Abstract
The endoplasmic reticulum and the other organelles of the eukaryotic cell are membrane-bound structures that carry out specialized functions. In this chapter, we discuss strategies that the cell has adopted to link and coordinate the different activities occurring within its various organelles as the cell carries out its physiological role.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
A defining feature of eukaryotic cells is the compartmentalization of various cellular functions specialized within organelles. These organelles are separated from each other by membranes and provide distinct protected environments where the cell can carry out various specialized functions at greater efficiencies by populating these structures with a unique set of proteins and lipids that can partition the required set of metabolites. On the other hand, segregation of functions poses a problem regarding substrate and metabolite exchange among membrane-bound compartments as well as intracellular communication that is essential for coordination of cellular metabolic activities. Recent developments have provided insights into the strategies employed by the cell to overcome the problem.
2 The ER
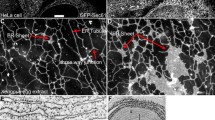
The ER is an extensive system of membranes arranged as a “net-like” network that in many cases occupies most of the interior of the cell (Fig. 4.1). This organelle houses a variety of critical ATP-requiring functions, such as maintenance of cellular homeostasis, synthesis of membrane-associated, luminal and secreted proteins, correct folding of proteins and glycoproteins, post-translational modification of proteins, lipid and steroid synthesis, Ca2+ storage and signaling, to name a few [1,2,3]. In some cellular systems, the ER provides specialized functions such as support of excitation-contraction in muscle (sarcoplasmic reticulum, SR), detoxification as well as lipid synthesis (smooth ER, SER) such as in hepatic and intestinal cells, support of protein secretory functions such as in pancreatic and liver cells (rough ER, RER), and activation of cells of the immune and nervous systems (Ca2+ signaling).
Cellular reticular network and interorganellar membrane contacts sites. The figure depicts the joining of ER membranes to cellular organelles to form the cellular reticular network from the plasma membrane to the nuclear envelope. The membrane contact sites enable rapid distribution and exchange of substrates, metabolites and signalling molecules to support proteostasis and lipidostasis. Calr calreticulin, Casq calsequestrin, Canx calnexin, PDI protein disulfide isomerase, RER rough endoplasmic reticulum, SER smooth endoplasmic reticulum
To perform these diverse functions, the ER engages a wide assortment of multifunctional integral membrane and luminal chaperones, folding enzymes and sensor molecules [4]. Chaperones are specialized proteins that assist in folding of polypeptides and in the assembly of multi-subunit proteins while folding enzymes accelerate folding process by supporting posttranslational modifications of newly synthesized proteins [5, 6]. Folding sensors play a key role in recognition of the properly folded or malfolded proteins [7]. These proteins are not only involved in ensuring the fidelity of protein folding, posttranslational modifications of newly synthesized proteins, but also contribute to storage of Ca2+ ions, facilitation of Ca2+ signaling, lipid and steroid synthesis and modification, and many other cellular roles beyond those occurring within the ER lumen [1, 8,9,10,11]. The ER also contains membrane-associated proteins that support cellular lipid and steroid synthesis [8, 9]. Importantly, ER resident chaperones and folding enzymes are major luminal ER Ca2+ binding proteins, and together with the inositol-1,4,5-trisphosphate (InsP3) receptor/Ca2+ release channel, the ER Ca2+-ATPase (SERCA) pump and ER-associated Ca2+ sensors stromal interacting molecule (STIM) proteins, are vital for Ca2+-based intracellular communication [2, 12, 13]. The ER membrane also contains an assortment of integral membrane kinases, which together with the ER-resident chaperones and folding enzymes, regulate ER stress responses allowing cellular adaptation to many challenges originating from environmental, metabolic and intrinsic demands [14, 15]. Some organelles, such as mitochondria and nucleus, are also equipped with additional specialized stress coping mechanisms [16,17,18,19,20]. These mechanisms work in concert with ER stress coping strategies to preserve or regain cellular homeostasis and prevent cellular dysfunction.
3 ER and Ca2+ Homeostasis
The ER is the main site of intracellular Ca2+ storage and physiological source of Ca2+ for intracellular signaling [2]. Disruption of Ca2+ homeostasis within the ER as well as release of Ca2+ from ER stores activate transcriptional and translational cascades that produce components involved in key pathways, such as the unfolded protein response (UPR), protein folding, ER-associated degradation (ERAD), expansion of the membrane systems as well as apoptosis [2, 12, 13]. Many of the ER proteins involved in Ca2+ binding and signaling also participate in nearly all critical ER functions [4, 21,22,23,24,25]. These proteins collectively make up the luminal ER Ca2+ stores, maintaining the total Ca2+ concentration in the ER within the μM to mM range, which is critical for the preservation of the integrity and survival of the cell. Among these ER resident Ca2+ binding proteins, calreticulin represents the major protein responsible for sequestering approximately 50% of luminal ER Ca2+ [26]. Other ER chaperones, GRP94 and GRP78 (also known as immunoglobulin binding protein or BiP) have relatively low capacity for binding Ca2+ and contribute approximately 30% of the total luminal ER Ca2+ store by virtue of their abundance [27,28,29]. The remaining balance of the luminal ER Ca2+ store is bound to proteins such as ER resident oxidoreductases, a PDI-like family of proteins [30,31,32]. In some cell types, such as muscle cells, calsequestrin is the major Ca2+ binding protein in the lumen of SR [33, 34].
4 ER-Plasma Membrane Connection: Store-Operated Calcium Entry
After the release of Ca2+ from the ER stores, the cell engages a recovery system to restore cellular Ca2+ levels to the initial state and replenish luminal ER Ca2+ stores via the action of the SERCA pump. Depletion of luminal ER Ca2+ stores leads to activation of store-operated Ca2+ entry (SOCE) mechanism [13, 35,36,37], a concept initially proposed by Putney [38]. SOCE is an excellent example of coordination of Ca2+ signaling between the ER lumen, the plasma membrane and the extracellular environment. This process involves the ER-resident Ca2+ sensors STIM proteins and plasma membrane Ca2+ channels known as Calcium Release-Activated Calcium Modulator (ORAI). STIM senses reductions in luminal ER Ca2+ stores, resulting in oligomerization of STIM and complex formation with ORAI [37, 39, 40] bringing plasma membrane Ca2+ channels close to ER membrane embedded Ca2+ pumps. This ORAI-STIM interaction effectively causes the influx of Ca2+ from the extracellular milieu, increasing cytoplasmic Ca2+ levels and subsequently refilling of the luminal ER Ca2+ stores.
