Abstract
Macrophages are innate immune cells that sense the presence of pathogens through conserved pattern recognition receptors, which include TLR4. Activation of TLR4 by bacterial lipopolysaccharide induces the expression of thousands of genes that function to initiate inflammation and coordinate innate and adaptive immune responses. Transcriptional activation of TLR4-responsive genes is mediated by signal-dependent transcription factors, such as NFκB, which bind to DNA regulatory elements termed enhancers. Recent findings indicate that macrophage enhancers are actively transcribed in concert with nearby genes. Similar observations have been reported for other cell types, raising the general question of whether enhancer transcription and/or the resulting enhancer RNAs (eRNAs) are of functional importance. Here, we review the use of macrophage activation as an experimental system for addressing these questions and highlight areas for future research.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Macrophage
- Enhancer
- Promoter
- eRNA
- mRNA
- Transcription
- TLR
- Chromatin
- Histone methylation
- Histone acetylation
- Nucleosome
- NFkB
1 Macrophage Activation as a Model System
Macrophages are myeloid lineage cells, which play essential roles in innate and adaptive immune responses and contribute to diverse aspects of tissue homeostasis (Wynn et al. 2013). Importantly, many of the transcriptional programs required for appropriate responses to pathogens also contribute to the pathogenesis of numerous chronic inflammatory diseases, which include atherosclerosis, diabetes, arthritis, and cancer. The innate immune response to pathogens is triggered by the interaction of pathogen-associated molecular patterns with pattern recognition receptors, which include members of the toll-like receptor (TLR) family (Medzhitov and Horng 2009; Takeuchi and Akira 2010). TLR4 recognizes the lipopolysaccharide (LPS) component of gram-negative bacteria, and is arguably the most intensively studied pattern recognition receptor (Beutler 2000). TLR4 ligation results in activation of several latent, signal-dependent transcription factors, including NF-kappaB (NF-κB), interferon regulatory factors (IRFs), AP-1 factors, and STAT factors, which act in a combinatorial manner to both positively and negatively regulate the expression of thousands of genes (Medzhitov and Horng 2009; Smale 2012) (Fig. 12.1). This response has been intensively studied at the level of genomics (e.g., Escoubet-Lozach et al. 2011; Ghisletti et al. 2010; Kaikkonen et al. 2013; Ostuni et al. 2013), proteomics (e.g., Meissner et al. 2013), and lipidomics (e.g., Maurya et al. 2013) in macrophages.
TLR4-dependent gene expression. Toll-like receptor 4 (TLR4) is a pattern recognition receptor that is activated by lipopolysaccharide (LPS). Ligation of TLR4 results in transmission of an inflammatory signal through Myd88 and TRF, resulting in activation of latent, signal-dependent transcription factors, including NF-κB, interferon regulatory factors (IRFs), AP-1 factors, and STAT factors. These factors act in a combinatorial manner to both positively and negatively regulate the expression of thousands of genes
The pathogen response of the mouse macrophage provides a powerful system for applying genomics and associated modeling approaches to the understanding of how transcriptional regulatory elements are selected from the genome and enable complex cell-specific programs of gene expression. First, mouse macrophages are exquisitely sensitive to LPS and other TLR4-specific analogues, such as Kdo2 lipid A (KLA). In addition to the large number of genes affected by TLR4 signaling, the dynamic range of transcriptional activation and repression exceeds three orders of magnitude for the most highly regulated genes. The response has both immediate–early and secondary phases based on the initial actions of latent transcription factors, which drive expression of cytokines such as TNF, and type I interferons, which induce expression and/or activities of a subsequent set of transcription factors. Although some genes, such as TNF and IL1b, are activated by TLR4 signaling in many cell types, a substantial component of the transcriptional response to TLR4 ligation is cell type specific. Therefore, the TLR4 response provides a powerful means of investigating the basis by which the genome is read to result in a signal-dependent, temporally orchestrated, and cell-specific response at the level of transcription.
Second, features of mouse macrophages themselves make them highly tractable for performing genomics analysis. As primary cells, they do not have the caveats associated with genomic alterations frequently associated with cell lines. Upward of 30 million thioglycollate-elicited macrophages (TGEMs) can be obtained from a single mouse. Importantly, more than 96 % of plated TGEMs express defining macrophage markers such as CD14 and CD11b, indicating a high degree of homogeneity. The relatively straightforward method for isolation has enabled simultaneous examination of multiple experimental conditions using all currently available and relevant genomics technologies, including ChIP-Seq, RNA-Seq, GRO-Seq, 5′-GRO-Seq, and HiC assays. Importantly, total and macrophage-specific loss-of-function alleles are available for many genes of interest, and siRNA knockdowns work well in TGEMs (e.g., Escoubet-Lozach et al. 2011; Heinz et al. 2010, 2013; Kaikkonen et al. 2013; Lam et al. 2013).
2 Studying Enhancers with High-Throughput Sequencing
As a result of this proclivity toward high-throughput studies, mouse macrophages are well suited for the investigation of the genome-wide phenomena such as enhancer selection and activation. Enhancers are regions of the genome that are able to regulate transcription from a distance, acting to increase or decrease transcriptional activity at target genes by looping into close proximity with promoters (Ong and Corces 2011). Enhancers act as platforms for the binding of many different transcription factors, as well as for the recruitment of the Mediator complex, cohesin, acetyltransferases, and RNA polymerase II (Pol II), all of which cooperate to regulate transcription at gene promoters.
With the advent of high-throughput sequencing, the ability to locate and characterize enhancers increased tremendously. Crucially, Heintzman et al. (2007) described a particular pattern of histone methylation that acts as a signature for enhancers genome wide, making it possible to map the enhancer landscapes of a variety of cells and organisms by looking for regions with high enrichment of H3K4 mono- and di-methylation (H3K4me1/2) and low enrichment of H3K4 tri-methylation (H3K4me3) (Fig. 12.2). While this histone signature does not guarantee that a given region is a functional enhancer, subsequent studies have shown that the majority of functional enhancers are indeed H3K4me1-high and H3K4me3-low (Heintzman et al. 2007; Heinz et al. 2010; Rada-Iglesias et al. 2011).
