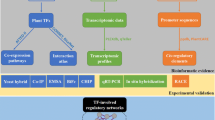
Abstract
Plants are endowed by a resilient developmental program enabling them growth, reproductive success, and response to many environmental challengers which are primarily modulated at transcriptional level via a myriad interactions of transcription-associated proteins (TAPs). TAPs are categorized into three groups based on the roles in transcriptional regulation such as transcription factors (TFs), transcriptional regulators (TRs), and putative TAPs. TFs modulate diverse panels of biological processes via binding cis-regulatory elements in promoter regions of target genes, and plant genomes are estimated to ascribe ~7 % of their coding genes to TFs. Today, the use of next-generation sequencing (NGS) technologies complemented with various bioinformatics resources allowed to systematically identify and classify TFs at genome-scale. Accordingly, many specialized plant TF databases have been also developed including AGRIS, PlnTFDB, PlantTFDB, GRASSIUS, PlantTFcat, TreeTFDB, SoyTFKB, RIKEN, SoyDB, RiceSRTFDB, and IT3F.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction to Transcription Factors
Plants are sessile but astonishing organisms. They are endowed by a resilient developmental program enabling them growth, reproductive success, and response to many environmental challengers (Udvardi et al. 2007). All these events are primarily modulated at gene transcriptional level via complex interactions of various transcription-associated proteins (TAPs ). Plant TAP proteins could be broadly categorized into three groups based on their roles in transcriptional regulation such as transcription factors (TFs), transcriptional regulators (TRs), and putative TAPs. TFs directly activate or suppress the transcription of target genes via binding their cis-regulatory elements in upstream or promoter regions (Franco-Zorrilla and Solano 2016). TRs include the general transcription initiation factors, coactivators or corepressors, and chromatin remodeling factors (Zheng et al. 2016). Putative TAPs contain the other unknown function proteins possibly related with transcriptional regulation (Richardt et al. 2007). Transcription factors modulate very diverse panels of biological processes in plants from embryogenesis, inflorescence development, maintenance of homeostasis, and plant morphology to response of internal or external stimuli (Yamasaki et al. 2013). Plant genomes are estimated to ascribe about 7% of their coding sequences to TFs, indicating the intricacy of transcriptional regulation (Udvardi et al. 2007). So, over the past decades, huge efforts have been realized to elucidate the roles of TFs in biological processes. TFs could possess one or more DNA-binding domains (DBDs ) which specifically bind to cis-regulatory elements in upstream regions/promoters of target genes to mediate the binding of RNA polymerase before transcription initiation (Lin et al. 2014). For instance, an ethylene response factor 1 (ERF1) from Arabidopsis possesses a single AP2 domain directing to bind to GCC box in promoter regions of ethylene-responsive genes, whereas AINTEGUMENTA (ANT) from Arabidopsis has two AP2 domains (Ohme-Takagi and Shinshi 1995; Nole-Wilson and Krizek 2000). Besides, TFs may also have an auxiliary domain to facilitate DNA binding, for example, auxin response factor (ARF) harbors a “B3” DNA-binding domain as well as contains an auxiliary domain “auxin/indole-3-acetic acid (Aux/IAA). ” ARF binds to TGTCTC-containing auxin responsive elements via interaction of auxiliary Aux/IAA domain (Ulmasov et al. 1997; Guilfoyle et al. 1998). TFs are mainly classified into TF families based on their protein domains. In Arabidopsis, 2,296 TFs were classified into 58 TF families (Jin et al. 2013). In rice about 2,478 putative TF genes were classified in 84 families covering nearly 6% of protein-coding genes (Priya and Jain 2013). In maize, 2,538 putative TF genes were distributed in 64 families representing the ~6% of protein-coding genes (Lin et al. 2014). However, interestingly soybean possessed 5,671 putative TF genes clustered in 63 families, including about 12 % of protein-coding genes (Schmutz et al. 2010). The high number of TFs in soybean was partly attributed to the whole-genome duplication (WGD ) events occurred about 13 million years ago (MYO ) (Schmutz et al. 2010). Based on estimations TFs comprise a notable portion (~7%) of protein-coding genes in a plant genome (Udvardi et al. 2007). Thus, their genome-wide identification is usually based on the availability of sequenced genomes. Herein, to have a comparative insights about TF families and their distribution, eight selected plant species with sequenced genomes from two dicots (A. thaliana and Glycine max), two monocots (Oryza sativa indica and Zea mays), two tree species (Populus trichocarpa and Eucalyptus grandis), and two lower plants (Chlamydomonas reinhardtii and Physcomitrella patens) have been investigated (Table 1). TF families have been searched in a comprehensive specialized plant TF database PlantTFDB which contains 320,370 TFs from 165 plant species classified into 58 families (Jin et al. 2013; also refer to Table 3). Database employs a family assignment rule according to which there are three types of domain a TF belongs to such as DNA-binding domain, auxiliary domain, and forbidden domain. Generally, a DNA-binding domain correctly assigns itself to a family. However, in some cases an auxiliary domain is needed to classify TF correspondingly. In addition, forbidden domains are employed to eliminate the sequences with DNA-binding domain but no transcription activity. Based on these rules, 2,296 identified TFs (1,717 loci) in Arabidopsis were classified into 58 families, 6,150 TFs (3,747 loci) in soybean classified into 57 families, 1,891 TFs (1891 loci) in rice classified into 56 families, 3308 TFs (2,289 loci) in maize classified into 56 families, 4,287 TFs (2,466 loci) in poplar classified into 58 families, 2,163 TFs (1,731 loci) in Eucalyptus classified into 56 families, 230 TFs (206 loci) in Chlamydomonas classified into 29 families, and 3,930 TFs (1,156 loci) in Physcomitrella classified into 53 families. In light of identified TF and loci in eight plants, it could be speculated that most TF genes could have been occurred through small and/or large-scale genomic duplications, particularly soybean. Besides, a lower plant green algae C. reinhardtii also demonstrated a clear divergence from other plants in terms of TF families. Among TFs, basic helix-loop-helix (bHLH) family was the most widely distributed TF, also implicating its functional diversity. All taken together, specialized TF databases appear to provide very useful information from functional and evolutionary aspects.
