Abstract
The challenge of fungal measurements in indoor environments is complex. Almost all studies that have used several methods for the assessment of fungal exposure have only observed moderate or weak correlation between them. These variations can be explained by the fungal life cycle with differences in spore release and the variation in the characteristics of spores of different species, and with differences in the target molecules used by the various fungal exposure assessment methods. Therefore, the use of different analysis methods will provide a different perspective on the stages of fungal growth and quantity.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Fungi
- indoor
- biomass markers
- glucan
- ergosterol
- quantitative PCR
- NAHA
- microscopy
- cultivation
- real-time monitoring
1 Introduction
Since we spend more than 90% of our time in indoor environments and breathe about 10 m3 of air every day (Dacarro et al., 2003) our proximity and interaction with indoor microbes, including fungi, is considerable. It has been established that indoor air quality is one of the most important factors that influence our general quality of life. Indoor air pollution can result in health problems and even in an increase in human mortality (Kanchongkittiphon et al., 2015, Mendell et al., 2011, Heseltine and Rosen, 2009). With respect to fungal contamination of indoor environments, the issue of moisture damage and dampness problems in buildings and the associated microbial proliferation and health problems observed in building occupants is the central issue.
The interaction between humans and indoor microbes is complex, stemming from the fact that the microbes’ amounts, activity, physiology and diversity depend on both human activities and operational conditions of the indoor environment. To deal with the complexity of the diverse effects of mold on health, various different measurement strategies have been developed to assess indoor fungal contamination. Depending on the purpose of the investigation, indoor samples, including air, dust and surface materials, are analyzed to detect either fungal particles or specific fungal compounds. Fungal spores and/or fungal fragments may be detected with the aim of assessing fungal exposure as such, while cell components and metabolites of molds known to be associated with adverse health effects may be specifically quantified to assess health risks.
2 Analytical Methods for Measuring Fungal Concentrations
Measurements of fungal concentrations in indoor air and on materials are important in determining the sources and nature of fungal contamination which in turn helps in estimating the risks associated with exposures to fungal particles in moisture damaged buildings. Fungi grow differently on different building materials and surfaces and are affected by the diverse and changing conditions in the indoor environment. Methods that accurately estimate their amounts and also provide qualitative information on the nature of the fungi present are of interest.
The quantitative measurements assess how many fungal cells or how much fungal biomass are present on material surfaces or in the air, being determined by techniques such as the culture-based methods, microscopic counts and molecular methods, the latter targeting chemical cell wall markers of fungal biomass or fungal DNA. These methods determine the extent of growth, sporulation and total number of cells, or biomass (Krause et al., 2003). Total fungal biomass has been used as a surrogate of the overall fungal exposure and can be determined using chemical markers that are found in the fungi. Such chemical markers include ergosterol (Szponar et al., 2003), N-acetylhexosaminidase (NAHA), (Reeslev et al., 2003, Rylander et al., 2010), 1 → 3-β- glucan (Foto et al., 2005) and extracellular polysaccharides (EPS) (Douwes et al., 1999; Noterman and Soentoro, 1986). Each one of the methods is thought to provide a different perspective of fungal quantities since they evaluate specific responses of the various stages of fungal growth.
2.1 Cultivation Method for Determining Fungal Growth
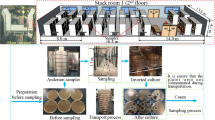
Traditional quantification of fungi is based on the determination of the number and type of colony forming units (CFU), providing quantitative and qualitative data on viable and culturable fungi from different types of samples. This method is useful in identifying fungi down to the genus level using microscopy of individual colonies (Burge et al., 1999). It may help in providing useful information to confirm an environmental source for an outbreak investigation. Counting culturable microorganisms not only allows for a quantitative, but also qualitative assessment of exposure by identifying the genus of fungi since not all fungi pose the same hazard. After cultivation, fungi may be identified to the genus or species level using morphological criteria and microscopy, Matrix Assisted Laser Desorption/Ionization-Time of Flight Mass Spectrometry (MALDI-TOF-MS) (De Carolis et al., 2012) or DNA based methodologies.
Cultivation method has its inherent limitations, such as the inability of particular media to satisfy the specific growth requirements of certain fungal species (Douwes et al., 2003). The choice of growth media used can contribute to substantial variability of the types and amounts of species that are cultured. For example, malt extract agar (MEA) has high sugar content and water activity allowing fast growing fungal species to flourish on its surface. Dichloran glycerol 18 on the other hand allows detection of a more diverse fungal flora, however excluding fungi which require high water activity (Chao et al., 2002, Wu et al., 2000). Cultivation method lacks the ability to detect non-culturable and dead microorganisms, cell debris and microbial components, although all of those may be of health relevance (Green et al., 2005). It has been demonstrated that by using the cultivation method, concentrations of viable fungal cells detected in relation to the total amount of fungal cells present vary widely, depending on the type of sample (<1–100%) (Lee et al., 2006, Meklin et al., 2004, Toivola et al., 2002). The total quantity and diversity of fungal cells are usually drastically underestimated by culture-based methods (Bridge and Spooner, 2001, Douwes et al., 2003). In addition to these limitations, the cultivation method is laborious and requires a long incubation time (minimum of 1 week) to detect fungal growth (Douwes et al., 2003). The method shows poor precision and is variable among replicate samples (Eduard and Halstensen, 2009, Mensah-Attipoe et al., 2016a). Since culture-based methods measure only a fraction of sampled fungi, CFU counted cannot completely characterize the fungal spectrum that might influence human health and well-being. For all of these reasons, other, culture independent approaches have been developed to identify and enumerate fungi in indoor samples (Viegas et al., 2012, Eduard and Halstensen, 2009).
2.2 Microscopic Spore Counting
To circumvent some of the limitations associated with culture-based techniques, methods that detect both viable and non-viable cells using microscopic counting have been developed (Palmgren, 1986, Bauer et al., 2008, Ho et al., 2005, Sattler et al., 2001). Simple light microscopy may be used to count microorganisms, but counting is based only on morphological recognition, which may result in severe measurement errors (Douwes et al., 2003). For example, identification of the fungal spores is often difficult: only a small number of fungal spore types can be identified with confidence at genus level, and important genera, such as Aspergillus and Penicillium, cannot be differentiated. In epifluorescence microscopy, dyes such as acridine orange which stain the spores’ DNA, are used to help in counting spores. The dye makes the spore fluoresce when viewed under the microscope (Thorne et al., 1994, Palmgren, 1986). The epifluorescence microscopic spore count method has its own limitations which include the masking of spores by large particles and the inability of some spores to absorb the dye (Burge, 1995). In addition, the spore counting approach is time consuming, laborious and not trivial to perform. The method is, however, relatively cheap to perform and may provide a general indication of atypical indoor fungal growth (Douwes et al., 2003).
