Abstract
Amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD) are two devastating and lethal neurodegenerative diseases seen comorbidly in up to 15% of patients. Despite several decades of research, no effective treatment or disease-modifying strategies have been developed. We now understand more than before about the genetics and biology behind ALS and FTD, but the genetic etiology for the majority of patients is still unknown and the phenotypic variability observed across patients, even those carrying the same mutation, is enigmatic. Additionally, susceptibility factors leading to neuronal vulnerability in specific central nervous system regions involved in disease are yet to be identified. As the inherited but dynamic epigenome acts as a cell-specific interface between the inherited fixed genome and both cell-intrinsic mechanisms and environmental input, adaptive epigenetic changes might contribute to the ALS/FTD aspects we still struggle to comprehend. This chapter summarizes our current understanding of basic epigenetic mechanisms, how they relate to ALS and FTD, and their potential as therapeutic targets. A clear understanding of the biological mechanisms driving these two currently incurable diseases is urgent—well-needed therapeutic strategies need to be developed soon. Disease-specific epigenetic changes have already been observed in patients and these might be central to this endeavor.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Amyotrophic lateral sclerosis
- Epigenetic modifications
- Frontotemporal dementia
- Methylation
- RNA-mediated regulation
1.1 Introduction
Amyotrophic lateral sclerosis (ALS) is the most prevalent motor neuron disease, causing progressive degeneration of upper and lower motor neurons in 2–3 individuals per 100,000 worldwide [1, 2]. Clinically, ALS is characterized by rapidly progressing muscle weakness and spasticity leading to paralysis and eventually respiratory failure [3]. Patients present with symptoms at a mean age of 55 years and die within 2–4 years [4].
Patients with ALS are also frequently affected by the second most common form of early-onset dementia known as frontotemporal dementia (FTD). While patient symptoms will start with either ALS or FTD, 13–15% of patients will eventually develop symptoms for the other disease [5,6,7,8,9]. This strong link between ALS and FTD was observed before researchers were able to link them genetically. FTD has a prevalence of 15–22 individuals per 100,000 worldwide [10] and is characterized by disturbances in behavior, personality, and language as a result of neurodegeneration in the frontal and temporal lobes. Cognitive impairments are usually detected around age 58, and it is fatal within 6–10 years after symptoms onset [11].
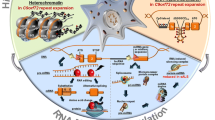
Historically, researchers have extensively searched for the underlying genetic etiologies for these conditions, yet, despite strong efforts to uncover genetic causes, the vast majority of ALS/FTD patients have no known genetic cause and no family history. The discovery of the C9orf72 DNA GGGGCC (G4C2) repeat expansion in ALS and FTD patients brought to light the possibility of RNA-mediated toxicity in disease which could be controlled by epigenetic mechanisms. In this chapter, we will summarize the major epigenetic regulation mechanisms. Then, we will review the current knowledge of epigenetic regulation as it relates to ALS and FTD (Fig. 1.1).
Schematic representation of the major epigenetic regulatory mechanisms and their interactions. The major epigenetic regulatory mechanisms are framed. Red stars mark epigenetic changes known to be involved in ALS/FTD. Orange stars indicate that altered expression of these RNAs has been reported in ALS/FTD. Purple spheres represent 5mC marks while green spheres represent 5hmC marks
1.2 Epigenetic Mechanisms
The epigenome is generally defined as the collection of heritable but dynamic mechanisms that largely dictate access to the DNA for templated functions such as gene transcription, DNA synthesis, and repair without altering the primary nucleotide sequence; there is, however, a valid argument that chimeric DNA, which does modify the primary nucleotide sequence, is facilitated by the epigenome. These intricacies regarding how far-reaching the epigenome’s involvement is in nucleotide sequence will be continually clarified as the field matures. Here, we will focus on general regulation processes.
Post-transcriptional regulation of RNA function and metabolism—by noncoding RNAs (ncRNAs), such as microRNA, or chemical modifications to ribonucleotides—is now also considered part of epigenetic control. Whereas an individual’s genome is mostly established at conception, the individual’s inherited epigenome continues to change throughout embryonic, fetal, and postnatal life in response to cell-intrinsic mechanisms and environmental input. Epigenetic changes are cell-specific and are presumed to be adaptive responses to internal and external stimuli.
As next-generation sequencing is becoming more commonplace, the epigenome is garnering greater attention in many fields, from developmental biology to environmental epidemiology, because it offers a potential mechanistic link between the environment and phenotype. For ALS/FTD researchers, epigenetics is a particularly interesting field that may explain the clinical variability within families sharing ALS/FTD-causing genetic variants, and the cause of disease in patients affected sporadically.
Epigenetic control of genomic functions occurs by regulating nucleosome position and density via histone chaperones and chromatin remodeling complexes [12]. Nucleosomes are the fundamental repeating unit of chromatin, and consist of nucleoprotein complexes containing two copies each of four core histone proteins (H2A, H2B, H3, and H3) and 147 base pairs of DNA wrapped 1.67 turns counterclockwise around this globular structure [13]. Key signals directing nucleosome positioning and chromatin structure include chemical modifications to the DNA and histones, as well as ncRNA species. Epigenetic regulation is also exerted via post-transcriptional regulation of RNA function and metabolism. This section will briefly review our current understanding of these epigenetic mechanisms.
1.2.1 Nuclear DNA Methylation
DNA methylation is the biochemical addition of a methyl group to a nucleotide, often resulting in altered transcription factor binding, chromatin remodeling, and ultimately altered gene expression. Methylation of the carbon 5 on cytosine (5mC) is the most studied and best understood DNA modification.
5mC most frequently occurs at cytosine-guanine nucleotide sequences (mCG), often referred to as CpG sites. Genomic regions enriched in CpG sites, referred to as CpG islands, are often associated with gene promoters. Whether located near transcription initiation sites or distant from annotated promoters, CpG islands play a central role in destabilizing nucleosomes and recruiting proteins that initiate chromatin remodeling [14,15,16]. Generally, methylation in a CpG island is associated with the reduced expression of a nearby gene.
Recent research has found 5mC marks at cytosine-adenine sequences (mCA). It is believed that mCA may be particularly important in the central nervous system, as the number of mCA and mCG marks detected in the adult brain are approximately equal [17]. Although both mCA and mCG are associated with transcriptional repression, early evidence suggests mCA may be particularly important for more precise transcription regulation—1–2-fold expression changes compared to 100–1000-fold changes observed with mCG [18].
DNA methylation is catalyzed by DNA methyltransferases (DNMTs) 1, 3A, and 3B. While DNMT3A and DNMT3B are associated with de novo methylation, DNMT1 is mainly responsible for maintaining methylation across cell generations by methylating the daughter strand of DNA during cell division, thus preventing hemimethylation of double-stranded DNA [19]. Because of DNMT1’s maintenance function, DNA methylation is considered a fairly stable epigenetic mark. Further research into the differentiating roles of DNMT3A and DNMT3B are warranted, as early evidence suggests DNMT3A may be primarily responsible for mCA methylation. Its proper regulation may be especially critical in the brain [20,21,22].
Bisulfite conversion is currently the preferred method to detect cytosine methylation. Briefly, bisulfite ion deaminates unmethylated cytosines, which result in its chemical conversion to uracil upon alkaline desulfonation [23]. As bisulfite converts 5mCs much more slowly, there is a selective conversion favoring unmethylated cytosines. After DNA amplification, all unconverted cytosines are considered methylated. Importantly, standard bisulfite conversion does not differentiate methylated cytosines from cytosines that have been demethylated, which is the topic of the next section.
1.2.2 Nuclear DNA Demethylation
Although DNA methylation is considered a stable epigenetic change, it is now well understood that it is closely and dynamically regulated not only by epigenetic “writers,” such as DNMTs, but also by “eraser” mechanisms, which keep CpGs unmethylated. Importantly, demethylation is of particular interest in neurodegeneration research because, among all organs, its footprints are most abundant in the brain and further increase with age [24].
DNA demethylation can occur as a passive or active process. Passive demethylation occurs when the process of copying methylation marks onto the daughter strand of DNA is impaired. As a result, future cell generations will lose methylation marks by dilution. Active demethylation is mediated by TET family of dioxygenases, which catalyze the oxidation of 5mC in a stepwise manner, resulting in 5-hydroxymethylcytosine (5hmC), 5-formylcytosine (5fC), and 5-carboxylcytosine (5caC). Oxidation is followed by replication-dependent dilution or thymine DNA glycosylase-mediated base excision and repair, resulting in full demethylation and restoration to unmodified cytosine [25]. 5fC- and 5caC-containing regulatory elements show very limited overlap and are therefore believed to play distinct regulatory roles. 5caC sites are more active than 5fC sites, and both are more active than regions decorated by 5hmC [26].
Our current understanding is that DNA methylation and demethylation processes must be highly orchestrated to appropriately regulate the transcriptome. Further research is necessary to elucidate the exact functions of 5hmC, 5fC and 5caC as it is still difficult to estimate the extent to which these events occur and appreciate their full potential as therapeutic targets.
1.2.3 Mitochondrial DNA Methylation
While nuclear DNA (nDNA) methylation is well recognized, mitochondrial DNA (mtDNA) methylation is still controversial. Using restriction enzyme cleavage, early studies estimated that about 2–5% of mtDNA CCGG sequences are fully methylated whereas the remainder of mtDNA is unmethylated [27]. Mass spectrometry confirmed the presence of methylated bases in human mtDNA [28]. Later, using affinity-based methods, both 5mC and 5hmC modifications were found in the D-loop region of human mtDNA at both CpG and non-CpG dinucleotides [29, 30]. Other studies used bisulfite conversion coupled with sequencing or pyrosequencing and claimed that mtDNA methylation may be as high as 2–18% in the D-loop region [31, 32]. Recent reports however suggest that mtDNA methylation levels may have been greatly overestimated due to the circular structure of mtDNA, which affects bisulfite conversion efficiency [33,34,35]. Nonetheless, the reported presence of methyltransferases DNMT1 and DNMT3A inside mitochondria suggests that methylation may take place in mtDNA [30, 36]. Since nDNA and mtDNA interact to modulate the transcriptome [37, 38], future studies need to better characterize mtDNA modifications and their potential role in the regulation of gene transcription.
1.2.4 RNA Methylation
Epigenetic regulation goes beyond transcription. RNA is also under epigenetic regulation and can be methylated or otherwise modified. In fact, methylation occurs in most RNA classes, but the exact function of RNA modifications remains unclear due to limitations in detection and quantification.
Whereas DNA methylation has mainly been observed at cytosines, RNA methylation can occur at cytosine (m5C), adenine (m6A), and guanine (m7G). m7G (N7-methylguanine) at the 5′ cap structure is the most widely studied RNA methylation and is necessary for the translation of most messenger RNAs (mRNAs). m7G cap also mediates nuclear transport of some mRNAs, preserves mRNA stability by protecting it from degradation, and facilitates other processing such as the addition of poly(A) tails to mRNAs [39, 40]. m7G cap methylation is reversible and can either go through the process of “decapping,” where the entire m7G cap is removed [41], or potentially through m7G demethylation.
