Abstract
Understanding brain function requires experimental approaches that decipher the information coded by individual neurons. Calcium imaging with genetically encoded calcium indicators (GECIs) is a promising method that can visualize the spatiotemporal activity patterns of brain cells. Recent advances in protein engineering have greatly improved the properties of fluorescent GECIs, and they now have high flexibility for imaging defined cell populations over the course of months. The action spectra of single-wavelength GECIs have been extended by the development of color-shifted fluorescence probes, which increase the potential for multicolor imaging of different cell structures and, more importantly, can be integrated into optogenetics experiments with photoactivatable proteins. In particular, red-shifted GECIs are highly important for imaging deeper tissues because longer-wavelength light can reduce tissue scattering and background autofluorescence. This chapter mainly describes recent advances in the engineering of red GECIs, and highlights important neuroscience applications in optical monitoring and the manipulation of neuronal activity.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Calcium imaging
- Genetically encoded calcium indicator
- Red fluorescent protein
- R-CaMP
- ChR2
- Cortical neurons
1 Genetically Encoded Calcium Indicators (GECIs) for Monitoring the Spatiotemporal Dynamics of Neuronal Populations
Monitoring where, when, and how individual cells are active in the brain is a central requirement in neuroscience research. Over the past decades, electrophysiological recordings have been used extensively for capturing neuronal dynamics at a single-cell resolution both in vitro and in vivo. Advances in multi-channel unit recordings of animal behavior allows the simultaneous recording of up to hundreds of neurons, which have provided major contributions to the discovery of activity patterns in a variety of brain circuits (Buzsáki 2004). However, the drawbacks of microelectrode recording are the lack of sufficient spatial resolution to resolve the precise location of cells and the difficulty of tracking identified cells over consecutive recording sessions because of electrode drifting, cell death, and gliosis around the tips of electrodes.
Compared with electrophysiological techniques, optical imaging holds greater promise for probing the activity patterns of large groups of neurons with identified physical positions over extended periods of time (Gobel and Helmchen 2007; Takahashi et al. 2010b; Grienberger and Konnerth 2012). In particular, calcium fluorescent imaging has been used to obtain reliable measurement of neuronal activity with a high signal-to-noise ratio (SNR) because action potential firing is coupled tightly to large and rapid changes in the intracellular calcium concentration. Multicell calcium imaging is performed by loading cells with membrane-permeant forms of synthetic calcium indicators such as Fura-2, Oregon Green BAPTA (OGB)-1, and Fluo-4 (Regehr and Tank 1991; Yuste et al. 1992; Dombeck et al. 2007; Chen et al. 2011; Takahashi et al. 2010a). The major advantages of these organic probes are as follows: (i) they are bright, high-affinity, and have calcium-dependent fluorescence changes with large dynamic ranges; and (ii) the bulk loading procedure is relatively simple, i.e., simply incubating living tissues with dye-containing solution. Despite the widespread availability of synthetic indicators, they are generally unsuitable for labeling specific cell populations, which limits the examination of detailed circuit dynamics. Furthermore, it is technically challenging to visualize identical neuron populations over days and weeks due to the rapid clearance of indicators from the intracellular spaces.
Genetically encoded calcium indicators (GECIs) have the potential to overcome these technical limitations (Tian et al. 2012). GECIs are proteins; therefore, they can be expressed by delivering their genes into cells via viral vectors (Ziv et al. 2013; Osakada et al. 2011), electroporation techniques (Yamada et al. 2011; Ohkura et al. 2012b), or by constructing transgenic animal lines (Chen et al. 2012; Zariwala et al. 2012). GECIs have several remarkable advantages over synthetic indicators. First, GECIs can be targeted to specific cell types or specific subcellular compartments using cell-type specific promoters or cellular targeting sequences. Second, GECIs can be tracked repeatedly to analyze the neuronal activity of identical cell populations over the course of multiple recording sessions. Third, GECIs can be easily applied to mature neurons, whereas the loading efficiency of synthetic indicators generally declines as the age of neurons increases. These properties should be essential for unveiling the development, organization, and maintenance of neural circuit dynamics.
