Abstract
Proteins catalysing the transport of molecules across biological membranes are vital for organisms in all kingdoms of life. These proteins are needed for the uptake of nutrients and the efflux of signalling molecules and toxic compounds. In the human opportunistic pathogen Candida albicans, efflux proteins can translocate antifungal drugs, such as fluconazole, and confer drug resistance. There are two main families of membrane proteins involved in drug transport, the major facilitator superfamily (MFS) and the ATP-binding cassette (ABC) proteins. Both types of protein possess multiple membrane spanning α-helices in transmembrane domains (TMDs), and ABC proteins, in addition, contain cytosolic nucleotide-binding domains (NBDs) involved in ATP hydrolysis. ABC proteins, and to a lesser extent MFS proteins, have broad substrate specificities that are determined by the structure and arrangement of the transmembrane α-helices. The multidrug transporter most often associated with drug resistance of C. albicans clinical isolates is ABC protein Cdr1. This is a pleiotropic drug resistance (PDR) ABC protein with domain arrangement NBD1-TMD1-NBD2-TMD2 and unique large extracellular loops between transmembrane α-helices that may be important for pump function. There is no crystal structure for Cdr1, but X-ray structures of related proteins indicate that unique interactions between conserved elements of the TMDs and the NBDs are important for pump function.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- ATP-binding cassette (ABC)
- Major facilitator superfamily (MFS)
- Chemosensitization
- Suppressor mutation
- Drug resistance
18.1 Introduction
Fungi possess a variety of integral membrane proteins, the functions of which are to transport substances across biological membranes (Perlin et al. 2014). They are present in many cellular membranes, including vacuolar, mitochondrial, peroxisomal and endoplasmic reticulum membranes, with many important transporters located in the plasma membrane. The plasma membrane is a vital permeability barrier for cellular homeostasis. This barrier stops the free flow of ions, small molecules and macromolecules across the membrane and maintains chemical and electrochemical gradients between the inside and the outside of the fungal cell. In order to transport molecules across this barrier, the fungal cell utilises integral membrane proteins. If the molecule moves down a concentration gradient facilitated by membrane protein channels or pores, then the transport protein simply accelerates passive diffusion, and no energy is required for this process. In many instances, however, molecules are transported against a concentration gradient, and energy is required.
Energy-dependent transport is carried out predominantly by proteins belonging to one of two large protein families, either the ATP-binding cassette (ABC) transporter family, or the major facilitator superfamily (MFS). ABC transporters are primary active transporters that use the energy released by ATP hydrolysis to power transport, whereas MFS proteins are secondary active transporters that utilise electrochemical gradients (e.g. Na+ or H+ gradients established by primary active transporters) across the membrane as their energy source. Fungi generate many compounds that are toxic to the cell unless effluxed such as secondary metabolites or endpoints of metabolic pathways including ethanol and organic acids. Fungi also encounter xenobiotics from their environment, which are kept outside the cell by multiple efflux proteins in plasma membranes. Multidrug efflux proteins transport a variety of substrates, and the range of substrates for different pumps can overlap. It is interesting to note that fungi that inhabit a variety of environmental niches, such as Cryptococcus neoformans, contain a large repertoire of 33 ABC (Kovalchuk and Driessen 2010) and 150–170 MFS proteins (Janbon et al. 2014). Another example of species that inhabit a variety of niches are Aspergillus spp. with 50–70 ABC proteins (Kovalchuk and Driessen 2010) and 275–356 MFS transporters (Coleman and Mylonakis 2009). In contrast, the fungus Candida albicans, which is almost entirely associated with mammalian hosts, possesses a more limited range of 26 ABC (Kovalchuk and Driessen 2010) and 85–95 MFS transporters (Coleman and Mylonakis 2009; Gaur et al. 2008), possibly because of its exposure to a smaller range of xenobiotics.
A complication of treating fungal infections with antifungal drugs is that fungal genomes encode transporters that are able to export these drugs out of the cells thereby rendering the fungi resistant. It has been found that the strong selective pressure exerted by antifungal drugs can select for mutations in transcriptional regulators that result in over-expression of ABC or MFS transporters. For example, the high-level azole resistance of C. albicans has been shown to be due to the over-expression of MFS protein Mdr1 or, most frequently, the over-expression of the ABC proteins Cdr1 and Cdr2 (Cannon et al. 2009). Over-expression of Mdr1 can be caused by gain-of-function mutations in transcription factor Mrr1 (Dunkel et al. 2008), whereas the over-expression of Cdr1 and Cdr2 is frequently caused by gain-of-function mutations in transcription factor Tac1 (Coste et al. 2007; Coste et al. 2004). This chapter describes the multidrug transporters of C. albicans, focusing on the clinically more significant ABC proteins, and current knowledge of structure–function relationships for these proteins.
