Abstract
Despite advances in clinical cancer chemotherapy, such as reduced toxicities and prolonged progression-free survival, cures remain elusive due to eventual acquired drug resistance. To study ways of avoiding and/or overcoming the complicated issue of acquired drug resistance, optimal cancer models must be used. Patient-derived xenograft (PDX) tumor models provide an excellent opportunity to study resistance as they reflect the complex nature of patients’ malignancies. While only a few of such studies have been published, the exact method of modeling resistance is surprisingly varied. This chapter illustrates various approaches of using PDX models to develop and study acquired resistance to cancer chemotherapies.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
Cancer remains a major cause of death worldwide. Along with advancing cancer prevention, more effective treatments are desperately required. Despite significant strides over the past 30 years, resistance to systemic therapies remains a massive obstacle [1]. Diverse mechanisms of acquired resistance to cancer chemotherapies have been discovered [2]. Moreover, multiple mechanisms of resistance may exist against a single therapy [3,4,5,6,7,8,9]. Despite being able to identify some of the mechanisms underlying drug resistance, many of them are still unknown. Furthermore, biomarkers to predict how resistance will occur and optimal subsequent treatment methods remain to be determined in the majority of cancer cases. Resistance to a therapy may be innate or acquired [10]. To combat innate resistance, improved patient stratification strategies for therapies are required. To combat acquired resistance, a better understanding of the numerous mechanisms of resistance that can arise, and how to avoid or overcome them, is imperative.
To study a phenomenon as complex as acquired therapy resistance, optimal tumor models are required. Models based on cultured cancer cell lines have been shown to poorly predict patients’ responses to drugs, in part due to a lack of tumor heterogeneity, lack of three-dimensional tissue architecture and stromal support, and lack of dynamic and naturally occurring physiological phenomena such as hypoxia [11]. In contrast, patient-derived xenograft (PDX) tumor models have been reported to recapitulate major complexities of patients’ malignancies, including their responses to therapies [8, 9, 12,13,14]. Furthermore, such models can be leveraged to study the heterogeneity within a patient’s tumor; a single tumor sample may be split and implanted into multiple biological replicates, each potentially representing an array of subclones. The PDX models may also lend themselves to molecular characterizations of tumors during the course of treatment. Such procedures are not yet routinely performed in patients due to clinical challenges in deciding which tumor site to re-biopsy, as well as ethical issues such as the invasiveness of the biopsy procedure [15]. With PDXs, however, multiple biological replicates can be implanted allowing for subsets to be studied at various time points of the treatment regimen. As such, PDXs lend themselves as excellent models for studying the development of resistance to therapies.
We recently carried out a PubMed search based on “patient-derived xenograft models” and “resistance” which led to numerous studies, the majority of which describe experimental models of innate resistance. Often these studies involve treatment of multiple PDX models with a therapy aimed at determining what is molecularly different between the responders and the nonresponders. They then generally use these differences to predict the responses of another cohort to the same therapeutic [16,17,18,19]. As these studies fringe on “biomarker studies,” they will not be the major focus of this chapter. The same PubMed search yielded a handful of studies of which at least one section investigated acquired resistance using PDXs. We selected 15 of these investigations for comparative analysis as they cover the scope of methods and motivations (summarized in Table 8.1) for studying acquired therapy resistance by PDX models.
Therapeutic Doses for Development of Drug Resistance
One of the first steps in studying acquired resistance is to identify a model that is sensitive to the therapy of interest. Often more than one drug dosage will be explored during this initial PDX screening phase, in an effort to determine the optimal dose that produces tumor sensitivity while avoiding mouse toxicity; a theoretical example of such a study is presented in Fig. 8.1. From these initial screening studies, a dose can be determined for future investigations, such as modeling acquired resistance. Such initial drug screening studies are not always published together with the acquired resistance studies. In the case of vemurafenib, Das Thakur et al. [20] reported their initial screening study in the same paper as their resistance study. They investigated the growth-inhibitory activity of vemurafenib at three separate doses, 5, 15, and 45 mg/kg and found that the lowest dose was ineffective (similar to Dose A in Fig. 8.1), whereas the two higher doses were able to shrink the tumors. Surprisingly, 15 mg/kg was almost as effective as the higher dose. However, it became apparent that 45 mg/kg was the more effective dose as it maintained minimal disease, whereas tumors treated at 15 mg/kg began to escape inhibition after approximately 1 month (similar to Dose C in Fig. 8.1). As such, the authors decided to use the highest dose for their resistance modeling (similar to Dose D in Fig. 8.1).