Enhancer and promoter transcription. Enhancers (yellow boxes) are marked by mono- and dimethylation of H3K4, while gene promoters (green boxes) are marked by di- and trimethylation of H3K4. General transcription factors (GTFs) bind at both enhancers and promoters, whereas lineage-determining transcription factors (LDTFs) and signal-dependent transcription factors (SDTFs) bind primarily at enhancers, resulting in cell type-specific transcriptional activity. RNA polymerase II (Pol II) is recruited at both enhancers and promoters, generating mRNA along gene bodies and enhancer RNAs (eRNAs) at enhancers
This genome-wide mapping of enhancers associated with specific chromatin signatures has led to the recognition that enhancers are distinct from cell type to cell type, even when gene expression is not (Heintzman et al. 2009). Further, recent studies have indicated that the cell type specificity of enhancers drives cell type-specific gene expression (Heinz et al. 2010; Kaikkonen et al. 2013; Thurman et al. 2012; Visel et al. 2009). Investigations of how similar signaling cascades and transcription factors can establish distinct sets of enhancers in different cell types led to the formulation of a hierarchical model of enhancer selection and function. The hierarchical model relies on two classes of transcription factors—lineage-determining transcription factors (LDTFs) and signal-dependent transcription factors (SDTFs) (Garber et al. 2012; Mullen et al. 2011; Soufi et al. 2012; Trompouki et al. 2011). LDTFs are the relatively small set of transcription factors crucial to a particular lineage, and these factors collaborate in a combinatorial manner to compete with nucleosomes to bind DNA in a cell type-specific manner. The binding of these factors is proposed to establish open regions of chromatin that are associated with cis-active regulatory regions. SDTFs, which are activated in response to cell signals and are often shared across cell types, are then able to bind to the enhancers pre-established by LDTFs (Ghisletti et al. 2010; Heinz et al. 2010; Mullen et al. 2011; Trompouki et al. 2011). This multi-step model explains how SDTFs like NF-kappaB and the glucocorticoid receptor, which are widely expressed, can produce such different responses from cell type to cell type.
In the macrophage system, molecular and genetic studies of the process of differentiation from hematopoietic stem cells indicate essential roles of PU.1, C/EBPα/β and AP-1 family members as LDTFs (Heinz et al. 2010). In line with this, forced expression of PU.1 and C/EBPα in fibroblasts is sufficient to promote reprogramming to macrophage-like cells, albeit with low efficiency (Feng et al. 2008). Although PU.1, C/EBPs, and AP-1 factors function as LDTFs in macrophages, these lineage-determining functions are not exclusive to macrophages. For example, PU.1 is also a lineage-determining transcription factor for B cells (Klemsz et al. 1990; Scott et al. 1994), and C/EBPα is required for differentiation of other cell types, which include adipocytes (Cristancho and Lazar 2011; Herrera et al. 1989). Studies of the genome-wide locations of PU.1 in macrophages and B cells indicated cell-similar binding patterns at promoters but cell-specific binding patterns at distal inter- and intra-genic locations (Heinz et al. 2010). At these locations, PU.1 frequently co-localized with C/EBP and AP-1 factors in macrophages, and with B cell lineage-determining factors in B cells. Gain- and loss-of-function experiments demonstrated that co-binding of PU.1 with alternate lineage-determining factors was co-dependent. For example, loss of function of PU.1 resulted in loss of C/EBPα binding at sites where the two proteins co-bound within ~100 bp. Conversely, PU.1 binding at a subset of B cell-specific enhancers was dependent on co-expression and binding of early B cell factor (EBF). In contrast, the binding of PU.1 in macrophages was not dependent on the nearby binding of SDTFs, whereas the binding of SDTFs was dependent on the binding of PU.1 (Heinz et al. 2010). This co-dependence of LDTFs was shown to be in effect genome wide, as binding motifs for PU.1 that were disturbed by polymorphisms in multiple strains of inbred mice abrogated not just PU.1 binding but binding of C/EBP as well as the SDTF NF-kappaB (Heinz et al. 2013).
3 Transcription at Enhancers
In addition to a histone signature and transcription factor binding, recent studies in neurons and cancer cells revealed that widespread transcription characterizes enhancer regions. That is, in addition to driving transcription at gene promoters, enhancers themselves are transcribed. RNA Polymerase II (Pol II) was known to bind at several well-characterized enhancers including the beta-globin enhancer (Koch et al. 2008; Szutorisz et al. 2005), but these were thought to be idiosyncratic occurrences. However, Kim et al. showed widespread binding of Pol II to 12,000 neuronal enhancers marked by both H3K4me1 and the transcriptional co-activator CBP (Kim et al. 2010). High-throughput sequencing of total RNA showed that Pol II was active at these locations and generated nuclear bi-directional transcripts, called enhancer-RNAs (eRNAs), originating from the center of the CBP-identified enhancers. The identified eRNAs were further shown to positively correlate with expression of nearby genes, implying that eRNA synthesis occurs preferentially at active enhancers.
In parallel, De Santa et al. described the same phenomenon in macrophages subject to LPS stimulation, and found that 70 % of extragenic Pol II binding sites overlapped with histone marks indicative of enhancers (De Santa et al. 2010). The transcribed enhancers generated transcripts that were poly-adenylated but not spliced or exported from the nucleus. The eRNAs were also very low in abundance as compared with mRNAs; quickly induced, as eRNA synthesis preceded downstream gene synthesis at several loci investigated via RT-PCR; and transient, as they were highly susceptible to depletion via actinomycin D treatment.
This transience makes eRNAs difficult to capture with traditional RNA-Seq. However, the advent of global nuclear run-on sequencing (GRO-Seq) made possible the in-depth study of non-coding RNA kinetics across the whole genome (Core et al. 2008). Briefly, GRO-Seq takes advantage of a nuclear run-on reaction to tag nascent RNAs as they are assembled by Pol II. These tagged nascent transcripts are then sequenced, giving a real-time picture of transcription within the cell. Whereas RNA-Seq measures expression levels of stable, spliced RNA species, GRO-Seq returns data on rates of active transcription, of both coding and non-coding RNA species. Using GRO-Seq in coordination with the enhancer histone signature, it was estimated that ~18 % of transcripts in mouse macrophages were produced at enhancer-like regions, comprising almost a third of unannotated transcripts (Allison et al. 2014).
In MCF7 cells, Hah et al. used GRO-Seq to demonstrate extensive bidirectional transcription at enhancers (Hah et al. 2011), and showed further that the expression level of eRNAs was responsive to estrogen treatment. eRNAs in fact made up the largest class of transcripts that were initiated proximal to an estrogen receptor alpha binding site. Similarly, GRO-Seq was used in LNCaP cells to show widespread changes in enhancer transcription in response to FoxA1 and androgen receptor (AR) binding (Wang et al. 2011). As in Hah et al., expression levels in eRNAs were associated with transcription factor binding events and correlated with the expression levels of nearby genes.
