Abstract
Two critical stages of a plant improvement programme are (1) the creation of genetic variation and (2) discriminating this variation through progeny evaluation. In each selection stage, a large number of genotypes are evaluated from which a few incrementally superior individuals are either progressed to the next selection stage or released as commercial cultivars. As progeny testing is expensive, and with many genotypes to screen, it is essential to optimise the experimental design and plot technique and statistical analyses of genetic tests.
In the early stages of selection, genotypes are often planted in trials in small, partly replicated, single-row plots due to limited sources of planting material and land requirements for field testing due to the large number of entries to test. Such trials are then subject to variation arising from spatial variability and interplot competition, which reduces the accuracy and confidence of identifying elite genotypes. Spatial variability and interplot competition may seriously affect the estimates of genetic merit and, hence, reduce genetic progress unless they are accounted for in either the trial design or analysis process. Although there are many analysis methods to individually model spatial variability or interplot competition, there are only a few studies that jointly account for both sources of bias. An approach which partitions spatial variability into global trend and extraneous variation and allows for both genotypic and residual level competition is described in this chapter.
As the selection cycle progresses, with only the superior individuals advanced to the next stage, genotypes may be tested across a number of trial locations and possibly over several years. Combining data from these many different sources and levels of imbalance to estimate genetic parameters is possible through a linear mixed model that uses residual maximum likelihood to estimate variance components and best linear unbiased prediction to estimate the random effects. There are many different multiplicative methods available to analyse data from multi-environment trials. The advantages and disadvantages of these methods are described in this chapter. One such method, the factor analytic approach, is widely used throughout plant breeding programs as it can partition spatial variability in local and global trends and extraneous variation and account for heterogeneity of residual variance simultaneously.
Such methods described in this chapter provide options for tropical crop plant breeders to improve and optimise the design and analysis of genetic experiments.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Best linear unbiased predictions (BLUPs)
- Linear mixed models
- Multi-environment trials (MET)
- Spatial variation
- Variogram
- Genotype by environment interaction (GxE)
- Randomised complete block design
- Row-column design
- Partially replicated design
- Heritability
1 Introduction
Breeding programmes deal with large number of activities , including evaluation of hundreds or thousands of genotypes and selection of the best individuals to comprise the next generation of individuals or to be released as new cultivars. Genetic testing is an expensive task that constitutes the largest activity performed in any breeding programme. Phenotyping of genotypes is particularly demanding on small breeding programmes, such is the case of most tropical crops, and for this reason, all activities that aim to maximise (or optimise) the use and quality of the information generated from genetic tests are critical. The basis for this evaluation and selection originates from data and information generated from field and greenhouse experiments, so these need to be carefully planned and analysed.
Genetic tests can be optimised through three different ways: (1) design of experiments, (2) implementation and measurement of trials and (3) statistical analysis. Appropriate selection of the experimental design, their implementation and then their statistical analyses can yield considerable benefits resulting in greater precision of estimates of genetic parameters leading to increased genetic gains from successful selections, and better operational decisions that depend on information obtained from genetic tests , such as heritability , genotype-by-environment interactions , trait-to-trait correlations , etc. Such optimisation can be classified into ‘a priori’ and ‘a posteriori’: the former related to actions that are implemented before the experiment is established (i.e. at the design stage), while ‘a posteriori’ are those actions that are critical to implement once the experiment is established and often relate to tools to be used in for statistical analysis.
There is a plethora of classical and modern literature on the ideal characteristics of a wide array of experimental designs. However, no single design will suit all experimental objectives and environmental conditions found in field tests around the world. Hence, the choice of the ‘best’ design must be made carefully. Statistical and computational tools can be used to generate experimental layouts with great efficiency, where, as always, the principles of replication, randomisation and blocking are critical (for more details about these principles, see Welham et al. (2014)).
Randomised complete block (RCB) designs are the most frequently used in plant breeding. Blocking is important to minimise variability, and this design is effective when within-replicate (or block) variability is relatively small. Where there is large site heterogeneity, or when there are many genotypes to be evaluated, other experimental designs can be more efficient. For example, incomplete block (IB) designs allow for a better control of site heterogeneity by specifying smaller compartments that include a (planned) subset of the genotypes to be tested. IB designs are often generated by implementing an alpha design, a particular class of IB design where the number of genotypes (or entries) is a multiple of block size (John and Williams 1995). Another efficient alternative is the use of row-column (RC) designs that consider both row and columns within a replicate as complete or incomplete blocks. Both of these designs provide greater control of site heterogeneity and can be generated using an array of public and commercial software. For more details about the use and analysis of these designs in the context of plant breeding, we recommend Williams et al. (2002). Other efficient design options include the use of restricted randomisation such as latinisation , nested structures and spatial designs (Whitaker et al. 2002), which can increase the efficiency of the experiments.