2 Recent Improvements in Green GECIs
Two important classes of GECIs have been engineered: (i) the Förster resonance energy transfer (FRET) type indicators such as a series of Cameleons (Nagai et al. 2004; Horikawa et al. 2010; Miyawaki et al. 1997), D3cpVenus (Palmer and Tsien 2006), and TN-XXL (Mank et al. 2008); and (ii) the single-wavelength type indicators, such as a series of GCaMPs (Nakai et al. 2001; Ohkura et al. 2005, 2012b; Tian et al. 2009, 2012; Chen et al. 2013) and flash-pericam (Nagai et al. 2001). Both types of GECIs comprise a calcium-binding domain such as calmodulin or troponin C and either one or two fluorescent proteins. These protein structures undergo changes in their fluorescence intensity, depending on the binding of calcium ions. Of these GECIs, GCaMPs are the most widely used sensors in a variety of brain regions and organisms, and they have been improved repeatedly to obtain faster kinetics and larger dynamic ranges of fluorescence changes. In particular, determinations of the X-ray crystal structures of GCaMPs have facilitated more rational improvements in the properties of probes. Recently, structure-based and random mutagenesis and semi-rational library screening have recently produced highly sensitive green calcium probes, including GCaMP3 (Tian et al. 2009), GCaMP4.1(Shindo et al. 2010), G-GECO (Zhao et al. 2011), and GCaMP5 (Akerboom et al. 2012). The latest version of GCaMPs is a family of GCaMP6 (Ohkura et al. 2012b; Chen et al. 2013), which have been developed by two independent groups (for further details, see Chap. 9 by Ohkura and Nakai). Such a new generation of calcium probes significantly outperforms other existing GECIs in the detection of neuronal activity at a sensitivity level similar to or better than those of synthetic calcium dyes. Green GECIs have been shown to be applicable to the long-term in vivo imaging of orientation-tuned activity in visual cortical neurons (Chen et al. 2013) and place-selective firing in hippocampal pyramidal neurons (Ziv et al. 2013) over days and weeks.
3 Development and Application of Red GECIs
3.1 Requirements for Red GECIs
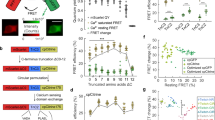
Despite the ongoing efforts to improve GECIs, the majority of single-wavelength probes were still limited to green fluorescent proteins (GFPs) or yellow fluorescent proteins (YFPs) that emit fluorescence at a wavelength of 500–550 nm. This restricted emission spectrum prevents the use of GECIs in cells that have already been labeled with a green fluorescent probe. Furthermore, recent advances in optogenetic manipulation using light-modulated microbial opsins (Yizhar et al. 2011; Nagel et al. 2003; Chow et al. 2010; Zhang et al. 2007; Boyden et al. 2005) such as channelrhodopsin (ChR)-2 require the combinatorial use of GECIs and photoactivatable proteins in an identical cell population. However, the use of GECIs to visualize opsin-expressing cells without photoactivation is technically challenging because of the large overlap in their broad excitation spectra (GCaMPs 470–510 nm; ChR2 430–500 nm). Thus, GCaMP3 and ChR2 can only be combined in conditions where the laser power is lowered strictly to excite GCaMP3 alone but not ChR2 (Guo et al. 2009). The development of GECIs with different absorption and emission spectra from GFPs should resolve these technical limitations and allow GECIs to be employed together with optogenetic tools. A number of fluorescent proteins have been developed, but few multicolor probes that exhibit clear fluorescent changes in response to calcium ions. Several research groups have recently engineered color-shifted GECIs by combining the fluorescent proteins and a calcium-binding protein (Zhao et al. 2011; Akerboom et al. 2012) (Fig. 10.1). Among these, red-shifted indicators are particularly desirable because longer-wavelength light is less susceptible to tissue scattering, blood absorption, phototoxicity, and background autofluorescence. These conditions are appropriate for in vivo live cell imaging from deeper tissues. This section describes the recent development and application of red-emitting GECIs in neuroscience research.
