Abstract
Microscopy became a scientific investigation method in the seventeenth century with the application of the first build microscopes on biological samples [1, 2]. Soon it became a popular method to stain samples in order to visualise particular (cellular and subcellular) structures [3]. These stains, either based on absorption or fluorescence, have limitations in respect to their specificity and are often toxic to cells, which limits investigations to short intervals or even dead samples. In 1987 the idea came up to use a fluorescent protein that was discovered 25 years before [4], in particular a green fluorescent protein (GFP) form the medusa Aequorea victoria to label cells and cellular structures [5]. With the sequencing and cloning of GFP, a so-called ‘green revolution’ started, which led to regular usage of fluorescent proteins as markers or sensors (for details see below) in the majority of cellular research in physiology, microbiology, pharmacology, molecular biology, anatomy, cell biology, biophysics and many other biomedical fields. Although the expression of the fluorescent proteins and their optical investigation can already be regarded as optogenetic tools, this term was only applied when the optical properties of proteins were used to manipulate cells. The best-known example of such a protein is the channelrhodopsin, a light-gated ion channel [6, 7]. When this ion channel is expressed in a membrane and illuminated with light of the appropriate wavelength, the channel will be activated and opened, which results in passive transportation of ions across the membrane and a change of the membrane potential. However, within this chapter we consider both aspects, the observation and the manipulation as optogenetic tools. To use the optogenetic tool, the genes of these proteins need to be transferred into the cells to allow the expression of the protein. For an overview of gene delivery into target cells, see [8].
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
General Introduction
Microscopy became a scientific investigation method in the seventeenth century with the application of the first build microscopes on biological samples [1, 2]. Soon it became a popular method to stain samples in order to visualise particular (cellular and subcellular) structures [3]. These stains, either based on absorption or fluorescence, have limitations in respect to their specificity and are often toxic to cells, which limits investigations to short intervals or even dead samples. In 1987 the idea came up to use a fluorescent protein that was discovered 25 years before [4], in particular a green fluorescent protein (GFP) form the medusa Aequorea victoria to label cells and cellular structures [5]. With the sequencing and cloning of GFP, a so-called ‘green revolution’ started, which led to regular usage of fluorescent proteins as markers or sensors (for details see below) in the majority of cellular research in physiology, microbiology, pharmacology, molecular biology, anatomy, cell biology, biophysics and many other biomedical fields. Although the expression of the fluorescent proteins and their optical investigation can already be regarded as optogenetic tools, this term was only applied when the optical properties of proteins were used to manipulate cells. The best-known example of such a protein is the channelrhodopsin, a light-gated ion channel [6, 7]. When this ion channel is expressed in a membrane and illuminated with light of the appropriate wavelength, the channel will be activated and opened, which results in passive transportation of ions across the membrane and a change of the membrane potential. However, within this chapter we consider both aspects, the observation and the manipulation as optogenetic tools. To use the optogenetic tool, the genes of these proteins need to be transferred into the cells to allow the expression of the protein. For an overview of gene delivery into target cells, see [8].
Quantification of Genetically Encoded Biosensors with a Special Emphasis on Förster Resonance Energy Transfer
The design of genetically encoded biosensors can follow several concepts. One of the early approaches was bimolecular fluorescence complementation [9] that can report protein-protein interactions but does not allow the read-out of fast dynamic processes like excitation-contraction coupling. For biosensors requiring fast kinetics such as calcium- or membrane potential sensors, two engineering strategies have been established: circular permutated fluorescent proteins (CPFP) and sensors taking advantage of the Förster Resonance Energy Transfer (FRET). Both concepts rely on the conformational change of the sensing domain, which induces a spatial rearrangement of the fluorescent protein(s). CPFPs can be described as a switch from a protonated to a deprotonated fluorophore [10] by restricting solvent access to the fluorophore and stabilizing it in an ionized form [11]. This stabilization is supported by extensive contacts between the structure inducing the conformational change and the fluorophore moieties [10]. Apart from the design approach, the application of CPFPs has a significant impact on the recording strategy. CPFPs are often used as equivalents to single-excitation, single-emission small-molecule dye probes. However, some of the CPFPs can also be used in a ratiometric excitation mode, e.g., ratiometric pericam. Despite the relative ease of recording when using the single-excitation single-emission mode, these Ca2+ indicators are associated with a number of shortcomings because the measured intensity changes can alternatively be a result of sample movement, inhomogeneous sensor distribution, or changing autofluorescence [12, 13]. In addition to such complications, pH-sensitivity, irreversible or reversible photoconversion, and rather complex intensity relationships can contribute to falsified interpretations of their intensity changes [14].
A real quantitative approach independent on the sensor concentration can be performed taking advantage of the Förster Resonance Energy Transfer (FRET).
FRET is a quantum mechanical effect between two chemical moieties when one is fluorescing and close enough (normally a few nanometers) to another light-absorbing moiety [15, 16]. Already in 1996 variations of the GFP appeared with the clear intention to design biosensors based on FRET [17]. A decent number of FRET-based biosensors have been designed ever since, including sensors to detect cardiac excitation coupling (ECC), like Ca2+-indicators and membrane potential sensors (for details see below). In most of these application cases, FRET-based biosensors are composed of donors and acceptors that are in tandem covalently connected and thus with fixed donor/acceptor stoichiometry. A continuous challenge is the quantitative microscopic measurement of FRET, because at the wavelength of the donor excitation also a fraction of the acceptor is likely to be excited, and the emission of donor and acceptor is widely overlapping. The difference between the measurement of the pure fluorescence ratio between donor and acceptor and a quantitative FRET analysis in terms of apparent FRET-efficiency (see below) is depicted in Fig. 1. Therefore we provide a short description on quantitative FRET measurements.
Comparison of Ca2+ transients calculated as the yellow fluorescent protein (YFP)/cyan fluorescent protein (CFP) ratio and apparent Förster resonance energy transfer (FRET) efficiency. (Aa) Adult cardiac myocytes expressing YC3.6. The cells were electrically paced, and the resulting CFP and YFP fluorescence were compared after CFP excitation and displayed as the YFP/CFP ratio. (Ab) The same Ca2+ transients as in (Aa) but displayed as the apparent FRET ratio. (Ac) Diagram depicting the signal:noise (S/N) ratio for the two presentation methods. (B) Cardiac myocytes expressing YC3.6 were electrically stimulated and analysed. Two representative myocytes displaying substantially different signal transients when calculating simple YFP/CFP fluorescence ratios are shown (dotted lines). Using the same raw data, we calculated the apparent FRET efficiency; the resulting signal transients (solid lines) confirm the expected similar behavior of the two myocytes. (C) High-speed recordings of YC3.6 expressing cardiomyocytes. (Ca) Image series of a Ca2+ upstroke at an image recording frequency of 100 Hz. Each pair of consecutive images was used to calculate the image of the apparent FRET ratio. (Cb) Example trace of consecutive Ca2+ transients of a single cell recorded at 100 Hz. The multiplicity of infection for all YC3.6 transductions was 4 × 104 pfu/1500 cells. E indicates the FRET efficiency; and fDA, fraction of the FRET sensor fluorescent entities. This figure is reproduced from [14], with permission from Wolters Kluwer
The fluorescence from donor and acceptor fluorophores undergoing FRET is the superposition of five quantities: the donor fluorescence from (i) free donor and (ii) donor-acceptor complexes and the acceptor fluorescence from (iii) free acceptor, (iv) directly excited acceptor in donor acceptor complexes and (v) acceptor emission resulting from energy transfer from a donor within a donor-acceptor complex. Due to this complexity, it is apparent that the direct determination of FRET efficiency from spectrally resolved intensity-based measurements is not possible without additional information. To handle the problem that an acceptor emission channel is often polluted by the bleed-through of donor emission, Youvan et al. introduced a filter based method to extract acceptor fluorescence intensity changes while correcting for donor direct acceptor excitation and donor bleed-through [18].