In contrast to the extensively studied 5mC in DNA, the m5C (N5-methylcytosine) RNA modification has not been thoroughly explored. While it is still unclear how m5C is regulated by DNMTs, a study suggested that mouse Dnmt2 RNA methyltransferase may be required for epigenetic heredity [42]. Early studies also demonstrated that m5C affects interactions of long noncoding RNAs (lncRNAs) with chromatin-associated protein complexes [43].
The most abundant mRNA modification is m6A (N6-methyladenine). Dominissini et al. found that silencing m6A methyltransferase—mostly acting at highly conserved long internal exons, stop codons, and 3′UTRs—alters gene expression and alternative splicing patterns [44]. The same group conducted affinity-based m6A profiling and identified proteins that bind specifically to m6A. Their findings suggest that RNA methylation, through the binding of these m6A-specific binding proteins, may disturb RNA binding proteins’ affinity to associate with partner unmethylated RNAs. Importantly, these m6A-binding proteins recruit additional factors that facilitate functions such as alternative splicing, nuclear export, and mRNA stability [40, 44]. One example is the m6A-binding protein YTHDC1, which facilitates the export of methylated mRNA from the nucleus to the cytoplasm in vitro. Knockdown of YTHDC1 results in retention of nuclear m6A-containing mRNA, where transcripts accumulate in the nucleus and are depleted from the cytoplasm [45]. Of interest, m6A has been the first reversible modification identified in coding and noncoding RNAs after the discovery that the fat mass and obesity-associated protein (FTO) act as a m(6)A demethylase [46,47,48], suggesting that, analogous to reversible DNA, reversible RNA methylation may affect gene expression and downstream regulation of RNA-related cellular pathways.
1.2.5 RNA-Mediated Regulation
While 70–90% of the genome is transcribed, only 1–3% of these transcripts encode proteins [49, 50]. As such, the transcriptome is mainly composed of ncRNAs having infrastructural and regulatory functions. These transcripts are gaining increased attention, not only for their recognized roles in transcriptional and post-transcriptional regulation, but also for their targeted effects on gene expression, making them attractive therapeutic targets.
ncRNAs are divided into short (<30 nucleotides) and long ncRNAs (>200 nucleotides), and each subtype of ncRNA fulfills very specific regulatory roles. Specifically, short RNAs (sRNAs) are sub-categorized into micro-RNAs (miRNAs), piwi-interacting RNAs (piRNAs), and small interfering RNAs (siRNAs). lncRNAs are generally sub-classified according to their proximity to protein coding genes—sense, anti-sense, bidirectional, intronic, and intergenic. Apart from their size, sRNAs and lncRNAs have largely different functions in transcription regulation. Two new classes of ncRNAs have also been recently recognized: enhancer RNAs (eRNAs) and promoter-associated RNAs (PARs).
Mature miRNAs are single-stranded 20–24 nucleotide sequences derived from precursor miRNA (pre-miRNAS). These are ~70 nucleotide transcripts with distinct hairpin structures initially derived from nuclear primary miRNA (pri-miRNAs). Mature miRNAs pair with complementary sequences on target mRNA transcripts through the 3′ untranslated region (3′UTR) and repress the targeted mRNA translation after recruiting the RNA-induced silencing complex (RISC) [51, 52]. Mature miRNAs may have several target mRNAs, but their effect on their targets is a modest 1–2-fold change in expression [53]. Although miRNAs seem to initiate modest changes in their targets, changes in expression of miRNAs themselves associate with widespread mRNA expression changes, indicating that miRNAs are central to global expression regulation [53]. In fact, a large screening of the mammalian genome found hundreds of mRNAs with conserved pairing to specific miRNAs, with an enrichment of genes involved in transcriptional regulation [54]. Specifically, several miRNAs target DNMTs that can have far-reaching impact on global methylation levels, which has been implicated in several cancers and autoimmune diseases [55]. The story becomes more complicated when one considers that miRNA expression can in turn be modulated by DNA methylation and histone modifications [55, 56], suggesting that transcriptional regulation needs to be well orchestrated by an epigenetic–ncRNA feedback loop. This highlights the complexity of epigenetics and the interdependence of these mechanisms to regulate gene expression.
piRNAs are 24–31-nucleotide sequences able to form complexes with Piwi proteins of the Argonaute family. They are part of a complex population of small RNAs highly enriched in male gonads [57]. Their main function is to silence transposable elements. These mobile elements are autonomous sequences of DNA that replicate and insert themselves into the genome, potentially introducing detrimental DNA damage. Some results also suggest a role for piRNA in transcriptional regulation and deadenylation-mediated mRNA degradation [58].
Mature siRNAs are 20–24 nucleotide sequences that modulate a given gene’s expression after binding to its complementary nucleotide sequence through a process called RNA interference (RNAi). siRNAs mediate post-transcriptional silencing in a way similar to miRNAs, and may also suppress transposon activity in a way similar to piRNAs [59, 60].
lncRNAs are sequences longer than 200 nucleotides characterized by low nuclear expression and low conservation across species. One exception is highly conserved large intergenic noncoding RNAs (lincRNAs), which recruit histone-modifying complexes and transcription factors to transcriptionally modulate targeted chromosomal regions [61, 62]. The majority of lncRNAs are transcribed as complex networks of sense and anti-sense transcripts overlapping protein-coding genes.
eRNAs are sequences of about 800 nucleotides on average that are derived from regions enriched in RNA Polymerase II and transcriptional co-regulators. What makes the genomic sites encoding eRNAs different from those of other lncRNAs is their specific histone methylation signature at histone 3 lysine 4 (H3K4), which is typical of enhancer sites. Similar to lincRNAs, eRNAs are evolutionarily conserved but have a short half-life. eRNA expression levels correlate with the expression of nearby genes, and is thus dynamically regulated upon signaling [63, 64]. eRNAs are believed to act as transcriptional activators.
PARs are short half-life transcripts that can be classified as short or long ncRNAs—their size range from 16 to over 200 nucleotides. PARS are either expressed near transcription start sites or from elements of the promoter [65], from both strands and in divergent orientation with respect to the transcription start site. Most PARS associate with highly expressed genes while being weakly expressed themselves. Increasing evidence demonstrates that PARs may associate with both transcriptional activation and repression [66,67,68,69].
1.2.6 Histone Modifications
As discussed above, the density of nucleosomes determines the accessibility of DNA to protein complexes performing templated functions. Heterochromatin contains tightly wound DNA, where gene transcription is repressed. Euchromatin is a looser conformation of the DNA, which is conducive to gene transcription. Chromatin conformation is dynamic and is regulated in part by covalent posttranslational histone modifications, which mainly occur at amino acid residues within unstructured histone “tails.” These modifications include acetylation and methylation of lysines, methylation of arginines, and phosphorylation of serines and threonines, among others. Specific functions have been assigned to several modifications, which led to the formulation of the histone code hypothesis [70]. It is our current understanding of the roles of histone modifications that their combinations and association with transcription factors and other chromatin regulators define epigenetic states, which can be discovered and assigned to genomic regions by machine learning [71].
Enzymes such as histone acetyltransferases (HATs), histone methyltransferases (HMTs), protein arginine methyltransferases (PRMTs), and kinases facilitate the addition of chemicals on amino acid residues (writers). For instance, HATs brings in a negative charge that neutralizes the positive charge on the histones. This chemical change decreases the interaction of the N termini of histones with the negatively charged phosphate groups of DNA, resulting in a more relaxed structure. This structure allows transcription factors to access DNA and initiate gene transcription. Other enzymes such as histone deacetylases (HDACs), lysine demethylases (KDMs), phosphatases, and deubiquitylases catalyze the removal of these epigenetic marks (erasers). As such, the process of histone acetylation is reversed by HDACs. Acetyl group removal leads to a more condensed chromatin state and transcriptional gene silencing. Proteins containing DNA methyl-binding domains, chromodomains, bromodomains, and Tudor domains recognize histone modifications and recruit other chromatin modifiers and remodeling proteins to ultimately regulate DNA-dependent processes (readers).
Functional groups affixed to histones can also initiate chromatin remodeling through cis or trans effects. This way, histone marks can modulate the chromatin by directly affecting histone-histone and histone-DNA interactions, or by recruiting non-histone proteins via specific binding domains that recognize particular modifications [72].
1.3 Epigenetic Changes in Amyotrophic Lateral Sclerosis (ALS) and Frontotemporal Dementia (FTD)
The 2011 discovery that an expanded hexanucleotide (G4C2) repeat within a noncoding region of the C9orf72 gene causes both ALS and FTD highlighted the genetic link between these two diseases and recognized this mutation as the most common genetic cause of ALS and FTD identified to date [73,74,75], yet more than 80% of all ALS and FTD cases remain genetically unexplained [76]. While other unknown genetic mutations are certainly at play, more researchers are recognizing that epigenetic changes may contribute to these two diseases based on their potential to (1) explain the phenotypic variability observed across family members carrying the same ALS/FTD-associated mutation, and (2) explain why the majority of patients develop disease without any family history of ALS or FTD. Very few epigenetic studies have been conducted on ALS and FTD before the finding of the pathogenic C9orf72 G4C2 repeat expansion in 2011. This section summarizes the current knowledge on the epigenetics of ALS and FTD. A full summary can also be found in Table 1.1.
1.3.1 ALS/FTD Epigenetic Studies
1.3.1.1 Nuclear DNA methylation /demethylation
Approximately a decade ago, researchers interested in better understanding the etiology of ALS and FTD started interrogating epigenetic mechanisms using cell models, animal models, and biospecimen obtained from patients.
Early studies on ALS evaluated blood and brain promoter DNA methylation status of a few genes known to be implicated in disease including SOD1, VEGF, and SLC1A2 [136, 137], but no changes in methylation were detected.
The methylation status of SOD1, FUS, TARDBP, and C9orf72 promoters has also been evaluated from the blood of ALS patients carrying not fully penetrant SOD1 mutations, but again, no methylation variations have been detected at these specific regulatory regions [77]. Of interest, Coppede et al. used an enzyme-linked immunosorbent assay (ELISA) to also evaluate global methylation levels in ALS patients carrying not fully penetrant SOD1 mutations and observed a significant overall DNA methylation increase [77].
Similarly, others compared brain methylation levels of sporadic ALS (sALS) patients to control cases using Affymetrix GeneChip Human Tiling 2.0R Arrays and identified 38 differentially methylated genomic regions in patients. Further analysis of these 38 regions shed light on specifically altered biological pathways involved in calcium homeostasis, neurotransmission, and oxidative stress [78].
Using ELISA assays, Figueroa-Romero et al. identified global 5mC and 5hmC changes in sALS patients’ spinal cords—these changes were not observed in blood [79]. Then, using high-throughput microarrays, the same group conducted genome-wide 5mC and expression profiling and identified loci-specific differentially methylated and expressed genes. The 112 genes identified were highly associated with immune and inflammation responses [79].
