Abstract
The focus of this chapter is on gene families encoding enzymes of phenylpropanoid pathway in common bean. The introductory section contains a short overview of the phenylpropanoid pathway. Section 11.2 introduces major gene families encoding enzymes of this pathway in common bean, soybean, and Arabidopsis in the current annotations of their complete genome sequences (Phaseolus vulgaris v1.0, Glycine max Wm82.a2.v1, and Arabidopsis thaliana TAIR10) deposited in Phytozome 10.2. For each of the 21 enzyme classes, their functional annotations were based on the commonly used Pfam and KOG databases, while the number of genes in each family was based on Phytozome and KEGG databases. Section 11.3 describes cytochrome P450s involved in the phenylpropanoid pathway with particular emphasis on ten families included in the general (central) phenylpropanoid pathway, C4H (family CYP73A), in the lignin/lignan branch, C3H (family CYP98A) and F5H (family CYP84A), in the flavonoid/anthocyanin/proanthocyanidin branch, F3′H (family CYP75B), F3′5′H (family CYP75A), and FNS (family CYP93B), and in the isoflavonoid branch IFS (family CYP93C), I2′H (family CYP81E), F6H (family CYP71D), and D6aH (family CYP93A). The availability of the complete genome sequences enabled a thorough inventory of putative P450 genes encoding enzymes of this metabolic pathway. The P450 gene sequences from common bean were compared to homologs from Arabidopsis and soybean and confirmed with the information published for both soybean and common bean genomes. Cinnamate 4-hydroxylase (C4H) is the first P450 enzyme in the phenylpropanoid pathway and is described in detail in Sect. 11.4. It belongs to the relatively small CYP73A gene family. Genome locations and gene structures including cis-regulatory regions in 5′UTRs (5′ regulatory sequences) are detailed for this family in common bean. In addition, the expression patterns of these genes in different tissues (Phytozome 10.2) and syntenic relationships (Plant Genome Duplication Database) between common bean and soybean were examined. Finally, genes encoding the C4H enzyme in landrace G19833 (Andean gene pool, Phytozome 10.2) and in cultivar OAC Rex (Mesoamerican gene pool) were compared and searched for polymorphisms. These sequence differences can be used to develop C4H gene-based marker(s) to explore the roles of these genes in various processes such as lignin or anthocyanin biosynthesis.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Common bean
- Cytochrome P450 superfamily
- C4H gene
- Genome sequence
- In silico
- Phenylpropanoid pathway
- Synteny
In this chapter:
-
Common names in plants: Arabidopsis (Arabidopsis thaliana), soybean (Glycine max), common bean (Phaseolus vulgaris )
-
Chromosome-based locus (gene model) identifier (Phytozome)
11.1 Introduction
As sessile organisms, plants produce numerous secondary metabolites to overcome biotic and abiotic stressors, to attract pollinators and nitrogen-fixing microorganisms, and to communicate with other plants (Koes et al. 2005; Noel et al. 2005; Moura et al. 2010; Agati et al. 2012, 2013; Baxter and Stewart 2013). Many of these compounds are synthesized by the phenylpropanoid pathway, which is likely one of the most studied pathways in plants. It is relatively well understood and was extensively reviewed (Goujon et al. 2003; Raes et al. 2003; Wang 2011; Falcone Ferreyra et al. 2012; Petrussa et al. 2013). Individual branches of the pathway have been thoroughly characterized. Most of the enzymes that catalyze individual steps of the pathway have been identified, and the genes coding for them have been isolated in a number of plant species, including Arabidopsis and soybean (Graham et al. 2008; Fraser and Chapple 2011).
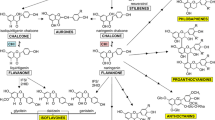
The core (general or central) pathway consists of three steps, including (1) the conversion of the aromatic amino acid phenylalanine into trans-cinnamic acid, which is catalyzed by phenylalanine ammonia-lyase (PAL); (2) the conversion of trans-cinnamic acid into p-coumaric acid, catalyzed by cinnamate 4-hydroxylase (C4H); and (3) the transformation of p-coumaric acid into p-coumaroyl-CoA, catalyzed by 4-coumarate:CoA ligase (4CL). The compound p-coumaroyl-CoA serves as a starting point for several branches of the phenylpropanoid pathway leading to biosynthesis of lignin, lignans, coumarins, stilbenes, flavonoids, anthocyanin, condensed tannins (proanthocyanidins), and isoflavonoids (Vogt 2010; Cheynier et al. 2013). These products have important functions not only for plant survival, growth, and development but they could also be powerful supplements to the human diet. For example, lignans, stilbenes, and isoflavonoids have been associated with the reduced onset/development of certain chronic disease in humans, including some forms of cancer and heart diseases (Cassidy et al. 2000; Chen et al. 2006; Adlercreutz 2007; Xiao 2008; Brunetti et al. 2013) (Fig. 11.1).
Cytochrome P450s involved in the phenylpropanoid pathway. The positions of ten enzymes and locus (gene model) identifiers (https://phytozome.jgi.doe.gov/pz/portal.html) in the pathway in common bean (blue), soybean (red), and Arabidopsis (black) are indicated; 1. Core phenylpropanoid pathway: cinnamate 4-hydroxylase (C4H, CYP73A); 2. Lignin/lignans branch: coumarate 3-hydroxylase (C3H, CYP98A) and ferulic acid 5-hydroxylase (F5H, CYP84A); 3. Anthocyanins/condensed tannins branch: flavonoid 3′-hydroxylase (F3′H, CYP75B), flavonoid 3′,5′-hydroxylase (F3′,5′H, CYP75A) and flavone synthase (FNS, CYP93B); 4. Isoflavonoid branch: isoflavone synthase (IFS, CYP93C), isoflavone 2′-hydroxylase (I2′H, CYP81E), flavonoid 6-hydroxylase (F6H, CYP71D), and 3,9-dihydroxypterocarpan 6a-monooxygenase (D6aH, CYP93A)
Lignin biosynthesis is a two-step process. First, monolignol is synthesized through a series of hydroxylations, O-methylations, and conversions of side-chain carboxyl into p-coumaryl, coniferyl, and sinapyl alcohols (Humphreys and Chapple 2002; Boerjan et al. 2003; Vanholme et al. 2010; Weng and Chaple 2010; Labeeuw et al. 2015). A second step involves monolignol polymerization by peroxidases (PER), laccases (LAC), and dirigent proteins (DP). In a reversible reaction, hydroxycinnamoyl-CoA:shikimate/quinate hydroxycinnamoyltransferase (HCT) converts p-coumaroyl-CoA and caffeoyl-CoA into their corresponding shikimate/quinate esters, which are then transformed by coumarate 3-hydroxylase (C3H) into their corresponding caffeoyl esters (Schoch et al. 2001). Caffeoyl-CoA O-methyltransferase (CCoAOMT) catalyzes methylation of caffeoyl-CoA to generate feruloyl-CoA. Cinnamoyl-CoA reductase (CCR) converts hydroxycinnamoyl-CoA esters into their corresponding aldehydes, and cinnamyl-alcohol dehydrogenase (CAD) catalyzes the conversion of cinnamyl aldehydes into their corresponding alcohols. Ferulic acid 5-hydroxylase (F5H) converts ferulic acid into 5-hydroxyferulic acid. F5H is also known as coniferaldehyde 5-hydroxylase (CAld5H), since the enzyme preferably transforms coniferaldehyde and/or coniferyl alcohol into synapaldehyde and/or sinapyl alcohol, respectively (Humphreys et al. 1999; Osakabe et al. 1999). Caffeic acid O-methyltransferase (COMT) converts 5-hydroxyconiferaldehyde and/or 5-hydroxyconiferyl alcohol into sinapaldehyde and/or sinapyl alcohol, respectively (Osakabe et al. 1999; Parvathi et al. 2001; Zubieta et al. 2002). COMT was previously thought to be a bifunctional enzyme, methylating caffeic and 5-hydroxyferulic acids.
Chalcone synthase (CHS) is the first enzyme in the flavonoid/anthocyanin branch of the phenylpropanoid pathway. It catalyzes the biosynthesis of chalcone from one molecule of p-coumaroyl-CoA with three molecules of malonyl-CoA. This basic flavonoid structure is then transformed by a set of various isomerases, reductases, hydroxylases, Fe2+/2-oxoglutarate-dependent dioxygenases, and transferases into different flavonoids, including flavanones, flavones, flavonols, anthocyanins, and condensed tannins (Winkel-Shirley 2001; Ralston et al. 2005; Ferrer et al. 2008; Saito et al. 2013). CHS and chalcone isomerase (CHI) catalyze the two-step condensation, producing a colorless flavanone (naringenin), which is then oxidized by flavanone 3-hydroxylase (F3H) into the colorless dihydroflavonol (dihydrokaempferol). Subsequent hydroxylation of this compound (at the 3′ or 5′ position of the B-ring), catalyzed by flavonoid 3′-hydroxylase (F3′H) and flavonoid 3′,5′-hydroxylase (F3′5′H), produces dihydroquercetin and dihydromyricetin. These two enzymes (F3′H and F3′5′H) can also hydroxylate flavanone (naringenin) to produce eriodictyol and pentahydroxy-flavanone, which are then hydroxylated by F3H into dihydroquercetin and dihydromyricetin, respectively. The next step in the pathway is the conversion of the three dihydroflavonols (dihydroquercetin, dihydrokaempferol, and dihydromyricetin). These compounds can be transformed into flavonols (kaempferol, quercetin, and myricetin) by flavonol synthases (FLS). Dihydroflavonol 4-reductase (DFR) converts dihydroflavonols into leucoanthocyanidins (colorless flavan-3,4-diols: leucocyanidin, leucopelargonidin, and leucodelphinidin), which are then oxidized by anthocyanin synthase [ANS, also known as leucoanthocyanidin dioxygenase (LDOX)] into colored but unstable anthocyanidins [cyanidin (red-magenta), pelargonidin (orange), and delphinidin (purple-mauve)]. Stable anthocyanins (colored) are produced by glycosylation of these compounds by the UDP-glucose:flavonoid 3-O-glucosyl transferases (UFGT). Some anthocyanins (cyanidin-3-glucoside and delphinidin-3-glucoside) may be further methylated by methyltransferases (MTs) to produce peonidin-3-glucoside and petunidin- or malvidin-3-glucoside, respectively.
Condensed tannins are synthesized through two branches of the anthocyanin pathway. The reduction of leucocyanidin to catechin (2,3-trans flavan-3-ols) is catalyzed by leucoanthocyanidin reductase (LAR), and the conversion of cyanidin into epicatechin (2,3-cis flavan-3-ols) is driven by anthocyanidin reductase (ANR). The subsequent steps catalyzed by polyphenol oxidases and condensing enzymes possibly take place in vacuoles.
Legume-specific isoflavonoids are produced through two branches of the isoflavonoid pathway having major reactions in common. The branch leading to the isoflavone genistein uses the same naringenin intermediate, which is synthesized in the flavonoid/anthocyanin branch of the phenylpropanoid pathway by a two-step condensation catalyzed by CHS and CHI (common to majority of plants) (Lozovaya et al. 2007). On the other hand, isoflavone daidzein is synthesized through the co-action of CHS and legume-specific chalcone reductase (CHR), yielding isoliquiritigenin (trihydroxychalcone), which is then transformed into liquiritigenin (dihydroxyflavanone), a core intermediate of this branch of the isoflavonoid pathway (Austin and Noel 2003). Isoflavone synthase [IFS, also known as 2-hydroxyisoflavanone synthase (2-HIS)] converts flavanone (naringenin or isoliquiritigenin) into 2-hydroxyisoflavanones (through an aryl migration of the aromatic B-ring from C-2 to C-3 position and hydroxylation in position C-2) (Steele et al. 1999; Jung et al. 2000), which are then dehydrated (formation of a double bond between C-2 and C-3) to the corresponding isoflavones (genistein and daidzein) by 2-hydroxyisoflavanone dehydratase (HID) (Akashi et al. 2005; Shimamura et al. 2007). They are further modified by isoflavonoid-specific enzymes to produce major phytoalexins, including medicarpin, biochanin A, glyceollin, pisatin, and maackiain (Latunde-Dada et al. 2001; Lozovaya et al. 2007; Artigot et al. 2013).
