Abstract
Morphological studies on brains have recently entered a new phase of circuit analysis identified under the newly designated area of connectomics, the study of brain wiring diagrams exact at synapse level that can now be produced by means of electron microscopy and automated reconstruction. The most comprehensive examples come from the brains of invertebrates with few neurons, which Nature provides in great abundance especially among marine larval invertebrates. Two complete examples, the nematode C. elegans and the larva of the ascidian Ciona intestinalis, are now published; others are in the pipeline. Each species has its advantages and champions, especially clearly so in Drosophila, which offers outstanding opportunities for functional analysis of complex behaviours using genetics-based methods. Collectively‚ these offer an ultimate prospect for the causal analysis of behaviour. In addition, the availability of multiple connectomes from behaviourally different species will reveal features of the network design that are common to all, and that enable comparison with networks from different levels of biological organization, as well as with those from networks that have evolved from human technologies.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Powerful Genetic Methods
- Meinertzhagen
- Connectome Data
- Connectome Studies
- Aversive Olfactory Conditioning
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
Morphological studies on brains have recently entered a new phase of circuit analysis identified under the newly designated area of connectomics, the study of exact brain wiring diagrams (Lichtman and Sanes 2008). A connectome is of course no new idea, merely one now finally enabled in select systems of neurons by efficient digital imaging and computer-aided reconstruction tools. Additionally, it relies on an enhanced range of electron imaging methods, including scanning block-face microscopy, SBFM (Denk and Heinz 2004) and focused ion beam milling (FIB)-SEM (Knott et al. 2008, 2011) as well as more traditional serial-section EM, ssEM methods (e.g. Fahrenbach 1984; Hall 1995; Harris et al. 2006), that all have sufficient resolution to identify clearly the synaptic contacts formed between identified neurons. Neuron identification in those cases is therefore usually enabled by 3-D reconstruction of neuron shapes, and comparisons between these and profiles from light microscopy. Obvious as it may be to some, not all are convinced by the approach, even when the contra arguments of its detractors have been robustly countered (Morgan and Lichtman 2013).
The history of connectome reports is rather sparsely populated by studies, most of which have taken advantage of small brains with few neurons. This epilogue summarizes such studies, highlighting ways in which invertebrate brains serve as a foundation for connectome studies, in parallel with alternative approaches on vertebrate brains, especially in the retina (e.g. Helmstaedter et al. 2013; Ding et al. 2016). Without doubt, pride of place among invertebrates goes to the first complete connectome report from Caenorhabditis elegans (White et al. 1986). But even this had its own forerunner, in the subway maps of Ascaris neurons published by Richard Goldschmidt nearly 80 years earlier (1908, 1909), and cleverly avoided patterns of rewiring in species with entirely different feeding habits (Bumbarger et al. 2013). As for genomes, the importance attributed to the connectome by its chief architect, Sydney Brenner, lies in it being complete, accurate and permanent, insofar as these absolutes can be determined. The initial version of the connectome (White et al. 1986) has in fact undergone some steps in its completion but these are very minor (Varshney et al. 2011), while comparisons between the reconstructions of two animals have demonstrated features of its accuracy both in the hermaphrodite (Durbin 1987), and in another comparison, with the posterior CNS of the adult male (Jarrell et al. 2012). Major early contributions on different systems were also made using photographic imaging, especially by the group of Cyrus Levinthal and colleagues at Columbia University, who pioneered in the development of approaches to automate serial image capture using pre-digital methods (e.g. Ware and LoPresti 1975), and examined amongst other systems, the entire nervous system of the rotifer Asplanchna brightwelli (Ware 1971) and the visual system of the crustacean Daphnia magna (Macagno et al. 1973).