The close association of eRNA production and gene expression levels raises the question of whether enhancer transcription or eRNAs themselves play a functional role. The act of transcription by Pol II has been associated with the conference of histone acetylation and the maintenance of open chromatin at extragenic regions (Travers 1999), as well as the deposition of methyl marks at histones (Gerber and Shilatifard 2003; Xiao et al. 2003), but initial reports of eRNAs did not assess the relevance of these effects at enhancers genome wide. In their study of LPS-treated macrophages, De Santa et al. found that transcription of an enhancer near the Ccl5 promoter was associated with increased acetylation in a manner sensitive to actinomycin D treatment (De Santa et al. 2010), but did not thoroughly establish the importance of transcription at this particular enhancer or enhancers generally in the recruitment of acetylation marks.
4 An Order of Events for Enhancer Transcription
Studies in primary mouse macrophages have helped elucidate the possible functions of enhancer transcription by taking advantage of the finding that TLR4 signaling induced the selection of thousands of “latent” or “de novo” enhancer-like regions in the genome. These de novo enhancers are defined by the new acquisition of H3K4me1 and/or H3K4me2 (Kaikkonen et al. 2013; Ostuni et al. 2013), marking enhancers that did not exist in the basal state but rather appeared upon cell activation. In the studies of Ostuni et al., “latent” enhancers were defined by (1) the lack of H3K4me1, H3K27Ac, and PU.1 in unstimulated cells; and (2) the presence of an LPS-induced H3K4me1 peak. There were ~500 such enhancers identified after 4 h of LPS stimulation, and ~1,000 after 24 h. The appearance of latent enhancers was not unique to LPS stimulation or TLR4 signaling, and several other stimuli, including TNFa and interleukin 1-beta (IL-1b), each induced a set of latent enhancers. Notably, Ostuni et al. did not find any histone mark or chromatin feature that allowed pre-identification of the latent enhancers in untreated conditions.
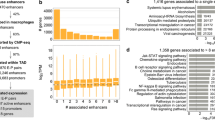
Kaikkonen et al. described a similar set of enhancers in macrophages by performing H3K4me2 ChIP-Seq of MNase-treated chromatin obtained following 0, 1, 6, 24, and 48 h of Kdo2-Lipid A (KLA) treatment (Kaikkonen et al. 2013). KLA, an LPS-analogue, induced ~32,000 inter- and intragenic locations marked by the enhancer histone signature prior to treatment, referred to as “pre-existing” enhancers. These regions were highly enriched for motifs recognized by PU.1, C/EBP, and AP-1 factors, consistent with previous findings (Ghisletti et al. 2010; Heinz et al. 2010), and were significantly correlated with the expression levels of nearby genes (Fig. 12.3). Notably, ~3,000 previously unmarked regions, termed “de novo” enhancers, gained H3K4me2 upon KLA stimulation. In contrast, ~1,000 regions lost this mark following KLA treatment. Gain and loss of H3K4me2 at enhancer-like regions was highly correlated with expression of nearby genes.
Relationship of changes in eRNA expression to changes in mRNA expression. Heat map illustrating the relationship between signal-dependent changes in eRNAs and corresponding changes in expression of the nearest gene. Each column represents the amount of nascent RNA detected at a specific genomic location as a function of time following treatment of macrophages with a TLR4 agonist. Columns in the bottom panel correspond to locations of signal-regulated enhancers. Columns in the upper panel correspond to genomic locations of the nearest mRNA encoding gene. Red indicates upregulation of nascent RNA, and green represents downregulation. The pattern illustrates that eRNA expression is significantly correlated with expression of the nearest mRNA in stimulated macrophages (Reproduced from Kaikkonen et al. 2013, with permission)
The emergence of de novo enhancers in response to a temporally precise signal allowed Kaikkonen et al. to investigate the sequence of events that occur as closed chromatin transitions to active enhancer-like regions. ChIP-Seq experiments for histone acetylation (H3K2, H4K5, H4K8, and H4K12); for the LDTFs PU.1 and C/EBPα; and for the p65 (RelA) component of the SDTF NF-kappaB as a function of time following KLA treatment were conducted in parallel with GRO-Seq to define nascent RNA transcripts as a measure of transcriptional output. These experiments resulted in several important observations relevant to the general hierarchical model. First, p65 binding was primarily observed to occur at pre-existing enhancers characterized by high enrichment for PU.1, C/EBP, and AP-1 motifs. Co-binding of PU.1 and C/EBPα was confirmed at a high percentage of these sites by ChIP-Seq studies, though an increase in histone acetylation was associated with binding of p65.
In contrast to pre-existing enhancers, motif analysis of de novo H3K4me2-marked regions returned AP-1 and NF-kappaB motifs as the most highly enriched sequence elements. Importantly, however, C/EBP and PU.1 were also highly enriched. Consistent with these findings, while PU.1 or C/EBPα binding was absent under basal conditions, these factors were recruited to a large fraction of the de novo enhancer-like regions concomitantly with p65 within 1 h of KLA treatment. Loss-of-function studies indicated that in addition to the dependence of PU.1 binding on the nuclear entry of p65, DNA binding of p65 was dependent on PU.1 at locations where the two factors bound to closely spaced motifs. Therefore, at these locations, p65 acted as both a signal-dependent and collaborative transcription factor to facilitate the selection of new functional enhancers.
The collaborative binding of p65, PU.1, and C/EBP at de novo enhancers was temporally linked to the acquisition of histone acetylation and the initiation of enhancer transcription. Unexpectedly, these events substantially preceded the acquisition of H3K4me1 and H3K4me2. In addition to this temporal relationship, the length of eRNA transcripts was highly correlated with the distribution of H3K4me1 and H3K4me2 (Kaikkonen et al. 2013), suggesting that enhancer transcription was linked to the writing of these marks. To investigate this possibility, Kaikkonen et al. assessed the effects of inhibiting Pol II elongation on the H3K4me2 status after KLA treatment. Two different elongation inhibitors were used: the cyclin-dependent kinase (cdk) inhibitor flavopiridol, which at low concentrations preferentially inhibits the Cdk9 activity of P-TEFb (Chao and Price 2001); and IBET151, a selective inhibitor of BET (bromodomain and extra terminal domain) protein binding to acetylated histones, which disrupts the recruitment of P-TEFb complexes to acetylated histones that are dependent on BRD4 (Dawson et al. 2011; Nicodeme et al. 2010). Both drugs affected the elongation of KLA-induced nascent transcripts as evidenced by a decrease in the cumulative GRO-Seq tags beyond the transcription start site (TSS), with the effect of flavopiridol being more pronounced. Inhibition of eRNA elongation by IBET151 and flavopiridol was correlated with a decrease in the deposition of H3K4me2 at ~40 % and ~70 % of de novo enhancers. The effectiveness of drug treatment on reducing eRNA expression at individual enhancers was significantly correlated with a corresponding local reduction of H3K4me1 and H3K4me2. Similar effects were observed with three other inhibitors of Pol II. KLA-induced gain in H3K4me2 observed at many pre-existing enhances was blocked by inhibition of Pol II elongation.
