Abstract
Nickel oxide nanoparticles (NiO-NPs) are increasingly used and concerns have been raised on its toxicity. Although a few studies have reported the toxicity of NiO-NPs, a comprehensive understanding of NiO-NPs toxicity in human cells is still lagging. In this study, we integrated transcriptomic approach and genotoxic evidence to depict the mechanism of NiO-NPs toxicity in human hepatocellular carcinoma (HepG2) cells. DNA damage analysis was done using comet assay, which showed 26-fold greater tail moment in HepG2 cells at the highest concentration of 100 μg/ml. Flow cytometric analysis showed concentration dependent enhancement in intracellular reactive oxygen species (ROS). Real-time PCR analysis of apoptotic (p53, bax, bcl2) and oxidative stress (SOD1) genes showed transcriptional upregulation. Transcriptome analysis using qPCR array showed over expression of mRNA transcripts related to six different cellular pathways. Our data unequivocally suggests that NiO-NPs induces oxidative stress, DNA damage, apoptosis and transcriptome alterations in HepG2 cells.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
10.1 Introduction
In recent years’ nanotechnology has exhibited exponential growth in various sectors to accomplish market commodities with higher prospective applications [1]. At least thousand consumer products are available which contains nanoparticles (NPs), ranging from everyday household items to medical diagnostic tools, imaging, drug delivery and aerospace engineering [2, 3]. Compared to the bulk counterpart, the small size and large specific surface area of NPs endow them with high chemical reactivity and intrinsic toxicity. Such unique physiochemical properties of NPs draw global attention of scientists and environmental watchdogs to keep concern about NPs potential risks and adverse effect on human health [4]. NPs find route to human body via skin penetration, ingestion, inhalation or injection and interact with cellular organelles for longer time period [5]. Consequently, NPs have been found to effortlessly interact with cells and organs by various mechanisms [6]. Since methodologies for exposure assessment are non-consistent, the toxicological research on NPs is still lagging. Therefore, in order to plug the gap between development and toxicity of NPs, a major effort is needed to study the effects of exposure to NPs.
In this study, we have selected nickel oxide nanoparticles (NiO-NPs) owing to its increasing application in ceramic material, catalysts, electronic component and biosensors [7,8,9]. Despite its wide use, NiO-NPs has raised concerns about its adverse effects on the environment and human health. NiO-NPs generated from welding fumes during the coastal region developments were considered as a potential nano-pollution source in coastal seawaters (IARC Monographs on the Evaluation of Carcinogenic Risks to Humans). Direct aerial emission of NiO-NPs has the tendency to pollute surface waters through leakages, spills and indirect storm-water runoff from land [10]. The metallic Ni-NPs has been recently used to catalyze the reversible hydration of CO2 to carbonic acid, which is holding extreme importance in CO2 capture technologies and mineralization processes. These advantages led to its utilization to point flue sources like air-conditions outlets on top of building or power plants [11, 12]. Being the 24th most abundant element in the Earth crust, nickel compounds (NiSO4, NiO, nickel hydroxides and crystalline nickel) are well known as an environmental pollutant and classified as carcinogenic agents to humans (Group 1) by the International Agency for Research on Cancer (IARC) [13]. The in vivo studies on NiO-NPs have been mostly focused on pulmonary pathology. Female Wistar rats intratracheally instilled with NiO-NPs exhibited a significant increase in the bronchiolar alveolar lavage fluid (BLAF), activation of IL-1β, IFN-Ƴ, MIP-2 and histological changes [14]. A short-term exposure of rats to 500 cm2/ml of NiO-NPs induced polymorphonuclear neutrophils in the BALF [15]. Inhalation exposure of rats with NiO-NPs in nebulizer chamber exhibited biopersistance of NPs in lungs and inflammatory responses [16]. Long term intratracheally instillation of NiO-NPs in rats exhibited increased vacuolization in alveolar macrophages and CINC-1 concentrations [17]. Wistar rats instilled with NiO-NPs after 4 days of exposure showed eosinophilic and neutrophilic inflammation, along with release of eotaxin and cellular disintegration by the release of Ni ions [18]. Female Wistar rats exposed by pharyngeal route to NiO-NPs showed enhanced proinflammatory cytokines, LDH, lymphocytes, polymorphonuclear leukocytes in BALF [19]. DNA damage and low expression of HO-1 and Nrf2 proteins were observed in Male Sprague Dawley rats when intratracheally instilled with Ni-NPs for two weeks. In addition, the animals showed alterations in the normal morphology of lungs, liver and kidneys [20]. Ultrafine-size particles and NiO-NPs of nickel compounds have greater bioavailability and toxicity as compared to its fine-size nickel compounds [21]. We have recently reported that NiO-NPs induces liver toxicity, cytogenetic anomalies and apoptosis via p53, MAPK, caspase 8 and 3 signalling in rats [22]. Another recent study in the same line expressed NiO-NPs genotoxicity, chromosomal aberrations, DNA damage in lymphocytes, liver and kidney of female rats [23]. Zebrafish exposed to NiO-NPs for longer time showed higher bioaccumulation and toxicity [24]. It is well documented and established that solubilization of Ni2+ from NiO-NPs plays vital role in inducing toxicity in animal, invertebrate, cell line and plant [18, 25,26,27].
