Abstract
The circadian clock is an endogenous oscillator with a 24-h period. Although delayed feedback repression was proposed to lie at the core of the clock more than 20 years ago, the mechanism for making delay in feedback repression in clock function has only been demonstrated recently. In the mammalian circadian clock, delayed feedback repression is mediated through E/E′-box, D-box, and RRE transcriptional cis-elements, which activate or repress each other through downstream transcriptional activators/repressors. Among these three types of cis-elements, transcriptional negative feedback mediated by E/E′-box plays a critical role for circadian rhythms. A recent study showed that a combination of D-box and RRE elements results in the delayed expression of Cry1, a potent transcriptional inhibitor of the E/E′-box. The overall interconnection of these cis-elements can be summarized as a combination of two oscillatory motifs: one is a simple delayed feedback repression where only an RRE represses an E/E′-box, and the other is a repressilator where each element inhibits another in turn (i.e., E/E′ box represses an RRE, an RRE represses a D-box, and a D-box represses an E/E′ box). Experimental verification of the roles of each motif as well as post-transcriptional regulation of the circadian oscillator will be the next challenges.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Circadian Clock in Mammals
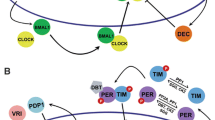
In mammals the master clock is located in the suprachiasmatic nucleus (SCN). Transcript analyses have indicated that circadian clocks are not restricted to SCN, but are found in several tissues including the liver (Yamazaki et al. 2000) and cultured cells such as rat fibroblasts Rat-1 (Balsalobre et al. 1998), mouse fibroblasts NIH3T3 (Tsuchiya et al. 2003), or human osteosarcoma U-2OS cells (Isojima et al. 2009; Vollmers et al. 2008). Therefore, circadian rhythms are driven by cell-autonomous oscillators. Studies across species have elucidated the conserved feature of molecular mechanisms underlying circadian rhythms: at the core of the clock lies a transcriptional/translational negative feedback loop. For example, in mice the transcription factors CLOCK and BMAL1 dimerize and activate transcription of the Per and Cry genes. PER and CRY proteins accumulate in the cytosol become phosphorylated and return to the nucleus where they inhibit the activity of CLOCK and BMAL1. The turnover of PER and CRY proteins leads to a new cycle of activation by CLOCK and BMAL1 via E/E′-box (Dunlap 1999; Griffin et al. 1999; Kume et al. 1999; Reppert and Weaver 2002; Young and Kay 2001). In this process, PER and CRY form a negative feedback loop that inhibits their own transcription. However, reciprocal activation of positive (CLOCK and BMAL1) and negative (PER and CRY) regulators in a negative feedback loop is not sufficient: there must be a delay or immediate self-inhibition of CRY and PER would result in the stable lower expression of these factors rather than oscillation. What molecular mechanism imposes this time delay? This chapter summarizes the transcription network of the mammalian circadian clock and provides insights into how the network together with post-translational regulation of clock proteins works as a delayed negative feedback loop.
2 Identification of the Circadian Transcriptional Network
2.1 Transcriptional Network Based on Three Clock-Controlled Elements
2.1.1 The E/E′-Box, the D-Box, and the RRE
The overall topology of mammalian circadian transcription network can be understood by the combination of three clock-controlled elements (CCEs), short consensus DNA sequences typically located near the promoter region of clock genes. These CCEs are called the E/E′-box (CACGT(T/G)) (Gekakis et al. 1998; Hogenesch et al. 1997; Ueda et al. 2005; Yoo et al. 2005), the D-box (DBP response element) (TTATG(C/T)AA) (Falvey et al. 1996; Ueda et al. 2005), and the RRE [RevErbA response element, also called as ROR response element (RORE)] [(A/T)A(A/T)NT(A/G)GGTCA] (Harding and Lazar 1993; Preitner et al. 2002; Ueda et al. 2002, 2005).
By performing transcriptome analysis, expression of 24-h periodic genes was reported in cultured cells (Grundschober et al. 2001), the SCN (Panda et al. 2002; Ueda et al. 2002), and other tissues such as heart (Storch et al. 2002), liver (Panda et al. 2002; Storch et al. 2002; Ueda et al. 2002), aorta (Rudic et al. 2005), adipose tissues (Zvonic et al. 2006), calvarial bone (Zvonic et al. 2007), and hair follicle (Akashi et al. 2010). Although there are differences in the rhythmicity of circadian-expressed genes in each tissue, the following mammalian clock genes most commonly have circadian oscillation: Period1 (Per1), Per2, Per3, Dec1 (Bhlhb2), Dec2 (Bhlhb3), Cryptochome1 (Cry1), Clock, Npas2, Bmal1 (Arntl), Dbp, E4bp4 (Nfil3), RevErbAa (Nr1d1), RevErbAb (Nr1d2), and Rora. The temporal expression of each gene is controlled by a different combination of CCEs. Evolutionary conserved E/E′-boxes are located in the noncoding regions of nine genes (Per1, Per2, Cry1, Dbp, Rorγ, RevErbAa, RevErbAb, Dec1, and Dec2), D-boxes are contained in eight genes (Per1, Per2, Per3, Cry1, RevErbAa, RevErbAb, Rorα, and Rorβ), and RREs in six genes (Bmal1, Clock, Npas2, Cry1, E4bp4, and Rorc). The expressed gene product positively or negatively regulates transcription activity by acting on CCEs: CCEs and these clock genes form a closed network structure (Fig. 1) as described below.