For early generation variety trials where large numbers of genotypes are often tested, there may be insufficient planting material to replicate all genotypes. One of the most widely used designs is the use of grid plots where checks (or control genotypes) are repeated several times arranged in a block or incomplete block, depending on the experimental design implemented. Test genotypes are unreplicated and allocated at random to the remaining plots. Examples of this are the various augmented block designs developed by Federer (1956) and Federer and Raghavarao (1975). In an alternative approach, Cullis et al. (2006) proposed the use of partially replicated (p-rep) designs in which a subset of the test genotypes are replicated two or more times, and these are arranged in a resolvable spatial design. Then, the unreplicated test genotypes are randomly allocated to the remaining plots. For a fixed amount of resources, Cullis et al. (2006) found that p-rep designs result in a greater genetic gain than augmented designs.
The second optimisation of genetic testing focuses on the implementation of and measurement within a field design. Here, it is important to observe carefully all operational aspects of field testing, including documentation, labelling, site preparation and crop maintenance. One aspect that is critical here refers to preparing the site in such a way that environmental heterogeneity is minimised. This applies to all soil selection and preparation before planting and its management while the trial is active. In addition, to ensure the best quality of the phenotypic data originating from these trials, adequate definitions of response variables and clarity and consistency on measurement protocols are critical. Any actions that decrease experimental noise will increase the precision of the estimation of genetic parameters and, therefore, increase heritability estimates.
It is also important to collect the most accurate and reliable data that will be used to make decisions on which genotypes are rejected, advanced or ultimately commercially released. For example, Australian sugarcane breeders evaluate genotypes on the basis of their relative economic genetic value for traits of commercial importance – how much value would a genotype add to industry profitability if grown commercially (Wei et al. 2006).
Having collected the data, the genetic tests can be optimised through statistical analysis. This has been an area that has had several important advances over the last few decades. Of special interest for plant breeding is the use of linear mixed models (LMM) that combine estimation procedures such as residual maximum likelihood (REML) to estimate variance components and to predict random effects (or best linear unbiased predictions, BLUP ). LMMs are an extension to the traditional linear models (LM) that allow for more flexible assumptions such as correlations among experimental units (e.g. temporal correlation) and among effects (e.g. by considering the numerator relationship matrix of genetic effects or BLUP ) and heterogeneity of variances (e.g. different error variances for each block or site).
Modern analysis of complex and unbalanced data to obtain parameters, such as site-to-site and trait-to-trait genetic correlations, is possible by fitting LMMs that estimate variance components. Spatial analysis (Gilmour et al. 1997) of field experiments is a useful tool that incorporates the co-ordinates of the experimental units (plots or plants) into the LMM to account for physical proximity by modelling the error structure (i.e. correlations among observations), something that can be extended easily to also model competition among neighbouring plants. This is also particularly important with augmented and p-rep designs where spatial analysis allows for extracting better genetic information from the unreplicated test genotypes.
The greatest benefit of LMMs is that it is possible to combine data from many sources, with different levels of unbalance, into a complex model that will maximise the use of this information to estimate genetic parameters. For example, multi-environment trials (MET) use information from several trials, where not all genotypes are present in all sites, and for each site, there might be different numbers of replicates and precision and therefore heritabilities . These trials are evaluated together into a single LMM to estimate overall breeding values and genetic correlations among sites.
Many statistical tools can be implemented ‘a posteriori’ given a field dataset. One of these is post hoc blocking, where, for a given experimental layout (say a RCB design), a new blocking structure is superimposed on top of the original, and a new linear model is fitted as if the superimposed blocking structure belonged to the original design (Gezan et al. 2006). This tool increases the precision of estimates of heritability and of the predicted genetic values at little extra cost by only marginally increasing the complexity of the analysis.
In the next sections, some of these modern statistical approaches, such as interplot competition and spatial analysis, will be first defined and then illustrated in more detail.
2 Accounting for Interplot Competition
In early-stage selection trials , most plant improvement programmes face the challenge of finding a few incrementally superior individuals from among a large number of lines produced by cross-pollination (Stringer et al. 2011). Due to limitations on planting material and space for field testing, genotypes are often planted in trials in small, partly replicated, single-row plots. Such trials are subject to variation arising from spatial variability and interplot competition, which makes the identification of elite genotypes problematic. Unless accounted for, spatial variability and interplot competition may seriously affect the estimates of genetic merit and, hence, reduce genetic progress.