Concerning the in vitro studies, recent reports suggest NiO-NPs as neurotoxic in SH-SY5Y neuroblastoma cells and cytotoxic for human breast carcinoma cells (MCF7) [28, 29]. NiO-NPs induced HIF-1α transcription factor followed by upregulation of its target NRDG1 (Cap43) in human lung epithelial (H460) cells [21]. In the same line, NiO-NPs induced oxidative stress and cytotoxicity in human alveolar epithelial cells (A549) has also been reported [30]. HepG2 cells exposed to NiO-NPs resulted in cytotoxicity and apoptosis responses via reactive oxygen species (ROS) generation, which is likely to be mediated through bax/bcl-2 pathway [31].
Despite these facts, a systematic interpretation on the underlying mechanism of NiO-NPs induced hepatotoxicity is scarce. In this context, NiO-NPs has been reported to induce cytotoxicity and apoptotic cell death in HepG2 cells via bax/blc2 pathway [31]. However, authors did not explain the vital queries on HepG2 transcriptomic profile. To decipher these unattended queries, we have provided a concrete evidence on hepatotoxicity under in vitro condition. Primary human hepatocytes have been considered as gold standard model for xenobiotic metabolism and cytotoxicity studies [32]. However, the complexity in isolation procedures, short life-span, inter-individual variability, cost effectiveness and rare availability of fresh human liver samples, constitute serious limitations for the use of aforesaid in vitro systems in screening [33]. Such constrains were run-over by immortalized liver-derived cell lines, owing to their unlimited availability and phenotypic stability. A first alternative is the widely used HepG2 cells, as these cells are highly differentiated and display many of the genotypic features of normal liver cells [34]. HepG2 can be used to screen the cytotoxic potential of new chemical entities at the lead generation phase and imitate the normal metabolic pathway in vivo [35, 36]. In this study, we have selected HepG2 cells as a model system for studying the hepatotoxic effects of NiO-NPs.
Consequently, the current study was aimed to evaluate molecular mechanism of NiO-NPs in vitro toxicity in HepG2 cells by the measurement of (i) intracellular ROS generation (ii) DNA damage (iii) transcriptional activation of array of genes related to human stress and toxicity pathways.
10.2 Materials and Methods
10.2.1 NiO-NPs Characterization
NiO-NPs (Cat. No. 637130) was purchased from sigma chemical company (St. Louis, MO, USA). A stock of NiO-NPs (1 mg/ml) was prepared in MQ water and sonicated for 20 min at 40 W. TEM analysis of NiO-NPs were done by dropping the stock solution on copper grids and subjected to microscopic analysis at 200 KeV (JEM-2100 F, JEOL, Japan).
10.2.2 Cell Culture and NiO-NPs Exposure
Human liver hepatocellular carcinoma (HepG2) cells were cultured in DMEM with 10% FBS, antibiotic-antimycotic solution and incubated at 37 °C with 5% CO2. HepG2 cells were seeded in 96 and 6-well plates and allowed to attach with the surface for 24 h prior to NiO-NPs treatment. Before each experiment, the ultra sonicated NiO-NPs (25, 50 and 100 μg/ml) solutions added to cell culture and grown for 24 h. Control groups were not added with NiO-NPs.
