Abstract
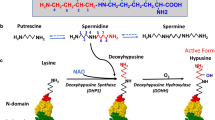
Although polyamines exert various effects on nucleic acids and macromolecular synthesis as polycations, spermidine is covalently incorporated into a single protein, eukaryotic initiation factor 5A (eIF5A), through a unique posttranslational modification. In this reaction, the aminobutyl moiety of spermidine is conjugated to a specific lysine residue of eIF5A to form an unusual amino acid, hypusine [N ε-(4-amino-2-hydroxybutyl)-lysine]. It occurs by two enzymatic steps catalyzed by deoxyhypusine synthase (DHS) and deoxyhypusine hydroxylase (DOHH). Hypusine synthesis occurs exclusively in eIF5A and is essential for eukaryotic cell proliferation. Although only a small percentage of the total spermidine in cells is used for hypusine formation, cells cannot survive/grow when hypusinated eIF5A falls below a critical level. Inactivation of the eIF5A gene or DHS gene is lethal in yeast and in mouse, further indicating the vital role of hypusinated eIF5A. eIF5A has been proposed to promote translation of a subset of cellular mRNAs. Indeed, recent evidence suggests that eIF5A facilitates translation at the elongation step, particularly at multiple strings of proline residues. A model of eIF5A docked in the ribosome reveals the hypusine directed toward the peptidyl transferase center. Thus, the hypusine modification defines a link between polyamines and cell growth, through promotion of translation.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
The polyamines, putrescine, spermidine and spermine, are ubiquitous natural compounds that are required for eukaryotic cell growth and survival. With their primary and secondary amino groups protonated at physiological pH, these polycations interact with negatively charged macromolecules such as DNA, RNA, proteins, and phospholipids and influence their activities. Besides these polycationic functions, the polyamine spermidine is required as a donor of its butyl amine moiety in the posttranslational formation of hypusine in eukaryotic translation initiation factor 5A (eIF5A) (Scheme 10.1). This process converts an inactive eIF5A precursor to an active protein, a factor essential for protein synthesis and cell growth. Hypusine synthesis thus represents an indispensable function of polyamines in cell growth. In this chapter, we describe briefly the discovery of hypusine, identification of the hypusine-containing protein, biosynthetic pathway of hypusine and its inhibitors, and discuss the function, regulation, and mechanism of eIF5A in translation and cell proliferation. Because of space limitations, topics on the role of eIF5A isoforms in cancer [reviewed elsewhere (Caraglia et al. 2013; Park et al. 2014; Wang et al. 2013)] and other diseases, such as AIDS and diabetes, are not covered, and only a selection of references are given.
Pathways of polyamine metabolism, hypusine synthesis and eIF5A activation/inactivation. A simplified diagram of polyamine interconversion is shown on the left (vertically) and, on the right, a path leading to hypusine formation on eIF5A, catalyzed by DHS and DOHH , (horizontally) and eIF5A acetylation (vertically). eIF5A (Lys) eIF5A precursor, eIF5A (Dhp) eIF5A intermediate containing deoxyhypusine, eIF5A (Hpu) eIF5A active form containing hypusine, eIF5A(AcHpu) eIF5A containing acetylated hypusine, eIF5A(Hpu)(AcK47) hypusinated eIF5A acetylated at Lys47, SSAT1 spermidine/spermine N 1-acetyltransferase, PCAF P300/CBP-associated factor, HDAC6 histone deacetylase 6, SIRT2 sirtuin-2
1.1 Hypusine and Its Biosynthesis
A modified lysine, hypusine [N ε-(4-amino-2-hydroxybutyl)-lysine], named for its structural relationship to hydroxyputrescine and lysine, was first isolated from bovine brain extracts and the structure determined (Shiba et al. 1971). It was found to occur in all animal tissues, as the free amino acid as well as protein-bound form (Nakajima et al. 1971). In 1981, in lymphocytes cultured in a medium containing radioactive spermidine , one specific protein was radiolabeled. Hypusine was discovered to be a component of this labeled protein (Park et al. 1981), which was later identified as an eukaryotic translation initiation factor 4D (eIF-4D, current nomenclature, eIF5A ) (Cooper et al. 1983). eIF5A and hypusine exist in all eukaryotes, including yeast (for reviews, see Chen and Liu 1997; Park et al. 1993; Park 2006). Hypusine and its precursor, deoxyhypusine, also occur in Archaebacteria, but not in Eubacteria. Hypusine is formed only posttranslationally; thus, free hypusine in urine or tissue is presumed to be derived from the breakdown of eIF5A.
