Abstract
Radish is important as a root vegetable, a leafy vegetable, a fruit vegetable, an oil crop, and also as a cover plant. The economic importance and characteristics of radish differ between the East and the West of the world. In the East, there are radish cultivars having large roots with various shapes called “Asian big radish” and those grown for production of immature pods or oil seeds, whereas radish is a small vegetable grown within one month in the West. Asian big radish is expected to eventually become popular in the West. Radish belongs to the genus Raphanus, but is similar to the Brassica species except for the shape of pods and seeds. Despite their similarities, the order of genes in chromosomes is quite different between Raphanus and Brassica. Radish genome sequences have been published from three groups using similar cultivars, and therefore, collaboration for combining sequence data is considered to be effective for determination of more reliable genome sequences. Some radish lines have high salt tolerance and disease resistance different from Brassica crops. Radish also has a characteristic glucosinolate composition. Since radish can be crossed with Brassica species, it is also important as a genetic resource for Brassica crop breeding.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1.1 Introduction
Radish is a Brassicaceae crop, mainly used as a root vegetable. Roots of radish have large variations in size and shape. European small radish has a small round root ca. 2–3 cm in diameter, whereas “Sakurajima-daikon” in Japan has a large round root of more than 30 cm. Most widely grown radish cultivars in Asia have roots of cylindrical shape ca. 10 cm in diameter and ca. 40 cm in length, whereas “Moriguchi-daikon” in Japan has a long cylindrical root of more than 2 m with a diameter of ca. 3 cm. The color of radish roots is also various. The major Asian radish has a white thick root and is called “East Asian big long radish,” “white radish,” or “daikon” in Japanese (“Asian big radish” hereafter), whereas European small radish, which is commonly called “radish,” and the root surface of some landraces in Asia are red, although the red part develops from a hypocotyl. There are also purple, green, and black colors on the root surface in other landraces. Some cultivars in China are red or green inside the roots.
Young plants removed by thinning are used as a leafy vegetable. “Kosena-daikon,” a Japanese radish landrace, which has a small root, is a cultivar specialized as a leafy vegetable. Immature pods of rat’s tail radish, which is cultivated in Southeast Asia, e.g., Thailand and India, are used as a vegetable. The root of rat’s tail radish is not thick. A radish sprout called “Kaiware-daikon” is popular as a vegetable in Japan, and the recent vogue of Japanese cuisine has also made it popular in Western countries. Since radish seeds contain a high amount of oil, ca. 40%, radish is also produced as an oil crop (Ahuja et al. 1987). Furthermore, cultivation of radish as a cover crop to avoid soil erosion and to suppress weeds has become popular in the USA and Canada (Weil and Kremen 2007).
The species name of radish is Raphanus sativus, its genus being different from that of the major Brassicaceae oil crops and vegetables, i.e., Brassica. European small radish is similar to red turnip landraces in Brassica rapa. “Shogoin-daikon,” a Japanese radish landrace having a large round root, is very similar to “Shogoin-kabu,” a Japanese turnip landrace. It is not easy to distinguish radish from turnip at a vegetable market. The greatest morphological difference between radish and turnip is the shape of siliques (Fig. 1.1). Siliques of Brassicaceae plants have two parts, i.e., a beak formed from the sub-stylar region and a part having valves, called the valvar portion (Gomez-Campo 1980). Although Brassica and most Brassicaceae species have seeds in the valvar portion without seeds in the beak, radish has seeds in the beak and the valvar portion of radish is absent. Seeds of radish are arranged in a line in a silique and are much larger than those of Brassica and most other Brassicaceae species, which are arranged in two lines in a silique. Radish siliques do not dehisce and can float on water.
Raphanus sativus is closely related to Raphanus raphanistrum , which is a wild radish distributed as a weed in farmlands, roadsides, and coastal dunes in Europe, North America, and Australia. R. sativus has white or pale purple flowers, whereas R. raphanistrum has yellow flowers, although some lines of R. raphanistrum have white flowers. Siliques of R. raphanistrum are longer than those of R. sativus and can be fragmented into pieces containing a single seed. R. raphanistrum can be crossed with R. sativus, and hybrid progeny have normal seed fertility, indicating that these two species have the same genome. In Asian countries, R. raphanistrum is not common as a farmland weed, whereas R. sativus var. raphanistroides is widely distributed in coastal dunes. R . sativus var. raphanistroides can grow near the seaside but is rarely found in farmlands or riversides, suggesting that it has high salt tolerance with low competitive ability as a weed in farmlands or riversides.
Although radish and Brassica species are distantly related, intergeneric hybrids between them can be obtained and radish is a useful genetic resource for breeding of Brassica crops especially in breeding for disease resistance and stress tolerance. In this chapter, biological and economical characteristics of radish are outlined.