For this method, three measurements are required. First, at an excitation wavelength that directly excites the donor, the emission is measured in the acceptor and the donor channels, \( {F}^{ex_D,{em}_A} \) and \( {F}^{ex_D,{em}_D} \), respectively. Next, fluorescence is measured in the acceptor channel \( {F}^{ex_A,{em}_A} \) at an excitation wavelength that selectively excites acceptor. The amount of donor bleed-through into the acceptor channel is determined by a ‘donor-only’ measurement, which provides the calibration constant \( \beta ={F}_D^{ex_D,{em}_A}/{F}_D^{ex_D,{em}_D} \). In an ‘acceptor-only’ measurement, the extent to which the short wavelength excitation directly excites the acceptor is determined relative to the excitation at the longer wavelength. This results in the constant \( \alpha ={F}_A^{ex_D,{em}_A}/{F}_A^{ex_A,{em}_A} \). It should be noted that, while donor bleed-though into the acceptor channel is considered and corrected for, any acceptor bleed-through into the donor channel is neglected. Methods that use these three measurements are collectively known as ‘three-cube’ methods, due to the necessity of three different filter cubes to perform the measurements. Both calibration constants, α and β, are fluorophore and system specific and may vary with the performance of the device. It is also notable that the nF as provided by Eq. (1) does not represent FRET efficiency, rather it only provides a value that varies with FRET [19]. Specifically, this FRET index varies with donor and acceptor concentration. Several studies have attempted to address this problem by scaling nF by various, somewhat arbitrary combinations of the donor and acceptor signals, but no practical solution was achieved [20,21,22].
A quantitative solution to determine apparent FRET efficiency from measurements of acceptor intensity was proposed by Lakowicz [23] in the form of
where \( {\varepsilon}_D^{ex_D} \) and \( {\varepsilon}_A^{ex_D} \) are the extinction coefficients of the donor and acceptor at donor excitation. It should be noted that the fractional occupancy of acceptor f A in (2) is denoted with f D [23]. This equation cannot be directly applied as written, but can only be used in the hypothetical situation that the acceptor in a FRET sample can be measured in absence of donor. More recently, Hoppe and colleagues applied corrections for donor bleed-through and excitation crosstalk to Eq. (2) to quantify apparent FRET efficiency with sensitized emission measurements using a three cube microscope setup [22]. The resulting equation,
where \( \gamma ={\varepsilon}_A^{ex_D}/{\varepsilon}_D^{ex_D} \), still requires information about the relative excitability of donor and acceptor at donor excitation wavelength. By applying Eq. (3) to a reference measurement of donor-acceptor tandem construct with known characteristic FRET efficiency, as measured from fluorescence lifetime measurements, γ can be determined. Hoppe et al. further extended this approach by taking into account the donor quenching due to FRET and were able to calculate the donor-dependent apparent FRET efficiency [22]
as well as the donor acceptor ratio
To calculate Ef D and R t, the constant ξ, which contains information about the donor and acceptor quantum efficiencies and the device detection efficiency, must be determined in a manner similar to γ, by applying reference measurements of a known FRET efficiency construct and solving Eq. (4). Thus, in addition to quantifying α and β from ‘acceptor-only’ and ‘donor-only’ reference measurements similarly to Youvan et al. [18], an additional reference measurement of a donor-acceptor tandem construct with known FRET efficiency must be performed in order to determine γ and ξ. Furthermore, the excitation wavelengths and emission channels are constrained as in the calculation of nF, such that ex A does not excite the donor and no acceptor emission occurs in the donor channel.
Wlodarczyk et al. proposed a new method, linear unmixing FRET (lux-FRET) approach [24], to accomplish correction for bleed-through, crosstalk and unpaired fluorophores in two well separated steps: First, it takes into account bleed-through in a rigorous way by separating spectral components. Thereby it does not need to restrict data acquisition to well-separated spectral channels, but uses photons from the whole spectrum for the fitting process resulting in better efficiency of photon usage [25]. It should be also noted that the equivalent of the three-cube measurement is readily performed using two filter cubes for two excitations, with a beam splitter, which separates emission into two channels. Secondly, for the tandem sensor constructs simplified imaging modes can be employed to measure lux-FRET. For example, after an initial spectrally resolved dual excitation calibration, the user can perform repetitive single excitation wavelength measurements to quantify Ef D at high temporal resolution. Practically, lux-FRET can be performed on all microscopic devices offering spectral resolution as well as on filter based systems but also on spectrofluorometers.
Excitation-Contraction Coupling
The pumping function of the heart in general and the contraction of the cardiac myocytes in particular are basically characterised by a process called Excitation-Contraction Coupling (ECC). It describes the processing of an electrical signal into the mechanical force [26]. The key player in this mediation is calcium. While the extend of the contraction can be straight forward (optically) recorded by cell length measurements [27], genetically encoded sensors for calcium and voltage will be discussed in dedicated paragraphs below. In similarity the induction or manipulation of ECC by channelrhodopsin will be presented and finally examples from cellular cardiology provided.
GECIs
Genetically Encoded Calcium Indicators (GECI) can be classified as outlined above into two major groups based on their principle of function, these are Circular Permutated Fluorescence Proteins (CPFP) and FRET-based calcium sensors. For a direct comparison of a selection of GECIs with the popular small molecular Ca2+-indicator Fura-2 in cardiomyocytes see Fig. 2. Examples of CPFP based Ca2+ sensors include the families of pericams and GCaMPs. Despite the shortcomings of CPFP-based Ca2+ indicators, recent iteration of these probes, i.e., GCaMP3-6, offer the best signal-to-noise ratios [32]. Very recently new red-shifted GECIs based on the fluorescent proteins mApple (jRGeCO1a) and on mRuby (jRCaMP1’s) were introduced [33] in an attempt to reach the favourable properties of the GCaMP6 series. This goal was partly reached, but smaller maximal fluorescence changes upon Ca2+-binding, a relatively low absorption cross-section and a rather complex intracellular behaviour limit these new probes [33].
