Abstract
The nematode worm Caenorhabditis elegans has evolved a sophisticated innate immune response that responds to viral, bacterial, and fungal infections. Because of the lack of an adaptive immune response, including specialized immune cells, the worm has become a powerful model for studying highly conserved basic features of innate immune responses. In this chapter we discuss the signaling pathways that mediate the immune response and features of the worm’s defenses against specific pathogens, as they are currently understood. In particular, recent unique discoveries in C. elegans that have provided valuable details on the interconnectedness between innate immunity, organismal stress resistance, and longevity pathways are highlighted—particularly in the context of how a simple animal can respond to environmental assaults to preserve both its somatic tissue and genetic material.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Caenorhabditis elegans
- Innate immunity
- DNA damage
- MAP kinase
- Stress response
- Insulin signaling
- Toll-like receptor
- p38 signaling
- Viral defense
- RNA interference (RNAi)
- p53 tumor suppressor
- Surveillance immunity
- Cellular homeostasis
Introduction
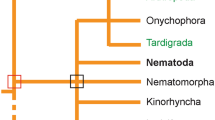
Caenorhabditis elegans is a small , generally free-living , non-parasitic nematode whose natural habitats include soil, compost, rotting fruit, and snails. The worm has been a powerful model for the study of many important biological processes, many of which can be very effectively transferred into studies of higher organisms. For example, the core apoptotic signaling pathway was first dissected in the worm (Horvitz and Lecture 2002), and a mutant screen revealed the first longevity-control pathway (Kenyon et al. 1993). Much of the power of C. elegans in basic research comes from its small size and easy handling in the laboratory, as well as a vast array of resources and infrastructure, including mutant and RNA interference libraries, a very active and open community of researchers, facilities for strain collection and distribution, and data curation. Another remarkable feature of the worm that increases its power as a model system is that the adult worms (which develop via the transition between four main larval stages), with the exception of the germline, are entirely post-mitotic. Following the fourth (and final) larval stage, further development consists only of growth, with no additional change in the number of somatic cells. Furthermore, the number and identity of the cells are invariable from worm to worm, and the full lineage for each cell, from fertilized oocyte to terminal differentiation, has been completely dissected. For this reason, some of the complexities of working with a multicellular organism in which cells turn over quickly are eliminated. The worm has been found to be susceptible to several human pathogens, including Pseudomonas aeruginosa, Serratia marcescens, Salmonella enterica, and Enterococcus faecalis, as well as fungi, including Cryptococcus neoformans (Marsh and May 2012). This feature of the worms’ response to pathogens spawned the productive and active field of nematode innate immunity. Furthermore, several nematode-specific pathogens have been identified, which allow the analysis of aspects of the immune response that may be part of the natural existence of the animals. These organisms include the bacteria Microbacterium nematophilum (Hodgkin et al. 2000), the fungus Drechmeria coniospora (Jansson 1994), and a positive-strand RNA virus (Orsay virus ) (Félix et al. 2011).
The C. elegans immune system (see the overview in Fig. 1) evolutionarily predates those of higher organisms and seems to be relatively simple, particularly in that it lacks an adaptive immune system. Furthermore, the worms have no specialized or mobile immune cells. While they possess three pairs of cells involved in detoxification (the coelomocytes), these cells have not been assigned any immune functions. Because the worm relies only on its innate immune response to resist and tolerate pathogens, the complex interactions that exist between innate and adaptive immune responses in higher organisms do not cloud the study of specific features of innate immunity.
Overview of the Caenorhabditis elegans innate immune response . The worm innate immune response consists of both cell-autonomous processes (a) and systemically disseminated responses (b). The transcription-based response can be stimulated via disruption of cellular processes (surveillance mediated immunity) or by direct cellular damage (damage-associated molecular pattern [DAMP]-mediated immunity). Signals from neurons appear to limit the immune response and DNA damage in the germ cells stimulates a robust immune response; however, most of the molecular mechanisms for these pathways remain to be clarified
Signaling Pathways in the Caenorhabditis elegans Innate Immune Response
The worm innate immune response generally occurs at the level of transcriptional regulation (Shivers et al. 2008) and is controlled by several signaling cascades, depending on the type and location of the pathogen challenge. To date, at least four pathways have been identified:
-
1.
p38/PMK-1 signaling
-
2.
ERK/MPK-1 signaling
-
3.
Insulin-like signaling (DAF-16 and DAF-2)
-
4.
DBL-1 pathway
The p38/PMK-1 Signaling Pathway
Genetic screens to identify mutants with increased sensitivity to P. aeruginosa uncovered the p38 mitogen-activated protein kinase (MAPK)–related pathway as an important regulator of immunity in the worm (Bolz et al. 2010; Kim et al. 2004; Troemel et al. 2006). The signaling cascade underlying this pathway consists of several players: the neuronal symmetry family member 1 (NSY-1), stress-activated protein kinase (SAPK)/extracellular signal–regulated kinase (ERK) kinase 1 (SEK-1), and PMK-1, the worm p38 homolog. Signal propagation occurs via sequential phosphorylation of SEK-1 by NSY-1 and PMK-1 by SEK-1 in a strictly linear fashion (Fig. 2). The primary downstream effector of the pathway is the transcription factor ATF-7, which, when activated, induces a large repertoire of putative immune factors (Pujol et al. 2001; Shivers et al. 2010). As discussed in more detail in section “Missing Links”, the nematode’s genome encodes only one Toll-like receptor (TLR) protein (TOL-1) , which has been only loosely associated with the immune response (Tenor and Aballay 2007). In mammals, all characterized TLRs seem to rely on adaptor proteins that contain Toll/interleukin-1 receptor (TIR) domains for the signal transduction that ultimately leads to the nuclear factor (NF)-κB-mediated pro-inflammatory response (i.e., TRAM [Trif-related adaptor molecule], TICAM [TIR domain-containing adapter molecule]/Trif [TIR-domain-containing adapter-inducing interferon-β], TIRAP [TIR domain-containing adapter protein]/Mal, and myeloid differentiation primary response gene 88 [MyD88]). A fifth TIR-containing protein, SARM (sterile alpha and TIR motif-containing protein), also exists and, while it is the least understood in mammals, it is also the only one that has a direct ortholog in C. elegans (tir-1) (Liberati et al. 2004). The tir-1 gene does, in fact, have important roles in worm immunity: in particular, it acts together with NSY-1 and SEK-1 in the p38 pathway . TIR-1, NSY-1, and SEK-1 can be co-immunoprecipitated and probably form a protein complex (Chuang and Bargmann 2004), and phosphorylation of p38/PMK-1 depends on tir-1 (Liberati et al. 2004); thus, it seems to be firmly situated in the p38 signaling pathway. Data suggest that TIR-1 is likely upstream in the pathway (Liberati et al. 2004), placing it in closer proximity to the initiating events of the signaling cascade; however, what signals lead to TIR-1 activation remain entirely unknown.