10.2.3 In Vitro DNA Damage Analysis by Comet Assay
The HepG2 cells exposed for 3 h with 25, 50 and 100 μg/ml of NiO-NPs were detached and centrifuged at 3000 rpm for 3 min to collect the pellets. The cells (4 × 104) from untreated and treated groups were suspended in 100 μl of Ca++ Mg++ free PBS and mixed with 100 μl of 1% low melting point agarose (LMA). The cell suspension (80 μl) was then layered on one-third frosted slides, pre-coated with normal melting agarose (NMA) (1% in PBS) and kept at 4 °C for 10 min. After gelling, a layer of 90 μl of LMA (0.5% in PBS) was added. After the solidification of agarose on slides, all of them were kept in lysis solution for overnight, followed by unwinding and electrophoresis at 24 V (300 mA) for 20 min. Cells were stained with ethidium bromide (20 μg/ml) and DNA damage were scored under fluorescence microscope.
10.2.4 ROS Measurements in HepG2 Cells
After the specified treatment, cells were trypsinized, pelleted and washed twice with cold PBS, followed by the resupension of cells in 500 μl PBS (Ca++ and Mg++ free) containing 5 μM of dichloro-dihydro-fluorescein diacetate (DCFH-DA) dye. All cells were incubated for 60 min at 37 °C in dark followed by washing and the fluorescence were recorded upon excitation at 488 nm at FL1 Log channel through 525 nm band-pass filter on Beckman Coulter flow cytometer (Coulter Epics XL/Xl-MCL, USA). Qualitative analysis of ROS in NiO-NPs treated cells were also done by staining the HepG2 cells with 5 μM of DCFH-DA for 60 min at 37 °C in CO2 incubator. Fluorescence images were captured on microscope equipped with fluorescent lamp (Nikon Eclipse 80i, Japan).
10.2.5 RT2 Profiler PCR Array Analysis
PCR array experiments were done with HepG2 cells exposed for 24 h with NiO-NPs (100 μg/ml). In brief, total RNA was isolated using the commercially available kit (RNeasy Mini Kit, Cat. No. 74106, Qiagen, USA), purification was done using iPrep™ PureLink™ kit (Invitrogen, USA) by Invitrogen® automated system. Purity of total RNA was verified by use of a Nanodrop 8000 spectrophotometer (Thermo Scientific, USA). The first-strand cDNA synthesis was performed with 1 μg of total RNA and 100 ng of oligo-p(dT)12-18 primer and MLV reverse transcriptase (GE Health Care, UK). Changes in the relative gene expression of 84 genes responsible for human stress and toxicity pathway were quantified using 96-well format of RT2 Profiler™ PCR Array (Cat. No. PAHS-003 A, SABiosciences Corporation, Frederick, MD). cDNA equivalent to 1 μg of total RNA was used for each array. The arrays were run on Roche® LightCycler® 480 (Roche Diagnostics, Rotkreuz, Switzerland) following the recommended cycling programs. Online software from SABiosciences Corporation, Frederick, MD, was used to analyze the expression data. NiO-NPs expression results were normalized to the average Ct value of five housekeeping genes (B2M, HPRT1, RPL13A, GAPDH and ACTB) and expressed with respect to the untreated control. RT-PCR array data were evaluated from at least three independent experiments and the resultant ΔCt values were combined to calculate the average fold regulation values. Genes that were significantly different for NiO-NPs versus control were determined by Students t-test (p < 0.05) by comparing the ΔCt values for the triplicate trials for each test sample with the ΔCt values for the control. Then PCR array data were validated by measuring the mRNA expression of some selected genes (P53, BAX, BCL2, SOD1) using real-time PCR analysis (Table 10.1).