Schematic representation of the transcriptional network of the mammalian circadian clock. (a) In vitro cycling assay. Cultured mammalian cells (Rat-1) were transfected with dLuc under the control of a clock-controlled element (CCE) and SV40 basic promoter (Ueda et al. 2005). (b) Representative circadian rhythms of bioluminescence from a wild-type Per1 E-box CCE fused to the SV40 basic promoter driving a dLuc reporter (left panel) and compared to bioluminescence rhythms driven by a Per1 D-box (center panel) and a RRE (right panel). Original figures are reproduced from Ueda et al. (2005). (c) Genes and CCEs are depicted as ellipsoids and rectangles, respectively. Transcriptional/translational activation is shown by arrows (→) and repression is depicted by arrows with flat ends (┤)
2.1.2 Transcription Regulation via the E/E′-Box and Clock Genes
The E/E′-box is positively regulated by Bmal1, Clock, and Npas2 and negatively regulated by Per1–3, Cry1–2, and Dec1–2. CRY and PER are hypothesized to autoregulate their own expression by repressing the heterodimeric complex of the basic helix–loop–helix (bHLH) PER-ARNT-SIM (PAS) domain transcriptional activators CLOCK and BMAL1, which bind to E/E′-box elements in the Cry1 and Per1–2 promoters. Although both positive regulators (Bmal1, Npas2, Clock) and negative regulators (Per1–3 and Cry1–2) have circadian rhythmic expression patterns, peak time of positive regulators are antiphase to that of negative regulators (delayed negative feedback).
2.1.3 Transcription Regulation via the D-Box and Clock Genes
The D-box is positively regulated by PAR-bZIP (proline- and acidic amino acid-rich basic leucine zipper) transcription factors (Dbp, Tef, and Hlf) and negatively by E4bp4. Like the E/E′-box, an antiphase relationship of gene expression between negative and positive regulators can be observed. In the D-box case, the expression phase of the positive regulator Dbp is similar to that of Per1, whereas the expression phase of the negative regulator E4bp4 is similar to that of Bmal1 (Mitsui et al. 2001).
2.1.4 Transcription Regulation via the RRE and Clock Genes
RRE is positively regulated by Rora, Rorb, and Rorc and negatively regulated by RevErbAa and RevErbAb. In the SCN, Rora and Rorb have circadian rhythms but not Rorc (Ueda et al. 2002). Liu et al. reported that RevErbAa and RevErbAb are functionally redundant and necessary for oscillation of the RRE-regulated gene Bmal1. By contrast, Rors contribute to Bmal1 amplitude, but are not required for generating oscillation (Liu et al. 2008).
2.1.5 Timing of Each CCE
The circadian timing at which each element becomes active for transcription can be monitored with an in vitro cell culture system in which a destabilized firefly luciferase (dLuc) reporter is driven by different clock-controlled promoters. After cells are synchronized (i.e., with dexamethasone, forskolin, or serum), oscillations in gene expression are recorded by bioluminescence (Nagoshi et al. 2004; Ueda et al. 2002, 2005; Welsh et al. 2004). Using this in vitro cycling assay, the “phase” of each CCE can be determined (Ueda et al. 2002, 2005) (Fig. 1). Note that the term “phase” used in this chapter represents relative peak timing of each circadian gene expression within single circadian cycle. Each CCE is responsible for the gene expression at distinct circadian times: the peak time of E/E′-box-driven expression is followed by D-box-driven expression after an interval of ~5 h. Then, RRE-driven expression follows D-box expression after ~8 h. E/E′-box-driven expression begins to appear again ~11 h after RRE-driven expression. In the case of the SCN, the subjective time drawn by each CCE can be illustrated as “morning-time” for the E/E′-box, “evening-time” for the D-box, and “nighttime” for the RRE (Ueda et al. 2005).
2.2 Importance of Gene Regulation via the E/E′-Box
2.2.1 Circadian Clock Perturbation via CCEs
The three CCEs have different impacts on cellular circadian rhythms: perturbation of E/E′-box regulation abolishes circadian rhythms; perturbation of RRE regulation has an intermediate but significant effect; and D-box disruption has almost no effect.
A study using Rat-1 cell showed this by overexpressing regulatory genes with repressive activity to different CCEs (Ueda et al. 2005) (Fig. 2). The Per2 promoter is regulated via an E/E′-box and a D-box, and the Bmal1 promoter is regulated via an RRE. When E/E′-box activity is perturbed by overexpression of the Cry1 gene, both Per2-promoter-driven reporter gene (Per2-dLuc) and Bmal1-promoter-driven reporter gene (Bmal1-dLuc) lose circadian rhythms. When an RRE is perturbed through RevErbAa overexpression, Bmal1-dLuc loses circadian rhythms and the amplitude of Per2-dLuc rhythmic expression is decreased. The impact of RRE perturbation through RevErbAa overexpression appeared to be more significant in mice liver. Kornmann et al. showed that liver-specific overexpression of RevErbAa abolishes the rhythmicity of PER2::Luc expression in liver explants (Kornmann et al. 2007). Contrary to the case of E/E′-box and RRE, D-box perturbation through E4bp4 overexpression causes both Per2-dLuc and Bmal1-dLuc transcriptional activity to have normal circadian rhythms (Ueda et al. 2005). These varying effects are difficult to explain by mere quantitative differences in the strength of the three repressors, which suggests that there is some qualitative difference between E/E′-box, D-box, and RRE regulation in circadian rhythmicity.
Importance of the E/E′-box. Effect of repression on each CCE. The E/E′-boxes, D-box, and RRE were repressed by overproduction of CRY1, E4BP4, and REVERBAa, respectively. The consequences of those repressions were monitored by bioluminescence from Per2 and Bmal1 promoter driving a destabilized luciferase (Per2-dLuc and Bmail1-dLuc). Original figures are reproduced from Ueda et al. (2005). The different shades of gray in the plot indicate different amounts of transfected vector
2.2.2 Circadian Feedback Repression: Heart of the Circadian Transcriptional Network
PER and CRY play key roles in the circadian clock transcriptional network by closing the negative feedback loop of E/E′-box regulation. CRY has stronger repressor activity than PER (Kume et al. 1999). Sato et al. reported that interference of CRY1’s repressor activity on E/E′-box-mediated transcription can abolish circadian transcriptional oscillations. They screened both human CLOCK and BMAL1 alleles that were insensitive to CRY1 repression but maintained normal transcriptional activity. Selected clones have normal transcriptional activities similar to wild type in the absence of CRY1, but have greater reporter activity in the presence of CRY1. By analyzing either Per2-dLuc or Bmal1-dLuc, they observed that cotransfection of either CLOCK or BMAL1 mutant alleles resulted in substantial impairment of circadian rhythmicity after one or two cycles of oscillation; cotransfection of both CRY-insensitive mutant CLOCK and BMAL1 together resulted in the loss of circadian promoter activity. This suggests that transcriptional repression of CLOCK/BMAL1 by CRY1 is required for circadian regulation via both an E/E′-box and an RRE (Sato et al. 2006) (Fig. 3).