Signaling in the Caenorhabditis elegans innate immune response. The worm immune response consists of three currently known signaling pathways, the p38 axis (C. elegans PMK-1), the DBL-1 axis, and the extracellular signal-regulated kinase (ERK) axis (C. elegans MPK-1). The outcomes of these pathways following pathogen exposure are the transcriptional induction of many genes thought to be involved in pathogen tolerance and resistance, although the functions of these genes remain mostly unknown (see text)
The ERK/MPK-1 Signaling Pathway
The p38 MPK signaling cascade seems to be the most important in terms of innate immune function in C. elegans; however, the ERK1/ERK2 MAPK homolog MPK-1 can also be activated by some pathogens, in particular infection by M. nematophilum leads to an MPK-1-dependent immune response (Gravato-Nobre et al. 2005; Hodgkin et al. 2000; Nicholas and Hodgkin 2004). The upstream factors in this signaling cascade are LIN-45 and MEK-2, which have also been assigned various functions in development and fertility (Fig. 2). Thus, it is clear that this signaling cascade is not exclusively dedicated to the immune response pathway, but instead regulates a plethora of processes ranging from stress responses to multiple aspects of the animal’s development. The most upstream element of the pathway and the terminal effector protein remain unknown.
Insulin-Like Signaling (DAF-16 and DAF-2)
The insulin-like signaling (IIS) pathway involving the insulin-like growth factor receptor DAF-2 and the forkhead box O (FOXO) transcription factor DAF-16 were first identified as regulators of the extraordinarily long-lived dauer larval stage (Kenyon et al. 1993), an alternative developmental fate of worms under starvation stress, as well as determinants of adult longevity in C. elegans. Subsequently, they have been shown to be part of a larger network of pathways that confer stress resistance, which is intimately intertwined with the worm’s immune response (Cezairliyan et al. 2013; Mahajan-Miklos et al. 1999). In the presence of its ligand DAF-28, DAF-2 is activated, which goes on to activate the phosphatidylinositol-3 OH kinase AGE-1 (Li et al. 2003; Malone et al. 1996). AGE-1 then catalyzes the conversion of phosphatidylinositol bisphosphate (PIP2) into phosphatidylinositol triphosphate (PIP3) (Tazearslan et al. 2009). PIP3 then binds to the AKT-1/AKT-2 complex to reveal two phosphorylation sites that are phosphorylated by the PDK-1 kinase (which also depends on PIP3 binding for its function). The AKT complex then phosphorylates the transcription factor DAF-16, which is blocked from entering the nucleus (Paradis and Ruvkun 1998). In contrast, in the presence of an antagonistic ligand (e.g., INS-1 [Insulin-like peptide]), the pathway is inactive and DAF-16 is not phosphorylated, leading to its translocation into the nucleus, where it activates stress response and putative antimicrobial genes. Genetic inactivation of daf-2 leads to the same outcome as DAF-16 remains constitutively hypophosphorylated. Not entirely unexpectedly, loss of daf-2 leads to pathogen resistance and this effect seems to be primarily rooted in the intestinal cells (Garsin et al. 2003; Hsin and Kenyon 1999; Libina et al. 2003; Lin et al. 2001). As is the case for ERK signaling, the outcomes of DAF-16 activation extend far beyond immune function, indicating that the pathway is not a dedicated immune pathway.
The DBL-1 Signaling Pathway
The gene dbl-1 encodes one of four transforming growth factor (TGF)-β-like ligands in C. elegans and is (in part) required for resistance to both P. aeruginosa and S. marcescens (Kurz and Tan 2004; Mallo et al. 2002). The DBL-1 protein binds to the DAF-4/SMA-6 heterodimeric receptor and, via the SMA-2/SMA-3/SMA-4 complex, controls gene expression levels (Fig. 2), while it also has diverse functions independent of immunity (e.g., body size regulation and structural patterning). In fact, loss of the sma genes leads to increased sensitivity to P. aeruginosa (Kurz and Tan 2004; Mallo et al. 2002; Roberts et al. 2010). Interestingly, TGF-β signaling in mammals leads to immunosuppression, demonstrating a remarkable divergence of function during evolution.
The C. elegans Viral Defense Strategy
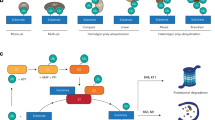
To date, only a single virus that can infect and replicate in C. elegans has been identified (Félix et al. 2011). This virus , called Orsay after the site of its discovery in France, is a member of the Nodaviridae family and is a positive-strand RNA virus. Infection leads to easily observable morphological defects in the worm’s intestine. The first indication of an antiviral response came from the observation that viral load was increased when factors of the RNAi pathway were deactivated, such as RDE-1, RDE-4, MUT-7 (RNaseD), and DRH-1 (Dicer) (Ashe et al. 2013; Félix et al. 2011; Guo et al. 2013). The implication of the RNAi pathway in viral defense (also supported by viral infection experiment using isolated worm cells) provided a sturdy scaffold for considering the evolutionary origins and conservation of the pathway, which is consistent with its function in plants. The current model for antiviral immunity in C. elegans (Fig. 3) proposes that the double-stranded RNA (dsRNA) intermediates produced during viral replication are bound by the dsRNA-binding complex RDE-1/RDE-4 responsible for initial detection and sequestration of exogenous dsRNAs. The canonical RNAi pathway is subsequently recruited: the dsRNA is passed to the DExD box RNA helicase DRH-1, which when interacting with RDE-1/RDE-4 unwinds the molecule to provide accessibility by the dicing complex. The Dicer homolog DCR-1 then produces small RNAs that go on to serve as templates for the RNA-directed RNA polymerase to produce a pool of secondary antisense small RNAs, which mediate the degradation of the full length viral RNA genome.