Interplot competition (also known as interference ) arises when a treatment or response on one experimental or measurement unit may affect the response on neighbouring units (Martin and Eccleston 2004) and is caused by both genetic and environmental sources (Magnussen 1989). It is difficult to quantify and there is no universal method to account for the competitive interactions among genotypes. As resource limitations generally preclude the use of multi-row plots to account for interplot competition, statistical approaches have been developed to adjust for competition in the design and analysis of field trials.
One alternative is to use appropriate experimental layouts, such as the neighbour-balanced (NB) designs suggested by Williams (1952), Street and Street (1987) and Azaïs et al. (1993). However, these designs are not practical where large numbers of genotypes are to be screened, due to the number of replicates required to achieve balance between neighbouring genotypes (Kempton 1982).
In regard to statistical analyses, Besag and Kempton (1986) presented two approaches to estimate interplot competition . Building on earlier work by Kempton (1982), they developed the phenotypic interference model , which is a simultaneous autoregressive approach where competition is assumed to be directly related to yields of neighbouring plots. This has been applied successfully to a wide range of crops including sugar beet (Kempton 1982; Durban et al. 2001), potatoes (Connolley et al. 1993), swedes (Bradshaw 1989) and trees (Resende et al. 2005).
The second model developed by Besag and Kempton (1986) is the treatment or genotypic interference model and was originally proposed by Pearce (1957). In this model, competition effects are associated with genotype differences in characteristics such as plant height, tillering ability, date to maturity and canopy size (Kempton and Lockwood 1984; Talbot et al. 1995). Here, competition effects are associated with the average genotypic value of the nearest neighbouring genotypes rather than the phenotypic response (Stringer et al. 2011). In addition, each treatment is assumed to have a direct effect and a neighbour effect on adjacent plots.
3 Incorporating Spatial Variation
In early-stage field trials, which are typically large, growing conditions may be quite variable across the trial area, leading to the phenomenon known as spatial variability. One of the oldest techniques available to minimise the effect of this variability is the method of check (or control) plots (Wiancko 1914) in which replicated plots are distributed over the trial site as checks and are used as a benchmark to assess the yields of test plots. It is assumed that the checks and test varieties show the same general pattern of response to soil fertility over a trial as the test varieties. If this is not true, then the method of check plots will actually increase the error of assessment (Kempton 1984a; Besag and Kempton 1986). An alternative approach, which may be more useful for dealing with small-scale variation , is spatial or nearest neighbour (NN) analysis where a plot parameter is adjusted by using information from immediate neighbours. Although Papadakis (1937) proposed the earliest NN method , it lacked efficiency.
Spatial methods were largely neglected by statisticians until Wilkinson et al. (1983) developed the smooth trend plus independent error model on which most spatial models have since been based (Stringer et al. 2012). Since then, there have been many alternative approaches, including the one-dimensional models of Gleeson and Cullis (1987) and the two-dimensional approaches of Cullis and Gleeson (1991). Spatial analysis has been successfully applied to early generation trials by Cullis et al. (1992), who found that the response to selection for the spatial method was greater than for check plot method proposed by Wiancko (1914). In all of these models, the covariance structure of the plot errors was modelled as a single component. These techniques were later extended by Gilmour et al. (1997), who demonstrated that modelling plot errors alone as a single process may not be appropriate in most cases, requiring the spatial variation to be partitioned into three components. This approach is currently used to analyse over 1000 cereal variety trials in Australia annually (Stringer et al. 2012) and has resulted in increased accuracy and precision in the estimates of genotype effects in a wide range of crops (Apiolaza et al. 2000; Dutkowski et al. 2002; Gilmour et al. 1997; Grondona et al. 1996; Qiao et al. 2000; Sarker et al. 2001; Silva et al. 2001; Singh et al. 2003).
