Abstract
The major problem when analyzing a metagenomic sample is to taxonomically annotate its reads in order to identify the species and their relative abundances. Many tools have been developed recently, however they are not always adequate for the increasing database volume. In this paper we propose an efficient method, called SKraken, that combines taxonomic tree and k-mers frequency counting. SKraken extracts the most representative k-mers for each species and filter out less representative ones. SKraken is inspired by Kraken, which is one of the state-of-art methods. We compare the performance of SKraken with Kraken on both real and synthetic datasets, and it exhibits a higher classification precision and a faster processing speed. Availability: https://bitbucket.org/marchiori_dev/skraken.
Access provided by CONRICYT-eBooks. Download conference paper PDF
Similar content being viewed by others
1 Introduction
Metagenomics is a study of the heterogeneous microbes samples (e.g. soil, water, human microbiome) directly extract from the natural environment with the primary goal of determining the taxonomical identity of the microorganisms residing in the samples. It is an evolutionary revise, shifting focuses from the individual microbe study to a complex microbial community. As already mentioned in [1, 2], the classical genomic-based approaches require the prior clone and culturing for the further investigation. However, not all bacteria can be cultured. The advent of metagenomics succeeded to bypass this difficulty.
The study of metagenomics is far more than just about labeling the species of the microbes. The analysis can reveal the presence of unexpected bacteria and viruses in a microbial sample, also allows the identification and characterization of bacterial and viral genomes at a level of detail not previously possible. For example, in the case of the human body, imbalances in the microbiome are related with many diseases, e.g. inflammatory bowel disease (IBD) [3] and colorectal cancer [4]. Furthermore it may aid the researchers to systematically understand and characterize the microbial communities: how genes influence each others activities in serving collective functions; grasp the collections of communities that composites the biosphere where the humans be a part, etc. It has already been applied in many areas like ecology, medicine, microbiology and some others mentioned in [3,4,5,6].
In this paper, we focus on the taxonomic classification, one branch of the metagenomics investigation, with a referenced database. Generally speaking, two techniques could be in consideration for the sake of classification task: (1) sequencing phylogenetic marker genes, e.g., 16S rRNA; (2) next generation sequencing (NGS) of all the genomic material in the sample. The use of marker genes requires amplification steps that can introduce bias in the taxonomic analysis. Moreover, not all bacteria can be identified by traditional 16S sequencing for its divergent gene sequences [7]. Indeed, We believe that the most effective and unbiased method to study microbial samples is via whole-genome NGS. However, the short length of NGS reads poses a number of challenges to the correctness of taxonomical classification for each read. Several methods and software tools are already available, but with the increasing throughput of modern sequencing technologies, the faster and more accurate algorithms are needed. These methods can be broadly divided into three categories: (1) sequence-similarity-based methods, (2) marker-based methods where certain specific marker sequences are used to identify the species. (3) sequence-composition-based methods, which are based on the nucleotide composition (e.g. k-mers usage).
The sequence-similarity-based methods work as searching reads according to the sequence similarity in the reference database, the popular examples are MegaBlast [8] and Megan [9]. They precisely identify reads from genomes within the reference database, while they are generally very slow, especially compared with composition-based methods. Marker-based methods try to use phylogenetic marker genes as a taxonomic reference [10,11,12]. For example, MetaPhlAn [12] is based on marker genes that are clade specific.
The fastest and most promising approaches belong to the composition-based one. Its trait can be summarized as follows: the genomes of reference organisms are modeled based on k-mers counts; the reads are searched throughout the reduced-version database. The most representative methods located within this category are Kraken [13], Clark [14] and Lmat [15]. As for the precision of these methods, they are as good as MegaBlast [8] (similarity-based method), nevertheless the processing speed is much faster. Thus, these methods are really capable to keep pace with the increasing throughput of modern sequencing instruments.
Recently the paper [16] has shown that Kraken [13] is one of the most promising tool in terms of classifying correctness and speed. It owes to the construction of the reference database, where each genome is expressed by k-mers (a piece of genome with length k), and the a taxonomic tree. More precisely, Kraken constructs a data structure that is an augmented taxonomic tree in which a list of significant k-mers is associated to each node, leafs and internal nodes. Given a node on this taxonomic tree, its list of k-mers is considered representative for the taxonomic label and will be used for the classification of metagenomic reads.