The Caenorhabditis elegans antiviral response . Protection against viral invasion is mediated by components of the RNA interference (RNAi)-mediated gene-silencing pathway . Following infection by an RNA-based virus, the RNA is bound and processed to yield antisense small interfering RNAs (siRNAs) against the viral genome, ultimately leading to degradation of the viral genetic material to limit the infection
Some mammalian viruses have mechanisms to avoid detection by the host immune system; for example, by blocking the major histocompatibility complex (MHC) class I antigen processing and presentation pathway to escape T-killer cells (Horst et al. 2011). In C. elegans, the Flock house virus protein B2 can robustly downregulate the RNAi machinery to increase the sensitivity of worms to Orsay virus (Guo and Lu 2013). Another shared element of mammalian and nematode antiviral pathways is the similarity between DRH-1 and the RIG-I (retinoic acid inducible gene I) RNA helicase, an important sensor of dsRNA in mammals (Ashe et al. 2013; Coffman et al. 2017; Guo et al. 2013). While the RNA-binding domains are highly similar, the proteins do have different functions: DRH-1 presents RNA to Dicer for processing, while RIG-I activates an inflammatory antiviral immune response. Nevertheless, it is plausible that the two antiviral responses are connected over the span of evolutionary time.
Neuronal Regulation of Immunity
The simple body plan and limited number and type of cells has led to functional multitasking in various cell types. As already mentioned, the intestinal cells are a key site of immune function; it turns out that neurons are also key players in worm immunity . C. elegans is an extremely powerful model for studying the nervous system, as the morphology, identity, and synaptic connectivity of all its 302 neurons is entirely understood; furthermore, a detailed catalog of the relevant neurotransmitters for most of the neurons has also been compiled. In addition to the production of antimicrobial factors, worms also seem to respond to pathogen exposure via neuron-driven behavioral programs, most notably pathogen avoidance. A polymorphism in the gene encoding the G-protein-coupled receptor (GPCR) protein NPR-1 caused decreased survival during P. aeruginosa infection by limiting the ability of the worms to avoid the pathogen (Reddy et al. 2009); however, it turned out that this is not the sole function of npr-1 in worm immunity. Worms lacking NPR-1 exhibited altered expression of intestinally expressed, PMK-1-regulated genes during infection (Styer et al. 2008). Remarkably, elimination of the sensory neurons AQR, RQR, and URX rescued this phenotype, suggesting that in the absence of NPR-1 these neurons become hyperactive and disturb immune pathways. Worms lacking these neurons are more pathogen resistant, suggesting that they have a negative regulatory function on the immune response.
The neuron-expressed GPCR OCTR-1 also plays a role in worm innate immunity (Sun et al. 2011). Through action in the ASH and ASI neurons, OCTR-1 suppresses the pathogen-dependent activation of PMK-1 and blocks the induction of a non-canonical UPR in distal tissues, indicating a non-cell-autonomous function. While the increased resistance of npr-1 mutant worms to P. aeruginosa involved alterations in pathogen avoidance behavior, octr-1 mutant worms exhibit increased pathogen resistance without such a behavioral change. How the neurons are stimulated by pathogens and the mechanisms underlying these phenotypes remain open questions in the field. Interestingly, the OCTR-1 protein is related to vertebrate adrenergic receptors that bind to their ligand noradrenalin. The outcome of this binding is a response to acute stress that can be accompanied by immune suppression (Aballay 2013).
The Interface Between Innate Immunity and DNA Damage
Study of the innate immune response of C. elegans has primarily focused on host–pathogen interactions; however, it is also now clear that the system can also respond to damaged self-DNA through a process called germline DNA damage-induced systemic stress resistance, or GDISR (Fig. 4) (Ermolaeva et al. 2013). C. elegans has been an especially valuable tool for dissecting the distinctive features and roles of DNA damage responses (DDRs) in the germline versus somatic tissues. In this discussion, we focus on germline processes, as they have so far been shown to be intricately intertwined with the innate immunity of C. elegans.
Non-cell-autonomous stress resistance following immune induction. DNA damage in germline cells leads to the activation of immune genes, stimulating the ubiquitin proteosome system (UPS). This enhanced activity confers systemic stress and pathogen resistance. The mechanism for the detection of the DNA damage and the intermediate signaling pathway leading to gene induction remains unknown
The majority of tissues in the adult worm are post-mitotic, as the cellular lineages are invariable and somatic development is generally completed by the last larval stage. The exception is the germline, which contains mitotic cells in a stem cell niche. Once cells leave the stem cell niche, they proceed through meiosis to generate mature germ cells—in hermaphrodites, the most common sex in worms—the production of sperm and oocytes are temporally separated during growth. In hermaphrodites, diakinesis-arrested oocytes are fertilized by sperm produced earlier during development to generate clonal offspring (Kimble and Crittenden 2005). Following DNA damage in germ cells, conserved cell cycle checkpoints are robustly activated to arrest mitotic proliferation, allowing time for DNA repair pathways to remove destabilizing lesions (Ahmed and Hodgkin 2000; Gartner et al. 2000). In a case where checkpoint activation cannot be resolved in a timely manner, apoptosis mediated by the CEP-1, the C. elegans homolog of the highly conserved p53 tumor suppressor , occurs through CEP-1-dependent transcriptional induction of the BH3-only-domain genes egl-1 and ced-13, the protein products of which then trigger the apoptosome (Derry et al. 2001; Hofmann et al. 2002; Schumacher et al. 2001, 2005).