In earlier times of TF studies, identification of TFs and their potential binding elements has been mainly studied through the conventional laboratory experiments (Wingender 1988). However today, the availability of sequenced genomes of many plant species has allowed us to systematically identify and classify the TFs at genome-scale (Riechmann et al. 2000; Schmutz et al. 2010; Priya and Jain 2013; Lin et al. 2014). The integration of next-generation sequencing (NGS) technologies along with various bioinformatics resources has also revolutionized the field of TF research. In addition, the introduction of various bioinformatics portals also paved the way for storage, analysis, and distribution of TFs efficiently. A number of specialized TF databases have been developed for plants including AGRIS (Yilmaz et al. 2011), PlnTFDB (Pérez-Rodríguez et al. 2009), PlantTFDB (Jin et al. 2013), GRASSIUS (Yilmaz et al. 2009), PlantTFcat (Dai et al. 2013), TreeTFDB (Mochida et al. 2013), SoyTFKB (Yu et al. 2016), RIKEN (Iida et al. 2005), SoyDB (Wang et al. 2010), RiceSRTFDB (Priya and Jain 2013), and IT3F (Bailey et al. 2008). Besides, many other general databases such as Green PhylDB (Rouard et al. 2010), Phytozome (Goodstein et al. 2012), PLAZA (Proost et al. 2015), ProFITS (Ling et al. 2010), AthaMap (Bülow et al. 2009), PLEXdb (Dash et al. 2012), and more others also provided various internal utilities to investigate the TFs from structural and functional aspects. However, only few databases such as PlnTFDB (Pérez-Rodríguez et al. 2009), PlantTFcat (Dai et al. 2013), ProFITS (Ling et al. 2010), GrassCoregDB in Grassius (Yilmaz et al. 2009), PlanTAPDB (Richardt et al. 2007) give annotation of transcription coregulators (TCs; Mannervik et al. 1999) which are proteins without DNA-binding domains but could bind to TFs or RNA polymerase II to mediate the gene regulation in plants.
2 General Bioinformatics Database Resources for Plant TF
Various biological databases with general contents provide internal tools or utilities to search TFs from various aspects based on their data types and structures. For example, Phytozome is a publicly available plant-based comparative genomics portal (Table 2; Goodstein et al. 2012). As of release v1, it allows access to 65 sequenced and annotated plant genomes. Individual genes has been annotated with PANTHER, KOG, PFAM, KEGG, and GO assignments. Different search options are available for TF exploration. TFs could be either directly searched via “Keyword search” or they could be retrieved based on templates which are predefined queries using PhytoMine interface of Phytozome . PLAZA is another plant-based comparative genomics portal harboring genomic data from different genome sequencing initiatives (Table 2; Proost et al. 2015). As of release PLAZA 3. 0, database deposits the genome sequences from 38 species. TFs could be queried via search option by selecting “TF family” or “Gene” options from the menu. In addition, links to other specialized databases are useful option for cross-validation. GreenPhylDB is other web resource for comparative and functional genomics in 37 different plant species (Table 2; Rouard et al. 2010). The “Transcription factors” option under “Gene Family lists” menu is specifically designed to display the list of transcription factor families. The number of TFs could be graphically displayed, and TF information could be mined. ProFITS is a database aiming to facilitate the studies on signal transduction systems in maize (Table 2; Ling et al. 2010). It also categorizes TF families and other transcriptional regulators. TFs could be searched from “Transcription factor” tab by browsing. AthaMap is a species-specific web resource dedicated to Arabidopsis providing a genome-wide map of putative TFs and small RNA binding sites (Table 2; Bülow et al. 2009). It contains a complete list of 211 TFs derived from published TF binding specificities available as proven single binding sites or alignment matrices. TFs could be explored using different search functions under “Tools” menu. PLEXdb is a combined gene expression database for plants and plant pathogens (Table 2; Dash et al. 2012). It currently provides a genotype to phenotype information from 14 different species. The expression profiles of TFs and their target genes can be analyzed using array and/or RNA-seq data on relevant experiments. ATTED-II is a plant co-expression database integrating various co-expression data sets and network analysis tools (Table 2; Aoki et al. 2015). As of release v8. 0, it harbors eight microarray and six RNA sequencing-derived co-expression data from seven dicot species, such as Arabidopsis, soybean, tomato, field mustard, medick, grape, and poplar, and from two monocot species of maize and rice. The interacting gene sets are inclined to be co-expressed, thereby dissecting a co-expression network of TFs could provide very useful information about functional gene relationships (Aoki et al. 2015). CORNET is a system biology portal for Arabidopsis and maize integrating co-expressions, regulatory interactions (e. g. , TFs), gene associations, protein-protein interactions (PPIs), and functional annotations (Table 2; De Bodt et al. 2012). Interactions of TFs among themselves and between TFs and their targets demonstrate intricate regulatory cascades; thus, a holistic approach like system biology is an effective way in understanding the complex regulatory networks. Multiple options in CORNET are available to construct the networks centralized around input genes or proteins.