1.2 Classification of Radish
Raphanus sativus and R. raphanistrum belong to the subtribe Raphaninae , which is different from the subtribe containing Brassica, Sinapis eruca, and Diplotaxis, i.e., Brassisineae . The morphology of silique is an important taxonomic criterion to separate Raphanus from Brassica. However, recent molecular taxonomic studies using DNA polymorphism data of chloroplast DNA (cpDNA) and nuclear DNA do not support the grouping of Brassica, Sinapis eruca, and Diplotaxis excluding Raphanus, but suggest that Raphanus is closely related to Brassica rapa and Brassica oleracea (see Chap. 2).
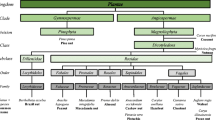
The tribe Brassiceae contains seven subtribes: Brassicinae, Cakilinae , Moricandiinae , Raphaninae, Savignyinae , Vellinae , and Zillinae . Molecular phylogeny based on cpDNA restriction site polymorphism has indicated that the tribe Brassiceae is separated into seven lineages, i.e., Rapa/Oleracea lineage, Nigra lineage, Crambe lineage, Cakile lineage, Vella lineage, Savingya lineage, and Zilla lineage. Cakile lineage, Vella lineage, Savingya lineage, and Zilla lineage correspond to Cakilinae , Vellinae, Savignyinae , and Zillinae , respectively, but species in Brassicinae, Raphaninae , and Moricandiinae are intermingled and reclassified into Rapa/Oleracea lineage containing B. rapa, B. oleracea, Eruca vesicaria, Erucastrum abyssinicum, Diplotaxis erucoides, Moricandia arvensis, etc., Nigra lineage containing Brassica nigra, Brassica flruticulosa, Sinapis arvensis, Sinapis alba , Erucastrum varium, etc., and Crambe lineage (Warwick and Black 1997; Warwick and Hall 2009). R. sativus and R. raphanistrum belong to the Rapa/Oleracea lineage. Phylogenetic studies using nucleotide sequence data of the nuclear S-locus related 1 (SLR1) gene (Inaba and Nishio 2002) and the nuclear ribosomal internal transcribed spacer (ITS) and chloroplast trnL intron sequences (Warwick and Sauder 2005) have shown a slightly different classification of species, but R. sativus has always been found to belong to a group of B. rapa and B. oleracea. These findings suggest that morphological change of the silique from the Brassica type to the Raphanus type in the Rapa/Oleracea lineage occurred independently from that in the Crambe lineage, the Cakile lineage, and the Zilla lineage, which also have siliques of the Raphanus type.
1.3 Genome of Radish
Radish is a monogenomic species having a chromosome number of gametes (n) of nine. Although it is monogenomic, most genes have three similar copies in a genome as in the Brassica monogenomic species, suggesting that genome triplication has occurred after divergence from an ancestor of Arabidopsis thaliana . The chromosome number is the same as that of B. oleracea, but the genome structure of radish is quite different from those of B. oleracea and closely related B. rapa, as has been suggested by a classical genome study analyzing meiotic chromosome pairing (Harberd and McArthur 1980; Mizushima 1980). Genome syntenies of radish with B. rapa or B. oleracea can be observed in short regions, but there are few overall syntenies (Li et al. 2011; Shirasawa et al. 2011), suggesting that genome rearrangements have occurred many times in the speciation processes of these species (Mun et al. 2015; see Chap. 5). This is in contrast to the relationships between rice and other Poaceae species (Bennetzen and Freeling 1997) and between tomato and other Solanum species (Peters et al. 2012).
Genome size of radish has been estimated to be from 526 (Arumuganathan and Earle 1991) to 573 Mb (Johnston et al. 2005) by flow cytometry. Unigene sequences of radish have been published in the RadishBase (http://bioinfo.bti.cornell.edu/cgi-bin/radish/index.cgi) (Shen et al. 2013). The sequences of the whole radish genome were first reported by Kitashiba et al. (2014) and are available in “Raphanus sativus Genome Data Base” (http://radish.kazusa.or.jp). Using short-read genomic sequences of 191.1 Gb (246.5 times the radish genome size estimated by them) obtained by next-generation sequencers and both end sequences of 20,736 BAC clones, 76,592 scaffolds of 402 Mb spanning 75.9% of estimated genomic size containing 61,572 predicted genes were obtained. To a high-density linkage map of 2553 DNA markers , 1345 scaffolds were assigned. Using next-generation sequencing data 121.8 times the radish genome size, 40,123 scaffolds spanning 393.3 Mb have been constructed (Mitsui et al. 2015). Jeong et al. (2016) have also read the radish genome sequences and assembled them into 10,674 scaffolds spanning 426 Mb. In these studies, similar cultivars of Asian big radish were used for determination of the genome sequence. Reassembling of sequences or overlapping of scaffolds obtained in these studies may enable construction of much longer scaffolds covering large parts of the radish genome. Although these studies have so far been performed with the support of different organizations, collaboration of these teams is expected (see Chaps. 3 and 4).
