Abstract
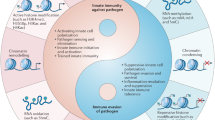
Multidisciplinary approaches combining microbiology, cell biology, and genetics have improved our understanding of bacterial diseases by elucidating mechanisms employed by bacteria to manipulate eukaryotic cellular processes. In parallel, research on epigenetics has increased our knowledge about eukaryotic gene expression by providing a mechanistic basis for the amazing plasticity of the genome in response to developmental and environmental cues. These two fields of research have now converged, providing information about the ways in which bacteria shape the epigenome and the mechanisms by which the epigenetic machinery allows the host to respond to colonization by pathogenic or commensal bacteria. The study of this cross talk has revealed remarkable diversity in the mechanisms of action of bacteria on chromatin and has identified epigenetic regulators involved in host responsiveness to bacteria. One powerful strategy used by intracellular pathogens (e.g., Anaplasma, Chlamydia, Ehrlichia, Legionella, Listeria, Mycobacteria, Mycoplasma, Shigella) is the secretion of nucleomodulins that manipulate chromatin structure in the host nucleus. The effects of this dialog are often limited in time, causing transient gene expression changes. However, increasing evidence suggests that certain epigenetic changes triggered by bacterial molecules are long-lasting, leading to the priming of transcriptional responses and the reprogramming of genes involved in inflammation or tolerance, with consequences for reinfection and polymicrobial infections. In addition, the effects of bacteria on the host epigenome may ultimately modify the identity of the cell by breaking epigenetic barriers, leading to cell differentiation, dedifferentiation, or trans-differentiation, thereby potentially contributing to tissue remodeling and emergence of complex diseases.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Infectious diseases
- Chromatin
- Microbiota
- Bacterial pathogenesis
- Cellular reprogramming
- Host–Pathogen interaction
- Nucleomodulin
- Patho-epigenetics
6.1 Introduction
Pathogenic bacteria have evolved a wide range of mechanisms for manipulating eukaryotic cell functions to their advantage (Diacovich and Gorvel 2010; Jimenez et al. 2016). In particular, by modifying the host cell transcriptional program, they disturb diverse cellular processes and take control of host defense systems. The commensal bacteria of the microbiota also affect host transcriptional gene networks by producing metabolites that influence the differentiation, proliferation, migration, and metabolic functions of mucosal cells (Brestoff and Artis 2013). Conversely, in conditions of microbial attack or colonization, host cells trigger various responses enabling them to tolerate or eliminate the invaders by mobilizing genes involved in key processes (e.g., immunity, cell death/survival, adhesion/motility, metabolism) (Jenner and Young 2005). Studies of the molecular basis of this cross talk are crucial for an understanding of infectious diseases and mucosal homeostasis. Research has long focused on the manipulation of transcription factors (e.g., NF-κB, FOS/JUN, IRFs, STATs, HIFs, SMADs) (Jenner and Young 2005; Bhavsar et al. 2007), through the bacteria-mediated deregulation of signaling pathways, or through posttranslational modifications (PTMs), activating, shutting down, or delocalizing these transcription factors. Another powerful means by which bacteria alter the expression of host genes has recently emerged from studies in different bacterial models: specific modifications of chromatin in the cell nucleus. This chapter will update a previous contribution dealing with the relationship between bacteria and chromatin regulation (Bierne et al. 2012) and will present new evidence for the epigenetic inheritance of bacterial imprints in cells and tissues. We will first recall the general principles of epigenetic regulation, and several examples will then be used to illustrate the diversity of mechanisms employed by bacteria and animal or human cells to mobilize the epigenetic machinery and modify the expression of susceptibility or resistance genes in the short or long term. The epigenetic control of adaptive immunity (Alvarez-Errico et al. 2015) and the effects of plant-associated bacteria on chromatin (Ma et al. 2011; Canonne and Rivas 2012; Holeski et al. 2012) will not be addressed here.
6.2 The Machinery of Chromatin Regulation
6.2.1 Chromatin Marks
In eukaryotic cells, DNA is wrapped around histone proteins to form nucleosomes, which are themselves packed with non-histone chromosomal proteins in the chromatin fiber. Chromatin condensation organizes and confines the genome into the tight space of the nucleus. More locally, the state of chromatin compaction plays a major role in nuclear processes by controlling the accessibility of DNA to the transcription, replication, and repair machineries. The regulation of chromatin structure is a dynamic process that involves DNA methylation (mostly on cytosines) (Klose and Bird 2006; Chen and Riggs 2011), histone PTMs (e.g., phosphorylation, methylation, acetylation, ubiquitination, SUMOylation, citrullination, ADP-ribosylation) (Kouzarides 2007; Sadakierska-Chudy and Filip 2015), and the sliding of nucleosomes along the DNA. Mechanisms for modifying and remodeling chromatin function together, controlling the formation of higher order chromatin structures that are either loosely packed and transcriptionally active (i.e., “euchromatin”) or highly condensed and transcriptionally silent (i.e., “heterochromatin”). Different combinations of histone PTMs and DNA methylation patterns form a code that controls transcription by affecting either chromatin structure itself or the recruitment of DNA-binding transcription activators or repressors. Several chromatin marks are known to be associated essentially with transcriptional activation (e.g., H3K4me, H3S10p, and H3K14ac; all abbreviations are listed in Table 6.1), whereas others are associated with repression (e.g., H3K9me, H3K27me, and deacetylated histones). However, it is often difficult to interpret a specific chromatin signature for the prediction of gene expression outcomes. Some genes may carry both repressive and activating histone marks, and RNA polymerase II may constitutively bind their proximal promoters, preparing the gene for efficient future transcription while remaining silent (Mikkelsen et al. 2007). Such active chromatin states at sites of repressed transcription may “poise” transcripts for rapid activation in cells in which a rapid change in expression levels is required, during immune and metabolic responses, for example (Cuddapah et al. 2010; Rye et al. 2014).
Chromatin modifications also contribute to the alternative splicing of pre-mRNA, making it possible for a single eukaryotic gene to encode several proteins with different functions (Allemand et al. 2008; Hnilicova and Stanek 2011). An additional level of complexity is added by noncoding RNAs and RNA-binding proteins (Turner and Morris 2010; Kaikkonen et al. 2011; Sadakierska-Chudy and Filip 2015). Most of the genomic DNA of eukaryotes is transcribed, but only 1–2% of transcripts encode proteins. The vast majority of RNAs are thus noncoding RNAs (ncRNA) of various sizes, generated from exons, introns, enhancers, or intergenic regions, in sense or antisense orientation (Mattick and Makunin 2005; Tisseur et al. 2011). Antisense RNAs and small ncRNAs play diverse roles in chromatin regulation by recruiting chromatin-modifying enzymes (Faghihi and Wahlestedt 2009; Kaikkonen et al. 2011; Cao 2014) and/or acting as scaffolds localizing genes to specific subnuclear regions (Yang et al. 2011; Schmitz et al. 2010). Last but not least, RNA can itself be modified, particularly by adenosine methylation (Dominissini et al. 2012), which affects messenger RNA localization, stability, splicing, and translation (Meyer and Jaffrey 2014; Dominissini et al. 2016). The role of the RNA world in the relationship between microbes and epigenetic regulation is an emerging field of research worthy of consideration in its own right. This chapter focuses exclusively on the role of histone and DNA modifications.
6.2.2 Chromatin Regulators
The molecular machinery controlling chromatin structure includes about 800 proteins with diverse functions (referenced in the EpiFactors database (Medvedeva et al. 2015); abbreviations for those mentioned here are listed in Table 6.1). ATP-dependent remodeling enzymes from the SWI/SNF, ISWI, CHD, and INO80/SWR families position the nucleosomes by catalyzing the movement of histone octamers relative to DNA, using energy from ATP hydrolysis to move, destabilize, evict, or reassemble nucleosomes (Langst and Manelyte 2015). Covalent modifications of chromatin are added or removed by a wide range of enzymes known as “writers”, such as histone kinases, acetyltransferases (HATs), and methyltransferases (HMTs), and “erasers”, such as histone phosphatases, deacetylases (HDACs), and demethylases (HDMs) (Fig. 6.1) (Zhou et al. 2011). In DNA methylation, the writers of 5-methylcytosine (5mC) are DNA methyltransferases (DNMTs), which either establish methylation (i.e., the “de novo” methyltransferases DNMT3a and DNMT3b) or copy methylation patterns onto the newly synthesized DNA strand during replication (i.e., the “maintenance” methyltransferase DNMT1). DNA methylation can be reversed passively, through a lack of 5mC copying during DNA replication, or actively through an active erasure process, involving intermediate chemical modifications of 5mC, followed by passive demethylation or DNA repair. Several groups of proteins, such as the ten-eleven translocation (TET) proteins and DNA glycosylases, have been shown to be involved in this complex process (Bhutani et al. 2011; Chen and Riggs 2011; Wu and Zhang 2011). 5mC and histone PTMs serve as signaling platforms for proteins known as “readers”, which interact, stabilize, or modify other chromatin components (Fig. 6.1). Histone readers are docked onto specific PTMs via chromatin-binding modules, such as bromodomains (BRD), bromo-adjacent homology domains (BAH), chromodomains, 14-3-3, Tudor, or PHD domains (Taverna et al. 2007). Methyl-cytosine readers include methyl-DNA-binding domains (MBD), SET and RING-associated domains (SRA), and some specific zinc finger motifs (Sasai and Defossez 2009; Liu et al. 2013; Buck-Koehntop and Defossez 2013).
The epigenetic machinery. Histone posttranslational modifications, histone sliding, and 5-cytosine methylation (5mC) control chromatin structure and gene expression. Chromatin is modified and remodeled by enzymes known as “writers,” “erasers,” and “remodelers.” Examples of chromatin enzymes and marks are shown. Activating marks include serine phosphorylation on lysine 10 (S10p) and acetylation on lysine 14 (K14ac) of histone H3 and methylation on lysine 4 (K4me) of histone H4. Repressive marks include dephosphorylation, deacetylation, and demethylation of the same residues, as well as methylation of lysine 9 (K9me) of H3. Cytosine methylation is a repressive mark at promoter sequences and an activating mark at gene bodies. Its erasure involves a complex pathway with chemical modifications of 5mC, followed by passive demethylation or DNA repair. Epigenetic marks are recognized and interpreted by protein modules known as “readers.” For instance, the bromodomain of the HAT p300 binds H3K14ac, the chromodomain of HP1 binds H3K9me, and the MBD domain of MBD1 binds 5mC. Writers, readers, erasers, and nucleosome remodelers act within large macromolecular complexes that open or close chromatin, leading to gene activation or repression. Cell signaling pathways triggered by external stimuli control interaction or stability of chromatin-activating or -repressive complex subunits and their combinatorial interaction with transcription factors (not shown). HAT, histone acetyltransferase; HMT, histone methyltransferase; HDAC, histone deacetylase; HDM, histone demethyltransferase; DNMT, DNA methyltransferases. All other abbreviations are listed in Table 6.1. Adapted from (Bierne et al. 2012)
Writers, readers, and erasers are often modular proteins with several properties. The enzyme p300/CBP illustrates this well: it acts as a writer (via its HAT module), a reader (via its bromodomain), and an adaptor (via other modules). In addition, writers, readers, erasers, and remodelers often function as subunits of large macromolecular complexes assembled with scaffold proteins. NurD is a paradigm of such chromatin-remodeling complexes (Fig. 6.2). It contains MTA scaffolding proteins (MTA1, MTA2, MTA3) that bridge subunits involved in nucleosome remodeling (CHD3, CHD4), histone deacetylation (HDAC1, HDAC2), and demethylation (LSD1), binding to other subunits and histones (RBBP4, RBBP7, GATAD2A, GATAD2B), and the targeting of methylated DNA (MBD2) and transcription factors (MBD3) (Lai and Wade 2011). The combinatorial assembly of these subunits determines the function of NuRD in genomic targeting and in the mediation of cell type-specific transcriptional regulations, such as the repression of tumor suppressor genes.
The NuRD and BAHD1 chromatin-repressive complexes. NuRD and BAHD1 complexes are examples of chromatin-associated macromolecular complexes involved in gene repression. Both contain scaffold proteins (MTA1/2/3 and BAHD1-MIER1/2/3, respectively) that bridge subunits involved in histone deacetylation (HDAC1, HDAC2), nucleosome remodeling (CHD3, CHD4), and binding to methylcytosine (5mC) (MBD2 and MBD1, respectively). The BAHD1 complex also contains a HMT subunit (e.g., G9a) that “writes” the H3K9me mark to which the heterochromatin protein HP1 binds. In addition, the BAH domain of BAHD1 is a reader of the H3K27me mark. The function and targeting of these complexes to specific loci depend on the combinatorial assembly of the different subunits with transcription factors (TF) in response to external signals. Upon Listeria infection, the BAHD1 complex assembles at promoters of a set of interferon-stimulated genes (as shown in Fig. 6.3). BAHD1 also controls expression of metabolic genes. Adapted from (Lakisic et al. 2016)
6.2.3 Signaling to Chromatin
The modular, multifunctional, and combinatorial nature of this regulation ensures extremely precise temporal and spatial control over chromatin structure (Ram et al. 2011). The vast array and different combinations of histone PTMs coordinate the sequential recruitment of complexes in a regulatory process that reinforces or reverses existing histone PTMs (Latham and Dent 2007; Lee et al. 2010; Suganuma and Workman 2011). For instance, the Polycomb repressive complexes PRC2 and PRC1 are sequentially recruited, first to “write” H3K27me3 and then to “read” this mark to induce the mono-ubiquitylation of histone H2A, ultimately leading to chromatin compaction at target genes (Margueron and Reinberg 2011). Such cross talk also takes place between histone PTMs and DNA methylation (Cedar and Bergman 2009; Du et al. 2015), via the interaction of histone and DNA modifiers and readers (Li et al. 2006; Vire et al. 2006; Fujita et al. 2003; Ichimura et al. 2005). Cooperation between chromatin-remodeling complexes and DNMTs is particularly important in the maintenance of a specific chromatin state (Cai et al. 2014). The spatial information required to guide chromatin regulators towards specific sites within the genome is provided by combinatorial interactions with DNA-bound transcriptional factors and/or ncRNAs.
Diverse PTMs induced by cell signaling pathways alter the interaction or stability of subunits of chromatin-associated complexes and transcription factors. Signal transduction information is thus translated into chromatin structure in response to various external signals (Mohammad and Baylin 2010; Arzate-Mejia et al. 2011). In particular, phosphorylation and sumoylation are important modifications in the function of chromatin-modifying complexes (Garcia-Dominguez and Reyes 2009; Baek 2011). For instance, phosphorylation of the transcription factor c-JUN by JNK kinase impairs the binding of c-JUN to the MBD3 subunit of the NuRD complex, thereby relieving repression of target genes (Aguilera et al. 2011). Several kinases of signal transduction pathways, such as JNK, MSK1/2, and IKKα, can directly phosphorylate histones (Baek 2011; Tiwari et al. 2011) or histone readers, such as HP1 (Hiragami-Hamada et al. 2011). The signaling molecules activated in cells in response to a wide range of stimuli thus have major effects on the language and syntax of chromatin, through their control of transcription factors and large chromatin-associated co-activator or co-repressor complexes (Fig. 6.1).
6.2.4 Epigenetic Inheritance
Some chromatin modifications remain stable in interphase cells and can be transmitted to daughter cells through mitosis, resulting in their persistence after the disappearance of the initiating signal. This transmission process, resulting in heritable changes in gene expression without altering the sequence of nucleotides in the DNA, defines epigenetic regulation (Riggs et al. 1996).
Epigenetic marks play a key role in cell differentiation, by enabling a cell to “remember” its transcriptional profile. Specific epigenetic signatures fix the identity of the cell while allowing it to respond to external signals. This plasticity explains how the DNA sequence of single cell, the zygote, can generate a huge number of different cell types (about 200 in the human body), most of which being highly differentiated and specialized, whereas others remaining undifferentiated and pluripotent for cell renewal. However, many of the epigenetic marks induced by cell signaling, DNA repair, or cell cycle transitions are short-lived and do not result in long-term memory. This has led to controversy concerning the use of the terms “epigenetic” and “epigenetic marks” to describe chromatin-associated processes and modifications that are not heritable. Adrian Bird has proposed a definition of epigenetic events accounting for both transient and stable modifications of epigenetic language: “a structural adaptation of chromosomal regions so as to register, signal or perpetuate altered activity states” (Bird 2007).