Biosynthesis of lignin, flavonoids/anthocyanins/proanthocyanidins, and isoflavonoids is under complex regulation. The expression of the lignin biosynthetic genes is coordinately regulated by a number of transcription factors. The majority of these genes contain a common AC cis-element, which is required for their expression in cells undergoing lignification. NST1/2/3 (NAC secondary wall thickening promoting factor 1/2/3) and Myb26/Myb83 transcription factors act as master switches to regulate biosynthesis of major secondary wall components, including cellulose, xylan, and lignin in Arabidopsis (Zhong and Ye 2009; Zhao and Dixon 2011; Hao and Mohnen 2014; Yoon et al. 2015). In Arabidopsis flavonoid pathway, genes for early biosynthetic enzymes (CHS, CHI, F3H, and F3′H) are regulated by the three functionally redundant R2R3-MYB transcription factors (MYB11, MYB12, and MYB111), while the activation of late biosynthetic genes is controlled by the R2R3-MYB/bHLH/WD40 (MBW) complex (Grotewold 2005; Hartman et al. 2005; Ramsey and Glover 2005; Gonzalez et al. 2008; Gou et al. 2011; Petrussa et al. 2013; Li et al. 2014; Xu et al. 2014, 2015). Genes of legume-specific isoflavonoid branch of phenylpropanoid pathway are regulated by a different set of transcription factors. For example, GmMYB176, a R1 MYB transcription factor, regulates CHS8 expression and isoflavonoid synthesis in soybean (Yi et al. 2010a, b; Dhaubhadel 2011). The constitutive over-expression of LjMYB14 was associated with the activation of dozen of genes coding for enzymes in the core phenylpropanoid pathway and isoflavonoid branch in Lotus japonicus (Shelton et al. 2012). At the same time, the expression of other transcription factors was altered resulting in coordinated down-regulation of the competing biosynthetic pathways.
Genes encoding the major enzymes of the phenylpropanoid pathway have been identified in a number of plant species (Tsai et al. 2006; Tohge et al. 2007; Xu et al. 2009). In most species, enzymes involved in the phenylpropanoid pathway are encoded by gene families of various sizes. For example, plants’ CADs can reduce various aldehydes, including those expressed in response to pathogens (Barakat et al. 2010; Miedes et al. 2014). The nine putative CAD genes that were identified in Arabidopsis are split into three classes based on protein phylogenetic analysis (Raes et al. 2003). Using Southern hybridization of genomic DNA, Ryder et al. (1987) identified six to eight CHS genes in common bean, some of them tightly clustered, which represented different loci, not allelic variation. The soybean CHS gene family consists of nine members (CHS1 to CHS9), some of which are clustered (Akada and Dube 1995; Yi et al. 2010a). They share a high degree of sequence similarity and play different roles in plant development and interactions with environment. Matsumura et al. (2005) mapped eight CHS genes on five linkage groups (A1, A2, B1, DIa, and K) in soybean. Duplicated CHS1 gene was associated with the suppressed seed coat pigmentation in yellow soybean (Senda et al. 2002).
Gene families arise from interspecific hybridization, polyploidization, and local duplication. Genome duplication results in biased gene content (Freeling 2009) and non-random divergence in gene expression (Casneuf et al. 2006; Wang et al. 2012, 2013). After a duplication event, the new gene copy (or the original copy) can retain the same function (subfunctionalization), undergo neo-functionalization, or become non-functional (loss of function) (Lynch and Conery 2000; Hanada et al. 2011; Barker et al. 2012). Gene clusters formed by gene duplication have been frequently found in multigene families, including plant specialized metabolism (Nutzmann and Osbourn 2014, 2015). For example, clusters encoding enzymes of all steps in lignin biosynthesis have been identified in the Eucalyptus grandis EST libraries (Harakava 2005). The authors also predicted co-localization of several phenylpropanoid pathway enzymes including PAL, C4H, 4CL, C3H, and F5H on the endoplasmic reticulum (ER) membrane. This may suggest the existence of metabolons involving P450 multienzyme complexes and channeling of pathway intermediates without their release into the general metabolic pool (Hrazdina and Wagner 1985; Winkel-Shirley 1999; Ralston and Yu 2006; Bassard et al. 2012).
The availability of complete genome sequences enabled genome-wide analyses of the phenylpropanoid pathway genes in several species (Naoumkina et al. 2010). Shi et al. (2010) identified 95 genes (ten gene families) associated with phenylpropanoid pathway in Populus trichocarpa and identified functional redundancy at the transcript level for six lignin biosynthetic genes [PAL, C4H, 4CL, HCT, CCoAOMT, CAld5H (F5H)]. Using an in silico approach, Costa et al. (2003) analyzed the organization and function of phenylpropanoid pathway gene network in Arabidopsis, while Lucheta et al. (2007) focused on genes encoding key enzymes in the flavonoid pathway in Citrus sinensis. Hamberger et al. (2007) conducted genome-wide analysis of phenylpropanoid pathway gene families in poplar and compared them to homologs in Arabidopsis and rice. The focus of these studies was on the genes of the core pathway and the lignin branch. To explore the evolution of phenylpropanoid pathway diversity, Tohge et al. (2013) compared 65 gene families involved in the pathway among 23 species, including Arabidopsis and soybean. Another evolutionary study was focusing on the isoflavonoid pathway (Chu et al. 2014). The research examined nine major isoflavonoid genes in seven plant species, including Arabidopsis, soybean, and common bean. Genes coding for PAL, C4H, 4CL, CHS, and CHI were identified in all analyzed species, while for CHR, IFS, IOMT (isoflavonoid O-methyltransferase), and IFR (isoflavonoid reductase) were confirmed to be legume-specific. Divergent evolutionary patterns were observed among different gene copies of centrally located branch-point enzymes (4CL, CHS, and CHI) regardless of the level of polymorphism or the evolutionary rate.
However, information about this important pathway in common bean is still fragmentary. In our previous study (Reinprecht et al. 2013), 35 phenylpropanoid pathway genes were cloned and mapped in silico in common bean genome (annotation Phaseolus vulgaris v1.0). The work also identified syntenic regions containing phenylpropanoid pathway genes in common bean and soybean (annotation Glycine max v1.1) (Reinprecht et al. 2013). In another study, 22 phenylpropanoid pathway genes have been mapped in the Bat93 × Jalo EEP558 (a core mapping resource for P. vulgaris) and OAC Rex × SVM Taylor recombinant inbred line (RIL) populations (Yadegari 2013). Currently, work on identifying an association between these genes and different seed phenolics in common bean using an association mapping approach is underway. Cytochrome P450 gene family encodes several key enzymes in the phenylpropanoid pathway. Alber and Ehlting (2012) reviewed P450s involved in lignin biosynthesis. The availability of the complete common bean genome sequence allowed Kumar et al. (2015) to identify members of this gene family. The focus of our work was to study gene families encoding enzymes of phenylpropanoid pathway in common bean, using an in silico approach.
11.2 Gene Families Encoding Enzymes of Phenylpropanoid Pathway in Common Bean
Currently, complete genome sequences for 55 plant species, including common bean (Schmutz et al. 2014; current annotation P. vulgaris v1.0), are deposited in Phytozome 10.3 (a comparative genomic database, available at http://phytozome.jgi.doe.gov/pz/portal.html; accessed 16 Nov 2015; Goodstein et al. 2012). This allowed us to study the complete gene families encoding enzymes of phenylpropanoid pathway in common bean, thus extending our previous work (Reinprecht et al. 2013). In particular, we examined their conservation and diversification through comparative analyses with previously sequenced soybean (Schmutz et al. 2010; current annotation G. max Wm82.a2.v1) and Arabidopsis (The Arabidopsis Genome Initiative 2000; Lamesch et al. 2012; current annotation Arabidopsis thaliana TAIR10) genomes. The basic information for the sequenced Arabidopsis, soybean, and common bean genomes is presented in Table 11.1.
Genome annotations for common bean (Schmutz et al. 2014), soybean (Schmutz et al. 2010), and Arabidopsis (The Arabidopsis Genome Initiative 2000) were obtained from Phytozome 10.2 (Goodstein et al. 2012). For each gene, identifiers and descriptions for all Pfam (Protein families), KEGG (Kyoto Encyclopedia of Genes and Genomes), GO (Gene Ontology), PANTHER (Protein ANalysis THrough Evolutionary Relationships), and KOG (EuKaryotic Orthologous Groups) classifications assigned to this gene can be found.
Table 11.2 contains the list and the number of putative genes in each of the major gene families encoding enzymes of the phenylpropanoid pathway in common bean, soybean, and Arabidopsis in the current annotations of their complete genome sequences (P. vulgaris v1.0, G. max Wm82.a2.v1, and A. thaliana TAIR10) deposited in Phytozome. For each of the 21 enzyme classes, their functional annotations were based on the Pfam and KOG databases (commonly used), while the number of genes in each family was based on Phytozome and KEGG databases. For example, with the KOG0222 search, four phenylalanine ammonia-lyase (PAL, EC:4.3.1.24) genes were identified in Arabidopsis, eight PAL genes were identified in soybean, and six PAL genes were found in common bean (Table 11.2). Several large gene families are involved in phenylpropanoid pathway, including the cytochrome P450 family.
11.3 The Role of Cytochrome P450 Superfamily in Phenylpropanoid Pathway
11.3.1 Cytochrome P450
Cytochromes P450 (CYPs) are ubiquitous monooxygenase enzymes involved in the oxidation of various substrates using oxygen and NADPH. Plant P450s play vital roles in metabolism and detoxification (Mizutani and Ohta 2010; Hamberger and Back 2013). They catalyze reactions in both primary metabolism and secondary metabolism and are involved in the biosynthesis of various metabolites, including fatty acids, sterols, hormones, phenylpropanoids, terpenoids, and signaling molecules. Chemical diversity across plant species is well correlated with the heterogeneity of the P450s (Mizutani and Sato 2011; Mizutani 2012; Sezutsu et al. 2013). They contain a heme cofactor, which absorbs light at 450 nm, and are named for this trait (Pigment absorbing at 450 nm), as well as their cellular localization. Plant P450s are typically membrane-bound to the cytoplasmic surface of the endoplasmic reticulum (ER) by a short N-terminal segment.
The P450s are one of the largest families of enzymes in plants and, in most of plant species, exist as a superfamily. The number of P450 genes is highly variable among plants (Nelson 2006) and represents 0.57–1.07% of the protein coding genes in various plant species [1.07% in Arabidopsis (246/23,000) (Nelson et al. 2004), 0.71% in soybean (332/46,500) (Guttikonda et al. 2010), and 0.78% in common bean (247/31,638) (Kumar et al. 2015)]. The large number of P450s in higher plants is due to gene duplication and diversification (Werck-Reichhart and Feyereisen 2000).