These processes are themselves cell autonomous ; however, it is now clear that the GDISR pathway leads to non-cell-autonomous effects via factors associated with the innate immune response (Ermolaeva et al. 2013). The systemic aspects of the DDR were initially observed in animals that were defective for global-genome nucleotide excision repair, which fail to remove UV light-induced lesions in germ cells, ultimately leading to accumulation of DNA damage. Quite remarkably, the somatic tissues of UV-exposed animals developed profound resistance to both heat and oxidative stress. Importantly, this effect was not specific to UV-induced damage, as DNA damage induced by ionizing radiation or hydroxyurea, as well as endogenously generated DNA double-strand breaks in pachytene germ cells, was sufficient to elicit the stress resistance phenotypes. This induction of stress resistance through damage of endogenous DNA suggests that damaged DNA can be recognized as a damage-associated molecular pattern (DAMP) by C. elegans. The molecular basis of GDISR was shown to depend on the activation of the ERK homolog MPK-1, which subsequently induced expression of a repertoire of immune genes. This added burden of such a broad induction of gene expression subsequently stimulated the ubiquitin proteasome system (UPS), which conferred systemic stress resistance. Importantly, this pathway is distinct from the p38/PMK-1 pathway discussed earlier and the first tendency may be to attribute the effect to CEP-1 activity; however, this was not the case and, in fact, no components of the canonical DNA damage checkpoint signaling were necessary for GDISR. Therefore, GDISR is a distinct response to DNA damage , independent of checkpoint signaling. A connection between UV irradiation and immunity has been demonstrated in human skin cells where UV irradiation leads to a complex and highly coordinated range of immune-associated processes, ranging from localized inflammation to systemic outcomes mediated by cytokines and other growth factors—highly reminiscent of GDISR. It is likely that further study of GDISR in C. elegans may lead to the discovery of fundamental features of such systemic responses and how damaged DNA is recognized as a DAMP. Immune reactions to DNA damage have also been reported following infection with bacteria such as Escherichia coli and Helicobacter pylori, which can both cause DNA damage in eukaryotic cells (Nougayrede et al. 2006; Toller et al. 2011), suggesting that GDISR—at least conceptually—may be an ancestral version of extant mammalian pathways.
Missing Links
A conspicuously missing component of innate immunity in C. elegans is a repertoire of mechanisms for the detection of pathogen-associated molecular patterns (PAMPs) and DAMPs. In higher organisms, a number of pathways have been characterized for the specific detection of a broad list of foreign elements (e.g., Toll-like signaling, cGAS-STING [cyclic GMP–AMP synthase–stimulator of interferon genes], among others). Despite efforts by a number of groups over many years, similar pathways have not been identified in the nematode.
The sole putative DAMP detection mechanism identified in the worm to date is DCAR-1, a GPCR protein that was previously assigned a function in chemosensory neurons (Zugasti et al. 2014). It was subsequently shown to be expressed in the epidermis, a major site of innate immune responses in the worm. In this tissue, DCAR-1 can detect the tyrosine derivative hydroxyphenyllactic acid (HPLA), which accumulates when worms experience wounding or fungal infection. In these situations, HPLA accumulates triggering a signaling cascade that culminates in the canonical p38 (PMK-1)-mediated innate immune response.
While the C. elegans gene encodes one TLR, TOL-1, it has not been assigned a role in detection; rather, it seems to be involved primarily in developmental processes and tissue maintenance (Pujol et al. 2001; Tenor and Aballay 2007). Furthermore, the genome also encodes a large collection of leucine rich repeat (LRR)-containing proteins. It is reasonable to conjecture that such genes may function in detection based on the well-characterized functions of LRRs in ligand binding in the Toll-like and nucleotide-binding oligomerization domain (NOD)-like receptors; however, while one LRR protein has been assigned a function in pathogen resistance (FSHR [follicle-stimulating hormone receptor]-1) (Miller et al. 2015; Powell et al. 2009), it has not been shown to function as a sensor.
As discussed in the section “The Interface Between Innate Immunity and DNA Damage”, worms induce a potent innate immune response to DNA damage in the germline that then confers systemic somatic stress resistance (GDISR) . Pathways for the detection of damaged DNA, including double-strand breaks, single-strand breaks, and stalled replication forks at sites of damage have been identified and well-characterized in C. elegans. Quite unexpectedly, such pathways were clearly shown not to be involved in the induction of this immune response (Ermolaeva et al. 2013).
Another significant gap in our understanding of worm innate immunity is the function of the specific proteins induced as part of the innate immune response. As discussed earlier, the C. elegans innate immune response is characterized in general by the transcriptional upregulation of a large regulon of genes that encode putative secreted factors, many of which have structures indicating that they may function as antimicrobial agents; however, to date we know very little about how they function to combat pathogenic challenges. The one example in which a specific function is coming into focus is a C-type lectin gene (of which the genome encodes hundreds), which is induced following exposure to the Gram-positive bacterial pathogen Bacillus thuringiensis (Pees et al. 2017). Mutants for several C-lectin genes were shown to have either decreased or, surprisingly, increased resistance to the pathogen . Specifically, loss of one gene in particular resulted in enhanced avoidance behavior, which prompts the worms to leave the lawns of the pathogen. Furthermore, the same mutant animals also had increased periods of feeding cessation; thus, in this case, an immune-regulated gene was shown to actually be a negative regulator of pathogen resistance via behavioral modulation. Even with this bit of insight, the roles not only of the C-lectin proteins but essentially of the other immune-regulated genes remain entirely unknown. The elucidation of their functions is particularly hampered by the dazzling similarities between many of the genes, likely resulting in robust redundancy. Overcoming this experimental challenge remains a stubborn block in furthering research in this area.
Surveillance-Mediated Immunity
As mentioned in the section “Missing Links”, our understanding of the sensors and effectors of the C. elegans innate immune response remains rudimentary, as several fundamental components have yet to be identified, most notably pathways for sensing PAMPs and DAMPs. One theory proposed to resolve this issue is that the nematode relies on an alternative approach for immune activation, independent of direct sensing, called “surveillance immunity” (Pukkila-Worley 2016)—similar in principle to the long-studied effect-triggered immunity in plants. The basis for surveillance immunity is that instead of monitoring for pathogens directly, the animals monitor for disruptions in endogenous processes that could be caused by the presence of a pathogen, for example, translation, cellular homeostasis , or structural integrity.