In vertebrates, there are two isoforms of STIM proteins, STIM1 and STIM2 [13]. STIM1 is activated by receptor-mediated ER Ca2+ release whereas STIM2 is involved in maintaining resting ER Ca2+ concentration [41]. Both isoforms are ubiquitously expressed and share common structural features. STIM1 and STIM2 contain a single transmembrane domain and two EF-hands, a helix-loop-helix structural motif characteristic of Ca2+-binding proteins [42], which are important for the Ca2+ sensing activity. In addition to the N-terminal EF-hands, STIM proteins contain the sterile α motif (SAM) domain and three coiled-coil domains (CC1-3) at their C-terminal end, which are exposed to the cytoplasm and important for interaction with ORAI [13]. ORAIs are plasma membrane proteins with four transmembrane domains, which function as a Ca2+ channel. The crystal structure of ORAI from Drosophila melanogaster revealed that the Ca2+ channel is comprised of a hexameric assembly of ORAI subunits arranged around a central ion pore [43]. In mammalian cells, there are three isoforms of ORAI proteins, namely ORAI1, ORAI2 and ORA3. All the ORAI isoforms support functional SOCE [44]. ORAI1 and ORAI3 form multimeric Ca2+ channels that are also regulated in a Ca2+ store-independent way by lipid messengers arachidonic acid and leukotriene C4 [45,46,47]. No interaction between STIM and ORAI is apparent at resting ER Ca2+ levels, but upon depletion of the luminal ER Ca2+, STIM molecules oligomerize due to conformation changes at the CC1 domain. The STIM multimers then recruit and gate the ORAI channels at ER and plasma membranes junctions. The ER also contains a negative regulator of SOCE, known as SOCE-associated regulatory factor (SARAF), that associates with STIM to facilitate slow Ca2+-dependent inactivation thus protecting cells from Ca2+ overfilling. STIM-ORAI signaling requires additional associated proteins including CRACR2A, STIMATE, junctate, POST, and septins [13]. Luminal ER oxidoreductase PDIA3 can also control STIM function, and consequently STIM1-induced SOCE, by binding to two conserved cysteines in the luminal domain of STIM1 [48]. Additionally, PDIA3 can regulate SERCA2b, a non-muscle isoform of SERCA, enabling further dynamic control of ER Ca2+ homeostasis [49]. The actions of PDIA3 illustrate how multiple ER activities, such as protein folding and posttranslational modification, are linked with Ca2+ transport and maintenance of ER Ca2+ homeostasis.
5 ER and Protein Quality Control
Considering that over 30% of proteins are synthesized and processed in the ER, it is not surprising that the ER has evolved a sophisticated protein quality control system to preserve proteostasis [1, 3, 10, 11, 50, 51]. Proteostasis (protein homeostasis) refers to optimal folding and function of proteome [51]. Proteostasis is accomplished by an elaborate mechanism that integrates key cellular processes, namely biosynthethic and degradative pathways as well as control of gene transcription. Disrupted proteostasis is detrimental for the survival of the cell and health of the organism. The cell has a wide repertoire of molecular chaperones, which include cytoplasmic heat shock proteins (HSPs) [52], a subset of glucose regulatory proteins (GRP78/BiP and GRP94), the lectin chaperones calreticulin, calnexin, and ER degradation-enhancing α-mannosidase-like protein [53, 54]. These chaperones are specialized with respect to their substrates. For example, calreticulin, calnexin and PDIA3 are the proteins that make up the core machinery responsible for ensuring quality control of newly synthesized glycoproteins [1, 3, 10, 11, 51]. The ER-associated oxidoreductases [protein disulphide isomerases (PDIs) and peptidyl prolyl isomerases (PPIs)], oligosaccharide transferases, glucosidases and mannosidases are responsible for protein modifications such as disulfide bond formation and N-linked glycosylation [1, 50, 55,56,57]. Depletion of luminal ER Ca2+ inhibits chaperone function causing a global ER stress and disrupted proteostasis [22, 58, 59].
6 ER and Cellular Stress Coping Response Strategies
Loss of nutrients/energy homeostasis is a universal feature defining the induction of ER stress and impacts on all aspects of cellular function. Ca2+ signaling may be the principal mechanism involved in recognizing, communicating the state of cellular reticular homeostasis, and coordinating the activities to multiple corrective strategies. Mitochondria and the nucleus also developed stress responses to prevent cellular dysfunction due to stress challenges, environmental and metabolic demands [16,17,18,19,20, 60]. Activation of the UPR leads to translational attenuation to prevent synthesis of new proteins in the ER, transcriptional induction of genes encoding chaperones, folding enzymes and other proteins involved in ERAD, controlled degradation of misfolded proteins, and activation of apoptosis if the cell is unable to re-establish ER homeostasis [14]. The UPR coping strategy in the ER is thought to be comprised of three separate pathways, each controlled by distinct ER-associated integral membrane sensor proteins: the ER kinase dsRNA-activated protein kinase-like ER kinase (PERK), activating transcription factor 6 (ATF6), and inositol-requiring enzyme 1 (IRE1) embedded in the ER membrane and complexed with GRP78/BiP in the lumen of the ER [14]. Additionally, a controlled self-digestion and degradation process known as autophagy, is stimulated to help promote cell survival by eliminating damaged cellular components [8, 61, 62]. Autophagy may also provide the cell with a short-term source of raw materials (such as amino acids and fatty acids) and energy substrates [63].