1.4 Characteristic Traits of Radish
Raphanus sativus is an allogamous species having self-incompatibility. Therefore, populations of radish landraces and wild radish have high genetic polymorphism, which is reflected by DNA polymorphism in a cultivar (see Chap. 6). Self-incompatibility can be overcome by bud pollination or high-concentration CO2 treatment (Niikura and Matsuura 2000), and selfed progeny can be obtained. However, they show inbreeding depression , making it difficult to obtain inbred lines.
Like self-incompatibility of Brassica species, S-receptor kinease (SRK) is the recognition molecule of the stigma, and SP11, also called SCR, is the recognition molecule of pollen (Okamoto et al. 2004). The genes of these recognition molecules, i.e., SRK and SP11/SCR, have multiple alleles and are inherited as one set, which is called S haplotype , by progeny. There are many S haplotypes in R. sativus (Sakamoto et al. 1998; see Chap. 13), and nucleotide sequences of SRK and SP11/SCR of some S haplotypes in R. sativus are similar to those of B. rapa, suggesting that S haplotypes possessed by an ancient species were inherited by species in both Raphanus and Brassica without great alteration of nucleotide sequences (Okamoto et al. 2004).
Most radish cultivars are vegetables having thick roots. Size, shape, and color of roots are important traits of radish. Quantitative trait loci (QTLs) controlling root shape and color have been reported (Tsuro et al. 2008; Hashida et al. 2013). However, root thickening is influenced by other traits, such as flowering . In our QTL analysis of root thickness using a progeny obtained by crossing between “Aokubi” having a white thick root with green color on the top and a cultivar of rat’s tail radish, a QTL having the highest LOD score corresponded to a QTL for bolting time (unpublished). This may be natural because early flowering is considered to have an adverse effect on root thickening. Transcriptome analysis of growing roots has been performed, and genes related to root thickening were identified (see Chaps. 8 and 9).
The color of root surface in radish is derived from anthocyanins . Pigments of red color and purple color of radish cultivars are pelargonidin and cyanidin , respectively. A hybrid between a red root line and a white root line was found to have purple roots, suggesting that the red and the white have knockout mutations in different genes responsible for cyanidin synthesis and the functional alleles in the red and the white acted as complementary genes. Alleles of the gene for flavonoid 3′-hydroxylase (F3′H) in red root cultivars have been revealed to have insertions of a Ty3/gypsy transposon or a helitron (Ozeki 2010). A gene for dihydroflavonol reductase (RsDFR) and a gene for anthocyanidin synthase (RsANS) are expressed in the epidermal tissues of roots of a red-skinned cultivar, whereas these genes are not expressed in those of a white-skinned cultivar (Park et al. 2011).
The large seed size is a remarkable trait of radish among Brassicaceae species. Seed size of radish is about five times in weight that of B. rapa. Due to the large seed size, cotyledons and hypocotyls of seedlings are larger than those of Brassica. The larger size of seedlings facilitates cultivation by direct sowing of seeds in the field and produces bigger sprouts than Brassica. The large seed trait is related to the shape of siliques. Amphidiploid plants of intergeneric hybrids between R. sativus and B. rapa have siliques of an intermediate type, which has both the beak and the valvar portion with a few seeds (Fig. 1.2). The amphidiploid plants are partially sterile, but a small number of seeds can be obtained. The size of the seeds is also intermediate between Raphanus and Brassica.
The pungency of radish is due to isothiocyanates , which are hydrolyzed products of glucosinolates . The content of glucosinolates has a great influence on the taste of grated fresh radish and radish salad. The major glucosinolate in radish roots is glucoraphasatin (also called 4-methylthio-3-butenyl glucosinolate, dehydroerucin), and variation of the glucosinolate composition is limited. However, there is a great variation in the contents of glucoraphasatin in Japanese radish cultivars (Ishida et al. 2015). QTLs controlling glucosinolate content in the root have been reported (Zou et al. 2013), and candidate genes responsible for the content have been inferred. Recently, a mutant having a high amount of glucoerucin without glucoraphasatin has been selected (Ishida et al. 2015), and the gene responsible for this mutation has been identified (Kakizaki et al. 2017).