Electron microscopy (EM) or scanning EM can also be used and it provides a better determination of spore counts and concentrations (Eduard et al., 1988, Karlsson and Malmberg, 1989). Like the simple light microscopy, identification and enumeration is done by morphology. There is an increased interest in identifying and differentiating particles from different sources but having similar appearances which often lead to difficulties in their quantification (Wittmaack et al., 2005). A good technique developed to circumvent this problem is the use of SEM microscopy coupled with energy dispersive X-ray spectroscopy (EDX). This analysis is based on determining the elemental composition of the particles after they have been identified with SEM. The elemental composition of biological particles differs from other particles and they also behave differently from non-biological particles. Based on this property, criteria for determining primary biogenic organic aerosols (PBOA) in atmospheric samples have been developed (Matthias-Maser and Jaenicke, 1991, 1994). These were based on the detection of minor amounts of K, P, S, Na and Ca (usually <10% of relative element of X-ray intensity of the particle). This criterion was recently adopted (Coz et al., 2010) to characterize PBOA in the atmosphere. Also a recent study (Mensah-Attipoe et al., 2016b) applied this method to differentiate fragments of biological origin from those of non-biological origin.
2.3 DNA-Based Methods
The last decades have seen a surge in the development of several culture-independent, molecular, DNA based techniques supplying many advantages over the traditional cultivation technique (Amann et al., 1995). The DNA-based method, like the total spore count method, detects both culturable and unculturable spores. DNA-based methods in addition, measure mycelial cells (Meklin et al., 2004, Gonzalez and Saiz-Jimenez, 2004, Herrera et al., 2009, Yamamoto et al., 2010). Those methods require an initial step of extracting DNA from an environmental sample prior to subsequent analysis. The molecular methods most often used in fungal studies include conventional or quantitative PCR (qPCR) specific for fungal species or groups (Haugland et al., 2004, Zeng et al., 2006), as well as ribosomal DNA amplicon sequencing or metagenome analysis (Tringe et al., 2008, Frohlich-Nowoisky et al., 2009, Liu et al., 2012, Adams et al., 2013a, Adams et al., 2013b, Dannemiller et al., 2014b, Yamamoto et al., 2014, Dannemiller et al., 2014a, Pitkaranta et al., 2008). In the pre-next generation sequencing (NGS) era, DNA fingerprinting methods were based on universal fungal PCR combined with denaturing or temperature gradient gel electrophoresis (DGGE, TGGE) (Gonzalez and Saiz-Jimenez, 2004), and terminal or conventional restriction fragment length polymorphism analysis (Buttner et al., 2007). Other methods include the molecular tracer methods (Elbert et al., 2007)
The advantages of using the DNA-based methods for detecting and quantifying fungi instead of cultivation-based methods and microscopic counts are the speed, accuracy, and analytical sensitivity of this approach and the possibility to detect and identify also dead or dormant microorganisms (Ettenauer et al., 2014). Detection is based on DNA, and therefore, not dependent on the viability of the microbe. The ability of DNA-based methods to detect dead or dormant cells is important in indoor studies since the main exposure hazards relating to indoor microbial contamination may not require viability. Thus, these techniques have enabled a reliable assessment of fungal communities associated with different materials such as wood, concrete, mineral wool, paper, or dust (Ettenauer et al., 2012, Piñar and Sterflinger, 2009). Quantitative PCR has provided valuable information on the occurrence and levels of the most common indoor fungi, and exhibits great potential at being able to provide quickly quantitative data on the occurrence of the studied organisms (Meklin et al., 2004, Pietarinen et al., 2008, Pitkäranta et al., 2011). Primers and probes are generally designed for the detection of a given genus (genus-specific primers), groups of genera (group-specific), or for the detection of a single species (species-specific primers). The 18S ribosomal RNA gene and internal transcribed spacer regions (ITS) can be used in the design of specific primers and probes because they contain sequences that are highly conserved between members of the same species or genus, for example, but are variable among different species or genera (Haugland et al., 2004).
There are a studies describing the exploitation of qPCR analytical methods in estimating the concentrations of individual species or groups of fungi in indoor dust and air samples (Vesper, 2007, Haugland et al., 2004, Meklin et al., 2004, Kaarakainen et al., 2009) and building materials (Pitkäranta et al., 2011, Pietarinen et al., 2008). For example, real-time PCR methods have been utilized to detect and quantify Cladosporium (Zeng et al., 2006) and Aspergillus (Goebes et al., 2007) at the genus level. Similar methods have been developed for targeting species, groups of species or genera of common indoor fungi such as Aspergillus, Cladosporium, Penicillium and Alternaria (Vesper et al., 2005, Meklin et al., 2007, Haugland et al., 2004). Applying multiple PCR assays make it possible to assess the presence and amounts of large groups of microorganisms. For instance, a quantitative PCR approach has been developed integrating measurement of 36 fungal species commonly associated with damp houses and background species, used to define an “environmental relative mouldiness index” (ERMI) for houses in the United States (Vesper, 2007). It is also important to mention in this context that PCR and qPCR are targeted approaches that detect only what the primer set is designed for, unlike cultivation or next generation sequencing methods, which are largely untargeted methods.
There are well known limitations inherent to DNA based approaches referring to biases in DNA extraction and PCR amplification that can be preferential of one over another species or taxon (see more detail below). Quantitation in qPCR can also have biases due to variation in target gene copy numbers in different fungal species and genera (Herrera et al., 2009). However, results may not be affected by these bias when primers and probes are used that target the ITS gene region and keep the copy numbers fairly stable across varying conditions (Herrera et al., 2009). In addition, when fungal spore suspensions of known concentrations are used in creating standard curves for individual species rather than genera or groups, the issue of target gene copy numbers is less pronounced.