Comparison of genetically encoded Ca2+ indicators expressed in adult ventricular myocytes with Fura-2-loaded cells. (A) Examples of Ca2+ transients measured with five different genetically encoded Ca2+ indicators and Fura-2 under otherwise identical experimental conditions (left, distribution of the fluorescence in myocytes 2 days after transduction; right, typical train of electrically evoked global Ca2+ transients). Although the sensor TN-XXL was expressed in cardiac myocytes, it did not report Ca2+ changes. Adult rat ventricular myocytes were isolated as previously described [28] and maintained in optimized culture conditions [29] for 3 days. Measurements were performed with a video-imaging set-up as previously described [30] at 34 °C. The multiplicity of infection (MOI) was in the range of 4 × 104 to 7 × 105 pfu per 1500 cells and determined as depicted in (B). Example of determination of adenoviral dose for the ratiometric pericam. The MOI marked with the red circle was used for virus transduction. (C) Statistical analysis of the signal:noise (S/N) ratio. (D) Statistical analysis of (a) upstroke duration and (b) decay time constant. (C) and (D), The cumulative data for ≈85 cells per indicator from three independent animal preparations. (E) Measurement of the genetically encoded Ca2+ indicator concentration in cardiac myocytes according to a previously described method [31]. (a) Calibration curve based on confocal measurements of solutions of known yellow fluorescent protein (YFP) concentrations. (b) Distribution of relative YFP fluorescence and YC3.6 concentration in virally transduced cardiac myocytes. This figure presents primary experimental results. RFU indicates relative fluorescence units. This figure is reproduced from [14], with permission from Wolters Kluwer
An alternative to single fluorophore-based Ca2+ indicators is the use of ratiometric Ca2+ sensors based on FRET between two different mutants of GFP, similar to the first set of GECI [34, 35]. Both approaches employ Ca2+ sensing by calmodulin (CaM) in combination with the Ca2+-dependent interaction of calmodulin and the CaM-binding peptide M13 from the myosin light chain kinase. In cameleons [35], CaM-M13 is sandwiched between CFP and YFP. In these probes, Ca2+ binding to CaM is translated into alterations of the steric arrangement between CFP and YFP by a twofold mechanism. First, Ca2+ binding to calmodulin itself leads to significant re-arrangement of the EF hands in the molecule. Such steric changes within the molecule result in changes in the CFP-YFP interaction. Nevertheless, the major intramolecular FRET originates from the process of intramolecular interaction of Ca2+-CaM with its binding partner M13. In this process, Ca2+-CaM almost “wraps” around the binding partner, and the entire Ca2+-sensing domain complex drastically changes its arrangement, resulting in a substantial alteration of the CFP-YFP interaction, eventually changing the energy transfer between these two fluorescent proteins (for details see FRET concept above). Additionally, calmodulin-protein interactions might induce cytosolic alterations of the probe properties, such side effects include Ca2+ responsiveness and intracellular mobility. Observations for genetically encoded Ca2+ probes suggest that the Ca2+-sensing domains of the molecules might be either modified by the host cell or altered through protein-protein interactions in the cytosol. There are numerous intracellular interaction partners for calmodulin [36], which include the apo-calmodulin and Ca2+-occupied forms. In both states, calmodulin can in fact serve as both receiver and donor for protein-protein interactions. Apart from changes in the dynamic range and Ca2+ affinity of the Ca2+ probe, such calmodulin-based sensors might even exert signaling effects, both as a receiver and distributor of subcellular signaling pathways. Apart from the calmodulin domain of the sensors, the intramolecular CaM-binding domain M13 has also been reported to interact with endogenous calmodulin [37]. However, this interaction has been abrogated in a new generation of Ca2+ sensors in which the intramolecular calmodulin-M13 interaction has been remodeled so that endogenous calmodulin and CaM cannot bind [38, 39]. A different approach is to avoid employing the ubiquitous calmodulin as a Ca2+ binding domain and to make use of the skeletal and cardiac Ca2+-binding protein Troponin C (TnC) [40, 41]. To transfer Ca2+ binding into fluorescent changes, genetically encoded Ca2+ probes based on TnC use the same approach as calmodulin-based sensors, i.e., FRET between CFP and YFP (or their variants) [37]. In addition to minimized intracellular interference, these indicators also display rapid on and off kinetics because the Ca2+-binding domain (TnC) originates from a very fast Ca2+ read-out system, i.e., the contractile machinery of skeletal and/or cardiac muscle. Early versions of these Ca2+ probes employed TnC from chicken skeletal muscle (TN-L15) and human cardiac muscle (TN-humTnC) and displayed favorable Ca2+ affinities of 1.2 μM and 0.47 μM, respectively [41]. Further molecular evolution yielded molecules with brighter fluorescence, better protein folding (CerTN-L15, Kd = 1.2 μM), and reduced Ca2+ affinities (TN-XL, Kd = 2.5 μM) [40]. Because TnC exhibits significant Mg2+ binding, its Mg2+ affinity was substantially reduced in more advanced versions of these Ca2+ probes, such as TN-XL which is characterized by a relatively high Kd for Ca2+ of 2.5 μM and an unusually high Hill coefficient of 1.7 [40]. With such a Kd, TN-XL preferentially detects the peak of the Ca2+ transients in cardiac myocytes. This restriction to just a part of the signal effectively narrows the apparent time course and reducing the signal-to-noise ratio [14].
GEVIs
Genetically encoded membrane potential sensors, fluorescent protein based voltage sensors or optogenetic voltage reporters are different synonyms for the same kind of membrane potential probes that are termed GEVI (Genetically Encoded Voltage Indicators) throughout this chapter. Beside all varieties throughout the genesis of GEVIs, they all share voltage sensing domains that are based on (or part of) an integral membrane protein. Therefore the GEVIs will be presented according to the different voltage detection principles (and not like the GECIs according to their fluorescence properties). Regarding the detection principle one can discriminate three different types of GEVIs: based on voltage sensitive conformational changes, microbial opsin-based and sensing non-linear optical properties of fluorescent proteins [42]. However, as a general statement, GEVIs are not yet as mature as GECIs.