Those abovementioned databases with general content but enabling regulatory element explorations with various internal utilities and functionalities are only some glimpse from flourishing a number of bioinformatics resources. Thus, development of new versatile resources is an emerging issue to further understand the transcription regulatory mechanisms from various aspects, particularly system levels.
3 Specialized Bioinformatics Database Resources for Plant TF and Regulatory Element Search
In addition to general databases, many specialized TF databases have been also developed for TF and regulatory element exploration in plants such as AGRIS (Arabidopsis Gene Regulatory Information Server), PlantTFDB (Plant Transcription Factor Database), PlnTFDB (Plant Transcription Factor Database), GRASSIUS (Grass Regulatory Information Server), PlantTFcat (Plant Transcription Factor Categorization and Analysis Tool), TreeTFDB (Tree Transcription Factor Database), PlanTAPDB (Plant Transcription Associated Protein Database), TOBFAC (Database of Tobacco Transcription Factors), ppdb (plant promoter database), and PlantCARE (Plant Cis-Acting Regulatory Element).
3.1 AGRIS (Arabidopsis Gene Regulatory Information Server)
AGRIS is an Arabidopsis-specific database resource providing information on promoter sequences, TFs, and their target genes (Table 3; Yilmaz et al. 2011). AGRIS contains three distinct databases such as AtcisDB, AtTFDB, and AtRegNet. AtcisDB (arabidopsis. med. ohio-state. edu/AtcisDB/) contains the information about 33,000 upstream regions of annotated Arabidopsis genes and describes the validation of cis-regulatory elements as experimentally or predicted. It is composed of different data types such as promoter sequence, TF binding site information, and associated annotations, and data can be searched by TAIR gene symbol or locus ID. AtTFDB (arabidopsis. med. ohio-state. edu/AtTFDB/) includes the information about 1,770 TFs grouped in 50 families based on domain conservancy. The users can search the database by using a specific locus ID or gene name or browsing the TF families. AtRegNet (arabidopsis. med. ohio-state. edu/grgx/) harbors 18,772 direct interactions between TFs and target genes. For example, employing AGRIS , interaction between BR-activated transcription factor (BZR1 ) and phytochrome-interacting factor4 (PIF4 ) was demonstrated to integrate the brassinosteroid and environmental responses (Oh et al. 2012).
3.2 PlantTFDB (Plant Transcription Factor Database)
PlantTFDB contains the 320,370 TFs from 165 plant species classified into 58 families (Table 3; Jin et al. 2013). Very extensive annotations have been provided for each identified TF such as functional domains, binding motifs, gene and plant ontologies, 3D structures, regulation information, curated functional description, interaction, expression information, references, and cross-link to various databases. The evolutionary relationships between TFs were provided by constructing the phylogenies and inferring the orthologous groups. Database could be searched using TF IDs and common names in “search” tab or providing sequences in BLAST. In addition, new portals are also internally available for regulation prediction and functional enrichment at PlantRegMap (Plant Transcriptional Regulatory Map) and for architecture and evolutionary features of transcriptional regulatory networks at ATRM (Arabidopsis Transcriptional Regulatory Map). For example, an Arabidopsis transcriptional regulatory map constructed with 388 TFs from 47 families showed the architectural heterogeneity in stress response and developmental subnetworks and demonstrated three types of new network motifs (Jin et al. 2015).
3.3 PlnTFDB (Plant Transcription Factor Database)
PlnTFDB covers the 28,193 protein models and 26,184 distinct protein sequences distributed in 84 gene families from 20 plant species (Table 3; Pérez-Rodríguez et al. 2009). It is an integrative database providing information on TFs and other TRs in completely sequenced and annotated plant species. For example, chickpea transcripts were queried against PlnTFDB TFs to identify all TF families in chickpea transcriptome using BLASTX (Garg et al. 2011). Each gene family is provided with a basic description complemented by literature reference as well as with domain alignment. TF/TR entries also cover information of expressed sequence tags (ESTs), domain architecture, 3D structures of homologue proteins, and other cross-links to various resources. The different species are also associated to each other with orthologous genes to facilitate the cross-species comparisons. Database search could be realized using sequence identifiers, blasting, or direct browsing.
