can be plated on fungicide-amended media, and growing colonies can be singly transferred to fresh media. For instance, in B. cinerea it is relatively easy to get spontaneous mutants resistant to dicarboximides, phenylpyrroles, anilinopyrimidines and QoIs (Faretra and Pollastro 1991, 1993a, b; De Miccolis Angelini et al. 2002, 2012a).
Mutations can be induced by exposing conidia or hyphae to chemical (e.g. N-methyl-N-nitro-n-nitrosoguanidine) or physical mutagens (e.g. UV light) causing established proportions of lethality, before incubation on selective media. Mutagenesis greatly increases the yield of mutants but may cause unwished mutations in the genome which may interfere with identification and analysis of gene(s) causing fungicide resistance. Physical or chemical mutagenesis have been used successfully to produce resistant mutants in numerous fungi, including B. cinerea (De Miccolis Angelini et al. 2002, 2010a, b, 2012a, b), F. graminearum NRRL 13383 (Becher et al. 2010), P. capsici and P. infestans (Young et al. 2001), Ustilago maydis (Orth et al. 1994) and V. inaequalis (Zheng et al. 2000). In fungi with multinucleate conidia, laboratory mutants are frequently heterokaryons containing both mutated and wild-type nuclei so that they are often phenotypically unstable and produce both resistant and wild-type progeny during subculturing. Hence, at least initially, the selective pressure exerted by fungicide is crucial for the stability of the resistance trait.
The availability of molecular techniques has made it possible to investigate the genomes of pathogenic fungi which are not amenable to classical Mendelian analysis of meiotic progeny. For instance, genetic differences between isolates can be detected by RFLP (restriction fragment length polymorphisms) or various PCR-based techniques suitable for evidencing SNPs (single nucleotide polymorphisms) and allelic variants (AS-PCR, allele-specific PCR). Individual genes can be dissected out of the genome, then cloned and sequenced or altered genetically and put back into the genome. Cells containing cloned genes can be used to obtain large amounts of protein for amino acid sequencing. In the last fifteen years, site-direct mutagenesis has been used for studies of fungicide resistance. For instance, N. crassa mutants in the osmosensing histidine kinase os-1 gene exhibit resistance to dicarboximides, aromatic hydrocarbons and phenylpyrroles. The os-1 mutants can be classified into two groups: type I are null mutants highly resistant to iprodione and fludioxonil and moderately sensitive to osmotic stress, and type II carry single amino acid changes and are moderately resistant to both fungicides and highly sensitive to osmotic stress. This suggests that Os1p is essential for the antifungal activity of these fungicides and that amino acid repeats have an important function in osmoregulation (Ochiai et al. 2001). Site-directed mutagenesis followed by gene replacement was used to introduce mutations in different codons of the β-tubulin gene of a carbendazim-sensitive field strain of Gibberella zeae. All the mutants were resistant to carbendazim, but the level of resistance was depending on the mutations (Qiu et al. 2011). Site-directed mutagenesis of the SdhB gene was applied to confirm that each of the mutations identified in field strains conferred resistance to boscalid in B. cinerea and partial cross resistance to other SDHIs (fluopyram, carboxin) (Lalève et al. 2014b).
