Abstract
Early structural studies on glycoproteins revealed bi-, tri-, and tetra-antennary N-glycans in which GlcNAc residues were linked to a conserved trimannosyl core, prompting the search for the GlcNAc-transferases that catalyzed the addition of each GlcNAc residue. Mannosyl (alpha-1,3-)-glycoprotein beta-1,2-N-acetylglucosaminyltransferase I (MGAT1), originally termed N-acetylglucosaminyltransferase I, abbreviated GlcNAc-TI, was the first N-glycan branching GlcNAc-transferase for which an assay was developed (Gottlieb et al. 1975; Stanley et al. 1975). MGAT1 catalyzes the transfer of GlcNAc from UDP-GlcNAc to the terminal α1,3-linked Man in Man5GlcNAc2Asn to initiate the synthesis of hybrid and complex N-linked glycans in multicellular organisms (reviewed in Kornfeld and Kornfeld 1985). It is not found in yeast or bacteria. The human gene MGAT1 resides on chromosome 5q35 (Kumar et al. 1992), covering 25.12 kb, from 180,242,651 to 180,217,536 (NCBI 37, August 2010) on the reverse strand (Thierry-Mieg and Thierry-Mieg 2006). The mouse gene, Mgat1, is on chromosome 11 (Pownall et al. 1992). Northern blot analyses revealed two transcripts of ˜2.9 and ˜3.3 kb present in most mammalian tissues, with the shorter transcript predominating in liver, and the longer transcript in brain (Yang et al. 1994; Yip et al. 1997). However, the human MGAT1 locus is complex with 30 introns, seven predicted alternative promoters, ten validated poly[A] addition sites >30 transcripts that encode 11 protein isoforms, with three containing the complete coding sequence (Thierry-Mieg and Thierry-Mieg 2006). The coding region is in a single exon and the Mgat1 gene is ubiquitously expressed.
Access provided by Autonomous University of Puebla. Download reference work entry PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
Introduction
Early structural studies on glycoproteins revealed bi-, tri-, and tetra-antennary N-glycans in which GlcNAc residues were linked to a conserved trimannosyl core, prompting the search for the GlcNAc-transferases that catalyzed the addition of each GlcNAc residue. Mannosyl (alpha-1,3-)-glycoprotein beta-1,2-N-acetylglucosaminyltransferase I (MGAT1), originally termed N-acetylglucosaminyltransferase I, abbreviated GlcNAc-TI, was the first N-glycan branching GlcNAc-transferase for which an assay was developed (Gottlieb et al. 1975; Stanley et al. 1975). MGAT1 catalyzes the transfer of GlcNAc from UDP-GlcNAc to the terminal α1,3-linked Man in Man5GlcNAc2Asn to initiate the synthesis of hybrid and complex N-linked glycans in multicellular organisms (reviewed in Kornfeld and Kornfeld 1985). It is not found in yeast or bacteria. The human gene MGAT1 resides on chromosome 5q35 (Kumar et al. 1992), covering 25.12 kb, from 180,242,651 to 180,217,536 (NCBI 37, August 2010) on the reverse strand (Thierry-Mieg and Thierry-Mieg 2006). The mouse gene, Mgat1, is on chromosome 11 (Pownall et al. 1992). Northern blot analyses revealed two transcripts of ∼2.9 and ∼3.3 kb present in most mammalian tissues, with the shorter transcript predominating in liver, and the longer transcript in brain (Yang et al. 1994; Yip et al. 1997). However, the human MGAT1 locus is complex with 30 introns, seven predicted alternative promoters, ten validated poly[A] addition sites, >30 transcripts that encode 11 protein isoforms, with three containing the complete coding sequence (Thierry-Mieg and Thierry-Mieg 2006). The coding region is in a single exon and the Mgat1 gene is ubiquitously expressed.