Inspired by this paradigm, in this paper we propose SKrakenFootnote 1, a tool for metagenomics reads classification that selects the most representative k-mers for each node in the taxonomic tree, though, filtering out uninformative k-mers. The main properties of SKraken can be profiled as: (i) an efficient detection of representative k-mers over the taxonomic tree; (ii) SKraken improves the precision over Kraken on both simulated and real metagenomic datasets without compromising the recall; (iii) benefit from the downsized database. As a consequence, SKraken requires less memory RAM and the classification speed increases w.r.t. Kraken. In the next section we will give an overview of Kraken and analyze how to improve the classification. SKraken is presented in Sect. 2.2. Both tools are tested on simulated and real metagenomic datasets in Sect. 3 and the conclusions are drawn in Sect. 4.
2 Method
2.1 An Overview of Kraken
In order to better understand our contribution let’s briefly present Kraken in the first place. Instead of utilizing the complete genome as reference, Kraken considers only its k-mers, as well as many other tools [14, 15], thus a genome sequence is alternatively represented by a set of k-mers, which plays a role of efficiently indexing a large volume of target-genomes database, e.g., all the genomes in RefSeq. This idea is stemmed from alignment-free methods [18] and some researchers have verified its availability in different applications. For instance, the construction of phylogenetic trees, traditionally is performed based on a multiple-sequence alignment technique, the process is be carried out on the whole genomes [19, 20], in practice it’s difficult to realize due to the improper length of whole genomes. The alignment-free technique with the usage of k-mers is the solver. Recently some variations of k-mers-based methods have been devised for the detection of enhancers in ChIP-Seq data [21,22,23,24,25] and entropic profiles [26, 27]. Recently, the assembly-free comparison of genomes and metagenomes based on NGS reads and k-mers counts has been investigated in [28,29,30,31]. For a comprehensive review of alignment-free measures and applications we refer the reader to [18].
Considering the taxonomic tree, taken from the complete NCBI taxonomic information, this data structure is extended by annotating each node with k-mers, including leaves and internal nodes. Every node is associated with a list of k-mers that are critical for the future classification. More precisely, given a dataset of target genomes, the construction of this annotated taxonomic tree is carried out by scanning the k-mers of each genome in the dataset. If the k-mer appears only in a given genome, it is associated only to one leaf node J (representing that genome) and the k-mers list of node J is updated. If the k-mer appears in more than one species, the k-mer is erased from those corresponding nodes and moved to the lowest common ancestor of these nodes, see Fig. 1 for an example. At the end of this step each k-mer belongs to only one node in the taxonomic tree.

(figure taken from [17]).
In this example the k-mer AGCCT, that is contained in the species 9 and 13, is moved to the lowest common ancestor, the family node 2

(figure taken from [13]).
An overview of the metagenomic reads classification of Kraken
Once this database of annotated k-mers has been constructed, Kraken can classify reads in a very efficient manner. Figure 2 reports an overview of the classification process. Later when we launch the classification procedure, with a given read, Kraken firstly decomposes the read into a list of its k-mers. Then each k-mer is searched in the augmented taxonomic tree, whenever it hits we increase the counter of the corresponding node. After all k-mers have been analyzed, we will classify the read by searching the highest-weight path in the taxonomic tree, from top to button.
2.2 SKraken: Selecting Informative k-mers
The most important step of Kraken is the construction of the augmented taxonomic tree, annotating by a list of k-mers per node in order to implement the classification. In this paper we propose SKraken that has an exclusive procedure, distinguishable from Kraken, we select and filter k-mers instead. Therefore, those uninformative k-mers will be pruned away from the augmented taxonomic tree.
It may occur that one k-mers appears in more than one species, like demonstrated in Fig. 1, sequence “AGCCT” belongs to two species. Kraken adds this k-mers into the ancestor node 2 (representing a taxonomic family) out of node 9 and node 13. Since this k-mers will be used in the classification step, we would like to be informative for the family node 2. However, the majority of species in this family, nodes 10, 11 and 12, do not contain this k-mers.

(figure taken from [17]).