Example of an initial drug screening study to determine optimal dose for subsequent experiments. The average tumor volume (y-axis) of five mouse cohorts was plotted over time (x-axis). The gray arrow represents the start of treatment for the four dosed cohorts. In this example Dose A has the lowest drug concentration, Dose D has the highest, and Dose B and C are in between
Shen et al. [21] also reported their preliminary drug sensitivity screening in the same paper as their acquired resistance study. Two doses, 12.5 or 25 mg/kg of crizotinib once a day for 21 days, were investigated. However, data from the lower dose cohort were shown for only one of the PDX models studied. Regardless, using the model that was most sensitive to crizotinib, the authors chose to continue to treat at the lower dose of 12.5 mg/kg for another month to develop resistance. Using our example shown in Fig. 8.1, the authors essentially explored Doses C and D and opted to use Dose C for developing resistance.
Other investigators used different ways of screening drug dosages to generate therapy resistance. Gaponova et al. performed a drug screen for STA-8666 at 75 mg/kg for over a month and achieved stable disease or tumor regression (similar to Dose D in Fig. 8.1); they then increased the dosage to 150 mg/kg which led to increased regression. However, two of the three tumors escaped therapeutic control and grew uncontrollably [22]. Micel et al. similarly did a drug screen of the MEK inhibitor TAK-733, examining 25 or 10 mg/kg, and then used 100 mg/kg to develop resistance [23].
Tumor Volume at Treatment Initiation
Tumor volume is often the main characteristic used to determine when to start PDX dosing. Some studies use rolling enrollment, i.e., treatment of individual mice is started whenever their tumors reach a specific volume (Fig. 8.2a). Treatment of mice can also be started when their tumors reach a prespecified average volume (Fig. 8.2b). Given the technical difficulties of regularly measuring orthotopically implanted tumors (aside from mammary fat pad implantations), studies using these models tend to rely more on timing rather than specific tumor volumes [24, 25]. Among the studies using subcutaneously implanted PDXs, however, the tumor volume at dosing initiation varies widely, regardless of which enrollment method is used.
A number of studies did not explicitly state at what tumor volume treatment was started. Some of these studies reported the growth curves, with actual tumor volumes on the y-axis, allowing readers to extrapolate the starting volumes [20, 21, 26]. Unfortunately, other studies either did not report a starting tumor volume and published graphs with transformed data [22, 27, 28] or stated parameters such as “during log phase,” which could cover a wide range of volumes [29]. Monsma et al. [30] reported that treatment was started when tumors were in exponential growth; however, they also gave an approximate tumor volume to indicate what this might mean [30]. Ter Brugge et al. [31], Micel et al. [23], Gao et al. [32], and Tentler et al. [33] specifically stated either the mean or the individual tumor volume criteria that they used for treatment initiation, all of which seem to have been chosen around 100–300 mm3 [23, 31,32,33]. Ciamporcero et al. [34] also reported the approximate tumor volumes for treatment initiation; however, this was reported in mm2 instead of mm3, making comparisons with other studies less direct [34].
Methods for Developing Drug Resistance
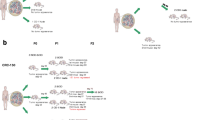
The overall method for developing a resistant model also varies from study to study. Parameters, such as constant versus escalating doses, constant versus cycled doses, and dosing across one or more passages, were varied across experiments. The majority of studies utilized continuous dosing at one concentration until resistance developed (Fig. 8.3a). The resistant tumors were either molecularly characterized or serially passaged to maintain a resistant model for further studies [20, 21, 23, 25, 26, 28, 29, 32, 33]. Ciamporcero et al. [34] also used continuous dosing; however, once tumors regrew under therapeutic pressure to double their initial volume, PDXs were considered “resistant” and randomized into experimental arms to determine the effectiveness of alternate therapies (Fig. 8.3b). Gaponova et al. [22] also employed continuous dosing to develop resistance, yet increased their dosage from 75 to 150 mg/kg STA-8666 after approximately 1 month (Fig. 8.3c). Cottu et al. [27] dosed their PDXs continuously, yet performed three further serial passages, with continued dosing, before they considered the model to be truly drug-resistant and used it for further studies (Fig. 8.3d).