In the hope of finding differently methylated regions that might act as modifiers of age of onset in ALS, Tremolizzo et al. evaluated DNA methylation levels in both early onset (<55 years) and late onset (>74 years) ALS patients. They found a global 25–30% increase in DNA methylation levels in whole blood that was independent of age of onset [80]. While no significant difference was found between early onset and late onset ALS, the methylation increase detected in blood is consistent with previous observation in the central nervous system (CNS) of ALS patients [36, 99].
Early epigenetic studies on FTD analyzed the progranulin-encoding gene (GRN) known to be mutated in 5–20% of familial FTD and 1–5% of sporadic FTD patients [138,139,140]. Two groups independently reported that increased GRN promoter methylation negatively correlates with GRN mRNA levels in FTD subjects [81, 82], a very interesting finding since GRN haploinsufficiency has been recognized as a major cause of FTD [141].
Global DNA methylation levels in blood were also assessed in tau-associated progressive supranuclear palsy (PSP) and FTD patients by Li et al. The major known risk locus for PSP and other neurodegenerative diseases is the H1 haplotype at 17q21.31, a genomic region in linkage disequilibrium with an inverted chromosomal sequence of about 970 kb [142,143,144]. It was shown that this disequilibrium resulted from an inversion at the H2 haplotype relative to the H1 human reference allele and from a lack of recombination between inverted and non-inverted chromosomes [145]. Li et al. found that the H1/H2 locus may affect the risk for tauopathies through methylation alterations not only at the MAPT locus, a region known to be mutated in FTD, but also in at least three neighboring genes. The 17q21.31-associated DNA methylation signature Li et al. identified was unique to tau-associated PSP patients and, to a lesser extent, to tau-associated FTD [83].
As DNA methyltransferases catalyze DNA methylation, researchers have been interested in assessing their expression in disease. Chestnut et al. found that protein levels of DNMT1 and DNMT3A were increased in the motor cortex of ALS patients [36]. Wong et al. went further and confirmed the presence of both Dnmt3a isoform in pure mitochondria of human cerebral cortex and mouse CNS, and 5mcs in mouse mitochondria of neurons and skeletal muscle myofibers, supporting an epigenetic regulation of brain mtDNA [84]. Wong et al. then evaluated mitochondrial Dnmt3a isoform levels in the CNS of different SOD1 transgenic mouse models including a line hemizygous for a low copy number of hSOD1 -p.G37R mutant allele, a line that expressed high levels of normal wild-type human SOD1 gene, and a line with skeletal muscle-restricted expression of hSOD1 -p.G37R, -p.G93A, and -wild-type variants. After studying all the lines at presymptomatic or early to middle stages of disease, they found that Dnmt3a protein levels in mouse skeletal muscle and spinal cord mitochondria were significantly reduced early in disease, this even before symptoms’ onset. They also found Dnmt1 bound to the outer mitochondrial membrane of the same mice. They observed that 5mC immunoreactivity became aggregated and sequestered into autophagosomes of transgenic mice motor neurons [84].
1.3.1.2 RNA-mediated regulation through miRNAs
It is well established that ncRNAs such as miRNAs are key regulators of RNA functions and metabolism. As such, many have been interested in assessing their potential contribution to ALS/FTD pathogenesis.
Dobrowolny et al. found that selective expression of the human SOD1 mutation p.G39A after injecting mouse muscles not only leads to hypomyelination in the sciatic nerve, it also alters spinal cord expression of miRNAs and mRNAs known to be involved in myelin homeostasis. This finding suggests that RNA and epigenetic alterations observed in motor neurons may result from changes initiated in neighboring non-neuronal cells [85].
Figueroa-Romero et al. then interrogated spinal cord tissues obtained from postmortem ALS patients and found that mature miRNA levels are globally reduced. They also identified altered microRNAs having RNA targets part of pathways previously associated with ALS. Knowing that TDP-43 is central to ALS and FTD pathogenesis—TDP-43 pathological signature is observed in about 97% of ALS and 50% of FTD patients [146, 147]—and that TDP-43 plays a central role in miRNA biogenesis [89, 148,149,150,151,152], the same group used transfected cells to determine whether TDP-43 mediates miRNA-induced regulation. They observed ALS-associated miRNA expression changes in response to nuclear clearance and cytoplasmic aggregation of TDP-43, suggesting that TDP-43 pathology may alter the expression or function of endogenous miRNAs and their downstream targets [86].
Several studies have identified alternatively expressed miRNAs in ALS and FTD. Marcuzzo et al. studied the brain of pre-symptomatic and late stage SOD1 p.G93A transgenic mice and found that expression levels of miR-9, miR-124a, miR-19a, and miR-19b were all altered in late stages of disease. Moreover, the expression analysis they conducted identified miRNA/target gene pairs that were differentially expressed in this mouse model [87]. Toivonen et al. found miR-206 altered in the blood of both SOD1 p.G93A transgenic mice and ALS patients [88], whereas Zhang et al. found levels of miR-9 decreased in induced pluripotent stem cells (iPSCs)-derived neurons of patients carrying either p.A90V or p.M337V TARDBP mutations [89].
Of interest, Jiao et al. found that miR-29b regulates human progranulin (hPGRN, GRN) through 3′UTR binding. They demonstrated in vitro that ectopic expression of miR-29b decreased hPGRN expression and knockdown of endogenous miR-29b increased it. Their findings suggest that miR-29b may possibly be therapeutically targeted to rescue the haploinsufficiency observed in FTD patients carrying a GRN mutation [90]. Moreover, expression of FTD-associated mutant CHMP2B in cerebral cortices of mice has initiated a decrease in miR-124 expression—brain-enriched miR-124 is especially important for the proper regulation of AMPA receptor (AMPAR) subunits. Ectopic expression of miR-124 in the prefrontal cortex of these mice restored AMPAR levels and rescued the behavioral deficits previously observed in the animals [91].
1.3.2 C9orf72 Epigenetic Studies
Decreased expression of one or multiple C9orf72 transcript variants has been observed in various human biospecimen carrying the pathological C9orf72 G4C2 repeat expansion. These biospecimen include frontal cortex, motor cortex, cerebellum, cervical spinal cord, lymphoblastoid cell lines, iPSCs and neurons differentiated from iPSCs, all obtained or derived from ALS and FTD patients [98, 153,154,155,156,157,158,159]. In an attempt to better understand the biological mechanism underlying the reduced C9orf72 expression in C9orf72-associated ALS and FTD (c9FTD/ALS) patients, methylation status of the regions flanking or encompassing the repeat expansion has been evaluated.
Using bisulfite sequencing and restriction enzyme assays, Rogaeva’s group found that the CpG island upstream of the G4C2 repeat expansion is hypermethylated in the brain and blood of about 36% of ALS and 17% of FTD cases [94, 96]. Lee’s group not only confirmed these results, but also uncovered that hypermethylation associates with reduced accumulation of intronic C9orf72 RNA and reduced burden of C9orf72-associated pathological signature (RNA foci and dipeptide repeat accumulation). They also found that demethylation increases cell vulnerability to oxidative and autophagic stress, suggesting that C9orf72 promoter hypermethylation may mitigate downstream molecular aberrations associated with the pathological G4C2 repeat expansion [93]. As the methods initially used to estimate DNA methylation in c9FTD/ALS cases could not differentiate 5mC from 5hmC, Esanov et al. were able to confirm the presence of 5hmC within the C9orf72 promoter in post-mortem brain tissues of hypermethylated patients [97]. This finding suggests that the previous estimates by Rogaeva’s and Lee’s groups included both 5mC and 5hmC modifications.
However, considering that all c9FTD/ALS patients show a 50% reduction in total C9orf72 RNA expression [73] and only approximately one third of patients are found with a hypermethylated CpG island, many details were missing. As such, Rogaeva’s group attempted to assess whether it is the repeat expansion that is hypermethylated in patients and consequently drives the reduced expression. For this purpose, they developed a new qualitative assay that was independently validated by a methylation-sensitive restriction enzyme assay, and found that the C9orf72 repeat expansion was indeed hypermethylated in all ALS and FTD cases carrying more than 50 G4C2 copies [95]. This finding was later confirmed by Bauer [92]. In addition, Belzil et al. investigated the brain of c9FTD/ALS patients and found that all patients carried repressive histone marks at the C9orf72 locus [98]. As such, methylation of the repeat expansion together with repressive histone marks at the C9orf72 locus likely explains the 50% C9orf72 reduced expression observed in c9FTD/ALS patients.
A recent multi-omic study aimed to better understand the molecular mechanisms initiating RNA misregulation in C9orf72-associated c9ALS and sALS combined RNA and DNA methylation data obtained from brain next-generation RNA sequencing (RNAseq) and reduced representation bisulfite sequencing (RRBS). They found an abundance of differentially methylated cytosines in c9ALS and sALS patients, including changes in many genes and biological pathways known to be involved in ALS. They also observed that c9ALS and sALS patients have generally distinct but overlapping brain DNA methylation profiles that differ from control individuals. Of importance, they found that the c9ALS- and sALS-affected genes and biological pathways have very similar biological functions, suggesting a conserved pathobiology between c9ALS and sALS [99].
Several studies aimed to assess whether DNA methylation is a clinical modifier of ALS and FTD but so far, few correlations have been identified. Among these, hypermethylation of the CpG island upstream of the C9orf72 repeat expansion has been found to correlate with shorter disease duration in ALS [96], but was found associated with longer disease duration and later age of death in FTD [160].
1.3.3 Potential Drivers of Epigenetic Changes
The epigenome is a dynamic, complex machinery that plays a critical role in coordinating cellular functions. It is constantly changing to address cellular needs or to react to environmental threats, such as infections. A prime example is heat-shock proteins. These proteins were discovered in the early 1960s by Ferrucio Ritossa when he noticed a “puffing” pattern—now known to be a sudden increase in RNA transcription—in Drosophila cells when one of his lab mates increased his incubator’s temperature [161]. Many discoveries have resulted from this unintended finding, but one of the most intriguing discoveries was that the “puff” Ritossa described was observable within 2–3 min of heat exposure, demonstrating how agile the epigenomic machinery is. Understanding not only which epigenetic modifications affect disease, but what drives these epigenetic changes is critical to better understanding human health and disease.
As demonstrated by Ritossa’s landmark discovery that “heat shock” can induce an immediate response from the epigenetic machinery, environmental influences are a clear driver. “Environment” has broad implications, however, and can include a cell’s internal or external influences, such as neighboring cells, as research has shown that epigenetic changes can be transmitted from cell to cell [162]. Many factors affect the dynamic interaction between environmental influences and the epigenome, including exercise, age, diet, and toxic exposures.
Researchers observed a clear example of environmental factors driving ALS in the indigenous Chamorro people of Guam, who experience high ALS incidence because their diet is enriched in cycad neurotoxins. The Chamorro diet includes the flying fox, which has high levels of cycad neurotoxins because it feeds on cycad seeds [100,101,102]. Additional studies have found associations between ALS and cycad neurotoxins or reactive oxygen species (ROS) [102,103,104, 163,164,165]. Exactly how these neurotoxins are driving disease is unknown, but epigenetic modifications are a primary suspect. Importantly, diet has also been shown to induce epigenetic changes across other diseases [105].