The methods developed by Gilmour et al. (1997), and later refined by Stefanova et al. (2009), partition spatial variation into three additive components. Atkin (2012) defines these as :
-
Local trend , reflecting small smooth changes due to parameters such as fertility, soil moisture and light
-
Nonstationary global trend which is usually aligned with the columns and rows of a field trial and associated with large-scale changes across the trial, for example, large-scale moisture or fertility gradients
-
Extraneous variation , which usually arises from management practices or experimental procedures that have recurrent patterns, such as spraying operations or serpentine harvesting (harvesting columns of rows in alternating directions)
Previous approaches at removing global trend involved first or second differencing of the data (Gleeson and Cullis 1987), but this often overcomplicated the model. Gilmour et al. (1997) recommend directly fitting nonstationary global trends through the use of polynomial or spline functions (Verbyla et al. 1999) to the row and column co-ordinates. The modelling approach developed by Gilmour et al. (1997) is a sequential approach and commences by including design factors such as replicates that reflect the trial design. Next, modelling local trend is undertaken using a first-order separable autoregressive process in the row and column directions. During the modelling procedure, diagnostic tools, such as the sample variogram and trellis plots , play a large role in determining what effects should be included in a model. This is always followed by formal assessment by using the Wald test for fixed effects and REML likelihood ratio test for random effects.
4 Modelling Competition and Spatial Variation
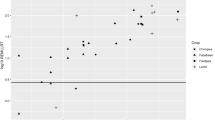
As indicated earlier, interplot competition arises when a treatment or response on one unit affects the response on neighbouring units (Martin and Eccleston 2004). For example, in sugarcane, estimates of cane yield are affected more by interplot competition than are estimates of sugar content when genotypes are evaluated in single-row plots (Fig. 1.1), because plants in adjoining plots compete for resources such as water, fertiliser and sunlight (Jackson and McRae 2001; McRae and Jackson 1998; Skinner 1961; Stringer and Cullis 2002). This often results in a negative correlation between neighbouring plots, biasing estimates of cane yield.
Although there are many approaches in the literature that individually model spatial variability or interplot competition , there are only a few studies that jointly account for both sources of bias. Durbán Reguera (1998) and Durban et al. (2001) presented one such approach. They used cubic smoothing splines to model spatial global trend together with the phenotypic interference model for competition (Stringer et al. 2011). Genotype effects were considered fixed and adjusted profile likelihood was used for parameter estimation (McCullagh and Tibshirani 1990). This model was limited by not considering genotypes to be random nor incorporating a spatial process to model local trend. In a small simulation study based on the Rothamsted downy mildew data, Durbán Reguera (1998) found that the profile likelihood gave biased estimates of the variance components and in some cases the competition parameter was also biased. However, when using McCullagh and Tibshirini’s adjustment to the profile likelihood, bias in the parameters of interest was small. Matassa (2003) developed a method combining both models from Besag and Kempton (1986) for interplot competition together with the methods of Gilmour et al. (1997) for spatial variability. Matassa’s approach was similar to Durbán Reguera (1998) and Durban et al. (2001) in that genotype effects were fixed. However, Matassa (2003) used marginal likelihood and profile likelihood for parameter estimation. On comparing the estimation procedures in a simulation study, Matassa (2003) found that the preferred method depended on what terms were included in the design matrix and also on the sign of the trend parameter.
Stringer et al. (2011) developed an alternative approach to jointly model spatial variability and interplot competition. They partitioned spatial variability into global trend and extraneous variation (Gilmour et al. 1997) and allowed for both genotypic (Besag and Kempton 1986) and residual level competition. Genotype effects were considered to be random, as recommended by Smith et al. (2001), and REML was used for parameter estimation. Stringer et al. (2011) presented two simultaneous autoregressive processes to model competition at the residual level. They recommended an equal-roots second-order autoregressive model for trials where competition is dominant and an equal-roots third-order autoregressive model where both competition and spatial variability exist.
In sorghum breeding trials in Australia, parental lines are evaluated in single-row plots where both interplot competition and spatial variability are present (Hunt et al. 2013). Hunt et al. (2013) extended the methods of Stringer et al. (2011) used for sugarcane clonal trials, by incorporating pedigree information into a LMM. This allowed Hunt et al. (2013) to partition total genetic effects into additive and nonadditive components for parent evaluation in the presence of both competition and spatial effects. The methods developed by Stringer et al. (2011) and Hunt et al. (2013) are routinely applied to sugarcane clonal and full-sib family (produced from biparental cross-pollination) trials from Queensland, Australia. In such trials, clones and families are evaluated in single-row plots and large spatial trends and interplot competition are regularly present. The presence of spatial variation and interplot competition effects in sugarcane family trials will probably have a similar effect on estimating additive genetic effects of sugarcane parents as with estimating total genetic or clonal effects and lead to biased estimates of breeding values (BV) for cane yield (in the presence of both spatial variation and interplot competition) and sugar content (in the presence of spatial variation only). In turn, this will bias the ranking of parents and so impact the outcomes of parental selection.
