Abstract
Pseudomonas syringae is well known as a model bacterial phytopathogen in the laboratory, environment, and the field. A focus on understanding mechanisms of virulence in planta has motivated extensive research into genetic, genomic, and evolutionary factors that influence disease. However, in recent years, appreciation has grown for the life cycle of P. syringae outside of the context of plant disease. This bacterial species survives and thrives across many environments, with its broad ecology shaped through interactions with phage, bacteria, fungi, and insects in addition to traditional host plants. Here we explore what is known about the genetic and genomic basis of these diverse ecological interactions. We highlight how both new and old approaches can be used to unify our understanding of these relationships and map a path forward enabled by high-throughput genomics.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Extracellular Polymeric Substance
- Effector Protein
- Extracellular Polymeric Substance Production
- Plant Immune Response
- Host Range Limitation
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
Pseudomonas syringae is a facultative bacterial phytopathogen of many (if not all) plant species, but strains can also survive and thrive outside of their plant hosts across many different environments (Hirano and Upper 2000). Given an abundance of recent reviews that focus largely on studies of pathogenicity (Block and Alfano 2011; Lindeberg et al. 2012; O’Brien et al. 2011; Studholme 2011), we take this opportunity to summarize and explore what is known about genomic features that influence ecological dynamics for P. syringae inside and outside of plant hosts. Recent sequencing efforts have yielded a wealth of data about genomic diversity throughout this species, but have also given rise to numerous overarching questions about evolutionary dynamics for P. syringae and related bacteria. We view this chapter as an incomplete roadmap that both summarizes current knowledge of the genomic basis for ecological diversity throughout the species and highlights unexplained patterns that arise from comparison across genomes.
3.1 Taxonomy
Taxonomic relationships across phytopathogens and other soil-associated microbes often change, with nomenclatural disputes sparking heated disagreements across disciplines. The focus of this chapter will be what is currently referred to as P. syringae but which includes other potential species such as P. savastanoi and P. cannabina (Ramos et al. 2012; Sarris et al. 2013; Young 2010). We reference genome sequences from within these isolates as a point of context for comparison, but support those that would chose to split off species names given appropriate phenotypic contexts. P. syringae as a whole contains upward of 50 different pathovars spanning pathogens of important crops as well as wild plants (O’Brien et al. 2011; Young 2010). In recent years, due to extensive environmental sampling, greater appreciation has been given to environmental isolates that are not known to be virulent on any host (and thus lack pathovar designations) but which phylogenetically cluster within P. syringae (Diallo et al. 2012a; Kniskern et al. 2010; Morris et al. 2010). One clade in particular, referred to below as MLST group 2C, stands out as having shifted its ecological niche compared to closely related phytopathogenic strains. For the moment, we consider these all to be P. syringae isolates and suggest that elucidation of the ecological roles for these environmental isolates is an intriguing future research direction.
Historically, P. syringae has been classified based on phenotypic responses to the LOPAT test, with positive interactions for levan synthesis and tobacco hypersensitive response and negative responses for arginine dihydrolase, oxidase production, and potato rot as distinguishing features (Young 2010). Strains may be further classified into pathovars based on phenotypic information concerning disease symptoms, hosts of isolation, and pathogenicity on a variety of alternative hosts, in addition to nutritional characteristics (Young 2010). While fruitful in providing a context for rapid and inexpensive classification, phenotype-based methods are prone to errors due to incorrect scoring of phenotypes and convergent evolution of phenotypes relevant for classification. For instance, the MLST group 2C clade tests negative for the tobacco hypersensitive response, but genome sequences indicate that they are a clear subclade within P. syringae (Diallo et al. 2012a). Moreover, there are multiple instances where pathovar designation is polyphyletic, with convergence of disease states on the same host (Baltrus et al. 2011; O’Brien et al. 2012). For these reasons, the focus of classification has shifted to genotypic methods that provide a much richer framework for interpreting evolutionary relationships and patterns between isolates.
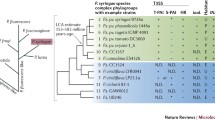
Genotypic classification of strains began with random PCR amplification-based methods, but the first leap forward occurred when Gardan and colleagues used DNA–DNA hybridization to classify isolates into eight genomospecies by genomic similarity (Gardan et al. 1999; Young 2010). More recently, multi-locus sequence type (MLST)-based comparisons have become the default technique for characterizing strains and pathovars (Almeida et al. 2010; Hwang et al. 2005; Morris et al. 2010). MLST characterization relies on sequencing portions of conserved “housekeeping” loci such as rpoD, gapA, gltA (also known as cit), and gyrB. To further facilitate rapid MLST comparisons, a database and Web-based server has been established (Almeida et al. 2010). According to these MLST comparisons, P. syringae strains can be subdivided into at least five distinct clades with complete genome sequences available for strains within three of these groups (group I: P. syringae pv. tomato DC3000, PtoDC3000; group 2: P. syringae pv. syringae B728a, PsyB728a; and group 3: P. syringae pv. phaseolicola 1448a, Pph1448a), draft genome sequences for isolates from the other two main groups (group 4: P. syringae pv. oryzae 1_6, Por1_6 and group 5: P. cannabina pv. alisalensis, Pcal), and numerous other strains throughout the phylogeny (Baltrus et al. 2011; O’Brien et al. 2011; Sarris et al. 2013; Studholme Studholme 2011). For the remainder of this chapter, we will refer to the strains by their MLST group designations where possible and secondarily by their genomospecies (see Fig. 3.1) and will focus on important insights to be gained and questions to be asked by focusing on genome evolution between and within MLST groups.
Phylogeny of Pseudomonas syringae strains and related species. Phylogeny is based on draft and whole-genome sequences and has been modified from (Sarris et al. 2013). MLST groups and genomospecies for each strain are shown at the right. In cases where MLST groups (IV) or genomospecies (4, 5, 6, 7) are not represented within the tree, numbers within the tree highlight estimated branching points for these groups
3.2 Genome Features
Although genomic features within P. syringae have been covered extensively elsewhere, to facilitate further discussion, we highlight a handful of interesting nuances within this chapter that have arisen from comparison across diverse strains. The average genomic content for most P. syringae strains is roughly 5–6 Mb, coding for slightly more than 5,000 open reading frames (O’Brien et al. 2011). Strain PtoDC3000 contains the most thoroughly vetted genome annotation, the product of extensive hand curation and multiple RNAseq experiments (Filiatrault et al. 2010, 2011), and we caution that one should always be wary when analyzing numerous draft genomes due to their fractured nature.
Genomic comparisons across P. syringae have already raised numerous ecological questions, the number of which will only increase with additional genomic information. For instance, group I strains contain approximately 500 additional genes compared to any of the other group, with some functional classes overrepresented within this suite compared to the remaining genomic distribution (Baltrus et al. 2011). While this pattern is unimpressive when compared to the considerable genomic diversity of plant-associated pseudomonads, it could signal important ecological differences when considered within the context of P. syringae as a species. Likewise, plasmids are found within most P. syringae isolates and likely play a primary role in structuring ecologically and virulence-related genomic diversity throughout the species, while also enabling rapid evolutionary shifts through horizontal transfer across strains. Of the complete genomes, PtoDC3000 and Pph1448a both contain two plasmids that house numerous virulence genes, but PsyB728a lacks such plasmids altogether (Buell et al. 2003; Feil et al. 2005; Joardar et al. 2005). Plasmids have also been observed within many other strains (Ma et al. 2007; Vivian et al. 2001). Since plasmids are notorious for containing repetitive elements, which confound draft genome assemblies, it is difficult to pinpoint which contigs within these incomplete sequences are associated with extrachromosomal elements. However, most strains with draft genomes do appear to contain genomic features that are correlated with the presence of mobile elements (Baltrus et al. 2011). Given the important role horizontal gene transfer plays in the evolution of virulence (Jackson et al. 2011; Sundin 2007), it remains to be seen what effects, if any, lack of plasmids has on strain evolutionary and ecological dynamics (Feil et al. 2005). One of the most interesting plasmids is found within a small number of group III strains from pathovar lachrymans (Baltrus et al. 2011). These strains have recently acquired an independently replicated, circular megaplasmid that increases total genomic content by ~1 Mb. Although it contains hundreds of additional genes little else is known about the functions of this element; however, that megaplasmids often code for niche-specific pathways implies that undescribed ecological variability exists within this strain (Harrison et al. 2010).