References
Anderson JB, Sirjusingh C, Ricker N (2004) Haploidy, diploidy and evolution of antifungal drug resistance in Saccharomyces cerevisiae. Genetics 168:1915–1923
Avenot H, Simoneau P, Iacomi-Vasilescu B, Bataillé-Simoneau N (2005) Characterization of mutations in the two-component histidine kinase gene AbNIK1 from Alternaria brassicicola that confer high dicarboximide and phenylpyrrole resistance. Curr Genet 47:234–243
Avila-Adame C, Köller W (2003) Characterization of spontaneous mutants of Magnaporthe grisea expressing stable resistance to the Qo-inhibiting fungicide azoxystrobin. Curr Genet 42:332–338
Becher R, Hettwer U, Karlovsky P, Deising HB, Wirsel SGR (2010) Adaptation of Fusarium graminearum to tebuconazole yielded descendants diverging for levels of fitness, fungicide resistance, virulence, and mycotoxin production. Phytopathology 100:444–453
Bertier L, Leus L, D’hondt L, de Cock AWAM, Höfte M (2013) Host adaptation and speciation through hybridization and polyploidy in Phytophthora. PLoS ONE 8, e85385
Bi Y, Chen L, Cai M, Zhu S, Pang Z, Liu X (2014) Two non-target recessive genes confer resistance to the anti-oomycete microtubule inhibitor zoxamide in Phytophthora capsici. PLoS ONE 9, e89336
Birky CW Jr (2001) The inheritance of genes in mitochondria and chloroplasts: laws, mechanisms, and models. Annu Rev Genet 35:125–148
Blatter RHE, Brown JKM, Wolfe MS (1998) Genetic control of the resistance of Erysiphe graminis f.sp. hordei to five triazole fungicides. Plant Pathol 47:570–579
Blum M, Gisi U (2008) Inheritance of resistance in Plasmopara viticola. In: Dehne HW, Gisi U, Kuck KH, Russell PE, Lyr H (eds) Modern fungicides and antifungal compounds V, BCPC, DPG, Braunschweig, Germany, pp 101–104
Blum M, Waldner M, Gisi U (2010) A single point mutation in the novel PvCesA3 gene confers resistance to the carboxylic acid amide fungicide mandipropamid in Plasmopara viticola. Fungal Genet Biol 47:499–510
Blum M, Gamper HA, Waldner M, Sierotzki H, Gisi U (2012) The cellulose synthase 3 (CesA3) gene of oomycetes: structure, phylogeny and influence on sensitivity to carboxylic acid amide (CAA) fungicides. Fungal Biol 116:529–542
Borck K, Braymer HD (1974) The genetic analysis of resistance to benomyl in Neurospora crassa. J Gen Microbiol 85:51–56
Brent KJ, Hollomon DW (2007a) Fungicide resistance in crop pathogens: how can it be managed? FRAC monograph no. 1. Global Crop Protection Federation, Brussels
Brent KJ, Hollomon DW (2007b) Fungicide resistance: the assessment of risk. FRAC monograph no. 2. Global Crop Protection Federation, Brussels
Brown AWA (1977) Epilogue: resistance as a factor in pesticide management. In: Mattson WJ (ed) The role or arthropods in forest ecosystems. Proceedings, 15th international congress of entomology, Washington, DC, 19–27 August 1976. Springer, New York
Brown JKM, Jessop AC, Thomas S, Rezanoor HN (1992) Genetic control of the response of Erysiphe graminis f.sp. hordei to ethirimol and triadimenol. Plant Pathol 41:126–135
Camps SM, Rijs AJ, Klaassen CH, Meis JF, O’Gorman CM, Dyer PS, Melchers WJ, Verweij PE (2012) Molecular epidemiology of Aspergillus fumigatus isolates harboring the TR34/L98H azole resistance mechanism. J Clin Microbiol 50:2674–2680
Catal M, King L, Tumbalam P, Wiriyajitsomboon P, Kirk WW, Adams GC (2010) Heterokaryotic nuclear conditions and a heterogeneous nuclear population are observed by flow cytometry in Phytophthora infestans. Cytometry A 77:769–775
Chapeland F, Fritz R, Lanen C, Gredt M, Leroux P (1999) Inheritance and mechanisms of resistance to anilinopyrimidine fungicides in Botrytis cinerea (Botryotinia fuckeliana). Pestic Biochem Physiol 64:85–100
Chin KM, Chavaillaz D, Kaesbohrer M, Staub T, Felsenstein FG (2001) Characterizing resistance risk of Erysiphe graminis f.sp. tritici to strobilurins. Crop Prot 20:87–96
Cookson B (2005) Clinical significance of emergence of bacterial antimicrobial resistance in the hospital environment. J Appl Microbiol 99:989–996