3.2 Repertoire of Red GECIs
3.2.1 R-GECO1
While neurophysiologists were awaiting new applications of GECIs, a pioneering study by Zhao et al. (2011) succeeded in creating color-shifted GECIs with unique excitation and emission spectra on the basis of a combination of single-wavelength fluorescent proteins. They used a variety of strategies to obtain efficient probes, including mutagenesis with an error-prone polymerase chain reaction (PCR) and digital fluorescence imaging of gene variants expressed in Escherichia coli colonies. Introducing targeted and random mutations into GCaMP3 yielded a blue calcium indicator, B-GECO1, and a series of high-sensitive green calcium indicators, G-GECOs. Part of the cfGFP in G-GECO1.1 was then replaced with an analogous cp version of a red fluorescent protein (mApple) (Shaner et al. 2008), which yielded red-shifted probes that are potentially calcium sensitive. Through the iterative evolution of variants created by error-prone PCR and randomization, the best optimized protein was finally identified as R-GECO1. The absorption and emission maxima of R-GECO1 were approximately 570 nm and 600 nm, respectively. On the basis of the calcium responses obtained from HeLa cells and cultured cells, the dynamic range of fluorescent changes in R-GECO1 appears to be almost similar to that in G-CaMP3.
3.2.2 R-CaMP1.07
The development of red-shifted GECIs by Zhao et al. (2011) motivated us to improve the probe further (Ohkura et al. 2012a). As an initial step, random mutations were introduced into a prototype molecule, R-GECO1, with the RSET tag at the N-terminus. Repeated rounds of in vitro screening yielded the best variant, R-CaMP1.01, which had mutations of K47V and T49V in the circularly permuted mApple domain. However, both the R-GECO1 and R-CaMP1.01 signals were found to be aggregated in the HeLa cell nucleus and the cytoplasm. This abnormal subcellular localization is probably related to the dysfunctioning of probes due to partial proteolysis (Tian et al. 2009). To prevent nuclear entry, R-CaMP1.01 was combined with a self-cleaving peptide, F2A, at the C-terminus. The resulting variant obtained, R-CaMP1.07, showed no signals of nuclear localization, and approximately two times larger fluorescent changes in response to adenosine triphosphate (ATP) application in HeLa cells compared with R-GECO1.
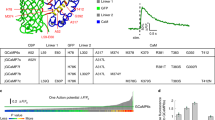
The performance of R-CaMP1.07 was also characterized in mammalian hippocampal CA3 pyramidal neurons in cultured slices. R-CaMP1.07 detected calcium transients induced by single spikes with 95 % probability. The signal amplitudes increased almost linearly up to six spikes with a frequency of 50 Hz (Fig. 10.2A). On average, R-CaMP1.07 yielded 1.5- to 2.0-fold larger dynamic responses (ΔF/F) and higher SNRs than R-GECO1 (Fig. 10.2B). The rise and decay time constants of the spike-induced calcium transients of R-CaMP1.07 were approximately 120 ms and 900 ms, respectively, which are almost identical to those of R-GECO1. According to the temporal kinetics, this sensor could resolve individual spikes in bursts with a frequency of up to 5 Hz (Sasaki et al. 2008). However, R-CaMP1.07 has not yet reached the affinity and speed of the commonly used green GECIs. It may be possible to optimize the kinetics of sensors to further resolve the temporal patterns of spikes during burst firing.