Luminal ER Ca2+ binding proteins, such as GRP78/BiP, and the Ca2+-dependent microRNA miR-322 have been identified as bonafide regulators of UPR [64, 65]. Calreticulin, a major ER Ca2+ binding protein, associates with ATF6 in a carbohydrate-dependent manner, and together with GRP78/BiP (also an ER Ca2+ binding protein), maintains ATF6 in its transcriptionally inactive membrane-bound precursor state [66]. PDIA5, under stress conditions, promotes ATF6α export from the ER and activation of its target genes [67]. PDIA6 is a regulator of IRE1 activity in response to depletion of luminal ER Ca2+ stores [64, 68]. The binding of PDIA6 with the luminal domain of IRE1α in a cysteine-dependent manner has been shown to enhance IRE1α activity [64, 68]. This effect is specific for the IRE1α branch since PDIA6 has no influence on the activity of the PERK branch of the UPR pathway [64, 68]. Depletion of luminal ER Ca2+ and activation of SOCE reduces the abundance of the microRNA miR-322, which in turn regulates PDIA6 mRNA stability and consequently IRE1α activity [64]. Other microRNAs have been shown to regulate components that maintain ER homeostasis, including channels that control Ca2+ fluxes across the ER membrane [69]. The expression of these microRNAs is sensitive to changes in ER Ca2+ homeostasis [69]. The luminal ER environment (amount of Ca2+, composition of ER resident proteins) together with non-coding RNAs including microRNAs cooperate to control UPR status and to maintain homeostasis within the entire cellular reticular network [64, 69, 70].
7 ER and Lipid Metabolism
Cellular membranes are comprised of proteins as well as lipids, which include phospholipids, glycolipids and sterols [71]. The lipid constituents are not evenly distributed throughout the membrane systems of the cell. The plasma membrane and organellar membranes have characteristic lipid compositions which defines their identity. For instance, unesterified cholesterol is typically abundant in the mammalian plasma membrane and rare in ER membranes. Cardiolipin is enriched in mitochondrial membranes. Membrane leaflets may display lipid asymmetry as observed in the plasma membrane where phosphatidylserine is present in greater abundance in the inner leaflet whereas phosphatidylethanolamine is more highly represented in the outer leaflet. These lipids serve not only a structural role but also as a source of lipid metabolites that act as signaling molecules such as InsP3, diacylglycerols and others [71]. Heterologous membrane systems do not readily mix or merge largely due to the physicochemical properties imparted by their distinct lipid compositions. It is thought that fusion of dissimilar membranes requires the action of specific proteins present at the sites of membrane interactions [71, 72]. The unique lipid composition of the plasma membrane and organellar membranes may also partly determine the characteristic proteome associated with these different membrane systems by virtue of specific lipid-protein interactions that promote the retention of specific proteins in certain types of membrane environments.
Membrane lipid homeostasis, or lipidostasis, involves regulated synthesis of certain lipid species and lipid quality control to remove damaged lipids [51]. The ER is the major site of membrane lipid synthesis, although other organelles participate in producing biosynthetic intermediates, as in the case of phosphatidylserine synthesis (involves mitochondria) [73] and cholesterol synthesis (involves peroxisomes) [74, 75]. The enzymes responsible for these reactions are either embedded in membranes or located in the lumen of the organelles. Conversion of cholesterol into other bioactive compounds, such as steroid hormones, vitamin D metabolites and bile acids, also occurs in the ER in conjunction with other organelles [76, 77]. Vitamin D is known as a major regulator of systemic Ca2+ homeostasis [78] and is also important in the maintenance of cell signaling pathways [12]. The secondary bile acid known as ursodeoxycholic acid is a potent proteostasis promoter [79]. Considering the intimate relationships between specific lipids and proteins that comprise the various membrane systems of the cell, we propose that the coordination of lipidostasis and proteostasis is a key aspect necessary for assembly of functional membrane units to preserve cellular function and viability [51]. A recent study established that specific amino acid residues within the transmembrane domain of SERCA make ordered contacts with membrane phosphatidylcholine residues that surround the transmembrane domain to facilitate Ca2+ transport into the cell [80]. Disrupted lipidostasis is likely to curtail SERCA function and thus impair cellular Ca2+ homeostasis.
The induction of the UPR has been observed to induce phosphatidylcholine synthesis. This was initially attributed to the activation of the IRE1 branch of the pathway since enforced expression of Xbp1s mRNA could recapitulate the effect of UPR induction [81]. The increase in the synthesis of phosphatidylcholine supports expansion of ER membranes and is likely a part of the response to resolve ER stress. Interestingly, there were no changes in the abundance of mRNA of genes that are known to participate in the synthesis of phosphatidylcholine, suggesting that induction of membrane lipid synthesis is accomplished via posttranscriptional mechanisms. ATF6 was also found capable of stimulating phosphatidylcholine synthesis in an Xbp1s-independent fashion [82]. It is not clear how these transcription factors augment phosphatidylcholine synthesis. Nevertheless, the findings support for the notion that lipidostasis is linked to proteostasis, considering that membrane assembly is a concerted process involving both lipids and proteins.