Next generation sequencing (NGS) approaches today are widely used in fungal ecological studies in different environments. NGS refers to a suite of different methods carried out on different sequencing platforms, and these methods include: sequencing of 16S rRNA and internal transcribed spacer region (ITS) amplicons for studies of the bacterial and fungal communities, respectively (amplicon sequencing); whole-genome sequencing for understanding an organism’s function; and metagenomics sequencing to understand the functioning of microbial communities (Cox et al., 2013). More recently, NGS – and here thus far almost exclusively amplicon sequencing of the fungal ITS regions – has been introduced to studies of fungal communities in indoor environments. Given constant improvements in sample throughput, resolution, costs per analysis and bioinformatics that tackle the large data amounts typically produced in NGS surveys (Metzker, 2010), the number of studies using NGS in indoor assessments is steadily increasing. Early efforts utilizing DNA extracted from house dust samples revealed a hitherto unknown richness and diversity of the indoor fungal flora (Amend et al., 2010, Pitkaranta et al., 2008). Further indoor NGS studies have dealt with the ecology and sources of fungi determined from indoor samples and also attempted to study health implications of fungal exposure (Adams et al., 2013b, 2013c, 2015, Dannemiller et al., 2014a, 2016a, 2016b, Lymperopoulou et al., 2016). Amplicon sequencing has obvious advantages over other assessment methods: it is a non-targeted method that permits detection of theoretically all fungal taxa present in a sample, independent of viability and culturability; the method is characterized by high resolution that allows determination of hundreds of different fungal taxa from indoor samples; and, in case of ITS sequencing, taxonomic allocation of the fungal sequences detected upon database comparisons is often possible to the species or at least the genus level. Investigations that have interest in knowing which fungal cells in a sample are metabolically active and alive versus dead, may need to use other approaches than amplicon sequencing, as this method can at current not efficiently distinguish what is dead and alive in a sample. Another issue is absolute quantification, desirable in e.g. determining human exposure levels. NGS data are typically presented as relative abundance of a given fungal taxon rather than total amount of the respective fungal cells in a sample. There are further technical limitations in amplicon sequencing that refer generally to DNA and PCR based approaches and that need to be carefully considered. Selectivity in DNA extraction as well as in the PCR amplification prior sequencing, leading to preferential detection of some taxa over others, and variation in target sequence copy number can introduce biases into NGS studies (Amend et al., 2010, Huber et al., 2009, Rastogi et al., 2009, von Wintzingerode et al., 1997). Nonetheless, next generation sequencing approaches have already revolutionized our view and interpretation of the role of bacterial and fungal communities in human health and disease, and similar advances can be expected from studies applying these approaches in indoor microbial assessments.
2.4 Methods Measuring Cell Wall Components and Other Indicators of Biomass
Microbial cell wall components, including fungal compounds, are discussed in Chap. 8 “Endotoxins, Glucans and Other Microbial Cell Wall Agents” and the reader is referred to this part of the book for more detailed information. There is a variety of approaches that assay cellular constituents, usually microbial cell wall agents, instead of counting culturable and/or non-culturable microbial particles. These methods typically are used to measure surrogates of total fungal biomass; however, in some cases the targeted compound itself may be of interest due to potential health implications upon exposure. More or less commonly used markers for the assessment of fungal biomass include ergosterol, (Miller et al., 1988, Szponar et al., 2003), fungal extracellular polysaccharides (EPS) and β-glucan (Sonesson et al., 1988, Douwes et al., 2000). Measuring the activity of N-acetylhexosaminidase enzyme (Rylander et al., 2010, Reeslev et al., 2003) is another way to estimate total fungal biomass. Microbial volatile organic compounds (MVOCs) produced by fungi have been proposed as markers of active fungal growth (Dillon et al., 2007, Moularat et al., 2008). Also measuring mycotoxins, i.e. toxic fungal secondary metabolites, from indoor samples is being done, mostly in a search for exposing agents involved in provoking adverse health effects in occupants of moisture damaged building. These latter two categories – MVOCs and mycotoxins – will not be discussed further here, as separate chapters in this book have been dedicated to these agents.
There are several advantages common to these assay methods. These include: the stability of most of the measured components, allowing – among others – longer sampling times for airborne measurements; and not restricting the assessment to only “alive” fungal material. Storing samples frozen prior to analyses is typically not an issue, unlike for example in cultivation based approaches. Standards can be used in many of the methods allowing sound quantification, and typically, sensitivity and specificity of the measurement approaches is high, even though there often is a tradeoff between those two (i.e. high sensitivity comes to the expense of lower specificity and vice versa). Major limitations are, however, that these methods do not allow for an identification of the fungal taxa present in a sample, but rather provide information on total fungal material present. Furthermore, these methods do not leave fungal isolates for further investigation where needed, a disadvantage common to all methods but cultivation.
2.5 Ergosterol
Ergosterol is the major sterol present within cell membranes of fungal spores and hyphae and is considered an indicator for total fungal biomass (Szponar et al., 2003). This agent has been assayed in indoor dust (Saraf et al., 1997), in building materials (Szponar et al., 2003, Gutarowska and Piotrowska, 2007), and indoor air (Park and Cox-Ganser, 2011). As just mentioned as a common limitation when measuring cell wall agents, also quantifying ergosterol does not provide information about the individual fungal species present in a sample. Ergosterol contents are measured by gas chromatography-mass spectrometry (Miller et al., 1988, Szponar et al., 2003). The amount of ergosterol measured from a particular fungal isolate depends on its surface area and growth conditions. It has been shown that ergosterol is somewhat labile and thus its concentration declines after the death of fungal spores and hyphae (Mille-Lindblom et al., 2004, Gutarowska and Piotrowska, 2007), though it is not well understood how long ergosterol stays stable within dead fungal cells or in different sample materials. Some studies have shown that levels of ergosterol correlate well with total spore counts (Mensah-Attipoe et al., 2016a). Although this method is highly specific and sensitive giving accurate estimate of fungal amounts, ergosterol determination in samples is not done routinely, as it is a rather expensive methodology requiring specific and costly infrastructure and expertise and time to process and analyze samples.
2.6 Fungal Extracellular Polysaccharides
These are stable carbohydrates that are produced and excreted during fungal growth and are suggested as a marker for fungal exposure. Measurements are usually done with immunoassays and as the polysaccharides confer antigenic specificity and differ somewhat between major fungal taxa it is in theory possible to target fungi at the genus level. For example, Douwes et al. (2003) quantified EPS from Aspergillus and Penicillium fungi (EPS-Asp/Pen) in house dust. The authors of that paper found a correlation between the EPS measurements and culturable fungal spore counts. EPS-Asp/Pen in house dust serves as a marker of a somewhat specific fungal exposure, but EPS as such is not suspected to be causally related with poor respiratory or other ill health in children or adults. The method is not widely used today, as the required immunoassays have been developed for research purposes only and are not commercially available.