Collectively, these findings suggest that enhancer H3K4me1/2 deposition is coupled to enhancer transcription, at least for de novo enhancers. Further, Kaikkonen et al. demonstrated that transcription-coupled H3K4 methylation at de novo enhancers was mediated by members of the Mll family of histone methyltransferases. These findings are consistent with the ability of Mlls to associate with the phosphorylated C terminal domain (CTD) of Pol II (Hughes et al. 2004; Krogan et al. 2003; MacConaill et al. 2006; Milne et al. 2005; Ng et al. 2003; Rana et al. 2011; Wood et al. 2003) and suggest that the progressive accumulation of H3K4 methylation at de novo enhancers results from their association with active forms of Pol II (Fig. 12.4). Whether this mechanism accounts for deposition of H3K4 methylation at pre-existing enhancers remains to be established, but these findings provide evidence for one functional consequence of enhancer transcription.
Models for activation of pre-existing and de novo enhancers. (a) Pre-existing enhancers are bound in the basal state by lineage-determining transcription factors (LDTFs) and are marked by H3K4me1 and H3K4me2. Many of these pre-existing enhancers are transcribed at a low level by Pol II. (b) Upon initiation of an inflammatory signal (such as TLR4 stimulation with LPS), signal-dependent transcription factors (SDTFs) are recruited to the open chromatin at pre-existing enhancers. Co-activators such as histone acetyl transferases (HATs) are recruited to the enhancer with the SDTFs, resulting in increased Pol II activity. (c) Pol II transcription is associated with increased dimethylation of H3K4 at enhancers via the recruitment of methyltransferases. (d) In contrast to pre-existing enhancers, de novo enhancers lack LDTF binding and open chromatin in the basal state. (e) Upon stimulation, SDTFs and LDTFs are both recruited and coordinately open chromatin to form de novo enhancers. Co-activators and HATs acetylate surrounding histones. (f) The acetylated, bound de novo enhancers are then able to recruit Pol II and initiate transcription of eRNAs. Transcription is followed by recruitment of methyltransferases and the deposition of di-methyl at H3K4
5 Enhancer Transcription as a Marker of Activity
The close temporal relationship between enhancer transcription, H3K4 methyl deposition, and gene transcription raises the question of how to interpret eRNA with respect to the enhancer signature derived from histone marks. While high levels of H3K4me1 with respect to H3K4me3 are considered characteristic of enhancer-like regions in the genome (Heintzman et al. 2007), this combination is not necessarily associated with enhancer activity. In the studies of Ostuni et al. the H3K4me1 signature associated with “latent” enhancers selected in response to LPS stimulation persisted after the transcriptional response to LPS subsided. These regions were associated with a more rapid response to subsequent stimulation, suggesting that the H3K4me1 mark provided a molecular memory that facilitated the second response (Ostuni et al. 2013). Kaikkonen et al. observed a similar persistence of H3Kme2 at de novo enhancers even after the transcriptional response of nearby genes had largely returned to baseline levels (Kaikkonen et al. 2013). Therefore, while both H3K4me1 and H3K4me2 are associated with enhancers, they do not necessarily reflect enhancer activity.
In contrast, Wang et al. found that enhancer activation was tightly coupled to eRNA production, as the gain and loss of AR binding was most closely correlated with eRNA synthesis rather than histone mark deposition or even the presence of the histone acetyltransferase p300 (Wang et al. 2011). Similarly, eRNA production as measured by GRO-Seq was highly correlated with nascent RNA production at the nearest mRNA encoding gene throughout the entire KLA time course (Kaikkonen et al. 2013). Providing further evidence for eRNA as a marker of active enhancers, Wu et al. studied tissue-specific RNA expression in mouse embryonic tissues (Wu et al. 2014). According to deep total RNA-Seq, previously validated enhancers were extensively transcribed, and eRNA marked a larger set of active enhancers than either H3K27Ac or p300. The enhancers marked by eRNA alone were subsequently tested for their ability to activate a lacZ reporter gene, and 8 out of 19 tested enhancers drove reporter expression in the predicted tissue-specific manner. Further, eRNAs have proven to be very sensitive indicators of transcription factor binding and activity. In addition to the correlation between AR binding and eRNA production seen by Wang et al., hundreds of eRNAs were observed to be responsive to p53 binding in p53-competent as compared with p53-null cells (Allen et al. 2014), and the effects of rosiglitazone treatment on adipocytes could be closely tracked via changes in eRNA expression levels at PPARgamma binding sites (Step et al. 2014).
In order to systematically determine the relationship between enhancer transcription and enhancer activity, Zhu et al. built a logistic regression model using 24 histone marks and p300 assayed by ChIP-Seq in conjunction with GRO-Seq from IMR90 cells (Zhu et al. 2013). The model revealed that the histone mark most predictive of eRNA synthesis was acetylation at H3K27 (H3K27Ac), which had previously been shown to be a mark of active enhancers (Creyghton et al. 2010). Models based on four histone marks achieved the highest area under the curve (AUC) value for predicting enhancer transcription, with 432 combinations of the 24 histone marks yielding AUC values within the top 5 % of all possible combinations. In addition to H3K27Ac, the activation marks H3K79me1, H3K9Ac, and H4K8Ac were positively associated with eRNA production. On the other hand, the repressive mark H3K27me3 was predictive of eRNA production with a negative coefficient. Despite the correlation of these histone marks with enhancer transcription, Zhu et al. found that eRNA was the single most predictive indicator of enhancer activity, with eRNA synthesis being more significantly associated with increased expression at nearby genes than histone marks in multiple cell types. These results are supported by a separate study by Pulakanti et al. where it was found that eRNA synthesis correlated with H3K27Ac deposition, hypomethylation, and occupancy of the DNA hydroxylase Tet1 in embryonic stem cells, all of which are traditional markers for transcriptional activity (Pulakanti et al. 2013).