The impairment of CRY-mediated repression. Coexpression of CLOCK/BMAL1 mutant heterodimers that are insensitive to CRY repression ablates circadian E-box and RRE activities in NIH3T3 cells. Plasmids expressing Flag-tagged CLOCK and BMAL1 alleles were transiently cotransfected with the Per2-dLuc (upper panel) or Bmal1-dLuc reporter plasmid into NIH3T3 cells (lower panel). Per2 or Bmal1 promoter activities in NIH3T3 cells transfected with single or double CRY1-insensitive CLOCK, and BMAL1 mutants (MT) were monitored over 5 (upper panel) or 6 days (lower panel). All reporter activities were normalized such that the median wild-type luciferase activity over the time course was 100 %. Original figures are reproduced from Sato et al. (2006)
3 Minimal Circuit of the Mammalian Circadian Clock
3.1 Two Delayed Negative Feedback Loops
How is the negative feedback to an E/E′-box delayed? Although there is an E′-box and an E-box in Cry1’s regulatory region (Fustin et al. 2009; Ueda et al. 2005), the peak of Cry1 expression is evening-time, which is substantially delayed relative to other genes with an E/E′-box (Fustin et al. 2009; Ueda et al. 2005). Cry1 has two functional RREs in one of its introns (Ueda et al. 2005) and also D-box in its promoter region (Ukai-Tadenuma et al. 2011). Ukai-Tadenuma et al. experimentally confirmed that the combination of daytime elements (D-box) and nighttime elements (RREs) within its intronic enhancer gives rise to Cry1’s delayed evening-time expression. Interestingly, the observed delayed expression was well explained by a simple phase-vector model that enabled artificially designed delayed expressions (Ukai-Tadenuma et al. 2011) (Fig. 4).
Phase-vector model. A new phase results from the combinatorial synthesis of two transcriptional regulators or two clock-controlled DNA elements, which can be illustrated to a first-order approximation by a phase-vector model. This combinatorial regulatory mechanism for generating new circadian phases of transcription represents a general design principle underpinning complex system behavior. Assume wave function f x (t) = A x cos(θ(t) + φ x ). The amplitude of wave A is represented by the length of a phase vector P, and the phase of wave φ is represented by the angle of P. The component waves f 1 and f 2 are displayed by phase vectors P 1 and P 2. P c is the summed phase vector of P 1 and P 2. Original graph is reproduced from Ukai-Tadenuma et al. (2011)
Based on this simple phase-vector model (Fig. 4), they generated an array of Cry1 constructs that have different phases and used these in a genetic complementation assay to restore circadian oscillation in arrhythmic Cry1 −/−:Cry2 −/− cells established from Cry1 −/−:Cry2 −/− double-knockout mice (van der Horst et al. 1999). These experiments reveal that substantial delay of Cry1 expression is required to restore single-cell-level rhythmicity and that prolonged delay of Cry1 expression can slow circadian oscillations (Fig. 5). These results suggest that phase delay in Cry1 transcription is required for mammalian clock function and these results provide formal proof that the design principle of the mammalian circadian clock transcriptional network is negative feedback with delay (Ukai-Tadenuma et al. 2011).
The biological relevance of delayed Cry1 expression in circadian clock function. (a) Per2-dLuc bioluminescence levels in transfected Cry1 −/−:Cry2 −/− cells. The Per2-dLuc reporter and a Cry1 expression construct were cotransfected into Cry1 −/−:Cry2 −/− cells. (b) Cry1 expression phases under different promoters (Ueda et al. 2005; Ukai-Tadenuma et al. 2011) that contain either a D-box (Cry1 promoter), RRE (Cry1 intron), or both (promoter + intron). (c) Substantial delay in feedback repression is required for mammalian clock function. The decreased delay dampens the amplitude of circadian oscillations (top panel), and the prolonged delay in feedback repression slows the frequency of circadian oscillations (bottom panel) compared to wild type (middle panel). Original figures are reproduced from Ukai-Tadenuma et al. (2011). Different trace shades represent results from triplicated experiments
Based on these results, they hypothesized that the transcriptional network can be simplified into a model consisting of two transcriptional activations and four transcriptional repressions on three regulatory DNA elements (Fig. 6). Notably, this diagram can be envisaged as a composite of two distinct oscillatory network motifs (1) a repressilator, which is composed of three repressions, and (2) a delayed negative feedback loop, which is composed of two activations and one repression. Both oscillatory network motifs include delayed feedback repression and can generate autonomous oscillations independently (Elowitz and Leibler 2000; Stricker et al. 2008).
The minimal circuit for the mammalian circadian transcriptional network. (a) The transcription network of the mammalian circadian clock (Ueda et al. 2005; Ukai-Tadenuma et al. 2011). (b) The minimal circuit (top panel) can be illustrated as a composite of two distinct oscillatory network motifs: a repressilator (bottom left panel) and a delayed negative feedback loop (bottom right panel). Transcriptional activation (arrows); transcriptional repression (arrows with flat ends); regulatory DNA elements (rectangles; E/E′-box, morning; D-box, daytime; RRE, nighttime). Original graph is reproduced from Ukai-Tadenuma et al. (2011)
3.2 Genetic Evidence for the Importance of CCEs
The minimal circuit model implies that all of three CCEs have substantial importance for the circadian oscillator. The importance of an E/E′-box-mediated regulation is manifested by the phenotypes of several clock gene-knockout mice. The circadian clock governs physiological phenomena like day–night variation of activity, so changes in behavioral rhythms reflect differences in the endogenous clock of mutant mice. Accordingly, disruption of Bmal1, a positive regulator of E/E′-box-mediated regulation, directly results in the loss behavioral rhythms in mice (Bunger et al. 2000; Shi et al. 2010). Disruption of Clock gene did not result in loss of behavioral rhythms (DeBruyne et al. 2007) probably because Clock and another gene Npas2 have redundant roles: Clock and Npas2 double-knockout mice have arrhythmic behavioral patterns (DeBruyne et al. 2007), while Npas2-disrupted mice have normal behavioral rhythms (Dudley et al. 2003). Loss of negative regulator of E/E′-box-mediated transcription also results in arrhythmic phenotypes. Both Per1 and Per2 disrupted mice have loss of circadian rhythmicity of behavioral activity (Bae et al. 2001; Zheng et al. 2001), and Cry1 −/−:Cry2 −/− mice have arrhythmic behavioral patterns (van der Horst et al. 1999; Vitaterna et al. 1999).