The rest of the gang. The diminutive brains of invertebrates, especially those of larval stages with few cells (Meinertzhagen 2016a), are part of Nature’s bounty for connectomic analyses. Marine invertebrate larvae, many with essential virtues of small size, suitable for EM, and complete transparency, suitable for LM, are rich in opportunity, but generally poor in the functional studies they enable. The brains of larval polychaetes, which show considerable diversity, have some of the smallest numbers and are especially well suited to connectomic studies. For example, the 48-h trochophore of the polychaete Spirobranchus with only ~36 neurons (Lacalli 1984), compares to the CNS of dwarf male Dinophilus gyrociliatus with a total of 68 neurons (Windoffer and Westheide 1988), and the larva of Platyneiris dumerilii, with 71 neurons for visual navigation (Randel et al. 2014, 2015); while the rotifer CNS has 196 brain cells in Asplanchna (Ware 1971). These are to be compared with the tadpole larva of the tunicate Ciona intestinalis, a basal deuterostome with ~330 cells of which 177 are neurons with 6618 synapses (Ryan et al. 2016), and the nematode C. elegans with 302 neurons and 6393 synapses in the hermaphrodite (White et al. 1986), for both of which an entire connectome has been completed. In addition, there is the immediate prospect of the entire connectome for the CNS of the first-instar larva of the fruit fly Drosophila melanogaster, for which excerpts have appeared (e.g. Ohyama et al. 2015).
To identify behavioural function in a synaptic circuit, neuron by neuron, targeting transgenes that either disable neuron function or selectively restore it in null mutants is possible in Drosophila (Simpson 2009; Meinertzhagen and Lee 2012) and other genetic models, but targeted single-cell photoablations are also possible in some other species.
Three overlapping motivations have mostly driven work in this area: first, the analysis of the neural substrate for interesting behaviours; second, the availability of powerful genetic methods that enable the targeting and functional analyses of identified neurons; and third, the attraction of simple nervous systems having few, morphologically simple cells favouring the construction of a complete connectome. Some species may indulge us in two of these three, but none has them all. For example, Drosophila has both powerful genetic methods (e.g. Luan et al. 2006; Pfeiffer et al. 2010; Jenett et al. 2012) and interesting behaviours, especially for vision (e.g. Heisenberg and Wolf 1984; Borst 2009; Silies et al. 2014), but its neurons branch profusely to yield many slender neurites that require special methods to reconstruct at EM level (e.g. Feng et al. 2015), albeit still relatively slowly and somewhat incompletely. The nematode C. elegans has both powerful genetic methods and a landmark connectome, made possible by the simple tubular features of its neurons, but its behavioural repertoire is less dynamic, compelling and completely analysed, especially in the depth dimension. It is difficult to know how to weight these three requirements, and whether the relative ease of 3-D reconstruction in, for example C. elegans or C. intestinalis, might eventually outweigh the disadvantages of either. For the moment, the prize goes to the construction of a single connectome in each species, whereas the eventual need is to reconstruct many, in order to seek their common features.
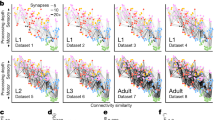
The brain as a network of networks. In addition to whole-brain connectome studies, considerable progress has been made in documenting smaller brain regions containing behaviourally interesting pathways. Early work on the connectome of individual neuropiles started most obviously with work on the first optic neuropile, or lamina, in various arthropods—notably the branchiopod crustacean D. magna (Macagno et al. 1973), the horseshoe crab Limulus polyphemus (Fahrenbach 1985), the housefly Musca domestica (Strausfeld and Campos-Ortega 1977) and fruit fly D. melanogaster (Meinertzhagen and O’Neil 1991; Meinertzhagen and Sorra 2001). These all capitalized on the parallel pathways endowed by compound eyes, and captured what was possible with limited ssEM. More recently, Randel et al. (2014) have reported the four-eye, 71-neuron sensory-motor circuits for visual navigation in Platyneiris that are comprehensively organized into circuits with 1106 connections. These have a high level of stereotypy between individuals (Randel et al. 2015).