3.4 GRASSIUS (Grass Regulatory Information Server)
GRASSIUS is a web portal including various resources related to the control of gene expression in grass species such as maize, rice, sorghum, sugarcane, and Brachypodium (Table 3; Yilmaz et al. 2009). Database currently contains 9,044 TFs, 579 coregulators, 149,075 promoter sequences, and 2,098 TF ORF clones. It harbors four different databases such as GrassTFDB , GrassCoRegDB , GrassPROMDB , and TFome Collection. GrassTFDB (grassius. org/grasstfdb. html) contains an extensive collection of TFs from maize, rice, sorghum, sugarcane, and Brachypodium. GrassCoRegDB (grassius. org/grasscoregdb. html) includes a collection of transcriptional regulator proteins. These proteins do not bind DNA at sequence-specific way. They act either through interacting with TFs or as chromatin modifiers releasing or restricting DNA accessibility. GrassPROMDB (grassius. org/grasspromdb. html) is a promoter sequence database for grass species covering the cis-regulatory elements . TFome Collection (grassius. org/tfomecollection. html) provides access to the grass TF ORFeome collection .
3.5 PlantTFcat (Plant Transcription Factor Categorization and Analysis Tool)
PlantTFcat is a web-based transcription factor and transcriptional regulator categorization and analysis tool (Table 3; Dai et al. 2013). It currently contains the information from a total of 108 published TF, TR, and chromatin regulator (CR) families. Database can be searched using protein or nucleic acid sequences in FASTA format or as pure sequence (without FASTA header). In many studies, PlantTFcat was employed in identification of TFs in various plant species including Vicia sativa (Panchal 2015), Phaseolus vulgaris (Patel et al. 2014a), Cicer arietinum (Patel et al. 2014b), Trigonellafoenum graecum (Patel et al. 2014c), Arachis hypogaea (Patel et al. 2015), Andrographis paniculata (Cherukupalli et al. 2016), Ananas comosus (Chen et al. 2016), Brassica napus (Shamloo-Dashtpagerdi et al. 2015), and some others.
3.6 TreeTFDB (Tree Transcription Factor Database)
TreeTFDB contains TFs from six economically valuable tree species such as papaya (Carica papaya), jatropha (Jatropha curcas), cassava (Manihot esculenta), poplar (Populus trichocarpa), castor bean (Ricinus communis), and grapevine (Vitis vinifera) to provide resource for comparative and functional genomics (Table 3; Mochida et al. 2013). The importance of specialized databases like TreeTFDB which is dedicated to tree species has also been emphasized in a mini review (Legué et al. 2014). As of ver. 1. 0, it includes 1,481 TF models of jatropha, 3,110 TF models of poplar, 2,638 TF models of cassava, 1,493 TF models of grapevine, 1,552 TF models of papaya, and 1,512 TF models of castor bean. A number of search options are also available for TF exploration such as using TF families, keyword, gene IDs, InterProScan result, GO terms, cis-motif (stress responsive), cis-motif (PLACE), cis-motif (hormone responsive), and employing blast options.
3.7 PlanTAPDB (Plant Transcription Associated Protein Database)
PlanTAPDB is a phylogeny-based web resource for transcription-associated proteins (TAPs) in a vast taxonomic range including algae and a moss (Table 3; Richardt et al. 2007). Database contains information on three categories of entries such as transcription factors (TFs), transcription regulators (TRs), and putative TAPs (PTs) that belong to one of 119 families (138 subfamilies). TAPs can be searched by family accession numbers and IDs, keyword search, and also being queried using BLAST.
3.8 TOBFAC (Database of Tobacco Transcription Factors)
TOBFAC is an integrative database dedicated to tobacco plant (Table 3; Rushton et al. 2008). It provides access to sequence, phylogeny, and various associated data for tobacco TFs. As of current, database includes 65 TF families each provided with literature information, domain architecture, list of genomic sequences, minimum number of genes, and other information. TOBFAC can be queried using various search and data retrieval options on main menu. Database provides very useful resources for tobacco studies. For example, in a custom oligo array design for transcriptome analyses in water-deficit tobacco, probe sequences were obtained from three different sources including TOBFAC TFs (Rabara et al. 2015). In many very recent studies, TOBFAC database has been also employed (Fu et al. 2013; Ogata et al. 2013; Xu et al. 2015).
3.9 ppdb (Plant Promoter Database)
ppdb is a plant promoter database providing information on core promoter structures such as TATA boxes, Y Patches, Initiators, CA and GA elements, transcription start sites (TSSs), and transcriptional regulatory elements from A. thaliana, Oryza sativa, Physcomitrella patens, and P. trichocarpa (Table 3; Hieno et al. 2013). For example, ABA-responsive promoter elements (ABREs) identified by ppdb were reported to investigate the relationship between stomatal closure and ABA signaling evolution (Lind et al. 2015). Database can be searched by using a gene name or a keyword. In addition, a “homologue gene search” option is available to compare the promoter structures of orthologous genes in specified plants.