Intense efforts are currently focused on unraveling the mechanisms by which transient epigenetic changes are converted into epigenetic inheritance, particularly given the great importance of these processes in regenerative medicine and research on complex diseases, such as cancer and metabolic and autoimmune diseases. Novel technologies, such as genome-wide epigenomics, chromosome conformation capture (3C), and super-resolution microscopy, have highlighted the complex three-dimensional organization of the genome with large chromatin domains (Guelen et al. 2008; Padeken and Heun 2014; Mattout et al. 2015), the formation of chromosomal loops (Kohwi-Shigematsu et al. 2012; Noordermeer and Duboule 2013), and a nonrandom subnuclear localization of chromatin-associated complexes (Wani et al. 2016). Diverse elements, including enhancers and insulators, regulate topological domains, their boundary regions, and gene looping (Noordermeer and Duboule 2013). The formation of boundaries blocking the spread of heterochromatin is particularly critical for the maintenance of stable gene expression patterns. Recent studies compiling data from a hundred of human epigenomes have highlighted the importance of examining chromatin at the megabase scale and of defining epigenetic profiles for regulatory elements located at some distance from promoters (Roadmap Epigenomics et al. 2015; Romanoski et al. 2015). Such chromatin signatures constitute epigenetic barriers to transcription factor-mediated reprogramming processes. Here, it is worth recalling a groundbreaking discovery in 2006: exogenous expression of a cocktail of transcription factors (Oct4, Sox2, Klf4, and Myc) is sufficient to turn any cell of the body into a pluripotent stem cell (iPS) (Takahashi and Yamanaka 2006). However, this nuclear reprogramming is an inefficient process. Recently, it was reported that depletion of the MBD3 subunit of the NuRD complex greatly improves the efficiency of reprogramming (Rais et al. 2013). This highlights the need to reset the epigenetic landscape of differentiated cells, so they can go back to pluripotency.
In conclusion, the epigenome can change rapidly in response to developmental, physiological, or environmental stimuli, but its stability is also important for the maintenance of cell identity. The mechanisms underlying this plasticity are highly sophisticated. Many studies have shown that bacterial products affect these mechanisms, through the activation of signaling cascades or the direct targeting of chromatin and chromatin regulators in the nucleus, as reviewed below through several examples (Table 6.2). Assessing the magnitude of these effects is an emerging fundamental question, which will also be illustrated here.
6.3 Bacterial Effects on the Host Epigenome
6.3.1 Lessons from Listeria and Anaplasma
Listeria monocytogenes is a food contaminant causing listeriosis, a serious disease for immunocompromised individuals, fetuses, and newborns. This facultative intracellular bacterium is a powerful model to study various aspects of the molecular interactions between pathogen and mammalian cells (Hamon et al. 2006), especially as it invades many different cell types and reaches various organs, such as the liver, spleen, placenta, and brain. In addition, it triggers a wide range of innate immune responses and a potent protective T-cell response (Pamer 2004). Anaplasma phagocytophilum, the causative agent of human granulocytic anaplasmosis, is another interesting bacterial model to address fundamental questions in cellular microbiology, as it displays a remarkable tropism for neutrophils. This obligate intracellular pathogen survives in the hostile environment of the neutrophil by abrogating key antimicrobial functions. This property is partly attributed to A. phagocytophilum’s ability to shape the transcriptional program of the host cell to its advantage (Borjesson et al. 2005; Lee et al. 2008; Sinclair et al. 2014). The studies on Listeria and Anaplasma have proven to be particularly suitable to identify mechanisms involved in chromatin modifications induced by microbial pathogens (Fig. 6.3).
Impacts of Listeria monocytogenes and Anaplasma phagocytophilum on chromatin regulation. (a) Detection of intracellular L. monocytogenes by pattern-recognition receptors (PRR) activates MAPK signaling pathways, leading to histone phosphorylation by MSK1/2 and histone acetylation by p300/CBP and transcriptional activation of pro-inflammatory genes. To control host genes, L. monocytogenes secretes effectors that activate signaling cascades or directly act on the chromatin-regulatory machinery. Toxin LLO induces dephosphorylation and deacetylation of histones via a signaling pathway involving potassium efflux at the plasma membrane. Invasin InlB activates the Met-PI3K pathway, leading to translocation of histone deacetylase SIRT2 into the nucleus and SIRT2-mediated H3K18 deacetylation and repression at a set of defense genes. In epithelial cells, L. monocytogenes infection induces interferon signaling pathways and recruitment of the BAHD1 repressive complex at interferon-stimulated genes by an unknown signal. When bacteria express the lntA gene, the nucleomodulin LntA enters the nucleus where it binds BAHD1, inhibits the BAHD1-HDAC1 silencing complex, restores H3K9 acetylation, and enhances the expression of ISGs. (b) A. phagocytophilum infection activates expression of HDAC1 and DNMT3a genes and induces genome-wide DNA hypermethylation. Anaplasma secretes the nucleomodulin AnkA that binds host DNA at AT-rich motifs overlapping with nuclear matrix attachment regions. One AnkA targeted locus is the CYBB promoter, where AnkA recruits HDAC1, leading to CYBB silencing. Adapted from (Lebreton et al. 2012) and (Rennoll-Bankert and Dumler 2012)
6.3.1.1 Histone Modifications as a Host Response to Bacterial Molecular Patterns
A first effect of L. monocytogenes infection on chromatin has been described in human umbilical vein endothelial cells (HUVECs) cells, in which sensing of cytosolic bacterial molecules by pattern-recognition receptors, such as NOD1, activates MAP-kinases (MAPK). The downstream signaling pathway activates MSK1/2-mediated H3S10 phosphorylation and increased binding of the HAT p300/CBP at the IL-8 gene promoter. The subsequent phosphorylation and acetylation of histones (H3S10p, H4K8ac, and H3K14ac) activate expression of this pro-inflammatory gene (Opitz et al. 2006; Schmeck et al. 2008). This is an illustration of how the host cell responds to an invading pathogen through local change of the chromatin structure at a defense gene (Fig. 6.3).
6.3.1.2 Histone Modifications Induced by Bacteria-Induced Specific Signaling
To control host responses, L. monocytogenes secretes specific effectors that dampen expression of a set of defense genes through activation of cellular signal transduction pathways (Fig. 6.3a). The pore-forming toxin Listeriolysin O (LLO) triggers potassium efflux by forming a pore at the plasma membrane. In human epithelial HeLa cells, this signal promotes a drastic and global deacetylation and dephosphorylation of histones and downregulates expression of a subset of immune genes, encoding for instance the inflammatory cytokine CXCL2, interferon regulatory factor 3 IFIT3, and phosphatase MKP2 (Hamon et al. 2007; Hamon and Cossart 2011). It is worthy to note that LLO also increases phosphorylation of the histone variant H2AX, a marker for DNA damage. The mechanism at play involves degradation of Mre11, a sensor of double-strand DNA breaks involved in DNA repair pathways (Samba-Louaka et al. 2014). Thus, signaling responses to LLO-mediated membrane perforation impact both the genome and epigenome. This suggests a mechanism by which LLO may prime the host cell for genetic and epigenetic changes before bacterial invasion.
Listeria invasion itself impacts chromatin regulation through the action of the internalization protein InlB, a ligand of the tyrosine kinase receptor c-Met. InlB–cMet interaction activates the PI3K-Akt pathway, leading to relocalization of the histone deacetylase SIRT2 from the cytoplasm to the nucleus. SIRT2 represses expression of a set of genes during Listeria infection by catalyzing H3K18 deacetylation at their transcription start sites (Fig. 6.3a). A significant number of these genes are implicated in transcription regulation (i.e., SMAD1, FOXM1, IRF2) and cell signaling (RASGRP1, MAPK14, PIK3R3, PTPNG, SOS1, VAV3, ABL1, CAMK26, MAP2K6, LEF1). The inactivation of the Sirt2 gene in a mouse model of listeriosis has demonstrated the importance of the SIRT2 regulation in Listeria infection (see Sect. 4.1). The link between Akt signaling and SIRT2 is intriguing and remains to be characterized.
6.3.1.3 Direct Control of the Chromatin-Regulatory Machinery by Bacterial Nucleomodulins: The LntA and AnkA Paradigms
Searching for L. monocytogenes effectors targeting intracellular organelles was an opportunity to discover a more active manner used by L. monocytogenes to subvert chromatin regulation processes. This approach led to the identification of Listeria-nuclear-targeted protein A (LntA), a small basic protein that translocates into the nucleus when expressed by intracellular Listeria. LntA interacts with a chromatin repressor, BAHD1 (Bierne et al. 2009; Lebreton et al. 2011), which is a core component of a chromatin-repressive complex that stimulates histone modifications, DNA methylation, and chromatin compaction (Libertini et al. 2015; Lakisic et al. 2016). The BAHD1-associated complex displays analogy with NuRD: it contains BAHD1 and MIER proteins that share structural features with the scaffold proteins MTAs of NuRD and bridge together chromatin writers, erasers, and readers mostly involved in gene repression (Fig. 6.2). As for NurD, the set of genes repressed by the BAHD1 complex depends on the cell type, as well as on the signal to which cells are submitted. Upon infection of epithelial cells with L. monocytogenes, BAHD1 represses Interferon-Stimulated Genes (ISGs) (Lebreton et al. 2011), which are important players in the innate immune response (Dussurget et al. 2014). When L. monocytogenes expresses lntA, the secreted factor LntA enters the nucleus and alleviates BAHD1 and HDAC1/2 binding to ISG promoters, leading to histone deacetylation and upregulation of ISG expression (Fig. 6.3a). LntA interacts directly with a central proline-rich region of BAHD1, via a surface patch containing a dilysine motif (K180/K181), located nearby a groove on the elbow region of LntA identified by crystallography (Lebreton et al. 2014). Mutation of this strategic dilysine abolishes LntA binding to BAHD1 and LntA-mediated stimulation of interferon responses upon infection. Inactivation or overexpression of lntA in bacteria, as well as knockdown of Bahd1 in the mouse (see Sect. 4.1), alters the infectious process in vivo (Lebreton et al. 2011). However, the signaling pathways that govern BAHD1 and LntA synthesis and the loading of these factors onto chromatin nearby ISGs are unknown. Thus, several questions remain to be addressed to understand how the LntA-BAHD1 interplay modulates the interferon (IFN) response in time and space during bacterial colonization of the host.
The study of LntA enabled to define the family of nucleomodulins, which encompasses bacterial effectors acting on nuclear processes after translocation into the nucleus (Bierne and Cossart 2012). A. phagocytophilum produces several nucleomodulins (Sinclair et al. 2015a). The extensive characterization of one of them, Ankyrin A (AnkA), has provided other conceptual advances on mechanisms by which bacterial actors may act on chromatin. AnkA is a large bacterial effector characterized by a central region containing ankyrin (Ank) repeats (Park et al. 2004; Garcia-Garcia et al. 2009b). Interestingly, Ank repeats are commonly found in eukaryotic proteins, notably in several nuclear proteins that bind transcription factors. Following its secretion in the cytoplasm by a bacterial type IV secretion system (T4SS), AnkA enters the granulocyte nucleus, binds stretches of AT-rich DNA, and alters transcription of antimicrobial defense genes. In particular, AnkA represses CYBB, which encodes the subunit beta (NOX2) of the NADPH oxidase (Garcia-Garcia et al. 2009b; Rennoll-Bankert and Dumler 2012). The mechanism at play involves binding of AnkA to DNA in the CYBB promoter region, direct recruitment of HDAC1 by AnkA, and deacetylation of H3 (Rennoll-Bankert et al. 2015) (Fig. 6.3b). As a consequence, the pathogen obtains a significant fitness advantage as it prevents superoxide anion production by the NADPH oxidase and associated bactericidal effects.
AnkA not only binds the CYBB locus. It also targets several DNA regions rich in AT nucleotides on distinct chromosomes (Park et al. 2004; Garcia-Garcia et al. 2009b). Remarkably, AnkA-binding sites overlap within matrix attachment regions (MARs) that serve as attachment sites for nuclear matrix proteins and mediate structural organization of the chromatin within the nucleus (Rennoll-Bankert et al. 2015). AnkA is in this way a functional mimic of the host MAR-binding protein SATB1, which is known to bind to the CYBB promoter and represses transcription early during myeloid differentiation by recruiting HDACs (Wang et al. 2010). SATB1 has a wide action on chromatin, as it contributes to the formation of nuclear architectural platforms that anchor hundreds of gene loci and control large-scale transcriptional reprogramming (Kohwi-Shigematsu et al. 2012). This opens the fascinating hypothesis that bacterial effectors like AnkA could act as global genome organizers both acting in cis (locally) and trans (at a distance) to a target gene. By controlling the dynamics of chromosomal looping, they may change the three-dimensional structure of chromatin (Sinclair et al. 2014). Recent mapping of AnkA binding sites on the neutrophil genome by ChIP-seq further supports this concept of microbial factors acting as genome “re-organizers” (Dumler et al. 2016). Also in line with this idea, there is evidence that BAHD1-mediated heterochromatin formation plays a role in the spatial architecture of the genome (Libertini et al. 2015). Thus, LntA-mediated inhibition of BAHD1 might also change the structure of large domains involved in the co-regulations of ISGs upon L. monocytogenes infection.
6.3.1.4 Deregulation of Epigenetic Factor Genes and Genome-Wide Mediated Epigenetic Changes
Numerous neutrophil genes are differentially expressed during A. phagocytophilum infection (Borjesson et al. 2005; Lee et al. 2008; Sinclair et al. 2014). Several of them are downregulated, coinciding with HDAC1 binding and H3 deacetylation at their promoters (Garcia-Garcia et al. 2009a, b). However, most of them are upregulated (Borjesson et al. 2005), in agreement with the complex effects of infection on host gene expression. It was recently shown that DNA methylation levels in the neutrophil genome are profoundly altered after 24 h of infection with A. phagocytophilum. In particular, many regions within 3 kb from gene transcriptional start and termination sites and at intron–exon junctions become hypermethylated. In addition, expression of the HDAC1 and DNMT3A genes is increased with infection (Garcia-Garcia et al. 2009a; Borjesson et al. 2005)(Fig. 6.3b). Overall, these findings highlight that Anaplasma infection induces large epigenomic changes as a result from the combined action of diverse mechanisms, including changes in expression of epigenetic factors and cross talk between these factors. Pharmacologic inhibition of histone deacetylases or DNA methyltransferases decreases Anaplasma intracellular survival (Garcia-Garcia et al. 2009a; Sinclair et al. 2015b), supporting the notion that broad epigenetic changes contribute to disease.
In summary, the Listeria and Anaplasma paradigms illustrate the diversity of mechanisms involved in modification of chromatin structure during bacterial infection, both at local and large genomic scales.
6.3.2 Chromatin Modifications Driven by Bacteria: Additional Examples
6.3.2.1 Histone Modifications
As shown for Listeria, bacteria in contact with eukaryotic cells have the ability to activate a large repertoire of host signaling pathways (e.g., MAPKs, NF-κB, and PI3K pathways) acting on histone kinases and acetylases (Yamamoto et al. 2003; Baek 2011). This is particularly the case of pro-inflammatory pathways. For instance, Moraxella catarrhalis, a saprophytic bacterium of the respiratory tract, and Bacteroides vulgatus, a commensal of the intestinal flora, induce inflammatory signaling cascades leading to phosphorylation/acetylation of H3 (Haller et al. 2003; Slevogt et al. 2006). The gastric pathogen Helicobacter pylori secretes the peptidyl prolyl cis-, trans-isomerase HP0175 that activates a TLR4-MAPK-MSK1 pathway leading to H3 phosphorylation and activation of the pro-inflammatory gene IL-6 in THP-1 monocytes (Pathak et al. 2006). Bacterial lipopolysaccharide (LPS) and flagellin trigger histone acetylation and phosphorylation events at the IL-8 gene downstream of the NF-κB pathway (Saccani et al. 2002; Schmeck et al. 2008).
However, inflammation is often counteracted by bacteria-driven mechanisms. In the case of B. vulgatus, this is performed by induction of the TGF-β1 anti-inflammatory pathway, which in turn induces H3 deacetylation and gene silencing via HDAC recruitment at pro-inflammatory gene promoters (Haller et al. 2003). This mechanism prevents B. vulgatus from eliciting a strong inflammatory response in the gut and contributes to its tolerance by the host. Bacterial toxins also dampen the host innate immune responses by inhibiting H3 phosphorylation/acetylation events. As described above for Listeria LLO (Fig. 6.3a), the pore-forming toxins PFO of Clostridium perfringens, PLY of Streptococcus pneumoniae, and aerolysin from Aeromonas hydrophila share a common mechanism that modulates histone marks, and subsequent gene expression, by acting on intracellular potassium levels (Hamon et al. 2007). Lethal toxin (LT) from Bacillus anthracis, the agent of anthrax, uses another mechanism by cleaving and inactivating MAPKKs, leading to disruption of MAPK signaling (Bardwell et al. 2004). In lung epithelial cells activated by TNF-α, LT-mediated MAPK inhibition promotes a decrease in the levels of H3S10p and H3K14ac at the promoters of IL-8 and KC genes (Raymond et al. 2009). In macrophages exposed to LT, MAPK inhibition induces expression of HDAC8, which results in a decrease of H3K27ac levels at one enhancer of the IL1-β gene and the subsequent repression of this gene (Ha et al. 2016).