The first voltage sensors solely comprising genetically encoded proteins (called FlaSh) fused a wtGFP to the C-terminus of the Drosophila Shaker K+-channel [43]. The second GEVI generated independently was based on the fourth transmembrane segment (S4) of the voltage gated K+ channel Kv2.1 coupled to a CFP/YFP FRET pair in sequence and was named VSFP1 [44]. This sensor was followed by a circular permutated version of the fluorescent protein [45]. The third GEVI type was called SPARC and comprised a GFP fused between domains I and II of the rat skeletal muscle Na+ channel [46]. Mainly the lack of distinct membrane localization of these three GEVIs [47] made them fail in biological applications. A new generation of GEVIs comprised self-contained voltage sensor domains, such as the voltage sensing domain of the Ciona intestinalis Voltage Sensor-containing Phosphatase (Ci-VSP) [48], or voltage sensor domain only proteins [49]. The Ci-VSP domain was chosen by two groups of the same institution (Brain Science Institute, RIKEN, Japan) that independently developed what is now termed VSFP2.x [50] and Mermaid [51]. Especially the VSFP2.x went through several optimisation steps like linker design (VSFP2.3) [52] or choice of fluorescent protein (Clover and mRuby2—VSFP-CR) [53]. However, both constructs, namely VSFP2.3 and Mermaid made it in independent studies into cardiac in vivo applications [42, 54, 55], which are compared in Fig. 3. All GEVIs so far share the property of a fast and a slow kinetic response component. While the fast component results from sensing currents within the voltage sensing domain, the slow component is a consequence of the voltage-dependent conformational change in the probe [58]. Seminal work in linker and fluorescent read-out optimization as well as employing voltage sensing domains of voltage-gated phosphatases of other species were performed to shift of the slow sensing component towards faster read-out kinetics [59, 60]. Again these sensors did not make it into biomedical applications because the intensity change per 100 mV voltage change was too low (0.3–0.5%). Based on Mermaid, an improved GEVI was designed using a similar rationale as for the VSFP3.x probes [61] but taking super ecliptic pHluorin [62] as the fluorescent protein. This sensor was named ArcLight and displayed a large fluorescence response of more than 30% per 100 mV voltage change [61, 63]. According to an initial report, the Accelerated Sensor of Action Potentials (ASAP1) is currently the best non-ratiometric GEVI in this group of voltage sensitive fluorescent proteins [64]. It is based on the voltage-sensitive phosphatase of chicken (Gallus gallus) and displays around 29% fluorescence intensity change per 100 mV voltage change [65]. In addition, the kinetic was also advantageous, the activation response of the fast component of 2.1 ± 0.2 ms represented approximately 60% of the total signal amplitude [64].
Transgenic mice expressing a Genetically Encoded Voltage Indicator (GEVI) for optical mapping of the heart. Comparison of mice expressing VSFP2.3 (left, all panels (a)) and Mermaid (right, all panels (b)). Although slightly different parameters are presented, both mice show consistent data. (A) Cardiac appearance of the GEVI expression. (a) View of the excised heart; (b) Cut open heart: left, short axis; right, long axis; (B) Isolated cells expressing the GEVI on the plasmalemma, including T-tubules. (a) Confocal section; (b) 3D reconstruction based on confocal recordings; (C) GEVIs neither alter morphologic nor functional cardiac parameters. (a) Echocardiographic based parameters of different VSFP2.3 mouse lines compared to WT and GCaMP2 mice [56]: top left, heart weight to body weight ratio (HW/BW); top right, fractional area shortening (FAS); bottom left, diastolic left ventricular inner diameter (LVIDd); bottom right, systolic left ventricular inner diameter (LVIDs). None of the mice lines showed any significant differences except for the comparison with GCaMP2 mice (n = 8 mice per genotype); (b) Magnetic resonance imaging based parameters of Mermaid mice compared to WT: top left, left ventricular mass (LVM); top middle, left ventricular stroke volume (LVSV); top right, left ventricular ejection fraction (LVEF); bottom left, heart rate; bottom middle, right ventricular ejection fraction (RVEF). None of the parameters showed significant differences between Mermaid and WT mice (n = 6 mice per genotype); (D) Patch-clamp related measurements in mice expressing GEVI. (a) Left: representative traces of CFP and YFP in response to a voltage step from −70 to +50 mV in cardiomyocytes expressing VSFP2.3. Right: YFP/CFP ratios in response to a voltage protocol as depicted in cardiomyocytes expressing VSFP2.3, the optical signals show a pronounced delay compared to the command voltage as was also shown for Mermaid in cardiomyocytes [57]; (b) Action potential (AP) properties of Mermaid mice compared to WT at a stimulation frequency of 5 Hz: left, AP amplitude; right, AP duration (APD) for 30% and 70% repolarization. None of the parameters showed significant differences between Mermaid and WT mice (n = 10 cells per genotype); (E) Proof-of-principle for Langendorff-perfused heart recordings of mice expressing GEVI. (a) Synchronous electrical cardiograms (ECG) and optical cardiograms (OCG) supplemented with representative images during 10 Hz electrical pacing via a point electrode; (b) Synchronous ECG and raw fluorescence traces (based on the region of interest as indicated by the red rectangle in the relative fluorescence (rel. fl.) image) of an autonomous beating heart (top traces) were subjected to a Fast Fourier Transformation (FFT, left graph). The FFT phase at the frequency of interest (beating frequency of the heart) was visualized for each pixel (bottom right) to map the temporal AP distribution over the heart. Items in the left column (all panels (a)) are reproduced from [54], with permission from Wolters Kluwer and items in the right column (all panels (b)) are reproduced from [42], with permission from MDPI AG
A completely different concept for GEVI design is based on the use of microbial opsins [66] and resulted in the development of sensors named PROBS and Arch [66, 67]. The latter one is derived from the rhodopsin protein, Archaerhodopsin 3 [67]. Microbial opsins bind retinal, a vitamin A-related organic chromophore, and have evolved naturally to function as transducers of light into cellular signals. These proteins are known as tools for optogenetic manipulation [68] (see also below). The natural occurring relationship between light and voltage can be reversed, so that membrane voltage changes are reported as an optical signal. In the initial construct of Arch, the light required for imaging activated a proton current resulting in a contra productive change of the membrane potential. Although a point mutation (D95N) abolished Archs’ capacity to elicit light-driven currents, it also impaired the temporal response [67]. A particular interesting variant of Arch(D95N) is it’s fusion with the GECI GCaMP5G leading to the dual-function Ca2+ and voltage indicator named CaViar [69], which was used to map membrane voltage and calcium in zebrafish heart. The microbial opsin-based GEVIs were improved ever since leading to new versions like QuasAr1 and QuasAr2 [63]. Although the QuasArs display a substantial change in fluorescence intensity per 100 mV change of membrane potential and a fast component of the activation response [63], its overall fluorescence intensity is 30- to 80-fold dimmer than GFP [65]. This might be the reason why QuasArs (at least in our hands) did not work on cardiac myocytes. The combination of fluorescent proteins with a fungal rhodopsin (Leptosphaeria maculans) to perform FRET resulted in the development of MacQ-GEVIs with a good responsiveness of around 20% per 100 mV of voltage change and an activation response of the fast component of 2.2 ± 0.2 ms representing approximately 74% of the total signal [70]. A very similar strategy was performed combining QuasAr2 with various fluorescent proteins from eGFP to mKate2 [65].