18.2 Multidrug Efflux Transporters of C. albicans
18.2.1 MFS Transporters
MFS transporters are integral membrane proteins with, most commonly, either 12 or 14 transmembrane spans (TMSs); exceptions do exist with one MFS transporter predicted to have 6 and another small MFS family predicted to have 24 TMSs (Law et al. 2008; Reddy et al. 2012). C. albicans possesses an estimated 95 MFS transporters (Gaur et al. 2008) that can be divided into 17 separate phylogenetic lineages. The four major MFS families are the SP (sugar porter), the DHA1 (drug: H+ antiporter 1), the DHA2 and the ACS (anion:cation symporter) family with 22, 22, 9 and 16 representatives, respectively (Gaur et al. 2008). SP-, DHA1- and ACS-type transporters are predicted to have 12 TMSs and DHA2 members have 14 TMSs. The C. albicans genome encodes between 26 (Costa et al. 2014) and 31 (Gaur et al. 2008) DHA1 and DHA2 MFS transporters. Few of these ORFs have been characterised and only one, Mdr1, is thought to be a clinically relevant MFS multidrug efflux pump. C. albicans Mdr1, a DHA1 protein (Fig. 18.1 and Table 18.1) also called Benr, was originally identified by its ability to confer benomyl and methotrexate resistance on Saccharomyces cerevisiae (Fling et al. 1991). It prefers generally less hydrophobic and less branched substrates than C. albicans Cdr1 (Puri et al. 2010), and it transports compounds such as fluconazole (FLC) and ketoconazole (KTZ), but not miconazole (MCZ) or itraconazole (ITC) (Lamping et al. 2007). High levels of MDR1 expression have been detected in certain azole-resistant clinical isolates of C. albicans (Sanglard et al. 1995; White et al. 2002; Wirsching et al. 2000). In general, MFS multidrug efflux transporters efflux a narrower range of substrates than ABC multidrug efflux proteins (Puri et al. 2010; Sa-Correia et al. 2009). Mdr1 expression is up-regulated in clinical isolates by gain-of-function mutations in transcription factor Mrr1 (Dunkel et al. 2008; Morschhauser et al. 2007). For further details on the transcriptional control of C. albicans MFS transporters see the next chapter. Another DHA1 MFS transporter with a potential role in drug resistance is Flu1 (Table 18.1). Although there is no evidence of Flu1 over-expression in azole-resistant clinical isolates, disruption of FLU1 in C. albicans caused susceptibility to mycophenolic acid but it had no effect on FLC sensitivity (Calabrese et al. 2000). Flu1 is thought to efflux the human fungicidal peptide histatin 5 (Hst 5), although Hst 5 did not induce over-expression of FLU1 mRNA (Li et al. 2013).
Basic topologies of MFS, and PDR and MDR ABC transporters. Cylinders indicate α-helices, the arrow indicates a β-sheet, ovals indicate cytosolic NBDs, and the plasma membrane (PM) separating the inside (in) from the outside (out) of the cell is represented by two dashed lines. a General topology of DHA1 MFS transporters such as C. albicans Mdr1 (Ca Mdr1). DHA1 MFS transporters typically comprise ~500 amino acids including 12 TMSs that are connected by short intracellular loops (ILs) or extracellular loops (ELs). The core MFS transporter fold consists of four TMS repeats (i.e. TMS1-3, TMS4-6, TMS7-9, TMS10-12) that originated from two separate tandem-duplications of a three TMS ancestor; i.e. TMS1/4/7/10, TMS2/5/8/11 and TMS3/6/9/12 share the same TMS ancestor (also see Fig. 18.2a). b The prototypic fungal PDR family multidrug efflux transporters S. cerevisiae Pdr5 and C. albicans Cdr1 (Ca Cdr1; top panel), are characterised by a topology with the NBDs NBD1 and NBD2 preceding TMD1 (TMS1-6) and TMD2 (TMS7-12), respectively. All other ABC transporter families, including the MDR family such as the human multidrug efflux pump HsABCB1 (Hs ABCB1; bottom panel), exhibit the typical ABC transporter topology with the TMDs preceding the respective NBDs
Two other genes with homology to DHA1 MFS transporters, NAG3 (also known as TMP1) and NAG4 (also known as TMP2), may also be involved in drug transport (Table 18.1). When these genes in C. albicans are disrupted the cells become more susceptible to cycloheximide (CHX) (Yamada-Okabe and Yamada-Okabe 2002) but their over-expression has not been associated with clinical drug resistance.