The P450 gene superfamily is characterized by enormous structural and functional diversity (Nelson et al. 2008; Nelson and Werck-Reinchhart 2011; Nagano 2014). Homology and phylogeny were used to group P450s into families (>40% amino acid sequence identity) and subfamilies (>55% amino acid sequence identity) (Nelson et al. 1996). Plant P450 proteins are numbered as CYP51, CYP71 to CYP99, and CYP701 to CYP772. They belong to ten clans (group of genes originated from a single ancestor), which are named by their lowest numbered member [six single-family clans (CYP51, CYP74, CYP97, CP710, CYP711, and CYP727) and four multiple-family clans (CYP71, CYP72, CYP85, and CYP86)] (Werck-Reichhart and Feyereisen 2000; Nelson et al. 2004; Schuler and Werck-Reinchhart 2003; Schuler et al. 2006). Following recommendations of a nomenclature committee (Nelson et al. 1996), the name of P450s consists of a CYP italicized root symbol, followed by a number of the family, a letter of the subfamily and ending by a number of the gene (e.g., CYP71D9—family 71, subfamily D, gene 9), which is determined by the order of identification regardless of the origin.
Initially, P450s were divided into a large A-type clade, which included members that are involved in secondary metabolism (clan CYP71) and several smaller, non-A-type clades, involved in primary metabolism (such as fatty acids and sterols) (Nelson 2006). The occurrence of large numbers of A-type P450s, compared to the non-A-type, suggests a rapid expansion of A-type P450 gene families in plants (Bak et al. 2011).
11.3.2 Clan CYP71—P450s Involved in the Phenylpropanoid Pathway
Based on the current genome annotations [Pfam:00067 (cytochrome P450) functional annotation at Phytozome 10.2; http://phytozome.jgi.doe.gov/pz/portal.html—accessed 26 June 2015], there are 249 P450 genes in A. thaliana TAIR10, 443 P450 genes in G. max Wm82.a.v1, and 264 P450 genes in P. vulgaris v1.0. However, the number of published P450s in these species is slightly different, 272 genes (including 28 pseudogenes) in Arabidopsis (Bak et al. 2011) and 247 genes (including 15 pseudogenes) in common bean (Kumar et al. 2015). P450s in common bean were classified into ten clans that contain 47 families. The largest CYP71 clan (A-type) consists of 19 families with 144 genes. The majority of the genes (>70%) contain a single intron, but more than 20% of the genes have two introns and only a small number of genes (4%) are intronless. In addition, over 80% of the introns are of the zero phase (intron sequence inserted between two successive codons).
It was reported that over 16 P450s are involved in the synthesis and metabolism of phenylpropanoids (Werck-Reichhart 1995). They are placed at the several key positions in the phenylpropanoid pathway, and their roles in phenylpropanoid metabolism were extensively reviewed. For example, Ehlting et al. (2006) and Alber and Ehlting (2012) focused on P450s involved in the core phenylpropanoid pathway and lignin branch, Ayabe and Akashi (2006) in flavonoid metabolism, while Tanaka (2006) and Tanaka and Brugliera (2013) reviewed the role of P450s in flower color.
Seven gene families that encode P450 enzymes involved in phenylpropanoid pathway, as identified in the current genome annotations in common bean, soybean, and Arabidopsis, are listed in Table 11.3. It should be noted, however, that the number of genes in analyzed genomes may change as more work on annotations is done. For example, the CYP71D family in soybean had 81 genes (including 39 pseudogenes) in G. max v1.0 (Nelson 2009) and 52 genes (including 16 pseudogenes) in G. max Wm82.a2.v1. Eleven gene sequences did not correspond between the two genome annotations.
We used the standard nomenclature of chromosome-based locus (gene model) identifiers in plant genome annotations and assemblies (Phytozome), which consists of four segments:
-
species [AT or At (A. thaliana), Glyma. (G. max), Phvul. (P. vulgaris)],
-
chromosome number [1 to 5 (A. thaliana), 01 to 20 (G. max), 001 to 011 (P. vulgaris)],
-
gene (G or g), and
-
five-digit code [A. thaliana—At2g37040 for phenylalanine ammonia-lyase 1 (PAL1)] or six-digit code [G. max (Glyma.03g181700, PAL1) and P. vulgaris (Phvul.001g177800, PAL1)], numbered from top to bottom of chromosome.
These gene families encode enzymes that catalyze various reactions in different branches of the phenylpropanoid pathway (Fig. 11.1), including
-
1.
core phenylpropanoid pathway: cinnamate 4-hydroxylase (C4H, CYP73A),
-
2.
lignin/lignan branch: coumarate 3-hydroxylase (C3H, CYP98A) and ferulic acid 5-hydroxylase (F5H, CYP84A),
-
3.
anthocyanin/condensed tannin branch: flavonoid 3′-hydroxylase (F3′H, CYP75B), flavonoid 3′,5′-hydroxylase (F3′,5′H, CYP75A), and flavone synthase (FNS, CYP93B), and
-
4.
isoflavonoid branch: isoflavone synthase (IFS, CYP93C), isoflavone 2′-hydroxylase (I2′H, CYP81E), flavonoid 6-hydroxylase (F6H, CYP71D), and 3,9-dihydroxypterocarpan 6a-monooxygenase (D6aH, CYP93A).
11.3.3 Gene Structure, Conserved Domains, and Motifs of P450s Involved in the Phenylpropanoid Pathway
Seven P450s families (clan CYP71) that encode enzymes in the phenylpropanoid pathway in common bean, soybean, and Arabidopsis contain 135 members, with one to 36 genes per family (Table 11.3). Most of these genes contain introns. Only one gene is intronless (Phvul.009g244000, CYP81E51). The number of introns ranges from one to four. The majority of the genes contain one (63%) or two introns (32%). The proteins that they encode range in size from 408 amino acids (Phvul.001g139500, CYP93A57) to 543 amino acids (Phvul.002g014800, CYP81E44). The protein sequences were aligned using Clustal Omega at EMBL-EBI (http://www.ebi.ac.uk/Tools/msa/clustalo/), and conserved regions were displayed with a sequence logo generated from the alignment using a Web-based WebLogo 3.4 (Crooks et al. 2004; available at http://weblogo.threeplusone.com/). All of the P450 sequences included the following domains: a heme-binding region (FxxGxRxCxG), a PERF motif (PERF/W), a K-helix region (KETRL) involved in defining the heme pockets and stabilizing the protein structure, and an I-helix region (AGxDT) involved in oxygen binding (Fig. 11.2).
Conserved domains and motif patterns of P450s, CYP71 clan involved in biosynthesis of various phenylpropanoids. P450 domains including a heme-binding region [cysteine (C*) residue is indicated by an asterisk (*)], PERF motif, K-helix and I-helix regions are indicated in red rectangles; the other regions (such as N-terminal region, proline-rich region, membrane anchor, and C-terminal region) are shown in black
11.3.4 Phylogenetic Analysis of P450s Involved in the Phenylpropanoid Pathway
The alignment and tree construction of 135 protein sequences (Table 11.3) from seven P450 gene families (clan CYP71) involved in the phenylpropanoid pathway were performed in MEGA6 (Tamura et al. 2013). These analyses were based on the full-length genes from the three genomes, with one nearly intact soybean C4H pseudogene included (indicated by P at the end of the CYP name—CYP73A88 P). A member from the soybean CYP81E family (CYP81E220de1b, Glyma.16g149200) is truncated (101 amino acids) and was not included in the tree construction.
The phylogenetic tree (Fig. 11.3) separates P450 protein sequences (clan 71) from the two species into seven families:
Protein sequences of the seven gene families from the clan CYP71 involved in the phenylpropanoid pathway in soybean and common bean. A neighbor-joining tree (Poisson model, complete deletion) was built using MEGA6. Soybean sequences are labeled in red, and common bean in blue; P at the end of CYP name indicates pseudogene (Glyma.10G275600-CYP73A88 P); shorter protein sequences are indicated by an asterisk (*); a truncated (101 amino acids) Glyma.16g149200-CYP81E220de1b was excluded from the tree construction
-
CYP71—CYP71D is a legume-specific cluster and contains 36 genes in soybean (and 16 pseudogenes, not included) and 21 genes in common bean (and four pseudogenes, not included). A single flavonoid 6-hydroylase (F6H) in common bean was clustered with three F6H proteins in soybean.
-
CYP73—CYP73A family contains four genes for cinnamic acid 4-hydroxylase (C4H) in soybean (including one pseudogene), three genes in common bean, and a single gene in Arabidopsis. The C4H cluster splits into class I and class II enzymes.
-
CYP75 family is split into two subfamilies. CYP75A consists of two genes for flavonoid 3′,5′-hydroxylase (F3′5′H) (and one pseudogene, not included) in soybean and two genes in common bean. There are no genes for F3′5′H in Arabidopsis. Subfamily CYP75B contains five genes for flavonoid 3′-hydroxylase (F3′H) (and one pseudogene, not included) in soybean, two genes in common bean, and a single gene for F3′H in Arabidopsis.
-
CYP81—CYP81E is a legume-specific cluster and consists of 12 genes coding isoflavone 2′-hydroxylase-like (I2′H) genes (and four pseudogenes, not included) in soybean and 12 genes (and two pseudogenes, not included) in common bean.
-
CYP84—CYP84A cluster contains three genes encoding ferulic acid 5-hydroxylase (F5H) (and one pseudogene, not included) in soybean, three genes in common bean, and two genes in Arabidopsis.
-
CYP93—The family is clustered into three subfamilies. CYP93A is a legume-specific subfamily. It consists of eight genes for 3,9-dihydropterocarpan 6a-monooxygenase (D6aH) (and two pseudogenes, not included) in soybean and seven genes (and one pseudogene, not included) in common bean. The CYP93B subfamily contains two genes encoding flavonoid synthase (FNS) in soybean and a single gene in common bean. There are no FNS genes in Arabidopsis. CYP93C is a legume-specific branch. It consists of two genes for isoflavone synthase (IFS) in soybean and three genes in common bean.
-
CYP98—CYP98A cluster consists of two genes for coumarate 3-hydroxylase (C3H) in soybean and single genes in common bean and Arabidopsis genomes, respectively. There are two additional pollen-specific CYP98As in Arabidopsis (CYP98A8 and CYP98A9; Matsuno et al. 2009—not included in tree construction).
11.3.5 Genome Organization of the Clan CYP71 Gene Families Involved in Phenylpropanoid Pathway in Common Bean
A common bean in silico map that contained genes coding for enzymes of phenylpropanoid pathway, including nine P450s, was developed previously (Reinprecht et al. 2013). The map was created by BLASTing the genomic sequences of the phenylpropanoid pathway genes against the whole common bean genome (P. vulgaris v1.0, Phytozome) using the starting nucleotide positions of the resulting alignments with the chromosome as the map positions for each of the gene sequences.
A similar approach was used to develop a common bean P450-based in silico map, which contains 144 P450, clan CYP71 genes. The mapping was initiated with 134 genes that were identified at Phytozome by searching for KOG0156 functional annotations (cytochrome P450 CYP2 subfamily). Selected gene sequences were BLASTed against the complete common bean genome sequence (Phytozome) to identify their locations. Gene identity was confirmed with the published common bean P450s (Kumar et al. 2015), and ten new sequences (not annotated as KOG0156 in Phytozome) were added to the map. Gene families involved in the phenylpropanoid pathway (shown in larger font, color-coded) were found throughout the common bean genome, except for chromosome Pv05 (Fig. 11.4).