Aside from variations on continuous dosing, some studies also employ cycling methods to develop drug resistance. Again, specifics such as number of passages and drug dosages vary across studies. Monsma et al. [30] treated PDX tumors of ~160 mm3 for 28 days, during which average tumor volumes regressed. After this treatment period, the researchers allowed three tumors to relapse (off treatment) and passaged them into a new cohort of mice and again allowed them to reach ~160 mm3 before treatment was continued. Upon the therapeutic rechallenge, the tumors exhibited decreased drug sensitivity and did not regress but continued to grow while under therapeutic pressure (albeit at a slower rate than untreated control tumors) (Fig. 8.4a). Vidal et al. [24] also cycled therapies across multiple passages to generate a resistant model; however, they increased the dose at each passage (Fig. 8.4b). The mice received one cycle of cisplatin at 2 mg/kg once a week for 3 weeks, and the tumors were then allowed to relapse. Tumors were then passaged into the next cohort of mice and allowed to again grow until intra-abdominal masses were palpable. These PDXs received one cycle of cisplatin at 3 mg/kg once a week for 3 weeks and were then allowed to relapse. This process was repeated for up to five passages and cycles with increasing doses (cycle 3, 3.5 mg/kg; cycle 4, 4 mg/kg; cycle 5, 5 mg/kg). After the fifth cycle, a stably shortened time to relapse was observed for the PDXs, and they were considered “cisplatin-resistant” and used for further studies. Lastly, Ter Brugge et al. [31] cycled tumors within a single passage (Fig. 8.4c). Mice were randomized when their tumors reached a volume of 100 mm3, and treatment was initiated at 200 mm3. Treatment was stopped if tumors regressed to <50% of the initial tumor volume and was resumed once the starting volume was regained.
Motivation for Developing Drug-Resistant PDX Models
Resistance was developed in PDXs across these studies for a number of reasons. While many of the studies using PDXs with acquired resistance had more than one focus (as depicted in Table 8.1), there seem to be four major themes. Studying the mechanism of acquired resistance was, as expected, the most common theme found in the studies reviewed. For some studies such as those by Nathanson et al. [28], Kopetz et al. [26], Shen et al. [21], and Ter Brugge et al. [31], resistant PDXs were generated to study novel mechanisms of resistance. Das Thakur et al. [20], Monsma et al. [29, 30], Ciamporcero et al. [34], Cottu et al. [27], and Vidal et al. [24] focused on determining optimal subsequent therapies to combat the acquired resistance that had developed. Gao et al. [32] and Zhao et al. [25] developed resistance in large part to demonstrate the benefit of using their models to study clinically relevant issues. Finally, Tentler et al. [33], Gaponova et al. [22], and Micel et al. [23] investigated the efficacy of novel therapies and wanted to determine whether acquired resistance might occur and through what mechanisms.
Perspectives and Future Directions
Determining how best to model acquired resistance by choosing a treatment regimen including drug concentration, PDX enrollment, and treatment timing can be a subjective matter. As such, some perspectives on each matter deserve a discussion.
The use of a lower dose that decreases tumor growth rate, but does not cause actual tumor shrinkage, may seem ineffective [21]. By the end of Shen et al.’s initial screen of 21 days, the tumor had actually continued to progress, albeit at a slower rate than the control tumors, suggesting that the optimal dose had not been achieved. The higher dose of crizotinib seems more optimal as it shrank tumors, and no toxicities were reported, thus making it the appropriate dose for modeling acquired resistance. However, achieving higher, optimal doses is not always possible in humans, and treatments that merely slow the growth of tumors may be clinically relevant.
Clearly reporting parameters such as tumor volume at treatment initiation is imperative as this criterion can have a large effect on a study’s outcome. Some treatments may be affected by the volume of the tumors. For example, large tumors may have more hypoxic and/or less vascularized regions, potentially decreasing effectiveness of the therapy employed. In contrast, small tumors may not be fully established in their implantation site, lacking supportive stroma that is observed in clinically detectable tumors, making them more vulnerable to treatments. As such, clearly stating important experimental parameters such as tumor volume at treatment initiation allows readers to draw their own conclusions regarding the reported results of the study.