Cools HJ, Fraaije BA (2013) Update on mechanisms of azole resistance in Mycosphaerella graminicola and implications for future control. Pest Manag Sci 69:150–155
Coplin D (1989) Plasmids and their role in evolution of plant pathogenic bacteria. Annu Rev Phytopathol 27:187–212
Cowen LE, Anderson JB, Kohn LM (2002) Evolution of drug resistance in Candida albicans. Annu Rev Microbiol 56:139–165
Crute IR, Harrison JM (1988) Studies on the inheritance of resistance to metalaxyl in Bremia lactucae and on the stability and fitness of field isolates. Plant Pathol 37:231–250
Cui W, Beever RE, Parkes SL, Weeds PL, Templeton MD (2002) An osmosensing histidine kinase mediates dicarboximide fungicide resistance in Botryotinia fuckeliana (Botrytis cinerea). Fungal Genet Biol 36:187–198
Davis RH (1966) Mechanisms of inheritance. 2. Heterokaryosis. In: Ainsworth GC, Sussman AS (eds) The fungi, an advanced treatise, vol 2, The fungal organism. Academic Press, New York, pp 567–588
De Miccolis Angelini RM, Santomauro A, De Guido MA, Pollastro S, Faretra F (2002) Genetics of anilinopyrimidine-resistance in Botryotinia fuckeliana (Botrytis cinerea). In: Abstract Book of the 6th European Conference on Fungal Genetics, Pisa, Italy, 6–9 April 2002
De Guido MA, De Miccolis Angelini RM, Pollastro S, Santomauro A, Faretra F (2007) Selection and genetic analysis of laboratory mutants of Botryotinia fuckeliana resistant to fenhexamid. J Plant Pathol 89:203–210
De Miccolis Angelini RM, Habib W, Rotolo C, Pollastro S, Faretra F (2010a) Selection, characterization and genetic analysis of laboratory mutants of Botryotinia fuckeliana (Botrytis cinerea) resistant to the fungicide boscalid. Eur J Plant Pathol 128:185–199
De Miccolis Angelini RM, Rotolo C, Pollastro S, Faretra F (2010b) Phenotypic and molecular characterization of fungicide-resistant field isolates of Botryotinia fuckeliana (Botrytis cinerea). In: Abstracts of the XV international botrytis symposium, Càdiz, Spain, 30 May–4 June 2010
De Miccolis Angelini RM, Rotolo C, Masiello M, Pollastro S, Ishii H, Faretra F (2012a) Genetic analysis and molecular characterisation of laboratory and field mutants of Botryotinia fuckeliana (Botrytis cinerea) resistant to QoI fungicides. Pest Manag Sci 68:1231–1240
De Miccolis Angelini RM, Pollastro S, Faretra F (2012b) Genetics of fungicide resistance in Botryotinia fuckeliana (Botrytis cinerea). In: Thind TS (ed) Fungicide resistance in crop protection: risk and management. CAB International, Wallingford, pp 237–250
Delp CJ, Dekker J (1985) Fungicide resistance: definitions and use of terms. Bull OEPP 15:333–335
Délye C, Laigret F, Corio-Costet MF (1997) A mutation in the 14 alpha-demethylase gene of Uncinula necator that correlates with resistance to a sterol biosynthesis inhibitor. Appl Environ Microbiol 63:2966–2970
D’Mello JP, Macdonald AM, Postel D, Hunter EA (1997) 3-Acetyl deoxynivalenol production in a strain of Fusarium culmorum insensitive to the fungicide difenoconazole. Mycotoxin Res 13:73–80
D’Mello JPF, MacDonald AMC, Postel D, Dijksma WTP, Dujardin A, Plactina CM (1998) Pesticide use and mycotoxin production in Fusarium and Aspergillus phytopathogens. Eur J Plant Pathol 104:741–751
D’Mello JP, Macdonald AM, Briere L (2000) Mycotoxin production in a carbendazim-resistant strain of Fusarium sporotrichioides. Mycotoxin Res 16:101–111
Doukas EG, Markoglou AN, Ziogas BN (2008) Biochemical and molecular study of triazole-resistance and its effect on ochratoxin production by Aspergillus ochraceus Wilh. In: Abstracts of the 9th international congress of plant pathology. ICPP, Torino, Italy, 24–29 August 2008. J Plant Pathol 90:S2.319
Dry IB, Yuan KH, Hutton DG (2004) Dicarboximide resistance in field isolates of Alternaria alternata is mediated by a mutation in a two-component histidine kinase gene. Fungal Genet Biol 41:102–108