Although radish generally has high salt tolerance, such tolerance of R. sativus var. raphanistroides is especially high (Nasu et al. 2012). Although genetic analysis of salt tolerance of R. sativus var. raphanistroides has not yet advanced, the genes for salt tolerance of radish will be useful for breeding of Brassica crops. In the production of radish, high-temperature stress has become a serious problem. The core of a radish root turns reddish brown by the high-temperature stress, resulting in unmarketable products. However, since there is a variation in sensitivity to the high-temperature stress, it should be possible to develop a tolerant cultivar.
After bolting , radish roots become fibrous and unsuitable for markets. Therefore, a late bolting trait is preferred. One the other hand, rat’s tail radish or cultivars for oil production are requested to flower even in tropical regions. For floral induction, vernalization is required, but rat’s tail radish can flower without vernalization. There is a large variation in vernalization requirement in radish as in many other winter crops. In a QTL analysis of bolting time using a progeny obtained by a cross between “Aokubi” and rat’s tail radish, a QTL having a significant LOD score was detected in a region containing an FLC gene (see Chap. 11).
Although clubroot caused by Plasmodiophora brassicae is a serious problem in the production of Brassica vegetables, Japanese and Korean cultivars of radish are generally tolerant to clubroot. Radish can be used as a source of resistance genes for the breeding of Brassica vegetables (Akaba et al. 2009). A QTL controlling clubroot resistance has been mapped on LG1 (Kamei et al. 2010), corresponding to Rs5 (Kitashiba et al. 2014, Chap. 3). In radish cultivation, Fusarium yellow caused by Fusarium oxysporum is one of the most serious diseases. Leaves wilt and the vascular tissue in a root is browned by this disease. Some cultivars and landraces have relatively high resistance to Fusarium yellow (see Chap. 12).
Radish is recalcitrant in tissue and cell cultures because of its low plant regeneration ability. There have been few reports on successful protoplast culture, anther or isolated microspore culture (Takahata et al. 1996), and plant transformation (Park et al. 2005; Cho et al. 2008). The difficulty of plant transformation makes it impossible to demonstrate the function of isolated genes in radish. Development of an efficient plant transformation technique is indispensable for basic and applied studies in radish. Since there must be genetic variation in plant regeneration ability, identification of cultivars or lines having high regeneration ability would be the first step for the development of in vitro culture techniques.
1.5 Use of Radish as a Vegetable, a Feed, and Materials
Although radish was once the most cultivated vegetable in Japan, its production has been decreasing because consumption of radish pickles has recently decreased. Exploring novel ways of utilization is required for increasing the economic value of radish. European small radish is used for salad after slicing or rough cutting. Asian big radish has not been used for salad, but shredded white radish, which is a garnish of sliced raw fish, is increasingly used in a salad. Since crispiness is important for the shredded radish, white radish cannot be replaced by turnip roots. The white radish root can be stored and is inexpensive, and therefore, it is a good material for use in salad.
Tempura and shabu-shabu, popular Japanese dishes, are garnished with grated radish, which contains amylase and other enzymes to help digestion. Grated radish is also served with grilled fish or dried young sardines. Grated radish is pungent because of its high content of isothiocyanates . By disruption of cells by grating, glucosinolates are hydrolyzed by the enzyme myrosinase , which is stored in the vacuoles of myrosin cells. Isothiocyanates have anti-bacterial and anti-fungal activity, and some isothiocyanates have been revealed to have anti-carcinogenic activity (Ishida et al. 2015). The isothiocyanate produced from the major glucosinolate in roots, i.e., 4MTB-GSL, has also been reported to be able to reduce cell proliferation and induce apoptosis in cancer cell lines with very limited toxicity toward normal human T-lymphocytes (Papi et al. 2008). Due to the increasing popularity of Japanese cuisine, consumption of grated radish will most probably increase.
A popular boiled dish containing white radish is “oden,” which is served even in convenience stores in Japan. In Chinese cuisine, Asian big radish is mainly served in boiled dishes. Although digestive enzymes and isothiocyanates in radish roots are lost, pieces of radish roots absorb soup, resulting in good taste. There are various types of radish pickles in Japan, e.g., takuanzuke, bettarazuke, asazuke, and fukujinzuke, but consumption of radish pickles has decreased. In Korea, Asian big radish is used as a material of kimuchi.
Radish sprouts, called “kaiware-daikon,” are consumed with sushi. Consumption of kaiware-daikon has decreased in Japan after the incident of bacteria O157 tainting of kaiware-daikon in 1996. As fresh green vegetables, only kaiware-daikon, cucumber, and labiate leaves are served with sushi. The international popularity of sushi may result in increased consumption of kaiware-daikon. As in radish roots, kaiware-daikon contains 4MTB-GSL, a major glucosinolate (Barillari et al. 2005; Papi et al. 2008).