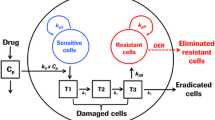
Lastly, reporting individual growth curves for all replicates in a study provides potentially useful information to readers. For example, if all replicates develop resistance, a highly distributed subclone (Fig. 8.5a) or therapy-induced event, such as epigenetic reprogramming or increased transcription of pro-survival genes (Fig. 8.5b), may be responsible for the resistance. If only a few replicates develop resistance, perhaps only those replicates contained rare subclones with innate mechanisms of resistance, such as mutations within the target pathway (Fig. 8.5c). At the very least, authors should state how many replicates actually developed resistance. Furthermore, the growth kinetics of tumors developing resistance may be informative too. Slow-growing “resistant” tumors may simply be demonstrating a decrease in sensitivity to drug and may actually be inhibited by higher doses, whereas quickly growing “resistant” tumors may have acquired novel mechanisms of resistance that completely overcome any therapy-specific effect regardless of dose. Among the manuscripts reviewed, approximately half showed either all replicates [20, 22, 27, 31] or at least showed examples of individual growth curves that did develop resistance [26, 29, 33]. Some of the subcutaneous PDX studies did not show any growth curves for the development of acquired resistance and only reported on downstream applications of the resistant model(s) [23, 32].
Schema demonstrating how various mechanisms of resistance may impact the number of biological replicates that actually develop resistance in a study. (a) Acquired resistance is due to a highly distributed subclone with innate resistance. (b) Acquired resistance is due to a therapy-induced cellular event such as epigenetic and/or transcriptional reprogramming. (c) Acquired resistance is due to only a few rare subclones with innate resistance
Conclusions
PDXs provide versatile cancer models for studying multiple facets of acquired drug resistance. It has been shown that, following exposure to pharmacological pressure, they develop resistance via similar mechanisms and with similar timelines as patients’ tumors [25, 32]. PDXs have been used to study novel mechanisms of resistance to current therapies [21, 26, 28, 31] and to investigate what to do once resistance develops [24, 27, 34]; they also have been used to try to stay ahead of the game and determine the propensity of tumors to acquire resistance to novel therapies [22, 23, 33]. PDXs are even becoming incorporated into clinical trial studies: PDXs with acquired resistance have been used in lieu of post-progression biopsies to determine potential mechanisms of resistance observed in a clinical trial [26]. Given the versatility of these models to explore clinically relevant issues in a timely manner, we believe that PDX incorporation into the clinic will become more common in the future.
Abbreviations
- PDX:
-
Patient-derived xenograft
References
Stewart BW, Wild CP. World Cancer Report 2014.
Hu X, Zhang Z. Understanding the genetic mechanisms of cancer drug resistance using genomic approaches. Trends Genet. 2016;32(2):127–37. doi:10.1016/j.tig.2015.11.003.
Braun DA, Burke KP, Van Allen EM. Genomic approaches to understanding response and resistance to immunotherapy. Clin Cancer Res. 2016;22(23):5642–50. doi:10.1158/1078-0432.CCR-16-0066.
Gillis NK, McLeod HL. The pharmacogenomics of drug resistance to protein kinase inhibitors. Drug Resist Updat. 2016;28:28–42. doi:10.1016/j.drup.2016.06.008.
Hotte SJ. Addressing taxane resistance in metastatic castration-resistant prostate cancer: a focus on chaperone proteins. Future Oncol. 2017;13(4):369–79. doi:10.2217/fon-2016-0279.
Paoletti C, Larios JM, Muniz MC, Aung K, Cannell EM, Darga EP, et al. Heterogeneous estrogen receptor expression in circulating tumor cells suggests diverse mechanisms of fulvestrant resistance. Mol Oncol. 2016;10(7):1078–85. doi:10.1016/j.molonc.2016.04.006.
Shi H, Zhang W, Zhi Q, Jiang M. Lapatinib resistance in HER2+ cancers: latest findings and new concepts on molecular mechanisms. Tumour Biol. 2016; doi:10.1007/s13277-016-5467-2.
Stewart EL, Mascaux C, Pham NA, Sakashita S, Sykes J, Kim L, et al. Clinical utility of patient-derived xenografts to determine biomarkers of prognosis and map resistance pathways in EGFR-mutant lung adenocarcinoma. J Clin Oncol. 2015a;33(22):2472–80. doi:10.1200/JCO.2014.60.1492.
