Abstract
CRISPR-Cas systems provide adaptive immunity against viruses and plasmids in bacteria and archaea. Interference is mediated by small non-coding CRISPR RNAs (crRNAs) that guide the Cas machinery towards complementary nucleic acids for sequence-specific cleavage. Several recent studies have shown that CRISPR-encoded immunity can increase the breadth and depth of phage resistance in bacteria, and can provide a barrier to acquisition of undesirable genetic elements, notably plasmid-encoded antibiotic resistance genes. Further, the adaptive and inheritable nature of those idiosyncratic chromosomal loci provide valuable genetic polymorphism which can be leveraged for typing purposes, proprietary strain tagging, ecological surveys, and epidemiological studies. The ability to readily transfer functional CRISPR-Cas systems across even distant bacteria, and re-program their endonuclease activity make them amenable to genetic engineering and useful for genome editing. These features, in combination with recent breakthroughs in unravelling the molecular underpinnings of the CRISPR mechanism of action have paved the way for several applications in a diversity of industrial and biotechnological areas.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
11.1 Introduction
While warfare is continuously being waged between microbes and their viral counterparts, arguably endlessly, a novel weapon is occasionally discovered in the arsenal, which might be exploited by humans to shift the balance between the conflicting parties. The raging battle between bacteria used as starter cultures in the food industry for the fermentation of milk into appetizing products such as yogurt and cheese, and their predatory bacteriophages has lasted for centuries, ever since the need to store surplus milk has arisen. Recently, CRISPR-Cas systems were shown to provide adaptive resistance against viruses of bacteria and archaea, and numerous studies have documented their functional properties, characterizing the molecular underpinnings of their biochemical mechanism of action. These studies have set the stage for leveraging those versatile molecular systems in a variety of technological applications.
The historical path that the CRISPR field has taken has been discussed in detail previously (see Chap. 1), and the occurrence, distribution, and evolution of those loci outlined (see Chaps. 2 and 3), with approximately 46 % of bacteria and 90 % of archaea carrying CRISPR loci, including many model and industrially relevant organisms. Notwithstanding the various types of CRISPR-Cas systems that have been established in the literature (see Chaps. 3, 5, 6, and 7), there are many elements that are somewhat conserved across CRISPR-Cas systems, both in mechanism of action and in function(s) that set the stage for a wide array of technological applications.
Although the field might be considered by many in its infancy, the CRISPR literature and citation rates reflect both the quantity and quality of the work that has been performed over the past decade. Further, the ability to potentially translate this work into tangible applications can be somewhat measured by the intellectual property activity, as monitored by a number of patent application deposits. To date, 12 patents related to CRISPR uses and applications have been published (see Table 11.1).
These patents span several distinct areas and types of applications, notably the detection and typing of bacterial strains, the development of phage resistance, and the use of CRISPR-Cas systems for interference and cleavage of nucleic acids. We highlight below a number of documented and potential applications of CRISPR-Cas systems.
11.2 Resistance Against Viruses
A variety of roles have been attributed to the diverse CRISPR-Cas systems within the last 10 years, including DNA repair and biofilm inhibition (Babu et al. 2011; Palmer and Whiteley 2011). Nevertheless, it has been quickly and broadly accepted that resistance against bacteriophages (phages), and more generally against viruses, is the primary and most common role of these small RNA-based interference systems. Hypothesized in 2005 (Pourcel et al. 2005; Mojica et al. 2005; Bolotin et al. 2005), the antiviral activity of CRISPR-Cas was demonstrated shortly thereafter with a food-grade bacterium of industrial relevance (Barrangou et al. 2007; see Table 11.1, patent application WO/2007/025097). Indeed, large-scale dairy fermentations using Streptococcus thermophilus-containing starter cultures are occasionally impaired by lytic phages, compelling starter cultures companies to constantly devise strategies aimed at controlling phage populations in industrial settings.
11.2.1 “CRISPerization”: Phage Resistance Improvement Through Iterative Challenges
Traditionally, phages have been extensively used—intentionally or otherwise—to challenge sensitive bacterial strains, in order to select subpopulations named Bacteriophage-Insensitive Mutants (BIMs) that display increased viral resistance (Labrie et al. 2010). Besides providing naturally improved strains, such approaches have led to the identification of a variety of phage resistance mechanisms, notably those involved in the early steps (phage adsorption onto cell receptor(s), phage DNA injection within the cytoplasm) of the phage–host interaction (Sturino and Klaenhammer 2006). Although relatively easy to generate, BIMs generally show a weak and volatile protection against phages, mainly because phage populations evolve at a faster rate than their hosts. Furthermore, receptor mutations only provide resistance against a narrow spectrum of phages that use a conserved pathway for infection.
Some CRISPR-Cas systems have been shown to be responsive to viral challenge, either naturally (Barrangou et al. 2007; Deveau et al. 2008; van der Ploeg 2009; Mills et al. 2010; Cady et al. 2012; Erdmann and Garrett 2012) or following genetic engineering and priming (Datsenko et al. 2012; Swarts et al. 2012; Yosef et al. 2012). Specifically, in the acquisition stage, small pieces (called proto-spacers) of the viral nucleic acid may be integrated as new spacers in-between new repeats at the leader end of CRISPR array(s), thus providing adaptive immunity. The presence of such additional spacers, subsequently transcribed in order to interfere with any complementary sequence, confers an improved resistance to the surviving host cell.
Based on this CRISPR-Cas adaptive system, “CRISPerization” strategies have been developed to rationally and purposefully generate improved lineages in S. thermophilus (see Table 11.1, notably patent application WO/2008/108989, and Fig. 11.1). Provided sufficient—both in number and diversity—virulent phages are available, iterative phage challenges may be performed (endlessly?) to increase the level of resistance of the host strain, leading to a stacking of newly acquired spacers. Furthermore, by selecting genetically diverse and industrially relevant phages, subsequent challenges advantageously broaden the spectrum of resistance of the host strain. Due to the apparent randomness of proto-spacer uptake (though new data suggest the proto-spacer sampling process is not completely random; Datsenko et al. 2012) and the broad reservoir of proto-spacers within each phage genome (typically several hundreds in a 35 kb genome), distinct bacterial lineages with complementary resistances may be generated by using independently the same phages, in the same order or otherwise (see Fig. 11.1).