Databanks
IUBMB enzyme nomenclature: EC 2.4.1.101
Mannosyl (alpha-1,3-)-glycoprotein beta-1,2-N-acetylglucosaminyltransferase (MGAT1)
Species | Gene symbol | Uniprot ID | GenBank accession number | PDB accession number |
|---|---|---|---|---|
H. sapiens | MGAT1 | AAA75523 | M55621 | NA |
AAA52563 | ||||
O. cuniculus | Rabgnt1 | AAA31493 | M57301 | 1FO8, 1FOA, 2AM3 |
2AM4, 2AM5, 2APC | ||||
R. norvegicus | Ratnagt | BAA03807 | D16302 | NA |
Q09325 | AB012874(5′utr) | |||
AB012878(5′utr) | ||||
M. musculus | Mgat1 | AAA40478 | L07037 | NA |
AAA37698 | X77487(5′utr) | |||
C. griseus | Mgat1 | AAC52872 | U65791 | NA |
AF343963 | ||||
M. auratus | Mgat1 | AAD04130 | AF087456 | NA |
D. melanogaster | Mgat1 | AAF70177 | AF251495 | NA |
C. elegans | gly-12 | AAD03023 | AF082011 | NA |
gly-13 | AAD03022 | AF082010 | NA | |
gly-14 | AAD03024 | AF082012 | NA | |
A. thaliana | GlcNAcT-I | JC7084 | AJ243198 | NA |
N. tabacum | CGL | CAB70464 | NA | NA |
Name and History
Mannosyl (alpha-1,3-)-glycoprotein beta-1,2-N-acetylglucosaminyltransferase I has numerous names and abbreviations. It was first called UDP-GlcNAc:α-d-mannoside β2-N-acetylglucosaminyltransferase I, abbreviated as GlcNAc-TI (or GlcNAcT-I, GnTI or NAGT).
The convention at this time is to use the symbol designated by the HUGO gene nomenclature committee for a gene and the protein it encodes. Thus, the MGAT1 human gene encodes MGAT1 (mannosyl (alpha-1,3-)-glycoprotein beta-1,2-N-acetylglucosaminyltransferase I). The mouse protein is also MGAT1 but the gene is Mgat1. MGAT1 was the first GlcNAc-transferase activity identified in a crude cell extract to transfer GlcNAc from UDP-GlcNAc to glycoproteins containing the trimannoside, N-linked core (Manα1,3Manα1,6)Manβ1,4GlcNAcβ1,4GlcNAcβ1Asn. An assay for GlcNAc-TI was developed in cell mutants that were resistant to the cytotoxicity of several plant lectins (Stanley 1984). N-glycan containing acceptors derived by sequential glycosidase digestion of glycoproteins including α1-acid glycoprotein, fetuin, and IgG were tested in cell extracts with a variety of nucleotide-sugar donors. Several independent CHO mutants including PhaR1-1 (Stanley et al. 1975) and Clone 15B (Gottlieb et al. 1975), and a BHK mutant RicR 14 (Meager et al. 1975) had a specific reduction in the ability to transfer GlcNAc from UDP-GlcNAc to mannose terminating acceptors. When a more defined set of bi-antennary complex N-glycan glycopeptide acceptors was used, CHO cells were found to possess at least two GlcNAc-transferase activities (Narasimhan et al. 1977). MGAT1 was the activity missing from PhaR1-1 CHO cells assayed with UDP-GlcNAc and the trimannosyl glycopeptide (Manα1,3Manα1,6)Manβ1,4GlcNAcβ1,4GlcNAcβ1 as acceptor (Narasimhan et al. 1977). CHO cells lacking MGAT1 activity were subsequently termed Lec1 (Stanley 1983).
Structure
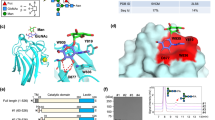
MGAT1 is a type II transmembrane protein of 447 amino acids (Kumar et al. 1990; Sarkar et al. 1991) that resides in the medial Golgi and functions primarily as a homodimer or heterodimer (Hassinen et al. 2011). It carries O-glycans but no N-glycans (Hoe et al. 1995). Crystal structures of the catalytic domain of rabbit MGAT1 alone (Unligil et al. 2000) and in complex with various substrate analogues (Gordon et al. 2006) have been deposited in PDB (see section “Databanks”). Point mutations that inactivate MGAT1 include the conversion of the conserved Cys at position 123 to Arg (Puthalakath et al. 1996), conversion of the conserved Gly at position 320 to Asp (Opat et al. 1998) and most recently, three inactivating mutations R415K, D291N, and P138L (Zhong et al. 2012). MGAT1 lacking the first 106 amino acids including the cytoplasmic, transmembrane, and stem domains is active, but removal of the C-terminal seven amino acids results in a 40 % reduction in activity (Sarkar et al. 1998). Missense mutations that dramatically increase the apparent Km of MGAT1 for both substrates were identified in Lec1A CHO mutants and include conversion of Asp 212 to Asn or Arg 303 to Tryp (Chen et al. 2001). There are also a number of deletion mutations (Chen and Stanley 2003; Zhong et al. 2012).