Like Brassica seed oil, radish seed oil contains a high amount of erucic acid , about 30–40% (Ahuja et al. 1987), which is considered to be harmful as a food. Erucic acid is known to be suitable for industrial use, such as for use as a lubricant. It is promoted to make hair velvety, and radish seed oil is marketed as cosmetics. Residue of oil extraction can be used for extraction of glucosinolates . The major glucosinolate in radish seeds has been reported to be glucoraphenin, which is hydrolyzed into sulforaphene (Barillari et al. 2005). Sulforaphene has been indicated to have 1.3–1.5 times higher activity to inhibit mutagenesis induced by cooked food mutagens than sulforaphane (Shishu 2009).
Availability of radish as a winter cover crop to avoid soil erosion and loss of soil nitrogen has been reported (Weil and Kremen 2007). Radish can alleviate soil compaction by drilling with thick roots, which are killed by cold temperature, resulting in holes. Radish can suppress weeds more effectively than other cover crops, e.g., rye and rape. Furthermore, cover crops of Brassica can reduce soil nematodes (Weil and Kremen 2007), and radish is also considered to be effective in control of nematodes.
1.6 Breeding of Radish and Use of Radish as a Breeding Material
Most modern cultivars of radish are hybrid cultivars. Old cultivars and landraces are mass selected cultivars having some genetic variations within a cultivar. Since inbreeding depression is remarkable, efforts to enhance uniformity of a cultivar result in decrease of plant vigor. As a root vegetable cultivar, uniformity of root shape and color is necessary, but some variations in leaf shape and other traits can be allowed. To develop a cultivar having high uniformity, hybrid breeding has been used (see Chap. 15). The parents of a hybrid cultivar are inbred lines. Although repeated selfings to develop inbred lines make plants less vigorous, some lines showing less inbreeding depression can be selected from a large number of inbred lines. These inbred lines are maintained and propagated by bud pollination by hand. Since the number of seeds obtained by one time of the hand pollination is small, double cross or three-way cross is preferred to single cross , which can provide more uniform progeny than double cross .
In the production of hybrid seeds, self-incompatibility or male sterility is used for avoiding self-fertilization. Self-incompatibility is sometimes unstable under some stress conditions, e.g., high temperature, and contamination of hybrid seeds by selfed seeds may increase. Since such contamination spoils the quality of hybrid seeds, examination of contaminated seeds using isozyme or DNA markers is indispensable. To reduce the percentage of selfed seeds, selection of parental lines having strong and stable self-incompatibility is required. The strength of self-incompatibility depends on S haplotypes . Therefore, a simple technique for S haplotype identification is required. The methods developed for identification of S haplotypes in B. rapa and B. oleracea can also be used in radish (Nishio et al. 1996; Sakamoto et al. 1998; Niikura and Matsuura 1998; Lim et al. 2002).
Although it is difficult to use male sterility in hybrid seed production by double cross , risk of contamination of selfed seeds in single cross or three-way cross is lower in the use of male sterility than that in the use of self-incompatibility. Furthermore, restoration of pollen fertility in male-sterile lines under some growing conditions, e.g., high temperature, can be easily noticed by visual inspection of plants. The source of the well-known cytoplasmic male sterility used in rapeseed breeding is radish, i.e., Ogura cytoplasm (see Chap. 7). The same cytoplasm has been found in landraces in radish, R. sativus var. raphanistroides , and R. raphanistrum (Yamagishi and Terachi 1996). Although a restorer gene is not necessary in hybrid breeding of root vegetables, a gene useful in the breeding of rapeseed, possibly also of rat’s tail radish, has been identified (Koizuka et al. 2003).
Mutation breeding of radish has not been successfully performed because it is difficult to distinguish induced mutations and variations present in radish cultivars. Furthermore, most mutated traits are controlled by recessive alleles and are not suitable for hybrid breeding, which combines dominant alleles of parents. However, all the landraces having characteristic traits have been developed by spontaneous mutations. Recently, a cultivar having a new composition of glucosinolate has been developed by selecting a spontaneous mutation in a landrace (Ishida et al. 2015). Techniques for reverse-genetic selection of mutants have been developed. Although TILLING (McCallum et al. 2000), which has been successfully used for reverse-genetic selection of mutants in many plant species, is not suitable for use in radish cultivars, which has many gene variations in a cultivar, mutant selection using a next-generation sequencer, e.g., KeyPoint ® of Keygene, may be available for use in mutation breeding of radish. Another cost-effective technique for reverse-genetic selection is required.