6 The Functionality of eRNAs
These findings suggest that enhancer function is in some way linked to enhancer transcription. To directly study this possibility, Lam et al. investigated the mechanisms by which the Rev-Erbα and Rev-Erbβ nuclear receptors functioned to repress gene expression in macrophages. RevErbα/β are atypical members of the nuclear receptor family in that they constitutively interact with NCoR/HDAC3 co-repressor complexes but are unable to interact with nuclear receptor co-activators. As a consequence, they function as active transcriptional repressors upon binding to Rev-Erb recognition motifs (Yin and Lazar 2005; Zamir et al. 1996). Genome-wide location analysis of biotin tagged (BLRP) Rev-Erbα and Rev-Erbβ in RAW macrophages indicated that >80 % of their high confidence binding sites in the genome were at enhancer-like regions characterized by high H3K4me1/low H3K4me3 and occupied by macrophage LDTFs including PU.1 (Lam et al. 2013). Evaluation of Rev-Erb target genes, such as Cx3cr1, in Rev-Erbα/β double knockout macrophages using GRO-Seq revealed significant increases not only in the Cx3cr1 mRNA but also a corresponding increase in eRNA production from an enhancer-like region 28 kb downstream of the Cx3cr1 TSS that is occupied by Rev-Erbβ. Using a method to measure nascent RNA at the site of transcriptional initiation, termed 5′-GRO-Seq, the major sites of initiation were observed at the Cx3cr1 promoter and at the −28 kb enhancer. Lam et al. found that initiation from both locations was greatly suppressed by overexpression of BLRP-Rev-Erbα, suggesting that the consequences of Rev-Erb binding to its enhancer are direct suppression of enhancer transcriptional initiation and indirect suppression of initiation from the Cx3cr1 promoter.
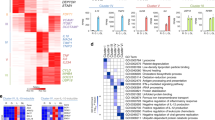
However, the importance of the initiation of transcription still left open the question of whether the transcripts generated at enhancers are themselves important for enhancer function independent of the process of transcription. In order to gain an understanding of the significance of the products of enhancer transcription, Lam et al. designed siRNAs and antisense oligonucleotides (ASOs) to target the plus and minus strands of Cx3cr1 and Mmp9 eRNAs. Both methods were capable of reducing eRNA transcript levels on the basis of quantitative PCR analysis of nuclear RNA (Lam et al. 2013). Notably, reduction of eRNA expression was associated with decreased expression of nearby, but not distant, genes. This is exemplified by the ASO knockdown experiment presented in Fig. 12.5. Using a combination of 5′-GRO-Seq and conventional GRO-Seq to define the origin and length of the Cx3cr1 minus strand eRNA, an overlapping series of ASOs was synthesized and tested for ability to knockdown eRNA expression. The majority of these ASOs reduced eRNA expression, with the effects of two of the most potent ASOs illustrated at the bottom of Fig. 12.5. Both ASOs significantly reduced Cx3cr1 eRNA expression and Cx3cr1 mRNA expression. In contrast, neither ASO significantly affected expression of distant genes, such as the Mmp9 and Csrnp1 genes. Thus, at least in the case of the selected Rev-Erb target enhancers, the production of eRNAs was important for associated gene expression levels.
Evidence for functional roles of eRNAs. (a) The top panel represents the Cx3cr1 28 kb distal enhancer and the experimental design for testing functional roles of eRNA. Shaded regions indicate locations of PU.1 binding, RevErb binding, and histone H3K4me1. Directional arrows represent eRNA transcription start sites defined by 5′-GRO-Seq. Antisense oligonucleotides (ASOs) were designed to tile eRNA generated along the minus strand of this enhancer region gene. (b) Two representative ASOs against the Cx3cr1 enhancer eRNA reduced expression of both the target eRNAs and the associated mRNA, as measured by qRT-PCR. The effect was specific to Cx3cr1 mRNA, such that Mmp9 mRNA and Csmp1 mRNA were unaffected by the ASOs (Adapted from Lam et al. 2013 with permission)
These results are supported by numerous studies outside macrophages. Melo et al. identified enhancers that produced eRNAs in a p53-dependent manner in immortalized human fibroblasts and MCF7 cells (Melo et al. 2013). The p53-bound enhancers were further shown to interact with distal p53-dependent gene promoters. Using siRNA, Melo et al. knocked down eRNAs at two p53 binding sites and demonstrated a concomitant loss of target mRNA as well as a reduction of Pol II at the promoters of the target genes. In primary human monocytes, Iiott et al. identified a set of LPS-inducible eRNAs whose expression correlated with that of nearby genes (NE et al. 2014). Many of these enhancers had NF-kappaB binding sites, suggesting they play an important role in transcriptional changes downstream of LPS stimulation via TLR4. To investigate the role of one particular enhancer near the highly induced IL1beta gene, Iiott et al. used locked nucleic acid (LNA)-based antisense inhibitors to suppress expression of the eRNA. The knockdown by LNAs (but notably not siRNAs) of the enhancer transcript resulted in reduced induction of IL1beta mRNA in response to LPS stimulation, demonstrating the importance of the eRNA at the IL1beta enhancer in the regulation of the target gene.
These loss-of-function studies demonstrated the importance of particular eRNAs in particular contexts but did not elucidate the mechanisms by which the eRNA was regulating target gene expression. One potential mechanism was elucidated by Li et al. in a study of estrogen receptor alpha (ER) binding in MCF7 cells (Li et al. 2013). Several enhancers that increased eRNA synthesis in response to E2 ligand treatment and subsequent ER binding were knocked down with siRNAs and LNAs. Induction of the proximal ER-dependent genes was inhibited, but ER binding and methylations at the enhancer were unaltered. A high-throughput variant of chromosome conformation capture, termed 3D-DSL (Harismendy et al. 2011), was used to show that under normal conditions, ER binding induced qualitative and quantitative changes in promoter-enhancer looping. However, the presence of the eRNA-targeting siRNAs or LNAs resulted in alterations in the looping architecture at the targeted loci, suggesting that the eRNA transcripts were instrumental in the orchestration of ligand-dependent chromosome conformation changes. Further supporting this model, eRNA knockdown resulted in loss of cohesin, a protein with a role in promoter-enhancer looping interactions (Hadjur et al. 2009; Kagey et al. 2010; Schmidt et al. 2010), recruitment at the enhancers and associated promoters.
Similar results were obtained by Hsieh et al. in a study of androgen receptor (AR) enhancers in a prostate cancer cell line (Hsieh et al. 2014). eRNAs were produced at an enhancer of the AR-dependent gene KLK3 that was also marked by H3K27Ac, H3K4me1, and AR itself. Both the enhancer and KLK3 were induced upon androgen treatment. Upon knockdown of the eRNA with siRNA, expression of both KLK3 and the nearby KLK2 was inhibited, although other KLK genes in the locus were unaffected. To assess the role of the eRNA in regulating the two KLK genes, Hsieh et al. performed RNA immunoprecipitation with the sense and antisense strands of the eRNA and found that both pulled down AR as well as Mediator1 (Med1), which has been shown to be involved in chromosomal looping (Chen et al. 2011). 3C-qPCR demonstrated that either siRNA knockdown of the eRNA or knockdown of Med1 resulted in reduced looping of the KLK2 promoter to the enhancer locus. However, knockdown of the eRNA did not affect AR or Pol II occupancy at the enhancer itself, implying that the functional role of the eRNA in establishing enhancer-promoter looping is downstream of enhancer assembly. Complicating this result slightly, Hsieh et al. also found that knockdown of the eRNA at the KLK3 enhancer resulted in reduced expression of several AR target genes in trans, raising the question of what indirect effects the eRNA knockdown might have.