An example of quality score q(GAACT)
To address this issue, for each k-mer, we define a scoring function that captures its representativeness within the taxonomic node. We recall that a k-mer is associated with only one node in the tree. Let’s define TaxID(m) that indicates the taxonomic node associated with the k-mer m. However, before the upward shift of k-mer, it is possible to occur in more than one species. We define NumSpecies(m) as the number of species that contains m. By construction TaxID(m) is the lowest common ancestor of all these species. Thus the species in which m appears, they are all leaf nodes of the subtree TaxID(m). We define TotSpecies(n) as the total number of species in the subtree routed in the node n. With these values we define q(m) the quality of a k-mer m as (equation from [17]):
Figure 3 shows an example of the quality q(GAACT). The quality of GAACT can also be interpreted as the percentage of species nodes that contains GAACT, i.e. NumSpecies(GAACT), with respect to the family node 2, i.e., TaxID(GAACT), in this case \(60\%\). Similarly, if we consider the example in Fig. 1, the quality of \(q(AGCCT)\,=\,\frac{NumSpecies(AGCCT)}{TotSpecies(TaxID(AGCCT))}\,=\,\frac{2}{5}\,=\,0.4\), that is \(40\%\). We can conclude as, a k-mer has an high quality can be considered representative for a given taxonomic node and the relative sub-tree, thus more likely will be informative for the classification. Based on these observations SKraken tries to screen out the uninformative k-mers by means of their quality value, and prunes the augmented taxonomic tree by removing the k-mers with a quality below a given threshold Q.
In order to compute the quality scores q(m) for all k-mers we have to evaluate NumSpecies(m) and TotSpecies(n) efficiently. Here is our setting: the construction of the augmented taxonomic tree of SKraken is divided into two steps. In the first step, given a set of target genomes, we scan the k-mers of each genome and build the augmented taxonomic tree, similarly to Kraken. Meanwhile, we set a variable to keep tracking NumSpecies(m), as soon as m is found in a new species we increment this variable. However, there can be genomes that are further classified as sub-species of a given species node. In order to compute the correct value of NumSpecies(m), we need to make sure that all genomes of a given species are processed before moving to next species. This can be tackled by scanning the input genomes in a particular order so that all genomes of a species and the sub-species, are processed at once. Another problem is the fact that a k-mer can appear in some sub-species of a given species node. When computing NumSpecies(m) we must ensure not to overcount these occurrences, and thus the corresponding variable is incremented only when m is found for the first time in a given species. All other occurrences of m within the same species will be discarded. At the end of the first step we have computed the augmented taxonomic tree, with all k-mers, and the corresponding values NumSpecies(m).
In the second step SKraken computes the quality values q(m) and filters uninformative k-mers. The number of leaf nodes that are the descendants of n, indicated as TotSpecies(n), can be obtained for all nodes in the tree with a post-order traversal of the taxonomic tree. Consequently all k-mers are processed and the corresponding qualities q(m) are computed. If q(m) is below a given input parameter Q, m is removed from the database.
Note that the size of the taxonomic tree is constant and much smaller with respect to the number of k-mers. The overall process depends only on the total number of k-mers and its size is linear to the input reference genomes. Once the augmented taxonomic tree is build, reads can be classified with the same procedure of Kraken.
3 Results
3.1 Datasets
Before the demonstration of result, we need to build a reference dataset. We conduct the experiments both on real and simulated datasets with different bacterial and archaeal genomes from NCBI RefSeq in order to capture a comprehensive understanding of the performance. The simulated and real datasets are acquired from the original paper of Kraken [13] as well as from other related studies [14, 32,33,34]. The simulated datasets represent five mock communities that are constructed from real sequencing data: MiSeq, HiSeq, Mix1, Mix2, simBA5. MiSeq and HiSeq metagenomes were built using 10 sets of bacterial whole-genome shotgun reads. Mix1 and Mix2 are based on the same species of HiSeq, but with two different abundance profiles.
The MiSeq dataset is particularly difficult to analyze because it contains five genomes from the Enterobacteriaceae family (Citrobacter, Enterobacter, Klebsiella, Proteus and Salmonella). The high sequence similarity of this family can elevate the difficulty of classification. The metagenome simBA5 was created by simulating reads from the complete set of bacterial and archaeal genomes in RefSeq, for a total of 1216 species. It contains reads with sequencing errors caused in the reading process and it was created with the exact purpose of measuring the stability with existing errors in a complex communities.