Because of the difficulty of plant regeneration, genetic engineering has not been used in radish breeding. However, many GMO cultivars have been developed in soybean, which is also recalcitrant in tissue and cell culture. The reason why such efforts have not been made in radish may be lower economic value of radish than that of soybean. Another problem of using genetic engineering in radish breeding may be wide distribution of wild radish in the world. Stress tolerance genes and insect resistance genes can be transmitted to wild plants, providing higher competitive ability. Since flowering is not necessary in root vegetable production, the use of a sterility gene, e.g., pollen sterility gene used in the seed production technology (DuPont-Pioneer), may be useful.
Intergeneric hybrids between R. sativus (as a female parent) and B. oleracea, named Raphanobrassica (Karpechenko 1924), and between B. rapa (as a female parent) and R. sativus, named Brassicoraphanus (Terasawa 1932), have long been studied (see Chap. 14). Efficiency of hybrid production without embryo culture between B. rapa and R. sativus depends on cultivars of B. rapa, and three QTLs controlling hybrid formation ability have been identified (Tonosaki et al. 2013). Crossability of R. sativus with Brassica suggests that Brassica genetic resources may be available for use in radish breeding. Since R. sativus has valuable traits for Brassica breeding, e.g., disease resistance (Akaba et al. 2009), nematode resistance (Lelivelt et al. 1993), stress tolerance, and a different composition of glucosinolates , the economic value of radish as a breeding material of several Brassica crop species can be emphasized.
References
Ahuja KL, Singh H, Raheja RK, Labana KS (1987) The oil content and fatty acid composition of various genotypes of cauliflower, turnip and radish. Plant Foods Hum Nutr 37:33–40
Akaba M, Kaneko Y, Hatakeyama K, Ishida M, Bang SW, Matsuzawa Y (2009) Identification and evaluation of clubroot resistance of radish chromosome using a Brassica napus–Raphanus sativus monosomic addition line. Breed Sci 59:203–206
Arumuganathan K, Earle ED (1991) Nuclear DNA content of some important species. Plant Mol Biol Rep 9:208–218
Barillari J, Cervellati R, Paolini M, Tatibouet A, Rollin P, Iori R (2005) Isolation of 4-methylthio-3-butenyl glucosinolate from Raphanus sativus sprouts (kaiware daikon) and its redox properties. J Agric Food Chem 53:9890–9896
Bennetzen J, Freeling M (1997) The unified grass genome: synergy in synteny. Genome Res 7:301–306
Cho MA, Min SR, Ko SM, Liu JR, Choi PS (2008) Agrobacterium-mediated genetic transformation of radish (Raphanus sativus L.). Plant Biotechnol 25:205–208
Gomez-Campo C (1980) Morphology and morpho-taxonomy of the tribe Brassiceae. In: Tsunoda S et al (eds) Brassca crops and wild allies. Japan Scientific Societies Press, Tokyo, pp 3–31
Harberd DJ, McArthur ED (1980) Meiotic analysis of some species and genus hybrids in the Brassiceae. In: Tsunoda S et al (eds) Brassca crops and wild allies. Japan Scientific Societies Press, Tokyo, pp 65–87
Hashida T, Nakatsuji R, Budahn H, Schrader O, Peterka H, Fujimura T, Kubo N, Hirai M (2013) Construction of a chromosome-assigned, sequence-tagged linkage map for the radish, Raphanus sativus L. and QTL analysis of morphological traits. Breed Sci 63:218–226
Inaba R, Nishio T (2002) Phylogenetic analysis of Brassiceae based on the nucleotide sequences of the S-locus related gene, SLR1. Theor Appl Genet 105:1159–1165
Ishida M, Kakizaki T, Morimitsu Y, Ohara T, Hatakeyama K, Yoshiaki H, Kohori J, Nishio T (2015) Novel glucosinolate composition lacking 4-methylthio-3-butenyl glucosinolate in Japanese white radish (Raphanus sativus L.). Thoer Appl Genet 128:2037–2046
Jeong Y-M, Kim N, Ahn B, Oh M, Chung W-H, Chung H, Jeong S, Lim K-B, Hwang Y-J, Kim G-B, Baek S, Choi S-B, Hyung D-J, Lee S-W, Sohn S-H, Kwon S-J, Jin M, Seol Y-J, Chae W, Choi K, Park B-S, Yu H-J, Mun J-H (2016) Elucidating the triplicated ancestral genome structure of radish based on chromosome-level comparison with the Brassica genomes. Theor Appl Genet 129:1357–1372