The two statistical models explained below (a basic RCB design model with and without modelling spatial variation) can be used to estimate additive genetic effects of sugarcane parents from family trials for parent selection (Atkin 2012) by applying some of the techniques used by Stringer et al. (2011) and Hunt et al. (2013).
4.1 Base Model: RCB Without Modelling Spatial Variation
Consider an experiment consisting of p trials that contains a total of m genotypes (families). Each trial is laid out in a rectangular array of r rows and c columns (n = r × c) (Fig. 1.1). Where the data are ordered as rows within columns, the mixed linear model for y (n×1) combined across trials is
where b (b×1) is a vector of fixed effects with the associated design matrix X (n×b); g (mp×1) contains the random genotype and genotype by environment effects of m entities in each of p trials with indicator matrix Z g (n×mp); u (d×1) contains the random replicate effects with associated design matrix Z u (n×d); and e (n×1) is a vector of plot error effects combined across trials. Vector b contains only an overall mean effect for each trial or more complex design structures.
Here, vector g is the random genotypic effect of unique parents (for each trial), where a sugarcane parent can be used as either a male or a female, or both. Using a biparental model (or a reduced animal model) (Mrode 2005; Quaas and Pollak 1980), vector g is then further partitioned into additive and nonadditive genetic effects as per Costa e Silva et al. (2004). The prediction of BVs described here is also applicable to the next model described below: the spatial model.
4.2 Spatial Model: RCB Plus Modelling Spatial Variation
The above model can be extended to include the partitioning of spatial variability, where vector b contains an overall mean for each trial, as well as trial-specific modelling due to global trend (Stefanova et al. 2009). Global trend is accommodated in the model by using design factors such as linear row and/or linear column effects or by fitting spline functions to the row and column co-ordinates (Verbyla et al. 1999). Vector u includes effects associated with the modelling of extraneous variation due to experimental procedures and blocking design factors specific to each trial or sub-trial (in cases where a trial comprised of two or more sub-trials). For each trial, vector e is further partitioned into a vector that represents a spatially dependent process and a vector of residual errors (Gilmour et al. 1997).
Local spatial trend is modelled using a first-order separable autoregressive (AR) process in the row (AR(1)) and column directions (AR(1)), as recommended by Cullis et al.(1998), Gilmour et al. (1997) and Grondona et al. (1996). After fitting the local trend, diagnostic tools such as the sample variogram and trellis plots (Gilmour et al. 1997) can be used to determine if global spatial trend and/or extraneous variation needed to be included in the model. An example of a theoretical variogram for an AR(1) × AR(1) process is given in Fig. 1.2. This variogram has a smooth appearance and an exponential increase in the row and column directions reaching a plateau giving it a ‘tabletop’ appearance. Departure from this smooth appearance indicates the presence of extraneous variation; similarly, if the sample variogram fails to reach a plateau in the row and/or column direction, this indicates the presence of a global trend that needs to be incorporated into the LMM (Stringer and Cullis 2002). An example of the presence of a global trend is given in Fig. 1.3 for which a linear row and column effect would then be fitted. The inclusion of these fixed effects is based on visual inspection of the sample variogram followed by a formal assessment using the Wald test (Agresti 1990). An example of extraneous variation is given in Fig. 1.4 for which a random row effect would be fitted. The inclusion of random effects is also based on visual inspection for the sample variogram , followed by the use of the likelihood ratio test to ascertain if the change in REML log-likelihood for random effects is significant.
Example of a theoretical variogram for an AR(1) × AR(1) process in the absence of both global and extraneous trend (From Stringer and Cullis (2002) – used with permission)
Example of a sample variogram for an AR(1) × AR(1) process indicating the presence of a global trend in the row and column direction (From Stringer and Cullis (2002) – used with permission)
Example of a sample variogram for an AR(1) × AR(1) process indicating the presence of extraneous variation (From Stringer and Cullis (2002) – used with permission)
These types of analyses are routinely performed in the Australian sugarcane breeding programme using the statistical package ASReml (Gilmour et al. 2006) and could be extended to other tropical crops where competition and spatial effects are often experienced in field trials.