CRISPerization process. The diagram displays the way in which CRISPR BIMs (bacteriophage insensitive mutants) can be selected following iterative exposure to phages (φA through φG), to generate multi-generational variants (V1–V5) that have acquired several new CRISPR spacers, eventually making them resistant to all phages used. Colored rectangles and other shapes represent CRISPR spacers newly acquired, with each color corresponding to the phage used in the challenge. WT, wild-type (parental) strain. Note that all phages do not need to be used in each lineage, as some spacers may be efficient against distinct phages sharing common sequences. This strategy can be enhanced by selecting CRISPR BIMs that have acquired spacers in multiple active CRISPR loci, that have acquired multiple spacers in a single round of phage exposure, and by selecting spacers that target highly conserved and/or functional sequences in phage genomes
CRISPerization by iterative challenges holds three major advantages over other phage resistance improvement strategies. First, the resulting variants are “natural microorganisms”, a trait which is currently critical to the food industry, notably in Europe. No genetic engineering is involved in the process, which is purely based on the generation, selection, and characterization of surviving subpopulations. Second, all variants that are obtained, whatever the number of iterations they underwent, are isogenic variants of the parental wild-type strain, that have maintained their valuable functional properties. Theoretically they only carry mutations (i.e., additional repeat-spacer units) in their CRISPR array(s), thus maintaining identical physiological and functional properties, another critical trait for the robustness of industrial applications. Obviously, combinations of various isogenic strains in rotation schemes are highly valuable in the dairy manufacturing environment, and provide increased phage resistance both in terms of depth of phage resistance and breadth of the phage resistance spectrum. While the industry historically relies on rotation strategies combining distinct phage resistance mechanisms and phenotypes, CRISPR-mediated phage resistance provides advantages both in terms of isogenic variants’ sustainable use, and stability of the chromosomally encoded resistance system, as opposed to plasmid-borne. Highly advanced “CRISPerized” strains can thus be considered as variants with an extended lifespan, which may eventually be immortalized. Finally, provided sufficient (both in number and diversity) spacers are acquired in controlled, laboratory conditions, it may become difficult—or even impossible—for the phages naturally occurring in the environment to circumvent the CRISPR-encoded immunity. CRISPerization through iterative challenges may be a clever way to get ahead in the alleged never-ending arms race between hosts and their predatory viruses.
11.2.2 Artificial Spacer Engineering
As opposed to natural spacer acquisition following viral challenge, additional spacers can be intentionally introduced into CRISPR arrays by using classical genetic engineering approaches. Only a relatively short segment (i.e., the size of a spacer) of the target nucleic acid sequence has to be known in order to build specific immunity against any complementary sequence (provided the associated proto-spacer adjacent motif is taken into account). A conservative, safe strategy is to “copy-paste” naturally occurring spacers (belonging to the same CRISPR-Cas system) between characterized strains. By extension, spacers can be designed entirely de novo prior to their integration between CRISPR repeats, so that it is virtually feasible to confer immunity against nucleic acid sequences that have never been observed yet. Engineered CRISPR arrays can also be an answer to strain improvement when no lytic virus is available for challenge, or when no virus is efficiently stimulating novel spacer acquisition.
We brought the first illustration of this approach in S. thermophilus in 2007 (Barrangou et al. 2007). Spacers S1 and S2, simultaneously acquired in the CRISPR1 locus of a BIM following a challenge with phage 858, were cloned from their host strain into a plasmid and transferred to the CRISPR1 locus of another strain, thereby transferring immunity against phage 858. De novo spacer engineering against phage Lambda was also performed in Escherichia coli (Brouns et al. 2008; Pougach et al. 2010; Sapranauskas et al. 2011), showing that CRISPR-Cas systems can be specifically engineered to contain particular spacers that target phage sequences and provide resistance against viruses that carry homologous sequences.
The major limit of artificial spacer engineering is the fact that not all sequences constitute efficient CRISPR spacers, due to the need, in some CRISPR-Cas systems, for a proto-spacer-associated motif (PAM) (Deveau et al. 2008; Horvath et al. 2008; Mojica et al. 2009). In such cases, the selection in a target nucleic acid of a sequence to be converted into a CRISPR spacer is constrained by the presence of an adequate PAM sequence.
11.2.3 Transfer Between Microorganisms
The propensity of CRISPR-Cas systems to be subjected to horizontal gene transfer has been documented for a while (Godde and Bickerton 2006; Horvath et al. 2009), and reflects the distribution and evolution of those systems, as discussed in Chaps. 2 and 3, respectively.
Finally, phage resistance may also be obtained through the transfer of a complete CRISPR-Cas system between strains (not necessarily belonging to the same species), as exemplified by Sapranauskas et al. (2011). After cloning of the S. thermophilus CRISPR3-Cas system on a plasmid, it was readily transferred into E. coli, and could provide resistance against phage and lower plasmid uptake propensity. The next major advance will be to assess whether functional systems can be transferred and/or engineered to provide nucleic acid interference in valuable, important, and model eukaryotic organisms, especially for agricultural, biotechnological, and medical applications (see Table 11.1, patent applications WO/2012/054726 and WO/2011/143124). As the visibility of the field increases, we expect that attempts will be made to engineer CRISPR-encoded interference in yeast, fungi, plants, and perhaps vertebrates.
11.3 Immunity Against Non-Viral Nucleic Acids
Although resistance against viruses is arguably the primary functional role of CRISPR-Cas systems, as it provides immunity against nucleic acids through base-pairing between spacer-derived crRNAs and complementary target sequences, DNA or RNA molecules other than virus-encoded may be subjected to interference. Indeed, similarity searches of spacer sequences within DNA databases generally show that, besides the large majority of matches with viral sequences, most of the matches correspond to plasmid sequences, followed by a minority of hits to chromosomal sequences (Horvath et al. 2008; Stern et al. 2010). The low occurrence of plasmid- and chromosome-derived spacers in CRISPR arrays may probably be considered as a side effect of the adaptive nature of CRISPR-Cas systems, whereby host genetic material (or the transcription products thereof), are perceived as “foreign” rather than “self” nucleic acid molecules. Thus, CRISPR/Cas systems may be exploited to provide non-viral immunity.