We also evaluated the performance of SKraken on a real stool metagenomic sample (SRR1804065) from the Human Microbiome Project. Because there is no ground truth for this dataset, we use BLAST with a sequence identity of 95% to find the reads that uniquely mapped to a genome and filter out all other reads. If two paired-end reads do not map to the same genome, we discard them. As a result, the real metagenomic sample contains 775 distinct species and 1053741 reads. A summary of the main characteristics of all simulated and real metagenomics datasets can be found in Table 1.
3.2 Evaluation Metrics
In order to compare the results we used the standard metrics of precision and recall. Given N the number of reads, Y the number of reads classified and X the number of reads correctly classified, we define precision as the fraction of correct assignments over the total number of assignments (X/Y), and recall as the ratio between the number of correct assignments and the number of reads (X/N). Note that our classification task will be unfolded from both genus and species level, where genus level is the parent of species level in the taxonomic tree. If the classification step selected a node at genus level, while we are evaluating the species level, even if the assigned genus-level node is the parent of the correct leaf node, we considered it as mislabeled. On the other hand, when we evaluate the genus-level, if the classification step selected a taxonomic node that is a descendant (leaf node, species level) of the correct (genus-level) node, we consider it to be correct.
When analyzing a metagenomic sample one needs to verify whether the estimated abundance ratios of species is similar to the known profile, here we adopt Pearson correlation that is a technique widely used. If the correlation value is close to 1, which means the estimated abundance ratio perfectly matches the known one. In the next sections, we will test the behavior of SKraken on these three aspects: precision, recall and Pearson correlation.
3.3 Comparison
For a better perception of the performance of Skraken, we comparison it with Kraken, since it is one of the cutting-edge tools as mentioned in [16].
For Kraken we use the default parameter \(k=31\), as suggest by the authors [13], it is the best balance between precision and recall. For SKraken we use the same value \(k=31\) and we test the performance by varying the tuning parameter Q.

the figure taken from [17].
Results on dataset Mix1 varying the filtering parameter Q,
We devised a series of tests with the variable parameter Q and the taxonomic level at which the classification is evaluated. In the first set we want to test how the filtering parameter Q impact the performance metrics. We run Kraken and SKraken on the dataset Mix1 and evaluate the classification accuracy at the species-level. The results are reported in Fig. 4. If the parameter \(Q=0\) (no filtering) all k-mers are kept as we expected, thus the performance of Kraken and SKraken are identical. As Q grows to \(100\%\), we can see that the precision improves from \(63\%\) to \(75\%\), whereas the recall remains constant, which implies the number of mistakenly-classified reads decreases. Another important observation is that the Pearson correlation with the known abundance ratios also increases.
Thus, we use the most stringent filtering (\(Q=100\%\)) to classify all dataset at the species-level. Figure 5 shows a summary of precision and recall for all simulated and real metagenomic datasets. This test confirms that SKraken is able to improve the precision on all datasets without compromising the recall. On simulated metagenomes the average precision increases from \(73\%\) of Kraken to \(81\%\) of SKraken. On the real metagenome, the accuracy of Kraken turns out to be \(91\%\), SKraken achieves \(96\%\).
In general the study of metagenomic sample requires an analysis based on the component of the genome, and for this reason researchers develop the studies at the lowest taxonomic level, species. However metagenomic reads can be mapped at a higher level, it’s also valuable to survey the classification performance at the genus-level. We performed a set of experiments similar to the ones in species-level, considering the genus taxonomic level for classification. At first we set filtering parameter \(Q=100\%\), and the results are shown in Fig. 6. If we observe the performance of Kraken at genus level we can see that are better than those at species level, as expected. Indeed, in the taxonomy tree, when the classification level is more specific (lower), the label assignment is more difficult. Moreover, it is possible that some mislabeled reads exist at species level, while at genus level they are correct due to its loose condition. The average precision of Kraken is \(96\%\) at genus-level and \(73\%\) at species-level, we may see how great it improves.