Stewart EL, Tan SZ, Liu G, Tsao MS. Known and putative mechanisms of resistance to EGFR targeted therapies in NSCLC patients with EGFR mutations-a review. Transl Lung Cancer Res. 2015b;4(1):67–81. doi:10.3978/j.issn.2218-6751.2014.11.06.
Gottesman MM. Mechanisms of cancer drug resistance. Annu Rev Med. 2002;53:615–27. doi:10.1146/annurev.med.53.082901.103929.
Choi SY, Lin D, Gout PW, Collins CC, Xu Y, Wang Y. Lessons from patient-derived xenografts for better in vitro modeling of human cancer. Adv Drug Deliv Rev. 2014;79–80:222–37. doi:10.1016/j.addr.2014.09.009.
Dodbiba L, Teichman J, Fleet A, Thai H, Starmans MH, Navab R, et al. Appropriateness of using patient-derived xenograft models for pharmacologic evaluation of novel therapies for esophageal/gastro-esophageal junction cancers. PLoS One. 2015;10(3):e0121872. doi:10.1371/journal.pone.0121872.
Nunes M, Vrignaud P, Vacher S, Richon S, Lievre A, Cacheux W, et al. Evaluating patient-derived colorectal cancer xenografts as preclinical models by comparison with patient clinical data. Cancer Res. 2015;75(8):1560–6. doi:10.1158/0008-5472.CAN-14-1590.
Wong NC, Bhadri VA, Maksimovic J, Parkinson-Bates M, Ng J, Craig JM, et al. Stability of gene expression and epigenetic profiles highlights the utility of patient-derived paediatric acute lymphoblastic leukaemia xenografts for investigating molecular mechanisms of drug resistance. BMC Genomics. 2014;15:416. doi:10.1186/1471-2164-15-416.
Meghan Campo ZP, Muzikansky A, Rizzo C, Gainor JF, Shaw AT, Engelman JA, Sequist LV, editors. The challenges of repeat biopsies at acquired resistance to EGFR TKIs. ASCO Annual Meeting. J Clin Oncol. 2016:e20571.
Chen H, Ye Q, Lv J, Ye P, Sun Y, Fan S, et al. Evaluation of trastuzumab anti-tumor efficacy and its correlation with HER-2 status in patient-derived gastric adenocarcinoma xenograft models. Pathol Oncol Res. 2015;21(4):947–55. doi:10.1007/s12253-015-9909-8.
Forest A, Amatulli M, Ludwig DL, Damoci CB, Wang Y, Burns CA, et al. Intrinsic resistance to cixutumumab is conferred by distinct isoforms of the insulin receptor. Mol Cancer Res. 2015;13(12):1615–26. doi:10.1158/1541-7786.MCR-15-0279.
Krumbach R, Schuler J, Hofmann M, Giesemann T, Fiebig HH, Beckers T. Primary resistance to cetuximab in a panel of patient-derived tumour xenograft models: activation of MET as one mechanism for drug resistance. Eur J Cancer. 2011;47(8):1231–43. doi:10.1016/j.ejca.2010.12.019.
Wu X, Zhang J, Zhen R, Lv J, Zheng L, Su X, et al. Trastuzumab anti-tumor efficacy in patient-derived esophageal squamous cell carcinoma xenograft (PDECX) mouse models. J Transl Med. 2012;10:180. doi:10.1186/1479-5876-10-180.
Das Thakur M, Salangsang F, Landman AS, Sellers WR, Pryer NK, Levesque MP, et al. Modelling vemurafenib resistance in melanoma reveals a strategy to forestall drug resistance. Nature. 2013;494(7436):251–5. doi:10.1038/nature11814.
Shen A, Wang L, Huang M, Sun J, Chen Y, Shen YY, et al. c-Myc alterations confer therapeutic response and acquired resistance to c-Met inhibitors in MET-addicted cancers. Cancer Res. 2015;75(21):4548–59. doi:10.1158/0008-5472.CAN-14-2743.
Gaponova AV, Nikonova AS, Deneka AY, Kopp MC, Kudinov AE, Skobeleva N, et al. A novel HSP90 inhibitor-drug conjugate to SN38 is highly effective in small cell lung cancer. Clin Cancer Res. 2016;22(20):5120–9. doi:10.1158/1078-0432.CCR-15-3068.