Faretra F, Pollastro S (1991) Genetic basis of resistance to benzimidazole and dicarboximide fungicides in Botryotinia fuckeliana (Botrytis cinerea). Mycol Res 95:943–951
Faretra F, Pollastro S (1993a) Genetics of sexual compatibility and resistance to benzimidazole and dicarboximide fungicides in isolates of Botryotinia fuckeliana (Botrytis cinerea) from nine countries. Plant Pathol 42:48–57
Faretra F, Pollastro S (1993b) Isolation, characterization and genetic analysis of laboratory mutants of Botryotinia fuckeliana (Botrytis cinerea) resistant to the phenylpyrrole fungicide CGA 173506. Mycol Res 97:620–624
Fernández-Ortuño D, Torés JA, de Vicente A, Pérez-García A (2008) Mechanisms of resistance to QoI fungicides in phytopathogenic fungi. Int Microbiol 11:1–9
Fillinger S, Ajouz S, Nicot PC, Leroux P, Bardin M (2012) Functional and structural comparison of pyrrolnitrin- and iprodione-induced modifications in the class III histidine-kinase Bos1 of Botrytis cinerea. PLoS ONE 7, e42520
Fillinger S, Leroux P, Auclair C, Barreau C, Al Hajj C, Debieu D (2008) Genetic analysis of fenhexamid-resistant field isolates of the phytopathogenic fungus Botrytis cinerea. Antimicrob Agents Chemother 52:3933–3940
Fincham JRS, Day PR, Radford A (1979) Fungal genetics, 4th edn. University of California Press, Berkeley, p 636
Georgopoulos SG (1988) Genetics and population dynamics. In: Delp CJ (ed) Fungicide resistance in North America. APS Press, St. Paul, pp 12–13
Gisi U (2012) Resistance to carboxylic acid amide (CAA) fungicides and anti-resistance strategies. In: Thind TS (ed) Fungicide resistance in crop protection: risk and management. CAB International, Wallingford, pp 96–103
Gisi U, Cohen Y (1996) Resistance to phenylamide fungicides: a case study with Phytophthora infestans involving mating type and race structure. Annu Rev Phytopathol 43:549–72
Gisi U, Sierotzki H (2008) Fungicide modes of action and resistance in downy mildews. Eur J Plant Pathol 122:157–167
Gisi U, Sierotzki H, Cook A, McCaffery A (2002) Mechanisms influencing the evolution of resistance to Qo inhibitor fungicides. Pest Manag Sci 58:859–867
Gisi U, Waldner M, Kraus N, Dubuis PH, Sierotzki H (2007) Inheritance of resistance to carboxylic acid amide (CAA) fungicides in Plasmopara viticola. Plant Pathol 56:199–208
Grasso V, Palermo S, Sierotzki H, Garibaldi A, Gisi U (2006) Cytochrome b gene structure and consequences for resistance to Qo inhibitor fungicides in plant pathogens. Pest Manag Sci 62:465–472
Griffiths AJF (1996) Mitochondrial inheritance in filamentous fungi. J Genet 75:403–414
Grindle M (1987) Genetics of fungicide resistance. In: Ford MG, Hollomon DH, Khambay BPS, Sawiki RM (eds) Combating resistance to xenobiotics – biological and chemical approaches. Ellis Horwood, Chichester, pp 75–93
Grindle M, Dolderson GH (1986) Effects of a modifier gene on the phenotype of a dicarboximide-resistant mutant of Neurospora crassa. Trans Br Mycol Soc 87:457–460
Grindle M, Faretra F (1993) Genetic aspects of fungicide resistance. In: Lyr H, Polter C (eds) Modern fungicides and antifungal compounds. Proceedings of the 10th international symposium, Reinhardsbrunn, Germany, May 1992. Ulmer, Stuttgart, Germany, pp 33–43
Guerineau M, Slonimski PP, Avner PR (1974) Yeast episome: oligomycin resistance associated with a small covalently closed non-mitochondrial circular DNA. Biochem Biophys Res Commun 61:462–469
Hawkins NJ, Cools HJ, Sierotzki H, Shaw MW, Knogge W, Kelly SL, Kelly DE, Fraaije BA (2014) Paralog re-emergence: a novel, historically contingent mechanism in the evolution of antimicrobial resistance. Mol Biol Evol 31:1793–1802
Heaney SP, Hall AA, Davies SA, Olaya G (2000). Resistance to fungicides in the QoI-STAR cross-resistance group: current perspectives. In: Proceedings of the British crop protection conference: pests and diseases, 13–16 November 2002, vol 2. BCPC, Farnham, Surrey, UK, pp 755–762