5 Further Approaches That Incorporate Genotype-by-Environment Interactions
Mixed model analyses of data from multi-environment trials (METs) can be used to partition the total variation into sources such as trial, genotype and genotype-by-environment (G×E) interactions . Although this provides an estimate of the magnitude of G×E, it does not provide any insight into the nature of G×E effects (Kempton 1984b). Multiplicative methods are particularly useful at describing G×E interactions and have been widely used in a fixed-effects setting. The earliest of these was the regression on mean model, where either the phenotypic values or interaction is regressed on environmental indices. This was first suggested by Yates and Cochran (1938) and enhanced by Finlay and Wilkinson (1963); however, this approach assumes genotypes respond linearly to environmental change (Flores et al. 1998). Freeman (1973) suggested the use of multiplicative methods in genetic analyses, and of these, the additive main effects and multiplicative interaction (AMMI) has been used very widely (Gauch and Zobel 1988). AMMI combines the additive analysis of variance for main effects with the multiplicative principal component analysis for the interaction. However, AMMI requires data to be balanced and, hence, it can be too restrictive for the analysis of MET data for most crops (Smith et al. 2001).
A key issue often neglected in G×E studies is the need to model plot-level residuals (Smith et al. 2001). Individual trials routinely exhibit spatial variability (Atkin 2012) as a correlation in residuals among neighbouring plots, and it is common for residual variances to differ among trials. Estimates of genotype main effects and G×E interactions may be biased if this is not accounted for (Cullis et al. 1998). An approach which overcomes these limitations was developed by Smith et al. (2001), in which spatial variability within a trial is partitioned into local and global trends and extraneous variation using the methods of Gilmour et al. (1997) (described previously), and the heterogeneity of residual variance among trials is accounted for. This is called the factor analytic model (FA) and implies that genetic effects are correlated between trials; hence, it allows the genetic variation at each environment to differ and allows for different covariances between pairs of environments (Smith et al. 2001). This flexibility requires a large number of variance components to be estimated. However, the particular FA model proposed by Smith et al. (2001) uses the algorithms of Thompson et al. (2003) by providing a parsimonious fit for parameter estimation. Therefore, the multiplicative mixed model with random genotype effects, i.e. the FA model (as used by Chapman et al. 2004), is currently considered the most appropriate approach for the analysis of G×E interactions in breeding programmes for many crops including sugarcane.
6 Final Remarks
The array of statistical tools available in quantitative genetics for the analysis of messy and complex data that originates from breeding trials is diverse, noisy and continuously evolving. The emergence of powerful statistical software that can deal with this data has allowed breeders to extract more information from each experiment, including aspects such as competition and spatial correlations. In addition, the availability of large quantities of molecular data to be incorporated into the linear mixed models , for example, by calculating observed relationships among genotypes based on molecular markers (VanRaden 2008), has widened the options to improve and optimise the design and analysis of genetic experiments. Here we have presented some tools, but these modern tools will constitute, in the near future, a daily part of all statistical analysis performed by many breeding programmes across the world.
References
Agresti A (1990) Categorical data analysis. Wiley, New York
Apiolaza LA, Gilmour AR, Garrick DJ (2000) Variance modelling of longitudinal height data from a Pinus radiata progeny test. Can J For Res 30:645–654
Atkin FC (2012) Using family data to maximise gentic gain from parent selection in a sugarcane breeding program. PhD Thesis, The University of Queensland
Azaïs JM, Bailey RA, Monod H (1993) A catalogue of efficient neighbour-designs with border plots. Biometrics 49:1252–1261
Besag J, Kempton R (1986) Statistical analysis of field experiments using neighbouring plots. Biometrics 42:231–251
Bradshaw JE (1989) Interplot competition in yield trials of swedes (Brassica napus ssp. rapifera L.) Euphytica 42:135–140