Drosophila is today’s star. Contemporary approaches to connectomics may show few bounds. Yet, even the tiny brain of Drosophila is too complex for us to imagine fully. In parallel with EM studies above, light microscopy in flies provides accounts for the cells and circuits that underlie visual behaviour (Musca: Douglass and Strausfeld 2003; Strausfeld and Nässel 1980; and Drosophila: Meinertzhagen 2014), for which there is a large body of quantitative behavioural studies (Heisenberg and Wolf 1984; Borst and Egelhaaf 1989; Borst and Haag 2002). The objectives at EM level may even be powerfully aided by wiring networks derived from light microscopy in Drosophila (Chiang et al. 2011; Shih et al. 2015). These are enabled in turn by new tools for neuroanatomy and neurogenetics (Pfeiffer et al. 2008), and 3-D registration (Peng et al. 2011). Thus, light microscopy identifies 58 tracts between 41 neuropiles and 6 hubs in Drosophila and a total of 16,000 neuron classes that fasciculate consistently within these (Chiang et al. 2011), upgraded to 43 local processing units (Shih et al. 2015). The individual connectomes for each of these can only be assembled piecemeal, in an initial strategy of divide and conquer followed by one of unification. In their entirety these obviously contribute unimaginable complexity, and the whole is only more manageable when each is considered apart from the others. Examples include not only the optic neuropiles (Takemura et al. 2008, 2013, 2015, 2017a; Shinomiya et al. 2014), but also the antennal lobe (Rybak et al. 2016; Tobin et al. 2017), mushroom body—alpha lobe (Takemura et al. 2017b) and calyx (Butcher et al. 2012). The FlyEM team at the HHMI Janelia Research Campus in fact proposes to complete the Drosophila connectome in exactly this manner, neuropile by neuropile, from 20 µm slices of an entire brain cut on a hot diamond knife (Hayworth et al. 2015) each then individually imaged by FIB-SEM, and the consecutive slices merged.
Simpler yet, but nevertheless still daunting, the CNS of a first-instar larval Drosophila is another candidate system for which detailed synaptic networks from specific systems are now available. The functional features of various anatomical connectomic circuits have received attention, for example for action selection in rolling behaviour (Ohyama et al. 2015), or the competitive interactions among neurons that mediate choice behaviour. Thus, selecting one behaviour over another, behavioural choice is mediated by reciprocally connected feedforward inhibitory interneurons, while maintaining a chosen behaviour is mediated by feedback disinhibition; sequence transitions are mediated by lateral disinhibition (Jovanic et al. 2016).
All Drosophila brain regions reconstructed to date reveal an unsuspected complexity of synaptic circuits. These are obvious in the adult fly, for the visual pathways in the medulla (Takemura et al. 2013, 2017a, b) and especially for the alpha 2 and 3 output lobes of the mushroom body (Takemura et al. 2017a, b), the substrate for aversive olfactory conditioning. The latter incorporate: direct connections from dopaminergic neurons (DAN) to the output neurons; universal anatomical connection of Kenyon cells (KC) to output neurons (whereas electrophysiology had indicated that only 30% are functionally connected); and KC to DAN connections. Once revealed anatomically, the functions of these and the many other connections in the connectome can of course be investigated using genetic, behavioural, and electrophysiological approaches, and it is these that may be remembered by posterity. But the fact that, despite intense behavioural investigations in the field of olfactory learning, circuits that had been neither detected by light microscopy nor proposed in any model of learning were revealed only by EM (Takemura et al. 2017a), illustrates clearly the predictive value of EM connectomics as an enabling technology, in revealing the entire envelope of possible behaviours, each in need of functional validation.
No less powerful, an even simpler case had in fact already been revealed, for UV phototaxis pathways in Drosophila (Gao et al. 2008). The history of their discovery illustrates the predictive power of connectomics. Initially, Gal4 reagents identified a rather nondescript distal amacrine cell in the medulla, called Dm8 (Fischbach and Dittrich 1989), and early ssEM identified that it received synaptic inputs from the UV-signalling photoreceptor neuron R7. These synapses were identified largely because Dm8’s neurites are somewhat coarse and could be reconstructed from EM with relative ease (Gao et al. 2008). Based only on this rather slender evidence for UV sensitivity, subsequent behavioural tests of UV phototaxis revealed that Dm8 neurons were in fact both necessary and sufficient for flies to exhibit phototaxis towards UV, in preference to green light (Gao et al. 2008). Thus, even though UV phototaxis is a low-level visual behaviour and Dm8 its rather nondescript substrate, this analysis provided a very clear precedent for the predictive power of connectomics in assigning an essential role to an identified neuron, in this case Dm8.