All previously described approaches using genetically encoded voltage sensors are based on native voltage sensing proteins that functionally rely on protonation or conformational changes, such as voltage dependent protonation of the retinal Schiff base or voltage dependent phosphatases. Their mechanical action towards conformational changes in the sensing domains induce steric alterations in the fluorescent proteins that are utilized to provoke and subsequently measure changes in fluorescence intensity. A different approach is to explore possible interactions between the membrane potential and the chromophore itself. The Stark effect caused by electric field changes is used in small molecular dye-based voltage sensors, e.g. [71]. However, for chromophores of fluorescent proteins this effect is too small to be detected by fluorescence microscopy. This highlights other properties of chromophores that have hardly been appreciated in the development of biosensors in general. These are the non-linear properties allowing the employment of second harmonic generation (SHG) in response to femtosecond pulsed infrared light. The general concept [72, 73] and initial attempts [73, 74] are summarized in [75] but they were not explored further.
Manipulation by Channelrhodopsins
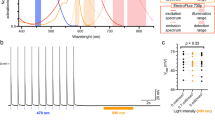
In contrast to the previous sections where light was used to read-out information, channelrhodopsins are proteins that translate light signals into control of cellular function. In some respect this is the genetically encoded variant of caged compounds as outlined in the chapter ‘Caged Compounds: Applications in Cardiac Muscle Research’ by Niggli and Shirokova within this book. Although there has been a strong interest in light sensitive proteins for decades, the break through came with the discovery and cloning of a faster type of microbial opsins, the channelrhodopsins [6, 7]. Similar as for the genetically encoded indicators initial utilisation and methodological development happened in the field of neuroscience [76]. Most cardiac applications of channelrhodopsins so far are based on the manipulation of tissue or the whole heart and are rather combined with electrical read-outs like multi electrode arrays (MEA) [77,78,79]. Both approaches are not in the focus of this book that sets the emphasis on microscopic methods. However, we investigated isolated adult cardiomyocytes of mice, where channelrhodopsin-2 was cardiac specific expressed (Fig. 4). As the images show, a 1 ms illumination of light induced contractions in both cells depicted, which shows that the activation of the channelrhodopsin induced an action potential in these cells. At the same time it becomes evident that illumination (as required for imaging approaches) with almost the entire visible spectrum (380–660 nm tested) induces constant openings of the channelrhodopsin. This prevented us from using the optical stimulation in combination with fluorescent probes, e.g., GECI and GEVI as described above.
Adult isolated cardiomyocytes from Chop2YFP mice [80] cross-breeded with a cardiac specific Cre mouse line [81] expressing Channelrhodopsin-2 and YFP were electrically paced as previously described [27] (field stimulation, control) or stimulated with 1 ms light pulses of the given wavelength in a microscopy platform (more, TILL Photonics, Germany) equipped with a monochromator (Polychrome V, TILL Photonics, Germany). For each recording the upper image depicts the cardiomyocytes at rest, whereas the lower one shows the maximal contraction. The graphs show the contractions with the colour codes introduced in the image of the control condition. The contraction of electrically paced cells is less pronounced as the light induced ones. This is caused by the fact that electrical pacing was in a steady state, while optical stimulation reflects post rest potentiation
In light of these properties the development of novel variants of spectrally shifted light gated ion channels, like C2-HR [82] or CheRiff [63], is reasonable. Especially hypochromic shifted spectral properties would open the green and red spectral range for the use of genetically encoded sensors as described above with the goal to perform an all optical observation of ECC from the cellular stimulation to the read-out of action potentials, Ca2+-signals and cellular contraction. Such an approach has recently been performed on human stem-cell derived cardiomyocytes as a proof of principle for cardiotoxicity screens (without cell length recordings) [83]. In this study of electically connected cells in culture, only a subset of cells expressed the channelrhodopsin variant CheRiff while another subset of cells expressed the combined Ca2+/voltage sensor CaViar. Such approaches need specialised microscopes as outlined in the chapter ‘Optical Sectioning Microscopy at ‘Temporal Super-Resolution’’ within this book.
Examples in Cellular Cardiology
Ca2+-Signals in the Nanodomains of Cardiac Myocytes
Although a classical application of genetically encoded Ca2+ indicators, the question of Ca2+ concentrations in nanodomains near Ca2+ transport proteins or in the fuzzy space has only recently been addressed following the introduction of a novel series of CPFPs, the GCaMP6s [32]. These kind of measurements are a functional complementation to the structural superresolution measurements as described in the chapter ‘Quantitative Super-Resolution Microscopy of Cardiomyocytes’ within this book. Based on the fastest version of the latest GECIs (GCaMP6f), C-terminal fusion proteins of tradin1 and junctin, GCaMP6f-T and GCaMP6f-J, respectively, allowed Ca2+ measurements exclusively in the dyadic cleft [84]. The recorded signals were termed ‘Ca2+ nanosparks’, because their calculated volume was approximately 50 times smaller than that of Ca2+ sparks. Although the focus of that paper [84] was methodology, the peak intensity (F/Fo) of the nanotransients was 7.8-fold higher than that due to global Ca2+ elevation.
Characterisation of the Cellular Cardiac Action Potential Duration and Shape
Conceptual studies of cardiotoxicity screens based on GEVI have been performed with ‘Mermaid’ [55, 85]. Mermaid compares well with small molecule dyes such as RH-237 or di-8-ANEPPS [86]—we even noticed a 25% higher change of the relative fluorescence ratio compared to the ratiometric read-out mode of di-8-ANEPPS [87], although the temporal kinetics of Mermaid was much slower than these chemical sensors. In adult cardiomyocytes, pharmaceutical prolongation of the action potential could be detected readily [85]. This prolongation of the action potential duration can be regarded as a cellular equivalent of the QT-interval prolongation in the ECG, which is a pro-arrythmogenic indicator [88]. Differentiating cardiomyocytes from embryonic or induced Pluripotent Stem cells (iPS-cells) for cardiac safety screens as well as for basic research is becoming increasingly popular [89, 90]. In a proof of principle study the combination of human iPS-cells and optogenetics has been successfully performed [83]. However, the stem cell derived cardiomyocytes contain a mixture of different phenotypes, like ventricular myocytes, atrial myocytes or myocytes of the conduction system. All subtypes have a different gene expression, but are morphologically indistinguishable. For the experimental design as well as for further differentiation, it is desirable to purify or just identify a particular subtype of cardiomyocytes. A method to discriminate the cell type is the shape of their action potential, which is characteristic for the subtypes mentioned above [91]. An elegant way to measure such an action potential is by means of a GEVI, was shown for ArcLight and VSFP-CR [42, 92], Fig. 5. Subtype-specific promoters are yet another way to measure such an action potential [94].