Deletion of calreticulin dramatically reduces luminal ER Ca2+ stores, and causes the extreme elevation of blood lipids in mice and intracellular lipid accumulation in worms [83]. Sterol response element binding proteins (SREBPs) are master regulators of lipid homeostasis by regulating the expression of genes involved in cholesterol and triacylglycerol metabolism [9, 84, 85]. There are three SREBP isoforms known as SREBP-1a, SREBP-1c (both encoded by the SREBP1 gene) and SREBP-2 (encoded by the SREBP2 gene). These transcription factors are synthesized as precursor ER integral membrane proteins. Structurally, the SREBPs are composed of a transcription factor domain located in the N-terminal region of the protein, a transmembrane domain, and a regulatory domain located in the C-terminal region of the protein that interacts with another ER membrane protein known as SREBP cleavage activating protein (SCAP). SREBP processing is triggered by the reduction of unesterified cholesterol concentration in the ER membrane [86]. When unesterified cholesterol is abundant in the ER membrane, the SREBP-SCAP complex is retained in ER. However, when the ER membrane is depleted of unesterified cholesterol, the SREBP-SCAP complex migrates to the Golgi apparatus where SREBP is sequentially processed by two Golgi resident proteases known as Site-1 and Site-2 proteases, respectively, to release its N-terminal fragment which is the active transcription factor. A recent discovery has added a twist in the complexity of this regulatory framework by elaborating the existence of a link between luminal ER Ca2+ homeostasis and cholesterol metabolism [83]. The direct reduction of the luminal ER Ca2+ caused the shrinkage of the unesterified cholesterol pool in the ER that triggers SREBP processing without altering the size of the total intracellular unesterified cholesterol pool [83]. This finding suggests that the size of the luminal ER Ca2+ pool may be involved in determining the basal sensitivity setpoint of the cholesterol sensing mechanism inherent to the SREBP processing pathway, and thus highlight the importance of luminal ER Ca2+ homeostasis in lipid metabolism.
8 ER and Membrane Contact Sites
The compartmentalization of specialized functions within discrete membrane bound organelles creates a challenge for efficient transport and exchange of molecules between compartments. Recent studies have documented the promiscuity of ER membranes for forming contacts with the plasma membrane and other organelles (mitochondria, peroxisomes, lysosomes, endosome, trans-Golgi, phagosomes, nuclear envelope) including lipid droplets [87,88,89,90,91,92,93,94,95]. These membrane contact sites occur between two heterologous membranes that are situated in very close proximity, approximately 10–30 nm, from each other and appear to be highly stable [36, 96, 97] except for STIM-ORAI (ER-plasma membrane) contacts which form transiently in the process of replenishing cellular Ca2+ stores. Contacts between other organelles, not involving the ER, have not been observed. It is also noteworthy that membrane contact sites between ER membranes with ribosomes (i.e., RER) and other organellar membranes are observed, implying possible functional heterogeneity in the membranes that make up the ER [93]. The joining of ER membranes to organellar membranes forms the cellular reticular network [14] characterized by interconnected membranes, tubules, vesicles and cisternae spanning the plasma membrane to the nuclear envelope linked together by membrane contacts sites that form portals that facilitates rapid passage of transport of substrates and metabolites (such as nutrients, biosynthetic intermediates, ATP) and signaling molecules (such as lipid messengers, Ca2+ ions) (Fig. 4.1) [14, 36, 94, 95, 98].
Recent studies have determined the identity of some of the molecules associated with membrane contacts sites [94, 99,100,101]. Proteins found at these structures include Ca2+ binding proteins (e.g., STIM, ORAI, SERCA, RyR), chaperones, transporters, and lipid binding and lipid transfer proteins. Membrane contact sites may represent highly specialized sites of function as suggested by studies on phospholipid synthesis [73, 102]. The existence of these structures may account for the rapid non-vesicular transport of various lipid metabolites between organelles, such as during biosynthesis of membrane lipids and cholesterol. Indeed, many of the proteins that have been characterized are lipid transfer proteins, whose substrates include fatty acids, phospholipids, sterols and their metabolites including those that serve as messengers [99, 103, 104]. In yeast, several specialized proteins associated with membrane contact sites between ER and mitochondria have been identified. These proteins form a functional unit referred to as ER-mitochondria encounter structure (ERMES) [96, 103, 105]. The equivalent functional unit in mammalian cells has not been described.
Furthermore, release of luminal ER Ca2+ generates areas of Ca2+ microdomains characterized by a high Ca2+ concentration and occur at the contact region between the ER and plasma membrane where Ca2+ channels on the plasma membrane open [37, 39, 40]. The formation of ER-membrane contacts and the complexity of Ca2+ signaling proteins in these microdomains accounts for the specific characteristics of discrete Ca2+ signals [89]. One of the best characterized membrane contacts are those formed between skeletal and cardiac muscle T (transverse)-tubules (extensions of the plasma membrane) and the SR [106,107,108,109]. These membrane contacts are the driving force to support excitation-contraction coupling in the muscle and the mechanisms behind the Ca2+-induced Ca2+ release in the cardiac muscle [110]. Membrane contacts between ER and mitochondria and Ca2+ microdomains that form between InsP3R and Ca2+ uniporters on the mitochondria support uptake of Ca2+ by mitochondria to match energy supply by Ca2+-dependent stimulation of oxidative phosphorylation [95, 111].
This versatile arrangement of contacts between ER and organellar membranes offers the cell an ability to influence and/or support highly specialized functions throughout the cellular reticular network. Impairment of the ER-organellar contact sites might be involved in the pathogenesis of diseases such as neurodegenerative diseases [51]. Understanding how the process involved in the formation, maintenance and function of ER-membrane contact sites are regulated will provide insight into the role of ER Ca2+ in coordinating gene expression and cellular function.
9 Conclusions
The ER is an extensive network of membranes that occupies a major proportion of the cell interior. The ER and the other organelles are characterized by unique proteome and lipidome which together provide ideal environments that enable specialized functions within the cell. Ca2+ homeostasis in general, and ER luminal Ca2+ in particular, is essential for the function of the ER and reticular network. Ca2+ is a key molecule involved in orchestrating the metabolic pathways occurring in different compartments of the cell. Internal or external factors that result in the loss of nutrient and energy homeostasis impose stress on cellular functions. Coping response strategies operate to alleviate and eliminate stress in the ER and other organelles to maintain proteostasis and lipidostasis within the entire cellular reticular network. ER membranes can form resilient contacts with the membranes of other organelles forming a complex cellular reticular network. Membrane contact sites enable rapid distribution and exchange of substrates, metabolites and signalling molecules to ensure optimal cellular function and homeostasis.