Johnston JS, Pepper AE, Hall AE, Chen ZJ, Hodnett G, Drabek J, Lopez R, Price HJ (2005) Evolution of genome size in Brassicaceae. Ann Bot 95:229–235
Kakizaki T, Kitashiba H, Zou Z, Li F, Fukino N, Ohara T, Nishio T, Ishida M (2017) A 2-oxoglutarate-dependent dioxygenase mediates the biosynthesis of glucoraphasatin in radish. Plant Physiol 173:1583–1593
Karpechenko GD (1924) Hybrids of Raphanus sativus L. x Brassica oleracea L. J Genet 14:375–396
Kamei A, Tsuro M, Kubo N, Hayashi T, Wang N, Fujimura T, Hirai M (2010) QTL mapping of clubroot resistance in radish (Raphanus sativus L.). Theor Appl Genet 120:1021–1027
Kitashiba H, Li F, Hirakawa H, Kawanabe T, Zou Z, Hasegawa Y, Tonosaki K, Shirasawa S, Fukushima A, Yokoi S, Takahata Y, Kakizaki T, Ishida M, Okamoto S, Sakamoto K, Shirasawa K, Tabata S, Nishio T (2014) Draft sequences of the radish (Raphanus sativus L.) genome. DNA Res 21:481–490
Koizuka N, Imai R, Fujimoto H, Hayakawa T, Kimura Y, Kohno-Murase J, Sakai T, Kawasaki S, Imamura J (2003) Genetic characterization of a pentatricopeptide repeat protein gene, orf587, that restores fertility in the cytoplasmic male-sterile Kosena radish. Plant J 34:407–415
Lelivelt CLC, Lang W, Dolstra O (1993) Intergeneric crosses for the transfer of resistance to the beet cyst nematode from Raphanus sativus to Brassica napus. Euphytica 58:111–120
Li F, Hasegawa Y, Saito M, Shirasawa S, Fukushima A, Ito T, Fujii H, Kishitani S, Kitashiba H, Nishio T (2011) Extensive chromosome homoeology among Brassiceae species were revealed by comparative genetic mapping with high-density EST-based SNP markers in radish (Raphanus sativus L.). DNA Res 18:401–411
Lim SH, Cho HJ, Lee SJ, Cho YH, Kim BD (2002) Identification and classification of S haplotypes in Raphanus sativus by PCR-RFLP of the S locus glycoprotein (SLG) gene and the S locus receptor kinase (SRK) gene. Theor Appl Genet 104:1253–1262
McCallum CM, Comai L, Greene EA, Henikoff S (2000) Targeted screening for induced mutations. Nat Biotechnol 18:455–457
Mitsui Y, Shimomura M, Komatsu K, Namiki N, Shibata-Hatta M, Imai M, Katayose Y, Mukai Y, Kanamori H, Kurita K, Kagami T, Wakatsuki A, Ohyanagi H, Ikawa H, Minaka N, Nakagawa K, Shiwa Y, Sasaki T (2015) The radish genome and comprehensive gene expression profile of tuberous root formation and development. Sci Rep 5:10835
Mizushima U (1980) Genome analysis in Brasica and allied genera. In: Tsunoda S et al (eds) Brassca crops and wild allies. Japan Scientific Societies Press, Tokyo, pp 89–106
Mun J-H, Chung H, Chung W-H, Oh M, Jeong Y-M, Kim N, Ahn BO, Park B-S, Park S, Lim K-B, Hwang Y-J, Yu H-J (2015) Construction of a reference genetic map of Raphanus sativus based on genotyping by whole-genome resequencing. Theor Appl Genet 128:259–272
Nasu S, Kitashiba H, Nishio T (2012) “Na-no-hana Project” for recovery from the Tsunami disaster by producing salinity-tolerant oilseed rape lines: Selection of salinity-tolerant lines of Brassica crops. J Integr Field Sci 9:33–37
Niikura S, Matsuura S (1998) Identification of self-incompatibility alleles (S) by PCR-RFLP in radish (Raphanus sativus L.). Euphytica 102:379–384
Niikura S, Matsuura S (2000) Genetic analysis of the reaction level of self-incompatibility to a 4% CO2 gas treatment in the radish (Raphanus sativus L.). Theor Appl Genet 101:1189–1193
Nishio T, Kusaba M, Watanabe M, Hinata K (1996) Registration of S alleles in Brassica campestris L by the restriction fragment sizes of SLGs. Theor Appl Genet 92:388–394
Okamoto S, Sato Y, Sakamoto K, Nishio T (2004) Distribution of similar self-incompatibility (S) haplotypes in different genera, Raphanus and Brassica. Sex Plant Reprod 17:33–39
Ozeki Y (2010) Study on the relationship between the anthocyanin molecular species and color phenotype of roots in the inbred lines of red radish (Raphanus sativus L.). Japan Food Chem Res Found Res Rep 16:33–39 (in Japanese with English summary)