The most striking support for epigenetic contribution is perhaps the reports of ALS-discordant monozygotic twins (monozygotic twins where one has disease and the other is unaffected), implicating environmental and epigenetic factors in disease [106,107,108]. One study identified monozygotic twins that both carry the C9orf72 repeat expansion, but only one has developed disease. The other study could not find a clear genetic factor that caused disease. A third study by Young et al. identified thousands of large between-twin differences at CpG sites in five monozygotic twin pairs. Young et al. conducted biological pathway analysis, which revealed that impairments in GABA signaling were common to all ALS individuals. Other altered pathways were also identified, including some relevant to ALS such as glutamate metabolism and the Golgi apparatus [108]. Importantly, Young et al. applied to their 450K data the Horvath algorithm of epigenetic age [166]—an aging clock of chromatin states derived from the characterization of 353 CpG sites—and found that ALS-affected twins were epigenetically older than their unaffected co-twins, confirming previous findings that ALS is characterized by accelerated brain aging [108, 167]. In all cases, the other twin may develop disease in time, but the question would still remain regarding why a significant time gap in onset exists.
Stress is also an environmental variable that has received increased attention in recent years. Both histone methylation and acetylation modifications have been observed in rodents because of stress after social defeat [168]—acute and chronic stress has been shown to activate and repress genes through histone modifications [169]. Interestingly, transposable elements are repressed during acute stress, as are hippocampal coding and noncoding RNA as a result of stress-induced histone modifications. These expression changes have been suggested to impair genomic stability and give rise to cognitive impairments [109,110,111,112,113,114].
ROS, a species of free radical, can be induced through environmental signals, causing oxidative stress and, ultimately, cause a range of epigenetic modifications altering gene expression [162, 170,171,172]. Heavy metals are believed to cause cellular stress and toxicity through ROS, driving protein denaturation and aggregation and preventing proteasomes from eliminating dysfunctional proteins [120]. One study used carboxy-DCFDA (5-(and-6)-carboxy-2′,7′-dichlorofluorescin diacetate) to quantify stress-induced ROS production from metal sulfates in human neurons [121] and found aluminum sulfate induced the most ROS.
Repetitive electromagnetic field exposure is also believed to trigger epigenetic changes, including DNA methylation and histone modifications, as was suggested by a study of a large cohort of workers [122]. Resistance welders had a higher incidence of Alzheimer’s Disease and ALS, potentially because they are exposed regularly to low frequency magnetic fields [122]. Other studies have suggested that exposure to other heavy metals such as mercury, lead, and selenium, plus pesticides and herbicides containing organophosphate may increase risk for ALS, though no clear association has been found [115,116,117,118,119, 123,124,125,126,127,128,129,130].
As science continues to demonstrate the environmental effects of some high-contact sports on mental health, additional studies to explore the increased ALS incidence in athletes that play American football and soccer [131, 173], and in war veterans [132, 133] are needed. These data further suggest that some ALS cases arise from environmental exposures, potentially from epigenetic consequences of violent jarring in the brain [134]. While various methods, including illicit drug use and ischemia from head injuries, have been proposed to increase ROS production and drive dementia [135], the exact molecular mechanism leading to ALS and FTD needs to be clearly mapped through future rigorous studies.
1.3.4 Therapeutic Potential
The ultimate goal in ALS and FTD research is to develop therapeutic interventions for these diseases, allowing those who are affected to live a long and healthy life. Here, we discuss potential effects of epigenetic therapeutics.
Targeted epigenetic modifiers capable of regulating expression of both the normal and mutant alleles are an exciting possibility. Although significant work must be done in this area, this concept was successfully demonstrated in 1994, suggesting epigenetic therapy is feasible [174]. Since that discovery, other research has been performed in neurodegeneration and cancer in an effort to translate this for clinical use [175, 176]. A recent ALS-specific study utilized bromodomain small molecule inhibitors to increase mRNA and pre-mRNA expression for the normal C9orf72 allele without destroying the epigenetic markers that repress expression of the expanded allele [177]. Increasing expression of the normal allele without increasing expression for the disease-causing allele is a significant achievement and demonstrates the reality of epigenetic therapy.
DNA methylation effects have been extensively evaluated across many diseases. One study found DNA methylation changes may be good ALS biomarkers for disease and potentially future therapeutic targets [79]. DNMTS have been shown to promote apoptosis and increase 5mC levels in motor neurons, and administering Dnmt inhibitors in a motor neuron-degenerative mouse model mitigated both apoptosis and 5mC levels in the motor neurons [36].
The therapeutic potential of oligonucleotides targeting miRNAs has also been evaluated by researchers using mouse models of ALS. Two groups showed that oligonucleotides able to inhibit either miR-155 or miR-29a extended the lifespan of SOD1 p.G93A transgenic mice [178, 179]. Similarly, Morel et al. found that injecting oligonucleotides targeting miR-124a in the same transgenic mouse model prevented the pathological loss of EAAT2/GLT1 (encoded by human SLC1A2), an astroglial glutamate transporter known to be implicated in ALS [180].
Researchers have shown that histone marks at the C9orf72 locus are associated with reduced gene expression in ALS and FTD patients carrying a repeat expansion when compared to controls [98]. They were then able to increase C9orf72 mRNA expression by treating patient-derived fibroblasts with 5-aza-2-deoxycytidine (a demethylating agent). While this study demonstrates a proof of concept, increasing the mutated allele might not be a good therapeutic approach for c9ALS/FTD, as results from others suggest that C9orf72-associated hypermethylation may actually be neuroprotective in patients [92, 93]. Nonetheless, similar epigenetic strategies have been developed for cancer therapy, where HDAC inhibitors have reversed the effects of cancer-induced epigenetic changes [181]. These techniques have also been applied in ALS both in vitro and in an animal model, and later proceeded to clinical trials. Specifically, sodium phenylbutyrate (NaPB) prolonged SOD1 p.G93A mouse survival [182], and was subsequently tested in a phase 2 clinical trial. The participants of this clinical trial tolerated this treatment well, and increased histone acetylation in participant blood samples [183]. These results present an exciting and realistic opportunity to treat ALS and FTD using targeted epigenetic therapeutics.
Given the epigenome’s dynamic and targetable nature combined with its apparent involvement in disease, it is a primary target for additional therapeutic efforts. As a field, researchers studying epigenetics in neurodegenerative diseases have made significant progress characterizing their involvement, but it is unclear whether the observed epigenetic modifications are driving disease or whether they are just a consequence. For example, researchers recently observed clear transcriptomic and epigenetic differences between c9ALS and sALS brains [99, 184], but it has not been shown whether reversing them would rescue neuronal health. If researchers can establish that the epigenetic dysregulation is driving these changes and that reversing them rescues neuronal behavior, epigenetic therapeutics would revolutionize ALS and FTD treatment.
References
Cronin S, Hardiman O, Traynor BJ. Ethnic variation in the incidence of ALS: a systematic review. Neurology. 2007;68:1002–7. https://doi.org/10.1212/01.wnl.0000258551.96893.6f.
Marin B, Boumediene F, Logroscino G, Couratier P, Babron MC, Leutenegger AL, et al. Variation in worldwide incidence of amyotrophic lateral sclerosis: a meta-analysis. Int J Epidemiol. 2017;46:57–74. https://doi.org/10.1093/ije/dyw061.
Bradley WG. Neurology in clinical practice. 3rd ed. Boston, MA: Butterworth-Heinemann; 2000.
Chio A, Logroscino G, Hardiman O, Swingler R, Mitchell D, Beghi E, et al. Prognostic factors in ALS: a critical review. Amyotroph Lateral Scler. 2009;10:310–23. https://doi.org/10.3109/17482960802566824.
Burrell JR, Kiernan MC, Vucic S, Hodges JR. Motor neuron dysfunction in frontotemporal dementia. Brain. 2011;134:2582–94. https://doi.org/10.1093/brain/awr195. awr195 [pii].
Giordana MT, Ferrero P, Grifoni S, Pellerino A, Naldi A, Montuschi A. Dementia and cognitive impairment in amyotrophic lateral sclerosis: a review. Neurol Sci. 2011;32:9–16. https://doi.org/10.1007/s10072-010-0439-6.
Gordon PH, Delgadillo D, Piquard A, Bruneteau G, Pradat PF, Salachas F, et al. The range and clinical impact of cognitive impairment in French patients with ALS: a cross-sectional study of neuropsychological test performance. Amyotroph Lateral Scler. 2011;12:372–8. https://doi.org/10.3109/17482968.2011.580847.
Lomen-Hoerth C, Anderson T, Miller B. The overlap of amyotrophic lateral sclerosis and frontotemporal dementia. Neurology. 2002;59:1077–9.
Ringholz GM, Appel SH, Bradshaw M, Cooke NA, Mosnik DM, Schulz PE. Prevalence and patterns of cognitive impairment in sporadic ALS. Neurology. 2005;65:586–90.
Knopman DS, Roberts RO. Estimating the number of persons with frontotemporal lobar degeneration in the US population. J Mol Neurosci. 2011;45:330–5. https://doi.org/10.1007/s12031-011-9538-y.
Hodges JR, Davies R, Xuereb J, Kril J, Halliday G. Survival in frontotemporal dementia. Neurology. 2003;61:349–54.
Pal S, Tyler JK. Epigenetics and aging. Sci Adv. 2016;2:e1600584. https://doi.org/10.1126/sciadv.1600584.
Luger K, Mader AW, Richmond RK, Sargent DF, Richmond TJ. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature. 1997;389:251–60. https://doi.org/10.1038/38444.
Illingworth RS, Gruenewald-Schneider U, Webb S, Kerr AR, James KD, Turner DJ, et al. Orphan CpG islands identify numerous conserved promoters in the mammalian genome. PLoS Genet. 2010;6:e1001134. https://doi.org/10.1371/journal.pgen.1001134.
Maunakea AK, Nagarajan RP, Bilenky M, Ballinger TJ, D’Souza C, Fouse SD, et al. Conserved role of intragenic DNA methylation in regulating alternative promoters. Nature. 2010;466:253–7. https://doi.org/10.1038/nature09165.
Saxonov S, Berg P, Brutlag DL. A genome-wide analysis of CpG dinucleotides in the human genome distinguishes two distinct classes of promoters. Proc Natl Acad Sci U S A. 2006;103:1412–7. https://doi.org/10.1073/pnas.0510310103.
He Y, Ecker JR. Non-CG methylation in the human genome. Annu Rev Genomics Hum Genet. 2015;16:55–77. https://doi.org/10.1146/annurev-genom-090413-025437.
Stroud H, Su SC, Hrvatin S, Greben AW, Renthal W, Boxer LD, et al. Early-life gene expression in neurons modulates lasting epigenetic states. Cell. 2017;171:1151–1164.e16. https://doi.org/10.1016/j.cell.2017.09.047.
Kinney SR, Pradhan S. Regulation of expression and activity of DNA (cytosine-5) methyltransferases in mammalian cells. Prog Mol Biol Transl Sci. 2011;101:311–33. https://doi.org/10.1016/B978-0-12-387685-0.00009-3.
