Abstract
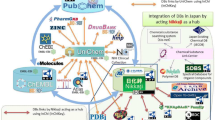
A large number of biological databases are currently in use by scientists. These databases employ different formats, many of which can be converted into resource description format (RDF), which can be subsequently queried using semantic web methods. These databases have “inter” and “intra” database relationships. RDF has an inherent graph structure that facilitates exploration of connections between data via graphical representations known as knowledge graphs. In this paper, we survey the existing methods that are in use to link biological databases and evaluate the effectiveness with which the available approaches can predict unknown links between entities in databases as a means of improving knowledge graphs.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Beckett, D., McBride, B.: RDF/XML Syntax Specification. W3C Work, pp. 1–56 (2003)
Belleau, F., et al.: Bio2RDF: towards a mashup to build bioinformatics knowledge systems. J. Biomed. Inform. 41(5), 706–716 (2008)
Momtchev, V., Peychev D., Primov, T., Georgiev, G.: Expanding the pathway and interaction knowledge in linked life data. In: Proceedings of International Semantic Web Challenge (2009)
Samwald, M., et al.: Linked Open drug data for pharmaceutical research and development. J. Cheminfo. 3 (2011). https://doi.org/10.1186/1758-2946-3-19
KaBOB: ontology-based semantic integration of biomedical databases. BMC Bioinform. 16, 126 (2015)
Ruttenberg, A., et al.: Life sciences on the semantic web: the neurocommons and beyond. Brief. Bioinfo. 10, 193–204 (2009)
Lauw, H., et al.: Homophily in the digital world: a live journal case study. IEEE Internet Comput. 14, 15–23 (2010)
Katz, L., et al.: A new status index derived from sociometric analysis. Psychometrika 18, 39–43 (1953)
Acar, E., et al.: Link prediction on evolving data using matrix and tensor factorizations. In: 2009 IEEE International Conference on Data Mining Workshops, pp. 262–269 (2009)
Jaccard, P., et al.: Étude comparative de la distribution florale dans une portion des Alpes et des Jura. Bull. del la Société Vaudoise des Sci. Nat. 37, 547–579 (1901)
Newman, M.E.J., et al.: Clustering and preferential attachment in growing networks. Phys. Rev. 64, 25102 (2001)
Liu, W., Lu, L.: Link prediction based on local random walk. EPL (Europhysics Lett.) 89, 58007 (2010)
Liben-Nowell, D., Kleinberg, J.: The link prediction problem for social networks. In: Proceedings of Twelfth Annual ACM International Conference Information and Knowledge Management, pp. 556–559 (2003)
Brin, S., Page, L.: The anatomy of a large-scale hypertextual Web search engine BT, Computer Networks and ISDN Systems. Comput. Netw. ISDN Syst. 30, 107–117 (1998)
Carroll, J.D., Chang, J.J.: Analysis of individual differences in multidimensional scaling via an n-way generalization of ‘Eckart-Young’ decomposition. Psychometrika 35, 283–319 (1970)
Harshman, R.: Foundations of the PARAFAC procedure: models and conditions for an ‘explanatory’ multimodal factor analysis. UCLA Work. Pap. Phonetics 16, 1–84 (1970)
Tucker, L.R.: The extension of factor analysis to three-dimensional matrices. In: Contributions to Mathematical Psychology, pp. 110–119 (1964)
Tucker, L.R.: PARAFAC2: mathematical and technical notes. UCLA Work. Pap. Phonetics 22, 30–44 (1972)
Hong, S.J., Harshman, R.: Shifted factor analysis, Part III: N-way generalization and application. J. Chemom. 17, 389–399 (2003)
Bro, R., et al.: Modeling multi-way data with linearly dependent loadings. J. Chemom. 23, 324–340 (2009)
Harshman, R.A., et al.: Shifted factor analysis? Part I: models and properties. J. Chemom. 17, 363–378 (2003)
Nickel, M., et al.: Factorizing YAGO. In: Proceedings of the 21st international conference on World Wide Web - WWW 2012, p. 271 (2012)
Nickel, M., Tresp, V.: Tensor factorization for multi-relational learning. Lecture Notes in Computer Science, pp. 617–621 (2013)
Nickel, M., et al.: A review of relational machine learning for knowledge graphs. Proc. IEEE 104, 11–33 (2016)
Jiang, X., et al.: Link prediction in multi-relational graphs using additive models. In: International Workshop on Semantic Technologies meet Recommender Systems and Big Data at the ISWC, pp. 1–12 (2012)
Riedel, S., et al.: Relation extraction with matrix factorization and universal schemas. In: Proceedings 2013 Conference of the North American Chapter of the Association Computational Linguistics Human Language Technologies, pp. 74–84 (2013)
Huang, Y., et al.: A scalable approach for statistical learning in semantic graphs. Semantic Web 5, 5–22 (2014)
Tresp, V., et al.: Materializing and querying learned knowledge. In: CEUR Workshop Proceedings (2009)
Richardson, M., Domingos, P.: Markov logic networks. In: Machine Learning, pp. 107–136 (2009)
Bishop, C.M.: Pattern Recognition and Machine Learning. Springer (2006). ISBN 0-387-31073-8
Nickel, M., Tresp, V., Kriegel, H.P.: A three-way model for collective learning on multi-relational data. In: Proceedings of the 28th International Conference on Machine Learning, pp. 809–816 (2011)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this paper
Cite this paper
Zaki, N., Tennakoon, C., Al Ashwal, H., Al Jaberi, A., Al Ameri, A. (2019). Methods of Creating Knowledge Graph by Linking Biological Databases. In: Fdez-Riverola, F., Mohamad, M., Rocha, M., De Paz, J., González, P. (eds) Practical Applications of Computational Biology and Bioinformatics, 12th International Conference. PACBB2018 2018. Advances in Intelligent Systems and Computing, vol 803. Springer, Cham. https://doi.org/10.1007/978-3-319-98702-6_7
Download citation
DOI: https://doi.org/10.1007/978-3-319-98702-6_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-98701-9
Online ISBN: 978-3-319-98702-6
eBook Packages: Intelligent Technologies and RoboticsIntelligent Technologies and Robotics (R0)