3.3 Genomic Factors that Limit Host Range of P. syringae
Although P. syringae has a wide host range as a species, each individual isolate only causes disease on a limited number of plants. Moreover, disease symptoms can vary from leaf and fruit spots to trunk cankers to galls depending on strain, pathovar, and host context (Arnold et al. 2011; Preston 2000; Ramos et al. 2012; Scortichini et al. 2012). Our current understanding of interactions between plant immune responses and bacterial virulence mechanisms can be described, at best, as complicated. However, at a primary level, P. syringae growth is limited by the environment within a plant host regardless of dedicated immune responses. For instance, it has long been known that isolates of P. syringae differ in their abilities to use a variety of substrates for nutrition. Plant species can vary dramatically in the quantitative and qualitative distributions of these substrates, and the importance of manipulating host metabolism seems to be a growing theme across phytopathogen species [i.e., Xanthomonas and SWEET genes (Verdier et al. 2012)]. It is likely that nutrient concentrations limit growth of P. syringae in planta, so different metabolite compositions within leaves could contribute to host range limitation (Rico and Preston 2008). With this idea in mind, it seems particularly noteworthy that multiple clades of P. syringae that invade woody plant species have convergently acquired pathways involved in the breakdown of catechols (Green et al. 2010; Marcelletti et al. 2011; Rodríguez-Palenzuela et al. 2010). Furthermore, Thlaspi caerulescens can hyperaccumulate zinc, nickel, and cadmium at high enough concentrations to prevent bacterial pathogenesis (Fones et al. 2010). In the absence of high concentrations of these metals, PcalES4326 is virulent on T. caerulescens. Transposon mutagenesis can be used to isolate mutants with higher metal tolerance, and these mutations enabled PcalES4326 to sustain virulence in T. caerulescens plants as concentration of toxic metals increased. Although this result is tantalizing and suggestive, it remains unclear how much natural variation exists for metal tolerance within P. syringae and how much of an impact such environmental context plays in structuring natural bacterial host ranges.
The first layer of induced plant immune responses against any phytopathogen involves dedicated receptor-like kinase proteins that directly recognize conserved peptides (termed pathogen-associated molecular patterns or PAMPs) triggering basal defenses (Thomma et al. 2011). Although we have learned extensive amounts about the molecular mechanisms underlying PAMP recognition, it is likely that this is only the tip of the iceberg and that there is much more to discover as to how these receptors vary in presence and specificity at the level of host phylogeny. Although the most highly reactive and widely studied PAMP within P. syringae is flagellin (specifically flg22), a library of other proteins has been implicated in triggering basal defenses (also known as pathogen-triggered immunity, PTI) (McCann et al. 2012; Thomma et al. 2011). Moreover, at least in a subset of plant species like tomato, other portions of flagellin (flg28) can also be recognized (Cai et al. 2011). Although natural variation in PAMP regions exists across P. syringae strains and pathovars, and these regions appear to have been the target of diversifying selection, the contribution of amino acid diversity of bacterial PAMPs to structuring host range remains unclear (McCann et al. 2012).
An emerging trend, however, is that receptor-like kinases that respond to specific peptides may not be conserved across all plant species, so that PAMP recognition may differ in a qualitative way across plant hosts (Segonzac and Zipfel 2011; Thomma et al. 2011). For example, bacterial elongation factor Tu (EfTu) can act as a PAMP (peptides are named elf18 and elf26) within Brassicaceae hosts, but does not in Solanaceous hosts like tobacco and tomato (Lacombe et al. 2010). Therefore, P. syringae isolates that are pathogenic on Brassicaceae must overcome recognition that isolates from other pathovars may not experience. The ability of such plant clade-specific receptors to structure host range is apparent, and indeed, genetic modification is already focused on moving specific PAMP receptors across plant families with the hopes of creating durable resistance (Lacombe et al. 2010). While the total number and phylogenetic distributions of PAMP receptors remain unknown, it is highly likely that there are additional receptors like EFR that can limit natural bacterial host range.
Immune responses to PTI include callose deposition, production of reactive oxygen species, and deployment of antimicrobial compounds (Jones and Dangl 2006; Nicaise et al. 2009). P. syringae strains must be able to cope with or avoid these stresses during infection, and it is possible that clade- or isolate-specific variation in proteins like catalase could mediate differential interactions inside of hosts. For instance, glucosinolate production within Arabidopsis strongly limits the growth of P. syringae (Fan et al. 2011). Interestingly, glucosinolate expression is not thought to be involved in the PTI response, as precursors are held in vacuoles until these containers are disrupted by mechanical forces like chewing insects. Bacterial isolates that are pathogenic on Arabidopsis have acquired multiple efflux systems through horizontal transfer that enable higher tolerance toward glucosinolates, but these systems are not found extensively throughout P. syringae. The overall contribution of such plant species with specific responses to host range limitation has not been thoroughly investigated across hosts and bacterial isolates.
Pseudomonas syringae uses a type III secretion system to translocate effector proteins into host cells, which is essential for virulence as described below, but recognition of these effector proteins (or their actions) by plant R genes can limit host range (referred to as effector-triggered immunity, ETI) (Jones and Dangl 2006; Thomma et al. 2011). The outcome of ETI is the plant hypersensitive response (HR), which involves localized cell death to limit pathogen growth and spread. At one level, ETI is responsible for establishing race structure within pathovars of P. syringae based on differential recognition of variable effector proteins across cultivars of a given plant species (Taylor et al. 1996). At another level, ETI can contribute to limiting the host range of pathovars across plant species. For instance, recognition of the effector AvrPtoB has been demonstrated to limit the growth and decrease symptoms of multiple pathovars of P. syringae on tomato (Chien et al. 2013; Lin and Martin 2007). Likewise, recognition of HopQ1 renders PtoDC3000 avirulent and may partially explain lack of growth of Pph1448a on tobacco (Ferrante et al. 2009; Wei et al. 2007). When considering the contribution of ETI to host range limitation overall, however, one must consider the distribution and conservation of known avirulence factors across all isolates within a pathovar and comparisons of diversity for R genes across plant species and populations. Effectors have been shown to trigger HR reactions in every tested cultivar of a plant species for only a handful of cases, but if such trends hold up with further sampling, these effector proteins may truly limit host range at the level of plant species (Arnold et al. 2001; Wroblewski et al. 2009). Consideration of how such interactions structure host range may be especially important when two pathovars have recently diverged to infect different hosts, as with HopC1 in pathovars Pgy and Pph (Baltrus et al. 2012), as a recent report suggests that ETI is the first stage of host differentiation between pathovars (Schulze-Lefert and Panstruga 2011).
3.4 Genomic Factors that Promote Host Range Expansion
All known phytopathogenic P. syringae isolates require a type III secretion system for virulence (O’Brien et al. 2011; Tampakaki et al. 2010). This TTSS was acquired by an immediate progenitor of all P. syringae and P. cannabina, likely from a clade within P. viridiflava (Araki et al. 2006; Sarris et al. 2013). Although localized recombination within the TTSS has been observed, the system as a whole has been vertically inherited since this introduction (Sarris et al. 2013). While avirulence due to TTEs limits host range in an R gene-dependent way, in the absence of ETI, these translocated proteins are absolutely essential for bacterial growth and disease progression in planta (Block and Alfano 2011; Lindeberg et al. 2012). Moreover, the actions of some TTE can cover up ETI triggered by other TTEs (Jones and Dangl 2006).
Each strain of P. syringae possesses a handful of conserved effector genes that, although they may not be functional within a strain, have been vertically inherited from an ancestor of all virulent P. syringae including hopI1, hopAH2, avrE, hopAA1, and hopM1 (Baltrus et al. 2011; O’Brien et al. 2011). Such conservation implies a foundational role for these proteins across a wide array of plant hosts. In addition to this small number of conserved effector proteins, each strain possesses from 3 to 36 (or more) additional effector proteins which are frequently lost through deletion or gained by horizontal gene transfer (Baltrus et al. 2011; Lindeberg et al. 2012). While an increasing amount of information about the precise functions of effector proteins has been summarized elsewhere, common themes emerge by comparing these isolates and exploring what these trends say about the evolution of virulence and host range.
Studies within one strain, PtoDC3000, across multiple host plants have demonstrated that effector proteins can be clustered into tiers based on function (Cunnac et al. 2011). Some TTEs can further be classified into redundant effector groups (REG) based on overlapping virulence function and ability to complement phenotypic virulence effects (Kvitko et al. 2009). Single knockouts of genes within a REG yield no (or very small) virulence defects, so that only by disrupting each member of a REG is virulence noticeably changed. Two main REGs have been identified at present within P. syringae: AvrE/HopM/HopR appears to disrupt secretion systems involved in plant defenses, while AvrPto/AvrPtoB disrupts perception and signaling after the recognition of bacterial PAMPs (Kvitko et al. 2009; Lin and Martin 2005; Badel et al. 2006). As a demonstration of effector tiers, Cunnac et al. (2011) showed that, if single members of either of these REGs were present within PtoDC3000, a variety of other type III effector proteins could quantitatively and interchangeably contribute to virulence. It is unclear how conserved REG functions are across strains and host plants. However, as Baltrus and colleagues showed, the placement of AvrPto from the soybean pathogen Pgy into a virulent bean pathogen (Pph1448a) could increase virulence on the target host species even though Pph1448a already contains AvrPtoB (Baltrus et al. 2012). It will be interesting to see how the REG concept and tiers of effector proteins are refined with future studies.