11.3.1 Plasmid Interference
Several reports in the literature document the ability of CRISPR-Cas systems to provide interference against plasmid DNA. It was first established in a milestone and elegant study in Staphylococcus epidermidis whereby CRISPR-encoded spacers lowered efficiency of plasmid uptake. This study also established that the primary CRISPR-Cas nucleic acid target is DNA (Marraffini and Sontheimer 2008). Subsequently, several studies showed that spacers can be acquired from plasmid sequences, and interfere with plasmid uptake (Garneau et al. 2010; Sapranauskas et al. 2011; Swarts et al. 2012; Datsenko et al. 2012; Jinek et al. 2012; Gasiunas et al. 2012).
11.3.2 Interference Against Other Mobile Elements
The documented ability of CRISPR-Cas systems to preclude plasmid uptake in S. epidermidis, S. thermophilus, and E. coli has set the stage for developing CRISPR-based systems that provide interference against mobile genetic elements. Given the elevated concerns about antibiotic resistance marker dissemination, especially in clinically relevant human pathogens, in combination with the circumstantial evidence which indicated a negative correlation between the occurrence of CRISPR-Cas systems and pathogenicity in Enterococcus (Palmer and Gilmore 2010) and Campylobacter (Schouls et al. 2003; Louwen et al. 2012), there is tremendous potential in leveraging CRISPR-mediated interference against antibiotic resistance genes. A recent report documenting the ability of S. thermophilus to naturally acquire spacers that target an antibiotic resistance gene (Garneau et al. 2010), in combination with the ability of the acquired spacers to preclude the uptake of plasmids that carry homologous DNA sequences, sets the stage for vaccination of bacterial strains against antibiotic resistance marker uptake. Similarly, because prophages can also readily mediate the transfer of pathogenic markers, CRISPR-encoded immunity can be used to reduce the pathogenic potential of a microorganism though reduction of its propensity to uptake novel DNA. This is consistent with the reported negative correlation between the occurrence of prophages and CRISPR spacers in Streptococcus pyogenes (Nozawa et al. 2011). As such, active CRISPR-Cas systems provide a natural means to select strains that are unlikely to uptake and disseminate antibiotic resistance markers and pathogenic traits. Likewise, the ability to engineer CRISPR-Cas systems with synthetic spacers provides an in vitro means to generate mutants that are refractory to the uptake of undesirable sequences. Indeed, recent reports show that CRISPR can prevent natural transformation and virulence marker acquisition in Streptococcus pneumoniae (Bikard et al. 2012), and influence mobilome diversity in Streptococcus agalactiae (Lopez-Sanchez et al. 2012).
Prokaryotic genome integrity and stability may be affected by the integration or excision of mobile genetic elements such as transposons and prophages. Spacers designed to target transposons and mobile genetic elements that mediate chromosomal rearrangements and shuffling could be used to increase chromosomal stability and integrity. Likewise, spacers targeting undesirable genes such as those coding for antibiotic resistance, toxins, and virulence factors could be used to generate “safer” strains.
11.4 CRISPR-Based Gene Regulation
Despite the genetic commonalities observed across the three CRISPR-Cas types, and the conservation of several mechanistic steps in various systems, in some Type III systems, at least, CRISPR targets RNA in vitro (Hale et al. 2009; Garrett et al. 2011). Accordingly, there is potential to use CRISPR-Cas systems for the regulation, transcriptional control, or regulation of transcript levels within a cell (Horvath and Barrangou 2010; see Table 11.1, patent application WO/2010/075424). A recent report illustrates the ability of CRISPR spacers to lower transcript levels, showing that a spacer homologous to the histidyl-tRNA synthetase sequence lowers His-tRNA levels (Aklujkar and Lovley 2010). A study documenting several examples of self-targeting spacers shows that this phenomenon may be under-appreciated (Stern et al. 2010). Likewise, several studies have implicated self-targeting CRISPR spacers in Pseudomonas aeruginosa lysogeny (Zegans et al. 2009; Cady and O’Toole 2011 ).
Analogies between CRISPR-mediated interference and RNA interference have been discussed in several reviews, and multiple studies have provided enough circumstantial evidence that crRNA can silence transcripts to pave the way for CRISPR-based mRNA targeting. Further, given the ancillary and emerging roles of CRISPR-Cas systems beyond foreign DNA defensive targeting (see Chap. 10), notably host regulatory and developmental processes, there are several RNA-targeting applications that can be developed. Given the tremendous interest in and the many successes of RNAi in eukaryotic systems, together with the growing importance of non-coding small RNAs in numerous biological functions, there is potential to harness the flexibility and modularity of CRISPR-Cas systems for RNA interference in bacteria and archaea (see Table 11.1, patent applications WO/2010/011961 and WO/2010/054108).
11.5 CRISPR-Based Strain Typing
A historical review of the CRISPR literature over time (see Chap. 1) clearly illustrates the potential of CRISPR loci for genotyping of bacteria. Several early studies that preceded the implication of CRISPR-Cas systems in adaptive immunity, notably spoligotyping in the early 1990s, have actually shown that these loci are both hypervariable, and provide a time-dependent iterative record of the environmental conditions to which a strain has been exposed. A milestone method describing the use of “direct repeat region” DNA sequences in the chromosome of Mycobacterium tuberculosis in 1993 (Groenen et al. 1993) had the foresight to observe that there was tremendous polymorphism across a diversity of strains in this particular region, and that sequence content could be digitized to monitor the epidemiology of clinical cases and samples of tuberculosis (Brudey et al. 2006). A similar approach was subsequently used and developed for Corynebacterium diphtheriae (Mokrousov et al. 2007, 2009). Undoubtedly, this is an insightful example of the contribution of genomics to the discovery of unknown, uncharacterized, occasionally un- or mis-annotated regions that nonetheless are hypervariable enough to provide a basis for genotyping. Indeed, the literature spans distant industrial or pathogenic bacteria across which CRISPR-based genotyping provides insights, notably M. tuberculosis (Abadia et al. 2010; Borile et al. 2011; Brudey et al. 2006; Groenen et al. 1993; Zhang et al. 2010), Yersinia pestis (Cui et al. 2008; Pourcel et al. 2005; Riehm et al. 2012; Vergnaud et al. 2007), C. diphtheriae (Mokrousov et al. 2007, 2009), P. aeruginosa (Cady et al. 2011), Legionella (D’Auria et al. 2010; Ginevra et al. 2012), S. pyogenes (Hoe et al. 1999; McShan et al. 2008), S. thermophilus (Horvath et al. 2008), Lactobacillus (see Table 11.1, patent WO/2006/073445), Propionibacterium acnes (Brüggemann et al. 2012), Erwinia amylovora (Rezzonico et al. 2011; McGhee and Sundin 2012), Campylobacter (Tasaki et al. 2012), Salmonella (Liu et al. 2011a, b; Fabre et al. 2012; Fricke et al. 2011; see Table 11.1, patent application WO/2009/115861), and pathogenic E. coli (Díez-Villaseñor et al. 2010; Delannoy et al. 2012).