The minimal structure shown in Fig. 6 implies that D-box and RRE also play an essential role to maintain the delay time for negative feedback. For example, the double knockout of RevErbAa and RevErbAb mice has arrhythmic behavioral phenotypes and arrhythmic clock gene expression (Bugge et al. 2012; Cho et al. 2012).
The importance of D-box transcriptional regulators is still unclear because no report shows that dysfunctional mice for D-box regulators have completely arrhythmic behavioral patterns. Lopez-Molina et al. reported that Dbp knockout mice have normal behavioral rhythms compared to wild type (Lopez-Molina et al. 1997). Hlf or Tef disrupted mice also have almost normal behavioral rhythms (Gachon et al. 2004). Even triple knockout of PAR-bZIP transcriptional factor mice have almost normal behavior rhythms (Gachon et al. 2004). Although E4bp4 knockout mice was constructed (Gascoyne et al. 2009), behavioral rhythms of the mice were not reported.
3.3 Generation of Various Phases by the Combination of CCEs
From DNA microarray data, more than 10 % of expressed genes have circadian rhythms with a wide range of peak timings (Delaunay and Laudet 2002); the distribution of peak timing is not limited to three circadian times corresponding to the expression timing of each CCE. How do these “intermediate” expression timings arise? One possibility is that the combination of three CCEs generates various circadian phases.
Ukai-Tadenuma, Kasukawa et al. adopted a synthetic approach to physically simulate the correlation between CCEs combinations and the peak timing of expression. They used three components: an artificial activator (dGAL4-VP16), an artificial repressor (dGAL4), and a dGAL4-VP16-driven reporter gene (dLuc) as an output (Fig. 7a). If the expression of artificial activator and repressor are controlled by different CCEs, then the output may vary according to a combination of the various peak timings of each CCE. By taking the peak expression timing of clock gene expression in mouse liver, phase of each CCE-driven gene expression can be related with subjective circadian time: E/E′-box-driven expression peak timing as “morning,” RRE-driven expression peak timing as “night,” and D-box-driven peak timing as “daytime.” They created “daytime” expression by the combination of E/E′-box (morning)-driven activator and RRE (night)-driven repressor (Fig. 7b). This is similar to transcriptional regulation via D-box control; D-box is activated by E/E′-box-controlled Dbp and repressed by RRE-controlled E4bp4, and output phase is “daytime.” Next, they created “night” by the combination of a D-box-driven activator and an E/E′-box-driven repressor (Fig. 7c). This is similar to an RRE with output phase “night”: RRE is regulated by D-box-driven activator (Rora) and E/E′-box-driven repressor (RevErbAa), though RevErbAa is also controlled by D-box. By combining these CCEs in different arrangements, Ukai-Tadenuma, Kasukawa et al. also generated additional phases (Fig. 7d), which are not identical to any of the original CCE timings (Ukai-Tadenuma et al. 2008).
Combinatorial regulation of circadian phases by a synthetic system. (a) The artificial transcription system. Activator and repressor are driven under clock-controlled elements (CCEs). Details are described in main text. (b, c) Promoter activities of an activator, repressor, and output in different artificial transcriptional circuits. The schemes summarize the representative promoter activities of each artificial circuit monitored by bioluminescence from NIH3T3 cells, where an activator, repressor, and output phases are indicated with their peak time (gray numbers). Morning activator under E′-box control and nighttime repressor under RRE control and (b) daytime activator under D-box control and morning repressor under E′-box control (c). (d) The relationship of the expression timings of the transcription factors and output. Various expression timing is generated from three basic phases (morning, daytime, and nighttime). Black lines indicate activation (arrows) and gray lines repression (arrows with flat ends). Original figures and graphs are reproduced from Ukai-Tadenuma et al. (2008)
4 Post-Translational Regulation, Another Layer of Delay or Another Oscillator?
4.1 Phosphorylation of PER
As we discussed above, accumulating evidence indicates that Cry1-mediated delayed negative feedback plays a critical role in the circadian transcription network. If so, is the network structure of transcription activator/inhibitor relationship sufficient for generating mammalian circadian properties? If we replace all transcription factors with artificial ones [such as GAL4-VP16 used in Ukai-Tadenuma et al. (2008)] but keep the network structure, could we reproduce a robust circadian system? Natural circadian systems, however, seem to be more complex than the transcription-translation network; post-translational regulation is also critical for circadian function (Gallego and Virshup 2007). In particular, phosphorylation of PERs by CKIδ/ε is one of the determinants of circadian period length (Lowrey et al. 2000; Toh et al., 2001; Xu et al., 2005). The first circadian mutant identified in mammal was the tau-mutant hamster, which has a shorter behavioral period length compared to a normal hamster (Ralph and Menaker 1988). Takahashi’s group identified the tau mutation in the CKIe gene and found that PER phosphorylation is lower in tau-mutant hamsters (Lowrey et al. 2000). The importance of PER phosphorylation by CKIδ/ε for circadian rhythms is also true in humans. Toh et al. discovered that familial advanced sleep-phase syndrome (FASPS) is caused by a mutation in the CKIδ/ε binding site of PER2 (Toh et al. 2001). Likewise, Xu et al. found that a mutation in CKIδ can also cause FASPS by modulating PER stability (Xu et al. 2005). Additionally, chemical biology approaches identified several compounds that shorten or lengthen circadian period (Chen et al. 2012; Hirota et al. 2008; Isojima et al. 2009). One remarkable example is a series of CKIδ/ε inhibitors, which can lengthen molecular clock period from 24 h to 48 h at the cellular level (Isojima et al. 2009).