The first identified and best understood surveillance pathway is involved in monitoring host translation. Many bacteria can produce protein toxins that interfere with efficient translation of mRNAs, including ToxA, produced by P. aeruginosa. This protein blocks polymerization of nascent peptides by blocking host elongation factor 2 function via ribosylation in intestinal cells following P. aeruginosa infection (Dunbar et al. 2012). Following this disruption, the transcription factor ZIP-2 (bZip transcription factor) accumulates and, in concert with the conserved protein CEBP-2, regulates an innate immune response (Estes et al. 2010; McEwan et al. 2012; Reddy et al. 2016). What is clear is that both the pathogen and toxin are functionally invisible to the animals and instead disruption of translational function stimulates the response. Quite remarkably, genetic ablation of host-encoded functions can induce a similar response, even in the absence of a pathogen, reinforcing this validity of this concept.
A conceptually similar pathway has also been reported in the mitochondria. Siderophores, toxins produced by pathogens (including P. aeruginosa), can interfere with mitochondrial homeostasis (Kirienko et al. 2015). The unfolded protein response in the mitochondria (UPRmt) helps to ensure mitochondrial function by inducing the expression of nuclear-encoded, mitochondrially targeted chaperone molecules. A central player in this pathway is the transcription factor ATFS-1. ATFS-1 is normally taken up by functionally intact mitochondria, thus limiting cytosolic levels; however, upon disruption of the mitochondria, this uptake is reduced, leading to cytosolic accumulation. Subsequently, the protein can enter the nucleus, where it induces a repertoire of genes encoding putative antimicrobial factors. ATFS-1 also enters the nucleus during P. aeruginosa infection, leading to the expression of genes that confer resistance to the infection (Nargund et al. 2012), while loss of ATFS-1 leads to reduced resistance (Pellegrino et al. 2014). Further work remains to fully understand the interplay between the UPRmt, ATFS-1, and bacterial infection, but as in the case of translation, this pathway provides a satisfying mechanism by which the animals can indirectly sense the presence of a pathogen.
In a large-scale study of the microbiome of the worm’s natural habitat , nearly 20% of isolates examined (a total of 560) induced mitochondrial stress (Liu et al. 2014). This observation strongly supports the broad usefulness of such a surveillance pathway in responding to bacterial challenges. Much work remains to be done to understand the implications and complexity of surveillance immunity, but the concept is already becoming a useful framework in which to consider the innate immune response of worms, given the gaps in the more conventional mechanistic pathways.
To What End: Immunity or General Stress Resistance?
Expression of genes associated with the C. elegans innate immune response can be controlled by pathways that are generally discussed in the context of distinct biological processes (i.e., MAPK signaling in response to pathogen infection and DNA damage and DAF-16 as part of the IIS pathway). Furthermore, activation of overlapping innate immune genes by DNA damage confers resistance not only to pathogens but also to heat and oxidative stresses (Ermolaeva et al. 2013). While the latter two cases seem to be secondary effects due to activation of the UPS driven by the enhanced expression of putative immune factors, rather than effected directly by the immune peptides, the net outcome of the activation of the innate immune response is enhanced stress resistance.
An important conceptual consideration is whether what is studied in C. elegans and labeled as “innate immunity” is rather a complex set of interconnected stress responses , which happen to confer pathogen resistance. The label applied to these responses certainly does not negate the value and usefulness of this field of research in C. elegans as broadly applicable biological processes have been clarified; however, oversimplification of the conceptualization of these responses could lead to missed opportunities for study and interpretation of results. Importantly, however, stress responses appear to comprise an essential component of not only ancestral but also of mammalian immune responses, such as when natural killer cells need to survive their own rampage against infections, during which they produce reactive oxygen species. The nematode might therefore turn out to be particularly instructive for the understanding of how the stress responses could balance the consequences of immune defenses. Given their intimate involvement in the regulation of longevity, stress response pathways could play a central role in alleviating the consequences of innate immunity driving the chronic inflammation during human aging. Deeper insight into the regulation of stress responses during the activation of innate immunity in C. elegans might therefore yield new conceptual avenues for counteracting the pathological consequences of chronic inflammation.
Future work on the responses of C. elegans to environmental challenges, from pathogens to chemicals, and even to radiation, will surely shed light on these questions and provide new and exciting avenues for further research.
References
Aballay A (2013) Role of the nervous system in the control of proteostasis during innate immune activation: insights from C. elegans. PLoS Pathog 9:e1003433. https://doi.org/10.1371/journal.ppat.1003433