Enzyme Activity Assay and Substrate Specificity
When MGAT1 was partially purified from rat liver, it was found to prefer the Man5 acceptor Manα1,3Manα1,6(Manα1,6Manα1,3)Manβ1,4GlcNAcβ1,4GlcNAcβ1Asn over the (Manα1,3Manα1,6)Manβ1,4GlcNAcβ1,4GlcNAcβ1Asn acceptor originally used (Oppenheimer and Hill 1981). Since glycoproteins from Lec1 CHO cells have Man5GlcNAc2Asn in place of complex N-glycans (Robertson et al. 1978; Tabas et al. 1978), the preference of MGAT1 for Man5GlcNAc2Asn was consistent with it being an important in vivo substrate of MGAT1 according to the following scheme: Fig. 17.1.
The addition of the β1,2-linked GlcNAc to Man5GlcNAc2Asn generates a substrate for a-mannosidase II (now termed MAN2A1) and initiates the synthesis of complex N-glycans through the subsequent action of MGAT2 (reviewed in Kornfeld and Kornfeld 1985). Alternatively, if α-mannosidase II does not act, a hybrid structure may be formed by the addition of Gal and potentially sialic acid to the β1,2-linked GlcNAc of GlcNAcβ1,2Man5GlcNAc2Asn. In mice lacking α-mannosidase II, complex N-glycans are formed in many tissues due to a complementing activity MAN2A2 that appears to be redundant with MAN2A1 (Akama et al. 2006). A mannosidase activity termed α-mannosidase III, that is enriched in Golgi fractions and removes mannose residues from Man5GlcNAc2Asn, was identified in mouse tissues and could generate Man3GlcNAc2Asn (which is known from the initial in vitro studies to be a substrate for MGAT1 (Gottlieb et al. 1975; Stanley et al. 1975; Narasimhan et al. 1977), thereby allowing α-mannosidase II-deficient mice to synthesize complex N-glycans. Acceptor specificity studies show that MGAT1 will not transfer GlcNAc to terminal α-mannose residues in a bi-antennary N-glycan in which the α1,6-mannose is substituted at the O-2 position or the β1,4-mannose is substituted at the O-4 position (Nishikawa et al. 1988).
Optimal assay conditions were determined for purified rabbit liver GlcNAc-TI as follows (Nishikawa et al. 1988): 0.25 mM Man5GlcNAc2Asn in 100 mM 2-(N-morpholino)ethanesulfonate acid (MES) pH 6.1, 0.5 mM UDP-14C-GlcNAc, 20 mM MnCl2, 5 mM AMP (as pyrophosphorylase inhibitor), 1.0 % Triton X-100, 100 mM GlcNAc (as hexosaminidase inhibitor), bovine serum albumin at 5 mg/ml (enzyme stabilizer), and enzyme (usually ∼0.1 mU) in a volume of 50 μl. After 30 min at 37° C, the reaction is terminated by the addition of ice-cold water containing 20 mM EDTA and the products fractionated on a 1 ml ion exchange column (AG-1 × 8, chloride form) eluted with water. The flow-through contains 14C-GlcNAcMan5GlcNAc2Asn product, unmodified Man5GlcNAc2Asn, and free 14C-GlcNAc generated by hydrolysis of UDP-14C-GlcNAc. Alternatively, products may be fractionated on a 1.5 ml column of concanavalin A (Con A)-Sepharose to which the 14C-GlcNAcMan5GlcNAc2Asn product binds and is eluted with 100 mM α-methylmannoside. Neither UDP-GlcNAc nor free GlcNAc bind to Con A, and thus background due to hydrolysis of UDP-GlcNAc is eliminated. The background of the MGAT1 assay is determined from labeled products generated by boiled enzyme, and from enzyme incubated with UDP-14C-GlcNAc in the absence of Man5GlcNAc2Asn acceptor.
To assay MGAT1 in cell or tissue extracts, the following conditions, optimized for nonionic detergent extracts of CHO cells may be used: Cells are washed three times in saline and extracted in 1.5 % NP-40 in cold distilled water containing protease inhibitors (75 μl detergent solution per 107 packed cells). After 10 min on ice, the extract is centrifuged at low speed to remove nuclei and 5–20 ml extract containing 50–100 μg protein is added to an assay tube on ice containing, in a final volume of 40 μl, 62.5 mM MES buffer pH 6.25, 25 mM MnCl2, 1 mM Man5GlcNAc2Asn, 1 mM UDP-3H-GlcNAc (specific activity ∼10,000 cpm per nmole), and 50–100 μg protein. After 30–90 min at 37° C, the reaction is terminated by the addition of 1 ml of cold Con A buffer (1 M Na acetate, 1 mM MnCl2, 1 mM MgCl2 and 1 mM CaCl2, pH 7.0). The mixture is subsequently passed over a 1.5 ml column of Con A-Sepharose, washed with ten column volumes of Con A buffer to remove UDP-3H-GlcNAc and 3H-GlcNAc, and the radiolabeled product 3H-GlcNAcMan5GlcNAc2Asn is eluted with a 6 ml aliquot of 200 mM α-methylmannoside in Con A buffer. The specific activity of GlcNAc-TI in CHO cell extracts is ∼5–10 nmol/h/mg protein. CHO mutants in the Lec1A group have a missense mutation that increases the Km for both substrates and have barely detectable GlcNAc-TI activity under these assay conditions. Their activity becomes normal however, if the pH of the assay is increased to 7.5, the UDP-GlcNAc concentration to 15 mM and the Man5GlcNAc2Asn concentration to 5 mM (Chaney and Stanley 1986; Chen et al. 2001).