Hermann D, Gisi U (2012) Fungicide resistance in oomycetes with special reference to Phytophthora infestans and phenylamides. In: Thind TS (ed) Fungicide resistance in crop protection: risk and management. CAB International, Wallingford, pp 133–140
Hermisson J, Pennings PS (2005) Soft sweeps: molecular population genetics of adaptation from standing genetic variation. Genetics 169:2335–2352
Hilber UW, Hilber-Bodmer M (1998) Genetic basis and monitoring of resistance of Botryotinia fuckeliana to anilinopyrimidines. Plant Dis 82:496–500
Hobbelen PH, Paveley ND, van den Bosch F (2014) The emergence of resistance to fungicides. PLoS One 9, e91910
Hollomon DW (1981) Genetic control of ethirimol resistance in a natural population of Erysiphe graminis f.sp. hordei. Phytopathology 71:536–540
Hollomon DW, Butters J, Clark J (1984) Genetic control of triadimenol resistance in barley powdery mildew. In: Proceedings of the British crop protection conference: pests and diseases, 19–22 November 1984, vol 2. BCPC, Croydon, UK, pp 477–482
Hollomon DW, Butters JA, Kendall SJ (1997) Mechanism of resistance to fungicides. In: Sjut V (ed) Molecular mechanisms of resistance to agrochemicals, vol 13, Chemistry of plant protection. Springer, Heidelberg, pp 1–20
Ishii H (2010) QoI fungicide resistance: current status and the problems associated with DNA-based monitoring. In: Gisi U, Chet I, Gullino ML (eds) Recent developments in management of plant diseases, plant pathology in the 21st century, vol 1. Springer, Dordrecht, pp 37–45
Ishii H (2012a) Resistance to QoI and SDHI fungicides in Japan. In: Thind TS (ed) Fungicide resistance in crop protection: risk and management. CAB International, Wallingford, pp 237–250
Ishii H (2012b) Resistance in Venturia nashicola to benzimidazoles and sterol demethylation inhibitors. In: Thind TS (ed) Fungicide resistance in crop protection: risk and management. CAB International, Wallingford, pp 21–31
Ishii H, Yano K, Date H, Furuta A, Sagehashi Y, Yamaguchi T, Sugiyama T, Nishimura K, Hasama W (2007) Molecular characterization and diagnosis of QoI resistance in cucumber and eggplant fungal pathogens. Phytopathology 97:1458–1466
Ishii H, Fountaine J, Chung WH, Kansako M, Nishimura K, Takahashi K, Oshima M (2009) Characterization of QoI resistant field isolates of Botrytis cinerea from citrus and strawberry. Pest Manag Sci 65:916–922
Kalamarakis AE, de Waard MA, Ziogas BN, Georgopoulos SG (1991) Resistance to fenarimol in Nectria haematococca var. cucurbitae. Pestic Biochem Physiol 40:212–220
Kappas A, Georgopoulos SG (1970) Genetic analysis of dodine resistance in Nectria haematococca (Syn. Hypomyces solani). Genetics 66:617–622
Karaoglanidis GS, Markoglou AN, Bardas GA, Doukas EG, Konstantinou S, Kalampokis JF (2011) Sensitivity of Penicillium expansum field isolates to tebuconazole, iprodione, fludioxonil and cyprodinil and characterization of fitness parameters and patulin production. Int J Food Microbiol 145:195–204
Katan T, Shabi E (1982) Characterization of a dicarboximide-fungicide-resistant laboratory isolate of Monilinia laxa. Phytoparasitica 10:241–245
Kim YS, Dixon EW, Vincelli P, Farman ML (2003) Field resistance to strobilurin (QoI) fungicides in Pyricularia grisea caused by mutations in the mitochondrial cytochrome b gene. Phytopathology 93:891–900
Knapova G, Schlenzig A, Gisi U (2002) Crosses between isolates of Phytophthora infestans from potato and tomato and characterization of F1 and F2 progeny for phenotypic and molecular markers. Plant Pathol 51:698–709
Kretschmer M (2012) Emergence of multi-drug resistance in fungal pathogens: a potential threat to fungicide performance in agriculture. In: Thind TS (ed) Fungicide resistance in crop protection: risk and management. CAB International, Wallingford, pp 251–267