Over time, the molecular methods that target CRISPR sequences have evolved. Initially, hybridization-based spoligotyping was developed in Mycobacterium and Corynebacterium, although results were highly dependent on the reference database, and solely known sequences could be targeted. Later on, Sanger-sequencing of CRISPR PCR amplicons, either completely or partially from the extremities, was developed and implemented for the genotyping of some species (see Fig. 11.2). Alternatives to sequencing were also assessed to compare and contrast CRISPR PCR amplicons, notably restriction fragment length polymorphism (RFLP) assays, capillary electrophoresis analysis, and melting curve analysis (Price et al. 2007). Nowadays, the ubiquitous and affordable natures of multiple sequencing technologies have rendered such approaches nearly obsolete. In fact, the pace of next-generation sequencing technologies development, in combination with the ever-increasing throughput and rapidly decreasing price, have opened new avenues for deep sequencing analysis of CRISPR amplicons and mixed population metagenomes. Currently, sequencing technologies have out-paced the development of fast, efficient, and convenient bioinformatic tools which provide the reconstruction of CRISPR loci and visualization of their content.
CRISPR-based typing schemes, Panel A: sequencing CRISPR arrays from both ends. For strain typing purposes, sequencing from both the leader and trailer (i.e., opposite to the leader) ends should always be preferred, when possible. Ancient spacers at the trailer end allow clustering of distantly related strains, while leader-end spacers, more recently acquired, differentiate closely related strains. In many cases, sequencing the whole CRISPR repeat-spacer array requires significant time and effort but adds only little information. If necessary, sequencing by walking across can be performed by designing primers within non-redundant spacer sequences. Panel B: one-sided CRISPR typing. When the sequences surrounding CRISPR arrays are polymorphic or unknown, sequencing is still possible from the conserved leader end, especially when cas genes are present. The PCR amplicon mix generated by using a reverse primer designed within the repeat sequence can be sequenced from the leader end and/or from internal spacers
11.6 Bacterial or Viral Strain Tracking
Further, the presence and diversity of CRISPR-Cas systems and their hypervariable spacer sequences in a diversity of industrially relevant bacteria provide a similar basis for genotyping of commercial strains, notably for lactic acid bacteria widely used as starter cultures in the dairy industry (Horvath et al. 2008, 2009; Barrangou and Horvath 2012). Even within a clonal population, active CRISPR loci are hypervariable and adaptive enough to track a strain over time, as shown in Leptospirillum isolated from acid mine drainage samples (Andersson and Banfield 2008; Tyson and Banfield 2008). Other metagenomics studies have shown that CRISPR loci can provide critical insights into population diversity and dynamics (Heidelberg et al. 2009; Held and Whitaker 2009; Anderson et al. 2011; Berg et al. 2012; Delaney et al. 2012; Garcia-Heredia et al. 2012; Pride et al. 2011, 2012; Rho et al. 2012; Stern et al. 2012). CRISPR spacer sequences may also be exploited to detect viral sequences or fish out viruses from complex, undefined ecosystems (Snyder et al. 2010). For metagenomic surveys, resolving CRISPR loci for mixed and occasionally complex microbial populations can unravel dynamics and ancestral relationships and occasionally reflect dramatic shifts and events such as selective bottlenecks. Nevertheless, it is important to keep in mind that such loci have variable typing potential across organisms given their broad range of (in-) activity and their highly variable distribution, occurrence, and propensity for horizontal gene transfer. Also, when multiple CRISPR loci are present within a chromosome, it is important to target a universal and polymorphic locus. Accordingly, their epidemiological potential has to be evaluated on a case-by-case basis, preferably using a broad and bio-geographically diverse set of strains and isolates.
11.7 Natural Genetic Tagging
In combination with increased phage resistance, CRISPR-Cas systems provide a tremendous avenue for the development of immortalized industrial workhorses which have highly desirable functional traits for the food supply chain, or that carry valuable biotechnological properties. In a competitive and global environment, although bacteria have been universally used as starter cultures in the food industry for centuries, it is increasingly critical to secure intellectual property and monitor the use of proprietary highly valuable strains.
A broadly used strategy is the deposit of characterized strains in strain banks, notably the culture collections that are official depositories under the Budapest Treaty (Budapest Treaty on the International Recognition of the Deposit of Microorganisms for the Purposes of Patent Procedure, signed on April 28, 1977). However, as strains evolve over time, and the origin and ownership of natural biological entities is difficult to define, it is important to secure intellectual property rights for the use of specific material for particular applications in specific fields. Accordingly, correctly and accurately defining a proprietary strain is critical, and CRISPR provides a unique natural means to generate mutants that have iteratively acquired a unique array of novel spacers in a human-defined order, directed manner, and selected way (see Table 11.1, patent application WO/2007/136815). Thus, iteratively selecting BIMs that have acquired novel CRISPR spacers following exposure to phage(s) (see Fig. 11.1) generates a natural (not genetically engineered) variant with a sequence tag (set of novel CRISPR spacers) which has an extremely remote probability to randomly arise in nature. This unique genetic watermark can subsequently be used to monitor the presence of a proprietary strain in any environment through simple and affordable Sanger sequencing of a CRISPR PCR amplicon.