Detection of spiking activity using red genetically encoded calcium indicators (GECIs) in hippocampal pyramidal cells. (a) Representative fluorescent traces (ΔF/F) in response to spike trains with a frequency of 50 Hz. Imaging was performed at 50 frames per second (fps) using a Nipkow-disk confocal microscope. The arrows indicate the timing of spikes. (b) Mean signal-to-noise ratios (SNRs) plotted against the number of action potentials in R-GECO1 (black) and R-CaMP1.07 (red) (Copyright (2014) PLOS. From Ohkura M et al. (2012a) with permission)
3.2.3 RCaMPs
In parallel with the study of mApple-based GECIs, Akerboom et al. (2012) independently engineered a series of multi-color GECIs. To create color-shifted GECIs, they first introduced mutations into the GFP chromophore of GCaMP3 through the mutation sites that are crucial for conversion to color-shifted fluorescent proteins (Heim et al. 1994; Heim and Tsien 1996; Ormo et al. 1996). Additional systematic mutations of these color-shifted GCaMP3 yielded blue, cyan, and yellow probes, which displayed substantial fluorescent changes in response to calcium (i.e., BCaMP, CyCaMP, and YCaMP, respectively). However, this strategy failed to produce red variants of GCaMP3. As an alternative pathway, part of cpEGFP in GCaMP3 was replaced with a circularly permuted version of mRuby (Kredel et al. 2009), a red fluorescent protein. The addition of random mutations and subsequent structure-guided optimization produced red-emitting calcium indicators, RCaMP1a-h (note that these are different probes from R-CaMP1.07, which was developed by Ohkura et al. [2012a]). In cultured rat hippocampal neurons, RCaMP1d, 1e, 1f, and 1 h were brighter, and they exhibited smaller fluorescent changes in response to neuronal activity than did R-GECO1. The kinetics of these RCaMPs were slower than those of R-GECO1. These results suggest that R-GECO1 may be more suitable for detecting neuronal activity than R-CaMPs. However, RCaMPs have significant advantages over R-GECO1, i.e., RCaMPs are more tolerant to photobleeching and, as described later, they are never photoactivated by blue and green wavelength light, which is a prerequisite for their combined use with optogenetics tools. In addition, it has been demonstrated that RCaMPs can be used together with GCaMP5 for visualization of the calcium activity with multicolor fluorescence from different organelles in a single cell or different cell types in a single preparation.
3.3 Application of Red GECIs to Optogenetics
The excitation spectra of red-shifted GECIs are distinct from the action wavelengths used for photostimulating ChR2 (Fig. 10.3B). By exploiting their spectra properties, red GECIs can be used in optogenetics experiments for monitoring and manipulating neural activity with multiple light wavelengths. Ohkura et al. (2012a) demonstrated that R-CaMP1.07 fluorescence imaging can be used to detect spike-induced calcium events triggered via ChR2 photoactivation with blue light (Fig. 10.3C). A longer duration of photostimulation evoked larger numbers of spikes, which were linearly correlated with the amplitudes of the fluorescent changes. No membrane potential changes were induced during the imaging of R-CaMP1.07 fluorescence with 568 nm light, indicating that ChR2 was not activated in these imaging conditions. These results demonstrate that R-CaMP1.07 may be suitable for monitoring calcium transients in response to optically evoked spikes. However, the drawback of the imaging system by Ohkura et al. (2012a) is that blue light illumination causes the contamination of photosignals, which prevents the direct recording of red fluorescent changes upon ChR2 photostimulation (as shown in the dashed line in Fig. 10.3A). Chang et al. (2012) overcame this issue by developing an optical system where a blue light pulse was applied during short intervals between image-acquisition events (Chang et al. 2012). This system allowed R-GECO calcium imaging without overlapping the photosignals generated by ChR2 photostimulation.
Optical monitoring and manipulation of neuronal activity. (a) Hippocampal pyramidal cell co-expressing R-CaMP1.07 and channelrhodopsin (ChR)-2. (b) Excitation (green) and emission (red) spectra of R-CaMP1.07 superimposed on the excitation spectrum of ChR2 (cyan). The spectrum range of excitation filter used for RCaMP1.07 imaging is shown as the green region. (c) Imaging of ChR2-triggered-action potentials by R-CaMP1.07. Photostimulation (470-nm) is indicated by the blue region. Putative calcium increases during the photostimulation are represented by broken lines. The number of action potentials was recorded using the current clamp mode (Copyright (2014) PLOS. From Ohkura et al. (2012a) with permission)
However, the results reported in these two previous studies have been questioned by the claim that mApple-based sensors, including R-GECO1 and R-CaMP1.07, inherit the property of photoreactivity to blue and green light; thus, implying that the ChR2-evoked fluorescent changes detected by these probes may be prone to artifacts. In contrast, RCaMPs produced by Akerboom et al. (2013) do not exhibit these photo-sensitive dynamics; therefore, it is possibly more suitable for use in artifact-free, integrated optogenetics experiments.