2.7 Beta-Glucan
Beta-glucans are polysaccharides found in the outer cell membrane of fungi, higher plants and some bacteria. In the fungal cell wall, glucans comprise a three-dimensional network of (1 → 3) and (1 → 6)-β D-linked anhydroglucose repeat units that are connected to other carbohydrates, proteins and lipids. (1 → 3)-β-D-glucan polymers can exist as a stable complex of three polymer strands forming a triple helix. The triple helical structure is generally considered to be the preferred form in nature (Young and Castranova, 2005). From a quantitative point of view, (1 → 3)-β-D-glucans are the main constituent, accounting for between 47% and 60% by weight of the cell wall (Young and Castranova, 2005). Measurements of (1 → 3)-β-D-glucan in indoor air have been done with a glucan-sensitive preparation of the Limulus amebocyte lysate (LAL) assay (Iossifova et al., 2009, 2007). While the LAL test is highly sensitive, it’s specificity for fungi is somewhat limited, as it reacts to some extent also to plant or bacterial material. Other methods for the analysis of fungal glucans have been based on antibodies. For example, Douwes et al. (1996, 1998) developed an inhibition enzyme immunoassay (EIA), which is specific to the (1 → 3)-glycosidic linkage and to water-insoluble glucans, but is less sensitive than the LAL test. Glucan immunoassays that are more specific to fungal material by targeting (1 → 3)-β glucan have been developed (Low et al., 2009). Generally, glucan content in fungal cells has been shown to vary according to the fungal species and dependent on the spore surface area (Iossifova et al., 2008), but appears to be relatively independent of growth conditions (Foto et al., 2004). Assessment of exposure to glucan is done both in homes and in occupational context in work places. For the latter, several epidemiological studies have reported glucans to have strong immunomodulating and inflammatory effects in occupational, high exposure settings. Douwes (2005) asserted from his review that the biological effects observed are not dependent on viability and that (1 → 3)-β-D-glucans from dead organisms may thus be equally relevant in causing potential health effects.
There have been mixed observation with exposures to (1 → 3)-β-glucan and the health effects, pointing also towards a beneficial role of glucan exposure. Some studies have suggested beneficial impacts on the development of immune system of infants (Iossifova et al., 2007, Schaub et al., 2006), lower prevalence of allergic sensitization in 2–4 year olds when exposed to (1 → 3)-β-glucan from mattress dust (Gehring et al., 2007) and inverse association with wheezing symptoms in children (Iossifova et al., 2007, 2009). Furthermore, (Tischer et al., 2011) found a difference in (1 → 3)-β-glucan effects between countries.
2.8 NAHA Enzyme Activity
Measurement of the activity of β-N-acetylhexosaminidase (NAHA) in fungi is another way of measuring fungal total biomass (Rylander et al., 2010, Reeslev et al., 2003). NAHA is present in both the growth and stationary phases of fungal growth (Rast et al., 2003, Reeslev et al., 2003) and its activity is reported to be relatively stable under appropriate storage conditions (Rylander, 2015). The activity measurement is based on a fluorescence labeled substrate which is cleaved by the enzyme that is only present in fungi. The amount of fluorescence detected is proportional to the amount of enzyme/biomass present, and this measurement approach has been translated into a commercially available product (Reeslev et al., 2003). By using enzyme activity as an indicator for fungal biomass, fungal growth present on a building material surface or the amount of fungal biomass in air can be determined. Since the method is fast and can be done onsite, it allows determining the extent of mould-affected materials and the efficacy of cleaning after remediation efforts in the field. Beta-N-acetylhexosaminidase activity has been shown to correlate well with the fungal molecules such as ergosterol and the phospholipid fatty acid 18:2ω6 in soil samples (Miller et al., 1988) and building materials (Mensah-Attipoe et al., 2016a). Significant correlations have been reported between NAHA and total spore counts in dust (Madsen, 2003, 2009) and building materials (Mensah-Attipoe et al., 2016a); fungal biomass (by gravimetric weight) of fungal species grown on nutrient agar; and ergosterol content of gypsum boards (Reeslev et al., 2003) and mineral wool contaminated by fungi (Mensah-Attipoe et al., 2016a). In a recent study (Mensah-Attipoe et al., 2015), the authors found a good correlation between NAHA enzyme activity and cultivation method.
The NAHA method is not able to differentiate between fungal species. Furthermore, levels of NAHA enzymes detected in air samples are usually very low and do not represent well the fungal amounts measured. To be able to achieve a high enough concentration in air, movement and agitation during sampling (usually in the form of “aggressive” blowing, i.e., resuspension of material) is required.
3 Real-Time Detection of Airborne Fungal Particles
It has been stated that a thorough understanding of the significance of microbial exposure in indoor environments is impaired by the methodological difficulties in identifying and enumerating various microbial components (Green et al., 2006). Traditional bioaerosol detection methods such as the Andersen impactor and filter sampling require a separate step for analysis after sampling and before concentration can be determined, which results in relatively low time resolution (Reponen et al., 2011, Górny et al., 2002). These methods are well-established for culture-based and microscopic analyses, and more recently also for DNA-based analyses in the case of filter sampling. Sample collection usually only lasts for a short period of time and thus samples collected by these methods reflect the concentration of the target organism only at the specific time of sampling (Reponen et al., 2007), while the great temporal as well as spatial variation in airborne fungal concentrations are well known. Detection of target bioaerosols in real time would allow understanding of emissions and the temporal variation of airborne concentrations. Therefore, real time detection techniques are needed in various fields, e.g., bioprocess monitoring (Ganzlin et al., 2007), health related applications (Elston, 2001), and in environmental, defense and public health (Davitt et al., 2005, Kanaani et al., 2007, Sivaprakasam et al., 2004, Méjean et al., 2004).
Direct reading instruments that measure particles in real-time have been used in several laboratory studies, e.g., optical particle counters (OPCs), which measure particle concentration in the size range of 0.3–20 µm based on their light scattering properties. The electrical low pressure impactor (ELPI), a multistage impactor, classifies aerosol samples into size fractions over a size range of 0.07–10 µm and the Aerodynamic Particle Sizer (APS) measures the dynamic size distribution of particles in the size range of 0.5–20 µm by determining the time-of-flight of individual particles in an accelerating flow field (Volckens and Peters, 2005). Particle concentrations in the size range of 0.02–1 µm can be measured using the P-Trak. This instrument measures the number concentration of particles by saturating them with either water or alcohol vapour and cooling them so that their enlarged particle sizes can be detected by optical methods. Although very useful in laboratory-based studies where other particles can be eliminated, especially particles within a certain size range, the above described instruments have limited utility in the assessment of bioaerosol exposures because they are not very specific since the process of distinguishing microbial and non-microbial particulate matter is complex (Green et al., 2011).
The Laser Induced Fluorescence (LIF) technique enables real-time detection of biological aerosol particles. LIF techniques have been developed to detect biological warfare agents (Hairston et al., 1997). The use of LIF techniques can give insight into the origin of the fluorescent spectral features and contribute to the interpretation of data obtained using other fluorescence-based techniques.