Preparation
MGAT1 was purified ∼64,000-fold to apparent homogeneity from a Triton X-100 extract of rabbit liver using three affinity chromatography steps on UDP-hexanolamine followed by two affinity chromatography steps on 5-Hg-UDP-GlcNAc (Nishikawa et al. 1988). Km values of purified rabbit liver MGAT1 for UDP-GlcNAc and Man5GlcNAc2Asn substrates were ∼0.04 mM and ∼2 mM, respectively. The Vmax was ∼16 μmol/min/mg protein and the specific activity was ∼20 μmol/min/mg protein. SDS-PAGE analysis revealed a major species of 45 kd and minor species of 50 and 54 kd. Since cleavage of glycosyltransferases often occurs in the Golgi at a position just beyond the stem region, and since mammalian MGAT1 is known to be O-glycosylated but not N-glycosylated (Hoe et al. 1995), the three forms probably represented O-glycosylated and/or proteolyzed forms of MGAT1. MGAT1 may also be produced in active form in bacteria, yeast, insect, plant, and mammalian cells. Plant, Drosophila, and C. elegans MGAT1 sequences include N-glycan Asn-X-Ser/Thr consensus sites, though their positions in the protein are not conserved, and it is not known whether they are utilized.
Biological Aspects
Since a lack of MGAT1 should not alter the synthesis or processing of N-glycans in the endoplasmic reticulum (ER), nor in the cis Golgi, glycoproteins are not expected be compromised in their interactions with ER chaperones like calnexin and calreticulin, and lysosomal enzymes will acquire their usual complement of Man-6-phosphate residues for targeting to lysosomes. Thus, a lack of MGAT1 affects N-glycan structures later in the secretory pathway leading to a dramatic alteration in the array of N-glycans expressed at the cell surface because all complex and hybrid N-glycans are replaced by Man5GlcNAc2Asn. While this change may not significantly affect biological or structural properties of a recombinant glycoprotein, it has a major effect on the tissue targeting of recombinant glycoproteins. Glycoproteins with oligomannosyl N-glycans are targeted to the reticuloendothelial system. In addition, if mammalian cells are stressed by reducing the serum concentration of culture medium, growth is slowed and growth factor signaling is reduced in the absence of branched, complex N-glycans (Song et al. 2010; Beheshti Zavareh et al. 2012).
Knockout, Knockdown, and Transgenic Organisms
Mice
In contrast to cells and plants (von Schaewen 1993), mammals have an absolute requirement for GlcNAc-TI during early embryogenesis. Mice with a null mutation in the Mgat1 gene die at E9.5 (Ioffe and Stanley 1994; Metzler et al. 1994). They are underdeveloped with fewer somites, a tubelike heart, an open neural tube, and some are altered in left-right symmetry. However, the cause of death of Mgat1 −/−embryos is not known. Maternal Mgat1 gene transcripts rescue the earliest embryos, and thus it is still not known whether hybrid or complex N-glycans are required for blastocyst formation or for implantation (Campbell et al. 1995; Ioffe et al. 1997).
In order to identify a cell type that requires GlcNAc-TI to develop or differentiate, Mgat1 −/−embryonic stem (ES) cells with an inert transgene were developed and tracked in E10 to E16.5 chimeric embryos by DNA/DNA in situ hybridization (Ioffe et al. 1996). These experiments showed that complex and/or hybrid N-glycans are essential for the formation of the organized layer of bronchial epithelium. Since heterozygote Mgat1 +/−WW6 cells also contributed very poorly to organized bronchial epithelium (Ioffe et al. 1996), it is possible that some form of lung disorder could arise in humans with only one active MGAT1 allele.