3.4 Note Regarding Red GECIs
It should be noted that the expression of GECIs alone may affect endogenous calcium-dependent signaling by buffering intracellular free calcium ions, thus leading to abnormal changes in cellular functions and behavioral phenotypes. For example, long-term and/or a high level of expression of GCaMP3 using in utero electroporation or viral infection caused nonfunctional indicators and abnormal physiology (Dombeck et al. 2010; Tian et al. 2009). In certain conditions, this concern may be excluded by demonstrating that GECI-positive cells in cultured slices do not show any changes in their intracellular and synaptic properties (Ohkura et al. 2012a, b), and GCaMP3-expressing transgenic mice generally display calcium transients in vivo without changing cellular functions (Chen et al. 2012). Overall, the most crucial factors are likely to be the accurate timing and magnitude of the expression of GECIs. Depending on the stability and affinity of sensors, the optimization of expression levels should be considered on a case-by-case basis. In addition, moderate levels of calcium indicators are crucial for obtaining the best performance with GECIs because excessively high concentrations of calcium indicators substantially affect the shapes of calcium transients with extended durations and smaller amplitudes (Helmchen et al. 1996).
The performance of red-shifted GECIs in rodent cells has been tested almost exclusively in slice preparations or cultured cells. However, little is known regarding whether these sensors may work in vivo with similar detection levels in rodent neurons. In general, the dynamic range of GECIs decreases in more intact preparations due to a variety of external factors such as pH, temperature, light scattering, and background autofluorescence (Tian et al. 2009; Ohkura et al. 2012b). In addition, the ability of detection by red GECIs in different neuron types has not yet been systematically examined. A number of physiological factors such as the electrical activity, the expression of calcium channels, and the endogenous calcium buffering capacity, vary considerably between neurons of different subtypes and among brain regions (Fierro and Llano 1996; Maeda et al. 1999). In other preparations, particularly in vivo rodents, the usefulness of each GECI should be confirmed and compared by individual end users. Further studies are needed to determine the optimal situations where each GECI shows the best sensitivity, dynamic range, and kinetics.
4 Concluding Remarks
Recent protein engineering has continuously improved the properties of GECIs in terms of their brightness, calcium affinity, response kinetics, and color expansion. In this trend, multicolor GECIs will facilitate new applications of optical imaging and activity manipulations, which have not been performed with existing experimental tools. To support these applications, gene delivery systems have been greatly improved to allow the expression of functional probes in a specific class of a neuronal population (Osakada et al. 2011; Chen et al. 2012; Zariwala et al. 2012). In addition, the microscopic instrumentation has improved recently, including multiphoton microscopy, high-speed scanning systems, Gallium Arsenide Phosphate (GaAsP) detectors, and portable integrated microscopy (Ghosh et al. 2011). This progress in biological and technological engineering should encourage neuroscientists to address a variety of fundamental questions regarding the neural mechanisms of behavior, learning, and diseases.