The best known and most widely used LIF device is the Ultra Violet aerodynamic particle sizer (UVAPS). Other devices include the BioScout and Wide Issue Bioaerosol Sensor (WIBS-3) and the waveband integrated bioaerosol sensor (WIBS-4). The basis by which all of these devices detect relies on the fluorescence of compounds such as reduced nicotinamide adenine dinucleotide phosphate (NAD(P)H), flavins, melanin, carotenoids, phenols, terpenoids, and DNA (Raimondi et al., 2009, O’Connor et al., 2011, Pöhlker et al., 2012, Saari et al., 2013, Frohlich-Nowoisky et al., 2009, Després et al., 2007), present in all living cells but at different proportions and measured at selective wavelengths. Thus, LIF enables the differentiation of bioaerosols from other particles because of their fluorescence capabilities (Hill et al., 2013, Pöhlker et al., 2012).
The UVAPS measures both aerodynamic particle size and autofluorescence of a single particle. The WIBS, on the other hand, measures optical size and the autofluorescence of bioaerosol particles (Gabey et al., 2010, Healy et al., 2014) by utilizing two excitation wavelengths and by detecting two bands of fluorescence. The BioScout measures both autofluorescence and optical particle size of single particles using continuous wave laser diode and light scattering. Saari et al. (2014) have used both BioScout and UVAPS to measure fungal spores in the laboratory and found the former device to be more sensitive. This indicates that the LIF devices have varying capabilities in detecting biological particles based on the wavelengths of light used and the type and amount of fluorescent compounds present in the samples. Data obtained from the use of these devices have usually been on fungal spores and not on small-sized fragments. This is because very little or no fluorescence is emitted from fungal fragments compared to larger spores (Kanaani et al., 2008, Saari et al., 2014).
Different fungal species have characteristic structures and more or less differing biochemical compositions which in turn could influence their autofluorescence. A variety of factors may affect the fluorescent properties of fungi under various conditions. These factors include the type of the fungal species under consideration, growth substrate, air velocity and age of the culture. A better understanding of the effects of these factors on the fluorescent properties of fungal spores measured with different LIF devices side-by-side would help with instrument calibration and ease the interpretation of LIF-based field results.
4 Conclusion
Depending on the purpose of a measurement, an analysis approach serving that purpose need to be chosen. For example, cultivation method accounts only for viable spores and cells that can grow on the culture media. DNA based methods and NGS give in-depth information of the fungi being assessed while ergosterol content, NAHA enzyme activity and other cell wall components display the total biomass of the fungi and indirectly estimate total exposure. Since the life cycle of the fungi is dynamic, the different methods employed will give insight into the different stages of the fungal growth.
References
Adams RI, Amend AS, Taylor JW et al (2013a) A unique signal distorts the perception of species richness and composition in high-throughput sequencing surveys of microbial communities: a case study of fungi in indoor dust. Microb Ecol 66(4):735–741
Adams RI, Bhangar S, Pasut W et al (2015) Chamber bioaerosol study: outdoor air and human occupants as sources of indoor airborne microbes. PLoS One 10(5):e0128022
Adams RI, Miletto M, Taylor JW et al (2013b) Dispersal in microbes: fungi in indoor air are dominated by outdoor air and show dispersal limitation at short distances. ISME J 7(7):1262–1273
Adams RI, Miletto M, Taylor JW et al (2013c) The diversity and distribution of fungi on residential surfaces. PLoS One 8(11):e78866
Amann RI, Ludwig W, Schleifer K (1995) Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev 59(1):143–169
Amend AS, Seifert KA, Bruns TD (2010) Quantifying microbial communities with 454 pyrosequencing: does read abundance count? Mol Ecol 19(24):5555–5565
Bauer H, Schueller E, Weinke G et al (2008) Significant contributions of fungal spores to the organic carbon and to the aerosol mass balance of the urban atmospheric aerosol. Atmos Environ 42(22):5542–5549
Bridge P, Spooner B (2001) Soil fungi: diversity and detection. Plant Soil 232(1-2):147–154
Burge H, Otten J, Fungi JM et al (1999) Bioaerosols: assessment and control. In: Anonymous American Conference of Governmental Industrial Hygienists (ACGIH), vol 19., p 1–13
Burge HA (1995) Bioaerosols. CRC Press
Buttner MP, Cruz P, Stetzenbach LD et al (2007) Evaluation of two surface sampling methods for detection of Erwinia herbicola on a variety of materials by culture and quantitative PCR. Appl Environ Microbiol 73(11):3505–3510
Chao HJ, Milton DK, Schwartz J et al (2002) Dustborne fungi in large office buildings. Mycopathologia 154(2):93–106
Cox MJ, Cookson WO, Moffatt MF (2013) Sequencing the human microbiome in health and disease. Hum Mol Genet 22(R1):R88–94
Coz E, Artíñano B, Clark LM et al (2010) Characterization of fine primary biogenic organic aerosol in an urban area in the northeastern United States. Atmos Environ 44(32):3952–3962
Dacarro C, Picco A, Grisoli P et al (2003) Determination of aerial microbiological contamination in scholastic sports environments. J Appl Microbiol 95(5):904–912
Dannemiller KC, Gent JF, Leaderer BP et al (2016a) Indoor microbial communities: influence on asthma severity in atopic and nonatopic children. J Allergy Clin Immunol 138(1):76–83-e1
Dannemiller KC, Mendell MJ, Macher JM et al (2014a) Next-generation DNA sequencing reveals that low fungal diversity in house dust is associated with childhood asthma development. Indoor Air 24(3):236–247
Dannemiller KC, Reeves D, Bibby K et al (2014b) Fungal High-throughput Taxonomic Identification tool for use with Next-Generation Sequencing (FHiTINGS). J Basic Microbiol 54(4):315–321
Dannemiller KC, Gent JF, Leaderer BP et al (2016b) Influence of housing characteristics on bacterial and fungal communities in homes of asthmatic children. Indoor Air 26(2):179–192