18.2.2 ABC Transporters
As their name suggests, ABC proteins contain nucleotide-binding domains (NBDs). These domains contain highly conserved amino acid motifs and are characterised by the Walker A and the Walker B motifs (Walker et al. 1982) and the hallmark ABC signature motif or C-loop. The 26 C. albicans ABC proteins (Braun et al. 2005) can be classified into the following subgroups: PDR (pleiotropic drug resistance), MDR (multidrug resistance), MRP (multidrug resistance-associated protein), ALDp (adrenoleukodystrophy protein), EF3 (elongation factor EF-3) and RL1 (RNAse L inhibitor) (Gaur et al. 2005). The EF3 and RL1 ABC proteins lack TMSs and so, despite the ability to hydrolyse ATP, are not involved in transport.
The two ABC proteins most often associated with drug resistance in clinical C. albicans isolates are Cdr1 and Cdr2 (Maebashi et al. 2001; Sanglard et al. 1995; White 1997) (Table 18.1). These are ‘full-size’ ABC transporters in that they contain two homologous halves each with a NBD and a TMD (Cannon et al. 2009). The two TMDs each contain six TMSs and together they are thought to form the substrate transport channel. Cdr1 and Cdr2 are PDR-type ABC proteins with an inverted order of the domains [NBD-TMD]2—only found in fungi and plants (Crouzet et al. 2006)—as opposed to the typical [TMD-NBD]2 arrangement found in all other ABC proteins (Fig. 18.1) (Lamping et al. 2010).
Cdr1 and Cdr2 have a broad substrate specificity and, when heterologously expressed in S. cerevisiae, they confer resistance to FLC, KTZ, MCZ, ITC, rhodamine 6G (R6G), CHX and cerulenin (CER) among many other compounds (Puri et al. 2010; Tanabe et al. 2011). Both Cdr1 and Cdr2 are often co-expressed at elevated levels in azole-resistant clinical C. albicans isolates because their promoters contain drug-responsive elements CGGN4CGG (de Micheli et al. 2002; Liu et al. 2007); the sequence recognised by transcription factor Tac1 (Coste et al. 2004). These azole-resistant isolates possess gain-of-function mutations in Tac1 (Coste et al. 2007; Coste et al. 2004). The use of Cdr1- and Cdr2-specific antibodies, Cdr1-specific inhibitors, and gene deletions, has shown that Cdr1 contributes more to FLC efflux than Cdr2 in FLC-resistant clinical isolates (Holmes et al. 2008; Tsao et al. 2009).
18.3 Topology of C. albicans Multidrug Efflux Transporters
18.3.1 MFS Transporter Mdr1
C. albicans Mdr1 has 12 predicted TMSs connected by hydrophilic loops (Fig. 18.1a) (Pasrija et al. 2007). Unlike other MFS transporters such as human glucose transporters GLUT1, -3 or -5 or Escherichia coli lactose transporter LacY, multidrug efflux transporter EmrD, and tetracycline transporter TetA, most fungal MFS transporters, including S. cerevisiae Flr1 and C. albicans Mdr1, Flu1, Nag3 and Nag4, have a ~100–160 amino acid N-terminal extension (Fig. 18.1a; data not shown), the function of which remains to be determined. Early topology studies of DHA1 transporters using reporter fusions revealed that the N- and C-termini of these proteins are cytoplasmic (Law et al. 2008). There is a degree of homology between the N- and C-terminal halves of MFS proteins indicating that they may have arisen by duplication of an ancestor protein with 6 TMSs (Paulsen and Skurray 1993). These two halves are separated by the extended cytoplasmic intracellular loop 3 (IL3; Fig. 18.1a), which permits a relatively large, rigid, movement between the two domains during the proposed ‘rocker-switch’-type substrate translocation through the centre of the transporter (Law et al. 2008; Yan 2013; Yan 2015). Further structural and functional analyses revealed that an even earlier duplication of a 3-TMS ancestor possibly gave rise to this 6 TMS precursor protein. Thus, the 12 TMS core of most MFS transporters consists of four inverted 3-TMS repeats (Fig. 18.1a) (Quistgaard et al. 2016; Yan 2015).