A second potential mechanism was highlighted by Mousavi et al. in an analysis of MyoD and MyoG binding activity in C2C12 skeletal muscle cells (Mousavi et al. 2013). At two enhancers, one proximal to MYOD1 and the other proximal to MYOG, knockdown of eRNAs with siRNA resulted in reduced recruitment of Pol II to both enhancers and their target genes. In line with this, binding of the transcription factor MyoD was reduced at the MYOG enhancer in response to knockdown of its eRNA, although this was not the case for the MYOD1 enhancer being studied. For both enhancers, knockdown of eRNAs resulted in reduced DNAse I accessibility at the target genes, although it was not clear how this effect was mediated, especially given that knockdown of the MYOD1 eRNA resulted in decreased accessibility at both MYOD1 and MYOG.
A third means by which eRNA might affect transcriptions was described in neurons responding to KCl-mediated membrane depolarization (Schaukowitch et al. 2014). Two enhancers proximal to the immediate early genes Arc and Gadd45b in neurons produce eRNA in response to stimulation. Strand-specific knockdown of the eRNAs with shRNA or LNAs resulted in reduced induction of their respective target genes, but, unlike the KLK3 locus, the looping of the enhancer to the promoter region was unaffected according to 3C assays. Similarly, recruitment of Med1 and members of the cohesion complex were unaffected. However, Schaukowitch et al. found that eRNA knockdown resulted in a marginally increased occupancy of the negative elongation factor (NELF) complex at the promoters of the target genes. NELF binds directly to Pol II and nascent RNA, and its release from the promoters of target genes is an important step in the unpausing of Pol II and subsequent mRNA elongation (Kwak and Lis 2013). In conjunction with increased NELF, there was a decrease in the elongating form of Pol II (phosphorylated at serine 5) with eRNA knockdown. Further, the eRNAs at Arc, Gadd45b, and c-fos were shown to pull down 1.5× to 2× more NELF in ultraviolet-crosslinked RNA immunoprecipitation assays, implying that the eRNA was binding directly to the RNA-binding domain of NELF. Thus, Schaukowitch et al. propose a model in which eRNA is able to destabilize the binding of NELF to nascent mRNAs, thereby allowing Pol II to elongate and continue to transcribe the mRNA.
7 Concluding Remarks
Enhancer transcription has emerged as a fascinating and quantitatively significant source of nuclear noncoding RNAs. Many questions remain regarding the general importance of enhancer transcription itself (apart from the eRNA product). At present, there is limited evidence that enhancer transcription is a major mechanism for deposition of H3K4me1/2 at enhancers other than the de novo enhancers studied by Kaikkonen et al. (2013). Although the proposed mechanism linking enhancer transcription to deposition of H3K4me1/2 via a PolII/MLL interaction is appealing, further studies of other classes of enhancers (e.g., those that are selected during transitions in cell differentiation) will be required to determine generality. In addition, while functional roles have been established for a small number of eRNAs, there is as yet no consistent picture of their mechanism(s) of action. Future studies are needed to examine questions such as the sequence determinants of activity and the identities of important interacting molecules, such as NELF. Macrophages are very likely to continue to be a robust experimental system for examination of these questions.
References
Allen MA, Andrysik Z, Dengler VL, Mellert HS, Guarnieri A, Freeman JA, Sullivan KD, Galbraith MD, Luo X, Kraus WL et al (2014) Global analysis of p53-regulated transcription identifies its direct targets and unexpected regulatory mechanisms. Elife 3:e02200
Allison KA, Kaikkonen MU, Gaasterland T, Glass CK (2014) Vespucci: a system for building annotated databases of nascent transcripts. Nucleic Acids Res 42:2433–2447
Beutler B (2000) Tlr4: central component of the sole mammalian LPS sensor. Curr Opin Immunol 12:20–26
Chao SH, Price DH (2001) Flavopiridol inactivates P-TEFb and blocks most RNA polymerase II transcription in vivo. J Biol Chem 276:31793–31799
Chen Z, Zhang C, Wu D, Chen H, Rorick A, Zhang X, Wang Q (2011) Phospho-MED1-enhanced UBE2C locus looping drives castration-resistant prostate cancer growth. EMBO J 30:2405–2419
Core LJ, Waterfall JJ, Lis JT (2008) Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322:1845–1848
Creyghton MP, Cheng AW, Welstead GG, Kooistra T, Carey BW, Steine EJ, Hanna J, Lodato MA, Frampton GM, Sharp PA et al (2010) Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc Natl Acad Sci U S A 107:21931–21936
Cristancho AG, Lazar MA (2011) Forming functional fat: a growing understanding of adipocyte differentiation. Nat Rev Mol Cell Biol 12:722–734
Dawson MA, Prinjha RK, Dittmann A, Giotopoulos G, Bantscheff M, Chan WI, Robson SC, Chung CW, Hopf C, Savitski MM et al (2011) Inhibition of BET recruitment to chromatin as an effective treatment for MLL-fusion leukaemia. Nature 478:529–533
De Santa F, Barozzi I, Mietton F, Ghisletti S, Polletti S, Tusi BK, Muller H, Ragoussis J, Wei CL, Natoli G (2010) A large fraction of extragenic RNA pol II transcription sites overlap enhancers. PLoS Biol 8:e1000384
Escoubet-Lozach L, Benner C, Kaikkonen MU, Lozach J, Heinz S, Spann NJ, Crotti A, Stender J, Ghisletti S, Reichart D et al (2011) Mechanisms establishing TLR4-responsive activation states of inflammatory response genes. PLoS Genet 7:e1002401
Feng R, Desbordes SC, Xie H, Tillo ES, Pixley F, Stanley ER, Graf T (2008) PU.1 and C/EBPalpha/beta convert fibroblasts into macrophage-like cells. Proc Natl Acad Sci U S A 105:6057–6062