Davitt K, Song Y, Patterson III W et al (2005) 290 and 340 nm UV LED arrays for fluorescence detection from single airborne particles. Opt Express 13(23):9548–9555
De Carolis E, Posteraro B, Lass-Flörl C et al (2012) Species identification of Aspergillus, Fusarium and Mucorales with direct surface analysis by matrix-assisted laser desorption ionization time-of-flight mass spectrometry. Clin Microbiol Infect 18(5):475–484
Després V, Nowoisky J, Klose M et al (2007) Characterization of primary biogenic aerosol particles in urban, rural, and high-alpine air by DNA sequence and restriction fragment analysis of ribosomal RNA genes. Biogeosciences 4(6):1127–1141
Dillon HK, Boling DK, Miller JD (2007) Comparison of detection methods for Aspergillus fumigatus in environmental air samples in an occupational environment. J Occup Environ Hyg 4(7):509–513
Douwes J, van der Sluis B, Doekes G et al (1999) Fungal extracellular polysaccharides in house dust as a marker for exposure to fungi: relations with culturable fungi, reported home dampness, and respiratory symptoms. J Allergy Clin Immunol 103:494–500
Douwes J (2005) (1→3)-β-D-glucans and respiratory health: a review of the scientific evidence. Indoor Air 15(3):160–169
Douwes J, Doekes G, Heinrich J et al (1998) Endotoxin and β (1 → 3)-Glucan in House Dust and the Relation with Home Characteristics: A Pilot Study in 25 German Houses. Indoor Air 8(4):255–263
Douwes J, Zuidhof A, Doekes G et al (2000) (1 → 3)-β-D-glucan and endotoxin in house dust and peak flow variability in children. Am J Resp Crit Care Med 162(4):1348–1354
Douwes J, Doekes G, Montijn R et al (1996) Measurement of beta (1→3)-glucans in occupational and home environments with an inhibition enzyme immunoassay. Appl Environ Microbiol 62(9):3176–3182
Douwes J, Thorne P, Pearce N et al (2003) Bioaerosol health effects and exposure assessment: progress and prospects. Ann Occup Hyg 47(3):187–200
Eduard W, Sandven P, Johansen BV et al (1988) Identification and quantification of mould spores by scanning electron microscopy (SEM): analysis of filter samples collected in Norwegian saw mills. Ann Occup Hyg 32(inhaled particles VI):447–455
Eduard W, Halstensen AS (2009) Quantitative exposure assessment of organic dust. Scand J Work Environ Health. Supplement (7):30.
Elbert W, Taylor P, Andreae M et al (2007) Contribution of fungi to primary biogenic aerosols in the atmosphere: wet and dry discharged spores, carbohydrates, and inorganic ions. Atmos Chem Phys 7(17):4569–4588
Elston DM (2001) Fluorescence of fungi in superficial and deep fungal infections. BMC Microbiol 1:21
Ettenauer J, Piñar G, Tafer H et al (2014) Quantification of fungal abundance on cultural heritage using real time PCR targeting the β-actin gene. Front Microbiol 5.
Ettenauer JD, Pinar G, Lopandic K et al (2012) Microbes on building materials – evaluation of DNA extraction protocols as common basis for molecular analysis. Sci Total Environ 439:44–53
Foto M, Vrijmoed L, Miller J et al (2005) A comparison of airborne ergosterol, glucan and Air-O-Cell data in relation to physical assessments of mold damage and some other parameters. Indoor Air 15(4):257–266
Foto M, Plett J, Berghout J et al (2004) Modification of the Limulus amebocyte lysate assay for the analysis of glucan in indoor environments. Anal Bioanal Chem 379(1):156–162
Frohlich-Nowoisky J, Pickersgill DA, Despres VR et al (2009) High diversity of fungi in air particulate matter. Proc Natl Acad Sci U S A 106(31):12814–12819
Gabey A, Gallagher M, Whitehead J et al (2010) Measurements and comparison of primary biological aerosol above and below a tropical forest canopy using a dual channel fluorescence spectrometer. Atmos Chem Phys 10(10):4453–4466
Ganzlin M, Marose S, Lu X et al (2007) In situ multi-wavelength fluorescence spectroscopy as effective tool to simultaneously monitor spore germination, metabolic activity and quantitative protein production in recombinant Aspergillus niger fed-batch cultures. J Biotechnol 132(4):461–468
Gehring U, Heinrich J, Hoek G et al (2007) Bacteria and mould components in house dust and children’s allergic sensitisation. Eur Respir J 29(6):1144–1153
Goebes MD, Hildemann LM, Kujundzic E et al (2007) Real-time PCR for detection of the Aspergillus genus. J Environ Monitor 9(6):599–609
Gonzalez JM, Saiz-Jimenez C (2004) Microbial diversity in biodeteriorated monuments as studied by denaturing gradient gel electrophoresis. J Separ Sci 27(3):174–180
Górny RL, Reponen T, Willeke K et al (2002) Fungal fragments as indoor air biocontaminants. Appl Environ Microbiol 68(7):3522–3531
Green BJ, Millecchia LL, Blachere FM et al (2006) Dual fluorescent halogen immunoassay for bioaerosols using confocal microscopy. Anal Biochem 354(1):151–153
Green BJ, Schmechel D, Summerbell RC (2011) Aerosolized fungal fragments. In: Anonymous fundamentals of mold growth in indoor environments and strategies for healthy living. Springer, p. 211–243
Green BJ, Sercombe JK, Tovey ER (2005) Fungal fragments and undocumented conidia function as new aeroallergen sources. J Allergy Clin Immunol 115(5):1043–1048
Gutarowska B, Piotrowska M (2007) Methods of mycological analysis in buildings. Build Environ 42(4):1843–1850
Hairston PP, Ho J, Quant FR (1997) Design of an instrument for real-time detection of bioaerosols using simultaneous measurement of particle aerodynamic size and intrinsic fluorescence. J Aerosol Sci 28(3):471–482
Haugland RA, Varma M, Wymer LJ et al (2004) Quantitative PCR analysis of selected Aspergillus, Penicillium and Paecilomyces Species. Syst Appl Microbiol 27(2):198–210
Healy D, Huffman J, O’Connor D et al (2014) Ambient measurements of biological aerosol particles near Killarney, Ireland: a comparison between real-time fluorescence and microscopy techniques. Atmos Chem Phys 14(15):8055–8069
Herrera ML, Vallor AC, Gelfond JA et al (2009) Strain-dependent variation in 18S ribosomal DNA Copy numbers in Aspergillus fumigatus. J Clin Microbiol 47(5):1325–1332
Heseltine E, Rosen J (2009) WHO guidelines for indoor air quality: dampness and mould. WHO Regional Office Europe.