10.3 Results
10.3.1 NiO-NPs Characterization
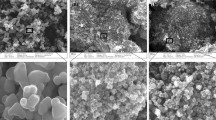
The size and morphology of NiO-NPs were measured by transmission electron microscopy (TEM). In TEM analysis, NiO-NPs appeared as an aggregate showing crystallite’s spheres. The particles size of NiO-NPs analyzed from six TEM images were determined to be 24.05 ± 2.9 nm (Fig. 10.1).
10.3.2 DNA Damage in HepG2 Cells
HepG2 cells exposed to NiO-NPs for 3 h resulted in DNA damage. The representative comet image from NiO-NPs (100 μg/ml) treatment clearly demonstrates the broken DNA liberated from the comet head (Fig. 10.2). NiO-NPs at 25, 50 and 100 μg/ml induced significant 25.1, 25.3 and 26.7-fold higher Olive tail moment (OTM) parameter of comet assay vis-à-vis the control showed a background of 0.28 ± 0.05 OTM (Table 10.2). The advantage of comet assay is that it is capable of analysing population of cells with various degrees of DNA damage. Nevertheless, the differences in distribution of DNA damage exist in the cell population. Variation in distribution of DNA damage by NiO-NPs exposure in-terms of frequency is shown in Fig. 10.2.
10.3.3 Quantitative and Qualitative Analysis of Intracellular ROS
A concentration dependent increase in the intracellular ROS generation in HepG2 cells, as evident by the shift of DCF peaks in treated groups (Fig. 10.3a). Compared to the 100% DCF fluorescence in control, cells treated with 25, 50 and 100 μg/ml of NiO-NPs showed significant 134%, 150% and 143% (p < 0.01) increase in ROS generation (Fig. 10.3a, Inset). Fluorescence images further validated the flow cytometric data by showing an enhanced level of DCF fluorescence in the NiO-NPs treated cells (Fig. 10.3b).
(a) Fluorescence enhancement of DCF indicating ROS production with increasing NiO-NPs concentrations in HepG2 cells analyzed by flow cytometry. Each histogram in inset represents the values of mean±SD of three independent experiments done in triplicate wells (**p < 0.01 vs. control). (b) Fluorescence microscopic images of treated cells showing an enhancement in green fluorescence of DCF in treated cells
10.3.4 qPCR Array of HepG2 Cells
HepG2 cells treated with NiO-NPs (100 μg/ml) for 24 h exhibited differential expression of genes in the RT2 profiler PCR array. The corresponding heat map suggested strong oxidative or metabolic stress, growth arrest and senescence, apoptosis signalling, proliferation and carcinogenesis, and activation of proinflammatory responses upon NiO-NPs exposure (Fig. 10.4). CYP2E1 gene in oxidative or metabolic stress group has exhibited a maximum of 3.4-fold up-regulation. Considerable number of genes in this pathway was up-regulated, and 1.4, 1.5 and 1.1-fold of maximum up-regulation has been recorded for HMOX1, SOD2 and SOD1 genes. Among the set of seven genes responsible for growth arrest and senescence, GDF15, DDIT3, GADD45A, MDM2 and P53 genes have exhibited 4.6, 2.4, 1.6, 1.2 and 1.2-fold up-regulation. TNFSF10, TNFRSF1A, CASP8 and NFKB1A genes in apoptosis signalling group showed maximum up-regulation of 1.8, 1.1, 1.4 and 1.0-fold, while the BCL2L1 expression was down-regulated to 1.1-fold. Within the proliferation and carcinogenesis pathway EGR1 showed 1.2-fold up-regulation. Among the proinflammatory genes, NOS2 was maximally up-regulated to 2.3-fold. HSPA6 gene in heat shock group, showed up-regulation of 2.5-fold. qPCR array data validation was done by measuring the expression of selective genes (P53, BAX, BCL2 and SOD1) by real-time PCR. The expressional analysis also showed 1.0, 1.2, and 1.1-fold up-regulation of P53, BAX and SOD1. BCL2 was found under expressed to 1.2-fold (Fig. 10.5).