How PER phosphorylation controls circadian period is still mysterious, but phosphorylation affects PER stability. PER protein is degraded by proteasome-mediated proteolysis when phosphorylation of PER triggers recruitment of βTrCP, a subunit of the SCF ubiquitin ligase (Eide et al. 2005; Shirogane et al. 2005). However, the FASPS mutation site is different from the region involved in βTrCP recognition of PER (Eide et al. 2005). Furthermore, several results imply that phosphorylation on FASPS-mutated site stabilizes PER protein (Shanware et al. 2011; Vanselow et al. 2006; Xu et al. 2007). Therefore, phosphorylation may regulate the stability of PER in multiple ways. Recent studies of Drosophila melanogaster PER and Neurospora crassa FRQ (a functional counterpart of PER) show that multisite phosphorylation induces conformational changes in these proteins (Chiu et al. 2011; Querfurth et al. 2011). A similar case might also be true for mammalian PER: phosphorylation may control the stability of mammalian PER by changing its global structure, not just by creating a recognition site for βTrCP at a specific location.
The stability control of PER also may contribute to delay for transcriptional negative feedback. Unlike other clock genes, the expression peak of Per1 and Per2 mRNA is ~4 h earlier than PER1/PER2 proteins (Pace-Schott and Hobson 2002). This delay between mRNA and protein may be one of the determinants of period length.
4.2 Stability Control of CRY in Circadian Oscillations
Recently, researchers noticed that not only PER but also CRY stability is important for clock period. In 2007, two lines of ENU-mutant mice with long behavioral rhythms were reported from different groups—Overtime (Siepka et al. 2007) and Afterhours (Godinho et al. 2007). Both the Ovt and Afh mutations are located in the same gene Fbxl3. Fbxl3 encodes an ubiquitin ligase E3 and controls CRY stability by inducing CRY protein ubiquitination and degradation (Godinho et al. 2007; Siepka et al. 2007). Delayed expression of CRY1 could be caused by the combinatorial effect of delayed transcription activation and active degradation. These data suggest that temporal control of clock gene products (like PER and CRY) is also important for generating circadian rhythms. Effects of the CKIεtau and Fbxl3Afh mutations are additive and independently contribute to circadian period (Maywood et al. 2011).
4.3 Post-Translational Oscillation of the Mammalian Circadian Clock
Phosphorylation-dependent degradation may be directly related to PER oscillation. Two reports showed that PER2 protein translated from constitutively expressed mRNA undergoes circadian oscillation (Fujimoto et al. 2006; Nishii et al. 2006). These studies imply that a layer of post-translational control blankets the transcription-translation circadian machinery. Consistent with this idea, several studies have shown that circadian rhythmicity is robust against fluctuations in oscillating transcriptional activity. For example, the expression pattern of Bmal1 and Clock can be constant throughout the circadian cycle (von Gall et al. 2003). Reducing the overall transcriptional activity only modestly affects the period length of circadian rhythms in cultured cells (Dibner et al. 2009). Even in for CRY, rhythmic expression is dispensable for circadian oscillation to a certain extent; weak circadian oscillations can be observed in Cry1−/−:Cry2−/−cells rescued by Cry1 under constant expression (Ukai-Tadenuma et al. 2011) or a constant supply of CRY proteins (Fan et al. 2007). Genetic studies in Drosophila show that flies with constant expression of PER maintain circadian rhythmicity (Ewer et al. 1988; Frisch et al. 1994; Vosshall and Young 1995; Yang and Sehgal 2001). Taken together, these results suggest that circadian oscillations do not necessarily depend solely on the transcriptional activity in the E/E′-box feedback loop, because post-translational control of clock proteins can compensate for loss of transcriptional rhythms.
A post-translational circadian oscillator was also found in the cyanobacterium circadian clock. Oscillations occur in the phosphorylation state of KaiC, a central component of cyanobacterial circadian clock, even after the termination of global transcriptional activity (Tomita et al. 2005). This KaiC-phosphorylation rhythm can be reconstituted in vitro by mixing KaiC and its regulatory factors KaiB and KaiC together with ATP (Nakajima et al. 2005). In mammals, a recent study discovered the presence of the circadian oscillations in the redox state of enucleated human red blood cells (O’Neill and Reddy 2011). The circadian oscillation in redox status of peroxiredoxin proteins is conserved from prokaryotes to eukaryotes (Edgar et al. 2012) and can regulate the neuronal activity of SCN (Wang et al. 2012). Although a core post-translational circadian oscillator in mammals remains to be identified, cooperation of transcription-translation oscillator and post-transcriptional oscillator would provide a more robust circadian timekeeping system. The investigation of compatible interactions between delayed negative feedback loops mediated by the CCEs and yet-unknown core post-translational oscillators will lead to a new understanding of mammalian circadian clocks.