Garber M, Yosef N, Goren A, Raychowdhury R, Thielke A, Guttman M, Robinson J, Minie B, Chevrier N, Itzhaki Z et al (2012) A high-throughput chromatin immunoprecipitation approach reveals principles of dynamic gene regulation in mammals. Mol Cell 47:810–822
Gerber M, Shilatifard A (2003) Transcriptional elongation by RNA polymerase II and histone methylation. J Biol Chem 278:26303–26306
Ghisletti S, Barozzi I, Mietton F, Polletti S, De Santa F, Venturini E, Gregory L, Lonie L, Chew A, Wei C-L et al (2010) Identification and characterization of enhancers controlling the inflammatory gene expression program in macrophages. Immunity 32:317–328
Hadjur S, Williams LM, Ryan NK, Cobb BS, Sexton T, Fraser P, Fisher AG, Merkenschlager M (2009) Cohesins form chromosomal cis-interactions at the developmentally regulated IFNG locus. Nature 460:410–413
Hah N, Danko CG, Core L, Waterfall JJ, Siepel A, Lis JT, Kraus WL (2011) A rapid, extensive, and transient transcriptional response to estrogen signaling in breast cancer cells. Cell 145:622–634
Harismendy O, Notani D, Song X, Rahim NG, Tanasa B, Heintzman N, Ren B, Fu XD, Topol EJ, Rosenfeld MG et al (2011) 9p21 DNA variants associated with coronary artery disease impair interferon-gamma signalling response. Nature 470:264–268
Heintzman ND, Stuart RK, Hon G, Fu Y, Ching CW, Hawkins RD, Barrera LO, Van Calcar S, Qu C, Ching KA et al (2007) Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat Genet 39:311–318
Heintzman ND, Hon GC, Hawkins RD, Kheradpour P, Stark A, Harp LF, Ye Z, Lee LK, Stuart RK, Ching CW et al (2009) Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459:108–112
Heinz S, Benner C, Spann N, Bertolino E, Lin YC, Laslo P, Cheng JX, Murre C, Singh H, Glass CK (2010) Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell 38:576–589
Heinz S, Romanoski CE, Benner C, Allison KA, Kaikkonen MU, Orozco LD, Glass CK (2013) Effect of natural genetic variation on enhancer selection and function. Nature 503(7477):487–492
Herrera R, Ro HS, Robinson GS, Xanthopoulos KG, Spiegelman BM (1989) A direct role for C/EBP and the AP-I-binding site in gene expression linked to adipocyte differentiation. Mol Cell Biol 9:5331–5339
Hsieh CL, Fei T, Chen Y, Li T, Gao Y, Wang X, Sun T, Sweeney CJ, Lee GS, Chen S et al (2014) Enhancer RNAs participate in androgen receptor-driven looping that selectively enhances gene activation. Proc Natl Acad Sci U S A 111:7319–7324
Hughes CM, Rozenblatt-Rosen O, Milne TA, Copeland TD, Levine SS, Lee JC, Hayes DN, Shanmugam KS, Bhattacharjee A, Biondi CA et al (2004) Menin associates with a trithorax family histone methyltransferase complex and with the hoxc8 locus. Mol Cell 13:587–597
Kagey MH, Newman JJ, Bilodeau S, Zhan Y, Orlando DA, van Berkum NL, Ebmeier CC, Goossens J, Rahl PB, Levine SS et al (2010) Mediator and cohesin connect gene expression and chromatin architecture. Nature 467:430–435
Kaikkonen MU, Spann NJ, Heinz S, Romanoski CE, Allison KA, Stender JD, Chun HB, Tough DF, Prinjha RK, Benner C et al (2013) Remodeling of the enhancer landscape during macrophage activation is coupled to enhancer transcription. Mol Cell 51:310–325
Kim TK, Hemberg M, Gray JM, Costa AM, Bear DM, Wu J, Harmin DA, Laptewicz M, Barbara-Haley K, Kuersten S et al (2010) Widespread transcription at neuronal activity-regulated enhancers. Nature 465:182–187
Klemsz MJ, McKercher SR, Celada A, Van Beveren C, Maki RA (1990) The macrophage and B cell-specific transcription factor PU.1 is related to the ets oncogene. Cell 61:113–124
Koch F, Jourquin F, Ferrier P, Andrau JC (2008) Genome-wide RNA polymerase II: not genes only! Trends Biochem Sci 33:265–273
Krogan NJ, Kim M, Tong A, Golshani A, Cagney G, Canadien V, Richards DP, Beattie BK, Emili A, Boone C et al (2003) Methylation of histone H3 by Set2 in Saccharomyces cerevisiae is linked to transcriptional elongation by RNA polymerase II. Mol Cell Biol 23:4207–4218
Kwak H, Lis JT (2013) Control of transcriptional elongation. Annu Rev Genet 47:483–508
Lam MT, Cho H, Lesch HP, Gosselin D, Heinz S, Tanaka-Oishi Y, Benner C, Kaikkonen MU, Kim AS, Kosaka M et al (2013) Rev-Erbs repress macrophage gene expression by inhibiting enhancer-directed transcription. Nature 498:511–515
Li W, Notani D, Ma Q, Tanasa B, Nunez E, Chen AY, Merkurjev D, Zhang J, Ohgi K, Song X et al (2013) Functional roles of enhancer RNAs for oestrogen-dependent transcriptional activation. Nature 498:516–520
MacConaill LE, Hughes CM, Rozenblatt-Rosen O, Nannepaga S, Meyerson M (2006) Phosphorylation of the menin tumor suppressor protein on serine 543 and serine 583. Mol Cancer Res 4:793–801
Maurya MR, Gupta S, Li X, Fahy E, Dinasarapu AR, Sud M, Brown HA, Glass CK, Murphy RC, Russell DW et al (2013) Analysis of inflammatory and lipid metabolic networks across RAW264.7 and thioglycolate-elicited macrophages. J Lipid Res 54:2525–2542
Medzhitov R, Horng T (2009) Transcriptional control of the inflammatory response. Nat Rev Immunol 9:692–703
Meissner F, Scheltema RA, Mollenkopf HJ, Mann M (2013) Direct proteomic quantification of the secretome of activated immune cells. Science 340:475–478
Melo CA, Drost J, Wijchers PJ, van de Werken H, de Wit E, Oude Vrielink JA, Elkon R, Melo SA, Leveille N, Kalluri R et al (2013) eRNAs are required for p53-dependent enhancer activity and gene transcription. Mol Cell 49:524–535
Milne TA, Dou Y, Martin ME, Brock HW, Roeder RG, Hess JL (2005) MLL associates specifically with a subset of transcriptionally active target genes. Proc Natl Acad Sci U S A 102:14765–14770
Mousavi K, Zare H, Dell’orso S, Grontved L, Gutierrez-Cruz G, Derfoul A, Hager GL, Sartorelli V (2013) eRNAs promote transcription by establishing chromatin accessibility at defined genomic loci. Mol Cell 51:606–617