The biosynthesis of hypusine (Scheme 10.1) is catalyzed by two specific enzymes, deoxyhypusine synthase (DHS) and deoxyhypusine hydroxylase (DOHH) (Park 2006; Wolff et al. 2007). DHS catalyzes an NAD-dependent cleavage of spermidine with the transfer of its 4-aminobutyl moiety to the terminal N of a specific lysine of eIF5A (Lys 50 in human, Lys 51 in yeast) to form an intermediate, deoxyhypusine residue. Importantly, neither putrescine nor spermine can substitute for spermidine as a substrate, emphasizing the critical role of spermidine in cell growth.
The second enzyme, DOHH, irreversibly adds a hydroxyl group to the side chain of deoxyhypusine (Park et al. 2006; Park 2006). Like DHS, it is entirely specific for its protein substrate. It is an Fe(II)-dependent monooxygenase with a superhelical structure and a reaction mechanism distinct from other known protein hydroxylases (Kim et al. 2006b).
1.2 Additional Posttranslational Modifications in eIF5A, Acetylation and Phosphorylation
Hypusine synthesis activates eIF5A and directs it into the cytoplasmic compartment (Lee et al. 2009). Localization of hypusinated eIF5A in the cytoplasm and its association with ribosomes is critical for its role in translation . Normally, eIF5A undergoes hypusine modification immediately after its translation, and this process is irreversible. Because of this irreversibility and the long half-life of eIF5A, it may be difficult to modulate the activity of hypusinated eIF5A rapidly. In a search for additional, reversible posttranslational modifications, it was found that eIF5A can be acetylated at two conserved lysine residues, K47 (Klier et al. 1995) and K68 (in the human sequence) (Kim et al. 2006a) and also at the hypusine residue (Lee et al. 2009, 2011). Acetylation of eIF5A at Lys47 by histone acetyltransferase, PCAF, (Ishfaq et al. 2012) (Scheme 10.1), would render eIF5A inactive (Cano et al. 2008), and directs it into the nuclei (Ishfaq et al. 2012). eIF5A can also be inactivated by acetylation at the hypusine residue by the spermidine /spermine acetyltransferase 1 (SSAT1) (Scheme 10.1) (Lee et al. 2011). Although such acetylation can be demonstrated in vitro and in cells, cellular levels of acetylated eIF5A are normally quite low, and their accumulation may become significant only upon induction of acetylating enzymes or inhibition of deacetylating enzymes.
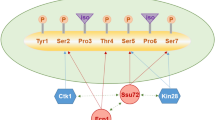
eIF5A proteins from Saccharomyces cerevisiae, Trichomonas vaginalis, and maize also undergo phosphorylation on specific Ser or Thr residues. Ser2 phosphorylation of maize eIF5A was shown to cause its sequestration in the nucleus. The significance of the nuclear accumulation of acetylated or phosphorylated eIF5A is unknown.
2 The Role of eIF5A in Cell Growth
A critical role for eIF5A in cell growth was first suggested by the observation that the hypusine -containing protein (later identified as eIF-4D/eIF5A) dramatically increased in lymphocytes upon activation with a mitogen (Cooper et al. 1983; Park et al. 1981). Hypusine-containing protein was found in other mammalian cells and a correlation between the rate of hypusine synthesis and growth was confirmed in rat hepatoma tissue culture cells (Gerner et al. 1986) and in NIH3T3 cells upon serum stimulation (Chen and Chen 1997b). Moreover, the hypusine synthesis rate was significantly elevated in Ras oncogene-transfected NIH3T3 cells compared to untransfected NIH3T3 cells (Chen and Chen 1997b). In contrast, hypusine synthesis was markedly reduced in human fibroblast cells undergoing senescence (Chen and Chen 1997a).
Convincing evidence for the essential role of eIF5A and hypusine modification on cell growth was derived from gene inactivation studies in yeast S. cerevisiae and mouse. Disruption of both eIF5A genes (Schnier et al. 1991) or a single DHS gene (Park et al. 1998; Sasaki et al. 1996) causes growth arrest and loss of viability in yeast. Similarly, inactivation of the eIF5A gene or the DHS gene in mouse leads to embryonic lethality at the early stage of gestation (E6.5) (Nishimura et al. 2012). The hydroxylation step of hypusine synthesis is not essential in yeast, as the DOHH deletion strain is viable. However, in higher eukaryotes, the second step appears to be important, because DOHH mutation leads to growth and developmental defects in Drosophila (Patel et al. 2009).