Ahmed S, Hodgkin J (2000) MRT-2 checkpoint protein is required for germline immortality and telomere replication in C. elegans. Nature 403:159–164. https://doi.org/10.1038/35003120
Ashe A, Bélicard T, Le Pen J, Sarkies P, Frézal L, Lehrbach NJ, Félix M-A, Miska EA (2013) A deletion polymorphism in the Caenorhabditis elegans RIG-I homolog disables viral RNA dicing and antiviral immunity. elife 2:e00994. https://doi.org/10.7554/eLife.00994
Bolz DD, Tenor JL, Aballay A (2010) A conserved PMK-1/p38 MAPK is required in Caenorhabditis elegans tissue-specific immune response to Yersinia pestis infection. J Biol Chem 285:10832–10840. https://doi.org/10.1074/jbc.M109.091629
Cezairliyan B, Vinayavekhin N, Grenfell-Lee D, Yuen GJ, Saghatelian A, Ausubel FM (2013) Identification of Pseudomonas aeruginosa phenazines that Kill Caenorhabditis elegans. PLoS Pathog 9:e1003101. https://doi.org/10.1371/journal.ppat.1003101
Chuang C-F, Bargmann CI (2004) A Toll-interleukin 1 repeat protein at the synapse specifies asymmetric odorant receptor expression via ASK1 MAPKKK signaling. Genes Dev 19:270–281. https://doi.org/10.1101/gad.1276505
Coffman SR, Lu J, Guo X, Zhong J, Jiang H, Broitman-Maduro G, Li W-X, Lu R, Maduro M, Ding S-W (2017) Caenorhabditis elegans RIG-I homolog mediates antiviral RNA interference downstream of dicer-dependent biogenesis of viral small interfering RNAs. MBio 8:e00264–e00217. https://doi.org/10.1128/mBio.00264-17
Derry WB, Putzke AP, Rothman JH (2001) Caenorhabditis elegans p53: role in apoptosis, meiosis, and stress resistance. Science 294:591–595. https://doi.org/10.1126/science.1065486
Dunbar TL, Yan Z, Balla KM, Smelkinson MG, Troemel ER (2012) C. elegans detects pathogen-induced translational inhibition to activate immune signaling. Cell Host Microbe 11:375–386. https://doi.org/10.1016/j.chom.2012.02.008
Ermolaeva MA, Segref A, Dakhovnik A, Ou H-L, Schneider JI, Utermöhlen O, Hoppe T, Schumacher B (2013) DNA damage in germ cells induces an innate immune response that triggers systemic stress resistance. Nature 501:416–420. https://doi.org/10.1038/nature12452
Estes KA, Dunbar TL, Powell JR, Ausubel FM, Troemel ER (2010) bZIP transcription factor zip-2 mediates an early response to Pseudomonas aeruginosa infection in Caenorhabditis elegans. Proc Natl Acad Sci U S A 107:2153–2158. https://doi.org/10.1073/pnas.0914643107
Félix M-A, Ashe A, Piffaretti J, Wu G, Nuez I, Bélicard T, Jiang Y, Zhao G, Franz CJ, Goldstein LD, Sanroman M, Miska EA, Wang D (2011) Natural and experimental infection of Caenorhabditis nematodes by novel viruses related to nodaviruses. PLoS Biol 9:e1000586. https://doi.org/10.1371/journal.pbio.1000586
Garsin DA, Villanueva JM, Begun J, Kim DH, Sifri CD, Calderwood SB, Ruvkun G, Ausubel FM (2003) Long-lived C. elegans daf-2 mutants are resistant to bacterial pathogens. Science 300:1921. https://doi.org/10.1126/science.1080147
Gartner A, Milstein S, Ahmed S, Hodgkin J, Hengartner MO (2000) A conserved checkpoint pathway mediates DNA damage--induced apoptosis and cell cycle arrest in C. elegans. Mol Cell 5:435–443
Gravato-Nobre MJ, Nicholas HR, Nijland R, O'Rourke D, Whittington DE, Yook KJ, Hodgkin J (2005) Multiple genes affect sensitivity of Caenorhabditis elegans to the bacterial pathogen Microbacterium nematophilum. Genetics 171:1033–1045. https://doi.org/10.1534/genetics.105.045716
Guo X, Lu R (2013) Characterization of virus-encoded RNA interference suppressors in Caenorhabditis elegans. J Virol 87:5414–5423. https://doi.org/10.1128/JVI.00148-13
Guo X, Zhang R, Wang J, Ding S-W, Lu R (2013) Homologous RIG-I-like helicase proteins direct RNAi-mediated antiviral immunity in C. elegans by distinct mechanisms. Proc Natl Acad Sci U S A 110:16085–16090. https://doi.org/10.1073/pnas.1307453110
Hodgkin J, Kuwabara PE, Corneliussen B (2000) A novel bacterial pathogen, Microbacterium nematophilum, induces morphological change in the nematode C. elegans. Curr Biol 10:1615–1618
Hofmann ER, Milstein S, Boulton SJ, Ye M, Hofmann JJ, Stergiou L, Gartner A, Vidal M, Hengartner MO (2002) Caenorhabditis elegans HUS-1 is a DNA damage checkpoint protein required for genome stability and EGL-1-mediated apoptosis. Curr Biol 12:1908–1918
Horst D, Verweij MC, Davison AJ, Ressing ME, Wiertz EJHJ (2011) Viral evasion of T cell immunity: ancient mechanisms offering new applications. Curr Opin Immunol 23:96–103. https://doi.org/10.1016/j.coi.2010.11.005
Horvitz HR, Lecture N (2002) Worms, life and death. pp 320–351. https://doi.org/10.1002/cbic.200300614
Hsin H, Kenyon C (1999) Signals from the reproductive system regulate the lifespan of C. Elegans. Nature 399(6734):362–366
Jansson HB (1994) Adhesion of conidia of Drechmeria coniospora to Caenorhabditis elegans wild type and mutants. J Nematol 26:430–435
Kenyon C, Chang J, Gensch E, Rudner A, Tabtiang R (1993) A C. elegans mutant that lives twice as long as wild type. Nature 366:461–464. https://doi.org/10.1038/366461a0
Kim DH, Liberati NT, Mizuno T, Inoue H, Hisamoto N, Matsumoto K, Ausubel FM (2004) Integration of Caenorhabditis elegans MAPK pathways mediating immunity and stress resistance by MEK-1 MAPK kinase and VHP-1 MAPK phosphatase. Proc Natl Acad Sci U S A 101:10990–10994. https://doi.org/10.1073/pnas.0403546101
Kimble J, Crittenden SL (2005) Germline proliferation and its control. Worm Book 1–14. https://doi.org/10.1895/wormbook.1.13.1