References
Akashi M, Soma H, Yamamoto T, Tsugitomi A, Yamashita S, Nishida E, Yasuda A, Liao JK, Node K (2010) Noninvasive method for assessing the human circadian clock using hair follicle cells. Proc Natl Acad Sci USA 107:15643–15648
Bae K, Jin X, Maywood ES, Hastings MH, Reppert SM, Weaver DR (2001) Differential functions of mPer1, mPer2, and mPer3 in the SCN circadian clock. Neuron 30:525–536
Balsalobre A, Damiola F, Schibler U (1998) A serum shock induces circadian gene expression in mammalian tissue culture cells. Cell 93:929–937
Bugge A, Feng D, Everett LJ, Briggs ER, Mullican SE, Wang F, Jager J, Lazar MA (2012) Rev-erbalpha and Rev-erbbeta coordinately protect the circadian clock and normal metabolic function. Genes Dev 26:657–667
Bunger MK, Wilsbacher LD, Moran SM, Clendenin C, Radcliffe LA, Hogenesch JB, Simon MC, Takahashi JS, Bradfield CA (2000) Mop3 is an essential component of the master circadian pacemaker in mammals. Cell 103:1009–1017
Chen Z, Yoo SH, Park YS, Kim KH, Wei S, Buhr E, Ye ZY, Pan HL, Takahashi JS (2012) Identification of diverse modulators of central and peripheral circadian clocks by high-throughput chemical screening. Proc Natl Acad Sci USA 109:101–106
Chiu JC, Ko HW, Edery I (2011) NEMO/NLK phosphorylates PERIOD to initiate a time-delay phosphorylation circuit that sets circadian clock speed. Cell 145:357–370
Cho H, Zhao X, Hatori M, Yu RT, Barish GD, Lam MT, Chong LW, DiTacchio L, Atkins AR, Glass CK, Liddle C, Auwerx J, Downes M, Panda S, Evans RM (2012) Regulation of circadian behaviour and metabolism by REV-ERB-alpha and REV-ERB-beta. Nature 485:123–127
DeBruyne JP, Weaver DR, Reppert SM (2007) CLOCK and NPAS2 have overlapping roles in the suprachiasmatic circadian clock. Nat Neurosci 10:543–545
Delaunay F, Laudet V (2002) Circadian clock and microarrays: mammalian genome gets rhythm. Trends Genet 18:595–597
Dibner C, Sage D, Unser M, Bauer C, d’Eysmond T, Naef F, Schibler U (2009) Circadian gene expression is resilient to large fluctuations in overall transcription rates. EMBO J 28:123–134
Dudley CA, Erbel-Sieler C, Estill SJ, Reick M, Franken P, Pitts S, McKnight SL (2003) Altered patterns of sleep and behavioral adaptability in NPAS2-deficient mice. Science 301:379–383
Dunlap JC (1999) Molecular bases for circadian clocks. Cell 96:271–290
Edgar RS, Green EW, Zhao Y, van Ooijen G, Olmedo M, Qin X, Xu Y, Pan M, Valekunja UK, Feeney KA, Maywood ES, Hastings MH, Baliga NS, Merrow M, Millar AJ, Johnson CH, Kyriacou CP, O’Neill JS, Reddy AB (2012) Peroxiredoxins are conserved markers of circadian rhythms. Nature 485:459–464
Eide EJ, Woolf MF, Kang H, Woolf P, Hurst W, Camacho F, Vielhaber EL, Giovanni A, Virshup DM (2005) Control of mammalian circadian rhythm by CKIε-regulated proteasome-mediated PER2 degradation. Mol Cell Biol 25:2795–2807
Elowitz MB, Leibler S (2000) A synthetic oscillatory network of transcriptional regulators. Nature 403:335–338
Ewer J, Rosbash M, Hall JC (1988) An inducible promoter fused to the period gene in Drosophila conditionally rescues adult per-mutant arrhythmicity. Nature 333:82–84
Falvey E, Marcacci L, Schibler U (1996) DNA-binding specificity of PAR and C/EBP leucine zipper proteins: a single amino acid substitution in the C/EBP DNA-binding domain confers PAR-like specificity to C/EBP. Biol Chem 377:797–809
Fan Y, Hida A, Anderson DA, Izumo M, Johnson CH (2007) Cycling of CRYPTOCHROME proteins is not necessary for circadian-clock function in mammalian fibroblasts. Curr Biol 17:1091–1100
Frisch B, Hardin PE, Hamblen-Coyle MJ, Rosbash M, Hall JC (1994) A promoterless period gene mediates behavioral rhythmicity and cyclical per expression in a restricted subset of the Drosophila nervous system. Neuron 12:555–570
Fujimoto Y, Yagita K, Okamura H (2006) Does mPER2 protein oscillate without its coding mRNA cycling? Post-transcriptional regulation by cell clock. Genes Cells 11:525–530
Fustin JM, O’Neill JS, Hastings MH, Hazlerigg DG, Dardente H (2009) Cry1 circadian phase in vitro: wrapped up with an E-box. J Biol Rhythms 24:16–24
Gachon F, Fonjallaz P, Damiola F, Gos P, Kodama T, Zakany J, Duboule D, Petit B, Tafti M, Schibler U (2004) The loss of circadian PAR bZip transcription factors results in epilepsy. Genes Dev 18:1397–1412
Gallego M, Virshup DM (2007) Post-translational modifications regulate the ticking of the circadian clock. Nat Rev Mol Cell Biol 8:139–148
Gascoyne DM, Long E, Veiga-Fernandes H, de Boer J, Williams O, Seddon B, Coles M, Kioussis D, Brady HJ (2009) The basic leucine zipper transcription factor E4BP4 is essential for natural killer cell development. Nat Immunol 10:1118–1124
Gekakis N, Staknis D, Nguyen HB, Davis FC, Wilsbacher LD, King DP, Takahashi JS, Weitz CJ (1998) Role of the CLOCK protein in the mammalian circadian mechanism. Science 280:1564–1569
Godinho SI, Maywood ES, Shaw L, Tucci V, Barnard AR, Busino L, Pagano M, Kendall R, Quwailid MM, Romero MR, O’Neill J, Chesham JE, Brooker D, Lalanne Z, Hastings MH, Nolan PM (2007) The after-hours mutant reveals a role for Fbxl3 in determining mammalian circadian period. Science 316:897–900
Griffin EA Jr, Staknis D, Weitz CJ (1999) Light-independent role of CRY1 and CRY2 in the mammalian circadian clock. Science 286:768–771
Grundschober C, Delaunay F, Puhlhofer A, Triqueneaux G, Laudet V, Bartfai T, Nef P (2001) Circadian regulation of diverse gene products revealed by mRNA expression profiling of synchronized fibroblasts. J Biol Chem 276:46751–46758