Abbreviations
- ATF6:
-
activating transcription factor 6
- BiP:
-
immunoglobulin binding protein
- ER:
-
endoplasmic reticulum
- ERAD:
-
ER-associated degradation
- ERMAS:
-
ER-mitochondria encounter structure
- GRP:
-
glucose regulated protein
- HSP:
-
heat shock protein
- InsP3:
-
inositol-1,4,5-trisphosphate
- IRE:
-
serine/threonine-protein kinase/endoribonuclease inositol-requiring enzyme
- ORAI:
-
Ca2+ release-activated Ca2+ channel
- PERK:
-
dsRNA-activated protein kinase-like ER kinase
- PDI:
-
protein disulfide isomerase
- RER:
-
rough ER
- SARAF:
-
SOCE-associated regulatory factor
- SCAP:
-
SREBP cleavage activating protein
- SER:
-
smooth ER
- SERCA:
-
sarcoplasmic/endoplasmic reticulum Ca2+-ATPase
- SOCE:
-
store-operated Ca2+ entry
- SR:
-
sarcoplasmic reticulum
- SREBP:
-
sterol-response element-binding protein
- STIM:
-
stromal-interacting molecule
- RyR:
-
ryanodine receptor
- UPR:
-
unfolded protein response
- Xbp1:
-
X-box binding protein 1
References
Hebert DN, Molinari M (2007) In and out of the ER: protein folding, quality control, degradation, and related human diseases. Physiol Rev 87:1377–1408
Krebs J, Agellon LB, Michalak M (2015) Ca2+ homeostasis and endoplasmic reticulum (ER) stress: an integrated view of calcium signaling. Biochem Biophys Res Commun 460:114–121
McCaffrey K, Braakman I (2016) Protein quality control at the endoplasmic reticulum. Essays Biochem 60:227–235
Coe H, Michalak M (2009) Calcium binding chaperones of the endoplasmic reticulum. Gen Physiol Biophys 28 Spec No Focus:F96–F103
Bose D, Chakrabarti A (2017) Substrate specificity in the context of molecular chaperones. IUBMB Life 69(9):647–659
Horowitz S, Koldewey P, Stull F, Bardwell JC (2017) Folding while bound to chaperones. Curr Opin Struct Biol 48:1–5
Caramelo JJ, Parodi AJ (2015) A sweet code for glycoprotein folding. FEBS Lett 589:3379–3387
Baiceanu A, Mesdom P, Lagouge M, Foufelle F (2016) Endoplasmic reticulum proteostasis in hepatic steatosis. Nat Rev Endocrinol 12:710–722
Horton JD, Goldstein JL, Brown MS (2002) SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver. J Clin Invest 109:1125–1131
Vincenz-Donnelly L, Hipp MS (2017) The endoplasmic reticulum: a hub of protein quality control in health and disease. Free Radic Biol Med 108:383–393
Volpi VG, Touvier T, D’Antonio M (2016) Endoplasmic reticulum protein quality control failure in myelin disorders. Front Mol Neurosci 9:162
Berridge MJ (2016) The inositol trisphosphate/calcium signaling pathway in health and disease. Physiol Rev 96:1261–1296
Soboloff J, Rothberg BS, Madesh M, Gill DL (2012) STIM proteins: dynamic calcium signal transducers. Nat Rev Mol Cell Biol 13:549–565
Groenendyk J, Agellon LB, Michalak M (2013) Coping with endoplasmic reticulum stress in the cardiovascular system. Annu Rev Physiol 75:49–67
Hetz C, Chevet E, Oakes SA (2015) Proteostasis control by the unfolded protein response. Nat Cell Biol 17:829–838
D’Amico D, Sorrentino V, Auwerx J (2017) Cytosolic proteostasis networks of the mitochondrial stress response. Trends Biochem Sci 42(9):712–725
Dicks N, Gutierrez K, Michalak M, Bordignon V, Agellon LB (2015) Endoplasmic reticulum stress, genome damage, and cancer. Front Oncol 5:11
Jovaisaite V, Auwerx J (2015) The mitochondrial unfolded protein response-synchronizing genomes. Curr Opin Cell Biol 33:74–81
Stepien KM, Heaton R, Rankin S, Murphy A, Bentley J, Sexton D, Hargreaves IP (2017) Evidence of oxidative stress and secondary mitochondrial dysfunction in metabolic and non-metabolic disorders. J Clin Med 6(7):E71
Szymanski J, Janikiewicz J, Michalska B, Patalas-Krawczyk P, Perrone M, Ziolkowski W, Duszynski J, Pinton P, Dobrzyn A, Wieckowski MR (2017) Interaction of mitochondria with the endoplasmic reticulum and plasma membrane in calcium homeostasis, lipid trafficking and mitochondrial structure. Int J Mol Sci 18(7):E1576
Baumann O, Walz B (2001) Endoplasmic reticulum of animal cells and its organization into structural and functional domains. Int Rev Cytol 205:149–214
Corbett EF, Michalak M (2000) Calcium, a signaling molecule in the endoplasmic reticulum? Trends Biochem Sci 25:307–311
High S, Lecomte FJ, Russell SJ, Abell BM, Oliver JD (2000) Glycoprotein folding in the endoplasmic reticulum: a tale of three chaperones? FEBS Lett 476:38–41
Jakob CA, Chevet E, Thomas DY, Bergeron JJ (2001) Lectins of the ER quality control machinery. Results Probl Cell Differ 33:1–17
Molinari M, Helenius A (2000) Chaperone selection during glycoprotein translocation into the endoplasmic reticulum. Science 288:331–333
Nakamura K, Zuppini A, Arnaudeau S, Lynch J, Ahsan I, Krause R, Papp S, De Smedt H, Parys JB, Müller-Esterl W, Lew DP, Krause K-H, Demaurex N, Opas M, Michalak M (2001) Functional specialization of calreticulin domains. J Cell Biol 154:961–972
Lievremont JP, Rizzuto R, Hendershot L, Meldolesi J (1997) BiP, a major chaperone protein of the endoplasmic reticulum lumen, plays a direct and important role in the storage of the rapidly exchanging pool of Ca2+. J Biol Chem 272:30873–33089