10.4 Discussion
An integrated approach was used to identify toxicity mechanism induced by NiO-NPs. In this study, low, medium and high (25, 50 and 100 μg/ml) doses of NiO-NPs has been chosen to expose the human liver cells. Lowest concentration was chosen with the aim to imitate the potential human exposure, on the other hand highest concentration was selected to reflect toxicological effects upon accidental exposure of NiO-NPs. In this line an enhanced level of ROS has been observed in NiO-NPs treated cells. These results corroborate with enhanced ROS level in NiO-NPs treated HepG2 and A549 cells [31, 37]. We suggest oxidative stress in HepG2 cells. The current study demonstrates that NiO-NPs can induce DNA damage after short exposure of 3 h, and corresponds with previous reports on DNA damage in HepG2 and WISH cells exposed to NiO, TiO2 and ZnFe2O3-NPs [31, 38, 39]. The appearance of comet tail with NiO-NPs exposure unequivocally suggest the impairment of DNA repair machinery. The enhancement in intracellular ROS and DNA damage data are in agreement with our recent report on NiO-NPs induced liver toxicity in rats [22]. Ni2+ is involved in ROS generation and accounted for inducing high level of damage via direct oxidative damage by H2O2 production [40]. Hence, the elevated toxicity and damage in our study could also be an additive oxidative action of Ni2+ released from NiO-NPs.
PCR array revealed that NiO-NPs treatment resulted in the up-regulation of genes related to different pathways. We found that TNFSF10, TNFRSF1A, CASP8 and NFKBIA genes in apoptosis signalling pathways were up-regulated. TNFSF10 and TNFRSF1A belong to the tumor necrosis factor receptor (TNFR) family and their up-regulation has been suggested to induce cell death [41]. Up-regulation of CASP8 expression has been linked to execute the apoptotic signaling mainly through extrinsic pathway [42]. Activation of the above genes strongly suggests the participation of death receptor-mediated TNFR family members to induce apoptosis via intrinsic as well as extrinsic pathways. TNFR genes can act through an autocrine pathway to induce cell growth arrest and apoptosis through NFKB activation [43]. Therefore, the NFKB pathway and related genes could also be an important molecular mechanism by which NiO-NPs induces apoptosis in HepG2 cells.
NiO-NPs treatment resulted in the up-regulation of EGR1, MDM2, GADD45A and DDIT3 genes. Although the induction of EGR1, a family of zinc finger transcription factors, is directly linked with oxidative stress per se, other condition like mitochondrial dysfunction contributes well in its up-regulation [44]. We have found in vivo mitochondrial dysfunction in rats exposed to NiO-NPs for 7 and 14 days [22]. Therefore, the observed mitochondrial dysfunction and oxidative stress data strongly substantiate the likelihood that NiO-NPs may function as an initiator to increase the expression of EGR1 in treated HepG2 cells. Up-regulation of DDIT3 and GADD45A transcripts can be correlated with the fact that under stressed condition EGR1 is known to initiate DDIT3 and GADD45 family genes by binding to 5′-flanking regions [45]. The oxidative stress related genes (SOD1, SOD2, GPX1 and HMOX1) were found up-regulated after NiO-NPs exposure. In view of the higher ROS generation by NiO-NPs, we suggest that cytoplasmic (SOD1), mitochondrial (SOD2), glutathione system (GPX1) and HMOX1 might have involved in scavenging the free radicals and cytoprotection. However, the excessive oxidative stress was beyond the attenuation capacity of these enzymes to subtle the DNA damage in treated cells. Up-regulation of above genes corresponds with increased expression of SOD, GPX, and HMOX1 in human cells, when treated with ZnO-NPs and polyphenolic compounds [46, 47]. NOS2 is a hallmark of inflammatory response and its up-regulation is governed by oxidative stress, metals and lipopolysaccharides [48]. NOS2 expression is in accordance with our previous work on ZnFe2O4-NPs, exhibiting its induction in WISH cells [38]. GDF15 overexpression corresponds well with p53-GDF15 link, and points towards its important role during inflammatory responses after NiO-NPs treatment [49]. Within the set of heat shock genes, HSPA6 was found highly up-regulated in NiO-NPs treated cells. Heat shock proteins (HSP) are highly conserved class of stress response proteins, which work as molecular chaperons to correct the protein conformation under stress condition to maintain cellular homeostasis and protect the cells from apoptotic cell death [50]. Nonetheless, the DNA damage in NiO-NPs treated HepG2 cells supports the view that HSP fails to intervene the apoptotic process, as depicted in the image (Fig. 10.6).