Other biological insights into the functions of complex N-glycans have come from the generation of tissue-specific Mgat1 gene deletion or knockdown using RNAi. Neuronal deletion of floxed Mgat1 was performed in Syn1-Cre recombinase transgenic mice (Ye and Marth 2004). The Syn1 rat promoter is expressed just after mid-gestation and is pan-neuronal at birth. Mice lacking MGAT1 in neurons were born but died between birth and up to ∼18 weeks with locomotor dysfunction, tremors, and paralysis before death. Deletion of Mgat1 in primary oocytes using ZP3-Cre caused fewer oocytes to ovulate, defects in preovulatory follicles and cumulus mass and ∼50 % of those that did ovulate developed poorly (Shi et al. 2004; Williams and Stanley 2009). A Stra8-iCre transgene was used to delete Mgat1 in spermatogonia and resulted in a block in spermatogenesis at the spermatid stage and infertility (Batista et al. 2012). Finally, knockdown of MGAT1 in prostate cancer cells reduces tumor progression markedly in terms of both prostate tumor size and metastasis (Beheshti Zavareh et al. 2012).
Drosophila
Interestingly, deletion of Drosophila MGAT1 causes fusion of the β-lobes of mushroom bodies in the central nervous system, severe locomotor defects, and shortened lifespan that is rescued by Mgat1 expression in neurons, which also increases lifespan in wild type flies (Sarkar et al. 2010; Schachter 2010).
C. elegans
The worm has three genes encoding MGAT1, gly-12, gly-13, and gly-14, and the triple knockout worm has no obvious phenotype although it essentially lacks N-glycan products of MGAT1 (Zhu et al. 2004).
Human Disease
Studies demonstrating that T cells in mice use complex N-glycans for regulation of their activation state led to a GWAS study of genes coding for GlcNAc-transferases of the N-glycan pathway in cohorts with multiple sclerosis. Intriguingly, disease-associated SNPs in MGAT1 were found to increase MGAT1 activity and thereby decrease N-glycan branching (Mkhikian et al. 2011). This leads to prolonged activation of T cells and may be an important factor in the development and/or progression of multiple sclerosis.
Future Perspectives
Crystal structures of the catalytic Golgi lumenal domain of rabbit GnT-I/MGAT1 in the presence and absence of UDP-GlcNAc analogues have allowed modeling of Manα1,3Manβ1 (Gordon et al. 2006). The missense mutations identified in Lec1A CHO mutants (Chen et al. 2001) and DUKX Lec1 mutants (Zhong et al. 2012) alter residues conserved in MGAT1 from plants through lower organisms and mammals that are important in metal binding and catalysis (Asp212) or stabilization of a structural element involved in UDP-GlcNAc binding and catalysis (R303, R415, D291, P138). It is now important to obtain a crystal structure with both UDP-GlcNAc and Man5GlcNAc2Asn bound to MGAT1 and to crystallize MGAT1 mutants with a point mutation that weakens or inactivates the enzyme. Crystal structures of MGAT1 from lower organisms will provide insight into enzyme mechanism as they are only about 30–40 % identical to mammalian MGAT1 in amino acid sequence.
Glycosylation engineering will remain very important as therapeutic recombinant antibodies continue to be developed. Lec1 CHO cells with Mgat1 mutations are available as single mutants or in combination with other glycosylation mutations for optimal glycosylation engineering (Stanley 1989). They can be used to produce recombinant glycoproteins that will target to the reticuloendothelial system or reduce the N-glycan heterogeneity of glycoproteins that prove difficult to crystallize. Recombinant glycoproteins produced in the Lec3.2.8.1 CHO mutant and treated with endoglycosidase H will have only GalNAc at O-glycan sites and only one GlcNAc at N-glycan sites. Six independent Lec1 mutants, including the line available from the American Type Culture Collection, each have a different mutation that leads to a premature stop codon (Chen and Stanley 2003), and no revertants have been isolated.
Another question of interest for the future is whether mammals have additional genes that encode an MGAT1 activity. C. elegans has three such genes – gly-12, gly-13, and gly-14 (Chen et al. 1999). All encode type II membrane proteins typical of Golgi glycosyltransferases. However, only gly-12 and gly-14 gave MGAT1 activity when expressed in insect cells. Whereas gly-12 and gly-13 are expressed ubiquitously in the adult, gly-14 is expressed only in gut cells. In mouse embryos, it is clear that no other gene product rescues Mgat1 −/−mouse embryos from death at E9.5 during embryogenesis. However, one or more genes related to Mgat1 could be expressed in the adult. Tissue-specific knockout of a floxed Mgat1 gene may reveal such complementary genes.