Kretschmer M, Leroch M, Mosbach A, Walker AS, Fillinger S, Mernke D, Schoonbeek HJ, Pradier JM, Leroux P, De Waard MA, Hahn M (2009) Fungicide-driven evolution and molecular basis of multidrug resistance in field populations of the grey mould fungus Botrytis cinerea. PLoS Pathog 5, e1000696
Lalève A, Fillinger S, Walker AS (2014a) Fitness measurement reveals contrasting costs in homologous recombinant mutants of Botrytis cinerea resistant to succinate dehydrogenase inhibitors. Fungal Genet Biol 67:24–36
Lalève A, Gamet S, Walker AS, Debieu D, Toquin V, Fillinger S (2014b) Site-directed mutagenesis of the P225, N230 and H272 residues of succinate dehydrogenase subunit B from Botrytis cinerea highlights different roles in enzyme activity and inhibitor binding. Environ Microbiol 16:2253–2266
Lasseron-De Falandre A, Daboussi MJ, Leroux P (1991) Inheritance of resistance to fenpropimorph and terbinafine, two sterol biosynthesis inhibitors, in Nectria haematococca. Phytopathology 81:1432–1438
Leroux P, Fritz R, Debieu D, Albertini C, Lanen C, Bach J, Gredt M, Chapeland F (2002) Mechanisms of resistance to fungicides in field strains of Botrytis cinerea. Pest Manag Sci 58:876–888
Lesemann SS, Schimpke S, Dunemann F, Deising HB (2006) Mitochondrial heteroplasmy for the cytochrome b gene controls the level of strobilurin resistance in the apple powdery mildew fungus Podosphaera leucotricha (Ell. & Ev.) E.S. Salmon. J Plant Dis Prot 113:259–266
Liu Y, Liu Z, Hamada MS, Yin YN, Ma ZH (2014) Characterization of laboratory pyrimethanil-resistant mutants of Aspergillus flavus from groundnut in China. Crop Prot 60:5–8
Ma Z, Michailides TJ (2005) Advances in understanding molecular mechanisms of fungicide resistance and molecular detection of resistant genotypes in phytopathogenic fungi. Crop Protect 24:853–863
Malandrakis AA, Markoglou AN, Nikou DC, Vontas JG, Ziogas BN (2006) Biological and molecular characterization of laboratory mutants of Cercospora beticola resistant to Qo inhibitors. Eur J Plant Pathol 116:155–166
Markoglou AN, Ziogas BN (1999) Genetic control of resistance to fenpropimorph in Ustilago maydis. Plant Pathol 48:521–530
Markoglou AN, Ziogas BN (2000) Genetic control of resistance to tridemorph in Ustilago maydis. Phytoparasitica 28:349–360
Markoglou AN, Ziogas BN (2001) Genetic control of resistance to the piperidine fungicide fenpropidin in Ustilago maydis. J Phytopathol 149:551–559
Markoglou AN, Malandrakis AA, Vitoratos AG, Ziogas BN (2006) Characterization of laboratory mutants of Botrytis cinerea resistant to QoI fungicides. Eur J Plant Pathol 115:149–162
Markoglou AN, Doukas EG, Ziogas BN (2008a) Phenylpyrrole-resistance and aflatoxin production in Aspergillus parasiticus Speare. Int J Food Microbiol 127:268–275
Markoglou AN, Vattis K, Dimitriadis K, Doukas EG, Ziogas BN (2008b) Effect of phenylpyrrole resistance mutations on mycotoxin production by Aspergillus carbonarius and Penicillium expansum. In: Abstracts of the 9th international congress of plant pathology. ICPP, Torino, Italy, pp 24–29 August 2008. J Plant Pathol 90:S2.322
Markoglou AN, Vitoratos AG, Doukas EG, Ziogas BN (2009) Phytopathogenic and mycotoxigenic characterization of laboratory mutant strains of Fusarium verticillioides resistant to demethylation inhibiting fungicides. In: Proceedings of the DPG-BCPC 3rd international symposium-plant protection and plant health in Europe, Berlin, Germany, 14–16 May 2009
Meyer RW, Parmeter JR (1968) Changes in chemical tolerance associated with heterokaryosis in Thanatephorus cucumeris. Phytopathology 58:472–475
Miguez M, Reeve C, Wood PM, Hollomon DW (2004) Alternative oxidase reduces the sensitivity of Mycosphaerella graminicola to QoI fungicides. Pest Manag Sci 60:3–7
Milgroom MG, Levln SA, Fry WE (1989) Population genetics theory and fungicide resistance. In: Leonard KJ, Fry WE (eds) Plant disease epidemiology: genetics, resistance, and management. McGraw-Hill Publishing Company, New York, pp 340–367