Distribution of cytochrome P450—clan CYP71 genes [locus (gene model) identifiers—Phytozome] in the common bean genome (identified on the right on bars). Genes belonging to families involved in the phenylpropanoid pathway are color-coded; P at the end of CYP name indicates a pseudogene; the orientation along the chromosome is indicated by a forward or reverse arrow. The starting nucleotide position of the resulting alignment with the chromosome was used as the map position for each P450 gene sequence (indicated on the left on bars)
Within the same family, P450s are usually grouped into clusters and the structure of the same P450 family is generally conserved (Nelson et al. 2004; Paquette et al. 2009). In the common bean genome, clustering of genes from the same family was noticed on the chromosomes Pv03 for family CYP93C (all three IFS genes) and Pv09 for family CYP81E (three I2′H genes). Some of the CYP71 genes are tandem arranged with at least four genes from the same subfamily in a row. Many of these clustered genes are found in the same orientation on four chromosomes [Pv01 (four CYP712B, all forward), Pv02 (four CYP71D, all forward), Pv04 (ten CYP82A, all forward; five CYP71AU, all reverse; five CYP736A, all reverse) and Pv06 (eight CYP71D, all reverse; four CYP79D, all forward)] but in a different orientation on three chromosomes [Pv03 (four CYP71D), Pv04 (four CYP81E), and Pv06 (four CYP71D)]. However, members of the large CYP71D subfamily clustered in the same orientation on chromosomes Pv02 (four) and Pv06 (eight) but in a different orientation on the chromosomes Pv03 (four) and Pv06 (four). Therefore, the subfamily distribution may not follow a regular pattern. Due to clustered organization, the 144 CYP71 P450 genes (Kumar et al. 2015) were not evenly distributed in the common bean genome. They ranged from two genes on the chromosome Pv05 to 25 genes on the chromosome Pv04 (Fig. 11.4).
11.4 Cinnamate 4-Hydroxylase (C4H, EC:1.14.13.11, CYP73A)
11.4.1 C4H Catalytic Reaction and Position in the Phenylpropanoid Pathway
Cinnamate 4-hydroxylase (trans-cinnamate 4-monooxygenase, C4H, EC:1.14.13.11, CYP73A) is the first P450 enzyme in the phenylpropanoid pathway. It is an ER membrane-bound P450 and belongs to the family of oxidoreductases that act on paired donors with incorporation of molecular oxygen. The enzyme catalyzes an irreversible (and rate-limiting) region-specific hydroxylation of the aromatic ring of trans-cinnamic acid (only at the 4-position or para position) to produce p-coumaric (hydroxycinnamic) acid (Fig. 11.5), a precursor for many phenylpropanoids including flavonoids, phytoallexins, and monolignols (Hahlbrock and Scheel 1989; Anterola and Lewis 2002; Lu et al. 2006). For activity, C4H requires molecular oxygen and a cytochrome P450 reductase (CPR).
Mizutani et al. (1997) isolated a cDNA and a genomic clone encoding cinnamate 4-hydroxylase from Arabidopsis (CYP73A5) and found its coordinated expression with PAL and 4CL genes. Mutations in this gene affected phenylpropanoid metabolism, growth, and development (Schilmiller et al. 2009). The gene was mapped to the lower arm of chromosome 2 and was highly expressed in all Arabidopsis tissues, especially in roots and lignifying cells (Bell-Lelong et al. 1997). Genes targeted by the same transcription factors tend to show similar expression patterns, which usually suggest relationships among the genes. Down-regulation of genes coding for PAL and C4H was associated with reduced lignin content and altered lignin composition in transgenic tobacco (Sewalt et al. 1997). The position of C4H in the phenylpropanoid pathway protein network is shown in Fig. 11.6a. Highly connected proteins have a stable steady-state distribution of gene expression (Fig. 11.6b).
Functional protein association network in Arabidopsis (action view) visualized on the STRING Web site (http://string-db.org/; accessed: 25 June 2015). a C4H is colored red, and modes of action are shown in different colors. Nodes directly linked to C4H are colored; b Co-expression of C4H with other phenylpropanoid pathway genes in Arabidopsis; locus AT1G15950 is a CCR1 gene
Separation of three common beans, four soybeans, and single Arabidopsis sequences into two groups (Fig. 11.7) confirmed earlier groupings of C4H into class I and class II proteins (Ehlting et al. 2006). This diversification occurred early in the evolution of vascular plants through gene duplication. Common bean and soybean have both classes of C4Hs, while Arabidopsis (Brassicaceae) contains only one gene encoding class I C4H. The alignment of C4H protein sequences (ClustalW2 at EMBL-EBI, available at http://www.ebi.ac.uk/Tools/msa/clustalw2/) revealed high conservation (60–98% identity) among the proteins (85–98% within five C4H class I proteins and 90% between two class II C4H proteins). However, when both monocots and dicots were compared, class I C4H was highly conserved (over 80% protein level), while class II C4Hs were more divergent (less than 70% protein level). This suggests that class I C4Hs “maintained an essential function that does not allow these genes to be lost or even changed much, and it is appealing to assume that this essential function is developmental lignification” (Alber and Ehlting 2012). Class II C4Hs are only present in some plant species, and the class seems to have more specialized functions.
The sequences of eight C4H proteins from common bean, soybean, and Arabidopsis were aligned using Clustal Omega (http://www.ebi.ac.uk/Tools/msa/clustalo/) and BoxShade (http://www.ch.embnet.org/software/BOX_form.html). The sequences were most divergent in their N-terminal membrane anchors. Conserved motifs found in plant P450s (Fig. 11.8, shown in bold) were present in all eight proteins, including proline-rich (PPGP) region, C helix (WrkmR), oxygen binding and activation I-helix (AAIETT), K-helix (EtlR), PERF motif (PeeFrPeRF), and heme-binding region (FgvGrRsCpG) at C-terminus. The only exception is soybean C4H (CYP73A88P) encoded by a pseudogene (Glyma.10g275600). It has truncated N-terminal region, and the generally highly conserved PERF motif has an arginine (R, Arg) to lysine (K, Lys) substitution (Fig. 11.8, highlighted).
Comparison of C4H protein sequences from common bean, soybean, and Arabidopsis. Conserved motifs and sequences are shown in bold. Secondary structures predicted for Arabidopsis C4H gene (At2g30490) are color-coded [shown at the top of sequences alignment, where H (blue) indicates alpha helices, E (red) represents extended (beta) strands, and C (pink) indicates random coils]
Secondary structures of C4H proteins were predicted by programs GOR (Garnier-Osguthorpe-Robson), IV (Garnier et al. 1996; https://npsa-prabi.ibcp.fr/cgi-bin/npsa_automat.pl?page=npsa_gor4.html), and Phyre2 (Protein Homology/analogY Recognition Engine V 2.0) (Kelley et al. 2015; http://www.sbg.bio.ic.ac.uk/phyre2/html/page.cgi?id=index). Transmembrane helices were predicted by program TMHMM-2.0 (TransMembrane prediction using Hidden Markov Models; Krogh et al. 2001; http://www.cbs.dtu.dk/services/TMHMM-2.0/). All proteins have secondary structures similar to the previously published P450s (Graham and Peterson 1999) including alpha helices (blue), beta sheets (red), and random coils (pink) (Fig. 11.9a). They consist of 36–45% alpha helices, 14–18% extended (or beta) strands, and 40–46% random coils. There is a slight difference between the classes of common bean and soybean C4H proteins. Class I C4H proteins contain higher percentages of alpha helices, while class II C4H proteins were predicted to have higher percentages of extended (or beta) strands and random coils. Membrane anchors were predicted for all proteins except for soybean C4H (CYP73A88P) encoded by the pseudogene Glyma.10g026000 (Fig. 11.9b). All C4H proteins are globular proteins as predicted by Phyre2 (Fig. 11.9c). Common bean and soybean C4Hs have tertiary structures similar to the previously identified CYP73A5 in Arabidopsis (At2g30490) and also contain an alpha-domain and a beta-domain (Rupasinghe et al. 2003).
Gene ontology (GO) annotations for C4H proteins (Table 11.4) were predicted using the protein function prediction (PFP), a sequence similarity-based protein function prediction server at Kihara Bioinformatics Laboratory (http://kiharalab.org/; Hawkins et al. 2009). PFP takes into account weakly similar sequences as well as GO term associations observed in known annotations.
11.4.2 CYP73A Gene Family—Structure and Genome Location of C4H Genes
C4Hs are encoded by the relatively small CYP73A gene family. It consists of three genes in common bean {Phvul.006g079700—CYP73A118, Phvul.007g026000—CYP73A15, and Phvul.008g247400—CYP73A [this P450 was incorrectly named as CYP73A2 in common bean (Kumar et al. 2015); however, CYP73A2 was identified in mung bean (Mizutani et al. 1993), Vigna radiata (previously Phaseolus aureus; recently moved from the genus Phaseolus to Vigna)], four genes (including one pseudogene, Glyma.10g275600—CYP73A88 P) in soybean, and a single gene in Arabidopsis (At2g30490; CYP73A5; REF3).
The gene is well conserved in plants, including soybean and common bean. It contains the Pfam domain (PF00067), found as a “duplication-resistant” gene (Paterson et al. 2006). The first C4Hs were identified in Jerusalem artichoke (Helianthus tuberosus—CYP73A1, GenBank accession Z17369; Teutsch et al. 1993) and mung bean (V. radiata—CYP73A2, GenBank accession L07634; Mizutani et al. 1993). Soybean C4H (CYP73A11), a class I C4H enzyme, was identified as an elicitor-induced cytochrome P450, using differential display of mRNA (Schopfer and Ebel 1998). In contrast, common bean C4H (CYP73A15) was identified as a class II C4H enzyme, whose expression was associated with differentiation (Nedelkina et al. 1999).
The genes coding for C4H in common bean, soybean, and Arabidopsis differ in their exon/intron structures. The exons are conserved, while introns are more variable. Genes encoding class II proteins in common bean and soybean consist of two exons separated by an intron of moderate size (354 and 463 bp, respectively). Both exons are split, resulting in four exons, in the two genes encoding class I C4Hs in soybean. These genes are characterized by a long intron 3 (1499 and 1272 bp, respectively). The class I C4H gene in Arabidopsis and the two genes in common bean all have three exons (Fig. 11.10).
11.4.3 Tissue-Specific Expression of Genes Encoding C4Hs
Using publicly available microarray data, Ehlting et al. (2008) created a tool for co-expression analysis of P450s in Arabidopsis. RNA sequencing (RNA-seq) atlases were developed for both soybean (Severin et al. 2010) and common bean (O’Rurke et al. 2014). Based on RNA-seq data (Phytozome 10), genes encoding C4H are differentially expressed in six common bean and soybean tissues (Fig. 11.11). In general, the expression of the genes encoding class I C4H enzymes [Phvul.008g247400 (CYP73A), Glyma.02g236500 (CYP73A11), and Glyma.14g205200 (CYP73A90)] compared to the class II enzymes [Phvul.007g026000 (CYP73A15) and Glyma.20g114200 (CYP73A87)] was higher in all tissues (flowers, pods, leaves, stems, roots, and nodules). Both common bean and soybean have two copies of genes encoding class I C4H enzymes. In both species, one of the genes (Phvul.008g247400 and Glyma.02g236500) is highly expressed in all tissues. The second copy of the genes (Glyma.14g205200 and Phvul.006g079700) is expressed at lower level. In soybean, Glyma.14g205200 had approximately half of the expression of Glyma.02g236500 in stems, roots, and nodules but very low expression in leaves, pods, and flowers. However, Phvul.006g079700 had very low expression in all common bean tissues compared to Phvul.008g247400 (Fig. 11.11).