Type III effectors are not the sole determinant of success and failure during infections by P. syringae as many strains also produce toxins and phytohormones that manipulate plant physiology (Bender et al. 1999; Melotto et al. 2006; Schellenberg et al. 2010), including polyketide synthase (PKS) and non-ribosomal peptide synthase (NRPS) pathways. One of the best studied of these PKS-derived phytohormone mimics is coronatine, which binds to jasmonic acid receptors, and is the product of two multi-gene pathways that can be horizontally transferred across strains (Alarcón-Chaidez et al. 1999; Melotto et al. 2006). While the best known function of coronatine involves stomatal manipulation, which can also be altered by the TTE AvrB or the toxin syringolin, additional evidence points to secondary virulence effects for coronatine after bacterial cells reach the apoplast (Geng et al. 2012; Melotto et al. 2006). One unexplained trend that has emerged from genomic comparisons across P. syringae is that group II strains contain a lower number of effector proteins on average than any of the other groups. Parsimonious reconstruction across TTE groups suggests that the ancestor of group II contained additional TTE, so that this trend is the outcome of gene loss [at a minimum loss of gene families hopR, hopAS, hopQ, and hopD, with more complicated histories for avrpto and hopAB (Baltrus et al. 2011)]. This pattern is not an artifact of sampling, but correlates with the presence of toxin pathways for syringolin, syringomycin, and syringopeptin. It is tantalizing to speculate that these strains have replaced virulence functions of TTE with those of toxins, especially because syringolin has recently been implicated both in manipulating stomatal function and in preventing ETI in PsyB728a (Schellenberg et al. 2010). Furthermore, an independent data point suggesting that toxins replace TTE function involves pv. pisi strains which appear to have secondarily lost all three of these toxins but have gained back numerous effector groups (Baltrus et al. 2011). It is difficult to know whether these toxins can each identically replace the “lost” effector groups as well as how toxins factor into strain specificity. Also of note when discussing phytohormone manipulation by P. syringae is that strains that form galls on woody plants contain additional copies of pathways to produce the phytohormone auxin, which directly contributes to plant tissue growth and is required for gall formation along with the TTSS (Rodríguez-Palenzuela et al. 2010). Other toxins include phaseolotoxin, mangotoxin, and tabtoxin, which are found in a limited number of strains, and all appear to contribute to disease symptoms like chlorosis but not necessarily growth in planta (Bender et al. 1999; Carrion et al. 2013; Groll et al. 2008).
A small subclade within group II strains (referred to as group IIC) has replaced the original TTSS with a phage sequence and has actually gained an alternative type III secretion system at a different location of the genome (Clarke et al. 2010; Mohr et al. 2008). While divergent in protein sequence from the canonical P. syringae TTSS, this second system most closely resembles structures used to translocate proteins across plant cells and these strains were shown to be able to grow in planta in a TTSS-dependent way (Clarke et al. 2010). Moreover, group IIC strains can be found epiphytically on plants across a wide range of environments and hosts (Diallo et al. 2012b; Kniskern et al. 2010). The precise function of this alternative type III secretion system is not known, but it could contribute to pathogenicity during this epiphytic stage, playing a more subtle ecological role than the canonical system, or may be used in an entirely different way during interactions with non-plant hosts. Of note is that this secondary system within group IIC does not appear to be as finely regulated at a transcriptional level when compared to the canonical system (Clarke et al. 2010). Lastly, some strains within groups III and IV have been shown to harbor a second type III secretion system within their genomes (O’Brien et al. 2011). This system does not appear to be involved in phytopathogenicity, and its true function is not known.
3.5 Expansion of Host Range Outside of Plants
Although the most widely studied aspects of P. syringae evolution and ecology involve phytopathogenesis, a growing body of work suggests that certain members of this species can survive and thrive in non-plant hosts (Stavrinides et al. 2009). Indeed, such interactions may be a critical aspect of environmental persistence and could structure ecological relationships between strains and species in previously unrecognized ways. For instance, P. syringae is routinely exposed to insects throughout the environment and can be isolated from inside-surface-sterilized insects that have fed on plants (Stavrinides et al. 2009). Insects are a prevalent part of the life cycles for numerous plant pathogens, but the potential for insects to act as vectors or hosts of P. syringae has received comparatively little attention (Nadarasah and Stavrinides 2011). However, multiple strains possess a demonstrated capacity to grow to high densities inside of pea aphids (Acyrthosiphon pisum) and can be deposited on new plants through excreted honeydew. Intriguingly, strain-specific entomopathogenicity has also been demonstrated, as PsyB728a but not PtoDC3000 has the potential to kill up to 95 % of aphids within 48 h of infection (Stavrinides et al. 2009).
The ecological importance and relevance of these interactions are currently unclear, but these early results suggest fascinating directions for future study. Firstly, hemipteran insects, like aphids, are excellent candidates for bacterial hosts/vectors because they feed on and excrete carbohydrate-rich plant sap. They also continually probe plant surfaces during feeding, which increases the chance of encountering epiphytic bacteria (Stavrinides et al. 2009). However, it is currently unknown whether P. syringae can successfully infect hemipteran insects in addition to aphids. Also unknown is the genetic and genomic basis of persistence inside of insects and entomopathogenicity. While Stavrinides et al. (2009) were able to demonstrate that a functioning flagellum is important for establishing insect infection, they also ruled out contributions from all recognizable entomotoxins within the genome sequence compared to other insect pathogens. Given the wealth of genomic information available for P. syringae, a promising future research direction could involve evolutionary and phylogenetic comparisons of entomopathogenicity. Such experiments could shed light on both the genetic basis and evolutionary mechanisms enabling insect killing. Are similar genes involved in virulence on plants versus insects, or is insect virulence a newly evolved trait? Do strains that are able to make use of insect hosts have specific adaptations associated with the phenotype?
P. syringae genomes also harbor multiple independent pathways that mediate interactions with fungi. Notably, many group II strains are demonstrated producers of toxins like syringomycin, and the bioassay for production of these molecules involves suppression of growth of the fungus Geotrichum candidum (Quigley and Gross 1994; Scholz-Schroeder et al. 2001). This toxin is assembled through an NRPS pathway and also contributes to plant infection and to disease symptom development. Another gene present within many P. syringae isolates, phcA, has been shown to induce a cell death response in Neurospora crassa, but an ecological role for this protein remains unclear (Wichmann et al. 2008). Lastly, most (if not all) P. syringae genomes contain at least one type VI secretion system (HSI-I) and associated effector molecules, while strains PtoDC3000 and P. cannabina pv. alisalensis contain a second type VI secretion system (HSI-II) (Records and Gross 2010; Sarris et al. 2010, 2013). Both HSI-I and HSI-II enable P. syringae to outcompete fungi, but very little is known about how such pathways structure natural communities (Haapalainen et al. 2012).
3.6 Competition with Other Bacteria
Syringopeptin and HSI-II can also be used to influence interactions between P. syringae and other bacterial species (Haapalainen et al. 2012; Scholz-Schroeder et al. 2001), and all isolates likely contain additional pathways to produce narrow spectrum antimicrobial compounds termed bacteriocins. Bacteriocins can take a variety of forms, but all are active only against strains closely related to the producer. Readers are directed to recent reviews (and references therein) on bacteriocins produced by phytopathogens, including P. syringae, for background on bacteriocin biology (Grinter et al. 2012; Holtsmark et al. 2008). In particular, bacteriocin activity has been demonstrated within multiple P. syringae isolates including pathovars syringae, morsprunorum, glycinea, phaseolicola, ciccaronei, and tomato (Barreteau et al. 2009; Garrett et al. 1966; Ghequire et al. 2012; Lavermicocca et al. 1999; Vidaver et al. 1972). Additionally, various bacteriocins have been predicted within the genomes of PsyB728a, Pph1448A, PtoDC3000, and P. syringae pv. aptata (Ghequire et al. 2012; Holtsmark et al. 2008; Parret and De Mot 2002).
Bacteriocins can influence the phyllosphere community as well as plant disease in several ways. Most directly, they could promote disease by providing potential pathogens with an epiphytic advantage by preferentially excluding closely related strains [which are likely to occupy a similar or identical niche (Wilson and Lindow 1994)]. Conversely, nonpathogenic strains could be used to reduce plant disease through preferential exclusion of pathogens. Indeed, Lavermicocca et al. (2002) found that application of a currently undetermined bacteriocin from P. syringae pv. ciccaronei reduced olive knot disease caused by P. savastanoi (Lavermicocca et al. 2002). Additionally, Garrett et al. (1966) found a general trend where P. syringae isolates recovered from citrus tended to produce bacteriocins active against isolates recovered from pear and vice versa. Whether such a pattern is common, is the result of long-term evolutionary forces, or is stochastic will require further research.
In addition to their direct killing effect, bacteriocins may influence the ecology of P. syringae by targeting receptors important for nutrient uptake, like TonB-dependent siderophore receptors (Davies and Reeves 1975; Denayer et al. 2007). Indeed, selection for bacteriocin resistance negatively affected the ability of Erwinia chrysanthemi (Dickeya dadantii) to cause soft rot in Saintpaulia plants, with a correlated loss of low-iron-induced outer membrane proteins (Expert and Toussaint 1985). In this way, bacteriocins could influence evolutionary patterns across P. syringae genomes, by directly altering selective pressures on targets like siderophore uptake receptors. While, to our knowledge, there are currently no receptors described as mediating bacteriocin sensitivity in P. syringae, identification of such receptors will significantly aid in understanding the influences of bacteriocins on P. syringae ecology.
3.7 Survival in the Environment
Pseudomonas syringae strains can survive epiphytically on the outside of plant leaves and can be isolated from environmental sites such as leaf litter, rivers, lakes, snowpack, and even from clouds (Morris et al. 2008, 2010). While some genetic factors that facilitate survival across such diverse habitats have been well studied [i.e., UV resistance (Kim and Sundin 2000; Zhang and Sundin 2004)], whole-genome analyses and intense sampling schemes have only raised additional ecological questions about persistence of P. syringae outside of disease-causing infections. Indeed, each genome contains many ECF sigma factors, regulatory targets of which are unknown, that appear to be dispensable for phytopathogenesis but could enable environmental survival (Oguiza et al. 2005; Thakur et al. 2013).