It is interesting to notice that besides immunity and inflammatory genes, changes in histone PTMs can also enable a pathogen to control expression of host genes involved in cell proliferation and death, as illustrated by the carcinogenic bacterium H. pylori. In gastric epithelial cells, this pathogen induces transient dephosphorylation of H3S10 and H3T3, as well as deacetylation of H3K23 (Fehri et al. 2009; Ding et al. 2010a). These modifications impact both the cell cycle (Fehri et al. 2009) and transcription of the oncogene c-JUN and heat shock gene hsp70 (Ding et al. 2010a). In addition, H. pylori-mediated pre-mitotic arrest involves dephosphorylation of H3S10 upon deregulation of the mitotic histone kinase VRK1, followed by rephosphorylation of H3S10 by an IKKα-dependent pathway. Furthermore, exposure of H. pylori to gastric epithelial cells promotes release of HDAC1 from the promoter of the cell cycle regulator gene p21 WAF, hyper-acetylation of H4, and increased expression of p21 WAF (Xia et al. 2008). These mechanisms may contribute to various H. pylori-associated gastric pathologies, including ulcers, mucosa-associated lymphoid tissue lymphoma, and cancer.
IFN responses are also modulated by diverse chromatin-based mechanisms. Like Listeria, Mycobacterium tuberculosis (Mtb), the causative agent of tuberculosis, controls histone-modifying multiprotein complexes at IFN-responsive genes (Lebreton et al. 2012). In macrophages infected with Mtb or exposed to Mtb components, such as the lipoprotein LpqH, genes induced in response to IFN-γ are partly repressed (Wang et al. 2005; Pennini et al. 2006). These genes include CIITA, coding for the master regulator of MHC class II genes, as well as some of its targets (e.g., HLA-DR). Activation of the TLR2-MAPK-dependent pathway upon Mtb infection stimulates recruitment of the transcriptional repressor C/EBP and histone deacetylation at the promoter of CIITA, antagonizing the nucleosome-remodeling activity of the SWI/SNF complex and downregulating CIITA expression (Pennini et al. 2007). Additionally, mycobacterial infection upregulates the expression of SIN3A, which encodes a core subunit of a HDAC-associated macromolecular complex (Wang et al. 2005) related to NuRD and BAHD1 complexes. Thus, to counteract IFN-γ-induced pathways, Mtb not only silences CIITA but also CIITA-regulated genes, such as HLA-DR, upon increased recruitment of SIN3A-HDACs to their promoters.
6.3.2.2 DNA Methylation
The importance of DNA methylation events associated with bacterial infections is also becoming increasingly appreciated. However, as for histone PTMs, alteration of 5mC patterns can result from an amalgam of bacteria-driven and host-driven effects on chromatin. Moreover, it can be difficult to connect gain or loss of this epigenetic mark to transcriptional changes. Indeed, the transcriptional effects of 5mC marks depend on their localization. Gain of DNA methylation is mainly coupled with transcriptional silencing at CpG-rich regions in promoters and enhancers and transcriptional activation at CpG-poor regions in gene bodies (Klose and Bird 2006; Chen and Riggs 2011). Interpreting the effect of bacteria on cytosine methylation is thus complex. This is illustrated by genome-wide studies of Mycobacterium-induced DNA methylation changes. On one hand, the response of human dendritic cells to Mtb infection is accompanied by both widespread de novo methylation and active demethylation primarily at enhancer elements (Pacis et al. 2015). On the other hand, the response of human THP-1 macrophages to Mtb infection is mostly accompanied by hypermethylation predominantly at cytosines present in a non-CpG dinucleotide context (Sharma et al. 2016). These differences may be explained by the use of different host cell models and infection times. Several DNA methylome maps have probably to be drawn in order to assess with robustness the dynamics of cytosine methylation during infection.
H. pylori infection also induces aberrant DNA methylation. In the human gastric mucosa, changes in 5mC patterns upon infection have been identified, strikingly at promoters of genes found methylated in gastric cancer cells (Maekita et al. 2006; Ding et al. 2010b; Hattori and Ushijima 2016). H. pylori-associated hypermethylation occurs for instance at the E-cadherin gene CDH1 (Chan et al. 2003), tumor suppressor genes (e.g., USF1/2 and WWOX (Bussiere et al. 2010; Yan et al. 2011), DNA repair genes [e.g., MLH1 (Yao et al. 2006)], as well as in CpG islands of miRNA genes (Ando et al. 2009). The ability of H. pylori to induce DNA methylation in the gastric mucosa was confirmed in the gerbil animal model, and, interestingly, this effect was relieved upon treatment with the immunosuppressor cyclosporin A (Niwa et al. 2010). Moreover, H. pylori-mediated inflammation triggers lymphocyte and macrophage infiltration, which appears to have a key role in induction of DNA methylation (Hur et al. 2011). It is currently believed that DNA methylation changes upon H. pylori infection are mostly the indirect consequence of the associated inflammatory responses (Hattori and Ushijima 2016). Signals from macrophages produced by chronic inflammation, such as IL-1β, TNF-α, or nitric oxide, may affect factors that protect DNA from methylation, such as TET proteins (Hattori and Ushijima 2016). It remains unclear whether H. pylori effectors contribute more directly to aberrant epigenetic changes during gastric cancer progression (Valenzuela et al. 2015). There is a growing number of studies showing that epigenetic changes, particularly in DNA methylation, are linked to an increased inflammatory response (Bayarsaihan 2011; Medzhitov and Horng 2009), as well as increased risk of chronic disease development and cancerization. The role of bacteria in shaping patho-epigenetic landscapes will be discussed in detail in Sect. 5.
Epithelia other than that of the stomach can undergo bacteria-induced DNA methylation changes. There is evidence that in the oral cavity bacterial-induced chronic infection and uncontrolled inflammatory response may trigger epigenetic modifications. As an illustration, periodontally inflamed gingival biopsies showed a significant increase in promoter methylation of the gene encoding the pro-inflammatory enzyme COX-2, compared with non-inflamed biopsy samples (Zhang et al. 2010). This would allow a chronic inflammatory stimulus to be tolerated, preventing unrestricted tissue destruction. Whether this is a bacteria-triggered phenomenon is unknown, but it is noteworthy that resident bacteria, such as Porphyromonas gingivalis, can induce hypermethylation of specific genes in gingival epithelial cells (Yin and Chung 2011).
In human uroepithelial cells, infection with uropathogenic Escherichia coli (UPEC) results in the upregulation of DNMT activity and DNMT1 expression and induces CpG methylation and downregulation of CDKN2A, a G1 cell cycle inhibitor regulator (Tolg et al. 2011). This may increase uroepithelial cell proliferation and pathogen persistence, by counteracting infection-stimulated host cell apoptosis. The placenta can also be targeted by bacteria-mediated epigenetic changes. Indeed, maternal infection with Campylobacter rectus induces hypermethylation of the imprinted IGF2 gene promoter in murine placental tissue (Bobetsis et al. 2007). This finding suggests that bacterial infections during pregnancy might epigenetically affect genes involved in fetal development.
Last but not least, there is evidence that nonpathogenic inhabitants of the gut shape the DNA methylome. The gene TLR4, which encodes a LPS-sensing receptor, is downregulated in intestinal epithelial cells, and a role of the commensal bacteria in TLR4 methylation and silencing is suspected (Takahashi et al. 2011). This is proposed to maintain intestinal homeostasis by preventing an excessive inflammatory reaction to the gut microbiota.
6.3.3 Bacterial Nucleomodulins
6.3.3.1 Nucleomodulins Acting Via Protein–Protein or Protein–DNA Interactions
As discussed above, L. monocytogenes LntA is a paradigm for nucleomodulins acting as inhibitor of HDAC-associated complexes, while A. phagocytophilum AnkA is a paradigm for nucleomodulins binding DNA and recruiting HDAC-associated complexes. So far, LntA orthologs have not been identified in other bacterial species, at least at the level of the primary protein sequence. In contrast, Ank-containing proteins are present in several human intracellular bacterial pathogens, such as Ehrlichia, Rickettsia, Orientia, Coxiella, and Legionella species. In particular, the protein Ank200 (or p200) from Ehrlichia chaffeensis (Wakeel et al. 2010) binds Alu-Sx elements located in promoters and introns of various human genes (Zhu et al. 2009). Several p200 target genes are strongly upregulated during infection, suggesting that p200 may affect gene transcription at a large genomic scale through mechanisms associated with Alu element gene regulation. Several other tandem-repeat containing proteins (TRPs) from E. chaffeensis may also enter into the nucleus (Luo et al. 2011; Luo and McBride 2012). Of those, the E. chaffeensis 32-kDa and 120-kDa tandem repeat proteins, TRP32 and TRP120, are nucleomodulins that binds host cell DNA particularly at G-rich motifs (Luo et al. 2011; Farris et al. 2016). Genes targeted by TRP120 are most frequently associated with transcriptional regulation, signal transduction, and apoptosis, whereas those targeted by TRP32 are linked to immune cell differentiation, chromatin remodeling, and RNA transcription. Interestingly, like many host nuclear factors, these nucleomodulins are subjected to post-translational modifications, TRP32 being phosphorylated and TRP120 sumoylated in host cells (Dunphy et al. 2014; Farris et al. 2016). Nucleomodulins may not only bind DNA and chromatin factors but also chromatin-anchoring factors. SinC, a protein secreted by Chlamydia psittaci via a type III secretion system (T3SS), exemplifies this potential mechanism. This effector targets the inner membrane of the nucleus in infected cells and may control chromatin interaction with the nuclear lamina (Mojica et al. 2015).
6.3.3.2 Nucleomodulins Acting as Epigenetic Modifiers
Several bacterial pathogens, and particularly those living in intracellular vacuolar compartments, can alter host chromatin structure by producing mimics of chromatin-modifying enzymes (Fig. 6.4). A first of such bacterial mimics, NUE, is a HMT discovered in the human pathogen Chlamydia trachomatis, based on its sequence similarities with eukaryotic lysine-specific methyltransferases containing a SET domain. After secretion by a T3SS, NUE enters the nucleus and associates with chromatin (Pennini et al. 2010). However, while NUE methylates mammalian histones in vitro, its target genes in the infected cell remain unknown.
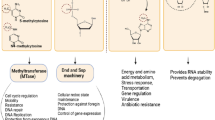
Bacterial nucleomodulins with enzymatic activities. Several bacterial pathogens inject bacterial effector proteins into the host nucleus, and some are enzymes that modify chromatin residues or regulators. 1. Legionella pneumophila, Burkholderia thailandensis, and Chlamydia trachomatis secrete SET domain containing effectors via type 3 (T3SS; Burkholderia; Chlamydia) or type 4 (T4SS; Legionella) secretion systems. L. pneumophila secretes a histone methyltransferase (LpSET) that has been assigned two functions: (i) in the nucleus, LpSET termed “RomA” trimethylates histone H3 at K14, causing a switch from acetylated to methylated H3K14 at specific gene promoters and thus transcriptional repression, and (ii) in the nucleolus, LpSET termed “LegAS4” binds HP1 at rDNA promoters and activates transcription by stimulating H3K4 methylation. B. thailandensis secretes a LegAS4-like protein (BtSET) that also activates rDNA transcription in the nucleolus. 2. C. trachomatis secretes the histone methyltransferase NUE that methylates host histones H2B, H3, and H4. Bacillus anthracis produces a histone H1 methyltransferase, BaSET. Mycoplasma hyorhinis Mhy1, Mhy2, and Mhy3 are nucleomodulins with CG- and GATC-specific cytosine methyltransferase activities. 3. Mycobacterium tuberculosis secretes at least three nucleomodulins: Rv1988 is a histone methyltransferase that methylates H3 on R42 in the nucleosome core; Rv3423 is a histone acetyltransferase; and Rv2966c is a DNA methyltransferases. 4. The Shigella flexneri T3SS effector OspF is a posttranslational modifier with a phosphothreonine lyase activity. OspF eliminylates MAP-kinases in the nucleus, leading to the downregulation of a subset of immunity genes. 5. The T3SS effector NleC from pathogenic E. coli is a metalloproteinase targeting the host histone acetyltransferase p300 for degradation. Ac Acetylation, Me methylation, P phosphorylation, E eliminylation. Adapted from (Bierne 2013)
Other SET domain-containing proteins were thereafter identified in Legionella pneumophila (Rolando et al. 2013; Li et al. 2013), Burkholderia thailandensis (Li et al. 2013), and Bacillus anthracis (Mujtaba et al. 2013) (Fig. 6.4). In L. pneumophila, LpSET is characterized as a bacterial HMT with dual functions. In L. pneumophila strain Paris, LpSET (named RomA: “Regulator of methylation A”) has been shown to act as a HMT that trimethylates K14 of H3 (H3K14me3), a modification that does not exist in mammals (Rolando et al. 2013). By promoting a burst of H3K14me3 genome wide, including at innate immune gene loci, RomA decreases H3K14 acetylation, which is an activating mark, thus leading to repression of host gene expression and playing an important role in bacterial replication inside macrophages. In a separate study performed with L. pneumophila strain Philadelphia, LpSET (named LegAS4) was reported to act in the nucleolus on the expression of ribosomal RNA (rRNA) (Fig. 6.4). Human cells contain several hundred rDNA genes organized in tandem repeats that are clustered into nucleolar organizer regions. The chromatin structure of rRNA genes plays a fundamental role in regulating transcription of rDNA loci. LegAS4 binds rDNA at promoter and intergenic-spacer regions, by interaction with the chromatin reader HP1. In vitro studies suggest that LegAS4 catalyzes dimethylation of histone H3 on lysine 4 (H3K4me2). Consistently, ectopic expression of LegAS4 in human cells is associated with increased levels of H3K4me2 at rDNA promoters and activation of the transcription of these genes (Li et al. 2013). B. thailandensis secretes a LegAS4-like protein (BtSET) that also activates rDNA transcription in the nucleolus. Stimulation of rDNA expression and increased 45S pre-RNA synthesis seems to contribute to bacterial replication, though the mechanism at play is not yet understood. Having different substrates and functions is a property that LpSET shares with eukaryotic HMTs. For instance, the H3K9 HMT G9a preferentially methylates K9 on histone H3 but can also methylate K27 and K56. This HMT predominantly represses genes at euchromatic regions but also acts as a positive activator of rDNA transcription (Yuan et al. 2007). Considering the number of SET domain proteins present in bacterial species that interact with eukaryotes, it is tempting to speculate that several bacteria might employ this strategy.
The agent of tuberculosis also modulates the host epigenetic machinery by secreting an original HMT, here methylating a noncanonical arginine located in the core of histone H3 (Yaseen et al. 2015) (Fig. 6.4). This effector, Rv1988, targets genes involved in defense against pathogens, including genes participating in the generation of reactive oxygen species (ROS). During infection, Rv1988 not only targets gene promoters for H3R42me2 but also putative regulatory regions. Aside from HMTs, MtB also secretes effectors with histone acetyltransferase [Rv3423.1 (Jose et al. 2016)] or DNA methyltransferase [(Rv2966c, (Sharma et al. 2015)] activity. It is interesting to notice that Rv2966c methylates cytosines present in a non-CpG context. Thus, MtB has evolved diverse strategies to directly manipulate chromatin in the nucleus. Mycoplasma species also produce nucleomodulins. For instance, three DNA methyltransferases have been identified in Mycoplasma hyorhinis, an intracellular commensal that can shift to an opportunist pathogen. M. hyorhinis produces Mhy1 and Mhy2, promoting CG methylation, and Mhy3 acting on GATC sites (Chernov et al. 2015). There is evidence that these bacterial DNMTs have the ability to translocate to the human cell nucleus and establish aberrant genome-wide methylation patterns. Yet, it remains to be proven that the host epigenome is reshaped in human cells naturally infected by M. hyorhinis.
Bacteria-induced epigenetic effects can also occur by specific modifications of epigenetic factors. The Shigella flexneri T3SS effector OspF nicely illustrates this mechanism. OspF is a phosphothreonine lyase that irreversibly modifies host MAPKs by eliminylation (Li et al. 2007; Brennan and Barford 2009). This enzymatic reaction converts a phosphothreonine residue into a dehydrobutyrine residue that can no longer be phosphorylated and hence locks the substrate in an inactive form. Inhibition of MAPK phosphorylation in the nucleus enables OspF to abrogate phosphorylation of histone H3 at a set of NF-κB-regulated promoters, thus impairing expression of a pool of pro-inflammatory genes (Arbibe et al. 2007). In addition, this effector alters the phosphorylation at S83 of the heterochromatin protein HP1-γ, demonstrating that in addition to histones, bacteria can control chromatin regulator PTMs (Harouz et al. 2014) (Fig. 6.4). Furthermore, OspF and another nuclear-targeted effector, OspB, interact with the human retinoblastoma protein Rb, which is known to bind several chromatin-remodeling factors (Zurawski et al. 2009). Shigella likely uses OspF–OspB synergy to downregulate host innate immunity via alteration of the chromatin structure at specific genes.
S. flexneri also secretes an effector acting as an E3 ubiquitin ligase, IpaH9.8, which targets several host cytosolic or nuclear proteins for proteasome-dependent degradation (Rohde et al. 2007). In the nucleus, IpaH9.8 disrupts the activity of a mRNA splicing factor, U2AF35, thus interfering with U2AF35-dependent splicing (Toyotome 2001; Seyedarabi et al. 2011) and impairing host inflammatory responses (Okuda et al. 2005). IpaH9.8 defines a family of bacterial effectors characterized by an N-terminal domain containing leucine-rich repeats (LRR) involved in substrate recognition and a C-terminal E3 ligase domain (Hicks and Galan 2010). The ortholog of IpaH9.8 in Salmonella enterica is SspH1, a nucleomodulin that targets for instance the host kinase PKN1 (Haraga and Miller 2003; Rohde et al. 2007). Yersinia pestis also encodes a LRR-containing nucleomodulin targeting host kinases, YopM (Benabdillah et al. 2004; Soundararajan et al. 2011). However, YopM is not itself a modifier but rather acts as a scaffolding protein that facilitates the formation of a complex between serine/threonine kinases RSK1 and PKN2 (McDonald et al. 2003; McCoy et al. 2010). Recent data suggest that the YopM causes enhanced phosphorylation of RSK1 in the nucleus, leading to enhanced transcription of immunosuppressive cytokines, such as IL-10. YopM intranuclear levels are dependent on its interaction with the DEAD-box helicase 3 (DDX3) (Berneking et al. 2016).