10.5 Conclusion
We conclude that NiO-NPs have the potential to alter the transcriptome of HepG2 cells. We observed that NiO-NPs generated ROS and these free radicals induce heavy oxidative stress, which has affected the cell survival and promoted DNA. Transcriptional analysis of PCR array revealed overall up-regulation of different pathway genes, suggesting a pleiotropic effect of NiO-NPs to induced HepG2 cell death. The analysis of transcriptome was helpful to reveal potential molecular mechanism underlying NiO-NPs induced effects on HepG2 cells. The observed toxicity in HepG2, corresponds well with our recent study on rat’s showing hepatotoxicity. Hence, NiO-NPs widespread application should be given meticulous attention for potential adverse biological effects.
References
Singh SP, Chinde S, Kamal SSK et al (2016) Genotoxic effects of chromium oxide nanoparticles and microparticles in Wistar rats after 28 days of repeated oral exposure. Environ Sci Pollut Res 23:3914–3924(2)
Tomankova K, Horakova J, Harvanova M et al (2015) Cytotoxicity, cell uptake and microscopic analysis of titanium dioxide and silver nanoparticles in vitro. Food Chem Toxicol 82:106–115
Kim T, Hyeon T (2014) Applications of inorganic nanoparticles as therapeutic agents. Nanotechnology 25:012001
Beaudrie CE, Kandlikar M, Satterfield T (2013) From cradle-to-grave at the nanoscale: gaps in U.S. regulatory oversight along the nanomaterial life cycle. Environ Sci Technol 47(11):5524–5534
Nel A, Xia T, Mädler L et al (2006) Toxic potential of materials at the nano level. Science 311:622–627
Nel AE, Mädler L, Velegol D et al (2009) Understanding biophysicochemical interactions at the nano-bio interface. Nat Mater 8:543–557
Salimi A, Sharifi E, Noorbakhsh A et al (2007) Direct electrochemistry and electrocatalytic activity of catalase immobilized onto electrodeposited nano-scale islands of nickel oxide. Biophys Chem 125(2–3):540–548
Mu Y, Jia D, He Y (2011) Nano nickel oxide modified non-enzymatic glucose sensors with enhanced sensitivity through an electrochemical process strategy at high potential. Biosens Bioelectron 26(6):2948–2952
Rao KV, Sunandana CS (2008) Effect of fuel to oxidizer ratio on the structure, micro structure and EPR of combustion synthesized NiO nanoparticles. J Nanosci Nanotechnol 8(8):4247–4253
Wiesner MR, Lowry GV, Alvarez P et al (2006) Assessing the risks of manufactured nanomaterials. Environ Sci Technol 40(14):4336–4345
Bhaduri GA, Siller L (2012) Carbon capture. United Kingdom Patent Application GB1208511.4
Bhaduri GA, Siller L (2013) Nickel nanoparticles catalyse reversible hydration of carbon dioxide for mineralization carbon capture and storage. Cat Sci Technol 3:1234e1239
Cameron KS, Buchner V, Tchounwou PB (2011) Exploring the molecular mechanisms of nickel-induced genotoxicity and carcinogenicity: a literature review. Rev Environ Health 26(2):81–92
Cho WS, Duffin R, Poland CA et al (2010) Metal oxide nanoparticles induce unique inflammatory footprints in the lung: important implications for nanoparticle testing. Environ Health Perspect 118(12):1699–1706
Lu S, Duffin R, Poland C et al (2009) Efficacy of simple short-term in vitro assays for predicting the potential of metal oxide nanoparticles to cause pulmonary inflammation. Environ Health Perspect 117(2):241–247
Oyabu T, Ogami A, Morimoto Y et al (2007) Biopersistence of inhaled nickel oxide nanoparticles in rat lung. Inhal Toxicol 19(1):55–58