3.10 PlantCARE (Plant Cis-Acting Regulatory Element)
PlantCARE is a web portal providing information on plant cis-regulatory elements , enhancers, and repressors and also a portal for tools in silico analysis of promoter sequences (Table 3; Lescot et al. 2002). Regulatory elements are demonstrated by consensus sequences, positional matrices, individual sites, and functional annotations for queried sequences. Database can be searched by submitting raw DNA sequence or FASTA file. In addition, other query options are also available for more guided searches using classifications, genes, name of factor, name of site, and referencia. PlantCARE is a very useful source for exploration of predicted or verified cis-regulatory elements in given sequences (Filiz et al. 2015; Tira-Umphon et al. 2015; Vatansever et al. 2016).
4 Conclusion and Perspectives
Biological processes are controlled at multiple levels; at transcriptional level it is regulated by TFs which modulate the expression of target genes via binding cis-regulatory elements in their promoter regions. Identification of TFs is a primary step in TF research which is usually achieved by a direct blasting against a particular plant genome or against a gene/protein database repository. Following TF identification , domain families of identified TFs are first needed to be confirmed using a domain search utility like Pfam (pfam. xfam. org/; Finn et al. 2016), InterProScan (ebi. ac. uk/interpro/search/sequence-search; Mitchell et al. 2014), PROSITE (prosite. expasy. org/; Sigrist et al. 2012), or NCBI Conserved Domain (ncbi. nlm. nih. gov/Structure/cdd/wrpsb. cgi; Marchler-Bauer et al. 2014). TFs with typical domain structures could be readily verified, but members with atypical domains could be difficult to identify due to the sequence divergence and reshuffling. In addition to DNA-binding domains, the presence of various other motif sequences in some TFs could also make more difficult for their identification. A statistical model known as hidden Markov model (HMM ) is commonly a used approach in search of typical and complex protein domain families (Finn et al. 2016). Besides, an iterative approach has been also reported to identify the complex domain motifs in TFs (Wang et al. 2012, 2016). However, it is an emerging issue to develop novel bioinformatics tools or approaches to efficiently identify the highly complex domain structures.
In addition, it is almost imperative to have insights about TF binding sites or cis-regulatory elements to dissect TF-based networks. For this, integration of various resources from expression, co-expression , protein-binding, and phylogenetic studies has become a powerful approach (Godoy et al. 2011; Franco-Zorrilla et al. 2014). To capture real interactions between TFs and their target sites, the chromatin immunoprecipitation (ChIP) , ChIP-microarray , ChIP-seq, and combined ChIP-seq-RNA-seq have been potential technologies so far reported (Buck & Lieb, 2004; Collas ; Kaufmann et al. 2010; Yang et al. 2013; Heyndrickx et al. 2014; Weirauch et al. 2014). Moreover, being parallel with ever-increasing biological data, the development of new databases enabling analysis and distribution of plant TFs has been fundamentally important in TF research. Some specialized and general content databases for TF research have been developed, but most of these databases are still far away from completeness due to such reasons: (1) verified and hypothetical data not clearly distinguished; (2) species-specific functional divergence ignored, particularly working with orthologues data; (3) data not sufficiently linked to relevant resources to cross-validate; (4) ambiguous entries present data not clearly annotated; (5) not regularly up-to-date; (6) less or lack of interoperability between TF resources; and (7)not included adequate analysis tools. Thus, it is a compelling demand to develop novel TF databases and enhance already ones to satisfy with abovementioned deficiencies and equip with more utilities and functionalities to improve TF research in a more promising way.
Abbreviations
- DBD:
-
DNA-binding domain
- TAP:
-
Transcription-associated proteins
- TF:
-
Transcription factor
- TR:
-
Transcriptional regulator
References
Aoki Y, Okamura Y, Tadaka S, Kinoshita K, Obayashi T (2015) ATTED-II in 2016: a plant coexpression database towards lineage-specific coexpression. Plant Cell Physiol 57(1):e5–e5
Bailey PC, Dicks J, Wang TL, Martin C (2008) IT3F: a web-based tool for functional analysis of transcription factors in plants. Phytochemistry 69(13):2417–2425