Voltage-Sensitive Fluorescent Protein Clover-mRuby2 (VSFP-CR) for phenotyping stem cell derived cardiomyocytes. (A) Overview of optically recorded (Di-8-ANEPPS) reference action potential phenotypes from induced pluripotent stem cell derived cardiomyocytes. This panel is reproduced from [93], with permission from John Wiley & Sons; (B) Recorded image series of a human stem cell derived cardiomyocyte expressing VSFP-CR (Lentiviral gene transfer). The false color ratio images are snapshots every 100 ms of a time series recorded at 500 frames per second using a scientific Complementary Metal-Oxide-Semiconductor (sCMOS) camera and point to the time course of the recorded action potential. Considering the temporal response of the GEVI (Genetically Encoded Voltage Indicators), the example shows most alikeness with an “intermediate” action potential with a tendency to the ventricular phenotype. Overlay of the raw ratio trace (black) and a smoothed trace (blue); (C) Original and processed traces of a train of recorded action potentials of an electrically paced (1 Hz) stem cell derived cardiomyocyte. (a) Raw traces of the spectral channels for Clover and mRuby2; (b) Overlay of the raw ratio trace (black) and a smoothed trace (blue). This figure is reproduced from [42], with permission from MDPI AG
Summary
Given the advantages and drawbacks discussed above, there are numerous applications for which GECI and GEVI could prove advantageous, particularly in light of the latest developments of GECI with increased sensitivity [32] and the large momentum, GEVI design and characterization has gained in recent years [42]. However, the selection of the appropriate genetically encoded indicator for a particular application remains challenging. In any case, the intrinsic properties of genetically encoded indicators have to match the dynamics of cardiac myocytes or their subcellular localization to obtain the best results.
The development of GECI and GEVI is far from being at its endpoint. Future research will lead to increased fluorescence yield, enlarged dynamic ranges, and faster sensors. The latter requirement seems to be essential to enable upstroke-based analysis methods of fast confocal recordings, as recently described for Ca2+ measurements [95, 96]. Furthermore, biocompatibility must be improved, particularly by reducing cross-talk with endogenous signalling pathways. There is also potential for broad expansion of the spectral range of genetically encoded indicators. There is still a particular demand for indicators with spectral properties shifted into the red or far-red spectral band and at the same time remaining in a limited spectral band. The indicators with bathochromic shifted wavelengths reduce the contribution of autofluorescence and enable better light penetration into the tissue, which is particularly important for in vivo imaging approaches with cellular resolution, while the distinct spectral width allows the combination of several biosensors. This, in the first place refers to the combination of GECI and GEVI, but also to other types of measurements, such as phosphorylation sensors [97, 98].
Beyond the future developments of optogenetics, their potential field of application is far from being fully exploited. Although the development of GECIs, GEVIs and channelrhodopsins was initially led by applications in neuroscience [99], the examples presented here on cardiac cells pave the way for an application in cardiac topics. It is the responsibility of researchers in the field of cardiovascular sciences to bridge the gap between the availability of optogenetic tools and their appropriate application in cardiovascular studies. We face the situation of a delay between the introduction of the genetic tools and their application, because of intermediate steps, including the generation of transgenic animals or viruses for gene transfer [8] and the establishment of a reproducible and robust read-out mode. In this context, we can expect that recently introduced indicators, novel sensors and light gated ion channels to be developed will replace the optogenetic tools so far used.
References
Swammerdam J. Bybel der natuur. London: C. G. Seyffert; 1737.
Scientists, A. C. O. D. The collected letters of Antoni Van Leeuwenhoek. Boca Raton, FL: CRC Press; 1996.
Milestones in light microscopy. Nat Cell Biol. 2009;11:1165.
Shimomura O, Johnson FH, Saiga Y. Extraction, purification and properties of aequorin, a bioluminescent protein from the luminous hydromedusan. Aequorea J Cell Compar Physiol. 1962;59:223–39.
Zimmer M. Glowing genes: a revolution in biotechnology. New York, NY: Prometheus Books; 2005.
Nagel G, et al. Channelrhodopsin-1: a light-gated proton channel in green algae. Science. 2002;296:2395–8.
Nagel G, et al. Channelrhodopsin-2, a directly light-gated cation-selective membrane channel. Proc Natl Acad Sci U S A. 2003;100:13940–5.
Kaestner L, Scholz A, Lipp P. Conceptual and technical aspects of transfection and gene delivery. Bioorg Med Chem Lett. 2015;25:1171–6.
Hu CD, Chinenov Y, Kerppola TK. Visualization of interactions among bZIP and Rel family proteins in living cells using bimolecular fluorescence complementation. Mol Cell. 2002;9:789–98.
Wang Q, Shui B, Kotlikoff MI, Sondermann H. Structural basis for calcium sensing by GCaMP2. Structure. 2008;16:1817–27.
Tsien RY. The green fluorescent protein. Annu Rev Biochem. 1998;67:509–44.
Coutinho V, Mutoh H, Knöpfel T. Functional topology of the mossy fibre-granule cell--Purkinje cell system revealed by imaging of intrinsic fluorescence in mouse cerebellum. Eur J Neurosci. 2004;20:740–8.
Díez-García J, Akemann W, Knöpfel T. In vivo calcium imaging from genetically specified target cells in mouse cerebellum. Neuroimage. 2007;34:859–69.
Kaestner L, et al. Genetically encoded Ca2+ indicators in cardiac myocytes. Circ Res. 2014;114:1623–39.
Förster T. Intermolecular energy migration and fluorescence. Ann Phys. 1948;437:55–75.
Tsien RY, Bacskai BJ, Adams SR. FRET for studying intracellular signalling. Trends Cell Biol. 1993;3:242–5.
Heim R, Tsien RY. Engineering green fluorescent protein for improved brightness, longer wavelengths and fluorescence resonance energy transfer. Curr Biol. 1996;6:178–82.
Youvan DC. Calibration of fluorescence resonance energy transfer in microscopy using genetically engineered GFP derivatives on nickel chelating beads. Biotechnol Alia. 2006;3:1–18. https://doi.org/10.1234/12345678.
Berney C, Danuser G. FRET or no FRET: a quantitative comparison. Biophys J. 2003;84:3992–4010.
Gordon GW, Berry G, Liang XH, Levine B, Herman B. Quantitative fluorescence resonance energy transfer measurements using fluorescence microscopy. Biophys J. 1998;74:2702–13.
Xia Z, Liu Y. Reliable and global measurement of fluorescence resonance energy transfer using fluorescence microscopes. Biophys J. 2001;81:2395–402.
Hoppe A, Christensen K, Swanson JA. Fluorescence resonance energy transfer-based stoichiometry in living cells. Biophys J. 2002;83:3652–64.
Lakowicz JR. Principles of fluorescence spectroscopy. New York, NY: Springer Science & Business Media; 2013.