Van PN, Peter F, Soling H-D (1989) Four intracisternal calcium-binding glycoproteins from rat liver microsomes with high affinity for calcium. No indication for calsequestrin-like proteins in inositol 1,4,5-trisphosphate-sensitive calcium sequestering rat liver vesicles. J Biol Chem 264:17494–17501
Waser M, Mesaeli N, Spencer C, Michalak M (1997) Regulation of calreticulin gene expression by calcium. J Cell Biol 138:547–557
Lebeche D, Lucero HA, Kaminer B (1994) Calcium binding properties of rabbit liver protein disulfide isomerase. Biochem Biophys Res Commun 202:556–561
Lucero HA, Kaminer B (1999) The role of calcium on the activity of ERcalcistorin/protein-disulfide isomerase and the significance of the C-terminal and its calcium binding. A comparison with mammalian protein-disulfide isomerase. J Biol Chem 274:3243–3251
Lucero HA, Lebeche D, Kaminer B (1998) ERcalcistorin/protein-disulfide isomerase acts as a calcium storage protein in the endoplasmic reticulum of a living cell. Comparison with calreticulin and calsequestrin. J Biol Chem 273:9857–9863
Faggioni M, Knollmann BC (2012) Calsequestrin 2 and arrhythmias. Am J Physiol Heart Circ Physiol 302:H1250–H1260
Lee D, Michalak M (2010) Membrane associated Ca2+ buffers in the heart. BMB Rep 43:151–157
Bhardwaj R, Hediger MA, Demaurex N (2016) Redox modulation of STIM-ORAI signaling. Cell Calcium 60:142–152
Saheki Y, De Camilli P (2017) Endoplasmic reticulum-plasma membrane contact sites. Annu Rev Biochem 86:659–684
Zhou Y, Cai X, Nwokonko RM, Loktionova NA, Wang Y, Gill DL (2017) The STIM-Orai coupling interface and gating of the Orai1 channel. Cell Calcium 63:8–13
Takemura H, Putney JW Jr (1989) Capacitative calcium entry in parotid acinar cells. Biochem J 258:409–412
Delpire E (2016) STIM and Orai proteins in calcium signaling: an AJP-cell physiology series of themed reviews. Am J Physiol Cell Physiol 310:C401
Tanwar J, Motiani RK (2017) Role of SOCE architects STIM and Orai proteins in cell death. Cell Calcium https://doi.org/10.1016/j.ceca.2017.06.002
Brandman O, Liou J, Park WS, Meyer T (2007) STIM2 is a feedback regulator that stabilizes basal cytosolic and endoplasmic reticulum Ca2+ levels. Cell 131:1327–1339
Kawasaki H, Kretsinger RH (2017) Structural and functional diversity of EF-hand proteins: Evolutionary perspectives. Protein Sci 26(10):1898–1920
Hou X, Pedi L, Diver MM, Long SB (2012) Crystal structure of the calcium release-activated calcium channel Orai. Science 338:1308–1313
DeHaven WI, Smyth JT, Boyles RR, Putney JW Jr (2007) Calcium inhibition and calcium potentiation of Orai1, Orai2, and Orai3 calcium release-activated calcium channels. J Biol Chem 282:17548–17556
Gonzalez-Cobos JC, Zhang X, Zhang W, Ruhle B, Motiani RK, Schindl R, Muik M, Spinelli AM, Bisaillon JM, Shinde AV, Fahrner M, Singer HA, Matrougui K, Barroso M, Romanin C, Trebak M (2013) Store-independent Orai1/3 channels activated by intracrine leukotriene C4: role in neointimal hyperplasia. Circ Res 112:1013–1025
Thompson JL, Shuttleworth TJ (2013) Exploring the unique features of the ARC channel, a store-independent Orai channel. Channels (Austin) 7:364–373
Zhang W, Zhang X, Gonzalez-Cobos JC, Stolwijk JA, Matrougui K, Trebak M (2015) Leukotriene-C4 synthase, a critical enzyme in the activation of store-independent Orai1/Orai3 channels, is required for neointimal hyperplasia. J Biol Chem 290:5015–5027
Prins D, Groenendyk J, Touret N, Michalak M (2011) Modulation of STIM1 and capacitative Ca2+ entry by the endoplasmic reticulum luminal oxidoreductase ERp57. EMBO Rep 12:1182–1188
Li Y, Camacho P (2004) Ca2+-dependent redox modulation of SERCA 2b by ERp57. J Cell Biol 164:35–46
Ellgaard L, McCaul N, Chatsisvili A, Braakman I (2016) Co- and post-translational protein folding in the ER. Traffic 17:615–638
Jung J, Michalak M, Agellon LB (2017) Endoplasmic reticulum malfunction in the nervous system. Front Neurosci 11:220
Balchin D, Hayer-Hartl M, Hartl FU (2016) In vivo aspects of protein folding and quality control. Science 353:aac4354
Bernasconi R, Molinari M (2011) ERAD and ERAD tuning: disposal of cargo and of ERAD regulators from the mammalian ER. Curr Opin Cell Biol 23:176–183
Qi L, Tsai B, Arvan P (2017) New insights into the physiological role of endoplasmic reticulum-associated degradation. Trends Cell Biol 27:430–440
Grek C, Townsend DM (2014) Protein disulfide isomerase superfamily in disease and the regulation of apoptosis. Endoplasmic Reticul Stress Dis 1:4–17
Parakh S, Atkin JD (2015) Novel roles for protein disulphide isomerase in disease states: a double edged sword? Front Cell Dev Biol 3:30
Trombetta ES, Parodi AJ (2003) Quality control and protein folding in the secretory pathway. Annu Rev Cell Dev Biol 19:649–676
Carafoli E, Krebs J (2016) Why calcium? How calcium became the best communicator. J Biol Chem 291:20849–20857
Gidalevitz T, Prahlad V, Morimoto RI (2011) The stress of protein misfolding: from single cells to multicellular organisms. Cold Spring Harb Perspect Biol 3(6):a009704