Molnar A, Hornok L, Pesti M (1985) The high level of benomyl tolerance in Fusarium oxysporum is determined by the synergistic interaction of two genes. Exp Mycol 9:326–333
Nakazawa Y, Yamada M (1997) Chemical control of grey mould in Japan. A history of combating resistance. Agrochem Japan 71:2–6
Ochiai N, Fujimura M, Motoyama T, Ichiishi A, Usami R, Horikoshi K, Yamaguchi I (2001) Characterization of mutations in the two-component histidine kinase gene that confer fludioxonil resistance and osmotic sensitivity in the os-1 mutants of Neurospora crassa. Pest Manag Sci 57:437–442
Ogden JE, Grindle M (1983) Changes in genetic constitution and sterol composition during growth of nystatin-resistant heterokaryons of Neurospora crassa. Genet Res 42:91–103
Orth AB, Sfarra A, Pell EJ, Tien M (1994) Characterization and genetic analysis of laboratory mutants of Ustilago maydis resistant to dicarboximide and aromatic hydrocarbon fungicides. Phytopathology 84:1210–1214
Oshima M, Fujimura M, Banno S, Hashimoto C, Motoyama T, Ichiishi A, Yamaguchi I (2002) A point mutation in the two-component histidine kinase BcOS-1 gene confers dicarboximide resistance in field isolates of Botrytis cinerea. Phytopathology 92:75–80
Otto SP, Gerstein AC (2008) The evolution of haploidy and diploidy. Curr Biol 18:R1121–4
Pang Z, Shao J, Chen L, Lu X, Hu J, Qin Z, Liu X (2013) Resistance to the novel fungicide pyrimorph in Phytophthora capsici: risk assessment and detection of point mutations in CesA3 that confer resistance. PLoS One 8, e56513
Peever TL, Milgroom MG (1992) Inheritance of triadimenol resistance in Pyrenophora teres. Phytopathology 82:821–828
Pollastro S, Faretra F, Santomauro A, Miazzi M, Natale P (1996) Studies on pleiotropic effects of mating type, benzimidazole-resistance and dicarboximide-resistance genes in near-isogenic strains of Botryotinia fuckeliana (Botrytis cinerea). Phytopathol Medit 35:48–57
Qiu J, Xu J, Yu J, Bi C, Chen C, Zhou M (2011) Localisation of the benzimidazole fungicide binding site of Gibberella zeae β2-tubulin studied by site-directed mutagenesis. Pest Manag Sci 67:191–198
Robinson HL, Riduct CJ, Sierotzki H, Gisi U, Brown JKM (2002) Isogamous, hermaphroditic inheritance of mitochondrion-encoded resistance to Qo inhibitor fungicides in Blumeria graminis f.sp. tritici. Fungal Genet Biol 36:98–106
Rumbou A, Gessler C (2006) Particular structure of Plasmopara viticola populations evolved under Greek island conditions. Phytopathology 96:501–509
Santomauro A, Pollastro S, De Guido MA, De Miccolis Angelini RM, Natale P, Faretra F (2000) A long-term trial on the effectiveness of new fungicides against grey mould on grapevine and on their influence on the pathogen’s population. In: Abstract Book of the XII international botrytis symposium. Reims, France, p P75
Shattock RC (1988) Studies on the inheritance of resistance to metalaxyl in Phytophthora infestans. Plant Pathol 37:4–11
Sierotzki H, Scalliet G (2013) A review of current knowledge of resistance aspects for the next-generation succinate dehydrogenase inhibitor fungicides. Phytopathology 103:880–887
Sierotzki H, Frey R, Wullschleger J, Palermo S, Karlin S, Godwin J, Gisi U (2007) Cytochrome b gene sequence and structure of Pyrenophora teres and P. tritici-repentis and implications for QoI resistance. Pest Manag Sci 63:225–33
Skinner W, Bailey A, Renwick A, Keon J, Gurr S, Hargreaves J (1998) A single amino-acid substitution in the iron-sulphur protein subunit of succinate dehydrogenase determines resistance to carboxin in Mycosphaerella graminicola. Curr Genet 34:393–398
Skylakakis G (1987) Changes in the composition of pathogen populations caused by resistance to fungicides. In: Wolfe MS, Caten CE (eds) Populations of plant pathogens: their dynamics and genetics. Blackwell, Oxford, pp 227–237
Steffens JJ, Pell EJ, Tien M (1996) Mechanisms of fungicide resistance in phytopathogenic fungi. Curr Opin Biotechnol 7:348–355