When applying \(Q=100\%\), SKraken has a higher precision than Kraken in almost every database, while it’s not the case with respect to recall. If we consider a less stringent threshold \(Q=25\%\) (see Fig. 7), we can obtain results that are in line with the previous experiments, with a moderate improvement in the precision and nearly unchanged recall. A possible explanation of the small gain in terms of precision is that the classification at the genus level is relatively easier, and Kraken has already very good performance. Since precision and recall depends on the number of reads classified and on the number of correct assignments, in order to have a complete picture, results are summarized in Table 2.
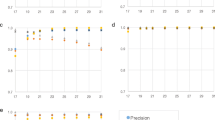
In the last experiment we test the ability to detect the correct abundance ratios in a metagenomic sample, we compared it at both genus level and species level of SKraken and Kraken, as displayed in Fig. 8. Both approaches have an extraordinary result in genus level, in particular, they virtually peaked in dataset HISeq, MIX1, MIX2 and SRR1804065. In dataset simBA5, SKraken increases the score from 0.92 to 0.97, since it is one of the most complex and realistic dataset, with 1216 species. If we compare these Pearson correlations with those of species level classification in general the values decrease confirming that it is more difficult to detect the correct species, rather than the genus. This is the case where the classification accuracy can benefit from a careful selection of discriminative k-mers. In fact in all dataset the correlation of SKraken is better than Kraken. Again, in one of the most difficult metagenome(simBA5) that contains 1216 species, the improvement is substantial from 0.61 of Kraken to 0.77 of SKraken.
All the results above have shown that SKraken enables to obtain a higher classifying precision without any expense of recall. In other words, more reads are correctly classified, and the estimated abundance ratios have a better match with the known profile. An important property of SKraken is that the impact on these metrics improves as the taxonomic level evaluated in the classification becomes lower and thus more difficult. Moreover, as the number of newly sequenced species grows, the probability that two non-related species share a given k-mer will grow. For this reason we conjecture that SKraken will be able to remove more uninformative k-mers as the number of sequenced genomes increases.
3.4 Memory and Speed
Besides the enhancement of precision, the screening processing has another welcomed “side effect”, it reduced the database size since the number of annotated k-mers decreases. The size of the database produced by Kraken, using all bacterial and archaeal complete genomes in NCBI RefSeq, is about 65 GB and it contains 5.8 billion k-mers. In Fig. 9 we evaluate the percentage of k-mers filtered by SKraken and the impact in memory for different values of threshold Q. As expected, the percentage of k-mers filtered grows with the threshold Q increases and it reaches the maximum of \(8.1\%\) with \(Q=100\). By construction, the impact in memory depends linearly on the number of k-mers to be indexed. When \(Q=100\), SKraken requires to index 5.3 billion k-mers in space of 60 GB.

(taken from [17]).
Percentage of k-mers filtered and database size as a function of the quality threshold Q
In Table 3 we report the classification speed as reads classified per minute, using these tools on a server equipped by Intel Xeon E5450 (12 MB Cache, 3.00 GHz) and 256 GB RAM, without multithreading. SKraken is tested, as a result being faster than Kraken on all datasets.
4 Conclusion
We have presented SKraken, an efficient and effective method for the genus-level and species-level classifications. It is based from Kraken, with the additional procedure where we detect the representative k-mers and filter the unqualified ones, which produces an improved dictionary. The experimental result has demonstrated that SKraken obtains a higher precision without compromising the recall, moreover it boosts the processing speed. As future direction of investigation it would be interesting to explore alternative way to define the k-mer scores incorporating other topological information of the tree of life.
Notes
- 1.
A preliminary version of this manuscript was published in [17].