Papi A, Orlandi M, Bartolini G, Barillari J, Iori R, Paolini M, Ferroni F, Fumo MG, Pedulli GF, Velgimigli L (2008) Cytotoxic and antioxidant activity of 4-methylthio-3-butenyl isothiocyanate from Raphanus sativus L. (kaiware daikon) sprouts. J Agric Food Chem 56:875–883
Park B-J, Liu Z, Kanno A, Kameya T (2005) Transformation of radish (Raphanus sativus L.) via sonication and vacuum infiltration of germinated seeds with Agrobacterium harboring a group 3 LEA gene from B. napus. Plant Cell Rep 24:494–500
Park NI, Xu H, Li X, Jang IH, Park S, Ahn GH, Lim YP, Kim SJ, Park SU (2011) Anthocyanin accumulation and expression of anthocyanin biosynthetic genes in radish (Raphanus sativus). J Agric Food Chem 59:6034–6039
Peters SA, Bargsten JW, Szinay D, van de Belt J, Visser RGF, Bai Y, de Jong H (2012) Structural homology in the Solanaceae: analysis of genomic regions in support of synteny studies in tomato, potato and pepper. Plant J 71:602–614
Sakamoto K, Kusaba M, Nishio T (1998) Polymorphism of the S-locus glycoprotein gene (SLG) and the S-locus related gene (SLR1) in Raphanus sativus L. and self-incompatible ornamental plants in the Brassicaceae. Mol Gen Genet 258:397–403
Shen D, Shu H, Huang M, Zheng Y, Li X, Fei Z (2013) RadishBase: a database for genomics and genetics of radish. Plant Cell Physiol 54(1–6):e3
Shirasawa K, Oyama M, Hirakawa H, Sato S, Tabata S, Fujioka T, Kimizuka-Takagi C, Sasamoto S, Watanabe A, Kato M, Kishida Y, Kohara M, Takahashi C, Tsuruoka H, Wada T, Sakai T, Isobe S (2011) An EST-SSR linkage map of Raphanus sativus and comparative genomics of the Brassicaceae. DNA Res 18:221–232
Shishu KIP (2009) Inhibition of cooked food-induced mutagenesis by dietary constituents: comparison of two natural isothiocyanates. Food Chem 112:977–981
Takahata Y, Komatsu H, Kaizuma N (1996) Microspore culture of radish (Raphanus sativus L.): influence of genotype and culture conditions on embryogenesis. Plant Cell Rep 16:163–166
Terasawa Y (1932) Tetraploid Bastarde von Brassica chinensis L. x Raphanus sativus L. Jpn J Genet 7:183–185 (In Japanese)
Tonosaki K, Michiba K, Bang SW, Kitashiba H, Kaneko Y, Nishio T (2013) Genetic analysis of hybrid seed formation ability of Brassica rapa in intergeneric crossings with Raphanus sativus. Theor Appl Genet 126:837–846
Tsuro M, Suwabe K, Kubo N, Matsumoto S, Hirai M (2008) Mapping of QTLs controlling root shape and red pigmentation in radish, Raphanus sativus L. Breed Sci 58:55–61
Warwick SI, Black LD (1997) Phylogenetic implications of chloroplast DNA restriction site variation in subtribes Raphaninae and Cakilinae (Brassicaceae, tribe Brassiceae). Can J Bot 75:960–973
Warwick SI, Hall JC (2009) Phylogeny of Brassica and wild relatives. In: Gupta SK (ed) Biology and breeding of crucifers. CRC Press, London, pp 19–36
Warwick SI, Sauder CA (2005) Phylogeny of tribe Brassiceae (Brassicaceae) based on chloroplast restriction site polymorphisms and nuclear ribosomal internal transcribed spacer and chloroplast trnL intron sequences. Can J Bot 83:467–483
Weil RR, Kremen A (2007) Thinking across and beyond disciplines to make cover crops pay. J Sci Food Agric 87:551–557
Yamagishi H, Terachi T (1996) Molecular and biological studies on male-sterile cytoplasm in the Cruciferae III. Distributtion of Ogura-type cytoplasm among Japanese wild radishes and Asian radish cultivars. Theor Appl Genet 93:325–332
Zou Z, Ishida M, Li F, Kakizaki T, Suzuki S, Kitashiba H, Nishio T (2013) QTL analysis using SNP markers developed by next-generation sequencing for identification of candidate genes controlling 4-methylthio-3-butenyl glucosinolate contents in roots of radish, Raphanus sativus L. PLoS ONE 8:e53541
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Nishio, T. (2017). Economic and Academic Importance of Radish. In: Nishio, T., Kitashiba, H. (eds) The Radish Genome. Compendium of Plant Genomes. Springer, Cham. https://doi.org/10.1007/978-3-319-59253-4_1
Download citation
DOI: https://doi.org/10.1007/978-3-319-59253-4_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-59252-7
Online ISBN: 978-3-319-59253-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)