As the in vivo polyamine functions in cell growth have remained obscure for decades, it was questioned whether hypusine formation represents the main or the sole factor in the polyamine requirement in eukaryotic cells. In yeast S. cerevisiae, it seems to be the case, because a mutant strain deficient in spermidine biosynthesis can grow at a nearly normal rate with <0.2 % of spermidine, consuming up to 54 % of cellular spermidine for hypusine synthesis (Chattopadhyay et al. 2008). In mammalian cells, the question was addressed by depletion of spermidine with inhibitors of polyamine biosynthesis and supplementation with spermidine analogues. In L1210 cells depleted of spermidine by an inhibitor of S-adenosylmethionine decarboxylase, only those closely related spermidine analogues that could serve as the substrate for DHS could support long-term growth in the absence of natural spermidine (Byers et al. 1992, 1994), indicating that hypusine synthesis is a core element of the polyamine requirement. Consistent findings were obtained in DU145 prostate cancer cells depleted of spermidine by treatment with α-DFMO (α-difluoromethyl ornithine), an irreversible inhibitor of ornithine decarboxylase (Hyvonen et al. 2007). In this study, the acute phase of cytostasis (within 6 days of α-DFMO treatment) could be reversed by all methylated analogues of spermidine and spermine, but long-term growth (>9 days) could only be supported by those analogues that can serve as a precursor for hypusine synthesis. These findings further suggest two elements of polyamine function in cells: the first, a polycationic function that can be fulfilled by various analogues of spermidine, and spermine, and the second, the function of supporting hypusine synthesis that requires a close structural similarity to spermidine. These two independent aspects of polyamine function in mammalian cell growth were also suggested in an independent study that showed growth inhibition of FM3A cells upon partial depletion of spermidine and spermine by treatment with polyamine biosynthesis inhibitors, before a decline in hypusinated eIF5A (Nishimura et al. 2005). More recently, inhibition of protein synthesis and growth was also observed in cells in which cellular spermidine and spermine were rapidly depleted by overexpression of polyamine catabolic enzyme, SSAT1, before any significant decrease in hypusinated eIF5A occurred (Mandal et al. 2013). These findings reinforce the notion that, in mammalian cells, polyamines have dual functions in promoting translation , as polycations, and as a component of hypusine in eIF5A.
3 Effects of Inhibition of eIF5A Modification
As hypusine is required for the activity of eIF5A , inhibitors were developed for inhibition of DHS and as antiproliferative agents. DHS has a narrow groove for spermidine binding, and the terminal amino groups of spermidine are anchored by the conserved acidic amino acids in the active site of the enzyme. Of many diamine and triamine derivatives tested, N 1-guanyldiaminoheptane (GC7) was the most potent inhibitor, with a K i value much lower than the K m for spermidine (Jakus et al. 1993). GC7 was effective in inhibiting deoxyhypusine synthesis in cells and caused cytostasis in mammalian cells (Park et al. 1994) and in various human cancer cell lines (Shi et al. 1996). It also displayed antitumor effects in an animal tumor model (Jasiulionis et al. 2007)
DOHH is a mono-oxygenase with a di-iron active center (Kim et al. 2006b; Park et al. 2006) and is inhibited by a panel of iron chelators, such as mimosine, ciclopiroxFootnote 1, or deferiprone.Footnote 2 These compounds caused an arrest in cell-cycle progression at the G1/S boundary, coincident with inhibition of deoxyhypusine hydroxylation (Hanauske-Abel et al. 1994). Ciclopirox inhibits endothelial cell growth and angiogenesis in vitro (Clement et al. 2002) and exerts antitumor effects in the MDA-231 xenograft in mice (Zhou et al. 2010). However, the possibility that these compounds can have other cellular targets in vivo cannot be ignored and complicates the interpretation with regard to the involvement of eIF5A .