References
Akerboom J, Chen TW, Wardill TJ et al (2012) Optimization of a GCaMP calcium indicator for neural activity imaging. J Neurosci 32(40):13819–13840
Boyden ES, Zhang F, Bamberg E et al (2005) Millisecond-timescale, genetically targeted optical control of neural activity. Nat Neurosci 8(9):1263–1268
Buzsáki G (2004) Large-scale recording of neuronal ensembles. Nat Neurosci 7(5):446–451
Chang YF, Arai Y, Nagai T (2012) Optogenetic activation during detector “dead time” enables compatible real-time fluorescence imaging. Neurosci Res 73(4):341–347
Chen X, Leischner U, Rochefort NL et al (2011) Functional mapping of single spines in cortical neurons in vivo. Nature 475(7357):501–505
Chen Q, Cichon J, Wang W et al (2012) Imaging neural activity using Thy1-GCaMP transgenic mice. Neuron 76(2):297–308
Chen TW, Wardill TJ, Sun Y et al (2013) Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499(7458):295–300
Chow BY, Han X, Dobry AS et al (2010) High-performance genetically targetable optical neural silencing by light-driven proton pumps. Nature 463(7277):98–102
Dombeck DA, Khabbaz AN, Collman F et al (2007) Imaging large-scale neural activity with cellular resolution in awake, mobile mice. Neuron 56(1):43–57
Dombeck DA, Harvey CD, Tian L et al (2010) Functional imaging of hippocampal place cells at cellular resolution during virtual navigation. Nat Neurosci 13(11):1433–1440
Fierro L, Llano I (1996) High endogenous calcium buffering in Purkinje cells from rat cerebellar slices. J Physiol 496(Pt 3):617–625
Ghosh KK, Burns LD, Cocker ED et al (2011) Miniaturized integration of a fluorescence microscope. Nat Methods 8(10):871–878
Gobel W, Helmchen F (2007) In vivo calcium imaging of neural network function. Physiology (Bethesda) 22:358–365
Grienberger C, Konnerth A (2012) Imaging calcium in neurons. Neuron 73(5):862–885
Guo ZV, Hart AC, Ramanathan S (2009) Optical interrogation of neural circuits in Caenorhabditis elegans. Nat Methods 6(12):891–896
Heim R, Tsien RY (1996) Engineering green fluorescent protein for improved brightness, longer wavelengths and fluorescence resonance energy transfer. Curr Biol 6(2):178–182
Heim R, Prasher DC, Tsien RY (1994) Wavelength mutations and posttranslational autoxidation of green fluorescent protein. Proc Natl Acad Sci U S A 91(26):12501–12504
Helmchen F, Imoto K, Sakmann B (1996) Ca2+ buffering and action potential-evoked Ca2+ signaling in dendrites of pyramidal neurons. Biophys J 70(2):1069–1081
Horikawa K, Yamada Y, Matsuda T et al (2010) Spontaneous network activity visualized by ultrasensitive Ca2+ indicators, yellow Cameleon-Nano. Nat Methods 7(9):729–732
Kredel S, Oswald F, Nienhaus K et al (2009) mRuby, a bright monomeric red fluorescent protein for labeling of subcellular structures. PLoS One 4(2):e4391. doi:10.1371/journal.pone.0004391
Maeda H, Ellis-Davies GC, Ito K et al (1999) Supralinear Ca2+ signaling by cooperative and mobile Ca2+ buffering in Purkinje neurons. Neuron 24(4):989–1002
Mank M, Santos AF, Direnberger S et al (2008) A genetically encoded calcium indicator for chronic in vivo two-photon imaging. Nat Methods 5(9):805–811
Miyawaki A, Llopis J, Heim R et al (1997) Fluorescent indicators for Ca2+ based on green fluorescent proteins and calmodulin. Nature 388(6645):882–887
Nagai T, Sawano A, Park ES et al (2001) Circularly permuted green fluorescent proteins engineered to sense Ca2+. Proc Natl Acad Sci U S A 98(6):3197–3202
Nagai T, Yamada S, Tominaga T et al (2004) Expanded dynamic range of fluorescent indicators for Ca2+ by circularly permuted yellow fluorescent proteins. Proc Natl Acad Sci U S A 101(29):10554–10559
Nagel G, Szellas T, Huhn W et al (2003) Channelrhodopsin-2, a directly light-gated cation-selective membrane channel. Proc Natl Acad Sci U S A 100(24):13940–13945
Nakai J, Ohkura M, Imoto K (2001) A high signal-to-noise Ca2+ probe composed of a single green fluorescent protein. Nat Biotechnol 19(2):137–141