Harding HP, Lazar MA (1993) The orphan receptor Rev-ErbA alpha activates transcription via a novel response element. Mol Cell Biol 13:3113–3121
Hirota T, Lewis WG, Liu AC, Lee JW, Schultz PG, Kay SA (2008) A chemical biology approach reveals period shortening of the mammalian circadian clock by specific inhibition of GSK-3beta. Proc Natl Acad Sci USA 105:20746–20751
Hogenesch JB, Chan WK, Jackiw VH, Brown RC, Gu YZ, Pray-Grant M, Perdew GH, Bradfield CA (1997) Characterization of a subset of the basic-helix-loop-helix-PAS superfamily that interacts with components of the dioxin signaling pathway. J Biol Chem 272:8581–8593
Isojima Y, Nakajima M, Ukai H, Fujishima H, Yamada RG, Masumoto KH, Kiuchi R, Ishida M, Ukai-Tadenuma M, Minami Y, Kito R, Nakao K, Kishimoto W, Yoo SH, Shimomura K, Takao T, Takano A, Kojima T, Nagai K, Sakaki Y, Takahashi JS, Ueda HR (2009) CKIepsilon/delta-dependent phosphorylation is a temperature-insensitive, period-determining process in the mammalian circadian clock. Proc Natl Acad Sci USA 106:15744–15749
Kornmann B, Schaad O, Bujard H, Takahashi JS, Schibler U (2007) System-driven and oscillator-dependent circadian transcription in mice with a conditionally active liver clock. PLoS Biol 5:e34
Kume K, Zylka MJ, Sriram S, Shearman LP, Weaver DR, Jin X, Maywood ES, Hastings MH, Reppert SM (1999) mCRY1 and mCRY2 are essential components of the negative limb of the circadian clock feedback loop. Cell 98:193–205
Liu AC, Tran HG, Zhang EE, Priest AA, Welsh DK, Kay SA (2008) Redundant function of REV-ERBalpha and beta and non-essential role for Bmal1 cycling in transcriptional regulation of intracellular circadian rhythms. PLoS Genet 4:e1000023
Lopez-Molina L, Conquet F, Dubois-Dauphin M, Schibler U (1997) The DBP gene is expressed according to a circadian rhythm in the suprachiasmatic nucleus and influences circadian behavior. EMBO J 16:6762–6771
Lowrey PL, Shimomura K, Antoch MP, Yamazaki S, Zemenides PD, Ralph MR, Menaker M, Takahashi JS (2000) Positional syntenic cloning and functional characterization of the mammalian circadian mutation tau. Science 288:483–492
Maywood ES, Chesham JE, Meng QJ, Nolan PM, Loudon AS, Hastings MH (2011) Tuning the period of the mammalian circadian clock: additive and independent effects of CK1epsilonTau and Fbxl3Afh mutations on mouse circadian behavior and molecular pacemaking. J Neurosci 31:1539–1544
Mitsui S, Yamaguchi S, Matsuo T, Ishida Y, Okamura H (2001) Antagonistic role of E4BP4 and PAR proteins in the circadian oscillatory mechanism. Genes Dev 15:995–1006
Nagoshi E, Saini C, Bauer C, Laroche T, Naef F, Schibler U (2004) Circadian gene expression in individual fibroblasts: cell-autonomous and self-sustained oscillators pass time to daughter cells. Cell 119:693–705
Nakajima M, Imai K, Ito H, Nishiwaki T, Murayama Y, Iwasaki H, Oyama T, Kondo T (2005) Reconstitution of circadian oscillation of cyanobacterial KaiC phosphorylation in vitro. Science 308:414–415
Nishii K, Yamanaka I, Yasuda M, Kiyohara YB, Kitayama Y, Kondo T, Yagita K (2006) Rhythmic post-transcriptional regulation of the circadian clock protein mPER2 in mammalian cells: a real-time analysis. Neurosci Lett 401:44–48
O’Neill JS, Reddy AB (2011) Circadian clocks in human red blood cells. Nature 469:498–503
Pace-Schott EF, Hobson JA (2002) The neurobiology of sleep: genetics, cellular physiology and subcortical networks. Nat Rev Neurosci 3:591–605
Panda S, Antoch MP, Miller BH, Su AI, Schook AB, Straume M, Schultz PG, Kay SA, Takahashi JS, Hogenesch JB (2002) Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 109:307–320
Preitner N, Damiola F, Lopez-Molina L, Zakany J, Duboule D, Albrecht U, Schibler U (2002) The orphan nuclear receptor REV-ERBalpha controls circadian transcription within the positive limb of the mammalian circadian oscillator. Cell 110:251–260
Querfurth C, Diernfellner AC, Gin E, Malzahn E, Hofer T, Brunner M (2011) Circadian conformational change of the Neurospora clock protein FREQUENCY triggered by clustered hyperphosphorylation of a basic domain. Mol Cell 43:713–722
Ralph MR, Menaker M (1988) A mutation of the circadian system in golden hamsters. Science 241:1225–1227
Reppert SM, Weaver DR (2002) Coordination of circadian timing in mammals. Nature 418:935–941
Rudic RD, McNamara P, Reilly D, Grosser T, Curtis AM, Price TS, Panda S, Hogenesch JB, FitzGerald GA (2005) Bioinformatic analysis of circadian gene oscillation in mouse aorta. Circulation 112:2716–2724
Sato TK, Yamada RG, Ukai H, Baggs JE, Miraglia LJ, Kobayashi TJ, Welsh DK, Kay SA, Ueda HR, Hogenesch JB (2006) Feedback repression is required for mammalian circadian clock function. Nat Genet 38:312–319
Shanware NP, Hutchinson JA, Kim SH, Zhan L, Bowler MJ, Tibbetts RS (2011) Casein kinase 1-dependent phosphorylation of familial advanced sleep phase syndrome-associated residues controls PERIOD 2 stability. J Biol Chem 286:12766–12774
Shi S, Hida A, McGuinness OP, Wasserman DH, Yamazaki S, Johnson CH (2010) Circadian clock gene Bmal1 is not essential; functional replacement with its paralog, Bmal2. Curr Biol 20:316–321
Shirogane T, Jin J, Ang XL, Harper JW (2005) SCFbeta-TRCP controls clock-dependent transcription via casein kinase 1-dependent degradation of the mammalian period-1 (Per1) protein. J Biol Chem 280:26863–26872
Siepka SM, Yoo SH, Park J, Song W, Kumar V, Hu Y, Lee C, Takahashi JS (2007) Circadian mutant overtime reveals F-box protein FBXL3 regulation of cryptochrome and period gene expression. Cell 129:1011–1023