Expression of common bean and soybean genes encoding cinnamic acid 4-hydroxylase (C4H) in six different tissues. FPKM (Fragments Per Kilobase of transcript per Million fragments mapped) data for expression levels of the genes were calculated from the RNA-seq data deposited at Phytozome 10.2 (available at http://phytozome.jgi.doe.gov/pz/portal.html)
Common bean C4H (CYP73A15) was characterized as a class II C4H enzyme, whose expression was more related to differentiation than the responses to stress (Nedelkina et al. 1999). Antisense and sense expression of cDNA coding for a truncated CYP73A15 gene from French bean led to a reduced and delayed production of lignin in tobacco (Blee et al. 2001). Three C4H genes were identified in the P. trichocarpa genome. Two of them (PtrC4H1 and PtrC4H2) were abundant in differentiating xylem, suggesting their importance in monolignol biosynthesis. Transcripts of PtrC4H3 had little or no expression in all examined tissues (Lu et al. 2006).
11.4.4 Cis-Regulatory Regions in 5′UTRs of C4H Genes
In order to understand the functions of individual members of the C4H multigene families, promoters of the common bean and soybean genes were analyzed and compared to Arabidopsis gene (At2g30490) promoter, which have known functions. Promoter sequences [1 kb of 5′ regulatory sequence upstream of the coding region (1 kb 5′UTR flanking region)] of C4H genes were retrieved from Phytozome (10.2) and aligned in Clustal Omega at EMBL-EBI (http://www.ebi.ac.uk/Tools/msa/clustalo/) to search for possible sequence similarities among these sequences in the two C4H classes. The analysis of the 5′ regulatory regions of C4H genes in Arabidopsis, soybean, and common bean C4H genes revealed a moderate degree of divergence in these regions (39–60% identity). Multiple sequence alignment was sent to ClustalW2_Phylogeny to produce a phylogenetic tree, which was visualized in TreeView. Based on the 5′UTR sequences, eight C4Hs were split into two clusters: a three-gene class I C4Hs (Phvul.008g247400, Glyma.02g236500, and Glyma.14g205200) and a two-gene class II C4Hs (Phvul.007g026000 and Glyma.20g114200) clusters. However, Arabidopsis (class I At2g30490), common bean (class I Phvul.006g079700), and soybean (class II pseudogene Glyma.10g275600) were not clearly included in any class (Fig. 11.12).
Phylogenetic tree of 5′ upstream region (5′UTR) sequences of the class I and class II C4H genes in Arabidopsis, common bean, and soybean. Arabidopsis sequences are labeled in black, soybean in red, and common bean are in blue; P at the end of the CYP name indicates pseudogene. Class II C4Hs are shown in boxes. * identifies the mostly highly expressed genes, and the number of asterisks indicates the relative levels
The 5′UTR sequence of C4H genes was analyzed for potential cis-acting regulatory elements using PlantCARE database (http://bioinformatics.psb.ugent.be/webtools/plantcare/html; Lescot et al. 2002). In total, 69 potential regulatory elements were identified in 5′UTR sequences of eight C4H genes (Fig. 11.13; Table 11.5). Twenty-six (38%) elements were present in four or more genes (Fig. 11.13, color-coded). In addition to the core TATA box and CAAT box (present in all genes), the list included a large number of light-responsive elements (27), as well as elements associated with tissue-specific expression (5), defense and stress responses (6), or hormonal responsiveness (9). A considerable number (14) of predicted regulatory elements were categorized as unknown function (Table 11.5), and two of these (AC II and unnamed_4) were present in all eight C4H genes.
Distribution of the putative cis-regulatory elements in the 5′ upstream regions (5′UTRs) in common bean, soybean, and Arabidopsis C4H genes, identified using PlantCARE database. The elements found in four or more genes are color-coded. Sequences and functions of elements are presented in Table 11.5
A fraction of identified regulatory elements was specific only to class I or class II C4H genes (Fig. 11.13; Table 11.5). Twenty-six elements (37.7%) were present only in class I C4H genes. Four of these elements were identified in all five class I C4H genes. The CGTCA-motif and the TGACG-motif are cis-acting elements involved in the MeJA responsiveness, while the functions of the unnamed_1 and unnamed_3 are unknown. In addition, MBS (a MYB binding site involved in drought inducibility) and O2 site (cis-acting regulatory element involved in regulation of zein metabolism) were identified in the 5′UTRs of all four legume class I C4H genes. Eleven elements (15.9%) were unique to the class II C4H genes. Three of these elements were identified in both soybean (Glyma.20g114200) and common bean (Phvul.007g026000) C4H genes. The MNF1 and CG motifs are light-responsive elements, while the function of the TATCCAT/C-motif is unknown. Lu et al. (2006) reported that four divergent C4H isoforms play distinct roles in P. trichocarpa. The divergent upstream sequences among the two group PtreC4H genes suggested that the mechanisms of gene regulation might be different.
The identification of the cis-acting sequences regulating differential expression of C4H genes and transcription factors that interact with these sequences in common bean, soybean, and Arabidopsis could lead to an understanding of the mechanism(s) of differential regulation of these highly similar genes in these plant species.
11.4.5 Syntenic Regions Containing Common Bean C4H Genes
The availability of the complete genome sequences for numerous plant species, including soybean (Schmutz et al. 2010) and common bean (Schmutz et al. 2014), allows the organization of the individual genomes to be studied, as well as enables comparison of the genomes at the nucleotide level. The size of the common bean genome (521 Mb) is approximately half of the size of the soybean genome (978 Mb). As a result of at least two rounds of polyploidization [~59 MYA (million years ago) and ~13 MYA], the soybean genome contains significant gene duplications and redundancy (Schmutz et al. 2010). In general, for any gene in common bean, two corresponding homologous genes could potentially be found in soybean. Moreover, because of the shared synteny between the two genomes, regions homologous to regions in two soybean chromosomes were found for all 11 common bean chromosomes, with a minor marker rearrangement and/or sequence orientation (Galeano et al. 2009; McClean et al. 2010; Reinprecht et al. 2013).
Synteny analysis was performed in Plant Genome Duplication Database (PGDD, available at http://chibba.agtec.uga.edu/duplication; Lee et al. 2013) against complete genome sequences available for 47 flowering plant species. Numerous syntenic regions (26–44) with other plant species were found for common bean, soybean, and Arabidopsis class I C4Hs. The blocks were of various sizes, ranging from 14 to 884 gene anchors. For example, common bean C4H on the chromosome Pv06, CYP73A118 (Phvul.006g079700), was syntenic to 44 regions in 31 different plant species including two regions in soybean, poplar, pear, watermelon, rice, kale, sacred lotus, and chickpea, three regions in Chinese cabbage, and four regions in kiwifruit (data not shown). In contrast, only five syntenic blocks were identified for common bean and soybean class II C4Hs. They were syntenic to each other and to another three legumes (Medicago truncatula, Cicer arietinum, and Cajanus cajan).
Several syntenic blocks containing C4H loci were identified among common bean, soybean, and Arabidopsis genomes (Table 11.6; Fig. 11.14). For example, Phvul.006g079700 (encoding common bean class I C4H) was syntenic to other four class I C4Hs: common bean Phvul.008g247400, soybean Glyma.02g236500 and Glyma.14g205200, and Arabidopsis At2g30490. Similarly, common bean class II C4H Phvul.007g026000 was syntenic to two soybean class II C4Hs: Glyma20g.114200 and Glyma.10g275600. They were contained in large syntenic blocks anchored by 641 and 561 genes, respectively (Table 11.6; Fig. 11.14, ). Synteny was also analyzed with SyMap v4.0 (Synteny Mapping and Analysis Program; available at http://www.symapdb.org; Soderlund et al. 2011) to produce circular alignments of multiple common bean and soybean chromosomes (Fig. 11.14, right).
Syntenic regions containing C4H loci in genomes of common bean and soybean. a. Class I C4H—Phvul.006g079700 (CYP73A118) and Phvul.008g247400 (CYP73A); b Class II C4H—Phvul.007g026000 (CYP73A15). Left—synteny identified in Plant Genome Duplication Database. Query locus is represented by a red arrow; blue arrows are other anchor genes in the region. Right—circular alignment of common bean and soybean chromosomes containing C4H loci
11.4.6 Sequence Polymorphisms in C4H Genes in Common Bean
Nucleotide polymorphisms for a number of phenylpropanoid pathway genes in various plant species have been described, including Arabidopsis (Savolainen et al. 2000; Aguade 2001; Wright et al. 2003) and maize (Brenner et al. 2010). In the current work, sequences of three C4H genes in the common bean landrace G19833 (Phytozome) were BLASTed against genome sequence of cultivar OAC Rex. The structure of C4H genes identified in OAC Rex was predicted with the HMM-based Fgenesh gene finder (Solovyev et al. 2006; available at http://linux1.softberry.com/berry.phtml?topic=fgenesh&group=programs&subgroup=gfind; accessed: 7 July 2015).
The C4H proteins in the two genotypes were very similar. The proteins encoded by the Phvul.006g079700 gene in G19833 and OAC Rex were identical. A single amino acid difference was identified at position 42 between OAC Rex (I) and G19833 (V) C4H proteins encoded by the Phvul.008g247400 gene (99.8% identity). OAC Rex and G19833 C4H proteins encoded by the Phvul.007g026000 gene were 99.1% identical. Differences were found in five amino acids at positions 4 (V in OAC Rex, F in G19833), 7 (N in OAC Rex, K in G19833), 18 (L in OAC Rex, S in G19833), 54 (K in OAC Rex, N in G19833), and 420 (I in OAC Rex, V in G19833) (data not shown).
The CH4 genomic sequences were also very similar between two common bean genotypes (97.2% identity for Phvul.006g079700, 98.5% identity for Phvul.008g247400, and 98.9% identity for Phvul.007g026000). However, by aligning the CH4 encoding sequences in the two bean genomes (G19833 and OAC Rex), polymorphism (SNPs, insertions, and deletions) was identified for all three C4H genes (Table 11.7; Fig. 11.15).
C4H gene sequence polymorphisms between common bean cultivar OAC Rex (UofG) and landrace G19833 (Phytozome v10.2). a Class I C4H—CYP73A118 (Phvul.006G079700; OAC Rex accession KU308554); in an alignment, E indicates exons (shown in capital letters) and I represents introns (shown in small letters); the sequence polymorphism in intron 2 (I2) is highlighted (shown in gray); b Class I C4H—CYP73A (Phvul.008G247400; OAC Rex accession KU308556); c Class II C4H—CYP73A15 (Phvul.007G026000; OAC Rex accession KU308555)
Although polymorphisms were detected in both the coding (one to 11 SNPs, shown in bold) and non-coding regions, the majority of the sequence differences that were identified occurred in the introns and UTRs. For example, the size difference of the Phvul.006g079700 gene (encoding class I C4H, CYP73A118) intron 2 (143 bp) in OAC Rex (272 bp) and G19833 (415 bp) can be used to develop gene-based marker(s). However, the usefulness of these polymorphisms as C4H gene-specific marker needs to be evaluated in additional germplasm from two common bean gene pools.
11.5 Conclusions
The availability of the whole genome sequences allowed us to identify gene families encoding major enzymes of the phenylpropanoid pathway in common bean, soybean, and Arabidopsis. The work focused on C4H, a cytochrome P450 that occupies an entry position in the pathway. Three genes encoding C4H proteins were identified in common bean genome compared to the four genes in soybean. The next step would be to functionally characterize these genes. The availability of the common bean genome sequence also makes it possible to identify and characterize the members of each gene family that are involved in the specific branches of the phenylpropanoid pathway. Furthermore, the identification of transcription factors that activate phenylpropanoid biosynthetic gene families could provide tools to potentially manipulate the amount of different phenylpropanoids in common bean.