Ohkura M, Matsuzaki M, Kasai H et al (2005) Genetically encoded bright Ca2+ probe applicable for dynamic Ca2+ imaging of dendritic spines. Anal Chem 77(18):5861–5869
Ohkura M, Sasaki T, Kobayashi C et al (2012a) An improved genetically encoded red fluorescent Ca2+ indicator for detecting optically evoked action potentials. PLoS One 7(7):e39933. doi:10.1371/journal.pone.0039933
Ohkura M, Sasaki T, Sadakari J et al (2012b) Genetically encoded green fluorescent Ca2+ indicators with improved detectability for neuronal Ca2+ signals. PLoS One 7(12):e51286. doi:10.1371/journal.pone.0051286
Ormo M, Cubitt AB, Kallio K et al (1996) Crystal structure of the Aequorea victoria green fluorescent protein. Science 273(5280):1392–1395
Osakada F, Mori T, Cetin AH et al (2011) New rabies virus variants for monitoring and manipulating activity and gene expression in defined neural circuits. Neuron 71(4):617–631
Palmer AE, Tsien RY (2006) Measuring calcium signaling using genetically targetable fluorescent indicators. Nat Protoc 1(3):1057–1065
Regehr WG, Tank DW (1991) The maintenance of LTP at hippocampal mossy fiber synapses is independent of sustained presynaptic calcium. Neuron 7(3):451–459
Sasaki T, Takahashi N, Matsuki N et al (2008) Fast and accurate detection of action potentials from somatic calcium fluctuations. J Neurophysiol 100(3):1668–1676
Shaner NC, Lin MZ, McKeown MR et al (2008) Improving the photostability of bright monomeric orange and red fluorescent proteins. Nat Methods 5(6):545–551
Shindo A, Hara Y, Yamamoto TS et al (2010) Tissue-tissue interaction-triggered calcium elevation is required for cell polarization during Xenopus gastrulation. PLoS One 5(2):e8897. doi:10.1371/journal.pone.0008897
Takahashi N, Sasaki T, Matsumoto W et al (2010a) Circuit topology for synchronizing neurons in spontaneously active networks. Proc Natl Acad Sci U S A 107(22):10244–10249
Takahashi N, Takahara Y, Ishikawa D et al (2010b) Functional multineuron calcium imaging for systems pharmacology. Anal Bioanal Chem 398(1):211–218
Tian L, Hires SA, Mao T et al (2009) Imaging neural activity in worms, flies and mice with improved GCaMP calcium indicators. Nat Methods 6(12):875–881
Tian L, Hires SA, Looger LL (2012) Imaging neuronal activity with genetically encoded calcium indicators. Cold Spring Harb Protoc 2012(6):647–656
Yamada Y, Michikawa T, Hashimoto M et al (2011) Quantitative comparison of genetically encoded Ca indicators in cortical pyramidal cells and cerebellar Purkinje cells. Front Cell Neurosci 5:18. doi:10.3389/fncel.2011.00018
Yizhar O, Fenno LE, Davidson TJ et al (2011) Optogenetics in neural systems. Neuron 71(1):9–34
Yuste R, Peinado A, Katz LC (1992) Neuronal domains in developing neocortex. Science 257(5070):665–669
Zariwala HA, Borghuis BG, Hoogland TM et al (2012) A Cre-dependent GCaMP3 reporter mouse for neuronal imaging in vivo. J Neurosci 32(9):3131–3141
Zhang F, Wang LP, Brauner M et al (2007) Multimodal fast optical interrogation of neural circuitry. Nature 446(7136):633–639
Zhao Y, Araki S, Wu J et al (2011) An expanded palette of genetically encoded Ca2+ indicators. Science 333(6051):1888–1891
Ziv Y, Burns LD, Cocker ED et al (2013) Long-term dynamics of CA1 hippocampal place codes. Nat Neurosci 16(3):264–266
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Japan
About this chapter
Cite this chapter
Sasaki, T. (2015). Probing Neuronal Activity Using Genetically Encoded Red Fluorescent Calcium Indicators. In: Yawo, H., Kandori, H., Koizumi, A. (eds) Optogenetics. Springer, Tokyo. https://doi.org/10.1007/978-4-431-55516-2_10
Download citation
DOI: https://doi.org/10.1007/978-4-431-55516-2_10
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-55515-5
Online ISBN: 978-4-431-55516-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)