Conditional knockout of the mouse Mgat1 gene in specific tissues will identify cell types that require complex or hybrid N-glycans for development or differentiation. Chimera experiments with Mgat1 −/−ES cells in Rag2 −/−blastocysts will determine whether T and/or B cells require complex or hybrid N-glycans to be generated or to function in immunity. The important association of MGAT1 SNPs with multiple sclerosis, a disease proposed to be autoimmune in origin (Mkhikian et al. 2011), provides a strong basis for analyzing MGAT1 SNPs in other autoimmune disorders. While humans with a mutation in one MGAT1 allele would not be expected to have developmental problems, they may have an altered susceptibility to lung disease (Ioffe et al. 1996) or other subtle problems like autism. No human MGAT1 mutation has yet been found to be the basis of a Congenital Disorder of Glycosylation. However, this could certainly occur, provided the mutation weakened, but did not inactivate, MGAT1 activity.
References
Akama TO, Nakagawa H, Wong NK, Sutton-Smith M, Dell A et al (2006) Essential and mutually compensatory roles of (alpha)-mannosidase II and (alpha)-mannosidase IIx in N-glycan processing in vivo in mice. Proc Natl Acad Sci USA 103:8983–8988
Batista F, Lu L, Williams SA, Stanley P (2012) Complex N-Glycans are essential, but core 1 and 2 mucin O-Glycans, O-fucose lycans, and NOTCH1 are dispensable, for mammalian spermatogenesis. Biol Reprod 86(179):1–12
Beheshti Zavareh R, Sukhai MA, Hurren R, Gronda M, Wang X, Simpson CD, Maclean N, Zih F, Ketela T, Swallow CJ, Moffat J, Rose DR, Schachter H, Schimmer AD, Dennis JW (2012) Suppression of cancer progression by MGAT1 shRNA knockdown. PLoS One 7:e43721
Boscher C, Dennis JW, Nabi IR (2011) Glycosylation, galectins and cellular signaling. Curr Opin Cell Biol 23:383–392
Campbell RM, Metzler M, Granovsky M, Dennis JW, Marth JD (1995) Complex asparagine-linked oligosaccharides in Mgat1-null embryos. Glycobiology 5:535–543
Chaney W, Stanley P (1986) Lec1A Chinese hamster ovary cell mutants appear to arise from a structural alteration in N-acetylglucosaminyltransferase I. J Biol Chem 261:10551–10557
Chen W, Stanley P (2003) Five Lec1 CHO cell mutants have distinct Mgat1 gene mutations that encode truncated N-acetylglucosaminyltransferase I. Glycobiology 13:43–50
Chen S, Zhou S, Sarkar M, Spence AM, Schachter H (1999) Expression of three Caenorhabditis elegans N-acetylglucosaminyltransferase I genes during development. J Biol Chem 274:288–297
Chen W, Unligil UM, Rini JM, Stanley P (2001) Independent Lec1A CHO glycosylation mutants arise from point mutations in N-Acetylglucosaminyltransferase I that reduce affinity for both substrates. Molecular consequences based on the crystal structure of GlcNAc-TI. Biochemistry 40:8765–8772
Gordon RD, Sivarajah P, Satkunarajah M, Ma D, Tarling CA et al (2006) X-ray crystal structures of rabbit N-acetylglucosaminyltransferase I (GnT I) in complex with donor substrate analogues. J Mol Biol 360:67–79
Gottlieb C, Baenziger J, Kornfeld S (1975) Deficient uridine diphosphate-N-acetylglucosamine:glycoprotein N-acetylglucosaminyltransferase activity in a clone of Chinese hamster ovary cells with altered surface glycoproteins. J Biol Chem 250:3303–3309
Grigorian A, Mkhikian H, Demetriou M (2012) Interleukin-2, Interleukin-7, T cell-mediated autoimmunity, and N-glycosylation. Ann NY Acad Sci 1253:49–57
Hassinen A, Pujol FM, Kokkonen N, Pieters C, Kihlstrom M, Korhonen K, Kellokumpu S (2011) Functional organization of Golgi N- and O-glycosylation pathways involves pH-dependent complex formation that is impaired in cancer cells. J Biol Chem 286:38329–38340
Hoe MH, Slusarewicz P, Misteli T, Watson R, Warren G (1995) Evidence for recycling of the resident medial/trans Golgi enzyme, N-acetylglucosaminyltransferase I, in ldlD cells. J Biol Chem 270:25057–25063