Kirienko NV, Ausubel FM, Ruvkun G (2015) Mitophagy confers resistance to siderophore-mediated killing by Pseudomonas aeruginosa. Proc Natl Acad Sci U S A 112:1821–1826. https://doi.org/10.1073/pnas.1424954112
Kurz CL, Tan M-W (2004) Regulation of aging and innate immunity in C. elegans. Aging Cell 3:185–193. https://doi.org/10.1111/j.1474-9728.2004.00108.x
Li W, Kennedy SG, Ruvkun G (2003) daf-28 encodes a C. elegans insulin superfamily member that is regulated by environmental cues and acts in the DAF-2 signaling pathway. Genes Dev 17:844–858. https://doi.org/10.1101/gad.1066503
Liberati NT, Fitzgerald KA, Kim DH, Feinbaum R, Golenbock DT, Ausubel FM (2004) Requirement for a conserved Toll/interleukin-1 resistance domain protein in the Caenorhabditis elegans immune response. Proc Natl Acad Sci U S A 101:6593–6598. https://doi.org/10.1073/pnas.0308625101
Libina N, Berman JR, Kenyon C (2003) Tissue-specific activities of C. elegans DAF-16 in the regulation of lifespan. Cell 115:489–502
Lin K, Hsin H, Libina N, Kenyon C (2001) Regulation of the Caenorhabditis elegans longevity protein DAF-16 by insulin/IGF-1 and germline signaling. Nat Genet 28:139–145
Liu Y, Samuel BS, Breen PC, Ruvkun G (2014) Caenorhabditis elegans pathways that surveil and defend mitochondria. Nature 508:406–410. https://doi.org/10.1038/nature13204
Mahajan-Miklos S, Tan MW, Rahme LG, Ausubel FM (1999) Molecular mechanisms of bacterial virulence elucidated using a Pseudomonas aeruginosa-Caenorhabditis elegans pathogenesis model. Cell 96:47–56
Mallo GV, Kurz CL, Couillault C, Pujol N, Granjeaud S, Kohara Y, Ewbank JJ (2002) Inducible antibacterial defense system in C. elegans. Curr Biol 12:1209–1214
Malone EA, Inoue T, Thomas JH (1996) Genetic analysis of the roles of daf-28 and age-1 in regulating Caenorhabditis elegans dauer formation. Genetics 143:1193–1205. https://doi.org/10.1016/0168-9525(96)81474-0
Marsh EK, May RC (2012) Caenorhabditis elegans, a model organism for investigating immunity. Appl Environ Microbiol 78:2075–2081. https://doi.org/10.1128/AEM.07486-11
McEwan DL, Kirienko NV, Ausubel FM (2012) Host translational inhibition by Pseudomonas aeruginosa Exotoxin A Triggers an immune response in Caenorhabditis elegans. Cell Host Microbe 11:364–374. https://doi.org/10.1016/j.chom.2012.02.007
Miller EV, Grandi LN, Giannini JA, Robinson JD, Powell JR (2015) The conserved G-protein coupled receptor FSHR-1 regulates protective host responses to infection and oxidative stress. PLoS One 10:e0137403–e0137416. https://doi.org/10.1371/journal.pone.0137403
Nargund AM, Pellegrino MW, Fiorese CJ, Baker BM, Haynes CM (2012) Mitochondrial import efficiency of ATFS-1 regulates mitochondrial UPR activation. Science 337:587–590. https://doi.org/10.1126/science.1223560
Nicholas HR, Hodgkin J (2004) The ERK MAP kinase cascade mediates tail swelling and a protective response to rectal infection in C. elegans. Curr Biol 14:1256–1261. https://doi.org/10.1016/j.cub.2004.07.022
Nougayrede JP, Homburg S, Taieb F, Boury M, Brzuszkiewicz E, Gottschalk G, Buchrieser C, Hacker J, Dobrindt U, Oswald E (2006) Escherichia coli induces DNA double-strand breaks in eukaryotic cells. Science 313:848–851. https://doi.org/10.1126/science.1127059
Paradis S, Ruvkun G (1998) Caenorhabditis elegans Akt/PKB transduces insulin receptor-like signals from AGE-1 PI3 kinase to the DAF-16 transcription factor. Genes Dev 12:2488–2498. https://doi.org/10.1101/gad.12.16.2488
Pees B, Kloock A, Nakad R, Barbosa C, Dierking K (2017) Enhanced behavioral immune defenses in a C. elegans C-type lectin-like domain gene mutant. Dev Comp Immunol 74:237–242. https://doi.org/10.1016/j.dci.2017.04.021
Pellegrino MW, Nargund AM, Kirienko NV, Gillis R, Fiorese CJ, Haynes CM (2014) Mitochondrial UPR-regulated innate immunity provides resistance to pathogen infection. Nature 516:414–417. https://doi.org/10.1038/nature13818
Powell JR, Kim DH, Ausubel FM (2009) The G protein-coupled receptor FSHR-1 is required for the Caenorhabditis elegans innate immune response. Proc Natl Acad Sci U S A 106:2782–2787. https://doi.org/10.1073/pnas.0813048106
Pujol N, Link EM, Liu LX, Kurz CL, Alloing G, Tan MW, Ray KP, Solari R, Johnson CD, Ewbank JJ (2001) A reverse genetic analysis of components of the toll signaling pathway in Caenorhabditis elegans. Curr Biol 11:809–821
Pukkila-Worley R (2016) Surveillance immunity: an emerging paradigm of innate defense activation in Caenorhabditis elegans. PLoS Pathog 12:e1005795–e1005795. https://doi.org/10.1371/journal.ppat.1005795
Reddy KC, Andersen EC, Kruglyak L, Kim DH (2009) A polymorphism in npr-1 is a behavioral determinant of pathogen susceptibility in C. elegans. Science 323:382–384. https://doi.org/10.1126/science.1166527
Reddy KC, Dunbar TL, Nargund AM, Haynes CM, Troemel ER (2016) The C. elegans CCAAT-enhancer-binding protein gamma is required for surveillance. Immunity 14:1581–1589. https://doi.org/10.1016/j.celrep.2016.01.055
Roberts AF, Gumienny TL, Gleason RJ, Wang H, Padgett RW (2010) Regulation of genes affecting body size and innate immunity by the DBL-1/BMP-like pathway in Caenorhabditis elegans. BMC Dev Biol 10:61. https://doi.org/10.1186/1471-213X-10-61
Schumacher B, Hofmann K, Boulton S, Gartner A (2001) The C. elegans homolog of the p53 tumor suppressor is required for DNA damage-induced apoptosis. Curr Biol 11:1722–1727
Schumacher B, Schertel C, Wittenburg N, Tuck S, Mitani S, Gartner A, Conradt B, Shaham S (2005) C. elegans ced-13 can promote apoptosis and is induced in response to DNA damage. Cell Death Differ 12:153–161