18.3.2 ABC Transporters Cdr1 and Cdr2
All ABC transporters consist of four core sub-domains: two TMDs that provide a channel for the passage of compounds across the lipid bilayer, and two cytoplasmic NBDs that provide the necessary energy for active transport (Fig. 18.1b). Prokaryotes commonly express these individual sub-domains as separate polypeptide chains, whereas eukaryotes usually express half-size transporters (TMD-NBD) that form functional homo- or hetero-dimers, or full-size transporters with all four sub-domains [TMD-NBD]2 expressed as a single polypeptide (Fig. 18.1b) (Hollenstein et al. 2007a, b; Jones et al. 2009). As mentioned above, PDR transporters like Cdr1 and Cdr2 are the only ABC sub-group with an inverted domain arrangement where the two NBDs precede their respective TMDs [NBD-TMD]2 (Fig. 18.1b). They also have a ~150 amino acid N-terminal extension rich in hydrophilic (S, T, N, Q) and charged (D, E, R, K) residues (Lamping et al. 2010), similar to the fungal MFS transporters mentioned above. This N-terminal domain may be important for protein–protein interactions that regulate PDR transporter function and/or expression. Our analysis of 244 PDR sequences also predicted four small (EL1, -2, -4, -5) and two large ELs (EL3, -6) with PDR family-defining helical and beta-sheet motifs near the exits of TMS5 and TMS11, and two small (IL2, -4) and two large ILs (IL1, -3) connecting 12 predicted TMSs (Fig. 18.1b) (Lamping et al. 2010). Plant and human PDR transporters exhibit the same topology with the same conserved EL3 and EL6 helical and beta-sheet motifs near the exits of TMS5 and TMS11 (Fig. 18.1b; unpublished data). MDR ABC proteins, like human ABCB1 (also known as P-glycoprotein or Pgp), also have 12 TMSs but, unlike PDR transporters such as Cdr1 and Cdr2, they have equally large IL1-IL4 that extend deep into the cytosol, and they do not have large ELs (Fig. 18.1b). As discussed below, this has significant structural and functional consequences for these different types of ABC exporter.
18.4 Structural Analyses of Multidrug Transporters
The structure of proteins, and protein complexes, provides pivotal clues about how proteins fulfil their function. There are several techniques that can generate structural information at atomic or near-atomic resolution—two of the most frequently used techniques being X-ray crystallography and cryo-electron microscopy (CryoEM) (Delmar et al. 2015; Ford et al. 2009; Vinothkumar 2015). There are hundreds of thousands of structures of soluble proteins deposited in the protein data bank (pdb; http://www.rcsb.org/pdb/home/home.do), but high-resolution structures for integral membrane proteins are much harder to obtain and account for less than 1% of all deposited structures. The amphiphilic nature of membrane proteins makes it very difficult to obtain enough mono-disperse protein and to produce the well-ordered three-dimensional (3-D) crystals that are required for X-ray crystallography. However, recent advances in CryoEM technology (i.e. improved detector technology and software analysis tools (Cheng et al. 2015; Delmar et al. 2015; Henderson 2015; Nannenga et al. 2013; Rigaud et al. 2000; Vinothkumar 2015)), which do not require well-ordered 3-D crystals for the structural analysis of proteins, has helped to increase the pace for solving membrane protein structures, including structures for MFS and ABC transporters.
18.4.1 MFS Transporters
There are currently 64 solved structures for 23 MFS transporters representing 10 different MFS transporter families and that display all four major pump conformations (inward open—inward occluded—outward occluded—outward open) of the recently proposed ‘clamp and switch model’ for MFS transporter function (Quistgaard et al. 2016).