Other types of bacterial modifiers acting in the nucleus include proteases and phosphatases. The T3SS effector NleC from enteropathogenic (EPEC) and enterohemorrhagic (EHEC) E. coli is a zinc metalloprotease that targets the HAT p300/CBP and decreases the abundance of this epigenetic factor in the nucleus (Fig. 6.4). Overexpression or knockdown of NleC impacts IL-8 secretion by EPEC, indicating that NleC contributes to dampening of inflammatory signaling during infection (Shames et al. 2011). The Gram-positive bacterium Streptococcus pyogenes expresses a serine/threonine phosphatase, SP-STP, which is secreted into host cells and targets the nucleus (Agarwal et al. 2012). There, it acts as a pro-apoptotic factor that induces apoptosis of pharyngeal cells, a hallmark of streptococcal infections, by influencing transcription of apoptotic genes and preventing the transcription of other genes, such as cytochrome p450.
6.3.4 Change in Expression and/or Activity of Epigenetic Regulators
As illustrated above with upregulation of HDAC1 in A. phagocytophilum-infected cells (Garcia-Garcia et al. 2009a) and SIN3a in Mtb-infected cells (Wang et al. 2005), some bacterial species positively or negatively modulate expression of epigenetic factors. Mtb infection induces HDAC1 expression in macrophages (Chandran et al. 2015). In contrast, the levels of HDAC1 and DNMT1 transcripts decrease in gingival epithelial cells treated with the oral pathogen Porphyromonas gingivalis (Yin and Chung 2011), and LPS from this bacterial species downregulates DNMT1, DNMT3a, and JMJD3 gene expression levels (de Camargo Pereira et al. 2013). M. catarrhalis also reduce HDAC1/2 expression in bronchial epithelial cells (Slevogt et al. 2006). In human urothelial cells, infection with UPEC results in the upregulation DNMT1 expression (Tolg et al. 2011), as well as of EZH2, encoding the H3K27 HMT of the Polycomb chromatin-repressive complex PRC2 (Ting et al. 2016). EZH2 plays a role in early host cell proliferative responses to infection. Bacteria can also produce metabolites, acting as inhibitors of chromatin-modifying enzymes, such as lactate and butyrate, proven to be potent inhibitors of HDACs (Latham et al. 2012).
6.3.5 Bacterial Molecular Patterns Acting on the Epigenetic Machinery
Certain bacterial-derived metabolites can modify the epigenome of host cells and in turn alter the development and function of the cell, either by acting on precursors of enzymatic reactions involved in chromatin modifications or by modulating the activity of epigenetic regulators (Alenghat and Artis 2014). In particular, commensals of the microbiota produce diet-dependent molecules that influence DNA methylation and histone acetylation. One such product is butyrate, a potent inhibitor of HDACs (Riggs et al. 1977). Butyrate exerts beneficial anti-inflammatory effects on the host, particularly on immune cells (Segain et al. 2000; Arpaia et al. 2013; Chang et al. 2014), possibly via epigenetic upregulation of anti-inflammatory genes (see Sect. 4.1). Such observations open the interesting possibility to use butyrate-producing probiotic bacteria as immunosuppressors (Licciardi et al. 2010). Interestingly, a recent study has shown that a bacterial quorum-sensing molecule can also act on HDACs. Prolonged exposure to 2-aminoacetophenone, which is excreted by the pathogen Pseudomonas aeruginosa, increases the expression and activity of HDAC1, resulting in hypoacetylation of H3K18 and attenuated expression of pro-inflammatory cytokine genes. This is proposed to promote host tolerance to infection (Bandyopadhaya et al. 2016).
6.4 Epigenetic Factors Engaged in Host Responses to Bacteria and Bacterial Imprinting
6.4.1 Lessons from BAHD1-, SIRT2-, and HDAC3-Deficient Mice
Considering the crucial role of epigenetic factors in embryonic development and cell differentiation, it is not surprising that knocking out their coding genes in the mouse often causes embryonic or perinatal lethality. Thus, so far only a few studies have addressed the role of epigenetic regulators in bacterial infections at the level of a mammalian organism. The use of Bahd1 haplo-deficient (heterozygous) mice (Lebreton et al. 2011) and Sirt2 knockout mice (Eskandarian et al. 2013) has confirmed the involvement of BAHD1 and SIRT2 in murine listeriosis. Following intravenous inoculation with L. monocytogenes, the spleens of these deficient mice were significantly less infected than those of wild-type littermates after 72 hours of infection. It was not possible to further refine the role of BAHD1 by infecting Bahd1 knockout mice, because the total ablation of Bahd1 induces a high neonatal mortality rate (Lakisic et al. 2016). However, it is worthy to note that it also causes restriction of placental growth, indicating a key role of BAHD1 in placental development. This finding opens the possibility that manipulation of the BAHD1 complex by Listeria contributes to the fetoplacental step of listeriosis. Bahd1 deficiency also leads to deregulation of carbohydrate and lipid metabolism, which may play roles in infection (Lakisic et al. 2016). Together these results support the notion that mutations in epigenetic regulatory genes influence the outcome of bacterial infections.
The use of HDAC conditional knockout mice permitted to investigate the functional roles of chromatin regulation in intestinal homeostasis and its cross talk with the microbiota. Intestinal epithelial cells (IECs) integrate numerous microbial signals from the intestinal microenvironment and respond to these stimuli by changing their transcriptional program. Interestingly, IEC-specific Hdac3-deficient mice show increased susceptibility to intestinal damage and inflammation and a change in microbial communities (i.e., dysbiosis) (Alenghat et al. 2013). Strikingly, when rendered germ free, Hdac3-conditional KO mice recover a normal intestinal barrier function, as observed in wild-type germ-free mice. These data indicate that HDAC3 has an important role in maintaining intestinal homeostasis and establishing normal host–commensal relationships. Ablating both Hdac1 and Hdac2 in murine IECs also alters the structure and functions of the gut, via a defect in cell differentiation and chronic intestinal inflammation (Turgeon et al. 2013). While a possible dysbiosis induced by this double mutation has not yet been addressed, it is possible that major alterations of the gut induced by the double Hdac1–Hdac2 mutation impacts the composition of the population of commensals. Thus, HDACs are likely to be key epigenetic programmers of the host in response to signals from the gut microbiota.
It is important to note that while loss of Hdac3 expression in IECs impairs microbiota-dependent intestinal barrier function, inhibition of HDACs by commensal bacteria-derived SCFAs, such as butyrate, generally protects from pathologic intestinal inflammation. However, rather than acting on epithelial cells, butyrate is described to inhibit HDAC function in intestinal immune cells, such as peripheral blood mononuclear cells (PBMCs) (Segain et al. 2000), Tregs (Arpaia et al. 2013), and macrophages (Chang et al. 2014). These results suggest that HDACs may have opposite effects in different intestinal cell populations, leading to either protective or pathologic immunity (Alenghat and Artis 2014).
6.4.2 Immune Tolerance and Toxin-Induced Resistance
Host–commensal mutual relationships require that commensal bacteria do not trigger an uncontrolled immune response, and thus become tolerated by the host immune system, while the latter efficiently eliminates invading pathogens. HDACs and other epigenetic regulators are involved in this tolerance. For instance, intestinal commensal bacteria induce DNA methylation at the gene encoding the main sensor of LPS, TLR4, leading to its downregulation in the large intestine (Takahashi et al. 2011). This is believed to maintain intestinal homeostasis by preventing an excessive inflammatory reaction to the gut microbiota. The commensal bacterium Bacteroides vulgatus triggers an anti-inflammatory response via recruitment of HDACs at pro-inflammatory gene promoters (Haller et al. 2003). Likewise, Bifidobacterium breve and Lactobacillus rhamnosus GG inhibit transcriptional activation of inflammatory bowel disease-causing factors through inhibition of histone acetylation and enhancement of DNA methylation (Ghadimi et al. 2012).
In the presence of opportunist pathogens, similar mechanisms may dampen uncontrolled inflammatory responses triggered by bacteria-induced chronic infections. For instance, in the oral cavity, periodontally inflamed gingival biopsies show a significant increase in promoter methylation of the gene encoding the pro-inflammatory enzyme COX-2, when compared with non-inflamed biopsy samples (Zhang et al. 2010). This would allow a chronic inflammatory stimulus to be tolerated, preventing unrestricted tissue destruction. Whether this is a bacteria-triggered phenomenon remains unknown, but it is remarkable that resident bacteria, such as P. gingivalis, can induce DNA hypermethylation of specific genes in gingival epithelial cells (Yin and Chung 2011).
When pathogenic species manage to cross epithelial barriers and to multiply in the blood, sustained exposure to microbial inflammatory products, such as LPS, leads to tissue damage, multi-organ dysfunction, septic shock, and death. To compensate these adverse effects, the immune system has developed post-septic immunosuppression (PSI) mechanisms that enable hematopoietic cells to become hypo-responsive to repeated stimulation by microbial insults. This compensatory anti-inflammatory response counteracts the harmful effects of sepsis but leaves individuals more susceptible to opportunistic infections for extended periods of time (weeks to years). Although PSI is a complex multifactorial process, the contribution of epigenetic regulation is recognized (McCall et al. 2010; Carson et al. 2011). One of the facets of PSI is LPS tolerance, in which LPS-elicited TLR4 responses are reprogrammed towards silencing of pro-inflammatory cytokine genes and expression of anti-inflammatory or antimicrobial mediators. LPS activation of TLR4 first elicits transcription of poised pro-inflammatory genes, which are rapidly derepressed and then returned to basal state within hours. Opening the chromatin at target genes during this acute phase involves histone phosphorylation and acetylation. However, sustained exposure to LPS or subsequent LPS challenge activates a pathway leading to permanent gene repression (Fig. 6.5). One mechanism was studied at the proximal promoters of TNF-α and IL1-β genes, where a change in the composition of the NF-κB transcription factor occurs (El Gazzar et al. 2008; Chen et al. 2009). LPS-mediated upregulation of RelB expression induces a shift from activating p65-p50 TF to repressive RelB-p50 TF. RelB interacts with H3K9 HMT G9a, leading to H3K9me2 and subsequent recruitment of HP1. The repressive complex formed by G9a and HP1 recruits DNMT3A/B, which induces de novo CpG methylation and assembly of silent, facultative heterochromatin (McCall et al. 2010). Interestingly, LPS stimulation has also been shown to leave epigenetic footprints on enhancers (Ostuni et al. 2013). As discussed in Sect. 2.4, the epigenetic topography of distal regulatory elements, such as enhancers and insulators, plays a key role in maintaining epigenetic memory. Thus, immune tolerance is likely to involve reshaping of the epigenetic landscape at both proximal and distal regions of pro-inflammatory genes.
LPS tolerance can last for weeks in humans. Tissue-resident macrophages, which appear to persist in the long term, may be the cells that support this memory (Perdiguero and Geissmann 2016). However, whether these cells keep an epigenetic memory along cell divisions is not yet proven. Furthermore, even if imprinted cells divide, why new cells from progenitors in the bone marrow do not restore an efficient immune system is an open question. A tempting hypothesis would be that epigenetic imprinting also occurs at the level of stem cells. This hypothesis needs to be investigated by analyzing the epigenome of stem cells isolated from animal models of sepsis. The reversal of heterochromatin to euchromatin at genes targeted for LPS-mediated repression is also a key issue to understand how “imprinted” immune cells return to homeostasis.
The reduced responsiveness to an effect of a bacterial product caused by prior exposure to this product has also been observed in the case of anthrax Lethal toxin. Some studies indicate that epigenetic modifications contribute to this toxin-induced tolerance (TIR) (Salles et al. 2003). Macrophages exposed to a sublethal dose of anthrax LT become refractory to subsequent cytolytic doses of toxin, and a subset of them retains this phenotype for up to six weeks. The histone deacetylase HDAC8 promotes TIR by changing the chromatin acetylation state of promoter regions of mitochondrial death genes, producing tolerance to a next intoxication (Ha et al. 2014).
6.4.3 Trained Innate Immunity
The priming of innate immune cells by a bacterial challenge can trigger effects opposite to that of tolerance, with cells mounting a faster and longer response than the initial response. This adaptive feature of innate immunity has been defined as “trained immunity” (Netea et al. 2016). A first observation of this phenomenon in humans has been reported upon vaccination with bacilli Calmette-Guérin (BCG). When compared to unvaccinated patients, vaccinated healthy volunteers mounted a more robust response to subsequent infection with unrelated pathogens, and this effect persisted three months after vaccination. The phenomenon was dependent on NOD2 but independent from T- and B-lymphocyte protection (Kleinnijenhuis et al. 2012). The mechanism at play involves BCG-induced epigenetic reprogramming of monocytes through the activating mark H3K4me3 (Kleinnijenhuis et al. 2012). The effect could be downregulated by vitamin A, which increases the levels of the silencing mark H3K9me3 (Arts et al. 2015). Likewise, training of monocytes with a fungal component, β-glucan, induces stable change in histone marks (Quintin et al. 2012) (Fig. 6.5). The list of genes whose expression is induced by this epigenetic reprogramming includes genes involved in glucose metabolism (Cheng et al. 2014). Interestingly, a study supports the notion that the functional programming of monocytes towards either enhanced (training) or decreased (tolerance) cytokine production depends on the nature of the bacterial ligand and of the subsequent activated pattern-recognition receptor (Ifrim et al. 2014).
6.4.4 Polymicrobial Infections and Viral Reactivation
A bacterial infection can also influence viral infections by reactivating latent viruses (Fig. 6.5). This kind of adverse effect is suspected to be associated, for instance, with periodontal pathogens colonizing the oral cavity, such as P. gingivalis. This opportunistic bacterium is proposed to be a risk factor for AIDS or Herpes, by reactivating latent Kaposi’s sarcoma-associated herpesvirus (KSHV) (Morris et al. 2007), human immunodeficiency virus (HIV), and Epstein–Barr virus (EBV) (Imai et al. 2009, 2011). It is proposed that the high production of butyrate by this bacterial species in gingival pockets reactivates viral genes maintained silent by HDAC-containing complexes. One effect of butyrate-mediated EBV reactivation is the increase of the expression of ZEBRA, a lytic gene transactivator (Imai et al. 2012). Butyrate and other SCFAs also downregulate expression of the HDAC SIRT1 and of the HMT EZH2 and SUV39H1, leading to histone hyperacetylation and reduction in the levels of H3K27me3 and H3K9me3, respectively (Ye and Karn 2015). P. gingivalis also elicits changes in the expression of gene encoding chromatin modifiers (as discussed in Sect. 3.4).
6.5 Bacterial Reprogramming of Cell and Tissue Fate
The ultimate effect of bacteria on the epigenome could manifest by a change of identity in the host cell itself. Several recent lines of evidence strongly support the existence of such a drastic change, which would occur when bacterial signals target stem cells or they are potent enough to disrupt the epigenetic barriers that maintain the differentiated cells in their locked state (Fig. 6.5).
6.5.1 Bacteria-Induced Cell Differentiation
The observation that the gut of germ-free mice is altered by defects in the maturation of the intestinal epithelium and of the immune and vascular systems of the gastrointestinal tract is a strong indication that the microbiota contributes to tissue morphogenesis (Sommer and Backhed 2013). The mechanisms are complex and the contribution of epigenetic regulation is not clearly established. Nevertheless, there is evidence that bacteria manipulate the stem cells that generate the different cell types of the intestinal epithelium at the bottom of the crypts. First, intestinal crypts of germ-free mice exhibit a slower turnover of the epithelial cells than conventional mice. Second, stem cells respond to bacterial patterns, such as muramyl-dipeptide (MDP) of the peptidoglycan (Nigro et al. 2014). The commensal microbiota also shapes the intestinal immune system by regulating T helper (TH) cell lineage differentiation (Furusawa et al. 2015). One mechanism involves butyrate secretion by the anaerobic commensal class of bacteria, Clostridia. In particular, butyrate enhances histone acetylation at the promoter of the master regulator of regulatory T cells, FOXP3 (Furusawa et al. 2013).
There is also evidence for pathogen-mediated targeting of cell differentiation pathways in the gut. For instance, the enteroinvasive species Salmonella enterica serovar Typhimurium can convert lymphoid follicle-associated enterocytes into intestinal epithelial microfold (M) cells, though activation of the Wnt/β-catenin signaling pathway mediated by the T3SS effector SopB (Tahoun et al. 2012). This is proposed to promote intestinal invasion by this pathogen.