Gabel HW, Kinde B, Stroud H, Gilbert CS, Harmin DA, Kastan NR, et al. Disruption of DNA-methylation-dependent long gene repression in Rett syndrome. Nature. 2015;522:89–93. https://doi.org/10.1038/nature14319.
Guo JU, Su Y, Shin JH, Shin J, Li H, Xie B, et al. Distribution, recognition and regulation of non-CpG methylation in the adult mammalian brain. Nat Neurosci. 2014;17:215–22. https://doi.org/10.1038/nn.3607.
Lister R, Mukamel EA, Nery JR, Urich M, Puddifoot CA, Johnson ND, et al. Global epigenomic reconfiguration during mammalian brain development. Science. 2013;341:1237905. https://doi.org/10.1126/science.1237905.
Darst RP, Pardo CE, Ai L, Brown KD, Kladde MP (2010) Bisulfite sequencing of DNA. Curr Protoc Mol Biol Chapter 7:Unit 7 9 1-17. doi:https://doi.org/10.1002/0471142727.mb0709s91
Hahn MA, Szabo PE, Pfeifer GP. 5-Hydroxymethylcytosine: a stable or transient DNA modification? Genomics. 2014;104:314–23. https://doi.org/10.1016/j.ygeno.2014.08.015.
Wu X, Zhang Y. TET-mediated active DNA demethylation: mechanism, function and beyond. Nat Rev Genet. 2017;18:517–34. https://doi.org/10.1038/nrg.2017.33.
Lu X, Han D, Zhao BS, Song CX, Zhang LS, Dore LC, et al. Base-resolution maps of 5-formylcytosine and 5-carboxylcytosine reveal genome-wide DNA demethylation dynamics. Cell Res. 2015;25:386–9. https://doi.org/10.1038/cr.2015.5.
Shmookler Reis RJ, Goldstein S. Mitochondrial DNA in mortal and immortal human cells. Genome number, integrity, and methylation. J Biol Chem. 1983;258:9078–85.
Infantino V, Castegna A, Iacobazzi F, Spera I, Scala I, Andria G, et al. Impairment of methyl cycle affects mitochondrial methyl availability and glutathione level in Down’s syndrome. Mol Genet Metab. 2011;102:378–82. https://doi.org/10.1016/j.ymgme.2010.11.166.
Bellizzi D, D’Aquila P, Scafone T, Giordano M, Riso V, Riccio A, et al. The control region of mitochondrial DNA shows an unusual CpG and non-CpG methylation pattern. DNA Res. 2013;20:537–47. https://doi.org/10.1093/dnares/dst029.
Shock LS, Thakkar PV, Peterson EJ, Moran RG, Taylor SM. DNA methyltransferase 1, cytosine methylation, and cytosine hydroxymethylation in mammalian mitochondria. Proc Natl Acad Sci U S A. 2011;108:3630–5. https://doi.org/10.1073/pnas.1012311108.
Byun HM, Barrow TM. Analysis of pollutant-induced changes in mitochondrial DNA methylation. Methods Mol Biol. 2015;1265:271–83. https://doi.org/10.1007/978-1-4939-2288-8_19.
Byun HM, Panni T, Motta V, Hou L, Nordio F, Apostoli P, et al. Effects of airborne pollutants on mitochondrial DNA methylation. Part Fibre Toxicol. 2013;10:18. https://doi.org/10.1186/1743-8977-10-18.
Hong EE, Okitsu CY, Smith AD, Hsieh CL. Regionally specific and genome-wide analyses conclusively demonstrate the absence of CpG methylation in human mitochondrial DNA. Mol Cell Biol. 2013;33:2683–90. https://doi.org/10.1128/MCB.00220-13.
Liu B, Du Q, Chen L, Fu G, Li S, Fu L, et al. CpG methylation patterns of human mitochondrial DNA. Sci Rep. 2016;6:23421. https://doi.org/10.1038/srep23421.
Mechta M, Ingerslev LR, Fabre O, Picard M, Barres R. Evidence suggesting absence of mitochondrial DNA methylation. Front Genet. 2017;8:166. https://doi.org/10.3389/fgene.2017.00166.
Chestnut BA, Chang Q, Price A, Lesuisse C, Wong M, Martin LJ. Epigenetic regulation of motor neuron cell death through DNA methylation. J Neurosci. 2011;31:16619–36. https://doi.org/10.1523/JNEUROSCI.1639-11.2011.
Doynova MD, Berretta A, Jones MB, Jasoni CL, Vickers MH, O’Sullivan JM. Interactions between mitochondrial and nuclear DNA in mammalian cells are non-random. Mitochondrion. 2016;30:187–96. https://doi.org/10.1016/j.mito.2016.08.003.
Rodley CD, Grand RS, Gehlen LR, Greyling G, Jones MB, O’Sullivan JM. Mitochondrial-nuclear DNA interactions contribute to the regulation of nuclear transcript levels as part of the inter-organelle communication system. PLoS One. 2012;7:e30943. https://doi.org/10.1371/journal.pone.0030943.
Glover-Cutter K, Kim S, Espinosa J, Bentley DL. RNA polymerase II pauses and associates with pre-mRNA processing factors at both ends of genes. Nat Struct Mol Biol. 2008;15:71–8. https://doi.org/10.1038/nsmb1352.
Liu J, Jia G. Methylation modifications in eukaryotic messenger RNA. J Genet Genomics. 2014;41:21–33. https://doi.org/10.1016/j.jgg.2013.10.002.
Liu H, Kiledjian M. Decapping the message: a beginning or an end. Biochem Soc Trans. 2006;34:35–8. https://doi.org/10.1042/BST20060035.
Kiani J, Grandjean V, Liebers R, Tuorto F, Ghanbarian H, Lyko F, et al. RNA-mediated epigenetic heredity requires the cytosine methyltransferase Dnmt2. PLoS Genet. 2013;9:e1003498. https://doi.org/10.1371/journal.pgen.1003498.
Amort T, Souliere MF, Wille A, Jia XY, Fiegl H, Worle H, et al. Long noncoding RNAs as targets for cytosine methylation. RNA Biol. 2013;10:1003–8. https://doi.org/10.4161/rna.24454.
Dominissini D, Moshitch-Moshkovitz S, Schwartz S, Salmon-Divon M, Ungar L, Osenberg S, et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature. 2012;485:201–6. https://doi.org/10.1038/nature11112.
Roundtree IA, Luo GZ, Zhang Z, Wang X, Zhou T, Cui Y, et al. YTHDC1 mediates nuclear export of N(6)-methyladenosine methylated mRNAs. Elife. 2017;6:pii: e31311. https://doi.org/10.7554/eLife.31311.
Jia G, Fu Y, He C. Reversible RNA adenosine methylation in biological regulation. Trends Genet. 2013;29:108–15. https://doi.org/10.1016/j.tig.2012.11.003.
Jia G, Fu Y, Zhao X, Dai Q, Zheng G, Yang Y, et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol. 2011;7:885–7. https://doi.org/10.1038/nchembio.687.
Zheng G, Dahl JA, Niu Y, Fedorcsak P, Huang CM, Li CJ, et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol Cell. 2013;49:18–29. https://doi.org/10.1016/j.molcel.2012.10.015.
Consortium EP. An integrated encyclopedia of DNA elements in the human genome. Nature. 2012;489:57–74. https://doi.org/10.1038/nature11247.
Knowling S, Morris KV. Non-coding RNA and antisense RNA. Nature’s trash or treasure? Biochimie. 2011;93:1922–7. https://doi.org/10.1016/j.biochi.2011.07.031.
Baek D, Villen J, Shin C, Camargo FD, Gygi SP, Bartel DP. The impact of microRNAs on protein output. Nature. 2008;455:64–71. https://doi.org/10.1038/nature07242.
Guo H, Ingolia NT, Weissman JS, Bartel DP. Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature. 2010;466:835–40. https://doi.org/10.1038/nature09267.
Gosline SJ, Gurtan AM, JnBaptiste CK, Bosson A, Milani P, Dalin S, et al. Elucidating microRNA regulatory networks using transcriptional, post-transcriptional, and histone modification measurements. Cell Rep. 2016;14:310–9. https://doi.org/10.1016/j.celrep.2015.12.031.
Lewis BP, Shih IH, Jones-Rhoades MW, Bartel DP, Burge CB. Prediction of mammalian microRNA targets. Cell. 2003;115:787–98.
Saito Y, Saito H, Liang G, Friedman JM. Epigenetic alterations and microRNA misexpression in cancer and autoimmune diseases: a critical review. Clin Rev Allergy Immunol. 2014;47:128–35. https://doi.org/10.1007/s12016-013-8401-z.
Saito Y, Liang G, Egger G, Friedman JM, Chuang JC, Coetzee GA, et al. Specific activation of microRNA-127 with downregulation of the proto-oncogene BCL6 by chromatin-modifying drugs in human cancer cells. Cancer Cell. 2006;9:435–43. https://doi.org/10.1016/j.ccr.2006.04.020.
Carmell MA, Girard A, van de Kant HJ, Bourc’his D, Bestor TH, de Rooij DG, et al. MIWI2 is essential for spermatogenesis and repression of transposons in the mouse male germline. Dev Cell. 2007;12:503–14. https://doi.org/10.1016/j.devcel.2007.03.001.
Rouget C, Papin C, Boureux A, Meunier AC, Franco B, Robine N, et al. Maternal mRNA deadenylation and decay by the piRNA pathway in the early Drosophila embryo. Nature. 2010;467:1128–32. https://doi.org/10.1038/nature09465.
Watanabe T, Takeda A, Tsukiyama T, Mise K, Okuno T, Sasaki H, et al. Identification and characterization of two novel classes of small RNAs in the mouse germline: retrotransposon-derived siRNAs in oocytes and germline small RNAs in testes. Genes Dev. 2006;20:1732–43. https://doi.org/10.1101/gad.1425706.
Yang N, Kazazian HH Jr. L1 retrotransposition is suppressed by endogenously encoded small interfering RNAs in human cultured cells. Nat Struct Mol Biol. 2006;13:763–71. https://doi.org/10.1038/nsmb1141.
Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature. 2009;458:223–7. https://doi.org/10.1038/nature07672.
Khalil AM, Guttman M, Huarte M, Garber M, Raj A, Rivea Morales D, et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc Natl Acad Sci U S A. 2009;106:11667–72. https://doi.org/10.1073/pnas.0904715106.
De Santa F, Barozzi I, Mietton F, Ghisletti S, Polletti S, Tusi BK, et al. A large fraction of extragenic RNA pol II transcription sites overlap enhancers. PLoS Biol. 2010;8:e1000384. https://doi.org/10.1371/journal.pbio.1000384.
Kim TK, Hemberg M, Gray JM, Costa AM, Bear DM, Wu J, et al. Widespread transcription at neuronal activity-regulated enhancers. Nature. 2010;465:182–7. https://doi.org/10.1038/nature09033.