References
Felczykowska, A., Bloch, S.K., Nejman-Faleczyk, B., Baraska, S.: Metagenomic approach in the investigation of new bioactive compounds in the marine environment. Acta Biochim. Pol. 59(4), 501–505 (2012)
Mande, S.S., Mohammed, M.H., Ghosh, T.S.: Classification of metagenomic sequences: methods and challenges. Briefings Bioinform. 13(6), 669–681 (2012)
Qin, J., Li, R., Raes, J., et al.: A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010)
Zeller, G., Tap, J., Voigt, A.Y., Sunagawa, S., Kultima, J.R., Costea, P.I., Amiot, A., Böhm, J., Brunetti, F., Habermann, N., Hercog, R., Koch, M., Luciani, A., Mende, D.R., Schneider, M.A., Schrotz-King, P., Tournigand, C., Tran Van Nhieu, J., Yamada, T., Zimmermann, J., Benes, V., Kloor, M., Ulrich, C.M., von Knebel Doeberitz, M., Sobhani, I., Bork, P.: Potential of fecal microbiota for early-stage detection of colorectal cancer. Mol. Syst. Biol. 10(11), 766 (2014)
Human Microbiome Project Consortium: Structure, function and diversity of the healthy human microbiome. Nature 486(7402), 207–214 (2012)
Said, H.S., Suda, W., Nakagome, S., Chinen, H., Oshima, K., Kim, S., Kimura, R., Iraha, A., Ishida, H., Fujita, J., Mano, S., Morita, H., Dohi, T., Oota, H., Hattori, M.: Dysbiosis of salivary microbiota in inflammatory bowel disease and its association with oral immunological biomarkers. DNA Res.: Int. J. Rapid Publ. Rep. Genes Genomes 21(1), 15–25 (2014)
Brown, C., Hug, L., Thomas, B., Sharon, I., Castelle, C., Singh, A., et al.: Unusual biology across a group comprising more than 15% of domain Bacteria. Nature 523(7559), 208–211 (2015)
Zhang, Z., Schwartz, S., Wagner, L., Miller, W.: A greedy algorithm for aligning DNA sequences. J. Comput. Biol. 7(1–2), 203–214 (2004)
Huson, D.H., Auch, A.F., Qi, J., Schuster, S.C.: Megan analysis of metagenomic data. Genome Res. 17, 377–386 (2007)
Caporaso, J.G., Kuczynski, J., Stombaugh, J., Bittinger, K., Bushman, F.D., Costello, E.K., Fierer, N., Pea, A.G., Goodrich, J.K., Gordon, J.I., Huttley, G.A., Kelley, S.T., Knights, D., Koenig, J.E., Ley, R.E., Lozupone, C.A., McDonald, D., Muegge, B.D., Pirrung, M., Reeder, J., Sevinsky, J.R., Turnbaugh, P.J., Walters, W.A., Widmann, J., Yatsunenko, T., Zaneveld, J., Knight, R.: Qiime allows analysis of high-throughput community sequencing data. Nat. Methods 7(5), 335–336 (2010)
Liu, B., Gibbons, T., Ghodsi, M., Treangen, T., Pop, M.: Accurate and fast estimation of taxonomic profiles from metagenomic shotgun sequences. BMC Genomics 12, P11 (2011)
Segata, N., Waldron, L., Ballarini, A., Narasimhan, V., Jousson, O., Huttenhower, C.: Metagenomic microbial community profiling using unique clade-specific marker genes. Nat. Methods 9, 811 (2012)
Wood, D., Salzberg, S.: Kraken: ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 15, R46 (2014)
Ounit, R., Wanamaker, S., Close, T.J., Lonardi, S.: Clark: fast and accurate classification of metagenomic and genomic sequences using discriminative k-mers. BMC Genomics 16(1), 1–13 (2015)
Ames, S.K., Hysom, D.A., Gardner, S.N., Lloyd, G.S., Gokhale, M.B., Allen, J.E.: Scalable metagenomic taxonomy classification using a reference genome database. Bioinformatics 29, 2253–2260 (2013)
Lindgreen, S., Adair, K.L., Gardner, P.: An evaluation of the accuracy and speed of metagenome analysis tools. Sci. Rep. 6, 19233 (2016)
Marchiori, D., Comin, M.: Skraken: fast and sensitive classification of short metagenomic reads based on filtering uninformative k-mers. In: Proceedings of the 10th International Joint Conference on Biomedical Engineering Systems and Technologies (BIOSTEC 2017), pp. 59–67 (2017)
Vinga, S., Almeida, J.: Alignment-free sequence comparison-a review. Bioinformatics 19, 513–523 (2003)