6.5.2 Bacteria-Induced Cell Dedifferentiation
The study of the behavior of human dermal fibroblasts (HDF) artificially infected with nonpathogenic lactic acid bacteria (LAD) led to an intriguing observation: these LAB-treated HDFs became clustered like embryoid spheres and lost their self-renewal ability (Ohta et al. 2012). In addition, LAB-incorporated cell clusters expressed a subset of pluripotent stem cell marker genes, such as NANOG, OCT3/4, and SOX2, while expression HOX genes, which control the body plan of an embryo, was decreased. Furthermore, LAB-incorporated cell clusters could transform into any of the derivatives of the three germ layers. The mechanism involved in this artificial cell reprogramming by LAD is unclear, but supports the concept that there is a potential for bacterial molecules to revert the host transcriptional program of differentiated cells towards pluripotency.
The study of Mycobacterium leprae (ML), the causative agent of human leprosy, supports this hypothesis. During infection of the peripheral nervous system, this pathogenic bacterium promotes an amazing reprogramming process on adult Schwann cells, by triggering their dedifferentiation into progenitor/stem cell-like cells (Masaki et al. 2013). By using this sophisticated strategy, bacteria disseminate to other niches without being detected by immune cells (Fig. 6.5). ML not only migrates within reprogrammed cells but also spreads the infection to skeletal and smooth muscles by re-differentiating stem cells into these tissues. Moreover, infected stem cells display immunomodulatory properties that promote recruitment of macrophages and formation of granuloma-like structures (Masaki et al. 2014). The mechanism of dedifferentiation of Schwann cells involves activation of differentiation/myelination and lineage-associated genes as well as silencing of numerous developmental genes. ML-induced cellular reprogramming also correlates with changes in DNA methylation supporting a key role of epigenetic regulation in this phenomenon (Masaki et al. 2013).
6.5.3 Bacteria-Induced Epithelial–Mesenchymal Transition and Oncogenesis
The existence of a link between bacteria-mediated aberrant somatic cell reprogramming and cancer is supported by the example of H. pylori, an important acquired risk factor for gastric cancer. Besides H. pylori-mediated effects on cell proliferation, DNA integrity, and DNA methylation (Ushijima and Hattori 2012), infection by this bacterium may also induce dedifferentiation of mature epithelial cells by changing the expression program of the stem cell signaling network. Recent evidence supports a role for H. pylori in inducing the so-called “intestinal metaplasia,” which transforms stomach cells to intestine-like cells via epithelial–mesenchymal transition (EMT) (Fig. 6.5) (Bessede et al. 2014). EMT is known to participate in different carcinogenesis processes and is involved in the generation of cancer stem cells. The bacterial secreted effector CagA promotes the EMT phenotype by activating the expression of master transcription factor genes regulating intestinal differentiation and maintenance.
The “Helicobacter paradigm” may be transposable to bacteria targeting other tissues. It is speculated that E. coli infection may be linked with bladder carcinoma risk (Tolg et al. 2011) and a set of intestinal bacteria might predispose to colon cancer (Sun 2010). More generally, deregulation of tumor-suppressor and/or stem cell-associated pathways (e.g., WNT, JAK-STAT, JNK, and NOTCH) upon genetic alteration and epigenetic reprogramming induced by bacteria is a possible cause of cancer development in epithelial niches.
6.6 Conclusions
The “patho-epigenetics” research field has been defined by Janos Minarovits as the elucidation of the epigenetic consequences of microbe–host interactions leading to pathogenesis (Minarovits 2009). We now propose to add the field of “probio-epigenetics” to cover studies addressing the beneficial effects of bacterial interactions with epigenetic factors. Both themes are developing rapidly as important emerging subtopics at the frontier of Epigenetics and Microbiology sciences. Deciphering the mechanisms underlying the plasticity of gene expression under the action of endogenous (from the microbiota) or exogenous (from food-associated or aerosol-transported bacteria), stimuli may have important impacts on health and disease. Patho-epigenetics may lead to new treatments against bacterial infectious diseases, at a time when pathogenic bacteria became a serious concern due to the emergence of drug resistances. In addition, epigenetic marks may be used as biomarkers to monitor latent infection, disease reactivation, or responses to treatment. Probio-epigenetics can help to characterize the benefits of commensal bacteria on health and to understand the deleterious effects of dysbiosis, which may promote pathological epigenetic signals. Indeed, it is recognized that alterations of microbial communities can cause immune dysregulation, leading to autoimmune disorders, and may contribute to metabolic diseases, neuropathies, and behavioral problems. Restoring the composition of altered intestinal microbiota with fecal flora transfer is now tested as a way to counteract these adverse effects. The interplay of bacteria and nutrients is also an important emerging field of research, due to the key role of metabolites on epigenetic regulation. Western food may change epigenetic patterns in the gut, favoring pathogens associated with intestinal inflammatory diseases. There is evidence that changes in nutritional habits, such as low intake in methyl donor molecules, promote abnormal epigenetic marks in a mouse model mimicking susceptibility to E. coli-mediated gut inflammation and Crohn’s disease (Denizot et al. 2015).
At the level of basic science, these investigations highlight the amazing molecular tools used by intracellular pathogens to manipulate chromatin and to fine-tune host gene expression. The number of nucleomodulins discovered in the bacterial world is rising steadily, in particular thanks to bioinformatic analysis. For instance, five of over 80 T4SS substrates of Coxiella burnetii, the agent of Q fever, are predicted to carry a nuclear localization signal. One of them, Cbu1314, has recently been confirmed as a protein binding to chromatin and controlling transcription of host cell genes (Weber et al. 2016). In the toolbox of bacterial molecules, bacterial metabolic by-products, such as SCFAs, are also prone to induce epimutations. Several of such molecules may emerge as novel epigenetic drugs. It is also striking that the study of bacterial factors promotes innovation in the epigenetics field. For instance, studies carried out in L. monocytogenes have led to the discovery of the BAHD1 chromatin-repressive complex (Bierne et al. 2009; Lebreton et al. 2011; Libertini et al. 2015; Lakisic et al. 2016) and of a nuclear function for the histone deacetylase SIRT2 (Eskandarian et al. 2013).
How bacterial modulation of the epigenetic information is spatio-temporally coordinated, at specific genome loci, according to time, cell type, and stimuli, is not fully understood. In addition, the impact of bacterial signals on the epigenetic profile of structural elements, such as enhancers, insulators, and DNA repeats, has been poorly addressed. Moreover, how expression and secretion of bacterial factors is controlled in the host and how chromatin writers and erasers become engaged in response to a cocktail of bacterial stimuli are important questions to be solved. In this regard, deciphering how chromatin modifications are spread throughout the genome (the “patho-epigenome” or the “probio-epigenome”) is likely to provide important clues. Technological advances in human genome-wide mapping of DNA methylation and histone modifications, as well as in systems biology, will help to make significant progress. Investigations will have to be performed at both the tissue and single cell levels, with the objective of analyzing precursors of cell lineages or differentiated cells with long life spans, in order to determine the “chromatin signature” of bacterial cues. As epigenetic processes can be reverted, elimination of patho-epigenetic changes induced by microbes may prevent chronic or latent infections, as well as some cancers and autoimmune diseases. This opens avenues for future research.
References
Agarwal S, Agarwal S, Jin H, Pancholi P, Pancholi V (2012) Serine/threonine phosphatase (SP-STP), secreted from Streptococcus pyogenes, is a pro-apoptotic protein. J Biol Chem 287(12):9147–9167. doi:10.1074/jbc.M111.316554
Aguilera C, Nakagawa K, Sancho R, Chakraborty A, Hendrich B, Behrens A (2011) c-Jun N-terminal phosphorylation antagonises recruitment of the Mbd3/NuRD repressor complex. Nature 469 (7329):231–235. doi:10.1038/nature09607
Alenghat T, Artis D (2014) Epigenomic regulation of host-microbiota interactions. Trends in immunology 35(11):518–525. doi:10.1016/j.it.2014.09.007
Alenghat T, Osborne LC, Saenz SA, Kobuley D, Ziegler CG, Mullican SE, Choi I, Grunberg S, Sinha R, Wynosky-Dolfi M, Snyder A, Giacomin PR, Joyce KL, Hoang TB, Bewtra M, Brodsky IE, Sonnenberg GF, Bushman FD, Won KJ, Lazar MA, Artis D (2013) Histone deacetylase 3 coordinates commensal-bacteria-dependent intestinal homeostasis. Nature 504(7478):153–157. doi:10.1038/nature12687
Allemand E, Batsche E, Muchardt C (2008) Splicing, transcription, and chromatin: a menage a trois. Curr Opin Genet Dev 18(2):145–151. doi:10.1016/j.gde.2008.01.006
Alvarez-Errico D, Vento-Tormo R, Sieweke M, Ballestar E (2015) Epigenetic control of myeloid cell differentiation, identity and function. Nat Rev Immunol 15(1):7–17. doi:10.1038/nri3777
Ando T, Yoshida T, Enomoto S, Asada K, Tatematsu M, Ichinose M, Sugiyama T, Ushijima T (2009) DNA methylation of microRNA genes in gastric mucosae of gastric cancer patients: its possible involvement in the formation of epigenetic field defect. Int J Cancer 124(10):2367–2374. doi:10.1002/ijc.24219
Arbibe L, Kim DW, Batsche E, Pedron T, Mateescu B, Muchardt C, Parsot C, Sansonetti PJ (2007) An injected bacterial effector targets chromatin access for transcription factor NF-kappaB to alter transcription of host genes involved in immune responses. Nat Immunol 8(1):47–56. doi:10.1038/ni1423
Arpaia N, Campbell C, Fan X, Dikiy S, van der Veeken J, deRoos P, Liu H, Cross JR, Pfeffer K, Coffer PJ, Rudensky AY (2013) Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504(7480):451–455. doi:10.1038/nature12726
Arts RJ, Blok BA, van Crevel R, Joosten LA, Aaby P, Benn CS, Netea MG (2015) Vitamin A induces inhibitory histone methylation modifications and down-regulates trained immunity in human monocytes. J Leukoc Biol 98(1):129–136. doi:10.1189/jlb.6AB0914-416R
Arzate-Mejia RG, Valle-Garcia D, Recillas-Targa F (2011) Signaling epigenetics: novel insights on cell signaling and epigenetic regulation. IUBMB Life 63(10):881–895. doi:10.1002/iub.557
Baek SH (2011) When signaling kinases meet histones and histone modifiers in the nucleus. Mol Cell 42(3):274–284. doi:10.1016/j.molcel.2011.03.022
Bandyopadhaya A, Tsurumi A, Maura D, Jeffrey KL, Rahme LG (2016) A quorum-sensing signal promotes host tolerance training through HDAC1-mediated epigenetic reprogramming. Nat Microbiol 1:16174. doi:10.1371/journal.ppat.1003024
Bardwell AJ, Abdollahi M, Bardwell L (2004) Anthrax lethal factor-cleavage products of MAPK (mitogen-activated protein kinase) kinases exhibit reduced binding to their cognate MAPKs. Biochem J 378(Pt 2):569–577. doi:10.1042/BJ20031382
Bayarsaihan D (2011) Epigenetic mechanisms in inflammation. J Dent Res 90(1):9–17. doi:10.1177/0022034510378683
Benabdillah R, Ls JM, Lützelschwab S, Demoinet E, Cornelis GR (2004) Identification of a nuclear targeting signal in YopM from Yersinia spp. Microbial pathogenesis 36(5):247–261. doi:10.1016/j.micpath.2003.12.006
Berneking L, Schnapp M, Rumm A, Trasak C, Ruckdeschel K, Alawi M, Grundhoff A, Kikhney AG, Koch-Nolte F, Buck F, Perbandt M, Betzel C, Svergun DI, Hentschke M, Aepfelbacher M (2016) Immunosuppressive Yersinia effector YopM binds DEAD box helicase DDX3 to control ribosomal S6 kinase in the nucleus of host cells. PLoS Pathog 12(6):e1005660. doi:10.1371/journal.ppat.1005660
Bessede E, Staedel C, Acuna Amador LA, Nguyen PH, Chambonnier L, Hatakeyama M, Belleannee G, Megraud F, Varon C (2014) Helicobacter pylori generates cells with cancer stem cell properties via epithelial-mesenchymal transition-like changes. Oncogene 33(32):4123–4131. doi:10.1038/onc.2013.380
Bhavsar AP, Guttman JA, Finlay BB (2007) Manipulation of host-cell pathways by bacterial pathogens. Nature 449(7164):827–834. doi:10.1038/nature06247
Bhutani N, Burns DM, Blau HM (2011) DNA demethylation dynamics. Cell 146(6):866–872. doi:10.1016/j.cell.2011.08.042
Bierne H (2013) Nuclear microbiology—bacterial assault on the nucleolus. EMBO Rep 14(8):663–664. doi:10.1038/embor.2013.105
Bierne H, Cossart P (2012) When bacteria target the nucleus: the emerging family of nucleomodulins. Cell Microbiol. doi:10.1111/j.1462-5822.2012.01758.x
Bierne H, Hamon M, Cossart P (2012) Epigenetics and bacterial infections. Cold Spring Harb Perspect Med 2(12):a010272. doi:10.1101/cshperspect.a010272
Bierne H, Tham TN, Batsche E, Dumay A, Leguillou M, Kerneis-Golsteyn S, Regnault B, Seeler JS, Muchardt C, Feunteun J, Cossart P (2009) Human BAHD1 promotes heterochromatic gene silencing. Proc Natl Acad Sci U S A 106(33):13826–13831. doi:10.1073/pnas.0901259106
Bird A (2007) Perceptions of epigenetics. Nature 447(7143):396–398. doi:10.1038/nature05913
Bobetsis YA, Barros SP, Lin DM, Weidman JR, Dolinoy DC, Jirtle RL, Boggess KA, Beck JD, Offenbacher S (2007) Bacterial infection promotes DNA hypermethylation. J Dent Res 86(2):169–174. doi:10.1177/154405910708600212
Borjesson DL, Kobayashi SD, Whitney AR, Voyich JM, Argue CM, Deleo FR (2005) Insights into pathogen immune evasion mechanisms: Anaplasma phagocytophilum fails to induce an apoptosis differentiation program in human neutrophils. J Immunol 174(10):6364–6372
Brennan DF, Barford D (2009) Eliminylation: a post-translational modification catalyzed by phosphothreonine lyases. Trends Biochem Sci 34(3):108–114. doi:10.1016/j.tibs.2008.11.005
Brestoff JR, Artis D (2013) Commensal bacteria at the interface of host metabolism and the immune system. Nat Immunol 14(7):676–684. doi:10.1038/ni.2640
Buck-Koehntop BA, Defossez PA (2013) On how mammalian transcription factors recognize methylated DNA. Epigenetics 8(2):131–137. doi:10.4161/epi.23632