Although epiphytic and apoplastic populations of P. syringae experience very different environments, the type III secretion system and associated effectors appear to play critical roles in epiphytic survival. Recent work by Yu et al. (2013) has begun to illuminate the traits harbored by P. syringae that contribute to both epiphytic and apoplastic fitness, as well as those traits that appear to be utilized in only one of the two environments. Confirming previous research that showed that motility was an important epiphytic trait, genes encoding flagellar components, chemosensory and chemotaxis proteins, rhlA, which encodes, and enzyme that produces 3-(3-hydroxyalkanouloxy) alkanoic acid (HAA), a swarming motility-enabling surfactant (Burch et al. 2012), were upregulated in the epiphytic environment compared to the apoplastic environment. Taken together, the data suggest that motility is important for the epiphytic colonization, but does not play a prominent role in apoplastic colonization [with the caveat that motility aids in initial invasion (Hattermann and Ries 1989; Panopoulos and Schroth 1974)]. Supporting the idea that group II strains have transitioned from relying on diverse TTEs to phytotoxins as mediators of virulence, secondary metabolites, including many NRPSs and PKSs, were all more highly expressed apoplastically than epiphytically. Interestingly, genes associated with mobile elements (prophage, insertion sequences, and transposases) were also more highly expressed within the apoplast than on the leaf surface. This finding correlates with recent findings in Pph1448A, where a genomic island (PPHGI-1) readily transferred between strains co-infiltrated into bean leaves, but did not readily transfer between strains co-inoculated into culture media (Lovell et al. 2009). Taken together with the knowledge that horizontal gene transfer has significantly contributed to evolution across P. syringae isolates, these results might suggest that gene transfer readily occurs between bacteria co-colonizing a leaf apoplast. As might be expected, metabolism was also influenced by association with the plant host. Pathways involved in the uptake and catabolism of GABA, phenylalanine, as well as other amino acids and sugars were induced in planta compared to basal conditions (Yu et al. 2013). Some pathways were more highly induced in the apoplast (GABA), while others were more highly induced epiphytically (phenylalanine). Perhaps one of the most dominant environmental forces inside and outside of leaves is water limitation, a trend which stands out in both phenotypic assays and transcriptome studies, but very little is known across strains about survival under water stress. While there is now a wealth of data regarding expression patterns in two important plant environments (the leaf surface and leaf interior), it will be interesting to compare such patterns across strains as well as plant hosts to determine which expression profiles are largely conserved or are more specific to a particular strain or plant host. For instance, are traits that are generally conserved across P. syringae pathovars equally conserved in their regulation?
A phenotype often associated with virulence and success in plants, the ability to attach to and aggregate on surfaces, is also required for P. syringae’s success in epiphytic and environmental habitats (Hirano and Upper 2000). Attachment and aggregation on surfaces not only prevent cells from being physically removed from a surface (i.e., being washed away), but also help P. syringae resist a variety of environmental stresses such as desiccation, exposure to UV light and reactive oxygen species, and competition from other microbes. However, the ability to aggregate on leaf surfaces is not required for in planta virulence and all strains are not equally proficient at forming environmental aggregations (Hirano and Upper 2000; Lindow et al. 1993).
Exactly how P. syringae attaches to surfaces is not well understood across strains, but can involve the secretion of extracellular polymeric substances (EPS) to form aggregations or biofilms (Hirano and Upper 2000; Laue 2006; Lindeberg et al. 2008). A variety of polymers have been proposed to be important in P. syringae biofilm formation. Epiphytic fitness has been best linked to production of the polysaccharide alginate (Laue 2006; Quiñones et al. 2005), the production of which is upregulated in epiphytic populations compared to apoplastic populations within strain PsyB728a (Yu et al. 2013), although other polysaccharides such as levan or cellulose may also be important (Laue 2006; Lindeberg et al. 2008; Ude et al. 2006).
A major problem with uncovering the genomic basis for surface aggregation in P. syringae is that many of the genes likely to be involved are widespread and conserved across strains. However, although regulation of EPS pathways seems to be tied to quorum sensing and cell density (Quiñones et al. 2005), differences in expression levels of pathways involved in bacterial aggregation may ultimately underlie phenotypic differences. For instance, while PtoDC3000 and the notable biofilm-forming human pathogen P. aeruginosa both possess full operons for cellulose production, they differ in regulatory genes for the operon (Lindeberg et al. 2008). Possibly, pairing transcriptome analyses with genomic analyses will help to identify adaptations useful in surface aggregation. This approach would be particularly useful when compared across strains that vary in environmental niches. Another area that remains relatively unexplored is the formation of aggregations on other environmental surfaces besides plant leaves. Environmentally isolated P. syringae strains have been found to use cellulose as a biofilm matrix (Ude et al. 2006), but it is not known whether they also use polysaccharides produced on leaves and how the production of these compounds may be regulated in diverse environments.
3.8 Dispersal
Global dispersal of P. syringae is thought to be intricately linked to the water cycle, where long-distance transfer takes place by cells or aggregations of bacteria being swept up into clouds and transported by meteorological forces (Morris et al. 2008). One of the most important pathways contributing to this dispersal process for many plant-associated bacteria is ice nucleation proteins (Morris et al. 2010). However, there is substantial variability in the protein composition and nucleation properties of ice crystals across strains, with some isolates like Pph1448a lacking production capabilities due to disruption by insertion elements. Moreover, some strains like PtoDC3000 display ice nucleation capabilities even though they lack inaZ (Feil et al. 2005; Hwang et al. 2005; Joardar et al. 2005).
Some pathovars of P. syringae may rely more heavily on seed-borne rather than environmental dispersal, and it is currently unknown how such ecological changes manifest at the level of the genome. For example, pathovar phaseolicola is readily dispersed through seeds and has a substantially smaller nutritional range than other strains like P. syringae pv. syringae B728a or pv. tomato DC3000 (Arnold et al. 2011; Rico and Preston 2008). These changes could be the product of selection for mutational deterioration of unused molecular pathways, but may also reflect differences in population size and genetic drift. Although just a correlation at this point, such changes in addition to the disruption of inaZ could explain why subclades within group III strains in particular appear to be under-sampled when it comes to environmental reservoirs of P. syringae (Morris et al. 2008, 2010). Also unclear are specific virulence factors that enable strains to facilitate seed dispersal.
That iron limitation is a strong selective force for environmental pseudomonad populations is reflected in the diverse array of siderophores that have evolved to scavenge this scarce element. Most P. syringae isolates studied to date have the genetic capability to produce two, if not more, high-affinity iron uptake systems (see below). The most widely studied siderophore produced by P. syringae is pyoverdine (Fig. 3.2), which is encoded by a NRPS pathway and imparts the characteristic fluorescence to strains of this species as well as related “fluorescent” pseudomonads (Visca et al. 2007). This class of siderophores has been studied for several decades, largely in Pseudomonas aeruginosa, where much has been learned with regard to the genetics, genomics, and biochemistry of pyoverdine production and transport [reviewed in Budzikiewicz (2004), Gross and Loper (2009), Meyer (2000), Visca et al. (2007)]. Interestingly, although three structurally different pyoverdines are produced among different strains of P. aeruginosa (Cornelis et al. 1989), which is mirrored by allelic diversity in the genes responsible for pyoverdine synthesis and uptake (Bodilis et al. 2009; Smith et al. 2005; Spencer et al. 2003), pyoverdines produced by different strains of P. syringae are largely identical (Bultreys and Gheysen 2000; Bultreys et al. 2001, 2003; Jülich et al. 2001). These observations suggest that while pyoverdine synthesis and uptake are under diversifying selection in P. aeruginosa, such selection does not occur in P. syringae. The difference in selection pressure between these two species is likely a result of differences between their ecologies.
Diverse colony morphologies and fluorescent pigment (pyoverdine) production across Pseudomonas syringae pathovars. Five microliters of overnight KB broth cultures was inoculated onto solid KB medium and incubated at room temperature for 48 h. Pictures of plates were taken without (top panels) or with UV illumination (bottom panels). MLST group I strains: Pan (pv. actinidiae), Pmp (pv. morsprunorum), Pto DC3 K (pv. tomato DC3000), Pla 106 (pv. lachrymans 106), and Pto T1 (pv. tomato T1); MLST group II strains: Psy B728a (pv. syringae B728a), Pac (pv. aceris), Cit7 (P. syringae Cit7), B48 (P. syringae B48), B15 (P. syringae B15), Ptt (pv. aptata), Ppi 1704 (pv. pisi), FF5 (pv. syringae FF5), 1212 (pv. syringae 1212), Pja (pv. japonica), and A2 (pv. syringae A2); MLST group III strains: Pmo (pv. mori), Pph 1448A (pv. phaseolicola 1448A), Pgy (pv. glycinea), Pla 107 (pv. lachrymans 107), Pta (pv. tabaci), and Pae (pv. aesculi); MLST group V Pcal (pv. alisalensis, but also known as P. syringae pv. maculicola ES4326), MLST group IV Por (pv. oryzae)
While pyoverdine produced across P. syringae pathovars is identical (or nearly so), and presumably the underlying genetics governing synthesis, export, and import is highly similar, the regulation of pyoverdine production and perception may differ between pathovars (see Fig. 3.1 for example). For instance, evidence suggests that production of pyoverdine and achromobactin in P. syringae pv. syringae 22d/93 was much more sensitive to culture conditions (presumably oxygen availability) than P. syringae pv. glycinea 1a/96 (Wensing et al. 2010). Indeed, the one clear example of pyoverdine playing a role in virulence toward its host comes from pv. tabaci, where production of pyoverdine was shown to play a signaling role (Taguchi et al. 2010). Taguchi et al. (2010) demonstrated that disruption of pyoverdine production influenced the regulation of EPS production, surfactant production, swarming motility, AHL production, antibiotic resistance, and tabtoxin production. Presumably, this global regulatory role of pyoverdine production is not generally conserved among all pathovars, or one would expect similar results to be observed by Jones and Wildermuth (2011).