Steinfeld U, Sierotzki H, Parisi S, Gisi U (2002) Comparison of resistance mechanisms to strobilurin fungicides in Venturia inaequalis. In: Lyr H, Russell PE, Dehne HW, Gisi U, Kuck KH (eds) Modern fungicides and antifungal compounds III. Proceedings of the 13th international reinhardsbrunn symposium, Reinhardsbrunn, Germany, May 2001. AgroConcept GmbH, Bonn, Germany, pp 167–176
Summers RW, Heaney SP, Grindle M (1984) Studies of a dicarboximide resistant heterokaryon of Botrytis cinerea. In: Proceedings, Brighton crop protection conference, Brighton, UK, pp 453–458
Taga M, Nakagawa H, Tsuda M, Ueyama A (1979) Identification of three different loci controlling kasugamycin resistance in Pyricularia oryzae. Phytopathology 69:463–466
Torriani SFF, Brunner PC, McDonald BA, Sierotzki H (2009) QoI resistance emerged independently at least 4 times in European populations of Mycosphaerella graminicola. Pest Manag Sci 65:155–162
van Tuyl JM (1977) Genetic aspects of resistance to imazalil in Aspergillus nidulans. Neth J Plant Path 83:169–176
Veloukas T, Kalogeropoulou P, Markoglou AN, Karaoglanidis GS (2014) Fitness and competitive ability of Botrytis cinerea field isolates with dual resistance to SDHI and QoI fungicides, associated with several sdhB and the cytb G143A mutations. Phytopathology 104:347–356
Wolfe MS (1982) Dynamics of the pathogen population in relation to fungicide resistance. In: Dekker J, Georgopoulos SG (eds) Fungicide resistance in crop protection. Centre for Agricultural Publishing and Documentation, Wageningen, pp 139–148
Yarden O, Katan T (1993) Mutations leading to substitutions at amino acids 198 and 200 of beta-tubulin that correlate with benomyl-resistance phenotypes of field strains of Botrytis cinerea. Phytopathology 83:1478–1483
Yoshimi A, Imanshi J, Gafur A, Tanaka C, Tsuda M (2003) Characterisation and genetic analysis of laboratory mutants of Cochliobolus heterostrophus resistant to dicarboximide and phenylpyrrole fungicides. J Gen Plant Pathol 69:101–108
Young DH, Spiewak SL, Slawecki RA (2001) Laboratory studies to assess the risk of development of resistance to zoxamide. Pest Manag Sci 57:1081–1087
Young DH, Kemmitt GM, Owen J (2005) A comparative study of XR-539 and other oomycete fungicides: similarity to dimethomorph and amino acid amides in its mechanism of action. In: Dehne HW, Gisi U, Kuck KH, Russell PE, Lyr H (eds) Modern fungicides and antifungal compounds IV. BCPC, Alton, pp 145–52
Zhang Y, Lamm R, Pillonel C, Lam S, Xu JR (2002) Osmoregulation and fungicide resistance: the Neurospora crassa os-2 gene encodes a HOG1 mitogen-activated protein kinase homologue. Appl Environ Microbiol 68:532–538
Zhang YJ, Yu JJ, Zhang YN, Zhang X, Cheng CJ, Wang JX, Hollomon DW, Fan PS, Zhou MG (2009) Effect of carbendazim resistance on trichothecene production and aggressiveness of Fusarium graminearum. Mol Plant Microbe Interact 22:1143–1150
Zheng D, Olaya G, Köller W (2000) Characterization of laboratory mutants of Venturia inaequalis resistant to the strobilurin-related fungicide kresoxim-methyl. Curr Genet 38:148–155
Zhu SS, Liu XL, Wang Y, Wu XH, Liu PF, Li JQ, Yuan SK, Si NG (2007) Resistance of Pseudoperonospora cubensis to flumorph on cucumber in plastic houses. Plant Pathol 56:967–975
Ziogas BN, Markoglou AN, Tzima A (2002) A non-Mendelian inheritance of resistance to strobilurin fungicides in Ustilago maydis. Pest Manag Sci 58:908–916
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Japan
About this chapter
Cite this chapter
De Miccolis Angelini, R.M., Pollastro, S., Faretra, F. (2015). Genetics of Fungicide Resistance. In: Ishii, H., Hollomon, D. (eds) Fungicide Resistance in Plant Pathogens. Springer, Tokyo. https://doi.org/10.1007/978-4-431-55642-8_2
Download citation
DOI: https://doi.org/10.1007/978-4-431-55642-8_2
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-55641-1
Online ISBN: 978-4-431-55642-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)