It would appear that all MFS transporters, irrespective of their substrate specificities and transport coupling mechanisms, share a common structural fold known as the MFS fold (Yan 2013). The canonical MFS fold comprises 12 TMSs which are organised into two discreetly folded N- and C-terminal domains displaying a two-fold pseudosymmetry (Yan 2013). Each domain further divides into a pair of inverted 3 + 3 TMS repeats, the basic functional unit of MFS proteins (Yan 2013). Interestingly, the corresponding TMSs (i.e. TMS1, -4, -7, -10; TMS2, -5, -8, -11; and TMS3, -6, -9, -12; blue, magenta and green, respectively; Fig. 18.2a) of two 3-TMS repeats are always positioned next to each other in opposite transmembrane orientations (Fig. 18.2a). The substrate-translocation cavity at the centre of the transporter is enclosed by the blue helices (TMS1, -4, -7 and -10; Fig. 18.2a). Major interactions between TMS1 and TMS7 near the extracellular space and between TMS4 and TMS10 near the cytosol are thought to control access of solutes to the substrate-binding cavity (Nomura et al. 2015; Quistgaard et al. 2016; Yan 2015). Some residues of the magenta helices (TMS2, -5, -8 and -11) that typically mediate the interface between the N- and C-terminal domains (Fig. 18.2a), also face the substrate-binding cavity and may participate in substrate binding/transport and coupling of substrate transport to the symport or antiport of H+ or other chemiosmotic solutes. The green helices (TMS3, -6, -9 and -12; Fig. 18.2a) are positioned on the outside of the protein, mostly stabilising MFS transporter structure. Most MFS transporters also have a conserved hallmark A-motif Gx3(D/E)(R/K)xG[x](R/K)(R/K) (x indicates any residue) between TMS2 and -3 and/or TMS8 and -9 (Quistgaard et al. 2016). These motifs are important for gating interactions in the four different MFS pump conformations (Quistgaard et al. 2016; Yan 2013; Yan 2015). Some drug: H+ MFS antiporter families, including C. albicans Mdr1, also have a conserved C-motif Gx3GPx2G near the end of TMS5 (Paulsen et al. 1996; Paulsen and Skurray 1993), which has been proposed to be involved in linking H+ translocation to export as it is only conserved among export proteins (Paulsen and Skurray 1993).
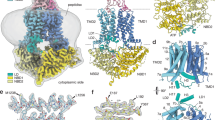
Structures of a typical MFS transporter and type I and type II ABC exporters. All structures were prepared with PyMOL Molecular Graphics System. The lipid bilayer is indicated with dashed grey lines. a E. coli multidrug efflux transporter EcEmrD (pdb: 2GFP; (Yin et al. 2006)), like C. albicans Mdr1 an H+/drug antiporter. Because all MFS transporters share the same core fold, it is likely that C. albicans Mdr1 folds like EcEmrD. The top panel in a shows how a typical MFS transporter contains four core subunits. Subunit I comprises TMS1/2/3, subunit II TMS4/5/6, subunit III TMS7/8/9 and subunit IV comprises TMS10/11/12. MFS transporters possibly experienced two separate gene duplications of a common 3 TMS ancestor. Thus, TMSs with the same colour (blue, magenta or yellow) are equivalent and ELs and ILs with the same colour (orange, yellow or grey) are also equivalent. The centre panel in a shows the entire EcEmrD molecule with the N-terminal half (TMS1-6) in green, the C-terminal half (TMS7-12) in pale blue, the IL3 linker between TMD1 and TMD2 in black and the conserved IL1 and IL4 highlighted in yellow. b and c Nucleotide-free conformations of type I and type II ABC exporters. b Type I Mus musculus multidrug efflux pump MmABCB1 (pdb: 3G5U; (Aller et al. 2009)); and c Type II Homo sapiens cholesterol ABC exporter HsABCG5-G8 (pdb: 5DO7; (Lee et al. 2016)). MmABCB1 is a single polypeptide, whereas HsABCG5-G8 is a heterodimer of two separate polypeptide chains. Their N- and C-terminal TMDs are green and pale blue and their N- and C-terminal NBDs magenta and red, respectively. The centre panels in b and c highlight the major contact points between the TMDs (ILs and coupling helices (Ch) in yellow) and the Q-loops (dark blue; conserved Qs are shown as red sticks) of the NBDs for type I (b) and type II (c) ABC exporters. The four ILs of type I exporters provide ‘asymmetric’ contact with the two NBD Q-loops (centre panel), whereas IL1 and a conserved positively charged cytosolic Ch preceding TMS1 provide ‘symmetric’ contact with their respective NBDs; i.e. the Ch and IL1 of ABCG5 