Chapman SC, Jackson PA, Rattey A (2004) Genotype by environment interactions across the Australian sugarcane industry. Proc Aust Soc Sugar Cane Technol 26:8 pp
Connolley T, Currie ID, Bradshaw JE, McNicol JW (1993) Inter-plot competition in yields of potatoes (Solanum tuberosum L.). Ann Appl Biol 123:367–377
Costa e Silva J, Borralho NMG, Potts BM (2004) Additive and non-additive genetic parameters from clonally replicated and seedling progenies of Eucalyptus globulus. Theoret Appl Genet 108:1113–1119
Cullis BR, Gleeson AC (1991) Spatial analysis of field experiments: an extension to two dimensions. Biometrics 47:1449–1460
Cullis BR, Gleeson AC, Thomson FM (1992) The response to selection of different procedures for the analysis of early generation variety trials. J Agric Sci 118:141–148
Cullis BR, Gogel BJ, Verbyla AP, Thompson R (1998) Spatial analysis of multi-environment early generation variety trials. Biometrics 54:1–18
Cullis BR, Smith AB, Coombes NE (2006) On the design of early generation variety trials with correlated data. J Agric Biol Environ Stat 11:381–393
Durbán Reguera ML (1998) Modelling spatial trends and local competition effects using semiparametric additive models. PhD Thesis, Department of Actuarial Mathematics and Statistics, Heriot-Watt University
Durban M, Currie ID, Kempton RA (2001) Adjusting for fertility and competition in variety trials. J Agric Sci 136:129–140
Dutkowski GW, Silva JCE, Gilmour AR, Lopez GA (2002) Spatial analysis methods for forest genetic trials. Can J For Res 32:2201–2214
Federer WT (1956) Augmented (or Hoonuiaku) designs. Hawaii Plant Rec 55:191–208
Federer WT, Raghavarao D (1975) On augmented designs. Biometrics 31:29–35
Finlay KW, Wilkinson GN (1963) The analysis of adaptation in a plant breeding programme. Aust J Agric Res 14:742–754
Flores F, Moreno MT, Cubero JI (1998) A comparison of univariate and multivariate methods to analyze G × E interaction. Field Crops Res 56:271–286
Freeman GH (1973) Statistical methods for the analysis of genotype-environment interactions. Heredity 31:339–354
Gauch HG Jr, Zobel RW (1988) Predictive and postdictive success of statistical analyses of yield trials. Theoret Appl Genet 76:1–10
Gezan SA, Huber DA, White TL (2006) Post-hoc blocking to improve breeding value prediction. Can J For Res 36:2141–2147
Gilmour AR, Cullis BR, Verbyla AP (1997) Accounting for natural and extraneous variation in the analysis of field experiments. J Agric Biol Environ Stat 2:269–293
Gilmour AR, Gogel BJ, Cullis BR, Thompson R (2006) ASReml user guide, 2.0 edn. VSN International Ltd, Hemel Hempstead
Gleeson AC, Cullis BR (1987) Residual maximum likelihood (REML) estimation of a neighbour model for field experiments. Biometrics 43:277–288
Grondona MO, Crossa J, Fox PN, Pfeiffer WH (1996) Analysis of variety yield trials using two-dimensional separable ARIMA processes. Biometrics 52:763–770
Hunt CH, Smith AB, Jordan DR, Cullis BR (2013) Predicting additive and non-additive genetic effects from trials where traits are affected by interplot competition. J Agric Biol Environ Stat 18:53–63
Jackson P, McRae TA (2001) Selection of sugarcane clones in small plots: effects of plot size and selection criteria. Crop Sci 41:315–322
John JA, Williams ER (1995) Cyclic and computer generated designs, Monographs on Statistics and Applied Probability, vol 38. Chapman & Hall, London, 255 pp
Kempton RA (1982) Adjustment for competition between varieties in plant breeding trials. J Agric Sci 98:599–611
Kempton RA (1984a) The design and analysis of unreplicated field trials. Vortrage Pflanzenzuchtung 7:219–242
Kempton RA (1984b) The use of biplots in interpreting variety by environment interactions. J Agric Sci 103:123–135
Kempton RA, Lockwood G (1984) Inter-plot competition in variety trials of field beans (Vicia faba L.). J Agric Sci 103:293–302
Magnussen S (1989) Inter-plant interactions and their influence on within and among plot variances. Scand J Forest Res 4:369–377
Martin RJ, Eccleston JA (2004) Variance-balanced designs under interference for dependent observations. J Stat Plan Inference 119:207–223
Matassa V (2003) Optimisation of experimental design and analysis for sugarcane variety trials. PhD Thesis, The University of Queensland
McCullagh P, Tibshirani R (1990) A simple method for the adjustment of profile likelihoods. J R Stat Soc Series B Stat Methodol 52:325–344
McRae TA, Jackson PA (1998) Competition effects in selection trials. Proc Aust Soc Sugar Cane Technol 20:154–161