Comin, M., Verzotto, D.: Whole-genome phylogeny by virtue of unic subwords. In: 2012 23rd International Workshop on Database and Expert Systems Applications (DEXA), pp. 190–194, September 2012
Sims, G.E., Jun, S.R., Wu, G.A., Kim, S.H.: Alignment-free genome comparison with feature frequency profiles (FFP) and optimal resolutions. Proc. Nat. Acad. Sci. 106, 2677–2682 (2009)
Antonello, M., Comin, M.: Fast alignment-free comparison for regulatory sequences using multiple resolution entropic profiles. In: Proceedings of the International Conference on Bioinformatics Models, Methods and Algorithms (BIOSTEC 2015), pp. 171–177 (2015)
Comin, M., Antonello, M.: On the comparison of regulatory sequences with multiple resolution entropic profiles. BMC Bioinf. 17(1), 130 (2016)
Comin, M., Verzotto, D.: Beyond fixed-resolution alignment-free measures for mammalian enhancers sequence comparison. IEEE/ACM Trans. Comput. Biol. Bioinf. 11(4), 628–637 (2014)
Goke, J., Schulz, M.H., Lasserre, J., Vingron, M.: Estimation of pairwise sequence similarity of mammalian enhancers with word neighbourhood counts. Bioinformatics 28(5), 656–663 (2012)
Kantorovitz, M.R., Robinson, G.E., Sinha, S.: A statistical method for alignment-free comparison of regulatory sequences. Bioinformatics 23, i249–i255 (2007)
Comin, M., Antonello, M.: Fast computation of entropic profiles for the detection of conservation in genomes. In: Ngom, A., Formenti, E., Hao, J.-K., Zhao, X.-M., van Laarhoven, T. (eds.) PRIB 2013. LNCS, vol. 7986, pp. 277–288. Springer, Heidelberg (2013). https://doi.org/10.1007/978-3-642-39159-0_25
Antonello, M., Comin, M.: Fast entropic profiler: an information theoretic approach for the discovery of patterns in genomes. IEEE/ACM Trans. Comput. Biol. Bioinf. 11(3), 500–509 (2014)
Schimd, M., Comin, M.: Fast comparison of genomic and meta-genomic reads with alignment-free measures based on quality values. BMC Med. Genomics 9(1), 41–50 (2016)
Comin, M., Leoni, A., Schimd, M.: Clustering of reads with alignment-free measures and quality values. Algorithms Mol. Biol. 10(1), 1–10 (2015)
Comin, M., Schimd, M.: Assembly-free genome comparison based on next-generation sequencing reads and variable length patterns. BMC Bioinf. 15(9), 1–10 (2014)
Ondov, B.D., Treangen, T.J., Melsted, P., Mallonee, A.B., Bergman, N.H., Koren, S., Phillippy, A.M.: Mash: fast genome and metagenome distance estimation using MinHash. bioRxiv (2016)
Girotto, S., Pizzi, C., Comin, M.: Metaprob: accurate metagenomic reads binning based on probabilistic sequence signatures. Bioinformatics 32(17), i567–i575 (2016)
Girotto, S., Comin, M., Pizzi, C.: Metagenomic reads binning with spaced seeds. Theor. Comput. Sci. 698, 88–99 (2017)
Girotto, S., Comin, M., Pizzi, C.: Higher recall in metagenomic sequence classification exploiting overlapping reads. BMC Genomics 18, 917 (2017)
Acknowledgement
The authors would like to thank the anonymous reviewers for their valuable comments and suggestions. This work was supported by the Italian MIUR project PRIN20122F87B2.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG, part of Springer Nature
About this paper
Cite this paper
Qian, J., Marchiori, D., Comin, M. (2018). Fast and Sensitive Classification of Short Metagenomic Reads with SKraken. In: Peixoto, N., Silveira, M., Ali, H., Maciel, C., van den Broek, E. (eds) Biomedical Engineering Systems and Technologies. BIOSTEC 2017. Communications in Computer and Information Science, vol 881. Springer, Cham. https://doi.org/10.1007/978-3-319-94806-5_12
Download citation
DOI: https://doi.org/10.1007/978-3-319-94806-5_12
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-94805-8
Online ISBN: 978-3-319-94806-5
eBook Packages: Computer ScienceComputer Science (R0)