Buck MJ, Lieb JD (2004) ChIP-chip: considerations for the design, analysis, and application of genome-wide chromatin immunoprecipitation experiments. Genomics 83(3):349–360
Bülow L, Engelmann S, Schindler M, Hehl R (2009) AthaMap, integrating transcriptional and post-transcriptional data. Nucleic Acids Res 37(suppl 1):D983–D986
Chen C, Zhang Y, Xu Z, Luan A, Mao Q, Feng J, Xie T, Gong X, Wang X, Chen H, He Y (2016) Transcriptome profiling of the pineapple under low temperature to facilitate its breeding for cold tolerance. PloS One 11(9):e0163315
Cherukupalli N, Divate M, Mittapelli SR, Khareedu VR, Vudem DR (2016) De novo assembly of leaf transcriptome in the medicinal plant Andrographispaniculata. Front Plant Sci 7:1203
Collas P (2010) The current state of chromatin immunoprecipitation. Mol Biotechnol 45(1):87–100
Dai X, Sinharoy S, Udvardi M, Zhao PX (2013) PlantTFcat: an online plant transcription factor and transcriptional regulator categorization and analysis tool. BMC Bioinforma 14(1):1
Dash S, Van Hemert J, Hong L, Wise RP, Dickerson JA (2012) PLEXdb: gene expression resources for plants and plant pathogens. Nucleic Acids Res 40(D1):D1194–D1201
De Bodt S, Hollunder J, Nelissen H, Meulemeester N, Inzé D (2012) CORNET 2. 0: integrating plant coexpression, protein–protein interactions, regulatory interactions, gene associations and functional annotations. New Phytol 195(3):707–720
Filiz E, Ozyigit II, Vatansever R (2015) Genome-wide identification of galactinol synthase (GolS) genes in Solanum lycopersicum and Brachypodium distachyon. Comput Biol Chem 58:149–157
Finn RD, Coggill P, Eberhardt RY, Eddy SR, Mistry J, Mitchell AL, Potter SC, Punta M, Qureshi M, Sangrador-Vegas A, Salazar GA, Tate J, Salazar GA (2016) The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res 44(D1):D279–D285
Franco-Zorrilla JM, Solano R (2016) Identification of plant transcription factor target sequences. BiochimicaetBiophysicaActa (BBA)-Gene Regulatory Mechanisms 1860(1):21–30
Franco-Zorrilla JM, López-Vidriero I, Carrasco JL, Godoy M, Vera P, Solano R (2014) DNA-binding specificities of plant transcription factors and their potential to define target genes. Proc Natl Acad Sci 111(6):2367–2372
Fu Y, Guo H, Cheng Z, Wang R, Li G, Huo G, Liu W (2013) NtNAC-R1, a novel NAC transcription factor gene in tobacco roots, responds to mechanical damage of shoot meristem. Plant Physiol Biochem 69:74–81
Garg R, Patel RK, Tyagi AK, Jain M (2011) De novo assembly of chickpea transcriptome using short reads for gene discovery and marker identification. DNA Res 18(1):53–63
Godoy M, Franco-Zorrilla JM, Pérez-Pérez J, Oliveros JC, Lorenzo Ó, Solano R (2011) Improved protein-binding microarrays for the identification of DNA-binding specificities of transcription factors. Plant J 66(4):700–711
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N, Rokhsar DS (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40(D1):D1178–D1186
Guilfoyle TJ, Ulmasov T, Hagen G (1998) The ARF family of transcription factors and their role in plant hormone-responsive transcription. Cell Mol Life Sci CMLS 54(7):619–627
Heyndrickx KS, Van de Velde J, Wang C, Weigel D, Vandepoele K (2014) A functional and evolutionary perspective on transcription factor binding in Arabidopsis thaliana. Plant Cell 26(10):3894–3910
Hieno A, Naznin HA, Hyakumachi M, Sakurai T, Tokizawa M, Koyama H, Sato N, Nishiyama T, Hasebe M, Zimmer AD, Lang D, Reski R, Rensing SA, Obokata J, Yamamoto YY (2013) ppdb: plant promoter database version 3. 0. Nucleic Acids Res 42:D1 D1188–D1192
Iida K, Seki M, Sakurai T, Satou M, Akiyama K, Toyoda T, Konagaya A, Shinozaki K (2005) RARTF: database and tools for complete sets of Arabidopsis transcription factors. DNA Res 12(4):247–256
Jin J, Zhang H, Kong L, Gao G, Luo J (2013) PlantTFDB 3. 0: a portal for the functional and evolutionary study of plant transcription factors. Nucleic Acids Res 42:D1182–D1187
Jin J, He K, Tang X, Li Z, Lv L, Zhao Y et al (2015) An Arabidopsis transcriptional regulatory map reveals distinct functional and evolutionary features of novel transcription factors. Mol Biol Evol 32(7):1767–1773
Kaufmann K, Muino JM, Østerås M, Farinelli L, Krajewski P, Angenent GC (2010) Chromatin immunoprecipitation (ChIP) of plant transcription factors followed by sequencing (ChIP-SEQ) or hybridization to whole genome arrays (ChIP-CHIP). Nat Protoc 5(3):457–472
Legué V, Rigal A, Bhalerao RP (2014) Adventitious root formation in tree species: involvement of transcription factors. Physiologiaplantarum 151(2):192–198
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30(1):325–327