Mrode RA (2005) Linear models for the prediction of animal breeding values, 2nd edn. CABI Publishing, Wallingford, 344 pp
Papadakis JS (1937) Méthode statistique pour des expériences sur champ. Bull Inst Amél Plante à Salonique 23:30 pp
Pearce SC (1957) Experimenting with organisms as blocks. Biometrika 44:141–149
Qiao CG, Basford KE, DeLacy IH, Cooper M (2000) Evaluation of experimental designs and analyses in wheat breeding trials. Theoret Appl Genet 100:9–16
Quaas RL, Pollak EJ (1980) Mixed model methodology for farm and ranch beef cattle testing programs. J Anim Sci 51:1277–1287
Resende MDV, Stringer J, Cullis B, Thompson R (2005) Joint modelling of competition and spatial variability in forest field trials. Braz J Math Stat 23:7–22
Sarker A, Singh M, Erskine W (2001) Efficiency of spatial methods in yield trials in lentil (Lens culinaris ssp. culinaris). J Agric Sci 137:427–438
Silva JCE, Dutkowski GW, Gilmour AR (2001) Analysis of early tree height in forest genetic trials is enhanced by including a spatially correlated residual. Can J For Res 31:1887–1893
Singh M, Malhotra RS, Ceccarelli S, Sarker A, Grando S, Erskine W (2003) Spatial variability models to improve dryland field trials. Exp Agric 39:151–160
Skinner JC (1961) Sugar cane selection experiments 2. Competition between varieties. Technical Communications. BSES, Brisbane
Smith A, Cullis B, Thompson R (2001) Analyzing variety by environment data using multiplicative mixed models and adjustments for spatial field trend. Biometrics 57:1138–1147
Stefanova KT, Smith AB, Cullis BR (2009) Enhanced diagnostics for the spatial analysis of field trials. J Agric Biol Environ Stat 14:392–410
Street AP, Street DJ (1987) Combinatorics of experimental design. Oxford University Press, Oxford, 400 pp
Stringer JK, Cullis BR (2002) Application of spatial analysis techniques to adjust for fertility trends and identify interplot competition in early stage sugarcane selection trials. Aust J Agric Res 53:911–918
Stringer JK, Cullis BR, Thompson R (2011) Joint modeling of spatial variability and within-row interplot competition to increase the efficiency of plant improvement. J Agric Biol Environ Stat 16:269–281
Stringer JK, Smith AB, Cullis BR (2012) Spatial analysis of agricultural field experiments. In: Hinkelmann K (ed) Design and analysis of experiments, Special designs and applications, vol 3. Wiley, Hoboken, pp 109–136
Talbot M, Milner AD, Nutkins MAE, Law JR (1995) Effect of interference between plots on yield performance in crop variety trials. J Agric Sci 124:335–342
Thompson R, Cullis B, Smith A, Gilmour A (2003) A sparse implementation of the average information algorithm for factor analytic and reduced rank variance models. Aust NZ J Stat 45:445–459
VanRaden PM (2008) Efficient methods to compute genomic predictions. J Dairy Sci 91:4414–4423
Verbyla AP, Cullis BR, Kenward MG, Welham SJ (1999) The analysis of designed experiments and longitudinal data by using smoothing splines. Appl Stat 48:269–311
Wei X, Jackson P, Cox M, Stringer J (2006) Maximising economic benefits to the whole sugarcane industry from the BSES-CSIRO sugarcane improvement program. Proc Aust Soc Sugar Cane Technol 28:181–186
Welham SJ, Gezan SA, Clark SJ, Mead A (2014) Statistical methods in biology: design and analysis of experiments and regression. Chapman and Hall/CRC, Boca Raton. 608 pp
Whitaker D, Williams ER, John JA (2002) CycDesigN: a package for the computer generation of experimental designs. Version 2.0. CSIRO, Canberra. 33 pp
Wiancko AT (1914) Use and management of check plats in soil fertility investigations. J Am Soc Agron 6:122–124
Wilkinson GN, Eckert SR, Hancock TW, Mayo O (1983) Nearest neighbour (NN) analysis of field experiments. J R Statist Soc B 45:151–211
Williams EJ (1952) Experimental designs for serially correlated observations. Biometrika 39:152–167
Williams ER, Matheson AC, Harwood CE (2002) Experimental design and analysis for tree improvement, 2nd edn. CSIRO Publishing, Collingwood, 214 pp
Yates F, Cochran WG (1938) The analysis of groups of experiments. J Agric Sci 28:556–580
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Stringer, J.K., Atkin, F.C., Gezan, S.A. (2017). Statistical Approaches in Plant Breeding: Maximising the Use of the Genetic Information. In: Genetic Improvement of Tropical Crops. Springer, Cham. https://doi.org/10.1007/978-3-319-59819-2_1
Download citation
DOI: https://doi.org/10.1007/978-3-319-59819-2_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-59817-8
Online ISBN: 978-3-319-59819-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)