Preker P, Nielsen J, Kammler S, Lykke-Andersen S, Christensen MS, Mapendano CK, et al. RNA exosome depletion reveals transcription upstream of active human promoters. Science. 2008;322:1851–4. https://doi.org/10.1126/science.1164096.
Affymetrix ETP, Cold Spring Harbor Laboratory ETP. Post-transcriptional processing generates a diversity of 5′-modified long and short RNAs. Nature. 2009;457:1028–32. https://doi.org/10.1038/nature07759.
Han J, Kim D, Morris KV. Promoter-associated RNA is required for RNA-directed transcriptional gene silencing in human cells. Proc Natl Acad Sci U S A. 2007;104:12422–7. https://doi.org/10.1073/pnas.0701635104.
Morris KV, Santoso S, Turner AM, Pastori C, Hawkins PG. Bidirectional transcription directs both transcriptional gene activation and suppression in human cells. PLoS Genet. 2008;4:e1000258. https://doi.org/10.1371/journal.pgen.1000258.
Wang X, Arai S, Song X, Reichart D, Du K, Pascual G, et al. Induced ncRNAs allosterically modify RNA-binding proteins in cis to inhibit transcription. Nature. 2008;454:126–30. https://doi.org/10.1038/nature06992.
Jenuwein T, Allis CD. Translating the histone code. Science. 2001;293:1074–80. https://doi.org/10.1126/science.1063127.
Ernst J, Kellis M. Chromatin-state discovery and genome annotation with ChromHMM. Nat Protoc. 2017;12:2478–92. https://doi.org/10.1038/nprot.2017.124.
Hake SB, Allis CD. Histone H3 variants and their potential role in indexing mammalian genomes: the “H3 barcode hypothesis”. Proc Natl Acad Sci U S A. 2006;103:6428–35. https://doi.org/10.1073/pnas.0600803103.
DeJesus-Hernandez M, Mackenzie IR, Boeve BF, Boxer AL, Baker M, Rutherford NJ, et al. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron. 2011;72:245–56. https://doi.org/10.1016/j.neuron.2011.09.011. S0896-6273(11)00828-2 [pii].
Rademakers R, van Blitterswijk M. Motor neuron disease in 2012: novel causal genes and disease modifiers. Nat Rev Neurol. 2013;9:63–4. https://doi.org/10.1038/nrneurol.2012.276. nrneurol.2012.276 [pii].
Renton AE, Majounie E, Waite A, Simon-Sanchez J, Rollinson S, Gibbs JR, et al. A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron. 2011;72:257–68. https://doi.org/10.1016/j.neuron.2011.09.010. S0896-6273(11)00797-5 [pii].
Renton AE, Chio A, Traynor BJ. State of play in amyotrophic lateral sclerosis genetics. Nat Neurosci. 2014;17:17–23. https://doi.org/10.1038/nn.3584. nn.3584 [pii].
Coppede F, Stoccoro A, Mosca L, Gallo R, Tarlarini C, Lunetta C, et al. Increase in DNA methylation in patients with amyotrophic lateral sclerosis carriers of not fully penetrant SOD1 mutations. Amyotroph Lateral Scler Frontotemporal Degener. 2017;19(1-2):93. https://doi.org/10.1080/21678421.2017.1367401.
Morahan JM, Yu B, Trent RJ, Pamphlett R. A genome-wide analysis of brain DNA methylation identifies new candidate genes for sporadic amyotrophic lateral sclerosis. Amyotroph Lateral Scler. 2009;10:418–29. https://doi.org/10.3109/17482960802635397.
Figueroa-Romero C, Hur J, Bender DE, Delaney CE, Cataldo MD, Smith AL, et al. Identification of epigenetically altered genes in sporadic amyotrophic lateral sclerosis. PLoS One. 2012;7:e52672. https://doi.org/10.1371/journal.pone.0052672.
Tremolizzo L, Messina P, Conti E, Sala G, Cecchi M, Airoldi L, et al. Whole-blood global DNA methylation is increased in amyotrophic lateral sclerosis independently of age of onset. Amyotroph Lateral Scler Frontotemporal Degener. 2014;15:98–105. https://doi.org/10.3109/21678421.2013.851247.
Banzhaf-Strathmann J, Claus R, Mucke O, Rentzsch K, van der Zee J, Engelborghs S, et al. Promoter DNA methylation regulates progranulin expression and is altered in FTLD. Acta Neuropathol Commun. 2013;1:16. https://doi.org/10.1186/2051-5960-1-16.
Galimberti D, D’Addario C, Dell’osso B, Fenoglio C, Marcone A, Cerami C, et al. Progranulin gene (GRN) promoter methylation is increased in patients with sporadic frontotemporal lobar degeneration. Neurol Sci. 2013;34:899–903. https://doi.org/10.1007/s10072-012-1151-5.
Li Y, Chen JA, Sears RL, Gao F, Klein ED, Karydas A, et al. An epigenetic signature in peripheral blood associated with the haplotype on 17q21.31, a risk factor for neurodegenerative tauopathy. PLoS Genet. 2014;10:e1004211. https://doi.org/10.1371/journal.pgen.1004211.
Wong M, Gertz B, Chestnut BA, Martin LJ. Mitochondrial DNMT3A and DNA methylation in skeletal muscle and CNS of transgenic mouse models of ALS. Front Cell Neurosci. 2013;7:279. https://doi.org/10.3389/fncel.2013.00279.
Dobrowolny G, Bernardini C, Martini M, Baranzini M, Barba M, Musaro A. Muscle Expression of SOD1(G93A) Modulates microRNA and mRNA Transcription Pattern Associated with the Myelination Process in the Spinal Cord of Transgenic Mice. Front Cell Neurosci. 2015;9:463. https://doi.org/10.3389/fncel.2015.00463.
Figueroa-Romero C, Hur J, Lunn JS, Paez-Colasante X, Bender DE, Yung R, et al. Expression of microRNAs in human post-mortem amyotrophic lateral sclerosis spinal cords provides insight into disease mechanisms. Mol Cell Neurosci. 2016;71:34–45. https://doi.org/10.1016/j.mcn.2015.12.008.
Marcuzzo S, Bonanno S, Kapetis D, Barzago C, Cavalcante P, D’Alessandro S, et al. Up-regulation of neural and cell cycle-related microRNAs in brain of amyotrophic lateral sclerosis mice at late disease stage. Mol Brain. 2015;8:5. https://doi.org/10.1186/s13041-015-0095-0.
Toivonen JM, Manzano R, Olivan S, Zaragoza P, Garcia-Redondo A, Osta R. MicroRNA-206: a potential circulating biomarker candidate for amyotrophic lateral sclerosis. PLoS One. 2014;9:e89065. https://doi.org/10.1371/journal.pone.0089065.
Zhang Z, Almeida S, Lu Y, Nishimura AL, Peng L, Sun D, et al. Downregulation of microRNA-9 in iPSC-derived neurons of FTD/ALS patients with TDP-43 mutations. PLoS One. 2013;8:e76055. https://doi.org/10.1371/journal.pone.0076055.
Jiao J, Herl LD, Farese RV, Gao FB. MicroRNA-29b regulates the expression level of human progranulin, a secreted glycoprotein implicated in frontotemporal dementia. PLoS One. 2010;5:e10551. https://doi.org/10.1371/journal.pone.0010551.
Gascon E, Lynch K, Ruan H, Almeida S, Verheyden JM, Seeley WW, et al. Alterations in microRNA-124 and AMPA receptors contribute to social behavioral deficits in frontotemporal dementia. Nat Med. 2014;20:1444–51. https://doi.org/10.1038/nm.3717.
Bauer PO. Methylation of C9orf72 expansion reduces RNA foci formation and dipeptide-repeat proteins expression in cells. Neurosci Lett. 2016;612:204–9. https://doi.org/10.1016/j.neulet.2015.12.018.
Liu EY, Russ J, Wu K, Neal D, Suh E, McNally AG, et al. C9orf72 hypermethylation protects against repeat expansion-associated pathology in ALS/FTD. Acta Neuropathol. 2014;128:525–41. https://doi.org/10.1007/s00401-014-1286-y.
Xi Z, Rainero I, Rubino E, Pinessi L, Bruni AC, Maletta RG, et al. Hypermethylation of the CpG-island near the C9orf72 G(4)C(2)-repeat expansion in FTLD patients. Hum Mol Genet. 2014;23:5630–7. https://doi.org/10.1093/hmg/ddu279.
Xi Z, Zhang M, Bruni AC, Maletta RG, Colao R, Fratta P, et al. The C9orf72 repeat expansion itself is methylated in ALS and FTLD patients. Acta Neuropathol. 2015;129:715–27. https://doi.org/10.1007/s00401-015-1401-8.
Xi Z, Zinman L, Moreno D, Schymick J, Liang Y, Sato C, et al. Hypermethylation of the CpG island near the G4C2 repeat in ALS with a C9orf72 expansion. Am J Hum Genet. 2013;92:981–9. https://doi.org/10.1016/j.ajhg.2013.04.017.
Esanov R, Belle KC, van Blitterswijk M, Belzil VV, Rademakers R, Dickson DW, et al. C9orf72 promoter hypermethylation is reduced while hydroxymethylation is acquired during reprogramming of ALS patient cells. Exp Neurol. 2015;277:171–7. https://doi.org/10.1016/j.expneurol.2015.12.022.
Belzil VV, Bauer PO, Prudencio M, Gendron TF, Stetler CT, Yan IK, et al. Reduced C9orf72 gene expression in c9FTD/ALS is caused by histone trimethylation, an epigenetic event detectable in blood. Acta Neuropathol. 2013;126:895–905. https://doi.org/10.1007/s00401-013-1199-1.
Ebbert MTW, Ross CA, Pregent LJ, Lank RJ, Zhang C, Katzman RB, et al. Conserved DNA methylation combined with differential frontal cortex and cerebellar expression distinguishes C9orf72-associated and sporadic ALS, and implicates SERPINA1 in disease. Acta Neuropathol. 2017;134:715–28. https://doi.org/10.1007/s00401-017-1760-4.
Banack SA, Cox PA. Biomagnification of cycad neurotoxins in flying foxes: implications for ALS-PDC in Guam. Neurology. 2003;61:387–9.
Bradley WG, Mash DC. Beyond Guam: the cyanobacteria/BMAA hypothesis of the cause of ALS and other neurodegenerative diseases. Amyotroph Lateral Scler. 2009;10(Suppl 2):7–20. https://doi.org/10.3109/17482960903286009.
Chiu AS, Gehringer MM, Welch JH, Neilan BA. Does alpha-amino-beta-methylaminopropionic acid (BMAA) play a role in neurodegeneration? Int J Environ Res Public Health. 2011;8:3728–46. https://doi.org/10.3390/ijerph8093728.
Dastur DK. Cycad toxicity in monkeys: clinical, pathological, and biochemical aspects. Fed Proc. 1964;23:1368–9.
Polsky FI, Nunn PB, Bell EA. Distribution and toxicity of alpha-amino-beta-methylaminopropionic acid. Fed Proc. 1972;31:1473–5.