Bussiere FI, Michel V, Memet S, Ave P, Vivas JR, Huerre M, Touati E (2010) H. pylori-induced promoter hypermethylation downregulates USF1 and USF2 transcription factor gene expression. Cell Microbiol 12(8):1124–1133. doi:10.1111/j.1462-5822.2010.01457.x
Cai Y, Geutjes EJ, de Lint K, Roepman P, Bruurs L, Yu LR, Wang W, van Blijswijk J, Mohammad H, de Rink I, Bernards R, Baylin SB (2014) The NuRD complex cooperates with DNMTs to maintain silencing of key colorectal tumor suppressor genes. Oncogene 33(17):2157–2168. doi:10.1038/Onc.2013.178
Canonne J, Rivas S (2012) Bacterial effectors target the plant cell nucleus to subvert host transcription. Plant Signal Behav 7(2):217–221. doi:10.4161/psb.18885
Cao J (2014) The functional role of long non-coding RNAs and epigenetics. Biol Proced Online 16:11. doi:10.1186/1480-9222-16-11
Carson WF, Cavassani KA, Dou Y, Kunkel SL (2011) Epigenetic regulation of immune cell functions during post-septic immunosuppression. Epigenetics 6(3):273–283
Cedar H, Bergman Y (2009) Linking DNA methylation and histone modification: patterns and paradigms. Nature Rev Genetics 10(5):295–304. doi:10.1038/nrg2540
Chan AO, Lam SK, Wong BC, Wong WM, Yuen MF, Yeung YH, Hui WM, Rashid A, Kwong YL (2003) Promoter methylation of E-cadherin gene in gastric mucosa associated with Helicobacter pylori infection and in gastric cancer. Gut 52(4):502–506
Chandran A, Antony C, Jose L, Mundayoor S, Natarajan K, Kumar RA (2015) Mycobacterium tuberculosis infection induces HDAC1-mediated suppression of IL-12B gene expression in macrophages. Front Cell Infect Microbiol 5:90. doi:10.3389/fcimb.2015.00090
Chang PV, Hao L, Offermanns S, Medzhitov R (2014) The microbial metabolite butyrate regulates intestinal macrophage function via histone deacetylase inhibition. Proc Natl Acad Sci U S A 111(6):2247–2252. doi:10.1073/pnas.1322269111
Chen ZX, Riggs AD (2011) DNA methylation and demethylation in mammals. J Biol Chem 286(21):18347–18353. doi:10.1074/jbc.R110.205286
Chen X, Yoza BK, El Gazzar M, Hu JY, Cousart SL, McCall CE (2009) RelB sustains IkappaBalpha expression during endotoxin tolerance. Clin Vaccine Immunol 16(1):104–110. doi:10.1128/CVI.00320-08
Cheng SC, Quintin J, Cramer RA, Shepardson KM, Saeed S, Kumar V, Giamarellos-Bourboulis EJ, Martens JH, Rao NA, Aghajanirefah A, Manjeri GR, Li Y, Ifrim DC, Arts RJ, van der Veer BM, Deen PM, Logie C, O’Neill LA, Willems P, van de Veerdonk FL, van der Meer JW, Ng A, Joosten LA, Wijmenga C, Stunnenberg HG, Xavier RJ, Netea MG (2014) mTOR- and HIF-1alpha-mediated aerobic glycolysis as metabolic basis for trained immunity. Science 345(6204):1250684. doi:10.1126/science.1250684
Chernov AV, Reyes L, Xu Z, Gonzalez B, Golovko G, Peterson S, Perucho M, Fofanov Y, Strongin AY (2015) Mycoplasma CG- and GATC-specific DNA methyltransferases selectively and efficiently methylate the host genome and alter the epigenetic landscape in human cells. Epigenetics 10(4):303–318. doi:10.1080/15592294.2015.1020000
Cuddapah S, Barski A, Zhao K (2010) Epigenomics of T cell activation, differentiation, and memory. Curr Opin Immunol 22(3):341–347. doi:10.1016/j.coi.2010.02.007
de Camargo Pereira G, Guimaraes GN, Planello AC, Santamaria MP, de Souza AP, Line SR, Marques MR (2013) Porphyromonas gingivalis LPS stimulation downregulates DNMT1, DNMT3a, and JMJD3 gene expression levels in human HaCaT keratinocytes. Clin Oral Investig 17(4):1279–1285. doi:10.1007/s00784-012-0816-z
Denizot J, Desrichard A, Agus A, Uhrhammer N, Dreux N, Vouret-Craviari V, Hofman P, Darfeuille-Michaud A, Barnich N (2015) Diet-induced hypoxia responsive element demethylation increases CEACAM6 expression, favouring Crohn’s disease-associated Escherichia coli colonisation. Gut 64(3):428–437. doi:10.1136/gutjnl-2014-306944
Diacovich L, Gorvel JP (2010) Bacterial manipulation of innate immunity to promote infection. Nat Rev Microbiol 8(2):117–128. doi:10.1038/nrmicro2295
Ding SZ, Fischer W, Kaparakis-Liaskos M, Liechti G, Merrell DS, Grant PA, Ferrero RL, Crowe SE, Haas R, Hatakeyama M, Goldberg JB (2010a) Helicobacter pylori-induced histone modification, associated gene expression in gastric epithelial cells, and its implication in pathogenesis. PLoS One 5(4):e9875. doi:10.1371/journal.pone.0009875
Ding SZ, Goldberg JB, Hatakeyama M (2010b) Helicobacter pylori infection, oncogenic pathways and epigenetic mechanisms in gastric carcinogenesis. Future Oncol 6(5):851–862. doi:10.2217/fon.10.37
Dominissini D, Moshitch-Moshkovitz S, Schwartz S, Salmon-Divon M, Ungar L, Osenberg S, Cesarkas K, Jacob-Hirsch J, Amariglio N, Kupiec M, Sorek R, Rechavi G (2012) Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485(7397):201–206. doi:10.1038/nature11112
Dominissini D, Nachtergaele S, Moshitch-Moshkovitz S, Peer E, Kol N, Ben-Haim MS, Dai Q, Di Segni A, Salmon-Divon M, Clark WC, Zheng GQ, Pan T, Solomon O, Eyal E, Hershkovitz V, Han D, Dore LC, Amariglio N, Rechavi G, He C (2016) The dynamic N-1-methyladenosine methylome in eukaryotic messenger RNA. Nature 530(7591):441–446. doi:10.1038/nature16998
Du J, Johnson LM, Jacobsen SE, Patel DJ (2015) DNA methylation pathways and their crosstalk with histone methylation. Nat Rev Mol Cell Biol 16(9):519–532. doi:10.1038/nrm4043
Dumler JS, Sinclair SH, Pappas-Brown V, Shetty AC (2016) Genome-wide Anaplasma phagocytophilum AnkA-DNA interactions are enriched in intergenic regions and gene promoters and correlate with infection-induced differential gene expression. Front Cell Infect Microbiol 20(6):97. doi:10.3389/fcimb.2016.00097
Dunphy PS, Luo T, McBride JW (2014) Ehrlichia chaffeensis exploits host SUMOylation pathways to mediate effector-host interactions and promote intracellular survival. Infect Immun 82(10):4154–4168. doi:10.1128/IAI.01984-14
Dussurget O, Bierne H, Cossart P (2014) The bacterial pathogen Listeria monocytogenes and the interferon family: type I, type II and type III interferons. Front Cell Infect Microbiol 4:50. doi:10.3389/fcimb.2014.00050
El Gazzar M, Yoza BK, Chen X, Hu J, Hawkins GA, McCall CE (2008) G9a and HP1 couple histone and DNA methylation to TNFalpha transcription silencing during endotoxin tolerance. J Biol Chem 283(47):32198–32208. doi:10.1074/jbc.M803446200
Eskandarian HA, Impens F, Nahori MA, Soubigou G, Coppee JY, Cossart P, Hamon MA (2013) A role for SIRT2-dependent histone H3K18 deacetylation in bacterial infection. Science 341(6145):1238858. doi:10.1126/science.1238858
Faghihi MA, Wahlestedt C (2009) Regulatory roles of natural antisense transcripts. Nat Rev Mol Cell Biol 10(9):637–643. doi:10.1038/nrm2738
Farris TR, Dunphy PS, Zhu B, Kibler CE, McBride JW (2016) Ehrlichia chaffeensis TRP32 is a nucleomodulin that directly regulates expression of host genes governing differentiation and proliferation. Infect Immun 84:3182–3194. doi:10.1128/IAI.00657-16
Fehri LF, Rechner C, Janssen S, Mak TN, Holland C, Bartfeld S, Bruggemann H, Meyer TF (2009) Helicobacter pylori-induced modification of the histone H3 phosphorylation status in gastric epithelial cells reflects its impact on cell cycle regulation. Epigenetics 4(8):577–586
Fujita N, Watanabe S, Ichimura T, Tsuruzoe S, Shinkai Y, Tachibana M, Chiba T, Nakao M (2003) Methyl-CpG binding domain 1 (MBD1) interacts with the Suv39h1-HP1 heterochromatic complex for DNA methylation-based transcriptional repression. J Biol Chem 278(26):24132–24138. doi:10.1074/jbc.M302283200
Furusawa Y, Obata Y, Fukuda S, Endo TA, Nakato G, Takahashi D, Nakanishi Y, Uetake C, Kato K, Kato T, Takahashi M, Fukuda NN, Murakami S, Miyauchi E, Hino S, Atarashi K, Onawa S, Fujimura Y, Lockett T, Clarke JM, Topping DL, Tomita M, Hori S, Ohara O, Morita T, Koseki H, Kikuchi J, Honda K, Hase K, Ohno H (2013) Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504(7480):446–450. doi:10.1038/nature12721
Furusawa Y, Obata Y, Hase K (2015) Commensal microbiota regulates T cell fate decision in the gut. Semin Immunopathol 37(1):17–25. doi:10.1007/s00281-014-0455-3
Garcia-Dominguez M, Reyes JC (2009) SUMO association with repressor complexes, emerging routes for transcriptional control. Biochim Biophys Acta 1789(6–8):451–459. doi:10.1016/j.bbagrm.2009.07.001
Garcia-Garcia JC, Barat NC, Trembley SJ, Dumler JS (2009a) Epigenetic silencing of host cell defense genes enhances intracellular survival of the rickettsial pathogen Anaplasma phagocytophilum. PLoS Pathog 5(6):e1000488. doi:10.1371/journal.ppat.1000488
Garcia-Garcia JC, Rennoll-Bankert KE, Pelly S, Milstone AM, Dumler JS (2009b) Silencing of host cell CYBB gene expression by the nuclear effector AnkA of the intracellular pathogen Anaplasma phagocytophilum. Infect Immun 77(6):2385–2391. doi:10.1128/IAI.00023-09
Ghadimi D, Helwig U, Schrezenmeir J, Heller KJ, de Vrese M (2012) Epigenetic imprinting by commensal probiotics inhibits the IL-23/IL-17 axis in an in vitro model of the intestinal mucosal immune system. J Leukoc Biol 92(4):895–911. doi:10.1189/jlb.0611286
Guelen L, Pagie L, Brasset E, Meuleman W, Faza MB, Talhout W, Eussen BH, de Klein A, Wessels L, de Laat W, van Steensel B (2008) Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453(7197):948–951. doi:10.1038/nature06947
Ha SD, Han CY, Reid C, Kim SO (2014) HDAC8-mediated epigenetic reprogramming plays a key role in resistance to anthrax lethal toxin-induced pyroptosis in macrophages. J Immunol 193(3):1333–1343. doi:10.4049/jimmunol.1400420
Ha SD, Reid C, Meshkibaf S, Kim SO (2016) Inhibition of interleukin 1beta (IL-1beta) expression by anthrax Lethal Toxin (LeTx) is reversed by Histone Deacetylase 8 (HDAC8) inhibition in murine macrophages. J Biol Chem 291(16):8745–8755. doi:10.1074/jbc.M115.695809
Haller D, Holt L, Kim SC, Schwabe RF, Sartor RB, Jobin C (2003) Transforming growth factor-beta 1 inhibits non-pathogenic Gram negative bacteria-induced NF-kappa B recruitment to the interleukin-6 gene promoter in intestinal epithelial cells through modulation of histone acetylation. J Biol Chem 278(26):23851–23860. doi:10.1074/jbc.M300075200
Hamon MA, Batsche E, Regnault B, Tham TN, Seveau S, Muchardt C, Cossart P (2007) Histone modifications induced by a family of bacterial toxins. Proc Natl Acad Sci U S A 104 (33):13467–13472. doi:10.1073/pnas.0702729104
Hamon M, Bierne H, Cossart P (2006) Listeria monocytogenes: a multifaceted model. Nat Rev Microbiol 4(6):423–434. doi:10.1038/nrmicro1413
Hamon MA, Cossart P (2011) K+ efflux is required for histone H3 dephosphorylation by Listeria monocytogenes listeriolysin O and other pore-forming toxins. Infect Immun 79(7):2839–2846. doi:10.1128/IAI.01243-10
Haraga A, Miller SI (2003) A Salmonella enterica serovar typhimurium translocated leucine-rich repeat effector protein inhibits NF-kappa B-dependent gene expression. Infect Immun 71(7):4052–4058
Harouz H, Rachez C, Meijer BM, Marteyn B, Donnadieu F, Cammas F, Muchardt C, Sansonetti P, Arbibe L (2014) Shigella flexneri targets the HP1gamma subcode through the phosphothreonine lyase OspF. EMBO J 33 (22):2606–2622. doi:10.15252/embj.201489244
Hattori N, Ushijima T (2016) Epigenetic impact of infection on carcinogenesis: mechanisms and applications. Genome Med 8(1):10. doi:10.1186/s13073-016-0267-2
Hicks SW, Galan JE (2010) Hijacking the host ubiquitin pathway: structural strategies of bacterial E3 ubiquitin ligases. Curr Opin Microbiol 13(1):41–46. doi:10.1016/j.mib.2009.11.008
Hiragami-Hamada K, Shinmyozu K, Hamada D, Tatsu Y, Uegaki K, Fujiwara S, Nakayama J (2011) N-terminal phosphorylation of HP1{alpha} promotes its chromatin binding. Mol Cell Biol 31(6):1186–1200. doi:10.1128/MCB.01012-10
Hnilicova J, Stanek D (2011) Where splicing joins chromatin. Nucleus 2(3):182–188. doi:10.4161/nucl.2.3.15876
Holeski LM, Jander G, Agrawal AA (2012) Transgenerational defense induction and epigenetic inheritance in plants. Trends Ecol Evol 27(11):618–626. doi:10.1016/j.tree.2012.07.011
Hur K, Niwa T, Toyoda T, Tsukamoto T, Tatematsu M, Yang HK, Ushijima T (2011) Insufficient role of cell proliferation in aberrant DNA methylation induction and involvement of specific types of inflammation. Carcinogenesis 32(1):35–41. doi:10.1093/carcin/bgq219
Ichimura T, Watanabe S, Sakamoto Y, Aoto T, Fujita N, Nakao M (2005) Transcriptional repression and heterochromatin formation by MBD1 and MCAF/AM family proteins. J Biol Chem 280(14):13928–13935. doi:10.1074/jbc.M413654200
Ifrim DC, Quintin J, Joosten LA, Jacobs C, Jansen T, Jacobs L, Gow NA, Williams DL, van der Meer JW, Netea MG (2014) Trained immunity or tolerance: opposing functional programs induced in human monocytes after engagement of various pattern recognition receptors. Clin Vaccine Immunol 21(4):534–545. doi:10.1128/CVI.00688-13
Imai K, Inoue H, Tamura M, Cueno ME, Inoue H, Takeichi O, Kusama K, Saito I, Ochiai K (2012) The periodontal pathogen Porphyromonas gingivalis induces the Epstein-Barr virus lytic switch transactivator ZEBRA by histone modification. Biochimie 94(3):839–846. doi:10.1016/j.biochi.2011.12.001
Imai K, Inoue H, Tamura M, Cueno ME, Takeichi O, Kusama K, Saito I, Ochiai K (2011) The periodontal pathogen Porphyromonas gingivalis induces the Epstein-Barr virus lytic switch transactivator ZEBRA by histone modification. Biochimie. doi:10.1016/j.biochi.2011.12.001
Imai K, Ochiai K, Okamoto T (2009) Reactivation of latent HIV-1 infection by the periodontopathic bacterium Porphyromonas gingivalis involves histone modification. J Immunol 182(6):3688–3695. doi:10.4049/jimmunol.0802906
Jenner RG, Young RA (2005) Insights into host responses against pathogens from transcriptional profiling. Nat Rev Microbiol 3(4):281–294. doi:10.1038/nrmicro1126
Jimenez A, Chen D, Alto NM (2016) How bacteria subvert animal cell structure and function. Annu Rev Cell Dev Biol. doi:10.1146/annurev-cellbio-100814-125227
Jose L, Ramachandran R, Bhagavat R, Gomez RL, Chandran A, Raghunandanan S, Omkumar RV, Chandra N, Mundayoor S, Kumar RA (2016) Hypothetical protein Rv3423.1 of Mycobacterium tuberculosis is a histone acetyltransferase. FEBS J 283(2):265–281. doi:10.1111/febs.13566
Kaikkonen MU, Lam MT, Glass CK (2011) Non-coding RNAs as regulators of gene expression and epigenetics. Cardiovasc Res 90(3):430–440. doi:10.1093/cvr/cvr097
Kleinnijenhuis J, Quintin J, Preijers F, Joosten LA, Ifrim DC, Saeed S, Jacobs C, van Loenhout J, de Jong D, Stunnenberg HG, Xavier RJ, van der Meer JW, van Crevel R, Netea MG (2012) Bacille Calmette-Guerin induces NOD2-dependent nonspecific protection from reinfection via epigenetic reprogramming of monocytes. Proc Natl Acad Sci U S A 109(43):17537–17542. doi:10.1073/pnas.1202870109