Comparing connectomes. The connectomes of different species and systems provide a rich data source for comparative computational studies. We need always to bear in mind that brains are the product of harsh behavioural selection over millions of years, and that evolutionary relationships, uncertain as these may be, are essential to interpreting connectome organization in ways not revealed by single-species connectomes (Hale 2014). Key to that organization is, of course, how the network functions.
The two comprehensive connectomes now available, the one in a widely analysed adult hermaphrodite C. elegans (White et al. 1986), a protostome, and the other in an entirely different deuterostome, the larva of C. intestinalis (Ryan et al. 2016), exhibit network similarities. Both comprise relatively few neurons and not surprisingly neither is a random nor a regular network. Instead, both constitute so-called ‘small-world’ networks, characterized by highly connected local sub-networks linked by fewer long-range connections (Watts and Strogatz 1998; Bassett and Bullmore 2006; Bezares-Calderón and Jékely 2016). These mirror many human-constructed networks, from power grids to social media (Watts and Strogatz 1998), and offer an intuitive appeal in their economy, supporting high levels of dynamic complexity while tending to minimize wiring costs (Bassett and Bullmore 2006). Those costs are rarely specified or quantified within a network, but have received treatment (Sterling and Laughlin 2015) in energetic rather than morphogenetic terms.
Network motifs, simple building blocks of connected elements, offer a further element of commonality among many forms of network. Recognition that network motifs can be identified not only in biological systems that range from food chains to genetic networks, but also in engineering and information-processing networks (Milo et al. 2002), provided an important insight to the interpretation of an entire connectome network. Thus, specific two-, three- and four-element motifs, especially—among the latter—bi-fan and bi-parallel patterns of interconnection, occur with a significantly higher frequency in the actual synaptic networks of the C. elegans connectome, than in matched randomized networks having the same single-node features (Milo et al. 2002). This preference suggests that such motifs confer special functional properties, yet the physiology of the motifs and their elements is rarely if ever known and unlikely to be the same in all cases. Of course, this lack merely emphasizes the need for functional studies that can indicate the operational features of individual motifs, in order to identify and define classes of networks and network homologies more closely. Connectomic data also highlight higher-order networks, for example, in the CNS of larval Drosophila (Jovanic et al. 2016; Schneider-Mizell et al. 2016), where they identify a particular role for disinhibition. Thus, interneurons involved in promoting distinct behaviours via disinhibition have extensive reciprocal connections (Jovanic et al. 2016). In a different example, among the visuomotor circuits of the polychaete Platyneiris, the reciprocal wiring reasoned to provide strong mutual inhibition between the circuits of the two eyes is interpreted to represent a network motif that enhances contrast detection (Randel et al. 2014). Essential knowledge of the neurotransmitter used by each neuron, and—even more important—the neurotransmitter receptors expressed by its postsynaptic partners, is required for all such interpretations, and represents the next stage in any functional analysis.
Opinions may differ on the relative utility of computational approaches to connectome networks, and few would deny the value of lower level information on the anatomy and physiology of synaptic circuits. The latter would at least help us identify synaptic pathway strengths. In some cases, synapse numbers may give a sufficient first impression, but where synapses vary in size the aggregate synapse size is also important in determining the area over which vesicle shedding and postsynaptic signalling can occur. Moreover, the processing depth of each element of a circuit is also crucial, because it identifies not only the synaptic gain resulting from the quantitative strength of its connection, but also the gain that the element inherits from, or is offset by, its upstream partners.