and ABCG8 are in contact with the Q-loop and a conserved E-helix (magenta; centre panel) of their own NBDs (centre right). Visible TMS helices are numbered in yellow (N-terminal TMD) and orange (C-terminal TMD). The bottom panels show cross-sections through the centres of the TMDs for MFS (a), type I (b) and type II (c) ABC transporters viewed from the extracellular side after tilting the TMDs depicted in the centre panels forward by 90˚. The N- and C-terminal TMDs are rainbow colour-coded and numbered in green (N-terminal TMD) or blue (C-terminal TMD), respectively. Each transporter has a unique TMS topology with unique arrangements of TMSs surrounding a centrally located substrate-binding pocket
18.4.2 ABC Transporters
ABC transporters are divided into three types of importers (type I, type II and energy-coupling factor (ECF) importers; Fig. 18.3), only found in prokaryotes, and two types of exporters (type I and type II exporters; Fig. 18.2b, c), found in all kingdoms of life. While the NBDs have generally well-conserved structures, their TMD folds vary significantly, as demonstrated in the centre and bottom panels in Figs. 18.2b, c and 18.3—possibly a reflection of their large substrate diversity (Jones et al. 2009; Kerr et al. 2010; Locher 2004; Rees et al. 2009). Many ABC transporters possess additional domains responsible for regulating transporter activity, necessary for protein–protein interactions or important for protein targeting to the correct membrane. Type I and type II ABC importers, for example, require an additional high-affinity substrate-binding protein component (yellow domains in Fig. 18.3a, b) that delivers a specific ligand to the transporter, while others have N-terminal TMD extensions that cause transporter dimerization and/or help target the transporter to its correct destination.
Structures of type I, type II and ECF-type ABC importers. Nucleotide-free conformations of three ABC importers with the same annotations used in Fig. 18.2b, c. a Type I Archaeoglobus fulgidus molybdate ABC importer ModAB2C2 (pdb: 2ONK; (Hollenstein et al. 2007a, b)). b Type II E. coli vitamin B12 ABC importer BtuC2D2F (pdb: 2QI9; (Hvorup et al. 2007)). c Lactobacillus brevis energy-coupling factor (EFC) ABC importer Ecf-folate transporter (pdb: 4HUQ; (Xu et al. 2013)). ModAB2C2 and BtuC2D2F each consist of five separate domains that are encoded by three separate genes, the periplasmic molybdate and vitamin B12 binding proteins ModA and BtuF (yellow), the N- (green) and C-terminal (pale blue) TMDs ModB and BtuC and the N- (magenta) and C-terminal (red) NBDs ModC and BtuD, respectively. In contrast, the L. brevis Ecf-folate transporter consists of four domains, each encoded by a separate gene: EcfA (magenta) and EcfA’ (red) are the N- and C-terminal NBDs, and EcfT (green) and the folate binding protein EcfS (pale blue) are the N- and C-terminal TMDs. The centre panels highlight the major contact points of the TMDs with the Q-loops of the NBDs. ModB-IL2 (L2 and L2’; left) and BtuC-IL3 (L3 and L3’; centre) provide the major contact points with the Q-loops of ModC and BtuD, respectively, ECF importers like the L. brevis Ecf-folate transporter, however, interact with the Q-loops of EcfA and EcfA’ via three intracellular coupling helices (CH1, CH2 and CH3) that are part of the large EcfT-IL2 (right). The bottom panels highlight the unique TMD folds of these three different types of ABC importer. Note how EcfS is toppled over towards EcfT—this conformation reflects the structure of the Ecf-folate transporter after substrate release
Due to the lack of any type II exporter structure, S. cerevisiae Pdr5 was initially modelled on the structures of type I exporters (Rutledge et al. 2011). However, the first type II ABC exporter structure for the open conformation of the human cholesterol transporter HsABCG5-G8 (Fig. 18.2c) has recently been solved (Lee et al. 2016). Perhaps not surprisingly, given the unique features characterising PDR transporters (Lamping et al. 2010), HsABCG5-G8 exhibited a unique TMD fold (bottom of Fig. 18.2c), quite different from that of type I exporter MmABCB1 (bottom of Fig. 18.2b). Lee and colleagues noticed a unique TMD-NBD cis-interface with novel connecting-helices (Ch; yellow) and E-helices (magenta) just before TMS1 and after the Q-loop of ABCG5 and ABCG8, respectively (centre panel; Fig. 18.2c). Conserved