Abstract
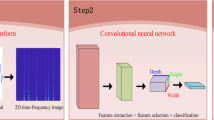
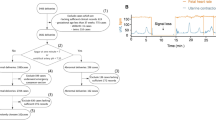
Electronic fetal monitoring (EFM) device which is used to record Fetal Heart Rate (FHR) and Uterine Contraction (UC) signals simultaneously is one of the significant tools in terms of the present obstetric clinical applications. In clinical practice, EFM traces are routinely evaluated with visual inspection by observers. For this reason, such a subjective interpretation has been caused various conflicts among observers to arise. Although the existing of international guidelines for ensuring more consistent assessment, the automated FHR analysis has been adopted as the most promising solution. In this study, an innovative approach based on deep convolutional neural network (DCNN) is proposed to classify FHR signals as normal and abnormal. The proposed method composes of three stages. FHR signals are passed through a set of preprocessing procedures in order to ensure more meaningful input images, firstly. Then, a visual representation of time-frequency information, spectrograms are obtained with the help of the Short Time Fourier Transform (STFT). Finally, DCNN method is utilized to classify FHR signals. To this end, the colored spectrograms images are used to train the network. In order to evaluate the proposed model, we conducted extensive experiments on the open CTU-UHB database considering the area under the receiver operating characteristic curve and other several performance metrics derived from the confusion matrix. Consequently, we achieved encouraging results.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Murray, H.: Antenatal foetal heart monitoring. Best Pract. Res. Clin. Obstet. Gynaecol. 38, 2–11 (2017)
Brown, R., Wijekoon, J.H.B., Fernando, A., Johnstone, E.D., Heazell, A.E.P.: Continuous objective recording of fetal heart rate and fetal movements could reliably identify fetal compromise, which could reduce stillbirth rates by facilitating timely management. Med. Hypotheses 83, 410–417 (2014)
van Geijn, H.P.: 2 Developments in CTG analysis. Baillieres Clin. Obstet. Gynaecol. 10, 185–209 (1996)
Ayres-de-Campos, D., Spong, C.Y., Chandraharan, E.: FIGO consensus guidelines on intrapartum fetal monitoring: Cardiotocography. Int. J. Gynecol. Obstet. 131, 13–24 (2015)
Tongsong, T., Iamthongin, A., Wanapirak, C., Piyamongkol, W., Sirichotiyakul, S., Boonyanurak, P., Tatiyapornkul, T., Neelasri, C.: Accuracy of fetal heart-rate variability interpretation by obstetricians using the criteria of the National Institute of Child Health and Human Development compared with computer-aided interpretation. J. Obstet. Gynaecol. Res. 31, 68–71 (2005)
Czabanski, R., Jezewski, J., Matonia, A., Jezewski, M.: Computerized analysis of fetal heart rate signals as the predictor of neonatal acidemia. Expert Syst. Appl. 39, 11846–11860 (2012)
Garabedian, C., Butruille, L., Drumez, E., Schreiber, E.S., Bartolo, S., Bleu, G., Mesdag, V., Deruelle, P., De Jonckheere, J., Houfflin-Debarge, V.: Inter-observer reliability of 4 fetal heart rate classifications. J. Gynecol. Obstet. Hum. Reprod. 46, 131–135 (2017)
Palomäki, O., Luukkaala, T., Luoto, R., Tuimala, R.: Intrapartum cardiotocography: the dilemma of interpretational variation. J. Perinat. Med. 34, 298–302 (2006)
Cömert, Z., Kocamaz, A.F.: Novel software for comprehensive analysis of cardiotocography signals CTG-OAS. In: KARCI, A. (ed.) International Conference on Artificial Intelligence and Data Processing (IDAP17), pp. 1–6. IEEE, Malatya (2017)
Bernardes, J., Ayres-de-Campos, D., Costa-Pereira, A., Pereira-Leite, L., Garrido, A.: Objective computerized fetal heart rate analysis. Int. J. Gynecol. Obstet. 62, 141–147 (1998)
Warrick, P., Hamilton, E., Macieszczak, M.: Neural network based detection of fetal heart rate patterns. In: IEEE International Joint Conference on Neural Networks, pp. 2400–2405 (2005)
Cömert, Z., Kocamaz, A.F.: Evaluation of fetal distress diagnosis during delivery stages based on linear and nonlinear features of fetal heart rate for neural network community. Int. J. Comput. Appl. 156, 26–31 (2016)
Magenes, G., Pedrinazzi, L., Signorini, M.G.: Identification of fetal sufferance antepartum through a multiparametric analysis and a support vector machine. In: 26th Annual International Conference of the IEEE Engineering in Medicine and Biology Society, pp. 462–465 (2004)
Monteiro-Santos, J., Gonçalves, H., Bernardes, J., Antunes, L., Nozari, M., Costa-Santos, C.: Entropy and compression capture different complexity features: the case of fetal heart rate. Entropy 19, 688 (2017)
Cömert, Z., Kocamaz, A.F.: Cardiotocography analysis based on segmentation-based fractal texture decomposition and extreme learning machine. In: 25th Signal Processing and Communications Applications Conference (SIU), pp. 1–4 (2017)