References
Adlercreutz H (2007) Lignans and human health. Crit Rev Clin Lab Sci 44:483–525
Agati G, Azzarello E, Pollastri S, Tattini M (2012) Flavonoids as antioxidants in plants: location and functional significance. Plant Sci 196:67–76
Agati G, Brunetti C, Di Ferdinando M, Ferrini F, Pollastri S, Tattini M (2013) Functional roles of flavonoids in photoprotection: new evidence, lessons from the past. Plant Physiol Biochem 72:35–45
Aguade M (2001) Nucleotide sequence variation at two genes of the phenylpropanoid pathway, the Fah1and F3Hgenes, in Arabidopsis thaliana. Mol Biol Evol 18:1–9
Akada S, Dube SK (1995) Organization of soybean chalcone synthase gene clusters and characterization of a new member of the family. Plant Mol Biol 29:189–199
Akashi T, Aoki T, Ayabe S (2005) Molecular and biochemical characterization of 2-hydroxyisoflavanone dehydratase. Involvement of carboxylesterase-like proteins in leguminous isoflavone biosynthesis. Plant Physiol 137:882–891
Alber A, Ehlting J (2012) Cytochrome P450s in lignin biosynthesis. Adv Bot Res 61:113–143
Anterola AM, Lewis NG (2002) Trends in lignin modification: a comprehensive analysis of the effects of genetic manipulations/mutations on lignification and vascular integrity. Phytochemistry 61:221–294
Artigot M-P, Dayde J, Berger M (2013) Expression of flavonoid 6-hydroxylase candidate genes in normal and mutant soybean genotypes for glycitein content. Mol Biol Rep 40:4361–4369
Austin MB, Noel JP (2003) The chalcone synthase superfamily of type III polyketide synthases. Nat Prod Rep 20:79–110
Ayabe S, Akashi T (2006) Cytochrome P450s in flavonoid metabolism. Phytochem Rev 5:271–282
Bak S, Beisson F, Hamberger B, Hofer R, Paquette S, Werck-Reichhart D (2011) Cytochrome P450. Arabidopsis Book 9:e0144. doi:10.1199/tab.0144
Barakat A, Bagniewska-Zadworna A, Frost CJ, Carlson JE (2010) Phylogeny and expression profiling of CAD and CAD-like genes in hybrid Populus (P. deltoides × P. nigra): evidence from herbivore damage for subfunctionalization and functional divergence. BMC Plant Biol 10:100. doi:10.1186/1471-2229-10-100
Barker MS, Baute GJ, Liu S-L (2012) Duplications and turnover in plant genomes. In: Wendel JF, Greilhuber J, Dolezal J, Leitch IJ (eds) Plant genome diversity, vol 1. Springer-Verlag, Wien, pp 155–169
Bassard J-E, Richert L, Geerinck J, Renault H, Duval F, Ullmann P, Schmitt M, Meyer E, Mutterer J, Boerjan W, De Jaeger G, Mely Y, Goossens A, Werck-Reichhart D (2012) Protein-protein and protein-membrane associations in the lignin pathway. Plant Cell 24:4465–4482
Baxter HL, Stewart N Jr (2013) Effects of altered lignin biosynthesis on phenylpropanoid metabolism and plant stress. Biofuels 4:635–650
Bell-Lelong DA, Cusumano JC, Meyer K, Chapple C (1997) Cinnamate-4-hydroxylase expression in Arabidopsis. Regulation in response to development and the environment. Plant Physiol 113:729–738
Blee K, Choi JW, O’Connell AP, Jupe SC, Schuch W, Lewis NG, Bolwell GP (2001) Antisense and sense expression of cDNA coding for CYP73A15, a class II cinnamate 4-hydroxylase, leads to a delayed and reduced production of lignin in tobacco. Phytochemistry 57:1159–1166
Boerjan W, Ralp J, Baucher M (2003) Lignin biosynthesis. Annu Rev Plant Biol 54:519–546. doi:10.1146/annurev.arplant.54.031902.134938
Brenner EA, Zein I, Chen Y, Andersen JR, Wenzel G, Ouzunova M, Eder J, Darnhofer B, Frei U, Barriere Y, Lubberstedt T (2010) Polymorphisms in O-methyltransferase genes are associated with stover cell wall digestibility in European maize (Zea mays L.). BMC Plant Biol 10:27. doi:10.1186/1471-2229-10-27
Brunetti C, Di Ferdinando M, Fini A, Pollastri S, Tattini M (2013) Flavonoids as antioxidants and developmental regulators: relative significance in plants and humans. Int J Mol Sci 14:3540–3555. doi:10.3390/ijms14023540
Cassidy A, Hanley B, Lamuela-Raventos RM (2000) Isoflavones, lignans and stilbenes—origins, metabolism and potential importance to human health. J Sci Food Agric 80:1044–1062
Casneuf T, De Bodt S, Raes J, Maere S, Van de Peer Y (2006) Nonrandom divergence of gene expression following gene and genome duplications in the flowering plant Arabidopsis thaliana. Genome Biol 7:R13
Chen J, Wang L, Thompson LU (2006) Flaxseed and its components reduce metastasis after surgical excision of solid human breast tumor in nude mice. Cancer Lett 234:168–175
Cheynier V, Comte G, Davies KM, Lattanzio V, Martens S (2013) Plant phenolics: recent advances on their biosynthesis, genetics, and ecophysiology. Plant Physiol Biochem 72:1–20
Chu S, Wang J, Cheng H, Yang Q, Yu D (2014) Evolutionary study of the isoflavonoid pathway based on multiple copies analysis in soybean. BMC Genet 15:76
Costa MA, Collins RE, Anterola AM, Cochrane FC, Davin LB, Lewis NG (2003) An in silico assessment of gene function and organization of the phenylpropanoid pathway metabolic networks in Arabidopsis thaliana and limitations thereof. Phytochemistry 64:1097–1112
Crooks GE, Hon G, Chandonia JM, Brenner SE (2004) WebLogo: a sequence logo generator. Genome Res 14:1188–1190
Dhaubhadel S (2011) Regulation of isoflavonoid biosynthesis in soybean seeds. In: Ng T-B (ed) Soybean—biochemistry, chemistry and physiology. InTech, http://www.intechopen.com
Ehlting J, Hamberger B, Milliom-Rousseau R, Werck-Reichhart D (2006) Cytochrome P450 in phenolic metabolism. Phytochem Rev 5:239–270
Ehlting J, Sauveplane V, Olry A, Ginglinger J-F, Provart NJ, Werck-Reichhart D (2008) An extensive (co-)expression analysis tool for the cytochrome P450 superfamily in Arabidopsis thaliana. BMC Plant Biol 8:47. doi:10.1186/1471-2229-8-47
Falcone Ferreyra ML, Rius SP, Casati P (2012) Flavonoids: biosynthesis, biological functions, and biotechnological applications. Front Plant Sci 3:222. doi:10.3389/fpls.2012.00222
Ferrer JL, Austin MB, Stewart C Jr, Noel JP (2008) Structure and function of enzymes involved in the biosynthesis of phenylpropanoids. Plant Physiol Biochem 46:356–370. doi:10.1016/j.plaphy.2007.12.009
Fraser CM, Chapple C (2011) The phenylpropanoid pathway in Arabidopsis. Arabidopsis Book 9:e0152. doi:10.1199/tab.0152
Freeling M (2009) Bias in plant gene content following different sorts of duplication: tandem, whole-genome, segmental or by transposition. Annu Rev Plant Biol 60:433–453
Galeano CH, Fernandez AC, Gomez M, Blair MW (2009) Single strand conformation polymorphism based SNP and Indel markers for genetic mapping and synteny analysis of common bean (Phaseolus vulgaris L.). BMC Genom 10:629. doi:10.1186/1471-2164-10-629
Garnier J, Gibrat JF, Robson B (1996) GOR method for predicting protein secondary structure from amino acid sequence. Methods Enzymol 266:540–553
Gonzalez A, Zhao M, Leavitt JM, Lloyd AM (2008) Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings. Plant J 53:814–827
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N, Rokhsar DS (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40:D1178–D1186
Gou J-Y, Felippes FF, Liu C-J, Weigel D, Wang J-W (2011) Negative regulation of anthocyanin biosynthesis in Arabidopsis by a miR156-targeted SPL transcription factor. Plant Cell 23:1512–1522
Goujon T, Sibout R, Eudes A, MacKay J, Jouanin L (2003) Genes involved in the biosynthesis of lignin precursors in Arabidopsis thaliana. Plant Physiol Biochem 41:677–687
Graham SE, Peterson JA (1999) How similar are P450s and what can their differences teach us? Arch Biochem Biophys 369:24–29
Graham T, Graham M, Yu O (2008) Genomics of secondary metabolism in soybean. In: Stacey G (ed) Genetics and genomics of soybean. Springer Science+Business Media, LLC, Berlin, pp 211–241
Grotewold E (2005) Plant metabolic diversity: a regulatory perspective. Trends Plant Sci 10:57–62
Guttikonda SK, Trupti J, Bisht NC, Chen H, An Y-QC, Pandey S, Xu D, Yu O (2010) Whole genome co-expression analysis of soybean cytochrome P450 genes identifies nodulation-specific P450 monooxygenases. BMC Plant Biol 12:243
Hahlbrock K, Scheel D (1989) Physiology and molecular biology of phenylpropanoid metabolism. Annu Rev Plant Physiol Plant Mol Biol 40:347–369
Hamberger B, Bak S (2013) Plant P450s as versatile drivers for evolution of species-specific chemical diversity. Philos Trans R Soc B 368:20120426
Hamberger B, Ellis M, Friedmann M, de Azevedo Souza C, Barbazuk B, Douglas CJ (2007) Genome-wide analyses of phenylpropanoid-related genes in Populus trichocarpa, Arabidopsis thaliana, and Oryza sativa: the Populus lignin toolbox and conservation and diversification of angiosperm gene families. Can J Bot 85:1182–1201
Hanada K, Sawada Y, Kuromori T, Klausnitzer R, Saito K, Toyoda T, Shinozaki K, Li W-H, Hirai MY (2011) Functional compensation of primary and secondary metabolites by duplicate genes in Arabidopsis thaliana. Mol Biol Evol 28:377–382
Hao Z, Mohnen D (2014) A review of xylan and lignin biosynthesis: foundation for studying Arabidopsis irregular xylem mutants with pleiotropic phenotypes. Crit Rev Biochem Mol Biol 49:212–241
Harakava R (2005) Genes encoding enzymes of the lignin biosynthesis pathway in Eucalyptus. Genet Mol Biol 28:601–607
Hartmann U, Sagasser M, Mehrtens F, Stracke R, Weisshaar B (2005) Differential combinatorial interactions of cis-acting elements recognized by R2R3-MYB, BZIP, and BHLH factors control light-responsive and tissue-specific activation of phenylpropanoid biosynthesis genes. Plant Mol Biol 57:155–171
Hawkins T, Chitale M, Luban S, Kihara D (2009) PFP: automated prediction of Gene Ontology functional annotations with confidence scores using protein sequence data. Proteins 74:566–582