Hill SC, Pan Y, Williamson C et al (2013) Fluorescence of bioaerosols: mathematical model including primary fluorescing and absorbing molecules in bacteria. Opt Express 21(19):22285–22313
Ho H, Rao CY, Hsu H et al (2005) Characteristics and determinants of ambient fungal spores in Hualien, Taiwan. Atmos Environ 39(32):5839–5850
Huber JA, Morrison HG, Huse SM et al (2009) Effect of PCR amplicon size on assessments of clone library microbial diversity and community structure. Environ Microbiol 11(5):1292–1302
Iossifova Y, Reponen T, Daines M et al (2008) Comparison of two analytical methods for detecting (1-3)-β-D-glucan in pure fungal cultures and in home dust samples. Open Allergy J 1:26–34
Iossifova YY, Reponen T, Ryan PH et al (2009) Mold exposure during infancy as a predictor of potential asthma development. Ann Allergy Asthma Immunol 102(2):131–137
Iossifova Y, Reponen T, Bernstein D et al (2007) House dust (1–3)-β-d-glucan and wheezing in infants. Allergy 62(5):504–513
Kaarakainen P, Rintala H, Vepsäläinen A et al (2009) Microbial content of house dust samples determined with qPCR. Sci Total Environ 407(16):4673–4680
Kanaani H, Hargreaves M, Ristovski Z et al (2007) Performance assessment of UVAPS: Influence of fungal spore age and air exposure. J Aerosol Sci 38(1):83–96
Kanaani H, Hargreaves M, Smith J et al (2008) Performance of UVAPS with respect to detection of airborne fungi. J Aerosol Sci 39(2):175–189
Kanchongkittiphon W, Mendell MJ, Gaffin JM et al (2015) Indoor environmental exposures and exacerbation of asthma: an update to the 2000 review by the Institute of Medicine. Environ Health Perspect 123(1):6–20
Karlsson K, Malmberg P (1989) Characterization of exposure to molds and actinomycetes in agricultural dusts by scanning electron microscopy, fluorescence microscopy and the culture method. Scand J Work Environ Health 353–359
Krause JD, Hammad YY, Ball LB (2003) Application of a fluorometric method for the detection of mold in indoor environments. Appl Occup Environ Hyg 18(7):499–503
Lee T, Grinshpun SA, Martuzevicius D et al (2006) Culturability and concentration of indoor and outdoor airborne fungi in six single-family homes. Atmos Environ 40(16):2902–2910
Liu CM, Kachur S, Dwan MG et al (2012) FungiQuant: a broad-coverage fungal quantitative real-time PCR assay. BMC Microbiol 12:255
Low SY, Hill JE, Peccia J (2009) DNA aptamers bind specifically and selectively to (1 → 3)-β-d-glucans. Biochem Biophys Res Commun 378(4):701–705
Lymperopoulou DS, Adams RI, Lindow SE (2016) Contribution of vegetation to the microbial composition of nearby outdoor air. Appl Environ Microbiol 82(13):3822–3833
Madsen A (2003) NAGase activity in airborne biomass dust and relationship between NAGase concentrations and fungal spores. Aerobiologia 19(2):97–105
Madsen AM, Schlunssen V, Olsen T et al (2009) Airborne fungal and bacterial components in PM1 dust from biofuel plants. Ann Occup Hyg 53(7):749–757
Matthias-Maser S, Jaenicke R (1991) A method to identify biological aerosol particles with radius> 0.3 μm for the determination of their size distribution. J Aerosol Sci 22:S849–S852
Matthias-Maser S, Jaenicke R (1994) Examination of atmospheric bioaerosol particles with radii> 0.2 μm. J Aerosol Sci 25(8):1605–1613
Méjean G, Kasparian J, Yu J et al (2004) Remote detection and identification of biological aerosols using a femtosecond terawatt lidar system. Appl Phys B 78(5):535–537
Meklin T, Haugland RA, Reponen T et al (2004) Quantitative PCR analysis of house dust can reveal abnormal mold conditions. J Environ Monitor 6(7):615–620
Meklin T, Reponen T, McKinstry C et al (2007) Comparison of mold concentrations quantified by MSQPCR in indoor and outdoor air sampled simultaneously. Sci Total Environ 382(1):130–134
Mendell MJ, Mirer AG, Cheung K et al (2011) Respiratory and allergic health effects of dampness, mold, and dampness-related agents: a review of the epidemiologic evidence. Environ Health Perspect 119(6):748
Mensah-Attipoe J, Reponen T, Salmela A et al (2015) Susceptibility of green and conventional building materials to microbial growth. Indoor Air 25(3):273–284
Mensah-Attipoe J, Reponen T, Veijalainen A et al (2016a) Comparison of methods for assessing temporal variation of growth of fungi on building materials. Microbiol 162(11):1895–1903
Mensah-Attipoe J, Saari S, Veijalainen A et al (2016b) Release and characteristics of fungal fragments in various conditions. Sci Total Environ 547:234–243
Metzker ML (2010) Sequencing technologies – the next generation. Nature Rev Genet 11(1):31–46
Mille-Lindblom C, von Wachenfeldt E, Tranvik LJ (2004) Ergosterol as a measure of living fungal biomass: persistence in environmental samples after fungal death. J Microbiol Methods 59(2):253–262
Miller J, Laflamme A, Sobol Y et al (1988) Fungi and fungal products in some Canadian houses. Internat Biodeterior 24(2):103–120
Moularat S, Robine E, Ramalho O et al (2008) Detection of fungal development in closed spaces through the determination of specific chemical targets. Chemosphere 72(2):224–232
Noterman S, Soentoro PS (1986) Immunological relationship of extra-cellular polysaccharide antigens produced by different mould species. Antonie Van Leeuwnhoek 52:393–401
O’Connor DJ, Iacopino D, Healy DA et al (2011) The intrinsic fluorescence spectra of selected pollen and fungal spores. Atmos Environ 45(35):6451–6458
Palmgren U (1986) Collection of airborne micro-organisms on Nuclepore filters, estimation and analysis – CAMNEA method. J Appl Bacteriol Oxford 61(5):401–406
Park J, Cox-Ganser JM (2011) Mold exposure and respiratory health in damp indoor environments. Front Biosci E 3:575–571
Pietarinen V, Rintala H, Hyvärinen A et al (2008) Quantitative PCR analysis of fungi and bacteria in building materials and comparison to culture-based analysis. J Environ Monitor 10(5):655–663
Piñar G, Sterflinger K (2009) Microbes and building materials. Building materials: properties, performance and applications. Nova Publishers, New York, p 163–188
Pitkäranta M, Meklin T, Hyvärinen A et al (2011) Molecular profiling of fungal communities in moisture damaged buildings before and after remediation – a comparison of culture-dependent and culture-independent methods. BMC Microbiol 11(1):1