P. syringae isolates may also contain one or several non-NRPS-derived siderophores including achromobactin (pv. actinidiae, pv. syringae, and pv. phaseolicola), yersiniabactin (pv. tomato, pv. actinidiae, and pv. phaseolicola), haemin (pv. actinidiae), enterobactin (pv. actinidiae and pv. aesculi), and citrate (Berti and Thomas 2009; Buell et al. 2003; Green et al. 2010; Jones and Wildermuth 2011; Owen and Ackerley 2011; Scortichini et al. 2012; Wensing et al. 2010). In addition to siderophore prediction from whole-genome analyses, yersiniabactin has either been detected directly using HPLC or is predicted to be encoded in several pathovars belonging to genomospecies 2, 3, 7, and 8 (Bultreys et al. 2001). The diversity of non-pyoverdine siderophores produced across P. syringae pathovars may be evolutionarily analogous to the diversity of pyoverdines produced across P. aeruginosa isolates.
In addition to receptors mediating uptake of endogenously produced siderophores, strains of P. syringae encode numerous receptors predicted to import non-native siderophores (Cornelis and Bodilis 2009; Cornelis and Matthijs 2002). Comparison of PtoDC3000, Pph1448A, and PsyB728a indicated that they share a common core set of 13 TonB-dependent uptake receptors, but collectively encode 29 such receptors. While cognate siderophores that interact with these receptors are largely unknown, it is tempting to speculate that this diversity is linked to each organism’s distinct ecology.
Siderophores have been demonstrated to play a role in epiphytic fitness, where they likely function to aid P. syringae in acquiring otherwise unavailable iron (Karamanoli et al. 2011; Wensing et al. 2010). Indeed, whole-cell biosensors employed by Joyner and Lindow indicated that, like many nutrients, biologically available iron is heterogeneously distributed over a bean leaf surface (Joyner and Lindow 2000). Additionally, plants that harbor greater amounts of leaf surface polyphenolics (including tannin, an iron chelator) induced greater production of pyoverdine in PsyB728a, as well as inhibited growth of an isogenic mutant unable to produce pyoverdine, than plants with lower levels of leaf surface polyphenolics (Karamanoli et al. 2011). In contrast to P. aeruginosa, where siderophore production contributes to virulence (Meyer et al. 1996; Takase et al. 2000), siderophore production by P. syringae appears to be largely dispensable for infection of plant hosts (Cody and Gross 1987; Jones and Wildermuth 2011; Owen and Ackerley 2011). This is likely a result of sufficient available iron within the leaf interior [see references cited in (Jones and Wildermuth 2011)]. Indeed, recent transcriptomic work in PsyB728a found that iron acquisition systems were largely uninduced in the plant apoplast when compared with an iron-limiting medium (Yu et al. 2013). While no work has directly addressed the role of siderophores in the life history of P. syringae beyond the phyllosphere, presumably these high-affinity iron uptake systems are important for these stages of its life cycle. Moreover, it may be that the alternate siderophores produced across pathovars are uniquely adaptive to each strain’s particular life cycle. Clearly, more experimental and comparative research is required to understand the adaptive basis for the diversity of alternate (non-pyoverdine) siderophores encoded by P. syringae pathovars.
While siderophores, and their associated uptake systems, are well understood for their role in acquiring scarce iron (and potentially other limiting metals) from the environment, one aspect of siderophore receptors that is completely unstudied is their role as receptors mediating phage and bacteriocin sensitivity. Similar to bacteriocins described above, phage–P. syringae interaction is likely to be an area where dedicated research is likely to improve our understanding of diversity-generating processes. Several prophage regions have been predicted in all of the fully sequenced P. syringae strains, though whether any of these regions is active (i.e., gives rise to an infective phage) is currently unreported in the literature. Previous research has indicated that some P. syringae strains harbor active prophage (Garrett and Crosse 1963; Minor et al. 1996; Nordeen et al. 1983; Prior et al. 2007; Sato 1983), but the lack of genome sequences for these strains and prophage limits understanding of the underlying biology. It is tempting to speculate that P. syringae–phage interactions may be important in several ways given the recent results indicating that genes encoded in a prophage region of PsyB728a are induced under environmentally relevant conditions (Hockett et al. 2013; Yu et al. 2013). Additionally, recent work has demonstrated that there is an abundance of infective phage that can be isolated from the interior of horse chestnut leaves, which can replicate on P. syringae hosts (Koskella et al. 2011). Future research should aid in answering questions about the possible role(s) of phages in affecting the ecology and evolution of P. syringae. For instance, are phages important contributors to genetic exchange among P. syringae pathovars, similar to plasmids?
3.9 Concluding Remarks
The plummeting cost of DNA sequencing has enabled an explosion of genomic data for P. syringae strains and related species. While these sequences have provided numerous insights into the mechanisms of pathogenicity and environmental persistence for a handful of strains, ecological dynamics and mechanisms of survival outside of plant hosts remain a black box for the species as a whole. Moreover, it is far too easy to fall into treating P. syringae strains monolithically and assume what is true for one strain is true for all. In reality, vast phenotypic diversity exists across this species and ecological interactions could dramatically differ even for close relatives. While realization of these differences can enable more thorough ecological explorations, genomic-scale comparisons must be matched by greater sampling of strains outside of plant hosts and experimental tests of phenotypes to understand what traits are critical for environmental survival, persistence, and dispersal. Each lineage of P. syringae exhibits a highly complex evolutionary trajectory where selection acts on survival inside and outside of plant hosts and only incorporation of lineage-specific ecologies and appreciation of differences between strains can enable a complete understanding of genomic patterns and evolutionary dynamics across this species.
References
Alarcón-Chaidez FJ, Peñaloza-Vázquez A, Ullrich M, Bender CL (1999) Characterization of plasmids encoding the phytotoxin coronatine in Pseudomonas syringae. Plasmid 42:210–220
Almeida NF, Yan S, Cai R, Clarke CR, Morris CE, Schaad NW et al (2010) PAMDB, a multilocus sequence typing and analysis database and website for plant-associated microbes. Phytopathology 100:208–215
Araki H, Tian D, Goss EM, Jakob K, Halldorsdottir SS, Kreitman M, Bergelson J (2006) Presence/absence polymorphism for alternative pathogenicity islands in Pseudomonas viridiflava, a pathogen of Arabidopsis. Proc Natl Acad Sci USA 103:5887–5892
Arnold DL, Jackson RW, Fillingham AJ, Goss SC, Taylor JD, Mansfield JW, Vivian A (2001) Highly conserved sequences flank avirulence genes: isolation of novel avirulence genes from Pseudomonas syringae pv. pisi. Microbiology 147:1171–1182
Arnold DL, Lovell HC, Jackson RW, Mansfield JW (2011) Pseudomonas syringae pv. phaseolicola: from “has bean” to supermodel. Mol Plant Pathol 12:617–622
Badel JL, Shimizu R, Oh HS, Collmer A (2006) A Pseudomonas syringae pv. tomato avrE1/hopM1 mutant is severely reduced in growth and lesion formation in tomato. Mol Plant Microbe Interact 19: 99–111
Baltrus DA, Nishimura MT, Dougherty KM, Biswas S, Muhktar S, Vicente JG et al (2012) The molecular basis of host specialization in bean pathovars of Pseudomonas syringae. Mol Plant Microbe Interact 25:877–888
Baltrus DA, Nishimura MT, Romanchuk A, Chang JH, Mukhtar MS, Cherkis K et al (2011) Dynamic evolution of pathogenicity revealed by sequencing and comparative genomics of 19 Pseudomonas syringae isolates. PLoS Pathog 7:e1002132