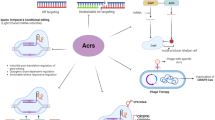
11.8 Cas Endonuclease Reprogramming and Restriction Enzyme Customization
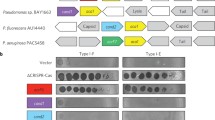
Two recent reports have shown that Cas-mediated DNA cleavage can be reprogrammed through crRNA design (Gasiunas et al. 2012; Jinek et al. 2012). Jinek et al. showed that both crRNA and tracrRNA direct DNA cleavage in S. pyogenes, and that a chimeric RNA can be engineered to redefine cleavage specificity. Gasiunas et al. showed that the S. thermophilus Cas9–crRNA ribonucleoprotein complex mediates specific DNA cleavage, and that the Cas9 HNH and RuvC domains nick the complementary and non-complementary DNA strands, respectively, ultimately generating a dsDNA cleavage. This is consistent with previous studies showing that Cas9 cleaves phage and plasmid dsDNA (Garneau et al. 2010; Sapranauskas et al. 2011; Magadán et al. 2012). The ability to nick either or both DNA strand(s) at (re-)programmable locations in a DNA sequence (see Fig. 11.3) opens new avenues for genome editing, stacking, shuffling, and engineering (Barrangou 2012). This essentially adds a new option to the genome engineering toolkit, in addition to zinc finger nucleases (ZFNs) and transcription activator-like effector nucleases (TALENs). Typically, genome engineering relies on site-specific endonucleases that trigger sequence modification by DNA-repair systems at the cleavage site. An advantage of Cas-crRNA-mediated cleavage is that specificity can be readily reprogrammed by customizing the crRNA sequence, rather than re-engineering cleavage proteins (ZFNs or TALENs) each time a new sequence has to be targeted (Gasiunas et al. 2012; Hale et al. 2012; Jinek et al. 2012).
Endonuclease customization. Cas9 endonuclease reprogramming. Any sequence containing at least one appropriate PAM can be cleaved specifically in its vicinity, at a precise location. In the example provided, the aim was to design a cleavage site within the E. coli 16S rDNA gene, between the variable regions V2 and V3, using the S. thermophilus CRISPR3-Cas system. Eight CRISPR3 PAM sequences (5′-NGNGG-3′, depicted as blue pentagons; Horvath et al. 2008) are found within this 183 bp region. The proper design and use of a chimeric crRNA targeting the proto-spacer (green rectangle), in combination with the Cas9 endonuclease, will lead to dsDNA cleavage within the proto-spacer, 3 nt upstream of the PAM (red arrows). Furthermore, the use of the wild-type Cas9, or RuvC- or HNH- mutants, will lead to a double-stranded, single (+)stranded, or single (−)stranded cleavage, respectively (Gasiunas et al. 2012)
11.9 Other Applications of CRISPR-Cas Systems
There are other ancillary and less documented roles and applications of CRISPR-Cas systems that remain to be substantiated and investigated, notably the potential that these loci have for the genesis of “large” amounts of small interfering RNAs (Djordjevic et al. 2012; see Table 11.1, patent application US20100076057), and the ability to generate and select “super phages” that circumvent CRISPR-encoded immunity for advanced biocontrol of microbial populations and phage therapy (see Table 11.1, patent application WO/2008/108989).
11.10 Conclusions and Perspectives
Overall, many intrinsic features of CRISPR-Cas systems provide avenues for applications that cover a broad spectrum, ranging from exploiting genetic hypervariability for typing and epidemiological purposes to increasing viral resistance, immunizing strains against the uptake of undesirable genetic material, through the generation of programmable RNA-guided endonucleases for genome engineering, editing, and stacking. Notwithstanding the tremendous potential of CRISPR loci in bacteria and archaea, it is critical to assess their potential for in vivo activity in eukaryotes to fully assess the potential of CRISPR-Cas systems for white biotechnology and next-generation synthetic biology.
As we reflect upon the past decade of CRISPR research, the impressive quality and quantity of manuscripts that have showcased their many powerful functionalities, in combination with the engaged and collegial CRISPR scientist community that has made the field so enjoyable and productive, it is obvious that the publication and citation rates of CRISPR manuscripts, together with the increased intellectual property activity, highlight the potential that these systems have for a diversity of applications.
Clearly, significant recent advances in phage resistance and strain typing have set the stage for extending the longevity of valuable industrial strains, and new epidemiological frameworks, respectively. For the latter, it is yet to be determined whether CRISPR loci can universally or broadly be used for typing of clinical isolates highly relevant for human health and disease. Nevertheless, we certainly hope the best is yet to come, and that many a talented and creative scientist will come up with innovative ways to harness the beauty and power of CRISPR-Cas systems for valuable and beneficial purposes.
References
Abadia E, Zhang J, dos Vultos T, Ritacco V, Kremer K, Aktas E, Matsumoto T, Refregier G, van Soolingen D, Gicquel B, Sola C (2010) Resolving lineage assignation on Mycobacterium tuberculosis clinical isolates classified by spoligotyping with a new high-throughput 3R SNPs based method. Infect Genet Evol 10:1066–1074
Aklujkar M, Lovley DR (2010) Interference with histidyl-tRNA synthetase by a CRISPR spacer sequence as a factor in the evolution of Pelobacter carbinolicus. BMC Evol Biol 10:230
Andersson AF, Banfield JF (2008) Virus population dynamics and acquired virus resistance in natural microbial communities. Science 320:1047–1050
Anderson RE, Brazelton WJ, Baross JA (2011) Using CRISPRs as a metagenomic tool to identify microbial hosts of a diffuse flow hydrothermal vent viral assemblage. FEMS Microbiol Ecol 77:120–133