Ioffe E, Stanley P (1994) Mice lacking N-acetylglucosaminyltransferase I activity die at mid-gestation, revealing an essential role for complex or hybrid N-linked carbohydrates. Proc Natl Acad Sci USA 91:728–732
Ioffe E, Liu Y, Stanley P (1996) Essential role for complex N-glycans in forming an organized layer of bronchial epithelium. Proc Natl Acad Sci USA 93:11041–11046
Ioffe E, Liu Y, Stanley P (1997) Complex N-glycans in MGAT1 null preimplantation embryos arise from maternal MGAT1 RNA. Glycobiology 7:913–919
Kornfeld R, Kornfeld S (1985) Assembly of asparagine-linked oligosaccharides. Annu Rev Biochem 54:631–664
Kumar R, Yang J, Larsen RD, Stanley P (1990) Cloning and expression of N-acetylglucosaminyltransferase I, the medial Golgi transferase that initiates complex N-linked carbohydrate formation. Proc Natl Acad Sci USA 87:9948–9952
Kumar R, Yang J, Eddy RL, Byers MG, Shows TB, Stanley P (1992) Cloning and expression of the murine gene and chromosomal location of the human gene encoding N-acetylglucosaminyltransferase I. Glycobiology 2:383–393, erratum Glycobiology (1999) 9:(8):ix
Meager A, Ungkitchanukit A, Nairn R, Hughes RC (1975) Ricin resistance in baby hamster kidney cells. Nature 257:137–139
Metzler M, Gertz A, Sarkar M, Schachter H, Schrader JW, Marth JD (1994) Complex asparagine-linked oligosaccharides are required for morphogenic events during post-implantation development. EMBO J J13:2056–2065
Mkhikian H, Grigorian A, Li CF, Chen HL, Newton B et al (2011) Genetics and the environment converge to dysregulate N-glycosylation in multiple sclerosis. Nat Commun 2(334):1–13
Narasimhan S, Stanley P, Schachter H (1977) Control of glycoprotein synthesis. LectiN-resistant mutant containing only one of two distinct N-acetylglucosaminyltransferase activities present in wild type Chinese hamster ovary cells. J Biol Chem 252:3926–3933
Nishikawa Y, Pegg W, Paulsen H, Schachter H (1988) Control of glycoprotein synthesis. Purification and characterization of rabbit liver UDP-N-acetylglucosamine:α-3-d-mannoside β-1,2-N-acetylglucosaminyltransferase I. J Biol Chem 263:8270–8281
Opat AS, Puthalakath H, Burke J, Gleeson PA (1998) Genetic defect in N-acetylglucosaminyltransferase I gene of a ricin-resistant baby hamster kidney mutant. Biochem J 336:593–598
Oppenheimer CL, Hill RL (1981) Purification and characterization of a rabbit liver α,1,3 mannoside β1,2 N-acetylglucosaminyltransferase. J Biol Chem 256:799–804
Pownall S, Kozak CA, Schappert K, Sarkar M, Hull E, Schachter H, Marth JD (1992) Molecular cloning and characterization of the mouse UDP-N-acetylglucosamine:α-3-D-mannoside β-1,2-N-acetylglucosaminyltransferase I gene. Genomics 12:699–704
Puthalakath H, Burke J, Gleeson PA (1996) Glycosylation defect in Lec1 Chinese hamster ovary mutant is due to a point mutation in N-acetylglucosaminyltransferase I gene. J Biol Chem 271:27818–27822
Robertson MA, Etchison JR, Robertson JS, Summers DF, Stanley P (1978) Specific changes in the oligosaccharide moieties of VSV grown in different lectiN-resistant CHO cells. Cell 13:515–526
Sarkar M, Hull E, Nishikawa Y, Simpson RJ, Moritz RL, Dunn R, Schachter H (1991) Molecular cloning and expression of cDNA encoding the enzyme that controls conversion of high-mannose to hybrid and complex N-glycans: UDP-N-acetylglucosamine: α-3-d-mannoside β-1,2-N-acetylglucosaminyltransferase I. Proc Natl Acad Sci USA 88:234–238
Sarkar M, Pagny S, Unligil U, Joziasse D, Mucha J, Glossl J, Schachter H (1998) Removal of 106 amino acids from the N-terminus of UDP-GlcNAc: α-3-d- mannoside β-1,2-N-acetylglucosaminyltransferase I does not inactivate the enzyme. Glycoconj J 15:193–197
Sarkar M, Iliadi KG, Leventis PA, Schachter H, Boulianne GL (2010) Neuronal expression of Mgat1 rescues the shortened life span of Drosophila Mgat11 null mutants and increases life span. Proc Natl Acad Sci USA 107:9677–9682
Schachter H (2010) Mgat1-dependent N-glycans are essential for the normal development of both vertebrate and invertebrate metazoans. Sem Cell Dev Biol 21:609–615