Barreteau H, Bouhss A, Fourgeaud M, Mainardi JL, Touze T, Gerard F et al (2009) Human- and plant-pathogenic Pseudomonas species produce bacteriocins exhibiting colicin M-like hydrolase activity towards peptidoglycan precursors. J Bacteriol 191:3657–3664
Bender CL, Alarcón-Chaidez F, Gross DC (1999) Pseudomonas syringae phytotoxins: mode of action, regulation, and biosynthesis by peptide and polyketide synthetases. Microbiol Mol Biol Rev 63:266–292
Berti AD, Thomas MG (2009) Analysis of achromobactin biosynthesis by Pseudomonas syringae pv. syringae B728a. J Bacteriol 191:4594–4604
Block A, Alfano JR (2011) Plant targets for Pseudomonas syringae type III effectors: virulence targets or guarded decoys? Curr Opin Microbiol 14:39–46
Bodilis J, Ghysels B, Osayande J, Matthijs S, Pirnay J-P, Denayer S et al (2009) Distribution and evolution of ferripyoverdine receptors in Pseudomonas aeruginosa. Environ Microbiol 11:2123–2135
Budzikiewicz H (2004) Siderophores of the Pseudomonadaceae sensu stricto (fluorescent and non-fluorescent Pseudomonas spp.). Fortschr Chem Org Naturst 87:81–237
Buell CR, Joardar V, Lindeberg M, Selengut J, Paulsen IT, Gwinn ML et al (2003) The complete genome sequence of the Arabidopsis and tomato pathogen Pseudomonas syringae pv. tomato DC3000. Pr Proc Natl Acad Sci USA 100:10181–10186
Bultreys A, Gheysen I (2000) Production and comparison of peptide siderophores from strains of distantly related pathovars of Pseudomonas syringae and Pseudomonas viridiflava LMG 2352. Appl Environ Microbiol 66:325–331
Bultreys A, Gheysen I, Maraite H, de Hoffmann E (2001) Characterization of fluorescent and nonfluorescent peptide siderophores produced by Pseudomonas syringae strains and their potential use in strain identification. Appl Environ Microbiol 67:1718–1727
Bultreys A, Gheysen I, Wathelet B, Maraite H, de Hoffmann E (2003) High-performance liquid chromatography analyses of pyoverdin siderophores differentiate among phytopathogenic fluorescent Pseudomonas species. Appl Environ Microbiol 69:1143–1153
Burch AY, Shimada BK, Mullin SWA, Dunlap CA, Bowman MJ, Lindow SE (2012) Pseudomonas syringae coordinates production of a motility-enabling surfactant with flagellar assembly. J Bacteriol 194:1287–1298
Cai R, Lewis J, Yan S, Liu H, Clarke CR, Campanile F et al (2011) The plant pathogen Pseudomonas syringae pv. tomato is genetically monomorphic and under strong selection to evade tomato immunity. PLoS Pathog 7:e1002130
Carrion VJ, Gutierrez-Barranquero JA, Arrebola E, Bardaji L, Codina JC, de Vicente A et al (2013) The mangotoxin biosynthetic operon (mbo) is specifically distributed within Pseudomonas syringae genomospecies 1 and was acquired only once during evolution. Appl Environ Microbiol 79:756–767
Chien C-F, Mathieu J, Hsu C-H, Boyle P, Martin GB, Lin N-C (2013) Nonhost resistance of tomato to the bean pathogen Pseudomonas syringae pv. syringae B728a Is due to a defective E3 ubiquitin ligase domain in AvrPtoB B728a. Mol Plant Microbe Interact 26:387–397
Clarke CR, Cai R, Studholme DJ, Guttman DS, Vinatzer BA (2010) Pseudomonas syringae strains naturally lacking the classical P. syringae hrp/hrc locus are common leaf colonizers equipped with an atypical type III secretion system. Mol Plant Microbe Interact 23:198–210
Cody YS, Gross DC (1987) Outer membrane protein mediating iron uptake via pyoverdinpss, the fluorescent siderophore produced by Pseudomonas syringae pv. syringae. J Bacteriol 169:2207–2214
Cornelis P, Bodilis J (2009) A survey of TonB-dependent receptors in fluorescent pseudomonads. Environ Microbiol Reports 1:256–262
Cornelis P, Matthijs S (2002) Diversity of siderophore-mediated iron uptake systems in fluorescent pseudomonads: not only pyoverdines. Environ Microbiol 4:787–798
Cornelis P, Hohnadel D, Meyer JM (1989) Evidence for different pyoverdine-mediated iron uptake systems among Pseudomonas aeruginosa strains. Infect Immun 57:3491–3497
Cunnac S, Chakravarthy S, Kvitko BH, Russell AB, Martin GB, Collmer A (2011) Genetic disassembly and combinatorial reassembly identify a minimal functional repertoire of type III effectors in Pseudomonas syringae. Proc Natl Acad Sci USA 108:2975–2980
Davies JK, Reeves P (1975) Genetics of resistance to colicins in Escherichia coli K-12: cross-resistance among colicins of group A. J Bacteriol 123:102–117
Denayer S, Matthijs S, Cornelis P (2007) Pyocin S2 (Sa) kills Pseudomonas aeruginosa strains via the FpvA type I Ferripyoverdine receptor. J Bacteriol 189:7663–7668
Diallo MD, Monteil CL, Vinatzer BA, Clarke CR, Glaux C, Guilbaud C, Desbiez CEC, Morris CE (2012a) Pseudomonas syringae naturally lacking the canonical type III secretion system are ubiquitous in nonagricultural habitats, are phylogenetically diverse and can be pathogenic. ISME J 6:1–11
Diallo MD, Monteil CL, Vinatzer BA, Clarke CR, Glaux C, Guilbaud C, Desbiez CEC, Morris CE (2012b) Pseudomonas syringae naturally lacking the canonical type III secretion system are ubiquitous in nonagricultural habitats, are phylogenetically diverse and can be pathogenic. ISME J 6:1325–1335
Expert D, Toussaint A (1985) Bacteriocin-resistant mutants of Erwinia chrysanthemi: possible involvement of iron acquisition in phytopathogenicity. J Bacteriol 163:221–227
Fan J, Crooks C, Creissen G, Hill L, Fairhurst S, Doerner P, Lamb C (2011) Pseudomonas sax genes overcome aliphatic isothiocyanate-mediated non-host resistance in Arabidopsis. Science 331:1185–1188
Feil H, Feil WS, Chain P, Larimer F, DiBartolo G, Copeland A et al (2005) Comparison of the complete genome sequences of Pseudomonas syringae pv. syringae B728a and pv. tomato DC3000. Proc Natl Acad Sci USA 102:11064–11069
Ferrante P, Clarke CR, Cavanaugh KA, Michelmore RW, Buonaurio R, Vinatzer BA (2009) Contributions of the effector gene hopQ1-1 to differences in host range between Pseudomonas syringae pv. phaseolicola and P.syringae pv. tabaci. Mol Plant Pathol 10:837–842
Filiatrault MJ, Stodghill PV, Bronstein PA, Moll S, Lindeberg M, Grills G et al (2010) Transcriptome analysis of Pseudomonas syringae identifies new genes, noncoding RNAs, and antisense activity. J Bacteriol 192:2359–2372
Filiatrault MJ, Stodghill PV, Myers CR, Bronstein PA, Butcher BG, Lam H et al (2011) Genome-wide identification of transcriptional start sites in the plant pathogen Pseudomonas syringae pv. tomato str. DC3000. PLoS One 6:e29335
Fones H, Davis CAR, Rico A, Fang F, Smith JAC, Preston GM (2010) Metal hyperaccumulation armors plants against disease. PLoS Pathog 6:e1001093
Gardan L, Shafik H, Belouin S, Broch R, Grimont F, Grimont PA (1999) DNA relatedness among the pathovars of Pseudomonas syringae and description of Pseudomonas tremae sp. nov. and Pseudomonas cannabina sp. nov. (ex Sutic and Dowson 1959). Int J Syst Bacteriol 49:469–478
Garrett CM, Crosse JE (1963) Observations on lysogeny in the plant pathogens Pseudomonas morsprunorum and P. syringae. J Appl Microbiol 26:27–34
Garrett CM, Panagopoulos CG, Crosse JE (1966) Comparison of plant pathogenic pseudomonads from fruit trees. J Appl Microbiol 29:342–356
Geng X, Cheng J, Gangadharan A, Mackey D (2012) The coronatine toxin of Pseudomonas syringae is a multifunctional suppressor of Arabidopsis defense. Plant Cell 24:4763–4774
Ghequire MGK, Li W, Proost P, Loris R, De Mot R (2012) Plant lectin-like antibacterial proteins from phytopathogens Pseudomonas syringae and Xanthomonas citri. Environ Microbiol Reports 4:373–380
Green S, Studholme DJ, Laue BE, Dorati F, Lovell H, Arnold D et al (2010) Comparative genome analysis provides insights into the evolution and adaptation of Pseudomonas syringae pv. aesculi on Aesculus hippocastanum. PLoS One 5:e10224
Grinter R, Milner J, Walker D (2012) Bacteriocins active against plant pathogenic bacteria. Biochem Soc Trans 40:1498–1502
Groll M, Schellenberg B, Bachmann AS, Archer CR, Huber R, Powell TK et al (2008) A plant pathogen virulence factor inhibits the eukaryotic proteasome by a novel mechanism. Nature 452:755–758
Gross H, Loper JE (2009) Genomics of secondary metabolite production by Pseudomonas spp. Nat Prod Rep 26:1408–1446
Haapalainen M, Mosorin H, Dorati F, Wu RF, Roine E, Taira S et al (2012) Hcp2, a secreted protein of the phytopathogen Pseudomonas syringae pv. tomato DC3000, is required for competitive fitness against bacteria and yeasts. J Bacteriol 194:4810–4822