Babu M, Beloglazova N, Flick R, Graham C, Skarina T, Nocek B, Gagarinova A, Pogoutse O, Brown G, Binkowski A, Phanse S, Joachimiak A, Koonin EV, Savchenko A, Emili A, Greenblatt J, Edwards AM, Yakunin AF (2011) A dual function of the CRISPR-Cas system in bacterial antivirus immunity and DNA repair. Mol Microbiol 79:484–502
Barrangou R (2012) RNA-mediated programmable DNA cleavage. Nat Biotechnol 30:836–838
Barrangou R, Horvath P (2012) CRISPR: new horizons in phage resistance and strain identification. Annu Rev Food Sci Technol 3:143–162
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P (2007) CRISPR provides acquired resistance against viruses in prokaryotes. Science 315:1709–1712
Berg Miller ME, Yeoman CJ, Chia N, Tringe SG, Angly FE, Edwards RA, Flint HJ, Lamed R, Bayer EA, White BA (2012) Phage-bacteria relationships and CRISPR elements revealed by a metagenomic survey of the rumen microbiome. Environ Microbiol 14:207–227
Bikard D, Hatoum-Aslan A, Mucida D, Marraffini LA (2012) CRISPR interference can prevent natural transformation and virulence acquisition during in vivo bacterial infection. Cell Host Microbe 12:177–186
Bolotin A, Quinquis B, Sorokin A, Ehrlich SD (2005) Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology 151:2551–2561
Borile C, Labarre M, Franz S, Sola C, Refrégier G (2011) Using affinity propagation for identifying subspecies among clonal organisms: lessons from M. tuberculosis. BMC Bioinf 12:224
Brouns SJ, Jore MM, Lundgren M, Westra ER, Slijkhuis RJ, Snijders AP, Dickman MJ, Makarova KS, Koonin EV, van der Oost J (2008) Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 321:960–964
Brudey K, Driscoll JR, Rigouts L et al (2006) Mycobacterium tuberculosis complex genetic diversity: mining the fourth international spoligotyping database (SpolDB4) for classification, population genetics and epidemiology. BMC Microbiol 6:23
Brüggemann H, Lomholt HB, Tettelin H, Kilian M (2012) CRISPR/cas loci of type II Propionibacterium acnes confer immunity against acquisition of mobile elements present in type I P. acnes. PLoS ONE 7:e34171
Cady KC, O’Toole GA (2011) Non-identity-mediated CRISPR-bacteriophage interaction mediated via the Csy and Cas3 proteins. J Bacteriol 193:3433–3445
Cady KC, White AS, Hammond JH, Abendroth MD, Karthikeyan RS, Lalitha P, Zegans ME, O’Toole GA (2011) Prevalence, conservation and functional analysis of Yersinia and Escherichia CRISPR regions in clinical Pseudomonas aeruginosa isolates. Microbiology 157:430–437
Cady KC, Bondy-Denomy J, Heussler GE, Davidson AR, O’Toole GA (2012) The CRISPR/Cas adaptive immune system of Pseudomonas aeruginosa mediates resistance to naturally occurring and engineered phages. J Bacteriol 194:5728–5738
Cui Y, Li Y, Gorgé O, Platonov ME, Yan Y, Guo Z, Pourcel C, Dentovskaya SV, Balakhonov SV, Wang X, Song Y, Anisimov AP, Vergnaud G, Yang R (2008) Insight into microevolution of Yersinia pestis by clustered regularly interspaced short palindromic repeats. PLoS ONE 3:e2652
Datsenko KA, Pougach K, Tikhonov A, Wanner BL, Severinov K, Semenova E (2012) Molecular memory of prior infections activates the CRISPR/Cas adaptive bacterial immunity system. Nat Commun 3:945
D’Auria G, Jiménez-Hernández N, Peris-Bondia F, Moya A, Latorre A (2010) Legionella pneumophila pangenome reveals strain-specific virulence factors. BMC Genomics 11:181
Delaney NF, Balenger S, Bonneaud C, Marx CJ, Hill GE, Ferguson-Noel N, Tsai P, Rodrigo A, Edwards SV (2012) Ultrafast evolution and loss of CRISPRs following a host shift in a novel wildlife pathogen, Mycoplasma gallisepticum. PLoS Genet 8:e1002511
Delannoy S, Beutin L, Burgos Y, Fach P (2012) Specific detection of enteroaggregative hemorrhagic Escherichia coli O104:H4 strains using the CRISPR locus as target for a diagnostic real-time PCR. J Clin Microbiol 50:3485−3492
Deveau H, Barrangou R, Garneau JE, Labonté J, Fremaux C, Boyaval P, Romero DA, Horvath P, Moineau S (2008) Phage response to CRISPR-encoded resistance in Streptococcus thermophilus. J Bacteriol 190:1390–1400
Díez-Villaseñor C, Almendros C, García-Martínez J, Mojica FJ (2010) Diversity of CRISPR loci in Escherichia coli. Microbiology 156:1351–1361
Djordjevic M, Djordjevic M, Severinov K (2012) CRISPR transcript processing: a mechanism for generating a large number of small interfering RNAs. Biol Direct 7:24
Erdmann S, Garrett RA (2012) Selective and hyperactive uptake of foreign DNA by adaptive immune systems of an archaeon via two distinct mechanisms. Mol Microbiol 85:1044−1056
Fabre L, Zhang J, Guigon G, Le Hello S, Guibert V, Accou-Demartin M, de Romans S, Lim C, Roux C, Passet V, Diancourt L, Guibourdenche M, Issenhuth-Jeanjean S, Achtman M, Brisse S, Sola C, Weill FX (2012) CRISPR typing and subtyping for improved laboratory surveillance of Salmonella infections. PLoS ONE 7:e36995
Fricke WF, Mammel MK, McDermott PF, Tartera C, White DG, Leclerc JE, Ravel J, Cebula TA (2011) Comparative genomics of 28 Salmonella enterica isolates: evidence for CRISPR-mediated adaptive sublineage evolution. J Bacteriol 193:3556–3568
Garcia-Heredia I, Martin-Cuadrado AB, Mojica FJ, Santos F, Mira A, Antón J, Rodriguez-Valera F (2012) Reconstructing viral genomes from the environment using fosmid clones: the case of haloviruses. PLoS ONE 7:e33802
Garneau JE, Dupuis MÈ, Villion M, Romero DA, Barrangou R, Boyaval P, Fremaux C, Horvath P, Magadán AH, Moineau S (2010) The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468:67–71
Garrett RA, Shah SA, Vestergaard G, Deng L, Gudbergsdottir S, Kenchappa CS, Erdmann S, She Q (2011) CRISPR-based immune systems of the Sulfolobales: complexity and diversity. Biochem Soc Trans 39:51–57
Gasiunas G, Barrangou R, Horvath P, Siksnys V (2012) Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc Natl Acad Sci U S A 109:E2579−E2586