4 The Mechanism of Action of eIF5A in Translation
eIF5A (eIF4D) was initially isolated as a factor that stimulates methionyl-puromycin synthesis, a model assay for the first peptide bond formation (Kemper et al. 1976). Although it was named as a translation initiation factor then, recent work has shown that eIF5A has a distinct effect on the elongation step of translation as measured by polysome profiles (Gregio et al. 2009; Saini et al. 2009). Dever and associates (Gutierrez et al. 2013) have reported evidence that eIF5A, similar to its bacterial orthologue EF-P (Doerfel et al. 2013; Ude et al. 2013), relieves ribosome stalling at consecutive proline residues and thereby facilitates translation elongation of proteins containing multiple proline residues. A docking model of eIF5A bound to the translating ribosome predicts its hypusine side chain directed toward the peptidyl transferase center, consistent with its proposed function in translation elongation (Gutierrez et al. 2013). Future efforts will be directed to identify more eIF5A target motifs and to elucidate the precise contribution of polyamine-derived side chain of hypusine in the peptidyl transferase reaction.
5 Concluding Remarks
In spite of abundant genetic and biochemical evidence for the essentiality of polyamines in eukaryotic organisms, their precise function was not well understood for decades. One missing link was found with the discovery of the hypusine pathway and the role of spermidine for this modification and thereby in translation and cell growth. Since the first isolation of hypusine as a chemical entity in 1971, of eIF5A in 1976, and the identification of eIF5A as the single cellular protein containing hypusine in 1983, it has taken decades to establish its pathway (Scheme 10.1) and to determine its role in translation elongation, presumably on specific eIF5A target motifs, including consecutive proline residues. Although eIF5A activity in translation has been characterized biochemically, it needs to be related to cellular changes at the level of proteome and phenotypes. Many cellular functions have been proposed for eIF5A isoforms (not mentioned in this chapter for reasons of space constraints), including their roles in nuclear export, mRNA turnover/NMD (nonsense-mediated decay), actin cytoskeletal organization, cell wall integrity, cell-cycle progression, apoptosis, autophagy, and intracellular protein trafficking. eIF5A has also been implicated in pathological conditions such as cancer, inflammation, human immunodeficiency virus (HIV)1 infection, and diabetes. It is as yet unclear whether the pleotropic effects are caused by changes in the cellular proteome resulting from a dysfunction of eIF5A in translation elongation or whether eIF5A is a multifunctional protein. Future investigations are warranted to solve the mystery of action of this novel protein.
Notes
- 1.
Approved anti-fungal drug.
- 2.
Approved anti-thalassemia drug.
Abbreviations
- DHS :
-
Deoxyhypusine synthase
- DOHH :
-
Deoxyhypusine hydroxylase
- EF-P:
-
Bacterial elongation factor P
- eIF5A :
-
Eukaryotic initiation factor 5A
- GC7:
-
N1-guanyl-1,7-diaminoheptane
- SSAT1:
-
Spermidine /spermine acetyltransferase 1
References
Byers TL, Ganem B, Pegg AE (1992) Cytostasis induced in L1210 murine leukaemia cells by the S-adenosyl-l-methionine decarboxylase inhibitor 5′-([(Z)-4-amino-2-butenyl]methylamino)-5′-deoxyadenosine may be due to hypusine depletion. Biochem J 287:717–724
Byers TL, Lakanen JR, Coward JK et al (1994) The role of hypusine depletion in cytostasis induced by S-adenosyl-l-methionine decarboxylase inhibition: new evidence provided by 1-methylspermidine and 1,12-dimethylspermine. Biochem J 303:363–368
Cano VS, Jeon GA, Johansson HE et al (2008) Mutational analyses of human eIF5A–1: identification of amino acid residues critical for eIF5A activity and hypusine modification. FEBS J 275:44–58
Caraglia M, Park MH, Wolff EC et al (2013) eIF5A isoforms and cancer: two brothers for two functions? Amino Acids 44:103–109