Cömert, Z., Kocamaz, A.F.: A study based on gray level co-occurrence matrix and neural network community for determination of hypoxic fetuses. In: International Artificial Intelligence and Data Processing Symposium (IDAP), pp. 569–573. TR (2016)
Cömert, Z., Kocamaz, A.F.: A study of artificial neural network training algorithms for classification of cardiotocography signals. Bitlis Eren Univ. J. Sci. Technol. 7, 93–103 (2017)
Spilka, J., Frecon, J., Leonarduzzi, R., Pustelnik, N., Abry, P., Doret, M.: Sparse support vector machine for intrapartum fetal heart rate classification. IEEE J. Biomed. Health Inform. 21(3), 664–671 (2016)
Cömert, Z., Kocamaz, A.F., Gungor, S.: Cardiotocography signals with artificial neural network and extreme learning machine. In: 24th Signal Processing and Communication Application Conference (SIU) (2016)
Sahin, H., Subasi, A.: Classification of the cardiotocogram data for anticipation of fetal risks using machine learning techniques. Appl. Soft Comput. 33, 231–238 (2015)
Cömert, Z., Kocamaz, A.F.: Comparison of machine learning techniques for fetal heart rate classification. Acta Phys. Pol. A 132, 451–454 (2017)
Bursa, M., Lhotska, L.: The use of convolutional neural networks in biomedical data processing. In: Bursa, M., Holzinger, A., Renda, M.E., Khuri, S. (eds.) Proceedings of Information Technology in Bio- and Medical Informatics: 8th International Conference, ITBAM 2017, Lyon, France, 28–31 August 2017, pp. 100–119. Springer, Cham (2017)
Krizhevsky, A., Sutskever, I., Hinton, G.E.: ImageNet classification with deep convolutional neural networks. In: Pereira, F., Burges, C.J.C., Bottou, L., Weinberger, K.Q. (eds.) Proceedings of the 25th International Conference on Neural Information Processing Systems, vol. 1, pp. 1097–1105. Curran Associates, Inc., USA (2012)
Chudáček, V., Spilka, J., Burša, M., Janků, P., Hruban, L., Huptych, M., Lhotská, L.: Open access intrapartum CTG database. BMC Pregnancy Childbirth 14, 16 (2014)
Cesarelli, M., Romano, M., Bifulco, P., Fedele, F., Bracale, M.: An algorithm for the recovery of fetal heart rate series from CTG data. Comput. Biol. Med. 37, 663–669 (2007)
Spilka, J., Georgoulas, G., Karvelis, P., Oikonomou, V.P., Chudáček, V., Stylios, C., Lhotská, L., Jankru, P.: Automatic evaluation of FHR recordings from CTU-UHB CTG database. In: Bursa, M., Khuri, S., Renda, M.E. (eds.) Proceedings of Information Technology in Bio- and Medical Informatics: 4th International Conference, ITBAM 2013, Prague, Czech Republic, 28 August 2013, pp. 47–61. Springer, Heidelberg (2013)
Kahaner, D., Moler, C., Nash, S.: Numerical Methods and Software. Prentice-Hall Inc., Upper Saddle River (1989)
Romano, M., Faiella, G., Bifulco, P., D’Addio, G., Clemente, F., Cesarelli, M.: Outliers detection and processing in CTG monitoring. In: Roa Romero, L.M. (ed.) XIII Mediterranean Conference on Medical and Biological Engineering and Computing 2013, MEDICON 2013, Seville, Spain, 25–28 September 2013, pp. 651–654. Springer, Cham (2014)
Nawab, S., Quatieri, T., Lim, J.: Signal reconstruction from short-time Fourier transform magnitude. IEEE Trans. Acoust. 31, 986–998 (1983)
Romano, M., Iuppariello, L., Ponsiglione, A.M., Improta, G., Bifulco, P., Cesarelli, M.: Frequency and time domain analysis of foetal heart rate variability with traditional indexes: a critical survey. Comput. Math Methods Med. 2016, 1–12 (2016)
Groome, L.J., Mooney, D.M., Bentz, L.S., Singh, K.P.: Spectral analysis of heart rate variability during quiet sleep in normal human fetuses between 36 and 40 weeks of gestation. Early Hum. Dev. 38, 1–9 (1994)
LeCun, Y., Bengio, Y., Hinton, G.: Deep learning. Nature 521, 436–444 (2015)
Yu, Y., Lin, H., Meng, J., Wei, X., Guo, H., Zhao, Z.: Deep transfer learning for modality classification of medical images. Information 8(3), 91 (2017)
Subha, V., Murugan, D.: Genetic Algorithm based feature subset selection for fetal state classification. J. Commun. Technol. Electron. Comput. Sci. 2, 13–17 (2015)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer International Publishing AG, part of Springer Nature
About this paper
Cite this paper
Cömert, Z., Kocamaz, A.F. (2019). Fetal Hypoxia Detection Based on Deep Convolutional Neural Network with Transfer Learning Approach. In: Silhavy, R. (eds) Software Engineering and Algorithms in Intelligent Systems. CSOC2018 2018. Advances in Intelligent Systems and Computing, vol 763. Springer, Cham. https://doi.org/10.1007/978-3-319-91186-1_25
Download citation
DOI: https://doi.org/10.1007/978-3-319-91186-1_25
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-91185-4
Online ISBN: 978-3-319-91186-1
eBook Packages: Intelligent Technologies and RoboticsIntelligent Technologies and Robotics (R0)