Ginevra C, Jacotin N, Diancourt L, Guigon G, Arquilliere R, Meugnier H, Descours G, Vandenesch F, Etienne J, Lina G, Caro V, Jarraud S (2012) Legionella pneumophila sequence type 1/Paris pulsotype subtyping by spoligotyping. J Clin Microbiol 50:696–701
Godde JS, Bickerton A (2006) The repetitive DNA elements called CRISPRs and their associated genes: evidence of horizontal transfer among prokaryotes. J Mol Evol 62:718–729
Groenen PM, Bunschoten AE, van Soolingen D, van Embden JD (1993) Nature of DNA polymorphism in the direct repeat cluster of Mycobacterium tuberculosis; application for strain differentiation by a novel typing method. Mol Microbiol 10:1057–1065
Hale CR, Zhao P, Olson S, Duff MO, Graveley BR, Wells L, Terns RM, Terns MP (2009) RNA-guided RNA cleavage by a CRISPR RNA-Cas protein complex. Cell 139:945–956
Hale CR, Majumdar S, Elmore J, Pfister N, Compton M, Olson S, Resch AM, Glover CV 3rd, Graveley BR, Terns RM, Terns MP (2012) Essential features and rational design of CRISPR RNAs that function with the Cas RAMP module complex to cleave RNAs. Mol Cell 45:292–302
Heidelberg JF, Nelson WC, Schoenfeld T, Bhaya D (2009) Germ warfare in a microbial mat community: CRISPRs provide insights into the co-evolution of host and viral genomes. PLoS ONE 4:e4169
Held NL, Whitaker RJ (2009) Viral biogeography revealed by signatures in Sulfolobus islandicus genomes. Environ Microbiol 11:457–466
Hoe N, Nakashima K, Grigsby D, Pan X, Dou SJ, Naidich S, Garcia M, Kahn E, Bergmire-Sweat D, Musser JM (1999) Rapid molecular genetic subtyping of serotype M1 group A Streptococcus strains. Emerg Infect Dis 5:254–263
Horvath P, Barrangou R (2010) CRISPR/Cas, the immune system of bacteria and archaea. Science 327:167–170
Horvath P, Romero DA, Coûté-Monvoisin AC, Richards M, Deveau H, Moineau S, Boyaval P, Fremaux C, Barrangou R (2008) Diversity, activity, and evolution of CRISPR loci in Streptococcus thermophilus. J Bacteriol 190:1401–1412
Horvath P, Coûté-Monvoisin AC, Romero DA, Boyaval P, Fremaux C, Barrangou R (2009) Comparative analysis of CRISPR loci in lactic acid bacteria genomes. Int J Food Microbiol 131:62–70
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337:816−821
Labrie SJ, Samson JE, Moineau S (2010) Bacteriophage resistance mechanisms. Nat Rev Microbiol 8:317–327
Liu F, Barrangou R, Gerner-Smidt P, Ribot EM, Knabel SJ, Dudley EG (2011a) Novel virulence gene and clustered regularly interspaced short palindromic repeat (CRISPR) multilocus sequence typing scheme for subtyping of the major serovars of Salmonella enterica subsp. enterica. Appl Environ Microbiol 77:1946–1956
Liu F, Kariyawasam S, Jayarao BM, Barrangou R, Gerner-Smidt P, Ribot EM, Knabel SJ, Dudley EG (2011b) Subtyping Salmonella enterica serovar enteritidis isolates from different sources by using sequence typing based on virulence genes and clustered regularly interspaced short palindromic repeats (CRISPRs). Appl Environ Microbiol 77:4520–4526
Lopez-Sanchez MJ, Sauvage E, Da Cunha V, Clermont D, Ratsima Hariniaina E, Gonzalez-Zorn B, Poyart C, Rosinski-Chupin I, Glaser P (2012) The highly dynamic CRISPR1 system of Streptococcus agalactiae controls the diversity of its mobilome. Mol Microbiol 85:1057–1071
Louwen R, Horst-Kreft D, de Boer AG, van der Graaf L, de Knegt G, Hamersma M, Heikema AP, Timms AR, Jacobs BC, Wagenaar JA, Endtz HP, van der Oost J, Wells JM, Nieuwenhuis EE, van Vliet AH, Willemsen PT, van Baarlen P, van Belkum A (2012) A novel link between Campylobacter jejuni bacteriophage defence, virulence and Guillain-Barré syndrome. Eur J Clin Microbiol Infect Dis. doi:10.1007/s10096-012-1733-4
Magadán AH, Dupuis MÈ, Villion M, Moineau S (2012) Cleavage of Phage DNA by the Streptococcus thermophilus CRISPR3-Cas System. PLoS ONE 7:e40913
Marraffini LA, Sontheimer EJ (2008) CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science 322:1843–1845
McGhee GC, Sundin GW (2012) Erwinia amylovora CRISPR elements provide new tools for evaluating strain diversity and for microbial source tracking. PLoS ONE 7:e41706
McShan WM, Ferretti JJ, Karasawa T, Suvorov AN, Lin S, Qin B, Jia H, Kenton S, Najar F, Wu H, Scott J, Roe BA, Savic DJ (2008) Genome sequence of a nephritogenic and highly transformable M49 strain of Streptococcus pyogenes. J Bacteriol 190:7773–7785
Mills S, Griffin C, Coffey A, Meijer WC, Hafkamp B, Ross RP (2010) CRISPR analysis of bacteriophage-insensitive mutants (BIMs) of industrial Streptococcus thermophilus—implications for starter design. J Appl Microbiol 108:945–955
Mojica FJ, Díez-Villaseñor C, García-Martínez J, Soria E (2005) Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J Mol Evol 60:174–182
Mojica FJ, Díez-Villaseñor C, García-Martínez J, Almendros C (2009) Short motif sequences determine the targets of the prokaryotic CRISPR defence system. Microbiology 155:733–740
Mokrousov I, Limeschenko E, Vyazovaya A, Narvskaya O (2007) Corynebacterium diphtheriae spoligotyping based on combined use of two CRISPR loci. Biotechnol J 2:901–906
Mokrousov I, Vyazovaya A, Kolodkina V, Limeschenko E, Titov L, Narvskaya O (2009) Novel macroarray-based method of Corynebacterium diphtheriae genotyping: evaluation in a field study in Belarus. Eur J Clin Microbiol Infect Dis 28:701–703
Nozawa T, Furukawa N, Aikawa C, Watanabe T, Haobam B, Kurokawa K, Mauyama F, Nakagawa I (2011) CRISPR inhibition of prophage acquisition in Streptococcus pyogenes. PLoS ONE 6:e19543
Palmer KL, Gilmore MS (2010) Multidrug-resistant enterococci lack CRISPR-cas. MBio 1:e00227–e00310
Palmer KL, Whiteley M (2011) DMS3-42: the secret to CRISPR-dependent biofilm inhibition in Pseudomonas aeruginosa. J Bacteriol 193:3431–3432
Pougach K, Semenova E, Bogdanova E, Datsenko KA, Djordjevic M, Wanner BL, Severinov K (2010) Transcription, processing and function of CRISPR cassettes in Escherichia coli. Mol Microbiol 77:1367–1379