Finally, no connectome is fixed. Even when completed comprehensively, synaptic partnerships and their connection strengths may change. Thus the literature on morphological plasticity in fly synaptic circuits (Meinertzhagen 2001, 2016b) reveals that there are many forms of plasticity among circuits that may appear fixed, but are evidently plastic. Such changes often go unacknowledged in the nervous systems of flies, which are generally assumed to have a fixed structural phenotype. In fact the fixity of synaptic circuits is at least partly the outcome of fixity in the conditions under which flies are usually raised in laboratory culture. More than this, the boundaries of connectome data are in many cases set by the developmental stage of the individual, especially in larval forms that are undergoing morphogenetic change. Even so, to the extent that the overall network remains unchanged, especially in Drosophila, such data will have archival as well as heuristic value, provided they can be made available in some accessible way for future generations. A related problem is therefore how to store such data, which rely on multi-terabyte datasets and need tools for rapid access and manipulability, needs and considerations that will not be treated in this perspective.
References
Bassett DS, Bullmore E (2006) Small-world brain networks. Neuroscientist 12:512–523
Bezares-Calderón LA, Jékely G (2016) Think small. eLIFE 5. pii: e22497. doi:10.7554/eLife.22497
Borst A (2009) Drosophila’s view on insect vision. Curr Biol 19:R36–R47
Borst A, Egelhaaf M (1989) Principles of visual motion detection. Trends Neurosci 12:297–306
Borst A, Haag J (2002) Neural networks in the cockpit of the fly. J Comp Physiol A Neuroethol Sens Neural Behav Physiol 188:419–437
Bumbarger DJ, Riebesell M, Rödelsperger C, Sommer RJ (2013) System-wide rewiring underlies behavioral differences in predatory and bacterial-feeding nematodes. Cell 152:109–119
Butcher NJ, Friedrich AB, Lu Z, Tanimoto H, Meinertzhagen IA (2012) Different classes of input and output neurons reveal new features in microglomeruli of the adult Drosophila mushroom body calyx. J Comp Neurol 520:2185–2201
Chiang AS, Lin CY, Chuang CC, Chang HM, Hsieh CH, Yeh CW, Shih CT, Wu JJ, Wang GT, Chen YC, Wu CC, Chen GY, Ching YT, Lee PC, Lin CY, Lin HH, Wu CC, Hsu HW, Huang YA, Chen JY, Chiang HJ, Lu CF, Ni RF, Yeh CY, Hwang JK (2011) Three-dimensional reconstruction of brain-wide wiring networks in Drosophila at single-cell resolution. Curr Biol 21:1–11. doi:10.1016/j.cub.2010.11.056
Denk W, Heinz H (2004) Serial block-face scanning electron microscopy to reconstruct three-dimensional tissue nanostructure. PLoS Biol 2:e329. doi:10.1371/journal.pbio.0020329
Ding H, Smith RG, Poleg-Polsky A, Diamond JS, Briggman KL (2016) Species-specific wiring for direction selectivity in the mammalian retina. Nature 535:105–110
Douglass JK, Strausfeld NJ (2003) Anatomical organization of retinotopic motion-sensitive pathways in the optic lobes of flies. Microsc Res Tech 62:132–150
Durbin RM (1987) Studies on the development and organisation of the nervous system of Caernorhabditis elegans. Doctoral thesis, University of Cambridge
Fahrenbach WH (1984) Continuous serial thin sectioning for electron microscopy. J Electron Microsc Techn 1:387–398
Fahrenbach WH (1985) Anatomical circuitry of lateral inhibition in the eye of the horseshoe crab, Limulus polyphemus. Proc R Soc Lond B 225:219–249
Feng L, Zhao T, Kim J (2015) neuTube 1.0: a new design for efficient neuron reconstruction software based on the SWC format. eNeuro 2(1). pii: ENEURO.0049-14.2014