Micel LN, Tentler JJ, Tan AC, Selby HM, Brunkow KL, Robertson KM, et al. Antitumor activity of the MEK inhibitor TAK-733 against melanoma cell lines and patient-derived tumor explants. Mol Cancer Ther. 2015;14(2):317–25. doi:10.1158/1535-7163.MCT-13-1012.
Vidal A, Munoz C, Guillen MJ, Moreto J, Puertas S, Martinez-Iniesta M, et al. Lurbinectedin (PM01183), a new DNA minor groove binder, inhibits growth of orthotopic primary graft of cisplatin-resistant epithelial ovarian cancer. Clin Cancer Res. 2012;18(19):5399–411. doi:10.1158/1078-0432.CCR-12-1513.
Zhao H, Thong A, Nolley R, Reese SW, Santos J, Ingels A, et al. Patient-derived tissue slice grafts accurately depict response of high-risk primary prostate cancer to androgen deprivation therapy. J Transl Med. 2013;11:199. doi:10.1186/1479-5876-11-199.
Kopetz S, Desai J, Chan E, Hecht JR, O’Dwyer PJ, Maru D, et al. Phase II pilot study of vemurafenib in patients with metastatic BRAF-mutated colorectal cancer. J Clin Oncol. 2015;33(34):4032–8. doi:10.1200/JCO.2015.63.2497.
Cottu P, Bieche I, Assayag F, El Botty R, Chateau-Joubert S, Thuleau A, et al. Acquired resistance to endocrine treatments is associated with tumor-specific molecular changes in patient-derived luminal breast cancer xenografts. Clin Cancer Res. 2014;20(16):4314–25. doi:10.1158/1078-0432.CCR-13-3230.
Nathanson DA, Gini B, Mottahedeh J, Visnyei K, Koga T, Gomez G, et al. Targeted therapy resistance mediated by dynamic regulation of extrachromosomal mutant EGFR DNA. Science. 2014;343(6166):72–6. doi:10.1126/science.1241328.
Monsma DJ, Cherba DM, Eugster EE, Dylewski DL, Davidson PT, Peterson CA, et al. Melanoma patient derived xenografts acquire distinct vemurafenib resistance mechanisms. Am J Cancer Res. 2015;5(4):1507–18.
Monsma DJ, Cherba DM, Richardson PJ, Vance S, Rangarajan S, Dylewski D, et al. Using a rhabdomyosarcoma patient-derived xenograft to examine precision medicine approaches and model acquired resistance. Pediatr Blood Cancer. 2014;61(9):1570–7. doi:10.1002/pbc.25039.
Ter Brugge P, Kristel P, van der Burg E, Boon U, de Maaker M, Lips E et al. Mechanisms of therapy resistance in patient-derived xenograft models of BRCA1-deficient breast cancer. J Natl Cancer Inst. 2016;108(11). doi:10.1093/jnci/djw148.
Gao H, Korn JM, Ferretti S, Monahan JE, Wang Y, Singh M, et al. High-throughput screening using patient-derived tumor xenografts to predict clinical trial drug response. Nat Med. 2015;21(11):1318–25. doi:10.1038/nm.3954.
Tentler JJ, Ionkina AA, Tan AC, Newton TP, Pitts TM, Glogowska MJ, et al. p53 family members regulate phenotypic response to aurora kinase a inhibition in triple-negative breast cancer. Mol Cancer Ther. 2015;14(5):1117–29. doi:10.1158/1535-7163.MCT-14-0538-T.
Ciamporcero E, Miles KM, Adelaiye R, Ramakrishnan S, Shen L, Ku S, et al. Combination strategy targeting VEGF and HGF/c-met in human renal cell carcinoma models. Mol Cancer Ther. 2015;14(1):101–10. doi:10.1158/1535-7163.MCT-14-0094.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Stewart, E.L., Tsao, M.S. (2017). Modeling Drug Resistance in PDX Models. In: Wang, Y., Lin, D., Gout, P. (eds) Patient-Derived Xenograft Models of Human Cancer . Molecular and Translational Medicine. Humana Press, Cham. https://doi.org/10.1007/978-3-319-55825-7_8
Download citation
DOI: https://doi.org/10.1007/978-3-319-55825-7_8
Published:
Publisher Name: Humana Press, Cham
Print ISBN: 978-3-319-55824-0
Online ISBN: 978-3-319-55825-7
eBook Packages: MedicineMedicine (R0)