Pourcel C, Salvignol G, Vergnaud G (2005) CRISPR elements in Yersinia pestis acquire new repeats by preferential uptake of bacteriophage DNA, and provide additional tools for evolutionary studies. Microbiology 151:653–663
Price EP, Smith H, Huygens F, Giffard PM (2007) High-resolution DNA melt curve analysis of the clustered, regularly interspaced short-palindromic-repeat locus of Campylobacter jejuni. Appl Environ Microbiol 73:3431–3436
Pride DT, Sun CL, Salzman J, Rao N, Loomer P, Armitage GC, Banfield JF, Relman DA (2011) Analysis of streptococcal CRISPRs from human saliva reveals substantial sequence diversity within and between subjects over time. Genome Res 21:126–136
Pride DT, Salzman J, Relman DA (2012) Comparisons of clustered regularly interspaced short palindromic repeats and viromes in human saliva reveal bacterial adaptations to salivary viruses. Environ Microbiol 14:2564−2576
Rezzonico F, Smits TH, Duffy B (2011) Diversity, evolution, and functionality of clustered regularly interspaced short palindromic repeat (CRISPR) regions in the fire blight pathogen Erwinia amylovora. Appl Environ Microbiol 77:3819–3829
Rho M, Wu YW, Tang H, Doak TG, Ye Y (2012) Diverse CRISPRs evolving in human microbiomes. PLoS Genet 8:e1002441
Riehm JM, Vergnaud G, Kiefer D, Damdindorj T, Dashdavaa O, Khurelsukh T, Zöller L, Wölfel R, Le Flèche P, Scholz HC (2012) Yersinia pestis lineages in Mongolia. PLoS ONE 7:e30624
Sapranauskas R, Gasiunas G, Fremaux C, Barrangou R, Horvath P, Siksnys V (2011) The Streptococcus thermophilus CRISPR/Cas system provides immunity in Escherichia coli. Nucleic Acids Res 39:9275–9282
Schouls LM, Reulen S, Duim B, Wagenaar JA, Willems RJ, Dingle KE, Colles FM, Van Embden JD (2003) Comparative genotyping of Campylobacter jejuni by amplified fragment length polymorphism, multilocus sequence typing, and short repeat sequencing: strain diversity, host range, and recombination. J Clin Microbiol 41:15–26
Snyder JC, Bateson MM, Lavin M, Young MJ (2010) Use of cellular CRISPR (clusters of regularly interspaced short palindromic repeats) spacer-based microarrays for detection of viruses in environmental samples. Appl Environ Microbiol 76:7251–7258
Stern A, Keren L, Wurtzel O, Amitai G, Sorek R (2010) Self-targeting by CRISPR: gene regulation or autoimmunity? Trends Genet 26:335–340
Stern A, Mick E, Tirosh I, Sagy O, Sorek R (2012) CRISPR targeting reveals a reservoir of common phages associated with the human gut microbiome. Genome Res 22:1985–1994
Sturino JM, Klaenhammer TR (2006) Engineered bacteriophage-defence systems in bioprocessing. Nat Rev Microbiol 4:395–404
Swarts DC, Mosterd C, van Passel MW, Brouns SJ (2012) CRISPR interference directs strand specific spacer acquisition. PLoS ONE 7:e35888
Tasaki E, Hirayama J, Tazumi A, Hayashi K, Hara Y, Ueno H, Moore JE, Millar BC, Matsuda M (2012) Molecular identification and characterization of clustered regularly interspaced short palindromic repeats (CRISPRs) in a urease-positive thermophilic Campylobacter sp. (UPTC). World J Microbiol Biotechnol 28:713–720
Tyson GW, Banfield JF (2008) Rapidly evolving CRISPRs implicated in acquired resistance of microorganisms to viruses. Environ Microbiol 10:200–207
van der Ploeg JR (2009) Analysis of CRISPR in Streptococcus mutans suggests frequent occurrence of acquired immunity against infection by M102-like bacteriophages. Microbiology 155:1966–1976
Vergnaud G, Li Y, Gorgé O, Cui Y, Song Y, Zhou D, Grissa I, Dentovskaya SV, Platonov ME, Rakin A, Balakhonov SV, Neubauer H, Pourcel C, Anisimov AP, Yang R (2007) Analysis of the three Yersinia pestis CRISPR loci provides new tools for phylogenetic studies and possibly for the investigation of ancient DNA. Adv Exp Med Biol 603:327–338
Yosef I, Goren MG, Qimron U (2012) Proteins and DNA elements essential for the CRISPR adaptation process in Escherichia coli. Nucleic Acids Res 40:5569–5576
Zegans ME, Wagner JC, Cady KC, Murphy DM, Hammond JH, O’Toole GA (2009) Interaction between bacteriophage DMS3 and host CRISPR region inhibits group behaviors of Pseudomonas aeruginosa. J Bacteriol 191:210–219
Zhang J, Abadia E, Refregier G, Tafaj S, Boschiroli ML, Guillard B, Andremont A, Ruimy R, Sola C (2010) Mycobacterium tuberculosis complex CRISPR genotyping: improving efficiency, throughput and discriminative power of ‘spoligotyping’ with new spacers and a microbead-based hybridization assay. J Med Microbiol 59:285–294
Acknowledgments
We are indebted to our DuPont colleagues, notably Patrick Boyaval, Christophe Fremaux, and Dennis Romero, and our many academic collaborators with whom we have had the privilege to share exciting times exploring CRISPR-cas systems, notably the Moineau laboratory at Laval, the Banfield laboratory at the University of California Berkeley, the Roberts laboratory at the Pennsylvania State University, the Dudley laboratory at the Pennsylvania State University, the Terns laboratory at the University of Georgia, the Levin laboratory at Emory, the Bhaya laboratory at Stanford University, the VerBerkmoes laboratory at ORNL, Eugene Koonin and Kira Makarova at NCBI, Eric Brown and Marc Allard at FDA, and our dedicated and talented teams. Work in Vilnius was funded by the European Social Fund under Global Grant measure Project R100.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Horvath, P., Gasiunas, G., Siksnys, V., Barrangou, R. (2013). Applications of the Versatile CRISPR-Cas Systems. In: Barrangou, R., van der Oost, J. (eds) CRISPR-Cas Systems. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-34657-6_11
Download citation
DOI: https://doi.org/10.1007/978-3-642-34657-6_11
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-34656-9
Online ISBN: 978-3-642-34657-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)