Fischbach K-F, Dittrich APM (1989) The optic lobe of Drosophila melanogaster. I. A Golgi analysis of wild-type structure. Cell Tiss Res 258:441–475
Gao S, Takemura S-Y, Ting C-Y, Huang S, Lu Z, Luan H, Rister J, Yang M, Hong S-T, Wang JW, Odenwald W, White B, Meinertzhagen IA, Lee C-H (2008) Neural substrate of spectral discrimination in Drosophila. Neuron 60:328–342
Goldschmidt R (1908) Das Nervensystem von Ascaris lumbricoides und megalocephala, I. Z wissenschaftliche Zool 90:73–136
Goldschmidt R (1909) Das Nervensystem von Ascaris lumbricoides und megalocephala, II. Z wissenschaftliche Zool 92:306–357
Hale ME (2014) Mapping circuits beyond the models: integrating connectomics and comparative neuroscience. Neuron 83:1256–1258
Hall DH (1995) Electron microscopy and three-dimensional image reconstruction. Methods Cell Biol 48:395–436
Harris KM, Perry E, Bourne J, Feinberg M, Ostroff L, Hurlburt J (2006) Uniform serial sectioning for transmission electron microscopy. J Neurosci 26:12101–12103
Hayworth KJ, Xu CS, Lu Z, Knott GW, Fetter RD, Tapia JC, Lichtman JW, Hess HF (2015) Ultrastructurally smooth thick partitioning and volume stitching for large-scale connectomics. Nat Methods 12:319–322
Heisenberg M, Wolf R (1984) Vision in Drosophila. Springer, Berlin
Helmstaedter M, Briggman KL, Turaga SC, Jain V, Seung HS, Denk W (2013) Connectomic reconstruction of the inner plexiform layer in the mouse retina. Nature 500:168–174
Jarrell TA, Wang Y, Bloniarz AE, Brittin CA, Xu M, Thomson JN, Albertson DG, Hall DH, Emmons SW (2012) The connectome of a decision-making neural network. Science 337:437–444
Jenett A, Rubin GM, Ngo TT, Shepherd D, Murphy C, Dionne H, Pfeiffer BD, Cavallaro A, Hall D, Myers EW, Iwinski ZR, Aso Y, DePasquale GM, Enos A, Hulamm P, Lam SC, Li HH, Laverty TR, Long F, Qu L, Murphy SD, Rokicki K, Safford T, Shaw K, Simpson JH, Sowell A, Tae S, Yu Y, Zugates CT (2012) A GAL4-driver line resource for Drosophila neurobiology. Cell Rep. 2:991–1001
Jovanic T, Schneider-Mizell CM, Shao M, Masson JB, Denisov G, Fetter RD, Mensh BD, Truman JW, Cardona A, Zlatic M (2016) Competitive disinhibition mediates behavioral choice and sequences in Drosophila. Cell 167(858–870):e19. doi:10.1016/j.cell.2016.09.009
Knott G, Marchman H, Wall D, Lich B (2008) Serial section scanning electron microscopy of adult brain tissue using focused ion beam milling. J Neurosci 28:2959–2964
Knott G, Rosset S, Cantoni M (2011) Focussed ion beam milling and scanning electron microscopy of brain tissue. J Vis Exp 53:e2588. doi:10.3791/2588
Lacalli TC (1984) Structure and organization of the nervous system in the trochophore larva of Spirobranchus. Philos Trans R Soc Lond B Biol Sci 306:79–135
Lichtman JW, Sanes JR (2008) Ome sweet ome: what can the genome tell us about the connectome? Curr Opin Neurobiol 18:346–353
Luan H, Peabody NC, Vinson CR, White BH (2006) Refined spatial manipulation of neuronal function by combinatorial restriction of transgene expression. Neuron 52:425–436
Macagno ER, Lopresti V, Levinthal C (1973) Structure and development of neuronal connections in isogenic organisms: variations and similarities in the optic system of Daphnia magna. Proc Natl Acad Sci USA 70:57–61
Meinertzhagen IA (2001) Plasticity in the insect nervous system. Adv Insect Physiol 28:84–167
Meinertzhagen IA (2014) The anatomical organization of the compound eye visual system. In: Dubnau J (ed) Handbook of behavior genetics of Drosophila melanogaster, vol 1. University Press, Cambridge, pp 1–19