Wlodarczyk J, et al. Analysis of FRET signals in the presence of free donors and acceptors. Biophys J. 2008;94:986–1000.
Thaler C, Koushik SV, Blank PS, Vogel SS. Quantitative multiphoton spectral imaging and its use for measuring resonance energy transfer. Biophys J. 2005;89:2736–49.
Bers DM. Cardiac excitation-contraction coupling. Nature. 2002;415:198–205.
Viero C, Kraushaar U, Ruppenthal S, Kaestner L, Lipp P. A primary culture system for sustained expression of a calcium sensor in preserved adult rat ventricular myocytes. Cell Calcium. 2008;43:59–71.
Kaestner L. et al. Isolation and genetic manipulation of adult cardiac myocytes for confocal imaging. J Vis Exp. 2009; (31).
Tian Q, et al. Functional and morphological preservation of adult ventricular myocytes in culture by sub-micromolar cytochalasin D supplement. J Mol Cell Cardiol. 2012;52:113–24.
Pahlavan S, et al. Gαq and Gα11 contribute to the maintenance of cellular electrophysiology and Ca2+ handling in ventricular cardiomyocytes. Cardiovasc Res. 2012;95:48–58.
Kang M, Walker J. Protein kinase C delta and epsilon mediate positive inotropy in adult ventricular myocytes. J Mol Cell Cardiol. 2005;38:753–64.
Chen T-W, et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature. 2013;499:295–300.
Dana H, et al. Sensitive red protein calcium indicators for imaging neural activity. Elife. 2016;5:413.
Romoser VA, Hinkle PM, Persechini A. Detection in living cells of Ca2+-dependent changes in the fluorescence emission of an indicator composed of two green fluorescent protein variants linked by a calmodulin-binding sequence. J Biol Chem. 1997;272:13270–4.
Miyawaki A, et al. Fluorescent indicators for Ca2+ based on green fluorescent proteins and calmodulin. Nature. 1997;388:882–7.
Cox JA. Calcium-calmodulin interaction and cellular function. J Cardiovasc Pharmacol. 1986;8(Suppl 8):S48–51.
Garaschuk O, Griesbeck O, Konnerth A. Troponin C-based biosensors: a new family of genetically encoded indicators for in vivo calcium imaging in the nervous system. Cell Calcium. 2007;42:351–61.
Palmer AE, Jin C, Reed JC, Tsien RY. Bcl-2-mediated alterations in endoplasmic reticulum Ca2+ analyzed with an improved genetically encoded fluorescent sensor. Proc Natl Acad Sci U S A. 2004;101:17404–9.
Palmer AE, et al. Ca2+ indicators based on computationally redesigned calmodulin-peptide pairs. Chem Biol. 2006;13:521–30.
Mank M, et al. A FRET-based calcium biosensor with fast signal kinetics and high fluorescence change. Biophys J. 2006;90:1790–6.
Heim N, Griesbeck O. Genetically encoded indicators of cellular calcium dynamics based on troponin C and green fluorescent protein. J Biol Chem. 2004;279:14280–6.
Kaestner L, et al. Genetically encoded voltage indicators in circulation research. Int J Mol Sci. 2015;16:21626–42.
Siegel MS, Isacoff EY. A genetically encoded optical probe of membrane voltage. Neuron. 1997;19:735–41.
Sakai R, Repunte-Canonigo V, Raj CD, Knöpfel T. Design and characterization of a DNA-encoded, voltage-sensitive fluorescent protein. Eur J Neurosci. 2001;13:2314–8.
Knöpfel T, Tomita K, Shimazaki R, Sakai R. Optical recordings of membrane potential using genetically targeted voltage-sensitive fluorescent proteins. Methods. 2003;30:42–8.
Ataka K, Pieribone VA. A genetically targetable fluorescent probe of channel gating with rapid kinetics. Biophys J. 2002;82:509–16.
Baker BJ, et al. Three fluorescent protein voltage sensors exhibit low plasma membrane expression in mammalian cells. J Neurosci Methods. 2007;161:32–8.
Murata Y, Iwasaki H, Sasaki M, Inaba K, Okamura Y. Phosphoinositide phosphatase activity coupled to an intrinsic voltage sensor. Nature. 2005;435:1239–43.
Ramsey IS, Moran MM, Chong JA, Clapham DE. A voltage-gated proton-selective channel lacking the pore domain. Nature. 2006;440:1213–6.
Dimitrov D, et al. Engineering and characterization of an enhanced fluorescent protein voltage sensor. PLoS One. 2007;2:e440.
Tsutsui H, Karasawa S, Okamura Y, Miyawaki A. Improving membrane voltage measurements using FRET with new fluorescent proteins. Nat Methods. 2008;5:683–5.
Mutoh H, et al. Spectrally-resolved response properties of the three most advanced FRET based fluorescent protein voltage probes. PLoS One. 2009;4:e4555.
Lam AJ, et al. Improving FRET dynamic range with bright green and red fluorescent proteins. Nat Methods. 2012;9:1005–12.
Chang Liao M-L, et al. Sensing cardiac electrical activity with a cardiac myocyte--targeted optogenetic voltage indicator. Circ Res. 2015;117:401–12.
Tsutsui H, Higashijima S-I, Miyawaki A, Okamura Y. Visualizing voltage dynamics in zebrafish heart. J Physiol (Lond). 2010;588:2017–21.
Tallini YN, et al. Imaging cellular signals in the heart in vivo: cardiac expression of the high-signal Ca2+ indicator GCaMP2. Proc Natl Acad Sci U S A. 2006;103:4753–8.
Tian Q, et al. Optical action potential screening on adult ventricular myocytes as an alternative QT-screen. Cell Physiol Biochem. 2011;27:281–90.
Villalba-Galea CA, et al. Charge movement of a voltage-sensitive fluorescent protein. Biophys J. 2009;96:L19–21.
Lundby A, Mutoh H, Dimitrov D, Akemann W, Knöpfel T. Engineering of a genetically encodable fluorescent voltage sensor exploiting fast Ci-VSP voltage-sensing movements. PLoS One. 2008;3:e2514.
Baker BJ, et al. Genetically encoded fluorescent voltage sensors using the voltage-sensing domain of Nematostella and Danio phosphatases exhibit fast kinetics. J Neurosci Methods. 2012;208:190–6.
Jin L, et al. Single action potentials and subthreshold electrical events imaged in neurons with a fluorescent protein voltage probe. Neuron. 2012;75:779–85.
Miesenbock G, De Angelis DA, Rothman JE. Visualizing secretion and synaptic transmission with pH-sensitive green fluorescent proteins. Nature. 1998;394:192–5.
Hochbaum DR, et al. All-optical electrophysiology in mammalian neurons using engineered microbial rhodopsins. Nat Methods. 2014;11:825–33.