Nishi K, Morimoto Y, Ogami A et al (2009) Expression of cytokine-induced neutrophil chemoattractant in rat lungs by intratracheal instillation of nickel oxide nanoparticles. Inhal Toxicol 21(12):1030–1039
Lee S, Hwang SH, Jeong J et al (2016) Nickel oxide nanoparticles can recruit eosinophils in the lungs of rats by the direct release of intracellular eotaxin. Part Fibre Toxicol 13(1):30
Jeong J, Kim J, Seok SH et al (2016) Indium oxide (In2O3) nanoparticles induce progressive lung injury distinct from lung injuries by copper oxide (CuO) and nickel oxide (NiO) nanoparticles. Arch Toxicol 90(4):817–828
Magaye R, Gu Y, Wang Y et al (2016) In vitro and in vivo evaluation of the toxicities induced by metallic nickel nano and fine particles. J Mol Histol 47(3):273–286
Pietruska JR, Liu X, Smith A et al (2011) Bioavailability, intracellular mobilization of nickel, and HIF-1alpha activation in human lung epithelial cells exposed to metallic nickel and nickel oxide nanoparticles. Toxicol Sci 124(1):138–148
Saquib Q et al (2017) p53, MAPKAPK-2 and caspases regulate nickel oxide nanoparticles induce cell death and cytogenetic anomalies in rats. Int J Biol Macromol.: pii: S0141-8130(17)31498-8. https://doi.org/10.1016/j.ijbiomac.2017.07.032
Dumala N et al (2017) Genotoxicity study of nickel oxide nanoparticles in female Wistar rats after acute oral exposure. Mutagenesis. https://doi.org/10.1093/mutage/gex007
Kovriznych JA, Sotnikova R, Zeljenkova D et al (2014) Long-term (30 days) toxicity of NiO nanoparticles for adult zebrafish Danio rerio. Interdiscip Toxicol 7(1):23–26
Kanold JM, Wang J, Brümmer F et al (2016) Metallic nickel nanoparticles and their effect on the embryonic development of the sea urchin Paracentrotus lividus. Environ Pollut 212:224–229
Duan WX, He MD, Mao L et al (2015) NiO nanoparticles induce apoptosis through repressing SIRT1 in human bronchial epithelial cells. Toxicol Appl Pharmacol 286(2):80–91
Faisal M, Saquib Q, Alatar AA et al (2013) Phytotoxic hazards of NiO-nanoparticles in tomato: a study on mechanism of cell death. J Hazard Mater 250–251:318–332
Abudayyak M, Guzel E, Özhan G (2017) Nickel oxide nanoparticles are highly toxic to SH-SY5Y neuronal cells. Neurochem Int 108:7–14. https://doi.org/10.1016/j.neuint.2017.01.017
Umaralikhana L, Jaffar MJM (2016) Antibacterial and anticancer properties of NiO nanoparticles by co-precipitation method. J Adv App Sci Res 1(4):24–35
Lu S et al (2015) Comparison of cellular toxicity caused by ambient ultrafine particles and engineered metal oxide nanoparticles. Part Fibre Toxicol 12:5
Ahamed M, Ali D, Alhadlaq HA et al (2013) Nickel oxide nanoparticles exert cytotoxicity via oxidative stress and induce apoptotic response in human liver cells (HepG2). Chemosphere 93(10):2514–2522
Guillouzo A et al (2007) The human hepatoma HepaRG cells: a highly differentiated model for studies of liver metabolism and toxicity of xenobiotics. Chem Biol Interact 168:66–73
Madan A et al (2003) Effects of prototypical microsomal enzyme inducers on cytochrome P450 expression in cultured human hepatocytes. Drug Metab Dispos 31:421–431
Sassa S, Sugita O, Galbraith RA et al (1987) Drug metabolism by the human hepatoma cell, HepG2. Biochem Biophys Res Commun 143:52–57
Gerets HHJ et al (2009) Selection of cytotoxicity markers for the screening of new chemical entities in a pharmaceutical context: a preliminary study using a multiplexing approach. Toxicol In Vitro 23:319–332
Liu X et al (2014) The role of lysosomes in BDE 47-mediated activation of mitochondrial apoptotic pathway in HepG2 cells. Chemosphere 124:10–21