Wang M, Kaufman RJ (2014) The impact of the endoplasmic reticulum protein-folding environment on cancer development. Nat Rev Cancer 14:581–597
Nakka VP, Prakash-babu P, Vemuganti R (2016) Crosstalk between endoplasmic reticulum stress, oxidative stress, and autophagy: potential therapeutic targets for acute CNS injuries. Mol Neurobiol 53:532–544
Zhang C, Syed TW, Liu R, Yu J (2017) Role of endoplasmic reticulum stress, autophagy, and inflammation in cardiovascular disease. Front Cardiovasc Med 4:29
Kaur J, Debnath J (2015) Autophagy at the crossroads of catabolism and anabolism. Nat Rev Mol Cell Biol 16:461–472
Groenendyk J, Peng Z, Dudek E, Fan X, Mizianty MJ, Dufey E, Urra H, Sepulveda D, Rojas-Rivera D, Lim Y, Kim do H, Baretta K, Srikanth S, Gwack Y, Ahnn J, Kaufman RJ, Lee SK, Hetz C, Kurgan L, Michalak M (2014) Interplay between the oxidoreductase PDIA6 and microRNA-322 controls the response to disrupted endoplasmic reticulum calcium homeostasis. Sci Signal 7:ra54
Leung AK, Sharp PA (2010) MicroRNA functions in stress responses. Mol Cell 40:205–215
Hong M, Luo S, Baumeister P, Huang JM, Gogia RK, Li M, Lee AS (2004) Underglycosylation of ATF6 as a novel sensing mechanism for activation of the unfolded protein response. J Biol Chem 279:11354–11363
Higa A, Taouji S, Lhomond S, Jensen D, Fernandez-Zapico ME, Simpson JC, Pasquet JM, Schekman R, Chevet E (2014) Endoplasmic reticulum stress-activated transcription factor ATF6alpha requires the disulfide isomerase PDIA5 to modulate chemoresistance. Mol Cell Biol 34:1839–1849
Eletto D, Eletto D, Dersh D, Gidalevitz T, Argon Y (2014) Protein disulfide isomerase A6 controls the decay of IRE1alpha signaling via disulfide-dependent association. Mol Cell 53:562–576
Khan S, Greco D, Michailidou K, Milne RL, Muranen TA, Heikkinen T, Aaltonen K, Dennis J, Bolla MK, Liu J, Hall P, Irwanto A, Humphreys K, Li J, Czene K, Chang-Claude J, Hein R, Rudolph A, Seibold P, Flesch-Janys D, Fletcher O, Peto J, dos Santos Silva I, Johnson N, Gibson L, Aitken Z, Hopper JL, Tsimiklis H, Bui M, Makalic E, Schmidt DF, Southey MC, Apicella C, Stone J, Waisfisz Q, Meijers-Heijboer H, Adank MA, van der Luijt RB, Meindl A, Schmutzler RK, Muller-Myhsok B, Lichtner P, Turnbull C, Rahman N, Chanock SJ, Hunter DJ, Cox A, Cross SS, Reed MW, Schmidt MK, Broeks A, Van't Veer LJ, Hogervorst FB, Fasching PA, Schrauder MG, Ekici AB, Beckmann MW, Bojesen SE, Nordestgaard BG, Nielsen SF, Flyger H, Benitez J, Zamora PM, Perez JI, Haiman CA, Henderson BE, Schumacher F, Le Marchand L, Pharoah PD, Dunning AM, Shah M, Luben R, Brown J, Couch FJ, Wang X, Vachon C, Olson JE, Lambrechts D, Moisse M, Paridaens R, Christiaens MR, Guenel P, Truong T, Laurent-Puig P, Mulot C, Marme F, Burwinkel B, Schneeweiss A, Sohn C, Sawyer EJ, Tomlinson I, Kerin MJ, Miller N, Andrulis IL, Knight JA, Tchatchou S, Mulligan AM, Dork T, Bogdanova NV, Antonenkova NN et al (2014) MicroRNA related polymorphisms and breast cancer risk. PLoS One 9:e109973
McMahon M, Samali A, Chevet E (2017) Regulation of the unfolded protein response by non-coding RNA. Am J Physiol Cell Physiol 313(3):C243–C254
van Meer G, Voelker DR, Feigenson GW (2008) Membrane lipids: where they are and how they behave. Nat Rev Mol Cell Biol 9:112–124
Chernomordik LV, Kozlov MM (2008) Mechanics of membrane fusion. Nat Struct Mol Biol 15:675–683
Kannan M, Lahiri S, Liu LK, Choudhary V, Prinz WA (2017) Phosphatidylserine synthesis at membrane contact sites promotes its transport out of the ER. J Lipid Res 58:553–562
Cerqueira NM, Oliveira EF, Gesto DS, Santos-Martins D, Moreira C, Moorthy HN, Ramos MJ, Fernandes PA (2016) Cholesterol biosynthesis: a mechanistic overview. Biochemistry 55:5483–5506
Kovacs WJ, Olivier LM, Krisans SK (2002) Central role of peroxisomes in isoprenoid biosynthesis. Prog Lipid Res 41:369–391
Agellon LB (2008) Metabolism and function of bile acids. In: Vance DE, Vance JE (eds) Biochemistry of lipids, lipoproteins and membranes, 5th edn. Elsevier, Amsterdam
Miller WL (2013) Steroid hormone synthesis in mitochondria. Mol Cell Endocrinol 379:62–73
Fleet JC (2017) The role of vitamin D in the endocrinology controlling calcium homeostasis. Mol Cell Endocrinol 453:36–45
Vega H, Agellon LB, Michalak M (2016) The rise of proteostasis promoters. IUBMB Life 68:943–954
Norimatsu Y, Hasegawa K, Shimizu N, Toyoshima C (2017) Protein-phospholipid interplay revealed with crystals of a calcium pump. Nature 545:193–198
Sriburi R, Jackowski S, Mori K, Brewer JW (2004) XBP1: a link between the unfolded protein response, lipid biosynthesis, and biogenesis of the endoplasmic reticulum. J Cell Biol 167:35–41
Bommiasamy H, Back SH, Fagone P, Lee K, Meshinchi S, Vink E, Sriburi R, Frank M, Jackowski S, Kaufman RJ, Brewer JW (2009) ATF6alpha induces XBP1-independent expansion of the endoplasmic reticulum. J Cell Sci 122:1626–1636