Remely M, Stefanska B, Lovrecic L, Magnet U, Haslberger AG. Nutriepigenomics: the role of nutrition in epigenetic control of human diseases. Curr Opin Clin Nutr Metab Care. 2015;18:328–33. https://doi.org/10.1097/MCO.0000000000000180.
Meltz Steinberg K, Nicholas TJ, Koboldt DC, Yu B, Mardis E, Pamphlett R. Whole genome analyses reveal no pathogenetic single nucleotide or structural differences between monozygotic twins discordant for amyotrophic lateral sclerosis. Amyotroph Lateral Scler Frontotemporal Degener. 2015;16:385–92. https://doi.org/10.3109/21678421.2015.1040029.
Xi Z, Yunusova Y, van Blitterswijk M, Dib S, Ghani M, Moreno D, et al. Identical twins with the C9orf72 repeat expansion are discordant for ALS. Neurology. 2014;83:1476–8. https://doi.org/10.1212/WNL.0000000000000886.
Young PE, Kum Jew S, Buckland ME, Pamphlett R, Suter CM. Epigenetic differences between monozygotic twins discordant for amyotrophic lateral sclerosis (ALS) provide clues to disease pathogenesis. PLoS One. 2017;12:e0182638. https://doi.org/10.1371/journal.pone.0182638.
Erwin JA, Marchetto MC, Gage FH. Mobile DNA elements in the generation of diversity and complexity in the brain. Nat Rev Neurosci. 2014;15:497–506. https://doi.org/10.1038/nrn3730.
Hunter RG, Gagnidze K, McEwen BS, Pfaff DW. Stress and the dynamic genome: steroids, epigenetics, and the transposome. Proc Natl Acad Sci U S A. 2015;112:6828–33. https://doi.org/10.1073/pnas.1411260111.
Hunter RG, McEwen BS, Pfaff DW. Environmental stress and transposon transcription in the mammalian brain. Mob Genet Elements. 2013;3:e24555. https://doi.org/10.4161/mge.24555.
Johnson R, Guigo R. The RIDL hypothesis: transposable elements as functional domains of long noncoding RNAs. RNA. 2014;20:959–76. https://doi.org/10.1261/rna.044560.114.
McEwen BS, Bowles NP, Gray JD, Hill MN, Hunter RG, Karatsoreos IN, et al. Mechanisms of stress in the brain. Nat Neurosci. 2015;18:1353–63. https://doi.org/10.1038/nn.4086.
Reilly MT, Faulkner GJ, Dubnau J, Ponomarev I, Gage FH. The role of transposable elements in health and diseases of the central nervous system. J Neurosci. 2013;33:17577–86.https://doi.org/10.1523/JNEUROSCI.3369-13.2013.
Bakir F, Damluji SF, Amin-Zaki L, Murtadha M, Khalidi A, al-Rawi NY, et al. Methylmercury poisoning in Iraq. Science. 1973;181:230–41.
Cicero CE, Mostile G, Vasta R, Rapisarda V, Signorelli SS, Ferrante M, et al. Metals and neurodegenerative diseases. A systematic review. Environ Res. 2017;159:82–94. https://doi.org/10.1016/j.envres.2017.07.048.
Combs GF Jr. Selenium in global food systems. Br J Nutr. 2001;85:517–47.
Fang F, Peters TL, Beard JD, Umbach DM, Keller J, Mariosa D, et al. Blood Lead, Bone Turnover, and Survival in Amyotrophic Lateral Sclerosis. Am J Epidemiol. 2017;186:1057–64. https://doi.org/10.1093/aje/kwx176.
Johnson FO, Atchison WD. The role of environmental mercury, lead and pesticide exposure in development of amyotrophic lateral sclerosis. Neurotoxicology. 2009;30:761–5. https://doi.org/10.1016/j.neuro.2009.07.010.
Migliore L, Coppede F. Environmental-induced oxidative stress in neurodegenerative disorders and aging. Mutat Res. 2009;674:73–84. https://doi.org/10.1016/j.mrgentox.2008.09.013.
Pogue AI, Jones BM, Bhattacharjee S, Percy ME, Zhao Y, Lukiw WJ. Metal-sulfate induced generation of ROS in human brain cells: detection using an isomeric mixture of 5- and 6-carboxy-2′,7′-dichlorofluorescein diacetate (carboxy-DCFDA) as a cell permeant tracer. Int J Mol Sci. 2012;13:9615–26. https://doi.org/10.3390/ijms13089615.
Hakansson N, Gustavsson P, Johansen C, Floderus B. Neurodegenerative diseases in welders and other workers exposed to high levels of magnetic fields. Epidemiology. 2003;14:420–6; . discussion 427–428. https://doi.org/10.1097/01.EDE.0000078446.76859.c9.
Cronin S, Greenway MJ, Prehn JH, Hardiman O. Paraoxonase promoter and intronic variants modify risk of sporadic amyotrophic lateral sclerosis. J Neurol Neurosurg Psychiatry. 2007;78:984–6. https://doi.org/10.1136/jnnp.2006.112581.
Diekstra FP, Beleza-Meireles A, Leigh NP, Shaw CE, Al-Chalabi A. Interaction between PON1 and population density in amyotrophic lateral sclerosis. Neuroreport. 2009;20:186–90. https://doi.org/10.1097/WNR.0b013e32831af220.
Matin MA, Hussain K. Striatal neurochemical changes and motor dysfunction in mipafox-treated animals. Methods Find Exp Clin Pharmacol. 1985;7:79–81.
Merwin SJ, Obis T, Nunez Y, Re DB. Organophosphate neurotoxicity to the voluntary motor system on the trail of environment-caused amyotrophic lateral sclerosis: the known, the misknown, and the unknown. Arch Toxicol. 2017;91:2939–52. https://doi.org/10.1007/s00204-016-1926-1.
Morahan JM, Yu B, Trent RJ, Pamphlett R. A gene-environment study of the paraoxonase 1 gene and pesticides in amyotrophic lateral sclerosis. Neurotoxicology. 2007;28:532–40. https://doi.org/10.1016/j.neuro.2006.11.007.
Saeed M, Siddique N, Hung WY, Usacheva E, Liu E, Sufit RL, et al. Paraoxonase cluster polymorphisms are associated with sporadic ALS. Neurology. 2006;67:771–6. https://doi.org/10.1212/01.wnl.0000227187.52002.88.
Sanchez-Santed F, Colomina MT, Herrero Hernandez E. Organophosphate pesticide exposure and neurodegeneration. Cortex. 2016;74:417–26. https://doi.org/10.1016/j.cortex.2015.10.003.
Valdmanis PN, Kabashi E, Dyck A, Hince P, Lee J, Dion P, et al. Association of paraoxonase gene cluster polymorphisms with ALS in France, Quebec, and Sweden. Neurology. 2008;71:514–20. https://doi.org/10.1212/01.wnl.0000324997.21272.0c. 71/7/514 [pii].
Chio A, Benzi G, Dossena M, Mutani R, Mora G. Severely increased risk of amyotrophic lateral sclerosis among Italian professional football players. Brain. 2005;128:472–6. https://doi.org/10.1093/brain/awh373.
Horner RD, Grambow SC, Coffman CJ, Lindquist JH, Oddone EZ, Allen KD, et al. Amyotrophic lateral sclerosis among 1991 Gulf War veterans: evidence for a time-limited outbreak. Neuroepidemiology. 2008;31:28–32. https://doi.org/10.1159/000136648.
Miranda ML, Alicia Overstreet Galeano M, Tassone E, Allen KD, Horner RD. Spatial analysis of the etiology of amyotrophic lateral sclerosis among 1991 Gulf War veterans. Neurotoxicology. 2008;29:964–70. https://doi.org/10.1016/j.neuro.2008.05.005.
Pupillo E, Poloni M, Bianchi E, Giussani G, Logroscino G, Zoccolella S, et al. Trauma and amyotrophic lateral sclerosis: a european population-based case-control study from the EURALS consortium. Amyotroph Lateral Scler Frontotemporal Degener. 2018;19(1-2):118. https://doi.org/10.1080/21678421.2017.1386687.
Szczygielski J, Mautes A, Steudel WI, Falkai P, Bayer TA, Wirths O. Traumatic brain injury: cause or risk of Alzheimer’s disease? A review of experimental studies. J Neural Transm (Vienna). 2005;112:1547–64. https://doi.org/10.1007/s00702-005-0326-0.
Oates N, Pamphlett R. An epigenetic analysis of SOD1 and VEGF in ALS. Amyotroph Lateral Scler. 2007;8:83–6. https://doi.org/10.1080/17482960601149160.
Yang Y, Gozen O, Vidensky S, Robinson MB, Rothstein JD. Epigenetic regulation of neuron-dependent induction of astroglial synaptic protein GLT1. Glia. 2010;58:277–86. https://doi.org/10.1002/glia.20922.
Baker M, Mackenzie IR, Pickering-Brown SM, Gass J, Rademakers R, Lindholm C, et al. Mutations in progranulin cause tau-negative frontotemporal dementia linked to chromosome 17. Nature. 2006;442:916–9. https://doi.org/10.1038/nature05016.
Gass J, Cannon A, Mackenzie IR, Boeve B, Baker M, Adamson J, et al. Mutations in progranulin are a major cause of ubiquitin-positive frontotemporal lobar degeneration. Hum Mol Genet. 2006;15:2988–3001. https://doi.org/10.1093/hmg/ddl241.
Rademakers R, Neumann M, Mackenzie IR. Advances in understanding the molecular basis of frontotemporal dementia. Nat Rev Neurol. 2012;8:423–34. https://doi.org/10.1038/nrneurol.2012.117.
Finch N, Baker M, Crook R, Swanson K, Kuntz K, Surtees R, et al. Plasma progranulin levels predict progranulin mutation status in frontotemporal dementia patients and asymptomatic family members. Brain. 2009;132:583–91. https://doi.org/10.1093/brain/awn352.
Baker M, Litvan I, Houlden H, Adamson J, Dickson D, Perez-Tur J, et al. Association of an extended haplotype in the tau gene with progressive supranuclear palsy. Hum Mol Genet. 1999;8:711–5.
Caffrey TM, Wade-Martins R. The role of MAPT sequence variation in mechanisms of disease susceptibility. Biochem Soc Trans. 2012;40:687–92. https://doi.org/10.1042/BST20120063.
Hoglinger GU, Melhem NM, Dickson DW, Sleiman PM, Wang LS, Klei L, et al. Identification of common variants influencing risk of the tauopathy progressive supranuclear palsy. Nat Genet. 2011;43:699–705. https://doi.org/10.1038/ng.859.
Stefansson H, Helgason A, Thorleifsson G, Steinthorsdottir V, Masson G, Barnard J, et al. A common inversion under selection in Europeans. Nat Genet. 2005;37:129–37.https://doi.org/10.1038/ng1508.
Arai T, Hasegawa M, Akiyama H, Ikeda K, Nonaka T, Mori H, et al. TDP-43 is a component of ubiquitin-positive tau-negative inclusions in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Biochem Biophys Res Commun. 2006;351:602–11.https://doi.org/10.1016/j.bbrc.2006.10.093.