Schachter H, Boulianne G (2011) Life is sweet! A novel role for N-glycans in Drosophila lifespan. Fly 5:18–24
Shi S, Williams SA, Seppo A, Kurniawan H, Chen W, Ye Z, Marth JD, Stanley P (2004) Inactivation of the Mgat1 gene in oocytes impairs oogenesis, but embryos lacking complex and hybrid N-glycans develop and implant. Mol Cell Biol 24:9920–9929
Song Y, Aglipay JA, Bernstein JD, Goswami S, Stanley P (2010) The bisecting GlcNAc on N-glycans inhibits growth factor signaling and retards mammary tumor progression. Cancer Res 70:3361–3371
Stanley P (1983) Selection of lectin-resistant mutants of animal cells. Methods Enzymol 96:157–184
Stanley P (1984) Glycosylation mutants of animal cells. Annu Rev Genet 18:525–552
Stanley P (1989) Chinese hamster ovary cell mutants with multiple glycosylation defects for production of glycoproteins with minimal carbohydrate heterogeneity. Mol Cell Biol 9:377–383
Stanley P, Narasimhan S, Siminovitch L, Schachter H (1975) Chinese hamster ovary cells selected for resistance to the cytotoxicity of phytohemagglutinin are deficient in a UDP-N-. acetylglucosamine-glycoprotein N-acetylglucosaminyltransferase activity. Proc Natl Acad Sci USA 72:3323–3327
Tabas I, Schlesinger S, Kornfeld S (1978) Processing of high mannose oligosaccharides to form complex type oligosaccharides on the newly synthesized polypeptides of the vesicular stomatitis virus G protein and the IgG heavy chain. J Biol Chem 253:716–722
Thierry-Mieg D, Thierry-Mieg J (2006) AceView: a comprehensive cDNA-supported gene and transcripts annotation. Genome Biol 7(Suppl 1):S12.1–14
Unligil UM, Zhou S, Yuwaraj S, Sarkar M, Schachter H, Rini JM (2000) X-ray crystal structure of rabbit N-acetylglucosaminyltransferase I, a key enzyme in the biosynthesis of N-linked glycans. EMBO J 19:5269–5280
von Schaewen A, Sturm A, O’Neill J, Chrispeels MJ (1993) Isolation of a mutant Arabidopsis plant that lacks N-acetylglucosaminyltransferase I and is unable to synthesize Golgi-modified complex N-linked glycans. Plant Physiol 102:1109–1118
Williams SA, Stanley P (2009) Oocyte-specific deletion of complex and hybrid N-glycans leads to defects in preovulatory follicle and cumulus mass development. Reproduction 137:321–331
Yang J, Bhaumik M, Liu Y, Stanley P (1994) Regulation of N-linked glycosylation. Neuronal cell-specific expression of a 5′ extended transcript from the gene encoding N-acetylglucosaminyltransferase I. Glycobiology 4:703–712
Ye Z, Marth JD (2004) N-glycan branching requirement in neuronal and post-natal viability. Glycobiology 14:547–558
Yip B, Chen SH, Mulder H, Hoppener JW, Schachter H (1997) Organization of the human b-1,2-N-acetylglucosaminyltransferase I gene (MGAT1), which controls complex and hybrid N-glycan synthesis. Biochem J 321:465–474
Zhong X, Cooley C, Seth N, Juo ZS, Presman E, Resendes N, Kumar R, Allen M, Mosyak L, Stahl M, Somers W, Kriz R (2012) Engineering novel Lec1 glycosylation mutants in CHO-DUKX cells: molecular insights and effector modulation of N-acetylglucosaminyltransferase I. Biotech Bioeng 109:1723–1734
Zhu S, Hanneman A, Reinhold VN, Spence AM, Schachter H (2004) Caenorhabditis elegans triple null mutant lacking UDP-N-acetyl-d-glucosamine:alpha-3-d-mannoside beta1,2-N-acetylglucosaminyltransferase I. Biochem J 382:995–1001
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Japan
About this entry
Cite this entry
Stanley, P. (2014). Mannosyl (Alpha-1,3-)- Glycoprotein Beta-1,2-N-Acetylglucosaminyltransferase (MGAT1). In: Taniguchi, N., Honke, K., Fukuda, M., Narimatsu, H., Yamaguchi, Y., Angata, T. (eds) Handbook of Glycosyltransferases and Related Genes. Springer, Tokyo. https://doi.org/10.1007/978-4-431-54240-7_129
Download citation
DOI: https://doi.org/10.1007/978-4-431-54240-7_129
Published:
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-54239-1
Online ISBN: 978-4-431-54240-7
eBook Packages: Biomedical and Life SciencesReference Module Biomedical and Life Sciences