Klose RJ, Bird AP (2006) Genomic DNA methylation: the mark and its mediators. Trends Biochem Sci 31 (2):89–97. doi:1016/j.tibs.2005.12.008
Kohwi-Shigematsu T, Kohwi Y, Takahashi K, Richards HW, Ayers SD, Han HJ, Cai S (2012) SATB1-mediated functional packaging of chromatin into loops. Methods 58(3):243–254. doi:10.1016/j.ymeth.2012.06.019
Kouzarides T (2007) Chromatin modifications and their function. Cell 128(4):693–705. doi:10.1016/j.cell.2007.02.005
Lai AY, Wade PA (2011) Cancer biology and NuRD: a multifaceted chromatin remodelling complex. Nat Rev Cancer 11(8):588–596. doi:10.1038/nrc3091
Lakisic G, Lebreton A, Pourpre R, Wendling O, Libertini E, Radford EJ, Le Guillou M, Champy MF, Wattenhofer-Donze M, Soubigou G, Ait-Si-Ali S, Feunteun J, Sorg T, Coppee JY, Ferguson-Smith AC, Cossart P, Bierne H (2016) Role of the BAHD1 chromatin-repressive complex in placental development and regulation of steroid metabolism. PLoS Genet 12(3):e1005898. doi:10.1371/journal.pgen.1005898
Langst G, Manelyte L (2015) Chromatin remodelers: from function to dysfunction. Genes (Basel) 6(2):299–324. doi:10.3390/genes6020299
Latham JA, Dent SY (2007) Cross-regulation of histone modifications. Nat Struct Mol Biol 14(11):1017–1024. doi:10.1038/nsmb1307
Latham T, Mackay L, Sproul D, Karim M, Culley J, Harrison DJ, Hayward L, Langridge-Smith P, Gilbert N, Ramsahoye BH (2012) Lactate, a product of glycolytic metabolism, inhibits histone deacetylase activity and promotes changes in gene expression. Nucleic Acids Res 40(11):4794–4803. doi:10.1093/nar/gks066
Lebreton A, Cossart P, Bierne H (2012) Bacteria tune interferon responses by playing with chromatin. Virulence 3(1):87–91
Lebreton A, Job V, Ragon M, Le Monnier A, Dessen A, Cossart P, Bierne H (2014) Structural basis for the inhibition of the chromatin repressor BAHD1 by the bacterial nucleomodulin LntA. MBio 5(1):e00775–e00713. doi:10.1128/mBio.00775-13
Lebreton A, Lakisic G, Job V, Fritsch L, Tham TN, Camejo A, Mattei P-J, Regnault B, Nahori M-A, Cabanes D, Gautreau A, Ait-Si-Ali S, Dessen A, Cossart P, Bierne H (2011) A bacterial protein targets the BAHD1 chromatin complex to stimulate type III interferon response. Science 331(6022):1319–1321
Lee HC, Kioi M, Han J, Puri RK, Goodman JL (2008) Anaplasma phagocytophilum-induced gene expression in both human neutrophils and HL-60 cells. Genomics 92(3):144–151. doi:10.1016/j.ygeno.2008.05.005
Lee JS, Smith E, Shilatifard A (2010) The language of histone crosstalk. Cell 142(5):682–685. doi:10.1016/j.cell.2010.08.011
Li T, Lu Q, Wang G, Xu H, Huang H, Cai T, Kan B, Ge J, Shao F (2013) SET-domain bacterial effectors target heterochromatin protein 1 to activate host rDNA transcription. EMBO Rep 14(8):733–740. doi:10.1038/embor.2013.86
Li H, Rauch T, Chen ZX, Szabo PE, Riggs AD, Pfeifer GP (2006) The histone methyltransferase SETDB1 and the DNA methyltransferase DNMT3A interact directly and localize to promoters silenced in cancer cells. J Biol Chem 281(28):19489–19500. doi:10.1074/jbc.M513249200
Li H, Xu H, Zhou Y, Zhang J, Long C, Li S, Chen S, Zhou J-M, Shao F (2007) The phosphothreonine lyase activity of a bacterial type III effector family. Science 315(5814):1000–1003. doi:10.1126/science.1138960
Libertini E, Lebreton A, Lakisic G, Dillies MA, Beck S, Coppee JY, Cossart P, Bierne H (2015) Overexpression of the heterochromatinization factor BAHD1 in HEK293 cells differentially reshapes the DNA methylome on autosomes and X chromosome. Front Genet 6:339. doi:10.3389/fgene.2015.00339
Licciardi PV, Wong SS, Tang ML, Karagiannis TC (2010) Epigenome targeting by probiotic metabolites. Gut Pathog 2(1):24. doi:10.1186/1757-4749-2-24
Liu Y, Zhang X, Blumenthal RM, Cheng X (2013) A common mode of recognition for methylated CpG. Trends Biochem Sci 38(4):177–183. doi:10.1016/j.tibs.2012.12.005
Luo T, Kuriakose JA, Zhu B, Wakeel A, McBride JW (2011) Ehrlichia chaffeensis TRP120 interacts with a diverse array of eukaryotic proteins involved in transcription, signaling, and cytoskeleton organization. Infect Immun 79(11):4382–4391. doi:10.1128/IAI.05608-11
Luo T, McBride JW (2012) Ehrlichia chaffeensis TRP32 interacts with host cell targets that influence intracellular survival. Infect Immun 80(7):2297–2306. doi:10.1128/IAI.00154-12
Ma KW, Flores C, Ma W (2011) Chromatin configuration as a battlefield in plant-bacteria interactions. Plant Physiol. doi:10.1104/pp.111.182295
Maekita T, Nakazawa K, Mihara M, Nakajima T, Yanaoka K, Iguchi M, Arii K, Kaneda A, Tsukamoto T, Tatematsu M, Tamura G, Saito D, Sugimura T, Ichinose M, Ushijima T (2006) High levels of aberrant DNA methylation in Helicobacter pylori-infected gastric mucosae and its possible association with gastric cancer risk. Clin Cancer Res 12(3 Pt 1):989–995. doi:10.1158/1078-0432.CCR-05-2096
Margueron R, Reinberg D (2011) The Polycomb complex PRC2 and its mark in life. Nature 469(7330):343–349. doi:10.1038/nature09784
Masaki T, McGlinchey A, Cholewa-Waclaw J, Qu J, Tomlinson SR, Rambukkana A (2014) Innate immune response precedes Mycobacterium leprae-induced reprogramming of adult Schwann cells. Cell Reprogram 16(1):9–17. doi:10.1089/cell.2013.0064
Masaki T, Qu J, Cholewa-Waclaw J, Burr K, Raaum R, Rambukkana A (2013) Reprogramming adult Schwann cells to stem cell-like cells by leprosy bacilli promotes dissemination of infection. Cell 152(1–2):51–67. doi:10.1016/j.cell.2012.12.014
Mattick JS, Makunin IV (2005) Small regulatory RNAs in mammals. Hum Mol Genet 14(1):R121–R132. doi:10.1093/hmg/ddi101
Mattout A, Cabianca DS, Gasser SM (2015) Chromatin states and nuclear organization in development—a view from the nuclear lamina. Genome Biol 16:174. doi:10.1186/s13059-015-0747-5
McCall CE, Yoza B, Liu T, El Gazzar M (2010) Gene-specific epigenetic regulation in serious infections with systemic inflammation. J Innate Immun 2(5):395–405. doi:10.1159/000314077
McCoy MW, Marre ML, Lesser CF, Mecsas J (2010) The C-terminal tail of Yersinia pseudotuberculosis YopM is critical for interacting with RSK1 and for virulence. Infect Immun 78(6):2584–2598. doi:10.1128/IAI.00141-10
McDonald C, Vacratsis PO, Bliska JB, Dixon JE (2003) The yersinia virulence factor YopM forms a novel protein complex with two cellular kinases. J Biol Chem 278(20):18514–18523. doi:10.1074/jbc.M301226200
Medvedeva YA, Lennartsson A, Ehsani R, Kulakovskiy IV, Vorontsov IE, Panahandeh P, Khimulya G, Kasukawa T, Consortium F, Drablos F (2015) EpiFactors: a comprehensive database of human epigenetic factors and complexes. Database (Oxford) 2015:bav067. doi:10.1093/database/bav067
Medzhitov R, Horng T (2009) Transcriptional control of the inflammatory response. Nature Rev Immunol 9(10):692–703. doi:10.1038/nri2634
Meyer KD, Jaffrey SR (2014) The dynamic epitranscriptome: N6-methyladenosine and gene expression control. Nat Rev Mol Cell Biol 15(5):313–326. doi:10.1038/nrm3785
Mikkelsen TS, Ku M, Jaffe DB, Issac B, Lieberman E, Giannoukos G, Alvarez P, Brockman W, Kim TK, Koche RP, Lee W, Mendenhall E, O'Donovan A, Presser A, Russ C, Xie X, Meissner A, Wernig M, Jaenisch R, Nusbaum C, Lander ES, Bernstein BE (2007) Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448(7153):553–560. doi:10.1038/nature06008
Minarovits J (2009) Microbe-induced epigenetic alterations in host cells: the coming era of patho-epigenetics of microbial infections. A review. Acta Microbiol Immunol Hung 56(1):1–19. doi:10.1556/AMicr.56.2009.1.1
Mohammad HP, Baylin SB (2010) Linking cell signaling and the epigenetic machinery. Nat Biotechnol 28(10):1033–1038. doi:10.1038/nbt1010-1033
Mojica SA, Hovis KM, Frieman MB, Tran B, Hsia RC, Ravel J, Jenkins-Houk C, Wilson KL, Bavoil PM (2015) SINC, a type III secreted protein of Chlamydia psittaci, targets the inner nuclear membrane of infected cells and uninfected neighbors. Mol Biol Cell 26(10):1918–1934. doi:10.1091/mbc.E14-11-1530
Morris TL, Arnold RR, Webster-Cyriaque J (2007) Signaling cascades triggered by bacterial metabolic end products during reactivation of Kaposi's sarcoma-associated herpesvirus. J Virol 81(11):6032–6042. doi:10.1128/JVI.02504-06
Mujtaba S, Winer BY, Jaganathan A, Patel J, Sgobba M, Schuch R, Gupta YK, Haider S, Wang R, Fischetti VA (2013) Anthrax SET protein: a potential virulence determinant that epigenetically represses NF-kappaB activation in infected macrophages. J Biol Chem 288(32):23458–23472. doi:10.1074/jbc.M113.467696
Netea MG, Joosten LA, Latz E, Mills KH, Natoli G, Stunnenberg HG, O’Neill LA, Xavier RJ (2016) Trained immunity: a program of innate immune memory in health and disease. Science 352(6284):aaf1098. doi:10.1126/science.aaf1098
Nigro G, Rossi R, Commere PH, Jay P, Sansonetti PJ (2014) The cytosolic bacterial peptidoglycan sensor Nod2 affords stem cell protection and links microbes to gut epithelial regeneration. Cell Host Microbe 15(6):792–798. doi:10.1016/j.chom.2014.05.003
Niwa T, Tsukamoto T, Toyoda T, Mori A, Tanaka H, Maekita T, Ichinose M, Tatematsu M, Ushijima T (2010) Inflammatory processes triggered by Helicobacter pylori infection cause aberrant DNA methylation in gastric epithelial cells. Cancer Res 70(4):1430–1440. doi:10.1158/0008-5472.CAN-09-2755
Noordermeer D, Duboule D (2013) Chromatin looping and organization at developmentally regulated gene loci. Wiley Interdiscip Rev Dev Biol 2(5):615–630. doi:10.1002/wdev.103
Ohta K, Kawano R, Ito N (2012) Lactic acid bacteria convert human fibroblasts to multipotent cells. PloS one 7(12):e51866. doi:10.1371/journal.pone.0051866
Okuda J, Toyotome T, Kataoka N, Ohno M, Abe H, Shimura Y, Seyedarabi A, Pickersgill R, Sasakawa C (2005) Shigella effector IpaH9.8 binds to a splicing factor U2AF35 to modulate host immune responses. Biochem Biophys Res Commun 333 (2):531–539. doi:10.1016/j.bbrc.2005.05.145
Opitz B, Puschel A, Beermann W, Hocke AC, Forster S, Schmeck B, van Laak V, Chakraborty T, Suttorp N, Hippenstiel S (2006) Listeria monocytogenes activated p38 MAPK and induced IL-8 secretion in a nucleotide-binding oligomerization domain 1-dependent manner in endothelial cells. J Immunol 176(1):484–490. doi:10.4049/jimmunol.176.1.484
Ostuni R, Piccolo V, Barozzi I, Polletti S, Termanini A, Bonifacio S, Curina A, Prosperini E, Ghisletti S, Natoli G (2013) Latent enhancers activated by stimulation in differentiated cells. Cell 152 (1–2):157–171. doi:10.1016/j.cell.2012.12.018
Pacis A, Tailleux L, Morin AM, Lambourne J, MacIsaac JL, Yotova V, Dumaine A, Danckaert A, Luca F, Grenier JC, Hansen KD, Gicquel B, Yu M, Pai A, He C, Tung J, Pastinen T, Kobor MS, Pique-Regi R, Gilad Y, Barreiro LB (2015) Bacterial infection remodels the DNA methylation landscape of human dendritic cells. Genome Res 25(12):1801–1811. doi:10.1101/gr.192005.115
Padeken J, Heun P (2014) Nucleolus and nuclear periphery: velcro for heterochromatin. Curr Opin Cell Biol 28:54–60. doi:10.1016/j.ceb.2014.03.001
Pamer EG (2004) Immune responses to Listeria monocytogenes. Nat Rev Immunol 4(10):812–823. doi:10.1038/nri1461
Park J, Kim KJ, K-S C, Grab DJ, Dumler JS (2004) Anaplasma phagocytophilum AnkA binds to granulocyte DNA and nuclear proteins. Cell Microbiol 6(8):743–751. doi:10.1111/j.1462-5822.2004.00400.x
Pathak SK, Basu S, Bhattacharyya A, Pathak S, Banerjee A, Basu J, Kundu M (2006) TLR4-dependent NF-kappaB activation and mitogen- and stress-activated protein kinase 1-triggered phosphorylation events are central to Helicobacter pylori peptidyl prolyl cis-, trans-isomerase (HP0175)-mediated induction of IL-6 release from macrophages. J Immunol 177(11):7950–7958. doi:10.4049/jimmunol.177.11.7950
Pennini ME, Liu Y, Yang J, Croniger CM, Boom WH, Harding CV (2007) CCAAT/enhancer-binding protein beta and delta binding to CIITA promoters is associated with the inhibition of CIITA expression in response to Mycobacterium tuberculosis 19-kDa lipoprotein. J Immunol 179 (10):6910–6918. doi:179/10/6910 [pii]
Pennini ME, Pai RK, Schultz DC, Boom WH, Harding CV (2006) Mycobacterium tuberculosis 19-kDa lipoprotein inhibits IFN-gamma-induced chromatin remodeling of MHC2TA by TLR2 and MAPK signaling. J Immunol 176(7):4323–4330. doi:10.4049/jimmunol.176.7.4323
Pennini M, Perrinet S, Dautry-Varsat A, Subtil A (2010) Histone methylation by NUE, a novel nuclear effector of the intracellular pathogen Chlamydia trachomatis. PLoS Pathog 6(7):e1000995. doi:10.1371/journal.ppat.1000995
Perdiguero EG, Geissmann F (2016) The development and maintenance of resident macrophages. Nat Immunol 17(1):2–8. doi:10.1038/ni.3341
Quintin J, Saeed S, Martens JH, Giamarellos-Bourboulis EJ, Ifrim DC, Logie C, Jacobs L, Jansen T, Kullberg BJ, Wijmenga C, Joosten LA, Xavier RJ, van der Meer JW, Stunnenberg HG, Netea MG (2012) Candida albicans infection affords protection against reinfection via functional reprogramming of monocytes. Cell Host Microbe 12(2):223–232. doi:10.1016/j.chom.2012.06.006
Rais Y, Zviran A, Geula S, Gafni O, Chomsky E, Viukov S, Mansour AA, Caspi I, Krupalnik V, Zerbib M, Maza I, Mor N, Baran D, Weinberger L, Jaitin DA, Lara-Astiaso D, Blecher-Gonen R, Shipony Z, Mukamel Z, Hagai T, Gilad S, Amann-Zalcenstein D, Tanay A, Amit I, Novershtern N, Hanna JH (2013) Deterministic direct reprogramming of somatic cells to pluripotency. Nature 502(7469):65–70. doi:10.1038/nature12587
Ram O, Goren A, Amit I, Shoresh N, Yosef N, Ernst J, Kellis M, Gymrek M, Issner R, Coyne M, Durham T, Zhang X, Donaghey J, Epstein CB, Regev A, Bernstein BE (2011) Combinatorial patterning of chromatin regulators uncovered by genome-wide location analysis in human cells. Cell 147(7):1628–1639. doi:10.1016/j.cell.2011.09.057
Raymond B, Batsche E, Boutillon F, Wu YZ, Leduc D, Balloy V, Raoust E, Muchardt C, Goossens PL, Touqui L (2009) Anthrax lethal toxin impairs IL-8 expression in epithelial cells through inhibition of histone H3 modification. PLoS Pathog 5(4):e1000359. doi:10.1371/journal.ppat.1000359
Rennoll-Bankert KE, Dumler JS (2012) Lessons from Anaplasma phagocytophilum: chromatin remodeling by bacterial effectors. Infect Disord Drug Targets 12(5):380–387
Rennoll-Bankert KE, Garcia-Garcia JC, Sinclair SH, Dumler JS (2015) Chromatin-bound bacterial effector ankyrin A recruits histone deacetylase 1 and modifies host gene expression. Cell Microbiol 17(11):1640–1652. doi:10.1111/cmi.12461
Riggs AD, Martienssen RA, Russo VEA (1996) Introduction. In Russo, VEA et al (ed) Epigenetic mechanisms of gene regulation. Cold Spring Harbor Laboratory Press, New York, p 1
Riggs MG, Whittaker RG, Neumann JR, Ingram VM (1977) n-Butyrate causes histone modification in HeLa and Friend erythroleukaemia cells. Nature 268(5619):462–464