Hrazdina G, Wagner GJ (1985) Metabolic pathways as enzyme complexes: evidence for the synthesis of phenylpropanoids and flavonoids on membrane associated enzyme complexes. Arch Biochem Biophys 237:88–100
Humphreys JM, Hemm MR, Chapple C (1999) New routes for lignin biosynthesis defined by biochemical characterization of recombinant ferulate 5-hydroxylase, a multifunctional cytochrome P450-dependent monooxygenase. Proc Natl Acad Sci U S A 96:10045–10050
Humphreys JM, Chapple C (2002) Rewriting the lignin roadmap. Curr Opin Plant Biol 5:224–229
Jung W, Yu O, Lau SM, O’Keefe DP, Odell J, Fader G, McGonigle B (2000) Identification and expression of isoflavone synthase, the key enzyme for biosynthesis of isoflavones in legumes. Nat Biotechnol 18:208–212
Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJE (2015) The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 10:45–858. doi:10.1038/nprot.2015.053
Koes R, Verweij W, Quattrocchio F (2005) Flavonoids: a colorful model for the regulation and evolution of biochemical pathways. Trends Plant Sci 10:236–242
Krogh A, Larsson B, von Heijne G, Sonnhammer EL (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305:567–580. doi:10.1006/jmbi.2000.4315
Kumar MS, Chakravarthy SS, Babu PR, Rao KV, Reddy VD (2015) Classification of cytochrome P450s in common bean (Phaseolus vulgaris L.). Plant Syst Evol 301:211–216
Labeeuw L, Martone LT, Boucher Y, Case RJ (2015) Ancient origin of the biosynthesis of lignin precursors. Biol Direct 10:23. doi:10.1186/s13062-015-0052-y
Lamesch P, Berardini TZ, Li D, Swarbreck D, Wilks C, Sasidharan R, Muller R, Dreher K, Alexander DL, Garcia-Hernandez M, Karthikeyan AS, Lee CH, Nelson WD, Ploetz L, Singh S, Wensel A, Huala E (2012) The Arabidopsis Information Resource (TAIR): improved gene annotation and new tools. Nucleic Acids Res 40:D1202–D1210
Latunde-Dada AO, Cabello-Hurtado F, Czittrich N, Didierjean L, Schopfer C, Hertkorni N, Werck-Reichhart D, Ebel J (2001) Flavonoid 6-hydroxylase from soybean (Glycine max L.), a novel plant P-450 monooxygenase. J Biol Chem 276:1688–1695
Lee TH, Tang H, Wang X, Paterson AH (2013) PGDD: a database of gene and genome duplication in plants. Nucleic Acids Res. doi:10.1093/nar/gks1104
Lescot M, Déhais P, Moreau Y, De Moor B, Rouzé P, Rombauts S (2002) PlantCARE: a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327
Li S (2014) Transcriptional control of flavonoid biosynthesis—fine-tuning of the MYB-bHLH-WD40 (MBW) complex. Plant Sig Behav 9:e27522
Lozovaya VV, Lygin AV, Zernova OV, Ulanov AV, Li S, Hartman GL, Widholm JM (2007) Modification of phenolic metabolism in soybean hairy roots through down regulation of chalcone synthase or isoflavone synthase. Planta 225:665–679
Lu S, Zhou Y, Li L, Chiang VL (2006) Distinct roles of cinnamate 4-hydroxylase genes in Populus. Plant Cell Physiol 47:905–914
Lucheta AR, Silva-Pinhati ACO, Basílio-Palmieri AC, Berger IJ, Freitas-Astúa J, Cristofani M (2007) An in silico analysis of the key genes involved in flavonoid biosynthesis in Citrus sinensis. Genet Mol Biol 30:819–831
Lynch M, Conery JS (2000) The evolutionary fate and consequences of duplicate genes. Science 290:151–1155
Matsumura H, Watanabe S, Harada K, Senda M, Akada S, Kawasaki S, Dubouzet EG, Minaka N, Takahashi R (2005) Molecular linkage mapping and phylogeny of the chalcone synthase multigene family in soybean. Theor Appl Genet 110:1203–1209
Matsuno M, Compagnon V, Schoch GA, Schmitt M, Debayle D, Bassard J-E, Pollet B, Hehn A, Heintz D, Ullmann P, Lapierre C, Bernier F, Ehlting J, Werck-Reichhart D (2009) Evolution of a novel phenolic pathway for pollen development. Science 324:1688–1692
McClean PE, Mamidi S, McConnell M, Chikara S, Lee R (2010) Synteny mapping between common bean and soybean reveals extensive blocks of shared loci. BMC Genom 11:184. doi:10.1186/1471-2164-11-184
Miedes E, Vanholme R, Boerjan W, Molina A (2014) The role of the secondary cell wall in plant resistance to pathogens. Front Plant Sci 5:358. doi:10.3389/fpls.2014.00358
Mizutani M (2012) Impacts of diversification of cytochrome P450 on plant metabolism. Biol Pharm Bull 35:824–832
Mizutani M, Ohta D (2010) Diversification of P450 genes during land plant evolution. Annu Rev Plant Biol 61:291–315
Mizutani M, Ohta D, Sato R (1997) Isolation of a cDNA and a genomic clone encoding cinnamate 4-hydroxylase from Arabidopsis and its expression manner in plants. Plant Physiol 113:755–763
Mizutani M, Sato F (2011) Unusual P450 reactions in plant secondary metabolism. Arch Biochem Biophys 507:94–203
Mizutani M, Ward E, DiMaio J, Ohta D Ryals J, Sato R (1993) Molecular cloning and sequencing of a cDNA encoding mung bean cytochrome P450 (P450C4H) possessing cinnamate 4-hydroxylase activity. Biochem Biophys Res Commun 190:875–880
Moura JC, Bonine CA, de Oliveira Fernandes Viana J, Dornelas MC, Mazzafera P (2010) Abiotic and biotic stresses and changes in the lignin content and composition in plants. J Integr Plant Biol 52:360–376. doi:10.1111/j.1744-7909.2010.00892.x
Nagano S (2014) Structural and functional diversity of cytochrome P450. In: Yamazaki H (ed) Fifty years of cytochrome P450 research. Springer, Japan, pp 95–106. doi:10.1007/978-4-431-54992-5_5
Naoumkina MA, Zhao Q, Gallego-Giraldo L, Dai X, Zhao PX, Dixon RA (2010) Genome-wide analysis of phenylpropanoid defence pathways. Mol Plant Pathol 11:829–846
Nedelkina S, Jupe SC, Blee KA, Schalk M, Werck-Reichhart D, Bolwell GP (1999) Novel characteristics and regulation of a divergent cinnamate 4-hydroxylase (CYP73A15) from French bean: engineering expression in yeast. Plant Mol Biol 39:1079–1090
Nelson DR (2006) Cytochrome P450 nomenclature, 2004. Method Mol Biol 320:1–10
Nelson DR (2009) The cytochrome P450 homepage. Hum Genomics 4:59–65
Nelson DR, Koymans L, Kamataki T, Stegeman JJ, Feyereisen R, Waxman DJ, Waterman MR, Gotoh O, Coon MJ, Estabrook RW, Gunsalus IC, Nebert DW (1996) P450 superfamily: update on new sequences, gene mapping, accession numbers and nomenclature. Pharmacogenetics 6:1–42
Nelson DR, Ming R, Alam M, Schuler MA (2008) Comparison of cytochrome P450 genes from six plant genomes. Trop Plant Biol 1:216–235
Nelson DR, Schuler MA, Paquette SM, Werck-Reichhart D, Bak S (2004) Comparative genomics of rice and Arabidopsis. Analysis of 727 cytochrome P450 genes and pseudogenes from a monocot and a dicot. Plant Physiol 135:756–772
Nelson DR, Werck-Reichhart D (2011) A P450-centric view of plant evolution. Plant J 66:194–211
Noel JP, Austin MB, Bomati EK (2005) Structure-function relationships in plant phenylpropanoid biosynthesis. Curr Opin Plant Biol 8:249–253
Nutzmann H-W, Osbourn A (2014) Gene clustering in plant specialized metabolism. Curr Opin Biotechnol 26:91–99
Nutzmann H-W, Osbourn A (2015) Regulation of metabolic gene clusters in Arabidopsis thaliana. New Phytol 205:503–510. doi:10.1111/nph.13189
O’Rourke JA, Iniguez LP, Fu F, Bucciarelli B, Miller SS, Jackson SA, McClean PE, Li J, Dai X, Zhao PX, Hernandez G, Vance CP (2014) An RNA-Seq based gene expression atlas of the common bean. BMC Genom 15:866
Osakabe K, Tsao CC, Li L, Popko JL, Umezawa T, Carraway DT, Smeltzer RH, Joshi CP, Chiang VL (1999) Coniferyl aldehyde 5-hydroxylation and methylation direct syringyl lignin biosynthesis in angiosperms. Proc Natl Acad Sci U S A 96:8955–8960
Paquette SM, Jensen K, Bak S (2009) A web-based resource for the Arabidopsis P450, cytochromes b5, NADPH-cytochrome P450 reductases, and family 1 glycosyltransferases (http://www.P450.kvl.dk). Phytochemistry 70:1940–1947
Parvathi K, Chen F, Guo D, Blount JW, Dixon RA (2001) Substrate preferences of O-methyltransferases in alfalfa suggest new pathways for 3-O-methylation of monolignols. Plant J 25:193–202
Paterson AH, Chapman BA, Kissinger JC, Bowers JE, Feltus FA, Estill JC (2006) Many genes and domain families have convergent fates following independent whole-genome duplication events in Arabidopsis, Oryza, Saccharomyces and Tetraodon. Trends Genet 22:597–602
Petrussa E, Braidot E, Zancani M, Peresson C, Bertolini A, Patui S, Vianello A (2013) Plant flavonoids-Biosynthesis, transport and involvement in stress responses. Int J Mol Sci 14:14950–14973. doi:10.3390/ijms140714950
Raes J, Rohde A, Christensen JH, Van de Peer Y, Boerjan W (2003) Genome-wide characterization of the lignification toolbox in Arabidopsis. Plant Physiol 133:1051–1071
Ralston L, Subramanian S, Matsuno M, Yu O (2005) Partial reconstruction of flavonoid and isoflavonoid biosynthesis in yeast using soybean type I and type II chalcone isomerases. Plant Physiol 137:1375–1388
Ralston L, Yu O (2006) Metabolons involving plant cytochrome P450s. Phytochem Rev 5:459–472
Ramsay NA, Glover BJ (2005) MYB-bHLH-WD40 protein complex and the evolution of cellular diversity. Trends Plant Sci 10:63–70
Reinprecht Y, Yadegari Z, Perry GE, Siddiqua M, Wright LC, McClean PE, Pauls KP (2013) In silico comparison of genomic regions containing genes coding for enzymes and transcription factors for the phenylpropanoid pathway in Phaseolus vulgaris L. and Glycine max L. Merr. Front Plant Sci 4:317. doi:10.3389/fpls.2013.00317
Rupasinghe S, Baudry J, Mary A, Schuler MA (2003) Common active site architecture and binding strategy of four phenylpropanoid P450s from Arabidopsis thaliana as revealed by molecular modeling. Protein Eng 16:721–731. doi:10.1093/protein/gzg094
Ryder TB, Hedrick SA, Bell JN, Liang XW, Clouse SD, Lamb CJ (1987) Organization and differential activation of a gene family encoding the plant defense enzyme chalcone synthase in Phaseolus vulgaris. Mol Gen Genet 210:219–233
Saito K, Yonekura-Sakakibara K, Nakabayashi R, Higashi Y, Yamazaki M, Tohge T, Fernie AR (2013) The flavonoid biosynthetic pathway in Arabidopsis: structural and genetic diversity. Plant Physiol Biochem 72:21–34