Meinertzhagen IA (2016a) Morphology of invertebrate neurons and synapses. In: Byrne JH (ed) Handbook of invertebrate neurobiology. Oxford University Press
Meinertzhagen IA (2016b) Connectome studies on Drosophila: a short perspective on a tiny brain. J Neurogenet 30:62–68
Meinertzhagen IA, Lee C-H (2012) The genetic analysis of functional connectomics in Drosophila. Adv Genet 80:99–151
Meinertzhagen IA, O’Neil SD (1991) Synaptic organization of columnar elements in the lamina of the wild type in Drosophila melanogaster. J Comp Neurol 305:232–263
Meinertzhagen IA, Sorra KE (2001) Synaptic organisation in the fly’s optic lamina: few cells, many synapses and divergent microcircuits. Progr Brain Res 131:53–69
Milo R, Shen-Orr S, Itzkovitz S, Kashtan N, Chklovskii D, Alon U (2002) Network motifs: simple building blocks of complex networks. Science 298:824–827
Morgan JL, Lichtman JW (2013) Why not connectomics? Nat Methods 10:494–500
Ohyama T, Schneider-Mizell CM, Fetter RD, Aleman JV, Franconville R, Rivera-Alba M, Mensh BD, Branson KM, Simpson JH, Truman JW, Cardona A, Zlatic M (2015) A multilevel multimodal circuit enhances action selection in Drosophila. Nature 520:633–639
Peng H, Chung P, Long F, Qu L, Jenett A, Seeds AM, Myers EW, Simpson JH (2011) BrainAligner: 3D registration atlases of Drosophila brains. Nat Methods 8:493–500
Pfeiffer BD, Jenett A, Hammonds AS, Ngo TT, Misra S, Murphy C, Scully A, Carlson JW, Wan KH, Laverty TR, Mungall C, Svirskas R, Kadonaga JT, Doe CQ, Eisen MB, Celniker SE, Rubin GM (2008) Tools for neuroanatomy and neurogenetics in Drosophila. P Natl Acad Sci USA 105:9715-9720
Pfeiffer BD, Ngo TT, Hibbard KL, Murphy C, Jenett A, Truman JW, Rubin GM (2010) Refinement of tools for targeted gene expression in Drosophila. Genetics 186:735–755
Randel N, Asadulina A, Bezares-Calderón LA, Verasztó C, Williams EA, Conzelmann M, Shahidi R, Jékely G (2014) Neuronal connectome of a sensory-motor circuit for visual navigation. eLIFE 3. doi:10.7554/eLife.02730
Randel N, Shahidi R, Verasztó C, Bezares-Calderón LA, Schmidt S, Jékely G (2015) Inter-individual stereotypy of the Platynereis larval visual connectome. eLIFE 4:e08069. doi:10.7554/eLife.08069
Ryan K, Lu Z, Meinertzhagen IA (2016) The CNS connectome of a tadpole larva of Ciona intestinalis highlights sidedness in the brain of a chordate sibling. eLIFE 5:e16962
Rybak J, Talarico G, Ruiz S, Arnold C, Cantera R, Hansson BS (2016) Synaptic circuitry of identified neurons in the antennal lobe of Drosophila melanogaster. J Comp Neurol 524:1920–1956
Schneider-Mizell CM, Gerhard S, Longair M, Kazimiers T, Li F, Zwart MF, Champion A, Midgley FM, Fetter RD, Saalfeld S, Cardona A (2016) Quantitative neuroanatomy for connectomics in Drosophila. eLIFE 5. pii: e12059
Shih CT, Sporns O, Yuan SL, Su TS, Lin YJ, Chuang CC, Wang TY, Lo CC, Greenspan RJ, Chiang AS (2015) Connectomics-based analysis of information flow in the Drosophila brain. Curr Biol 25:1249–1258
Shinomiya K, Karuppudurai T, Lin T-Y, Lu Z, Lee C-H, Meinertzhagen IA (2014) Candidate neural substrates for off-edge motion detection in Drosophila. Curr Biol 24:1–9
Silies M, Gohl DM, Clandinin TR (2014) Motion-detecting circuits in flies: coming into view. Ann Rev Neurosci 37:307–327
Simpson JH (2009) Mapping and manipulating neural circuits in the fly brain. Adv Genet 65:79–143
Sterling P, Laughlin S (2015) Principles of neural design. The MIT Press, London
Strausfeld NJ, Campos-Ortega JA (1977) Vision in insects: pathways underlying neural adaptation and lateral inhibition. Science 195:894–897