St-Pierre F, et al. High-fidelity optical reporting of neuronal electrical activity with an ultrafast fluorescent voltage sensor. Nat Neurosci. 2014;17:884–9.
Zou P, et al. Bright and fast multicoloured voltage reporters via electrochromic FRET. Nat Commun. 2014;5:4625.
Kralj JM, Hochbaum DR, Douglass AD, Cohen AE. Electrical spiking in Escherichia coli probed with a fluorescent voltage-indicating protein. Science. 2011;333:345–8.
Kralj JM, Douglass AD, Hochbaum DR, Maclaurin D, Cohen AE. Optical recording of action potentials in mammalian neurons using a microbial rhodopsin. Nat Methods. 2012;9:90–5.
Ernst OP, et al. Photoactivation of channelrhodopsin. J Biol Chem. 2008;283:1637–43.
Hou JH, Kralj JM, Douglass AD, Engert F, Cohen AE. Simultaneous mapping of membrane voltage and calcium in zebrafish heart in vivo reveals chamber-specific developmental transitions in ionic currents. Front Physiol. 2014;5:344.
Gong Y, Wagner MJ, Zhong Li J, Schnitzer MJ. Imaging neural spiking in brain tissue using FRET-opsin protein voltage sensors. Nat Commun. 2014;5:3674.
Kuhn B, Fromherz P, Denk W. High sensitivity of stark-shift voltage-sensing dyes by one- or two-photon excitation near the red spectral edge. Biophys J. 2004;87:631–9.
Bublitz G, King B, Boxer S. Electronic structure of the chromophore in green fluorescent protein. J Am Chem Soc. 1998;120:9370.
Rosell FI, Boxer SG. Polarized absorption spectra of green fluorescent protein single crystals: transition dipole moment directions. Biochemistry. 2003;42:177–83.
Khatchatouriants A, Lewis A, Rothman Z, Loew L, Treinin M. GFP is a selective non-linear optical sensor of electrophysiological processes in Caenorhabditis elegans. Biophys J. 2000;79:2345–52.
Kaestner L, Tian Q, Lipp P. In: Jung G, editor. Action potentials in heart cells. New York, NY: Springer; 2012. p. 163–82.
Entcheva E. Cardiac optogenetics. AJP Heart Circ Physiol. 2013;304:H1179–91.
Bruegmann T, et al. Optogenetic control of heart muscle in vitro and in vivo. Nat Methods. 2010;7:897–900.
Vogt CC, et al. Systemic gene transfer enables optogenetic pacing of mouse hearts. Cardiovasc Res. 2015;106:338–43.
Ambrosi CM, Klimas A, Yu J, Entcheva E. Cardiac applications of optogenetics. Prog Biophys Mol Biol. 2014;115:294–304.
Scheller A, Bai X, Kirchhoff F. The role of the oligodendrocyte lineage in acute brain trauma. Neurochem Res. 2017; https://doi.org/10.1007/s11064-017-2343-4.
Wiesen K, et al. Cardiac remodeling in Gαq and Gα11 knock out mice. Int J Cardiol. 2016;202:836–45.
Prigge M, et al. Color-tuned channelrhodopsins for multiwavelength optogenetics. J Biol Chem. 2012;287:31804–12.
Dempsey GT, et al. Cardiotoxicity screening with simultaneous optogenetic pacing, voltage imaging and calcium imaging. J Pharmacol Toxicol Methods. 2016;81:240. https://doi.org/10.1016/j.vascn.2016.05.003.
Shang W, et al. Imaging Ca2+ nanosparks in heart with a new targeted biosensor. Circ Res. 2014;114(3):412. https://doi.org/10.1161/CIRCRESAHA.114.302938.
Tian Q, et al. Optical measurement of action potential in adult ventricular myocytes. Biophys J. 2011;100:292a.
Haugland RP. Handbook of fluorescent probes and research products. Eugene, OR: Molecular Probes; 2002.
Kaestner L, Tian Q, Lipp P. Cardiac safety screens: molecular, cellular, and optical advancements. In: Lin CP, Ntziachistos V, editors. Biomedical optics III, vol. 8089. Munich: SPIE; 2011. p. 80890H-1–6.
Arrigoni C, Crivori P. Assessment of QT liabilities in drug development. Cell Biol Toxicol. 2007;23:1–13.
Sinnecker D, et al. Induced pluripotent stem cells in cardiovascular research. Rev Physiol Biochem Pharmacol. 2012;163:1. https://doi.org/10.1007/112_2012_6.
Matsa E, Burridge PW, Wu JC. Human stem cells for modeling heart disease and for drug discovery. Sci Transl Med. 2014;6:239ps6.
Dorn T, et al. Direct Nkx2-5 transcriptional repression of Isl1 controls cardiomyocyte subtype identity. Stem Cells. 2015;33:1113. https://doi.org/10.1002/stem.1923.
Leyton-Mange JS, et al. Rapid cellular phenotyping of human pluripotent stem cell-derived cardiomyocytes using a genetically encoded fluorescent voltage sensor. Stem Cell Rep. 2014;2:163–70.
Nagai T, Sawano A, Park ES, Miyawaki A. Circularly permuted green fluorescent proteins engineered to sense Ca2. Proc Natl Acad Sci U S A. 2001;98:3197–202.
Chen Z, et al. Subtype-specific promoter-driven action potential imaging for precise disease modelling and drug testing in hiPSC-derived cardiomyocytes. Eur Heart J. 2017;38:292–301.
Tian Q, Kaestner L, Lipp P. Noise-free visualization of microscopic calcium signaling by pixel-wise fitting. Circ Res. 2012;111:17–27.
Tian Q, Kaestner L, Schröder L, Guo J, Lipp P. An adaptation of astronomical image processing enables characterization and functional 3D mapping of individual sites of excitation-contraction coupling in rat cardiac muscle. Elife. 2017;6:665.
Violin JD, Zhang JX, Tsien RY, Newton AC. A genetically encoded fluorescent reporter reveals oscillatory phosphorylation by protein kinase C. J Cell Biol. 2003;161:899–909.
Schleifenbaum A, Stier G, Gasch A, Sattler M, Schultz C. Genetically encoded FRET probe for PKC activity based on pleckstrin. J Am Chem Soc. 2004;126:11786–7.
Knöpfel T, Gallero-Salas Y, Song C. Genetically encoded voltage indicators for large scale cortical imaging come of age. Curr Opin Chem Biol. 2015;27:75–83.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Kaestner, L., Zeug, A., Tian, Q. (2018). Optogenetic Tools in the Microscopy of Cardiac Excitation-Contraction Coupling. In: Kaestner, L., Lipp, P. (eds) Microscopy of the Heart. Springer, Cham. https://doi.org/10.1007/978-3-319-95304-5_5
Download citation
DOI: https://doi.org/10.1007/978-3-319-95304-5_5
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-95302-1
Online ISBN: 978-3-319-95304-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)