Pitkaranta M, Meklin T, Hyvarinen A et al (2008) Analysis of fungal flora in indoor dust by ribosomal DNA sequence analysis, quantitative PCR, and culture. Appl Environ Microbiol 74(1):233–244
Pöhlker C, Huffman J, Pöschl U (2012) Autofluorescence of atmospheric bioaerosols – fluorescent biomolecules and potential interferences. Atmos Meas Tech 5(1):37–71
Raimondi V, Agati G, Cecchi G et al (2009) In vivo real-time recording of UV-induced changes in the autofluorescence of a melanin-containing fungus using a micro-spectrofluorimeter and a low-cost webcam. Opt Express 17(25):22735–22746
Rast DM, Baumgartner D, Mayer C et al (2003) Cell wall-associated enzymes in fungi. Phytochemistry 64(2):339–366
Rastogi R, Wu M, Dasgupta I et al (2009) Visualization of ribosomal RNA operon copy number distribution. BMC Microbiol 9:208. doi:10.1186/1471-2180-9-208
Reeslev M, Miller M, Nielsen KF (2003) Quantifying mold biomass on gypsum board: Comparison of ergosterol and beta-N-acetylhexosaminidase as mold biomass parameters. Appl Environ Microbiol 69(7):3996–3998
Reponen T, Seo S, Grimsley F et al (2007) Fungal fragments in moldy houses: a field study in homes in New Orleans and Southern Ohio. Atmos Environ 41(37):8140–8149
Reponen T, Willeke K, Grinshpun S et al (2011) Biological particle sampling. In: Aerosol measurement: principles, techniques, and applications, 3rd Edn. 549–570
Rylander R (2015) β-N-Acetylhexosaminidase (NAHA) as a marker of fungal cell biomass – storage stability and relation to β-glucan. Int J Monitor Anal 3(4):205–209
Rylander R, Reeslev M, Hulander T (2010) Airborne enzyme measurements to detect indoor mould exposure. J Environ Monitor 12(11):2161
Saari S, Reponen T, Keskinen J (2014) Performance of two fluorescence-based real-time bioaerosol detectors: BioScout vs. UVAPS. Aero Sci Technol 48(4):371–378
Saari S, Putkiranta M, Keskinen J (2013) Fluorescence spectroscopy of atmospherically relevant bacterial and fungal spores and potential interferences. Atmos Environ 71:202–209
Saraf A, Larsson L, Burge H et al (1997) Quantification of ergosterol and 3-hydroxy fatty acids in settled house dust by gas chromatography-mass spectrometry: comparison with fungal culture and determination of endotoxin by a Limulus amebocyte lysate assay. Appl Environ Microbiol 63(7):2554–2559
Sattler B, Puxbaum H, Psenner R (2001) Bacterial growth in supercooled cloud droplets. Geophys Res Lett 28(2):239–242
Schaub B, Lauener R, von Mutius E (2006) The many faces of the hygiene hypothesis. J Allergy Clin Immunol 117(5):969–977
Sivaprakasam V, Huston A, Scotto C et al (2004) Multiple UV wavelength excitation and fluorescence of bioaerosols. Opt Express 12(19):4457–4466
Sonesson A, Larsson L, Fox A et al (1988) Determination of environmental levels of peptidoglycan and lipopolysaccharide using gas chromatography with negative-ion chemical-ionization mass spectrometry utilizing bacterial amino acids and hydroxy fatty acids as biomarkers. J Chromatog B: Biomed Sci Appl 431:1–15
Szponar B, Szponar A, Larsson L (2003) Direct assessment of microbial colonisation in damp houses by chemical marker analysis. Indoor Built Environ 12(4):251–254
Thorne PS, Lange JL, Bloebaum P et al (1994) Bioaerosol sampling in field studies: can samples be express mailed? Am Ind Hyg Assoc 55(11):1072–1079
Tischer C, Chen CM, Heinrich J (2011) Association between domestic mould and mould components, and asthma and allergy in children: a systematic review. Eur Respir J 38(4):812–824
Toivola M, Alm S, Reponen T et al (2002) Personal exposures and microenvironmental concentrations of particles and bioaerosols. J Environ Monitor 4(1):166–174
Tringe SG, Zhang T, Liu X et al (2008) The airborne metagenome in an indoor urban environment. PLoS One 3(4):e1862
Vesper S, Wymer L, Meklin T et al (2005) Comparison of populations of mould species in homes in the UK and USA using mould-specific quantitative PCR. Lett Appl Microbiol 41(4):367–373
Vesper S (2007) Development of an Environmental Relative Moldiness Index for US Homes. J Occup Environ Med 49(8):829
Viegas C, Viegas S, Monteiro A et al (2012) Comparison of indoor and outdoor fungi and particles in poultry units. WIT Transactions on Ecology and the Environment 162
Volckens J, Peters TM (2005) Counting and particle transmission efficiency of the aerodynamic particle sizer. J Aerosol Sci 36(12):1400–1408
von Wintzingerode F, Gobel UB, Stackebrandt E (1997) Determination of microbial diversity in environmental samples: pitfalls of PCR-based rRNA analysis. FEMS Microbiol Rev 21(3):213–229
Wittmaack K, Wehnes H, Heinzmann U et al (2005) An overview on bioaerosols viewed by scanning electron microscopy. Sci Total Environ 346(1):244–255
Wu P, Su HJ, Ho H (2000) A comparison of sampling media for environmental viable fungi collected in a hospital environment. Environ Res 82(3):253–257
Yamamoto N, Dannemiller KC, Bibby K et al (2014) Identification accuracy and diversity reproducibility associated with internal transcribed spacer-based fungal taxonomic library preparation. Environ Microbiol 16(9):2764–2776
Yamamoto N, Kimura M, Matsuki H et al (2010) Optimization of a real-time PCR assay to quantitate airborne fungi collected on a gelatin filter. J Biosci Bioeng 109(1):83–88
Young S, Castranova V (2005) Toxicology of 1-3-beta-glucans: glucans as a marker for fungal exposure. CRC Press
Zeng Q, Westermark S, Rasmuson-Lestander Å et al (2006) Detection and quantification of Cladosporium in aerosols by real-time PCR. J Environ Monitor 8(1):153–160
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Mensah-Attipoe, J., Täubel, M. (2017). Analysis Approaches for Fungi in Indoor Environmental Assessments. In: Viegas, C., Viegas, S., Gomes, A., Täubel, M., Sabino, R. (eds) Exposure to Microbiological Agents in Indoor and Occupational Environments. Springer, Cham. https://doi.org/10.1007/978-3-319-61688-9_6
Download citation
DOI: https://doi.org/10.1007/978-3-319-61688-9_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-61686-5
Online ISBN: 978-3-319-61688-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)