Lin JJ, Yu CP, Chang YM, Chen SCC, Li WH (2014) Maize and millet transcription factors annotated using comparative genomic and transcriptomic data. BMC Genomics 15(1):1
Lind C, Dreyer I, López-Sanjurjo EJ, von Meyer K, Ishizaki K, Kohchi T, Lang D, Zhao Y, Kreuzer I, Al-Rasheid KA, Ronne H, Reski R, Zhu JK, Geiger D, Hedrich R (2015) Stomatal guard cells co-opted an ancient ABA-dependent desiccation survival system to regulate stomatal closure. Curr Biol 25(7):928–935
Ling Y, Du Z, Zhang Z, Su Z (2010) ProFITS of maize: a database of protein families involved in the transduction of signalling in the maize genome. BMC Genomics 11(1):580
Mannervik M, Nibu Y, Zhang H, Levine M (1999) Transcriptional coregulators in development. Science 284(5414):606–609
Marchler-Bauer A, Derbyshire MK, Gonzales NR, Lu S, Chitsaz F, Geer LY, Geer RC, He J, Gwadz M, Hurwitz DI, Lanczycki CJ, Lu F, Marchler GH, Song JS, Thanki N, Wang Z, Yamashita RA, Zhang D, Zheng C, Bryant SH (2014) CDD: NCBI’s conserved domain database. Nucleic Acids Res 43:D222–D226
Mitchell A, Chang HY, Daugherty L, Fraser M, Hunter S, Lopez R, McAnulla C, McMenamin C, Nuka G, Pesseat S, Sangrador-Vegas A, Scheremetjew M, Rato C, Yong SY, Bateman A, Punta M, Attwood TK, Sigrist CJ, Redaschi N, Rivoire C, Xenarios I, Kahn D, Guyot D, Bork P, Letunic I, Gough J, Oates M, Haft D, Huang H, Natale DA, Wu CH, Orengo C, Sillitoe I, Mi H, Thomas PD, Finn RD (2014) The InterPro protein families database: the classification resource after 15 years. Nucleic Acids Res 43:D213–D221
Mochida K, Yoshida T, Sakurai T, Yamaguchi-Shinozaki K, Shinozaki K, Tran LSP (2013) TreeTFDB: an integrative database of the transcription factors from six economically important tree crops for functional predictions and comparative and functional genomics. DNA Res 20:151–162
Nole-Wilson S, Krizek BA (2000) DNA binding properties of the Arabidopsis floral development protein AINTEGUMENTA. Nucleic Acids Res 28(21):4076–4082
Ogata T, Kida Y, Tochigi M, Matsushita Y (2013) Analysis of the cell death-inducing ability of the ethylene response factors in group VIII of the AP2/ERF family. Plant Sci 209:12–23
Oh E, Zhu JY, Wang ZY (2012) Interaction between BZR1 and PIF4 integrates brassinosteroid and environmental responses. Nat Cell Biol 14(8):802–809
Ohme-Takagi M, Shinshi H (1995) Ethylene-inducible DNA binding proteins that interact with an ethylene-responsive element. Plant Cell 7(2):173–182
Panchal HJ (2015) De novo RNA Seq assembly and annotation of Vicia sativa L. (SRR403901). Genomics Appl Biol 6(2)
Patel SS, Shah DB, Panchal HJ (2014a) De novo RNA Seq assembly and annotation of Phaseolus Vulgaris L. (SRR1283084). Genomics Appl Biol 5(5)
Patel SS, Shah DB, Panchal HJ (2014b) De novo RNA Seq assembly and annotation of Cicer arietinum L. (SRR627764). Legum Genomics Genet, 5(5)
Patel S, Shah DB, Panchal HJ (2014c) De novo RNA Seq assembly and annotation of Trigonellafoenum-graecum L. (SRR066197). Legum Genomics Genet, 5(7)
Patel SS, Shah DB, Panchal HJ (2015) De novo transcriptome analysis of Arachis Hypogaea L. (SRR1212866). OMICS Res 5
Pérez-Rodríguez P, Riano-Pachon DM, Corrêa LGG, Rensing SA, Kersten B, Mueller-Roeber B (2009) PlnTFDB: updated content and new features of the plant transcription factor database. Nucleic Acids Res 38:D822–D827
Priya P, Jain M (2013) RiceSRTFDB: a database of rice transcription factors containing comprehensive expression, cis-regulatory element and mutant information to facilitate gene function analysis. Database 2013:bat027
Proost S, Van Bel M, Vaneechoutte D, Van de Peer Y, Inzé D, Mueller-Roeber B, Vandepoele K (2015) PLAZA 3. 0: an access point for plant comparative genomics. Nucleic Acids Res 43(D1):D974–D981
Rabara RC, Tripathi P, Choudhary MK, Timko MP, Shen QJ, Rushton PJ (2015) Transcriptome profiling of tobacco under water deficit conditions. Genomics Data 5:61–63
Richardt S, Lang D, Reski R, Frank W, Rensing SA (2007) PlanTAPDB, a phylogeny-based resource of plant transcription-associated proteins. Plant Physiol 143(4):1452–1466
Riechmann JL, Heard J, Martin G, Reuber L, Jiang CZ, Keddie J, Adam L, Pineda O, Ratcliffe OJ, Samaha RR, Creelman R, Pilgrim M, Broun P, Zhang JZ, Ghandehari D, Sherman BK, Yu G (2000) Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 290(5499):2105–2110
Rouard M, Guignon V, Aluome C, Laporte MA, Droc G, Walde C, Zmasek CM, Périn C, Conte MG (2010) GreenPhylDB v2. 0: comparative and functional genomics in plants. Nucleic Acids Res 39:D1095–D1102
Rushton PJ, Bokowiec MT, Laudeman TW, Brannock JF, Chen X, Timko MP (2008) TOBFAC: the database of tobacco transcription factors. BMC Bioinforma 9(1):1
Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T, Nelson W, Hyten DL, Song Q, Thelen JJ, Cheng J, Xu D, Hellsten U, May GD, Yu Y, Sakurai T, Umezawa T, Bhattacharyya MK, Sandhu D, Valliyodan B, Lindquist E, Peto M, Grant D, Shu S, Goodstein D, Barry K, Futrell-Griggs M, Abernathy B, Du J, Tian Z, Zhu L, Gill N, Joshi T, Libault M, Sethuraman A, Zhang XC, Shinozaki K, Nguyen HT, Wing RA, Cregan P, Specht J, Grimwood J, Rokhsar D, Stacey G, Shoemaker RC, Jackson SA (2010) Genome sequence of the palaeopolyploid soybean. Nature 463(7278):178–183
Shamloo-Dashtpagerdi R, Razi H, Ebrahimie E (2015) Mining expressed sequence tags of rapeseed (Brassica napus L. ) to predict the drought responsive regulatory network. Physiol Mol Biol Plants 21(3):329–340
Sigrist CJ, De Castro E, Cerutti L, Cuche BA, Hulo N, Bridge A, Bougueleret L, Xenarios I (2012) New and continuing developments at PROSITE. Nucleic Acids Res 41:D344–D347
Tira-Umphon A, Roustan JP, Chervin C (2015) The stimulation by ethylene of the UDP glucose-flavonoid 3-O-glucosyltransferase (UFGT) in grape tissues is independent from the MybA transcription factors. VITIS-J Grapevine Res 46(4):210
Udvardi MK, Kakar K, Wandrey M, Montanari O, Murray J, Andriankaja A, Zhang JY, Benedito V, Hofer JM, Chueng F, Town CD (2007) Legume transcription factors: global regulators of plant development and response to the environment. Plant Physiol 144(2):538–549
Ulmasov T, Hagen G, Guilfoyle TJ (1997) ARF1, a transcription factor that binds to auxin response elements. Science 276(5320):1865–1868
Vatansever R, Koc I, Ozyigit II, Sen U, Uras ME, Anjum NA et al (2016) Genome-wide identification and expression analysis of sulfate transporter (SULTR) genes in potato (Solanum tuberosum L. ) Planta 244:1167–1183
Wang Z, Libault M, Joshi T, Valliyodan B, Nguyen HT, Xu D, Stacey G, Cheng J (2010) SoyDB: a knowledge database of soybean transcription factors. BMC Plant Biol 10(1):14
Wang Y, Deng D, Zhang R, Wang S, Bian Y, Yin Z (2012) Systematic analysis of plant-specific B3 domain-containing proteins based on the genome resources of 11 sequenced species. Mol Biol Rep 39(5):6267–6282
Wang Y, Lu W, Deng D (2016) Bioinformatic landscapes for plant transcription factor system research. Planta 243(2):297–304
Weirauch MT, Yang A, Albu M, Cote AG, Montenegro-Montero A, Drewe P, Najafabadi HS, Lambert SA, Mann I, Cook K, Zheng H, Goity A, van Bakel H, Lozano JC, Galli M, Lewsey M, Huang E, Mukherjee T, Chen X, Reece-Hoyes JS, Govindarajan S, Shaulsky G, AJM W, Bouget FY, Ratsch G, Larrondo LF, Ecker JR, Hughes TR (2014) Determination and inference of eukaryotic transcription factor sequence specificity. Cell 158(6):1431–1443
Wingender E (1988) Compilation of transcription regulating proteins. Nucleic Acids Res 16(5 Pt B):1879
Xu F, Xiao W, Li J, Ding C, Li S, Liu W, Guo H (2015) NtGRAS-R1, a topping responsive transcription regulator in tobacco roots. Acta Physiol Plant 37(9):1–8
Yamasaki K, Kigawa T, Seki M, Shinozaki K, Yokoyama S (2013) DNA-binding domains of plant-specific transcription factors: structure, function, and evolution. Trends Plant Sci 18(5):267–276
Yang JH, Li JH, Jiang S, Zhou H, Qu LH (2013) ChIPBase: a database for decoding the transcriptional regulation of long non-coding RNA and microRNA genes from ChIP-Seq data. Nucleic Acids Res 41(D1):D177–D187
Yilmaz A, Nishiyama MY, Fuentes BG, Souza GM, Janies D, Gray J, Grotewold E (2009) GRASSIUS: a platform for comparative regulatory genomics across the grasses. Plant Physiol 149(1):171–180
Yilmaz A, Mejia-Guerra MK, Kurz K, Liang X, Welch L, Grotewold E (2011) AGRIS: the Arabidopsis gene regulatory information server, an update. Nucleic Acids Res 39(suppl 1):D1118–D1122
Yu Y, Wang N, Hu R, Xiang F (2016) Genome-wide identification of soybean WRKY transcription factors in response to salt stress. SpringerPlus 5(1):1–15
Zheng Y, Jiao C, Sun H, Rosli HG, Pombo MA, Zhang P, Banf M, Dai X, Martin G, Zhao PX et al. (2016) iTAK: a program for genome-wide prediction and classification of plant transcription factors, transcriptional regulators, and protein kinases. Mol Plant 9(12):1667–1670
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Filiz, E., Vatansever, R., Ozyigit, I.I. (2017). Bioinformatics Database Resources for Plant Transcription Factors. In: Hakeem, K., Malik, A., Vardar-Sukan, F., Ozturk, M. (eds) Plant Bioinformatics. Springer, Cham. https://doi.org/10.1007/978-3-319-67156-7_5
Download citation
DOI: https://doi.org/10.1007/978-3-319-67156-7_5
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-67155-0
Online ISBN: 978-3-319-67156-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)