Harrison PW, Lower RPJ, Kim NKD, Young JPW (2010) Introducing the bacterial “chromid”: not a chromosome, not a plasmid. Trends Microbiol 18:141–148
Hattermann DR, Ries SM (1989) Motility of Pseudomonas syringae pv. glycinea and its role in infection. Phytopathology 79:284–289
Hirano S, Upper C (2000) Bacteria in the leaf ecosystem with emphasis on Pseudomonas syringae—a pathogen, ice nucleus, and epiphyte. Microbiol Mol Biol Rev 64:624
Hockett KL, Burch AY, Lindow SE (2013) Thermo-regulation of genes mediating motility and plant interactions in Pseudomonas syringae. PLoS One 8:e59850
Holtsmark I, Eijsink VGH, Brurberg MB (2008) Bacteriocins from plant pathogenic bacteria. FEMS Microbiol Lett 280:1–7
Hwang MSH, Morgan RL, Sarkar SF, Wang PW, Guttman DS (2005) Phylogenetic characterization of virulence and resistance phenotypes of Pseudomonas syringae. Appl Environ Microbiol 71:5182–5191
Jackson RW, Vinatzer B, Arnold DL, Dorus S, Murillo J (2011) The influence of the accessory genome on bacterial pathogen evolution. Mob Genet Elements 1:55–65
Joardar V, Lindeberg M, Jackson RW, Selengut J, Dodson R, Brinkac LM et al (2005) Whole-genome sequence analysis of Pseudomonas syringae pv. phaseolicola 1448A reveals divergence among pathovars in genes involved in virulence and transposition. J Bacteriol 187:6488–6498
Jones AM, Wildermuth MC (2011) The phytopathogen Pseudomonas syringae pv. tomato DC3000 has three high-affinity iron-scavenging systems functional under iron limitation conditions but dispensable for pathogenesis. J Bacteriol 193:2767–2775
Jones JDG, Dangl JL (2006) The plant immune system. Nature 444:323–329
Joyner DC, Lindow SE (2000) Heterogeneity of iron bioavailability on plants assessed with a whole-cell GFP-based bacterial biosensor. Microbiology 146:2435–2445
Jülich M, Taraz K, Budzikiewicz H, Geoffroy V, Meyer JM, Gardan L (2001) The structure of the pyoverdin isolated from various Pseudomonas syringae pathovars. Z Naturforsch C 56:687–694
Karamanoli K, Bouligaraki P, Constantinidou HIA, Lindow SE (2011) Polyphenolic compounds on leaves limit iron availability and affect growth of epiphytic bacteria. Ann Appl Biol 159:99–108
Kim JJ, Sundin GW (2000) Regulation of the rulAB mutagenic DNA repair operon of Pseudomonas syringae by UV-B (290 to 320 Nanometers) radiation and analysis of rulAB-mediated mutability in vitro and in planta. J Bacteriol 182:6137–6144
Kniskern JM, Barrett LG, Bergelson J (2010) Maladaptation in wild populations of the generalist plant pathogen Pseudomonas syringae. Evolution 65:818–830
Koskella B, Thompson JN, Preston GM, Buckling A (2011) Local biotic environment shapes the spatial scale of bacteriophage adaptation to bacteria. Am Nat 177:440–451
Kvitko BH, Park DH, Velásquez AC, Wei C-F, Russell AB, Martin GB et al (2009) Deletions in the repertoire of Pseudomonas syringae pv. tomato DC3000 Type III secretion effector genes reveal functional overlap among effectors. PLoS Pathog 5:e1000388
Lacombe SEV, Rougon-Cardoso A, Sherwood E, Peeters N, Dahlbeck D, van Esse HP et al (2010) Interfamily transfer of a plant pattern-recognition receptor confers broad-spectrum bacterial resistance. Nat Biotechnol 28:365–369
Laue H (2006) Contribution of alginate and levan production to biofilm formation by Pseudomonas syringae. Microbiology 152:2909–2918
Lavermicocca P, Lonigro SL, Evidente A, Andolfi A (1999) Bacteriocin production by Pseudomonas syringae pv. ciccaronei NCPPB2355. Isolation and partial characterization of the antimicrobial compound. J Appl Microbiol 86:257–265
Lavermicocca P, Lonigro SL, Valerio F, Evidente A, Visconti A (2002) Reduction of olive knot disease by a bacteriocin from Pseudomonas syringae pv. ciccaronei. Appl Environ Microbiol 68:1403–1407
Lin N-C, Martin GB (2005) An avrpto/avrptoB mutant of Pseudomonas syringae pv. tomato DC3000 does not elicit Pto-mediated resistance and is less virulent on tomato. Mol Plant Microbe Interact 18:43–51
Lin N-C, Martin GB (2007) Pto- and Prf-mediated recognition of AvrPto and AvrPtoB restricts the ability of diverse Pseudomonas syringae pathovars to infect tomato. Mol Plant Microbe Interact 20:806–815
Lindeberg M, Cunnac S, Collmer A (2012) Pseudomonas syringae type III effector repertoires: last words in endless arguments. Trends Microbiol 20:199–208
Lindeberg M, Myers CR, Collmer A, Schneider DJ (2008) Roadmap to new virulence determinants in Pseudomonas syringae: insights from comparative genomics and genome organization. Mol Plant Microbe Interact 21:685–700
Lindow SE, Andersen G, Beattie GA (1993) Characteristics of insertional mutants of Pseudomonas syringae with reduced epiphytic fitness. Appl Environ Microbiol 59:1593–1601
Lovell HC, Mansfield JW, Godfrey SAC, Jackson RW, Hancock JT, Arnold DL (2009) Bacterial evolution by genomic island transfer occurs via DNA transformation In Planta. Curr Biol 19:1586–1590
Ma Z, Smith JJ, Zhao Y, Jackson RW, Arnold DL, Murillo J, Sundin GW (2007) Phylogenetic analysis of the pPT23A plasmid family of Pseudomonas syringae. Appl Environ Microbiol 73:1287–1295
Marcelletti S, Ferrante P, Petriccione M, Firrao G, Scortichini M (2011) Pseudomonas syringae pv. actinidiae draft genomes comparison reveal strain-specific features involved in adaptation and virulence to Actinidia species. PLoS One 6:e27297
McCann HC, Nahal H, Thakur S, Guttman DS (2012) Identification of innate immunity elicitors using molecular signatures of natural selection. Proc Natl Acad Sci USA 109:4215–4220
Melotto M, Underwood W, Koczan J, Nomura K, He SY (2006) Plant stomata function in innate immunity against bacterial invasion. Cell 126:969–980
Meyer JM, Neely A, Stintzi A, Georges C, Holder IA (1996) Pyoverdin is essential for virulence of Pseudomonas aeruginosa. Infect Immun 64:518–523
Meyer J-M (2000) Pyoverdines: pigments, siderophores and potential taxonomic markers of fluorescent Pseudomonas species. Arch Microbiol 174:135–142
Minor SM, Nordeen RO, Pachall R (1996) Partial characterization of bacteriophages of Pseudomonas syringae pv. tomato. Proc Ark Acad Sci 50:137–140
Mohr TJ, Liu H, Yan S, Morris CE, Castillo JA, Jelenska J, Vinatzer BA (2008) Naturally occurring nonpathogenic isolates of the plant pathogen Pseudomonas syringae lack a type III secretion system and effector gene orthologues. J Bacteriol 190:2858–2870
Morris CE, Sands DC, Vanneste JL, Montarry J, Oakley B, Guilbaud C, Glaux C (2010) Inferring the evolutionary history of the plant pathogen Pseudomonas syringae from its biogeography in headwaters of rivers in North America, Europe, and New Zealand. mBio 1: e00107-10-e00107-20
Morris CE, Sands DC, Vinatzer BA, Glaux C, Guilbaud C, Buffière A et al (2008) The life history of the plant pathogen Pseudomonas syringae is linked to the water cycle. ISME J 2:321–334
Nadarasah G, Stavrinides J (2011) Insects as alternative hosts for phytopathogenic bacteria. FEMS Microbiol Rev 35:555–575
Nicaise V, Roux M, Zipfel C (2009) Recent advances in PAMP-triggered immunity against bacteria: pattern recognition receptors watch over and raise the alarm. Plant Physiol 150:1638–1647
Nordeen RO, Morgan MK, Currier TC (1983) Isolation and partial characterization of bacteriophages of the phytopathogen Pseudomonas syringae. Appl Environ Microbiol 45:1890–1898
Oguiza JA, Kiil K, Ussery DW (2005) Extracytoplasmic function sigma factors in Pseudomonas syringae. Trends Microbiol 13:565–568
Owen JG, Ackerley DF (2011) Characterization of pyoverdine and achromobactin in Pseudomonas syringae pv. phaseolicola 1448a. BMC Microbiol 11:218
O’Brien HE, Thakur S, Guttman DS (2011) Evolution of plant pathogenesis in Pseudomonas syringae: a genomics perspective. Ann Rev Phytopathol 49:269–289
O’Brien HE, Thakur S, Gong Y, Fung P, Zhang J, Yuan L et al (2012) Extensive remodeling of the Pseudomonas syringae pv. avellanae type III secretome associated with two independent host shifts onto hazelnut. BMC Microbiol 12:141
Panopoulos NJ, Schroth MN (1974) Role of flagellar motility in the invasion of bean leaves by Pseudomonas phaseolicola. Phytopathology 64:1389–1397
Parret AHA, De Mot R (2002) Bacteria killing their own kind: novel bacteriocins of Pseudomonas and other gamma-proteobacteria. Trends Microbiol 10:107–112