Neumann M, Sampathu DM, Kwong LK, Truax AC, Micsenyi MC, Chou TT, et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science. 2006;314:130–3.
Buratti E, De Conti L, Stuani C, Romano M, Baralle M, Baralle F. Nuclear factor TDP-43 can affect selected microRNA levels. FEBS J. 2010;277:2268–81.https://doi.org/10.1111/j.1742-4658.2010.07643.x.
Emde A, Eitan C, Liou LL, Libby RT, Rivkin N, Magen I, et al. Dysregulated miRNA biogenesis downstream of cellular stress and ALS-causing mutations: a new mechanism for ALS. EMBO J. 2015;34:2633–51. https://doi.org/10.15252/embj.201490493.
Kawahara Y, Mieda-Sato A. TDP-43 promotes microRNA biogenesis as a component of the Drosha and Dicer complexes. Proc Natl Acad Sci U S A. 2012;109:3347–52. https://doi.org/10.1073/pnas.1112427109.
King IN, Yartseva V, Salas D, Kumar A, Heidersbach A, Ando DM, et al. The RNA-binding protein TDP-43 selectively disrupts microRNA-1/206 incorporation into the RNA-induced silencing complex. J Biol Chem. 2014;289:14263–71. https://doi.org/10.1074/jbc.M114.561902.
Li Z, Lu Y, Xu XL, Gao FB. The FTD/ALS-associated RNA-binding protein TDP-43 regulates the robustness of neuronal specification through microRNA-9a in Drosophila. Hum Mol Genet. 2013;22:218–25. https://doi.org/10.1093/hmg/dds420.
Almeida S, Gascon E, Tran H, Chou HJ, Gendron TF, Degroot S, et al. Modeling key pathological features of frontotemporal dementia with C9ORF72 repeat expansion in iPSC-derived human neurons. Acta Neuropathol. 2013;126:385–99. https://doi.org/10.1007/s00401-013-1149-y.
Ciura S, Lattante S, Le Ber I, Latouche M, Tostivint H, Brice A, et al. Loss of function of C9orf72 causes motor deficits in a zebrafish model of Amyotrophic Lateral Sclerosis. Ann Neurol. 2013;74:180. https://doi.org/10.1002/ana.23946.
Donnelly CJ, Zhang PW, Pham JT, Haeusler AR, Mistry NA, Vidensky S, et al. RNA toxicity from the ALS/FTD C9ORF72 expansion is mitigated by antisense intervention. Neuron. 2013;80:415–28. https://doi.org/10.1016/j.neuron.2013.10.015.
Fratta P, Poulter M, Lashley T, Rohrer JD, Polke JM, Beck J, et al. Homozygosity for the C9orf72 GGGGCC repeat expansion in frontotemporal dementia. Acta Neuropathol. 2013;126:401–9. https://doi.org/10.1007/s00401-013-1147-0.
Gendron TF, van Blitterswijk M, Bieniek KF, Daughrity LM, Jiang J, Rush BK, et al. Cerebellar c9RAN proteins associate with clinical and neuropathological characteristics of C9ORF72 repeat expansion carriers. Acta Neuropathol. 2015;130:559–73. https://doi.org/10.1007/s00401-015-1474-4. s00401-015-1474-4 [pii].
Gijselinck I, Van Langenhove T, van der Zee J, Sleegers K, Philtjens S, Kleinberger G, et al. A C9orf72 promoter repeat expansion in a Flanders-Belgian cohort with disorders of the frontotemporal lobar degeneration-amyotrophic lateral sclerosis spectrum: a gene identification study. Lancet Neurol. 2012;11:54–65. https://doi.org/10.1016/S1474-4422(11)70261-7. S1474-4422(11)70261-7 [pii].
Mori K, Weng SM, Arzberger T, May S, Rentzsch K, Kremmer E, et al. The C9orf72 GGGGCC repeat is translated into aggregating dipeptide-repeat proteins in FTLD/ALS. Science. 2013;339:1335–8. https://doi.org/10.1126/science.1232927. science.1232927 [pii].
Russ J, Liu EY, Wu K, Neal D, Suh E, Irwin DJ, et al. Hypermethylation of repeat expanded C9orf72 is a clinical and molecular disease modifier. Acta Neuropathol. 2015;129:39–52. https://doi.org/10.1007/s00401-014-1365-0.
Ritossa F. Discovery of the heat shock response. Cell Stress Chaperones. 1996;1:97–8.
Meaney MJ, Szyf M. Environmental programming of stress responses through DNA methylation: life at the interface between a dynamic environment and a fixed genome. Dialogues Clin Neurosci. 2005;7:103–23.
Caller TA, Doolin JW, Haney JF, Murby AJ, West KG, Farrar HE, et al. A cluster of amyotrophic lateral sclerosis in New Hampshire: a possible role for toxic cyanobacteria blooms. Amyotroph Lateral Scler. 2009;10(Suppl 2):101–8. https://doi.org/10.3109/17482960903278485.
Karlsson O, Roman E, Berg AL, Brittebo EB. Early hippocampal cell death, and late learning and memory deficits in rats exposed to the environmental toxin BMAA (beta-N-methylamino-L-alanine) during the neonatal period. Behav Brain Res. 2011;219:310–20. https://doi.org/10.1016/j.bbr.2011.01.056.
Purdie EL, Samsudin S, Eddy FB, Codd GA. Effects of the cyanobacterial neurotoxin beta-N-methylamino-L-alanine on the early-life stage development of zebrafish (Danio rerio). Aquat Toxicol. 2009;95:279–84. https://doi.org/10.1016/j.aquatox.2009.02.009.
Horvath S. DNA methylation age of human tissues and cell types. Genome Biol. 2013;14:R115. https://doi.org/10.1186/gb-2013-14-10-r115.
Zhang M, Tartaglia MC, Moreno D, Sato C, McKeever P, Weichert A, et al. DNA methylation age-acceleration is associated with disease duration and age at onset in C9orf72 patients. Acta Neuropathol. 2017;134:271–9. https://doi.org/10.1007/s00401-017-1713-y.
Tsankova NM, Berton O, Renthal W, Kumar A, Neve RL, Nestler EJ. Sustained hippocampal chromatin regulation in a mouse model of depression and antidepressant action. Nat Neurosci. 2006;9:519–25. https://doi.org/10.1038/nn1659.
Reul JM, Chandramohan Y. Epigenetic mechanisms in stress-related memory formation. Psychoneuroendocrinology. 2007;32(Suppl 1):S21–5. https://doi.org/10.1016/j.psyneuen.2007.03.016.
Griffiths BB, Hunter RG. Neuroepigenetics of stress. Neuroscience. 2014;275:420–35. https://doi.org/10.1016/j.neuroscience.2014.06.041.
Hunter RG, McEwen BS. Stress and anxiety across the lifespan: structural plasticity and epigenetic regulation. Epigenomics. 2013;5:177–94. https://doi.org/10.2217/epi.13.8.
Reul JM. Making memories of stressful events: a journey along epigenetic, gene transcription, and signaling pathways. Front Psych. 2014;5:00005. https://doi.org/10.3389/fpsyt.2014.00005.
Abel EL. Football increases the risk for Lou Gehrig’s disease, amyotrophic lateral sclerosis. Percept Mot Skills. 2007;104:1251–4. https://doi.org/10.2466/pms.104.4.1251-1254.
Wassenegger M, Heimes S, Riedel L, Sanger HL. RNA-directed de novo methylation of genomic sequences in plants. Cell. 1994;76:567–76.
Ho AS, Turcan S, Chan TA. Epigenetic therapy: use of agents targeting deacetylation and methylation in cancer management. Onco Targets Ther. 2013;6:223–32. https://doi.org/10.2147/OTT.S34680.
Veerappan CS, Sleiman S, Coppola G. Epigenetics of Alzheimer’s disease and frontotemporal dementia. Neurotherapeutics. 2013;10:709–21. https://doi.org/10.1007/s13311-013-0219-0.
Zeier Z, Esanov R, Belle KC, Volmar CH, Johnstone AL, Halley P, et al. Bromodomain inhibitors regulate the C9ORF72 locus in ALS. Exp Neurol. 2015;271:241–50. https://doi.org/10.1016/j.expneurol.2015.06.017.
Koval ED, Shaner C, Zhang P, du Maine X, Fischer K, Tay J, et al. Method for widespread microRNA-155 inhibition prolongs survival in ALS-model mice. Hum Mol Genet. 2013;22:4127–35. https://doi.org/10.1093/hmg/ddt261.
Nolan K, Mitchem MR, Jimenez-Mateos EM, Henshall DC, Concannon CG, Prehn JH. Increased expression of microRNA-29a in ALS mice: functional analysis of its inhibition. J Mol Neurosci. 2014;53:231–41. https://doi.org/10.1007/s12031-014-0290-y.
Morel L, Regan M, Higashimori H, Ng SK, Esau C, Vidensky S, et al. Neuronal exosomal miRNA-dependent translational regulation of astroglial glutamate transporter GLT1. J Biol Chem. 2013;288:7105–16. https://doi.org/10.1074/jbc.M112.410944.
Lakshmaiah KC, Jacob LA, Aparna S, Lokanatha D, Saldanha SC. Epigenetic therapy of cancer with histone deacetylase inhibitors. J Cancer Res Ther. 2014;10:469–78. https://doi.org/10.4103/0973-1482.137937.
Ryu H, Smith K, Camelo SI, Carreras I, Lee J, Iglesias AH, et al. Sodium phenylbutyrate prolongs survival and regulates expression of anti-apoptotic genes in transgenic amyotrophic lateral sclerosis mice. J Neurochem. 2005;93:1087–98. https://doi.org/10.1111/j.1471-4159.2005.03077.x.
Cudkowicz ME, Andres PL, Macdonald SA, Bedlack RS, Choudry R, Brown RH Jr, et al. Phase 2 study of sodium phenylbutyrate in ALS. Amyotroph Lateral Scler. 2009;10:99–106. https://doi.org/10.1080/17482960802320487.
Prudencio M, Belzil VV, Batra R, Ross CA, Gendron TF, Pregent LJ, et al. Distinct brain transcriptome profiles in C9orf72-associated and sporadic ALS. Nat Neurosci. 2015;18:1175–82. https://doi.org/10.1038/nn.4065.
Acknowledgments
We would like to thank Dr. Tamas Ordog, Director of the Epigenomics Translational Program at Mayo Clinic Center for Individualized Medicine for reviewing and providing critical input for this manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG, part of Springer Nature
About this chapter
Cite this chapter
Ebbert, M.T.W., Lank, R.J., Belzil, V.V. (2018). An Epigenetic Spin to ALS and FTD. In: Sattler, R., Donnelly, C. (eds) RNA Metabolism in Neurodegenerative Diseases. Advances in Neurobiology, vol 20. Springer, Cham. https://doi.org/10.1007/978-3-319-89689-2_1
Download citation
DOI: https://doi.org/10.1007/978-3-319-89689-2_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-89688-5
Online ISBN: 978-3-319-89689-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)