salt-bridges stabilised cis contacts between the TMDs (Ch and IL1) and NBDs (Q-loop and E-helix) of HsABCG5-G8 (centre panel in Fig. 18.2c) (Lee et al. 2016). We previously reported PDR transporter-specific positively charged helices just before TMS1 and TMS7 (Lamping et al. 2010) that align with the two Chs of ABCG5 and ABCG8 (data not shown), and we can confirm that all fungal PDR transporters also have the conserved E-helices adjacent to their respective Q-loops (Fig. 18.4b) suggesting that Pdr5 and Cdr1 have the same structural/functional constraints as HsABCG5-G8. Also of significant interest was how the unique extracellular domains of HsABCG5-G8 fold. The previously identified PDR transporter EL motifs (Lamping et al. 2010) are visualised in ABCG5 (PDRA = 557I-F565; PDRB = 569K-F580) and ABCG8 (EL6 motif = 584W-S591; EL6 helix = 595F-S609) as four separate helices (yellow; top panel of Fig. 18.2c). Both helices of each monomer bend towards each other and form a critical part of the TMD by dipping into the central binding cavity of the transporter (top panel; Fig. 18.2c).
Summary of Cdr1 mutagenesis studies. a Cdr1 TMD residues that have been shown to be important for Cdr1 efflux pump function (Kolaczkowski et al. 2013; Rawal et al. 2013; Shah et al. 2015) are identified. TMSs are shown as light grey boxes and residues that selectively affected transport of some, but not all, Cdr1 test substrates are shown as grey circles, and residues with severe functional defects (i.e. no drug efflux and reduced ATPase activity) are shown as black circles. A list of these residues is shown underneath the corresponding TMS or next to the corresponding IL. Residues G521 (centre of TMS1) and A1208 (centres of TMS7) that, when mutated, improved efflux of small compounds such as resazurin, an Snq2 substrate (Kolaczkowski et al. 2013), are shown as white circles and the six conserved EL3 and EL6 cysteines (Lamping et al. 2010) are indicated as yellow circles. b A graphical representation of the conserved Cdr1/Pdr5 NBD motifs highlighted in different colours with a list of the Walker A, Walker B, Q-loop, E-helix, D-loop, H-loop and ABC signature motif residues for Cdr1 and Pdr5 shown underneath. Red residues are critical for ATP-binding and/or hydrolysis as discussed in the text. * = non-canonical NBD motifs typical for PDR transporters
Most of the current knowledge about the transport and energy coupling of ABC exporters relates to studies with type I ABC exporters. S. aureus Sav1866, homolog of one of the best-studied multidrug efflux pumps, human Pgp, was one of the first type I ABC exporters for which the structure was solved (Dawson and Locher 2006; Dawson and Locher 2007). The structures obtained were for the closed conformation with the substrate-binding cavity accessible to the extracellular space. Most other type I ABC exporter structures solved since then have been for open conformations with the substrate-binding cavity open to the cytosol (Fig. 18.2b). These structures revealed important insights into how type I ABC exporters couple ATP-binding and hydrolysis with substrate transport. Typically, the first IL of each TMD (IL1 and IL3) is in close contact with its proximal NBD Q-loop (Q-loop1 and Q-loop2, respectively; Fig. 18.2b centre), whereas the second loop (IL2 and IL4) reaches across the opposite NBD by contacting Q-loop2 and Q-loop1, respectively (centre panel of Fig. 18.2b). Computer simulations of the transport cycle of type I exporters suggested large, rigid, conformational changes of the NBDs upon ATP-binding that move the two NBDs close together and the cylindrical, barrel-like, structure formed by the two TMDs, especially TMS4, -6, -10 and -11 that surround a central substrate-binding cavity (bottom panel of Fig. 18.2b), becomes exposed to the outer leaflet and the extracellular space (Aller et al. 2009; Dawson and Locher 2006; Jin et al. 2012; Li et al. 2014; Prajapati and Sangamwar 2014; Ravna et al. 2009; Rosenberg et al. 2003; Velamakanni et al. 2008; Ward et al. 2007; Ward et al. 2013). This twisting motion of the TMDs induced by movement of the hinge-like contacts between IL1-4 with the Q-loops of NBD1 and NBD2 (Fig. 18.2b) causes helix rotation (Helix Rotation Model) (Gutmann et al. 2010) of TMSs surrounding the substrate-binding cavity and thus changes the affinity of substrates so that they can be released to the cell