Strausfeld NJ, Nässel DR (1980) Neuroarchitectures serving compound eyes of Crustacea and insects. In: H Autrum (ed) Handbook of sensory physiology, vol VII/6B. Comparative physiology and evolution of vision in invertebrates. Springer, Berlin, pp 1–132
Takemura S, Lu Z, Meinertzhagen IA (2008) Synaptic circuits of the Drosophila optic lobe: the input terminals to the medulla. J Comp Neurol 509:493–513
Takemura S, Bharioke A, Lu Z, Nern A, Vitaladevuni S, Rivlin PK, Katz WT, Olbris DJ, Plaza SM, Winston P, Zhao T, Horne JA, Fetter RD, Takemura S, Blazek K, Chang L-A, Ogundeyi O, Saunders MA, Shapiro V, Sigmund C, Rubin GM, Scheffer LK, Meinertzhagen IA, Chklovskii DB (2013) A visual motion detection circuit suggested by Drosophila connectomics. Nature 500:175–181
Takemura S, Xu CS, Lu Z, Rivlin PK, Olbris DJ, Parag T, Plaza S, Zhao T, Katz WT, Umayam L, Weaver C, Hess H, Horne JA, Nunez J, Aniceto R, Chang L-A, Lauchie S, Nasca A, Ogundeyi O, Sigmund C, Takemura S, Tran J, Langille C, Le Lacheur K, McLin S, Shinomiya A, Chklovskii DB, Meinertzhagen IA, Scheffer LK (2015) Multi-column synaptic circuits and an analysis of their variations in the visual system of Drosophila. Proc Natl Acad Sci USA 112:13711–13716
Takemura S et al (2017a) EM reconstruction of α2 and α3 lobes of the mushroom body in adult Drosophila. eLIFE (submitted)
Takemura S, Nern A, Plaza S, Chklovskii DB, Scheffer LK, Rubin GM, Meinertzhagen IA (2017b) The comprehensive connectome of a neural substrate for ‘ON’ motion detection in Drosophila. Elife (under review)
Tobin W, Wilson R, Lee W-C (2017) Wiring variations that enable and constrain neural computation in a sensory microcircuit. eLIFE (submitted)
Varshney LR, Chen BL, Paniagua E, Hall DH, Chklovskii DB (2011) Structural properties of the Caenorhabditis elegans neuronal network. PLoS Comput Biol 7:e1001066
Ware R (1971) Computer aided nerve tracing in the brain of the rotifier, Asplanchna brightwelli. Ph.D. thesis, Massachusetts Institute of Technology, Boston, 213 pp
Ware RW, LoPresti V (1975) Three-dimensional reconstruction from serial sections. Int Rev Cytol 40:325–440
Watts DJ, Strogatz SH (1998) Collective dynamics of ‘small-world’ networks. Nature 393:440–442
White JG, Southgate E, Thomson JN, Brenner S (1986) The structure of the nervous system of the nematode Caenorhabditis elegans. Philos Trans R Soc Lond B Biol Sci 314:1–340
Windoffer R, Westheide W (1988) The nervous system of the male Dinophilus gyrociliatus (Polychaeta, Dinophilidae): II. Electron microscopical reconstruction of nervous anatomy and effector cells. J Comp Neurol 272:475–488
Acknowledgments
The author acknowledges various sources of support for his work summarized in this review, especially grant DIS-0000065 from the Natural Sciences and Engineering Research Council, for research on the larval nervous system of Ciona, and the FlyEM team at the Janelia Research Campus of HHMI for work on Drosophila. Dr. Kerrianne Ryan read an earlier version of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Meinertzhagen, I.A. (2017). Perspective: A New Era of Comparative Connectomics. In: Çelik, A., Wernet, M. (eds) Decoding Neural Circuit Structure and Function. Springer, Cham. https://doi.org/10.1007/978-3-319-57363-2_20
Download citation
DOI: https://doi.org/10.1007/978-3-319-57363-2_20
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-57362-5
Online ISBN: 978-3-319-57363-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)