Wang WA, Liu WX, Durnaoglu S, Lee SK, Lian J, Lehner R, Ahnn J, Agellon LB, Michalak M (2017) Loss of calreticulin uncovers a critical role for calcium in regulating cellular lipid homeostasis. Sci Rep 7:5941
Brown MS, Goldstein JL (1997) The SREBP pathway: regulation of cholesterol metabolism by proteolysis of a membrane-bound transcription factor. Cell 89:331–340
Shimano H (2001) Sterol regulatory element-binding proteins (SREBPs): transcriptional regulators of lipid synthetic genes. Prog Lipid Res 40:439–452
Radhakrishnan A, Goldstein JL, McDonald JG, Brown MS (2008) Switch-like control of SREBP-2 transport triggered by small changes in ER cholesterol: a delicate balance. Cell Metab 8:512–521
Barneda D, Christian M (2017) Lipid droplet growth: regulation of a dynamic organelle. Curr Opin Cell Biol 47:9–15
Daniele T, Schiaffino MV (2014) Organelle biogenesis and interorganellar connections: Better in contact than in isolation. Commun Integr Biol 7:e29587
Filadi R, Pozzan T (2015) Generation and functions of second messengers microdomains. Cell Calcium 58:405–414
Joshi AS, Zhang H, Prinz WA (2017) Organelle biogenesis in the endoplasmic reticulum. Nat Cell Biol 19:876–882
Nunes-Hasler P, Demaurex N (2017) The ER phagosome connection in the era of membrane contact sites. Biochim Biophys Acta 1864:1513–1524
Penny CJ, Kilpatrick BS, Eden ER, Patel S (2015) Coupling acidic organelles with the ER through Ca2+ microdomains at membrane contact sites. Cell Calcium 58:387–396
Phillips MJ, Voeltz GK (2016) Structure and function of ER membrane contact sites with other organelles. Nat Rev Mol Cell Biol 17:69–82
Prinz WA (2014) Bridging the gap: membrane contact sites in signaling, metabolism, and organelle dynamics. J Cell Biol 205:759–769
Prudent J, McBride HM (2017) The mitochondria-endoplasmic reticulum contact sites: a signalling platform for cell death. Curr Opin Cell Biol 47:52–63
Helle SC, Kanfer G, Kolar K, Lang A, Michel AH, Kornmann B (2013) Organization and function of membrane contact sites. Biochim Biophys Acta 1833:2526–2541
Orci L, Ravazzola M, Le Coadic M, Shen WW, Demaurex N, Cosson P (2009) From the Cover: STIM1-induced precortical and cortical subdomains of the endoplasmic reticulum. Proc Natl Acad Sci U S A 106:19358–19362
Jain A, Holthuis JCM (2017) Membrane contact sites, ancient and central hubs of cellular lipid logistics. Biochim Biophys Acta 1864:1450–1458
Chiapparino A, Maeda K, Turei D, Saez-Rodriguez J, Gavin AC (2016) The orchestra of lipid-transfer proteins at the crossroads between metabolism and signaling. Prog Lipid Res 61:30–39
Lang A, John Peter AT, Kornmann B (2015) ER-mitochondria contact sites in yeast: beyond the myths of ERMES. Curr Opin Cell Biol 35:7–12
Levine TP, Patel S (2016) Signalling at membrane contact sites: two membranes come together to handle second messengers. Curr Opin Cell Biol 39:77–83
Vance JE (1990) Phospholipid synthesis in a membrane fraction associated with mitochondria. J Biol Chem 265:7248–7256
Raturi A, Simmen T (2013) Where the endoplasmic reticulum and the mitochondrion tie the knot: the mitochondria-associated membrane (MAM). Biochim Biophys Acta 1833:213–224
Wong LH, Levine TP (2016) Lipid transfer proteins do their thing anchored at membrane contact sites… but what is their thing? Biochem Soc Trans 44:517–527
Michel AH, Kornmann B (2012) The ERMES complex and ER-mitochondria connections. Biochem Soc Trans 40:445–450
Barone V, Randazzo D, Del Re V, Sorrentino V, Rossi D (2015) Organization of junctional sarcoplasmic reticulum proteins in skeletal muscle fibers. J Muscle Res Cell Motil 36:501–515
Landstrom AP, Beavers DL, Wehrens XH (2014) The junctophilin family of proteins: from bench to bedside. Trends Mol Med 20:353–362
Louch WE, Nattel S (2017) T-tubular collagen: a new player in mechanosensing and disease? Cardiovasc Res 113(8):839–840
Manfra O, Frisk M, Louch WE (2017) Regulation of cardiomyocyte T-tubular structure: opportunities for therapy. Curr Heart Fail Rep 14:167–178
Eisner DA, Caldwell JL, Kistamas K, Trafford AW (2017) Calcium and excitation-contraction coupling in the heart. Circ Res 121:181–195
De Stefani D, Rizzuto R, Pozzan T (2016) Enjoy the trip: calcium in mitochondria back and forth. Annu Rev Biochem 85:161–192
Acknowledgements
Research in our laboratories is supported by grants from the Canadian Institutes of Health Research grants MOP-15291, MOP-15415, MOP-86750 and PS-153325.
Conflict of Interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG, part of Springer Nature
About this chapter
Cite this chapter
Agellon, L.B., Michalak, M. (2017). The Endoplasmic Reticulum and the Cellular Reticular Network. In: Krebs, J. (eds) Membrane Dynamics and Calcium Signaling. Advances in Experimental Medicine and Biology, vol 981. Springer, Cham. https://doi.org/10.1007/978-3-319-55858-5_4
Download citation
DOI: https://doi.org/10.1007/978-3-319-55858-5_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-55857-8
Online ISBN: 978-3-319-55858-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)