Chattopadhyay MK, Park MH, Tabor H (2008) Hypusine modification for growth is the major function of spermidine in Saccharomyces cerevisiae polyamine auxotrophs grown in limiting spermidine. Proc Natl Acad Sci USA 105:6554–6559
Chen ZP, Chen KY (1997a) Dramatic attenuation of hypusine formation on eukaryotic initiation factor 5A during senescence of IMR-90 human diploid fibroblasts. J Cell Physiol 170:248–254
Chen ZP, Chen KY (1997b) Marked elevation of hypusine formation activity on eukaryotic initiation factor 5A in v-HA-RAS transformed mouse NIH3T3 cells. Cancer Lett 115:235–241
Chen KY, Liu AY (1997) Biochemistry and function of hypusine formation on eukaryotic initiation factor 5A. Biol Signals 6:105–109
Clement PM, Hanauske-Abel HM, Wolff EC et al (2002) The antifungal drug ciclopirox inhibits deoxyhypusine and proline hydroxylation, endothelial cell growth and angiogenesis in vitro. Int J Cancer 100:491–498
Cooper HL, Park MH, Folk JE et al (1983) Identification of the hypusine-containing protein hy+ as translation initiation factor eIF-4D. Proc Natl Acad Sci USA 80:1854–1857
Doerfel LK, Wohlgemuth I, Kothe C et al (2013) EF-P is essential for rapid synthesis of proteins containing consecutive proline residues. Science 339:85–88
Gerner EW, Mamont PS, Bernhardt A et al (1986) Post-translational modification of the protein-synthesis initiation factor eIF-4D by spermidine in rat hepatoma cells. Biochem J 239:379–386
Gregio AP, Cano VP, Avaca JS et al (2009) eIF5A has a function in the elongation step of translation in yeast. Biochem Biophys Res Commun 380:785–790
Gutierrez E, Shin BS, Woolstenhulme CJ et al (2013) eIF5A promotes translation of polyproline motifs. Mol Cell 51:35–45
Hanauske-Abel HM, Park MH, Hanauske AR et al (1994) Inhibition of the G1-S transition of the cell cycle by inhibitors of deoxyhypusine hydroxylation. Biochim Biophys Acta 1221:115–124
Hyvonen MT, Keinanen TA, Cerrada-Gimenez M et al (2007) Role of hypusinated eukaryotic translation initiation factor 5A in polyamine depletion-induced cytostasis. J Biol Chem 282:34700–34706
Ishfaq M, Maeta K, Maeda S et al (2012) Acetylation regulates subcellular localization of eukaryotic translation initiation factor 5A (eIF5A). FEBS Lett 586:3236–3241
Jakus J, Wolff EC, Park MH et al (1993) Features of the spermidine-binding site of deoxyhypusine synthase as derived from inhibition studies. Effective inhibition by bis- and mono-guanylated diamines and polyamines. J Biol Chem 268:13151–13159
Jasiulionis MG, Luchessi AD, Moreira AG et al (2007) Inhibition of eukaryotic translation initiation factor 5A (eIF5A) hypusination impairs melanoma growth. Cell Biochem Funct 25:109–114
Kemper WM, Berry KW, Merrick WC (1976) Purification and properties of rabbit reticulocyte protein synthesis initiation factors M2Balpha and M2Bbeta. J Biol Chem 251:5551–5557
Kim SC, Sprung R, Chen Y et al (2006a) Substrate and functional diversity of lysine acetylation revealed by a proteomics survey. Mol Cell 23:607–618
Kim YS, Kang KR, Wolff EC et al (2006b) Deoxyhypusine hydroxylase is a Fe(II)-dependent, HEAT-repeat enzyme. Identification of amino acid residues critical for Fe(II) binding and catalysis [corrected]. J Biol Chem 281:13217–13225
Klier H, Csonga R, Joao HC et al (1995) Isolation and structural characterization of different isoforms of the hypusine-containing protein eIF-5A from HeLa cells. Biochemistry 34:14693–14702
Lee SB, Park JH, Kaevel J et al (2009) The effect of hypusine modification on the intracellular localization of eIF5A. Biochem Biophys Res Commun 383:497–502
Lee SB, Park JH, Folk JE et al (2011) Inactivation of eukaryotic initiation factor 5A (eIF5A) by specific acetylation of its hypusine residue by spermidine/spermine acetyltransferase 1 (SSAT1). Biochem J 433:205–213
Mandal S, Mandal A, Johansson HE et al (2013) Depletion of cellular polyamines, spermidine and spermine, causes a total arrest in translation and growth in mammalian cells. Proc Natl Acad Sci USA 110:2169–2174