Shivers RP, Pagano DJ, Kooistra T, Richardson CE, Reddy KC, Whitney JK, Kamanzi O, Matsumoto K, Hisamoto N, Kim DH (2010) Phosphorylation of the conserved transcription factor ATF-7 by PMK-1 p38 MAPK regulates innate immunity in Caenorhabditis elegans. PLoS Genet 6:e1000892. https://doi.org/10.1371/journal.pgen.1000892
Shivers RP, Youngman MJ, Kim DH (2008) Transcriptional responses to pathogens in Caenorhabditis elegans. Curr Opin Microbiol 11:251–256. https://doi.org/10.1016/j.mib.2008.05.014
Styer KL, Singh V, Macosko E, Steele SE, Bargmann CI, Aballay A (2008) Innate immunity in Caenorhabditis elegans is regulated by neurons expressing NPR-1/GPCR. Science 322:460–464. https://doi.org/10.1126/science.1163673
Sun J, Singh V, Kajino-Sakamoto R, Aballay A (2011) Neuronal GPCR controls innate immunity by regulating noncanonical unfolded protein response genes. Science 332:729–732. https://doi.org/10.1126/science.1203411
Tazearslan Ç, Ayyadevara S, Bharill P, Reis RJS (2009) Positive feedback between transcriptional and kinase suppression in nematodes with extraordinary longevity and stress resistance. PLoS Genet 5:e1000452. https://doi.org/10.1371/journal.pgen.1000452
Tenor JL, Aballay A (2007) A conserved Toll-like receptor is required for Caenorhabditis elegans innate immunity. EMBO Rep 9:103–109. https://doi.org/10.1038/sj.embor.7401104
Toller IM, Neelsen KJ, Steger M, Hartung ML, Hottiger MO, Stucki M, Kalali B, Gerhard M, Sartori AA, Lopes M, Muller A (2011) Carcinogenic bacterial pathogen Helicobacter pylori triggers DNA double-strand breaks and a DNA damage response in its host cells. Proc Natl Acad Sci U S A 108:14944–14949. https://doi.org/10.1073/pnas.1100959108
Troemel ER, Chu SW, Reinke V, Lee SS, Ausubel FM, Kim DH (2006) p38 MAPK regulates expression of immune response genes and contributes to longevity in C. elegans. PLoS Genet 2:e183. https://doi.org/10.1371/journal.pgen.0020183
Zugasti O, Bose N, Squiban B, Belougne JEROM, Kurz CLEO, Schroeder FC, Pujol N, Ewbank JJ (2014) Activation of a G protein–coupled receptor by its endogenous ligand triggers the innate immune response of Caenorhabditis elegans. Nat Immunol 15:833–838. https://doi.org/10.1038/ni.2957
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Glossary
- AGE-1
-
Ortholog of phosphoinositide 3-kinase (PI3K) p110 catalytic subunit.
- AKT-1/-2
-
Homologs of serine/threonine kinase Akt/PKB ortholog of the serine/threonine kinase Akt/PKB.
- ATF-7
-
Leucine zipper transcription factor; ortholog of CREB/activating transcription factors.
- ATFS-1
-
bZip transcription factor involved in UPRmt.
- CEBP-2
-
Ortholog of human CCAAT7 enhancer binding protein gamma (CEBPG).
- CED-13
-
BH3 domain-containing protein involved in apoptosis.
- CEP-1
-
Ortholog of human tumor suppressor p53.
- DAF-2
-
Receptor tyrosine kinase; insulin/insulin growth factor receptor ortholog.
- DAF-4
-
Serine/threonine kinase; ortholog of type II transforming growth factor (TGF)-β receptor.
- DAF-16
-
Forkhead box O (FOXO) transcription factor in insulin-mediated signaling.
- DAF-28
-
Beta-type insulin; homologous to human insulin.
- DBL-1
-
Member of transforming growth factor (TGF)-β super family.
- DCAR-1
-
Ortholog of human neuropeptide FF receptors (1 and 2) and pyroglutamylated RF amide peptide receptor.
- DCR-1
-
Ribonuclease involved in RNA interference.
- DRH-1
-
Dicer-related helicase involved in RNA interference.
- EGL-1
-
BH3 domain-containing protein involved in apoptosis.
- LIN-45
-
Ortholog of vertebrate RAF protein.
- MEK-2
-
Mitogen-activated protein kinase (MAPK) kinase involved in Ras-mediated signaling.
- MPK-1
-
Mitogen-activated protein kinase (MAPK); ortholog of human extracellular signal-regulated kinase (ERK).
- MUT-7
-
RNaseD homolog involved in RNA interference.
- NPR-1
-
G-protein-coupled neuropeptide receptor; homolog of mammalian neuropeptide Y receptor.
- NSY-1
-
Neuronal symmetry family member 1. Mitogen-activated protein kinase (MAPK) kinase; ortholog of mammalian ASK family of proteins.
- OCTR-1
-
G-protein-coupled receptor involved in neuronal signaling.
- PMK-1
-
Mitogen-activated protein kinase (MAPK); ortholog of human p38 MAPK, orthologous to human mi MAPK (OMIM:600289); MAPK, orthologous to human MAPK (OMIM:600289).
- RDE-1
-
Argonaute and PIWI family protein.
- RDE-4
-
Double-stranded RNA (dsRNA)-binding protein involved in RNA interference.
- SEK-1
-
Ortholog of human mitogen-activated protein kinase kinases (MAPKK) 3 and 6.
- SMA-2/-3/-4
-
Orthologs of SMAD proteins.
- SMA-6
-
Serine/threonine protein kinase; orthologous to type I transforming growth factor (TGF)-β receptors.
- TIR-1
-
Toll/interleukin-1 receptor domain adapter protein; ortholog of human SARM.
- TOL-1
-
Toll-like receptor protein.
- UPR mt
-
Mitochondrial unfolded protein response.
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG, part of Springer Nature
About this chapter
Cite this chapter
Williams, A.B., Schumacher, B. (2018). Nematoda: The Caenorhabditis elegans Model for Innate Immunity – Interactions Between Worms and Pathogens, and Their Responses to Immunogenic Damage. In: Cooper, E. (eds) Advances in Comparative Immunology. Springer, Cham. https://doi.org/10.1007/978-3-319-76768-0_5
Download citation
DOI: https://doi.org/10.1007/978-3-319-76768-0_5
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-76767-3
Online ISBN: 978-3-319-76768-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)