Savolainen O, Langley CH, Lazzaro BP, Freville H (2000) Contrasting patterns of nucleotide polymorphism at the alcohol dehydrogenase locus in the outcrossing Arabidopsis lyrata and the selfing Arabidopsis thaliana. Mol Biol Evol 17:645–655
Schilmiller AL, Stout J, Weng J-K, Humphreys J, Ruegger MO, Chapple C (2009) Mutations in the cinnamate 4-hydroxylase gene impact metabolism, growth and development in Arabidopsis. Plant J 60:771–782
Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T, Nelson W, Hyten DL, Song Q, Thelen JJ, Cheng J, Xu D, Hellsten U, May GD, Yu Y, Sakurai T, Umezawa T, Bhattacharyya MK, Sandhu D, Valliyodan B, Lindquist E, Peto M, Grant D, Shu S, Goodstein D, Barry K, Futrell-Griggs M, Abernathy B, Du J, Tian Z, Zhu L, Gill N, Joshi T, Libault M, Sethuraman A, Zhang XC, Shinozaki K, Nguyen HT, Wing RA, Cregan P, Specht J, Grimwood J, Rokhsar D, Stacey G, Shoemaker RC, Jackson SA (2010) Genome sequence of the palaeopolyploid soybean. Nature 463:178–183
Schmutz J, McClean PE, Mamidi S, Wu GA, Cannon SB, Grimwood J, Jenkins J, Shu S, Song Q, Chavarro C, Torres-Torres M, Geffroy V, Moghaddam SM, Gao D, Abernathy B, Barry K, Blair M, Brick MA, Chovatia M, Gepts P, Goodstein DM, Gonzales M, Hellsten U, Hyten DL, Jia G, Kelly JD, Kudrna D, Lee R, Richard MM, Miklas PN, Osorno JM, Rodrigues J, Thareau V, Urrea CA, Wang M, Yu Y, Zhang M, Wing RA, Cregan PB, Rokhsar DS, Jackson SA (2014) A reference genome for common bean and genome-wide analysis of dual domestications. Nat Genet 46:707–713
Schoch G, Goepfert S, Morant M, Hehn A, Meyer D, Ullmann P, Werck-Reichhart D (2001) CYP98A3 from Arabidopsis thaliana is a 3′-hydroxylase of phenolic esters, a missing link in the phenylpropanoid pathway. J Biol Chem 276:36566–36574
Schopfer CR, Ebel J (1998) Identification of elicitor-induced cytochrome P450s of soybean (Glycine max L.) using differential display of mRNA. Mol Gen Genet 258:315–322
Schuler MA, Werck-Reichhart D (2003) Functional genomics of P450s. Annu Rev Plant Biol 54:629–667
Schuler MA, Duan H, Bilgin M, Ali S (2006) Arabidopsis cytochrome P450s through the looking glass: a window on plant biochemistry. Phytochem Rev 5:205–237
Senda M, Jumonji A, Yumoto S, Ishikawa R, Harada T, Niizeki M, Akada S (2002) Analysis of the duplicated CHS1 gene related to the suppression of the seed coat pigmentation in yellow soybeans. Theor Appl Genet 104:1086–1091
Severin AJ, Woody JL, Bolon Y-T, Joseph B, Diers BW, Farmer AD, Muehlbauer GJ, Nelson RT, Grant D, Specht JE, Graham MA, Cannon SB, May GD, Vance CP, Shoemaker RC (2010) RNA-Seq atlas of Glycine max: a guide to the soybean transcriptome. BMC Plant Biol 10:160
Sewalt VJH, Ni W, Blount JW, Jung HG, Masoud SA, Howles PA, Lamb C, Dixon RA (1997) Reduced lignin content and altered lignin composition in transgenic tobacco down-regulated in expression of l-phenylalanine ammonia-lyase or cinnamate 4-hydroxylase. Plant Physiol 115:41–50
Sezutsu H, Le Goff G, Feyereisen R (2013) Origins of P450 diversity. Philos Trans R Soc B 368:20120428
Shelton D, Stranne M, Mikkelsen L, Pakseresht N, Welham T, Hiraka H, Tabata S, Sato S, Paquette S, Wang TL, Martin C, Bailey P (2012) Transcription factors of lotus: regulation of isoflavonoid biosynthesis requires coordinated changes in transcription factor activity. Plant Physiol 159:531–547
Shi R, Sun Y-H, Li Q, Heber S, Seferoff R, Chiang VL (2010) Towards a system approach for lignin biosynthesis in Populus trichocarpa: transcript abundance and specificity of the monolignol biosynthetic genes. Plant Cell Physiol 51:144–163
Shimamura M, Akashi T, Sakurai N, Suzuki H, Saito K, Shibata D, Ayabe S, Aoki T (2007) 2-hydroxyisoflavanone dehydratase is a critical determinant of isoflavone productivity in hairy root cultures of Lotus japonicus. Plant Cell Physiol 48:1652–1657
Soderlund C, Bomhoff M, Nelson WM (2011) SyMAP v3.4: a turnkey synteny system with application to plant genomes. Nucleic Acids Res 39:e68. doi:10.1093/nar/gkr123
Solovyev V, Kosarev P, Seledsov I, Vorobyev D (2006) Automatic annotation of eukaryotic genes, pseudogenes and promoters. Genome Biol 7:S10. doi:10.1186/gb-2006-7-s1-s10
Steele CL, Gijzen M, Qutob D, Dixon RA (1999) Molecular characterization of the enzyme catalyzing the aryl migration reaction of isoflavonoid biosynthesis in soybean. Arch Biochem Biophys 367:146–150
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetic analysis version 6.0. Mol Biol Evol 30:2725–2829
Tanaka Y (2006) Flower colour and cytochromes P450. Phytochem Rev 5:283–291. doi:10.1007/s11101-006-9003-7
Tanaka Y, Brugliera F (2013) Flower colour and cytochromes P450. Philos Trans R Soc Lond B Biol Sci 368:20120432. doi:10.1098/rstb.2012.0432
Teutsch HG, Hasenfratz M, Lesot A, Stoltz C, Garnier J-M, Jeltsch J-M, Durst F, Werck-Reichhart D (1993) Isolation and sequence of a cDNA encoding the Jerusalem artichoke cinnamate 4-hydroxylase, a major plant cytochrome P450 involved in the general phenylpropanoid pathway. Proc Natl Acad Sci U S A 90:4102–4106
The Arabidopsis Genome Initiative (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408:796–815
Tohge T, Watanabe M, Hoefgen R, Fernie AR (2013) The evolution of phenylpropanoid pathway in the green lineage. Crit Rev Biochem Mol Biol 48:123–152
Tohge T, Yonekura-Sakakibara K, Niida R, Watanabe-Takahashi A, Saito K (2007) Phytochemical genomics in Arabidopsis thaliana: a case study for functional identification of flavonoid biosynthesis genes. Pure Appl Chem 79:811–823
Tsai C-J, Harding SA, Tschaplinski TJ, Lindroth RL, Yuan Y (2006) Genome-wide analysis of the structural genes regulating defense phenylpropanoid metabolism in Populus. New Phytol 172:47–62. doi:10.1111/j.1469-8137.2006.01798.x
Vanholme R, Demedts B, Morreel K, Ralph J, Boerjan W (2010) Lignin biosynthesis and structure. Plant Physiol 153:895–905
Vogt T (2010) Phenylpropanoid biosynthesis. Mol Plant 3:2–20
Wang X (2011) Structure, function, and engineering of enzymes in isoflavonoid biosynthesis. Funct Integr Genomics 11:13–22
Wang Y, Tan X, Paterson AH (2013) Different patterns of gene structure divergence following gene duplication in Arabidopsis. BMC Genom 14:652
Wang Y, Wang X, Paterson AH (2012) Genome and gene duplications and gene expression divergence: a view from plats. Ann N Y Acad Sci 1256:1–14
Weng J-K, Chapple C (2010) The origin and evolution of lignin biosynthesis. New Phytol 187:273–285
Werck-Reichhart D (1995) Cytochromes P450 in phenylpropanoid metabolism. Drug Metabol Drug Interact 12:221–243
Werck-Reichhart D, Feyereisen R (2000) Cytochrome P450: a success story. Genome Biol 1:reviews3003.1–reviews3003.9
Winkel-Shirley B (1999) Evidence for enzyme complexes in the phenylpropanoid and flavonoid pathways. Physiol Plant 107:142–149
Winkel-Shirley B (2001) Flavonoid biosynthesis. A colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol 126:485–493
Wright SI, Lauga B, Charlesworth D (2003) Subdivision and haplotype structure in natural populations of Arabidopsis lyrata. Mol Ecol 12:1247–1263
Xiao CW (2008) Health effects of soy protein and isoflavones in humans. J Nutr 138:1244S–1249S
Xu Z, Zhang D, Hu J, Zhou X, Ye X, Reichel KL, Stewart NR, Syrenne RD, Yang X, Gao P, Shi W, Doeppke C, Sykes RW, Burris JN, Bozell JJ, Cheng Z-M, Hayes DG, Labbe N, Davis M, Stewart CN, Yuan JS (2009) Comparative genome analysis of lignin biosynthesis gene families across the plant kingdom. BMC Bioinform 10:S3
Xu W, Dubos C, Lepiniec L (2015) Transcriptional control of flavonoid biosynthesis by MYB-bHLH-WDR complexes. Trends Plant Sci 20:176–185
Xu W, Grain D, Bobet S, Le Gourrierec J, Thevenin J, Kelemen Z, Lepiniec L, Dubos C (2014) Complexity and robustness of the flavonoid transcriptional regulatory network revealed by comprehensive analyses of MYB-bHLH-WDR complexes and their targets in Arabidopsis seed. New Phytol 202:132–144. doi:10.1111/nph.12620
Yadegari Z (2013) Molecular mapping and characterization of phenylpropanoid pathway genes in common bean (Phaseolus vulgaris L.). Ph.D. thesis, University of Guelph, Guelph, Canada
Yi J, Derynck MR, Chen L, Dhaubhadel S (2010a) Differential expression of CHS7 and CHS8 genes in soybean. Planta 231:741–753
Yi J, Derynck MR, Li X, Telmer P, Marsolais F, Dhaubhadel S (2010b) A single repeat MYB transcription factor, GmMYB176, regulates CHS8 gene expression and affects isoflavonoid biosynthesis in soybean. Plant J 62:1019–1034
Yoon J, Choi H, An G (2015) Roles of lignin biosynthesis and regulatory genes in plant development. J Integr Plant Biol 57:902–912
Zhao Q, Dixon RA (2011) Transcriptional networks for lignin biosynthesis: more complex than we thought? Trends Plant Sci 16:227–233
Zhong R, Ye Z-H (2009) Transcriptional regulation of lignin biosynthesis. Plant Signal Behav 4:1028–1034
Zubieta C, Parvathi K, Ferrer J-L, Dixon RA, Noel JP (2002) Structural basis for the modulation of lignin monomer methylation by caffeic acid/5-hydroxyferulic acid3/5-O-methyltransferase. Plant Cell 14:1265–1277
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Reinprecht, Y., Perry, G.E., Peter Pauls, K. (2017). A Comparison of Phenylpropanoid Pathway Gene Families in Common Bean. Focus on P450 and C4H Genes. In: Pérez de la Vega, M., Santalla, M., Marsolais, F. (eds) The Common Bean Genome. Compendium of Plant Genomes. Springer, Cham. https://doi.org/10.1007/978-3-319-63526-2_11
Download citation
DOI: https://doi.org/10.1007/978-3-319-63526-2_11
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-63524-8
Online ISBN: 978-3-319-63526-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)