Roadmap Epigenomics C, Kundaje A, Meuleman W, Ernst J, Bilenky M, Yen A, Heravi-Moussavi A, Kheradpour P, Zhang Z, Wang J, Ziller MJ, Amin V, Whitaker JW, Schultz MD, Ward LD, Sarkar A, Quon G, Sandstrom RS, Eaton ML, Wu YC, Pfenning AR, Wang X, Claussnitzer M, Liu Y, Coarfa C, Harris RA, Shoresh N, Epstein CB, Gjoneska E, Leung D, Xie W, Hawkins RD, Lister R, Hong C, Gascard P, Mungall AJ, Moore R, Chuah E, Tam A, Canfield TK, Hansen RS, Kaul R, Sabo PJ, Bansal MS, Carles A, Dixon JR, Farh KH, Feizi S, Karlic R, Kim AR, Kulkarni A, Li D, Lowdon R, Elliott G, Mercer TR, Neph SJ, Onuchic V, Polak P, Rajagopal N, Ray P, Sallari RC, Siebenthall KT, Sinnott-Armstrong NA, Stevens M, Thurman RE, Wu J, Zhang B, Zhou X, Beaudet AE, Boyer LA, De Jager PL, Farnham PJ, Fisher SJ, Haussler D, Jones SJ, Li W, Marra MA, McManus MT, Sunyaev S, Thomson JA, Tlsty TD, Tsai LH, Wang W, Waterland RA, Zhang MQ, Chadwick LH, Bernstein BE, Costello JF, Ecker JR, Hirst M, Meissner A, Milosavljevic A, Ren B, Stamatoyannopoulos JA, Wang T, Kellis M (2015) Integrative analysis of 111 reference human epigenomes. Nature 518(7539):317–330. doi:10.1038/nature14248
Rohde JR, Breitkreutz A, Chenal A, Sansonetti PJ, Parsot C (2007) Type III secretion effectors of the IpaH family are E3 ubiquitin ligases. Cell Host Microbe 1(1):77–83. doi:10.1016/j.chom.2007.02.002
Rolando M, Sanulli S, Rusniok C, Gomez-Valero L, Bertholet C, Sahr T, Margueron R, Buchrieser C (2013) Legionella pneumophila effector RomA uniquely modifies host chromatin to repress gene expression and promote intracellular bacterial replication. Cell Host Microbe 13(4):395–405. doi:10.1016/j.chom.2013.03.004
Romanoski CE, Glass CK, Stunnenberg HG, Wilson L, Almouzni G (2015) Epigenomics: roadmap for regulation. Nature 518(7539):314–316. doi:10.1038/518314a
Rye M, Sandve GK, Daub CO, Kawaji H, Carninci P, Forrest AR, Drablos F, Fantom Consortium (2014) Chromatin states reveal functional associations for globally defined transcription start sites in four human cell lines. BMC Genomics 15:120. doi:10.1186/1471-2164-15-120
Saccani S, Pantano S, Natoli G (2002) p38-Dependent marking of inflammatory genes for increased NF-kappa B recruitment. Nat Immunol 3(1):69–75. doi:10.1038/ni748
Sadakierska-Chudy A, Filip M (2015) A comprehensive view of the epigenetic landscape. Part II: Histone post-translational modification, nucleosome level, and chromatin regulation by ncRNAs. Neurotox Res 27(2):172–197. doi:10.1007/s12640-014-9508-6
Salles, II, Tucker AE, Voth DE, Ballard JD (2003) Toxin-induced resistance in Bacillus anthracis lethal toxin-treated macrophages. Proc Natl Acad Sci U S A 100 (21):12426–12431. doi:10.1073/pnas.2134042100
Samba-Louaka A, Pereira JM, Nahori MA, Villiers V, Deriano L, Hamon MA, Cossart P (2014) Listeria monocytogenes dampens the DNA damage response. PLoS Pathog 10(10):e1004470. doi:10.1371/journal.ppat.1004470
Sasai N, Defossez PA (2009) Many paths to one goal? The proteins that recognize methylated DNA in eukaryotes. Int J Dev Biol 53(2–3):323–334. doi:10.1387/ijdb.082652ns
Schmeck B, Lorenz J, N'Guessan PD, Opitz B, van Laak V, Zahlten J, Slevogt H, Witzenrath M, Flieger A, Suttorp N, Hippenstiel S (2008) Histone acetylation and flagellin are essential for Legionella pneumophila-induced cytokine expression. J Immunol 181(2):940–947. doi:10.4049/jimmunol.181.2.940
Schmitz KM, Mayer C, Postepska A, Grummt I (2010) Interaction of noncoding RNA with the rDNA promoter mediates recruitment of DNMT3b and silencing of rRNA genes. Genes Dev 24(20):2264–2269. doi:10.1101/gad.590910
Segain JP, Raingeard de la Bletiere D, Bourreille A, Leray V, Gervois N, Rosales C, Ferrier L, Bonnet C, Blottiere HM, Galmiche JP (2000) Butyrate inhibits inflammatory responses through NFkappaB inhibition: implications for Crohn’s disease. Gut 47(3):397–403
Seyedarabi A, Sullivan JA, Sasakawa C, Pickersgill RW (2011) A disulfide driven domain swap switches off the activity of Shigella IpaH9.8 E3 ligase. FEBS Lett 584(19):4163–4168. doi:10.1016/j.febslet.2010.09.006
Shames SR, Bhavsar AP, Croxen MA, Law RJ, Mak SH, Deng W, Li Y, Bidshari R, de Hoog CL, Foster LJ, Finlay BB (2011) The pathogenic Escherichia coli type III secreted protease NleC degrades the host acetyltransferase p300. Cell Microbiol 13(10):1542–1557. doi:10.1111/j.1462-5822.2011.01640.x
Sharma G, Sowpati DT, Singh P, Khan MZ, Ganji R, Upadhyay S, Banerjee S, Nandicoori VK, Khosla S (2016) Genome-wide non-CpG methylation of the host genome during M. tuberculosis infection. Sci Rep 6:25006. doi:10.1038/srep25006
Sharma G, Upadhyay S, Srilalitha M, Nandicoori VK, Khosla S (2015) The interaction of mycobacterial protein Rv2966c with host chromatin is mediated through non-CpG methylation and histone H3/H4 binding. Nucleic Acids Res 43(8):3922–3937. doi:10.1093/nar/gkv261
Sinclair SH, Garcia-Garcia JC, Dumler JS (2015a) Bioinformatic and mass spectrometry identification of Anaplasma phagocytophilum proteins translocated into host cell nuclei. Front Microbiol 6:55. doi:10.3389/fmicb.2015.00055
Sinclair SH, Rennoll-Bankert KE, Dumler JS (2014) Effector bottleneck: microbial reprogramming of parasitized host cell transcription by epigenetic remodeling of chromatin structure. Front Genet 5:274. doi:10.3389/fgene.2014.00274
Sinclair SH, Yegnasubramanian S, Dumler JS (2015b) Global DNA methylation changes and differential gene expression in Anaplasma phagocytophilum-infected human neutrophils. Clin Epigenetics 7(1):77. doi:10.1186/s13148-015-0105-1
Slevogt H, Schmeck B, Jonatat C, Zahlten J, Beermann W, van Laak V, Opitz B, Dietel S, N'Guessan PD, Hippenstiel S, Suttorp N, Seybold J (2006) Moraxella catarrhalis induces inflammatory response of bronchial epithelial cells via MAPK and NF-kappaB activation and histone deacetylase activity reduction. Am J Physiol Lung Cell Mol Physiol 290(5):L818–L826. doi:10.1152/ajplung.00428.2005
Sommer F, Backhed F (2013) The gut microbiota—masters of host development and physiology. Nat Rev Microbiol 11(4):227–238. doi:10.1038/nrmicro2974
Soundararajan V, Patel N, Subramanian V, Sasisekharan V, Sasisekharan R (2011) The many faces of the YopM effector from plague causative bacterium Yersinia pestis and its implications for host immune modulation. Innate Immun 17(6):548–557. doi:10.1177/1753425910377099
Suganuma T, Workman JL (2011) Signals and combinatorial functions of histone modifications. Annu Rev Biochem 80:473–499. doi:10.1146/annurev-biochem-061809-175347
Sun J (2010) Enteric bacteria and cancer stem cells. Cancers 3(1):285–297. doi:10.3390/cancers3010285
Tahoun A, Mahajan S, Paxton E, Malterer G, Donaldson DS, Wang D, Tan A, Gillespie TL, O'Shea M, Roe AJ, Shaw DJ, Gally DL, Lengeling A, Mabbott NA, Haas J, Mahajan A (2012) Salmonella transforms follicle-associated epithelial cells into M cells to promote intestinal invasion. Cell Host Microbe 12(5):645–656. doi:10.1016/j.chom.2012.10.009
Takahashi K, Sugi Y, Nakano K, Tsuda M, Kurihara K, Hosono A, Kaminogawa S (2011) Epigenetic control of the host gene by commensal bacteria in large intestinal epithelial cells. J Biol Chem 286(41):35755–35762. doi:10.1074/jbc.M111.271007
Takahashi K, Yamanaka S (2006) Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126(4):663–676. doi:10.1016/j.cell.2006.07.024
Taverna SD, Li H, Ruthenburg AJ, Allis CD, Patel DJ (2007) How chromatin-binding modules interpret histone modifications: lessons from professional pocket pickers. Nat Struct Mol Biol 14(11):1025–1040. doi:10.1038/nsmb1338
Ting K, Aitken KJ, Penna F, Samiei AN, Sidler M, Jiang JX, Ibrahim F, Tolg C, Delgado-Olguin P, Rosenblum N, Bagli DJ (2016) Uropathogenic E.coli (UPEC) infection induces proliferation through Enhancer of Zeste Homologue 2 (EZH2). PLoS One 11(3):e0149118. doi:10.1371/journal.pone.0149118
Tisseur M, Kwapisz M, Morillon A (2011) Pervasive transcription – Lessons from yeast. Biochimie 93(11):1889–1896. doi:10.1016/j.biochi.2011.07.001
Tiwari VK, Stadler MB, Wirbelauer C, Paro R, Schubeler D, Beisel C (2011) A chromatin-modifying function of JNK during stem cell differentiation. Nat Genet 44(1):94–100. doi:10.1038/ng.1036
Tolg C, Sabha N, Cortese R, Panchal T, Ahsan A, Soliman A, Aitken KJ, Petronis A, Bägli DJ (2011) Uropathogenic E. coli infection provokes epigenetic downregulation of CDKN2A (p16INK4A) in uroepithelial cells. Lab Invest 91:825–836
Toyotome T (2001) Shigella protein IpaH9.8 Is secreted from bacteria within mammalian cells and transported to the nucleus. J Biol Chem 276(34):32071–32079. doi:10.1074/jbc.M101882200
Turgeon N, Blais M, Gagne JM, Tardif V, Boudreau F, Perreault N, Asselin C (2013) HDAC1 and HDAC2 restrain the intestinal inflammatory response by regulating intestinal epithelial cell differentiation. PLoS One 8(9):e73785. doi:10.1371/journal.pone.0073785
Turner AM, Morris KV (2010) Controlling transcription with noncoding RNAs in mammalian cells. Biotechniques 48(6):ix–xvi. doi:10.2144/000113442
Ushijima T, Hattori N (2012) Molecular Pathways: involvement of helicobacter pylori -triggered inflammation in the formation of an epigenetic field defect, and its usefulness as cancer risk and exposure markers. Clin Cancer Res. doi:10.1158/1078-0432.CCR-11-2011
Valenzuela MA, Canales J, Corvalan AH, Quest AF (2015) Helicobacter pylori-induced inflammation and epigenetic changes during gastric carcinogenesis. World J Gastroenterol 21(45):12742–12756. doi:10.3748/wjg.v21.i45.12742
Vire E, Brenner C, Deplus R, Blanchon L, Fraga M, Didelot C, Morey L, Van Eynde A, Bernard D, Vanderwinden JM, Bollen M, Esteller M, Di Croce L, de Launoit Y, Fuks F (2006) The Polycomb group protein EZH2 directly controls DNA methylation. Nature 439(7078):871–874. doi:10.1038/nature04431
Wakeel A, Zhu B, Yu XJ, McBride JW (2010) New insights into molecular Ehrlichia chaffeensis-host interactions. Microbes Infect 12(5):337–345. doi:10.1016/j.micinf.2010.01.009
Wang Y, Curry HM, Zwilling BS, Lafuse WP (2005) Mycobacteria inhibition of IFN-gamma induced HLA-DR gene expression by up-regulating histone deacetylation at the promoter region in human THP-1 monocytic cells. J Immunol 174(9):5687–5694. doi:10.4049/jimmunol.174.9.5687
Wang TY, Han ZM, Chai YR, Zhang JH (2010) A mini review of MAR-binding proteins. Mol Biol Rep 37 (7):3553–3560. doi:10.1007/s11033-010-0003-8
Wani AH, Boettiger AN, Schorderet P, Ergun A, Munger C, Sadreyev RI, Zhuang X, Kingston RE, Francis NJ (2016) Chromatin topology is coupled to Polycomb group protein subnuclear organization. Nat Commun 7:10291. doi:10.1038/ncomms10291
Weber MM, Faris R, McLachlan J, Tellez A, Wright WU, Galvan G, Luo ZQ, Samuel JE (2016) Modulation of the host transcriptome by Coxiella burnetii nuclear effector Cbu1314. Microbes Infect 18(5):336–345. doi:10.1016/j.micinf.2016.01.003
Wu H, Zhang Y (2011) Mechanisms and functions of Tet protein-mediated 5-methylcytosine oxidation. Genes Dev 25(23):2436–2452. doi:10.1101/gad.179184.111
Xia G, Schneider-Stock R, Diestel A, Habold C, Krueger S, Roessner A, Naumann M, Lendeckel U (2008) Helicobacter pylori regulates p21(WAF1) by histone H4 acetylation. Biochem Biophys Res Commun 369 (2):526–531. doi:1016/j.bbrc.2008.02.073
Yamamoto Y, Verma UN, Prajapati S, Kwak YT, Gaynor RB (2003) Histone H3 phosphorylation by IKK-alpha is critical for cytokine-induced gene expression. Nature 423(6940):655–659. doi:10.1038/nature01576
Yan J, Zhang M, Zhang J, Chen X, Zhang X (2011) Helicobacter pylori infection promotes methylation of WWOX gene in human gastric cancer. Biochem Biophys Res Commun 408(1):99–102. doi:10.1016/j.bbrc.2011.03.127
Yang L, Lin C, Liu W, Zhang J, Ohgi KA, Grinstein JD, Dorrestein PC, Rosenfeld MG (2011) ncRNA- and Pc2 methylation-dependent gene relocation between nuclear structures mediates gene activation programs. Cell 147(4):773–788. doi:10.1016/j.cell.2011.08.054
Yao Y, Tao H, Park DI, Sepulveda JL, Sepulveda AR (2006) Demonstration and characterization of mutations induced by Helicobacter pylori organisms in gastric epithelial cells. Helicobacter 11(4):272–286. doi:10.1111/j.1523-5378.2006.00408.x
Yaseen I, Kaur P, Nandicoori VK, Khosla S (2015) Mycobacteria modulate host epigenetic machinery by Rv1988 methylation of a non-tail arginine of histone H3. Nat Commun 6:8922. doi:10.1038/ncomms9922
Ye F, Karn J (2015) Bacterial short chain fatty acids push all the buttons needed to reactivate latent viruses. Stem Cell Epigenetics 2(1). doi:10.14800/sce.532
Yin L, Chung WO (2011) Epigenetic regulation of human beta-defensin 2 and CC chemokine ligand 20 expression in gingival epithelial cells in response to oral bacteria. Mucosal Immunol 4(4):409–419. doi:10.1038/mi.2010.83
Yuan X, Feng W, Imhof A, Grummt I, Zhou Y (2007) Activation of RNA polymerase I transcription by cockayne syndrome group B protein and histone methyltransferase G9a. Mol Cell 27(4):585–595. doi:10.1016/j.molcel.2007.06.021
Zhang S, Barros SP, Niculescu MD, Moretti AJ, Preisser JS, Offenbacher S (2010) Alteration of PTGS2 promoter methylation in chronic periodontitis. J Dent Res 89(2):133–137. doi:10.1177/0022034509356512
Zhou Y, Kim J, Yuan X, Braun T (2011) Epigenetic modifications of stem cells: a paradigm for the control of cardiac progenitor cells. Circ Res 109(9):1067–1081. doi:10.1161/CIRCRESAHA.111.243709
Zhu B, Nethery KA, Kuriakose JA, Wakeel A, Zhang X, McBride JW (2009) Nuclear translocated ehrlichia chaffeensis ankyrin protein interacts with a specific adenine-rich motif of host promoter and intronic alu elements. Infect Immun 77(10):4243–4255. doi:10.1128/IAI.00376-09
Zurawski DV, Mumy KL, Faherty CS, Mccormick BA, Maurelli AT (2009) Shigella flexneritype III secretion system effectors OspB and OspF target the nucleus to downregulate the host inflammatory response via interactions with retinoblastoma protein. Mol Microbiol 71(2):350–368. doi:10.1111/j.1365-2958.2008.06524.x
Acknowledgements
I apologize to colleagues whose work was not cited here. I acknowledge support from INRA, French Ligue Nationale Contre le Cancer (LNCC RS10/75-76 Bierne), French National Research Agency (ANR, grant EPILIS), and the iXcore Research Foundation.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Bierne, H. (2017). Cross Talk Between Bacteria and the Host Epigenetic Machinery. In: Doerfler, W., Casadesús, J. (eds) Epigenetics of Infectious Diseases. Epigenetics and Human Health. Springer, Cham. https://doi.org/10.1007/978-3-319-55021-3_6
Download citation
DOI: https://doi.org/10.1007/978-3-319-55021-3_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-55019-0
Online ISBN: 978-3-319-55021-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)