Nakajima T, Matsubayashi T, Kakimoto Y et al (1971) Distribution of hypusine, N 6-(4-amino-2-hydroxybutyl)-2,6-diaminohexanoic acid, in mammalian organs. Biochim Biophys Acta 252:92–97
Nishimura K, Murozumi K, Shirahata A et al (2005) Independent roles of eIF5A and polyamines in cell proliferation. Biochem J 385:779–785
Nishimura K, Lee SB, Park JH et al (2012) Essential role of eIF5A-1 and deoxyhypusine synthase in mouse embryonic development. Amino Acids 42:703–710
Park MH (2006) The post-translational synthesis of a polyamine-derived amino acid, hypusine, in the eukaryotic translation initiation factor 5A (eIF5A). J Biochem 139:161–169
Park MH, Cooper HL, Folk JE (1981) Identification of hypusine, an unusual amino acid, in a protein from human lymphocytes and of spermidine as its biosynthetic precursor. Proc Natl Acad Sci USA 78:2869–2873
Park MH, Wolff EC, Folk JE (1993) Hypusine: its post-translational formation in eukaryotic initiation factor 5A and its potential role in cellular regulation. Biofactors 4:95–104
Park MH, Wolff EC, Lee YB et al (1994) Antiproliferative effects of inhibitors of deoxyhypusine synthase. Inhibition of growth of Chinese hamster ovary cells by guanyl diamines. J Biol Chem 269:27827–27832
Park MH, Joe YA, Kang KR (1998) Deoxyhypusine synthase activity is essential for cell viability in the yeast Saccharomyces cerevisiae. J Biol Chem 273:1677–1683
Park JH, Aravind L, Wolff EC et al (2006) Molecular cloning, expression, and structural prediction of deoxyhypusine hydroxylase: a HEAT-repeat-containing metalloenzyme. Proc Natl Acad Sci USA 103:51–56
Park MH, Mandal S, Mandal A et al (2014) eIF5A and cancer. In: Parsyan A (ed) Translation and cancer: applications in medicine. Springer, New York, pp 223–232
Patel PH, Costa-Mattioli M, Schulze KL et al (2009) The Drosophila deoxyhypusine hydroxylase homologue nero and its target eIF5A are required for cell growth and the regulation of autophagy. J Cell Biol 185:1181–1194
Saini P, Eyler DE, Green R et al (2009) Hypusine-containing protein eIF5A promotes translation elongation. Nature (Lond) 459:118–121
Sasaki K, Abid MR, Miyazaki M (1996) Deoxyhypusine synthase gene is essential for cell viability in the yeast Saccharomyces cerevisiae. FEBS Lett 384:151–154
Schnier J, Schwelberger HG, Smit-McBride Z et al (1991) Translation initiation factor 5A and its hypusine modification are essential for cell viability in the yeast Saccharomyces cerevisiae. Mol Cell Biol 11:3105–3114
Shi XP, Yin KC, Ahern J et al (1996) Effects of N 1-guanyl-1,7-diaminoheptane, an inhibitor of deoxyhypusine synthase, on the growth of tumorigenic cell lines in culture. Biochim Biophys Acta 1310:119–126
Shiba T, Mizote H, Kaneko T et al (1971) Hypusine, a new amino acid occurring in bovine brain. Isolation and structural determination. Biochim Biophys Acta 244:523–531
Ude S, Lassak J, Starosta AL et al (2013) Translation elongation factor EF-P alleviates ribosome stalling at polyproline stretches. Science 339:82–85
Wang FW, Guan XY, Xie D (2013) Roles of eukaryotic initiation factor 5A2 in human cancer. Int J Biol Sci 9:1013–1020
Wolff EC, Kang KR, Kim YS et al (2007) Posttranslational synthesis of hypusine: evolutionary progression and specificity of the hypusine modification. Amino Acids 33:341–350
Zhou H, Shen T, Luo Y et al (2010) The antitumor activity of the fungicide ciclopirox. Int J Cancer 127:2467–2477
Acknowledgments
This research was supported by the Intramural Research Program of the NIH/NIDCR.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Japan
About this chapter
Cite this chapter
Wolff, E.C., Park, M.H. (2015). Role of the Polyamine Spermidine as a Precursor for Hypusine Modification in eIF5A. In: Kusano, T., Suzuki, H. (eds) Polyamines. Springer, Tokyo. https://doi.org/10.1007/978-4-431-55212-3_10
Download citation
DOI: https://doi.org/10.1007/978-4-431-55212-3_10
Published:
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-55211-6
Online ISBN: 978-4-431-55212-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)