Capasso L, Camatini M, Gualtieri M (2014) Nickel oxide nanoparticles induce inflammation and genotoxic effect in lung epithelial cells. Toxicol Lett 226(1):28–34
Saquib Q, Al-Khedhairy AA, Ahmad J et al (2013) Zinc ferrite nanoparticles activate IL-1b, NFKB1, CCL21 and NOS2 signaling to induce mitochondrial dependent intrinsic apoptotic pathway in WISH cells. Toxicol Appl Pharmacol 273(2):289–297
Saquib Q, Al-Khedhairy AA, Siddiqui MA et al (2012) Titanium dioxide nanoparticles induced cytotoxicity, oxidative stress and DNA damage in human amnion epithelial (WISH) cells. Toxicol In Vitro 26(2):351–361
Kawanishi S, Oikawa S, Inoue S et al (2002) Distinct mechanisms of oxidative DNA damage induced by carcinogenic nickel subsulfide and nickel oxides. Environ Health Perspect 110:789–791
Liu ZG (2005) Molecular mechanism of TNF signaling and beyond. Cell Res 15(1):24–27
Luzio A, Monteiro SM, Fontainhas-Fernandes AA et al (2013) Copper induced upregulation of apoptosis related genes in zebrafish (Danio rerio) gill. Aquat Toxicol 128–129:183–189
Kang JX, Liu J, Wang J et al (2005) The extract of huanglian, a medicinal herb, induces cell growth arrest and apoptosis by upregulation of interferon-beta and TNF-alpha in human breast cancer cells. Carcinogenesis 26(11):1934–1939
Han MH, Kim GY, Yoo YH et al (2013) Sanguinarine induces apoptosis in human colorectal cancer HCT-116 cells through ROS-mediated Egr-1 activation and mitochondrial dysfunction. Toxicol Lett 220(2):157–166
Frank MB, Yang Q, Osban J et al (2009) Frankincense oil derived from Boswellia carteri induces tumor cell specific cytotoxicity. BMC Complement Altern Med 9:6
Setyawati MI, Tay CY, Leong DT (2013) Effect of zinc oxide nanomaterials-induced oxidative stress on the p53 pathway. Biomaterials 34(38):10133–10142
Wang X, Stavchansky S, Kerwin SM et al (2010) Structure-activity relationships in the cytoprotective effect of caffeic acid phenethyl ester (CAPE) and fluorinated derivatives: effects on heme oxygenase-1 induction and antioxidant activities. Eur J Pharmacol 635(1–3):16–22
Lee JK, Sayers BC, Chun KS et al (2012) Multi-walled carbon nanotubes induce COX-2 and iNOS expression via MAP kinase-dependent and-independent mechanisms in mouse RAW264.7 macrophages. Part Fibre Toxicol 9:14
Yang H, Filipovic Z, Brown D et al (2003) Macrophage inhibitory cytokine-1: a novel biomarker for p53 pathway activation. Mol Cancer Ther 2(10):1023–1029
Wang X, Bynum JA, Stavchansky S et al (2014) Cytoprotection of human endothelial cells against oxidative stress by 1-(2-cyano-3,12-dioxooleana-1,9(11)-dien-28-oyl)imidazole (CDDO-Im): application of systems biology to understand the mechanism of action. Eur J Pharmacol 734:122–131
Acknowledgements
The authors are grateful to the Deanship of Scientific Research, King Saud University for funding through Vice Deanship of Scientific Research Chairs.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 The Author(s)
About this chapter
Cite this chapter
Saquib, Q. et al. (2018). Nickel Oxide Nanoparticles Induced Transcriptomic Alterations in HEPG2 Cells. In: Saquib, Q., Faisal, M., Al-Khedhairy, A., Alatar, A. (eds) Cellular and Molecular Toxicology of Nanoparticles. Advances in Experimental Medicine and Biology, vol 1048. Springer, Cham. https://doi.org/10.1007/978-3-319-72041-8_10
Download citation
DOI: https://doi.org/10.1007/978-3-319-72041-8_10
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-72040-1
Online ISBN: 978-3-319-72041-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)