Storch KF, Lipan O, Leykin I, Viswanathan N, Davis FC, Wong WH, Weitz CJ (2002) Extensive and divergent circadian gene expression in liver and heart. Nature 417:78–83
Stricker J, Cookson S, Bennett MR, Mather WH, Tsimring LS, Hasty J (2008) A fast, robust and tunable synthetic gene oscillator. Nature 456:516–519
Toh KL, Jones CR, He Y, Eide EJ, Hinz WA, Virshup DM, Ptacek LJ, Fu YH (2001) An hPer2 phosphorylation site mutation in familial advanced sleep phase syndrome. Science 291:1040–1043
Tomita J, Nakajima M, Kondo T, Iwasaki H (2005) No transcription-translation feedback in circadian rhythm of KaiC phosphorylation. Science 307:251–254
Tsuchiya Y, Akashi M, Nishida E (2003) Temperature compensation and temperature resetting of circadian rhythms in mammalian cultured fibroblasts. Genes Cells 8:713–720
Ueda HR, Chen W, Adachi A, Wakamatsu H, Hayashi S, Takasugi T, Nagano M, Nakahama K, Suzuki Y, Sugano S, Iino M, Shigeyoshi Y, Hashimoto S (2002) A transcription factor response element for gene expression during circadian night. Nature 418:534–539
Ueda HR, Hayashi S, Chen W, Sano M, Machida M, Shigeyoshi Y, Iino M, Hashimoto S (2005) System-level identification of transcriptional circuits underlying mammalian circadian clocks. Nat Genet 37:187–192
Ukai-Tadenuma M, Kasukawa T, Ueda HR (2008) Proof-by-synthesis of the transcriptional logic of mammalian circadian clocks. Nat Cell Biol 10:1154–1163
Ukai-Tadenuma M, Yamada RG, Xu H, Ripperger JA, Liu AC, Ueda HR (2011) Delay in feedback repression by cryptochrome 1 is required for circadian clock function. Cell 144:268–281
van der Horst GT, Muijtjens M, Kobayashi K, Takano R, Kanno S, Takao M, de Wit J, Verkerk A, Eker AP, van Leenen D, Buijs R, Bootsma D, Hoeijmakers JH, Yasui A (1999) Mammalian Cry1 and Cry2 are essential for maintenance of circadian rhythms. Nature 398:627–630
Vanselow K, Vanselow JT, Westermark PO, Reischl S, Maier B, Korte T, Herrmann A, Herzel H, Schlosser A, Kramer A (2006) Differential effects of PER2 phosphorylation: molecular basis for the human familial advanced sleep phase syndrome (FASPS). Genes Dev 20:2660–2672
Vitaterna MH, Selby CP, Todo T, Niwa H, Thompson C, Fruechte EM, Hitomi K, Thresher RJ, Ishikawa T, Miyazaki J, Takahashi JS, Sancar A (1999) Differential regulation of mammalian period genes and circadian rhythmicity by cryptochromes 1 and 2. Proc Natl Acad Sci USA 96:12114–12119
Vollmers C, Panda S, DiTacchio L (2008) A high-throughput assay for siRNA-based circadian screens in human U2OS cells. PLoS One 3:e3457
von Gall C, Noton E, Lee C, Weaver DR (2003) Light does not degrade the constitutively expressed BMAL1 protein in the mouse suprachiasmatic nucleus. Eur J Neurosci 18:125–133
Vosshall LB, Young MW (1995) Circadian rhythms in Drosophila can be driven by period expression in a restricted group of central brain cells. Neuron 15:345–360
Wang TA, Yu YV, Govindaiah G, Ye X, Artinian L, Coleman TP, Sweedler JV, Cox CL, Gillette MU (2012) Circadian rhythm of redox state regulates excitability in suprachiasmatic nucleus neurons. Science 337:839–842
Welsh DK, Yoo SH, Liu AC, Takahashi JS, Kay SA (2004) Bioluminescence imaging of individual fibroblasts reveals persistent, independently phased circadian rhythms of clock gene expression. Curr Biol 14:2289–2295
Xu Y, Padiath QS, Shapiro RE, Jones CR, Wu SC, Saigoh N, Saigoh K, Ptacek LJ, Fu YH (2005) Functional consequences of a CKIδ mutation causing familial advanced sleep phase syndrome. Nature 434:640–644
Xu Y, Toh KL, Jones CR, Shin JY, Fu YH, Ptacek LJ (2007) Modeling of a human circadian mutation yields insights into clock regulation by PER2. Cell 128:59–70
Yamazaki S, Numano R, Abe M, Hida A, Takahashi R, Ueda M, Block GD, Sakaki Y, Menaker M, Tei H (2000) Resetting central and peripheral circadian oscillators in transgenic rats. Science 288:682–685
Yang Z, Sehgal A (2001) Role of molecular oscillations in generating behavioral rhythms in Drosophila. Neuron 29:453–467
Yoo SH, Ko CH, Lowrey PL, Buhr ED, Song EJ, Chang S, Yoo OJ, Yamazaki S, Lee C, Takahashi JS (2005) A noncanonical E-box enhancer drives mouse Period2 circadian oscillations in vivo. Proc Natl Acad Sci USA 102:2608–2613
Young MW, Kay SA (2001) Time zones: a comparative genetics of circadian clocks. Nat Rev Genet 2:702–715
Zheng B, Albrecht U, Kaasik K, Sage M, Lu W, Vaishnav S, Li Q, Sun ZS, Eichele G, Bradley A, Lee CC (2001) Nonredundant roles of the mPer1 and mPer2 genes in the mammalian circadian clock. Cell 105:683–694
Zvonic S, Ptitsyn AA, Conrad SA, Scott LK, Floyd ZE, Kilroy G, Wu X, Goh BC, Mynatt RL, Gimble JM (2006) Characterization of peripheral circadian clocks in adipose tissues. Diabetes 55:962–970
Zvonic S, Ptitsyn AA, Kilroy G, Wu X, Conrad SA, Scott LK, Guilak F, Pelled G, Gazit D, Gimble JM (2007) Circadian oscillation of gene expression in murine calvarial bone. J Bone Miner Res 22:357–365
Acknowledgements
We thank Ms. Maki Ukai-Tadenuma and Drs. Arthur Millius and Rikuhiro Yamada for figure preparation and valuable comments.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Minami, Y., Ode, K.L., Ueda, H.R. (2013). Mammalian Circadian Clock: The Roles of Transcriptional Repression and Delay. In: Kramer, A., Merrow, M. (eds) Circadian Clocks. Handbook of Experimental Pharmacology, vol 217. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-25950-0_15
Download citation
DOI: https://doi.org/10.1007/978-3-642-25950-0_15
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-25949-4
Online ISBN: 978-3-642-25950-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)