Preston GM (2000) Pseudomonas syringae pv. tomato: the right pathogen, of the right plant, at the right time. Mol Plant Pathol 1:263–275
Prior ES, Andrews JA, Nordeen OR (2007) Characterization of bacteriophages of Pseudomonas syringae pv. tomato. Proc Ark Acad Sci 61:84–90
Quigley NB, Gross DC (1994) Syringomycin production among strains of Pseudomonas syringae pv. syringae: conservation of the syrB and syrD genes and activation of phytotoxin production by plant signal molecules. Mol Plant Microbe Interact 7:78–90
Quiñones B, Dulla G, Lindow SE (2005) Quorum sensing regulates exopolysaccharide production, motility, and virulence in Pseudomonas syringae. Mol Plant Microbe Interact 18:682–693
Ramos C, Matas IM, Bardaji L, Aragón IM, Murillo J (2012) Pseudomonas savastanoi pv. savastanoi: some like it knot. Mol Plant Pathol 13:998–1009
Records AR, Gross DC (2010) Sensor kinases RetS and LadS regulate Pseudomonas syringae type VI secretion and virulence factors. J Bacteriol 192:3584–3596
Rico A, Preston GM (2008) Pseudomonas syringae pv. tomato DC3000 uses constitutive and apoplast-induced nutrient assimilation pathways to catabolize nutrients that are abundant in the tomato apoplast. Mol Plant Microbe Interact 21:269–282
Rodríguez-Palenzuela P, Matas IM, Murillo J, López-Solanilla E, Bardaji L, Pérez-Martínez I, Rodríguez-Moskera ME, Penyalver R, López MM, Quesada JM, Biehl BS, Perna NT, Glasner JD, Cabot EL, Neeno-Eckwall E, Ramos C (2010) Annotation and overview of the Pseudomonas savastanoi pv. savastanoi NCPPB 3335 draft genome reveals the virulence gene complement of a tumour-inducing pathogen of woody hosts. Environ Microbiol 12:1–17
Sarris PF, Skandalis N, Kokkinidis M, Panopoulos NJ (2010) In silico analysis reveals multiple putative type VI secretion systems and effector proteins in Pseudomonas syringae pathovars. Mol Plant Pathol 11:795–804
Sarris PF, Trantas EA, Baltrus DA, Bull CT, Wechter WP, Yan S et al (2013) Comparative genomics of multiple strains of Pseudomonas cannabina pv. alisalensis, a potential model pathogen of both monocots and dicots. PLoS One 8:e59366
Sato M (1983) Phage-induction from lysogenic strains of Pseudomonas syringae pv. mori by the extract from mulberry leaves. Ann Phytopathol Soc Japan 49:259–261
Schellenberg B, Ramel C, Dudler R (2010) Pseudomonas syringae virulence factor syringolin A counteracts stomatal immunity by proteasome inhibition. Mol Plant Microbe Interact 23:1287–1293
Scholz-Schroeder BK, Hutchison ML, Grgurina I, Gross DC (2001) The contribution of syringopeptin and syringomycin to virulence of Pseudomonas syringae pv. syringae strain B301D on the basis of sypA and syrB1 biosynthesis mutant analysis. Mol Plant Microbe Interact 14:336–348
Schulze-Lefert P, Panstruga R (2011) A molecular evolutionary concept connecting nonhost resistance, pathogen host range, and pathogen speciation. Trends Plant Sci 16:117–125
Scortichini M, Marcelletti S, Ferrante P, Petriccione M, Firrao G (2012) Pseudomonas syringae pv. actinidiae: a re-emerging, multi-faceted, pandemic pathogen. Mol Plant Pathol 13:631–640
Segonzac C, Zipfel C (2011) Activation of plant pattern-recognition receptors by bacteria. Curr Opin Microbiol 14:54–61
Smith EE, Sims EH, Spencer DH, Kaul R, Olson MV (2005) Evidence for diversifying selection at the pyoverdine locus of Pseudomonas aeruginosa. J Bacteriol 187:2138–2147
Spencer DH, Kas A, Smith EE, Raymond CK, Sims EH, Hastings M et al (2003) Whole-genome sequence variation among multiple isolates of Pseudomonas aeruginosa. J Bacteriol 185:1316–1325
Stavrinides J, McCloskey JK, Ochman H (2009) Pea aphid as both host and vector for the phytopathogenic bacterium Pseudomonas syringae. Appl Environ Microbiol 75:2230–2235
Studholme DJ (2011) Application of high-throughput genome sequencing to intrapathovar variation in Pseudomonas syringae. Mol Plant Pathol 12:829–838
Sundin GW (2007) Genomic insights into the contribution of phytopathogenic bacterial plasmids to the evolutionary history of their hosts. Ann Rev Phytopathol 45:129–151
Taguchi F, Suzuki T, Inagaki Y, Toyoda K, Shiraishi T, Ichinose Y (2010) The siderophore pyoverdine of Pseudomonas syringae pv. tabaci 6605 is an intrinsic virulence factor in host tobacco infection. J Bacteriol 192:117–126
Takase H, Nitanai H, Hoshino K, Otani T (2000) Impact of siderophore production on Pseudomonas aeruginosa infections in immunosuppressed mice. Infect Immun 68:1834–1839
Tampakaki AP, Skandalis N, Gazi AD, Bastaki MN, Panagiotis FS, Charova SN et al (2010) Playing the “Harp”: evolution of our understanding of hrp/hrc genes. Ann Rev Phytopathol 48:347–370
Taylor J, Teverson D, Allen D (1996) Identification and origin of races of Pseudomonas syringae pv. phaseolicola from Africa and other bean growing areas. Plant Pathol 45:469–478
Thakur PB, Vaughn-Diaz VL, Greenwald JW, Gross DC (2013) Characterization of five ECF sigma factors in the genome of Pseudomonas syringae pv. syringae B728a. PLoS ONE 8:e58846
Thomma BPHJ, Nurnberger T, Joosten MHAJ (2011) Of PAMPs and effectors: the blurred PTI-ETI dichotomy. Plant Cell 23:4–15
Ude S, Arnold DL, Moon CD, Timms-Wilson T, Spiers AJ (2006) Biofilm formation and cellulose expression among diverse environmental Pseudomonas isolates. Environ Microbiol 8:1997–2011
Verdier V, Triplett LR, Hummel AW, Corral R, Cernadas RA, Schmidt CL et al (2012) Transcription activator-like (TAL) effectors targeting OsSWEET genes enhance virulence on diverse rice (Oryza sativa) varieties when expressed individually in a TAL effector-deficient strain of Xanthomonas oryzae. New Phytol 196:1197–1207
Vidaver AK, Mathys ML, Thomas ME, Schuster ML (1972) Bacteriocins of the phytopathogens Pseudomonas syringae, P. glycinea, and P. phaseolicola. Can J Microbiol 18(6):705–713
Visca P, Imperi F, Lamont IL (2007) Pyoverdine siderophores: from biogenesis to biosignificance. Trends Microbiol 15:22–30
Vivian A, Murillo J, Jackson RW (2001) The roles of plasmids in phytopathogenic bacteria: mobile arsenals? Microbiology 147:763–780
Wei C-F, Kvitko BH, Shimizu R, Crabill E, Alfano JR, Lin N-C et al (2007) A Pseudomonas syringae pv. tomato DC3000 mutant lacking the type III effector HopQ1-1 is able to cause disease in the model plant Nicotiana benthamiana. Plant J 51:32–46
Wensing A, Braun SD, Buttner P, Expert D, Völksch B, Ullrich MS, Weingart H (2010) Impact of siderophore production by Pseudomonas syringae pv. syringae 22d/93 on epiphytic fitness and biocontrol activity against Pseudomonas syringae pv. glycinea 1a/96. Appl Environ Microbiol 76:2704–2711
Wichmann G, Sun J, Dementhon K, Glass NL, Lindow SE (2008) A novel gene, phcA from Pseudomonas syringae induces programmed cell death in the filamentous fungus Neurospora crassa. Mol Microbiol 68:672–689
Wilson M, Lindow SE (1994) Coexistence among epiphytic bacterial populations mediated through nutritional resource partitioning. Appl Environ Microbiol 60:4468–4477
Wroblewski T, Caldwell KS, Piskurewicz U, Cavanaugh KA, Xu H, Kozik A et al (2009) Comparative large-scale analysis of interactions between several crop species and the effector repertoires from multiple pathovars of Pseudomonas and Ralstonia. Plant Physiol 150:1733–1749
Young JM (2010) Taxonomy of Pseudomonas syringae. J. Plant Pathol 92(1): S1-5–14
Yu X, Lund SP, Scott RA, Greenwald JW, Records AH, Nettleton D et al (2013) Transcriptional responses of Pseudomonas syringae to growth in epiphytic versus apoplastic leaf sites. Proc Natl Acad Sci USA 110:E425–E434
Zhang S, Sundin GW (2004) Long-term effect of mutagenic DNA repair on accumulation of mutations in Pseudomonas syringae B86-17. J Bacteriol 186:7807–7810
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Baltrus, D.A., Hendry, T.A., Hockett, K.L. (2014). Ecological Genomics of Pseudomonas syringae . In: Gross, D., Lichens-Park, A., Kole, C. (eds) Genomics of Plant-Associated Bacteria. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-55378-3_3
Download citation
DOI: https://doi.org/10.1007/978-3-642-55378-3_3
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-55377-6
Online ISBN: 978-3-642-55378-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)