Mullen AC, Orlando DA, Newman JJ, Loven J, Kumar RM, Bilodeau S, Reddy J, Guenther MG, DeKoter RP, Young RA (2011) Master transcription factors determine cell-type-specific responses to TGF-beta signaling. Cell 147:565–576
NE II, Heward JA, Roux B, Tsitsiou E, Fenwick PS, Lenzi L, Goodhead I, Hertz-Fowler C, Heger A, Hall N et al (2014) Long non-coding RNAs and enhancer RNAs regulate the lipopolysaccharide-induced inflammatory response in human monocytes. Nat Commun 5:3979
Ng HH, Robert F, Young RA, Struhl K (2003) Targeted recruitment of Set1 histone methylase by elongating Pol II provides a localized mark and memory of recent transcriptional activity. Mol Cell 11:709–719
Nicodeme E, Jeffrey KL, Schaefer U, Beinke S, Dewell S, Chung CW, Chandwani R, Marazzi I, Wilson P, Coste H et al (2010) Suppression of inflammation by a synthetic histone mimic. Nature 468:1119–1123
Ong CT, Corces VG (2011) Enhancer function: new insights into the regulation of tissue-specific gene expression. Nat Rev Genet 12:283–293
Ostuni R, Piccolo V, Barozzi I, Polletti S, Termanini A, Bonifacio S, Curina A, Prosperini E, Ghisletti S, Natoli G (2013) Latent enhancers activated by stimulation in differentiated cells. Cell 152:157–171
Pulakanti K, Pinello L, Stelloh C, Blinka S, Allred J, Milanovich S, Kiblawi S, Peterson J, Wang A, Yuan GC et al (2013) Enhancer transcribed RNAs arise from hypomethylated, Tet-occupied genomic regions. Epigenetics 8:1303–1320
Rada-Iglesias A, Bajpai R, Swigut T, Brugmann SA, Flynn RA, Wysocka J (2011) A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470:279–283
Rana R, Surapureddi S, Kam W, Ferguson S, Goldstein JA (2011) Med25 is required for RNA polymerase II recruitment to specific promoters, thus regulating xenobiotic and lipid metabolism in human liver. Mol Cell Biol 31:466–481
Schaukowitch K, Joo JY, Liu X, Watts JK, Martinez C, Kim TK (2014) Enhancer RNA facilitates NELF release from immediate early genes. Mol Cell 56:29–42
Schmidt D, Schwalie PC, Ross-Innes CS, Hurtado A, Brown GD, Carroll JS, Flicek P, Odom DT (2010) A CTCF-independent role for cohesin in tissue-specific transcription. Genome Res 20:578–588
Scott EW, Simon MC, Anastasi J, Singh H (1994) Requirement of transcription factor PU.1 in the development of multiple hematopoietic lineages. Science 265:1573–1577
Smale ST (2012) Transcriptional regulation in the innate immune system. Curr Opin Immunol 24:51–57
Soufi A, Donahue G, Zaret KS (2012) Facilitators and impediments of the pluripotency reprogramming factors’ initial engagement with the genome. Cell 151:994–1004
Step SE, Lim HW, Marinis JM, Prokesch A, Steger DJ, You SH, Won KJ, Lazar MA (2014) Anti-diabetic rosiglitazone remodels the adipocyte transcriptome by redistributing transcription to PPARgamma-driven enhancers. Genes Dev 28:1018–1028
Szutorisz H, Dillon N, Tora L (2005) The role of enhancers as centres for general transcription factor recruitment. Trends Biochem Sci 30:593–599
Takeuchi O, Akira S (2010) Pattern recognition receptors and inflammation. Cell 140:805–820
Thurman RE, Rynes E, Humbert R, Vierstra J, Maurano MT, Haugen E, Sheffield NC, Stergachis AB, Wang H, Vernot B et al (2012) The accessible chromatin landscape of the human genome. Nature 489:75–82
Travers A (1999) Chromatin modification by DNA tracking. Proc Natl Acad Sci U S A 96:13634–13637
Trompouki E, Bowman TV, Lawton LN, Fan ZP, Wu DC, DiBiase A, Martin CS, Cech JN, Sessa AK, Leblanc JL et al (2011) Lineage regulators direct BMP and Wnt pathways to cell-specific programs during differentiation and regeneration. Cell 147:577–589
Visel A, Blow MJ, Li Z, Zhang T, Akiyama JA, Holt A, Plajzer-Frick I, Shoukry M, Wright C, Chen F et al (2009) ChIP-seq accurately predicts tissue-specific activity of enhancers. Nature 457:854–858
Wang D, Garcia-Bassets I, Benner C, Li W, Su X, Zhou Y, Qiu J, Liu W, Kaikkonen MU, Ohgi KA et al (2011) Reprogramming transcription by distinct classes of enhancers functionally defined by eRNA. Nature 474:390–394
Wood A, Schneider J, Dover J, Johnston M, Shilatifard A (2003) The Paf1 complex is essential for histone monoubiquitination by the Rad6-Bre1 complex, which signals for histone methylation by COMPASS and Dot1p. J Biol Chem 278:34739–34742
Wu H, Nord AS, Akiyama JA, Shoukry M, Afzal V, Rubin EM, Pennacchio LA, Visel A (2014) Tissue-specific RNA expression marks distant-acting developmental enhancers. PLoS Genet 10:e1004610
Wynn TA, Chawla A, Pollard JW (2013) Macrophage biology in development, homeostasis and disease. Nature 496:445–455
Xiao T, Hall H, Kizer KO, Shibata Y, Hall MC, Borchers CH, Strahl BD (2003) Phosphorylation of RNA polymerase II CTD regulates H3 methylation in yeast. Genes Dev 17:654–663
Yin L, Lazar MA (2005) The orphan nuclear receptor Rev-erbalpha recruits the N-CoR/histone deacetylase 3 corepressor to regulate the circadian Bmal1 gene. Mol Endocrinol 19:1452–1459
Zamir I, Harding HP, Atkins GB, Horlein A, Glass CK, Rosenfeld MG, Lazar MA (1996) A nuclear hormone receptor corepressor mediates transcriptional silencing by receptors with distinct repression domains. Mol Cell Biol 16:5458–5465
Zhu Y, Sun L, Chen Z, Whitaker JW, Wang T, Wang W (2013) Predicting enhancer transcription and activity from chromatin modifications. Nucleic Acids Res 41:10032–10043
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Japan
About this chapter
Cite this chapter
Allison, K.A., Glass, C.K. (2015). Macrophage Activation as a Model System for Understanding Enhancer Transcription and eRNA Function. In: Kurokawa, R. (eds) Long Noncoding RNAs. Springer, Tokyo. https://doi.org/10.1007/978-4-431-55576-6_12
Download citation
DOI: https://doi.org/10.1007/978-4-431-55576-6_12
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-55575-9
Online ISBN: 978-4-431-55576-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)