exterior after transporter closure. Like efflux pump substrates, inhibitors also bind to the substrate-binding cavity (Aller et al. 2009) but, unlike Pgp substrates that can induce the ATPase activity of the efflux pump up to 100-fold by helping bring the two NBDs closer together and thus increase the affinity for ATP and induce their own transport—also referred to as the Induced Fit Model—inhibitors seem to keep the NBDs apart and freeze the transporter in an open conformation (Aller et al. 2009; Gutmann et al. 2010; Ma and Biggin 2013). ATP hydrolysis is believed to then reset the transporter in the inward facing conformation so that another transport cycle can begin. The Switch Model proposed that after ATP hydrolysis the two NBDs separate by >20 Å (Higgins and Linton 2004), while others suggested that the large gap separating the NBDs in the open conformation of most type I ABC exporter structures is not a biologically viable state. Some studies have shown that the NBDs indeed remain in contact at all times during an alternating ATP-binding and hydrolysis cycle (George and Jones 2012; Jones and George 2009; Sauna et al. 2007). In the Constant Contact Model, the ATP-bound active site is closed while the second ATP-binding site is empty and slightly further apart (George and Jones 2012). After transport, ATP hydrolysis and the release of Pi and ADP, the empty low-affinity site switches to a high-affinity site and another ATP hydrolysis cycle begins (Fribourg et al. 2014; George and Jones 2012; Ravna et al. 2009).
However, despite all these valuable contributions, details of the ATP kinetics of type I ABC exporters remain unresolved. Electron microscopy single particle analysis of E. coli MsbA and Pgp revealed remarkable differences between the two pumps, not only in their conformational changes in the presence of nucleotides and substrates, but also in the separation between their two NBDs. Therefore, perhaps more than one model is required to describe type I ABC exporter function (Moeller et al. 2015).
18.5 Structure–Function Activity Relationships Deduced from C. albicans Multidrug Transporter Mutagenesis
18.5.1 MFS Transporter Mdr1
Structural and functional analyses of C. albicans Mdr1 have indicated that TMS5 and IL3 residues are critical for drug transport. Alanine substitutions of 21 TMS5 residues showed clustering of functionally critical residues, including all conserved C-motif residues (G251, G255, P256, G259), to one face of TMS5 (Pasrija et al. 2007). The functional importance of IL3 residues was also established by site-directed mutagenesis (SDM) (Mandal et al. 2012). This region contains a number of charged residues, half of which were critical for Mdr1 function (Mandal et al. 2012). Yan proposed a model whereby IL3 helices provide a latch that strengthens the intracellular gate in the outward open conformation of GLUT1 and related SPs (Yan 2015). C. albicans Mdr1 also has remnants of the ‘hallmark’ A-motif (174-SPMSENAIFGRT-185; Paulsen et al. 1996) with three of the six conserved residues (i.e. E178, G183 and R184) found between TMS2 and -3, and, like most other MFS transporters, it has a conserved D235 at the end of TMS4 and a conserved 296-P-E-(T/S) motif at the cytosolic end of TMS6 (Nomura et al. 2015; Paulsen et al. 1996). Based on existing structures (Nomura et al. 2015; Quistgaard et al. 2016; Yan 2013; Yan 2015), Mdr1 R184 is likely to form important salt-bridges with D235 and E297 at different stages of the transport cycle. The critical contribution of E297 to Mdr1 drug transport was confirmed by Mandal and colleagues (Mandal et al. 2012).
An important question is why does Mdr1 have a wider substrate specificity than most other MFS pump homologues? MFS proteins often have conserved gating residues lining the central binding cavity, which are responsible for their substrate specificity. For example, a single-point mutation Q166E in TMS5 of the substrate-binding cavity of GLUT5 was enough to switch the substrate-binding preference from fructose to glucose (Nomura et al. 2015). Similarly, the E. coli multidrug efflux pump EmrD has two long helical regions composed of TMS4, -5, -10, -11 and slightly extended IL2 and IL4 that, together with a run of three positively charged residues at the end of TMS4, are proposed to act as a substrate-specificity filter (Yin et al. 2006). Another example of the importance of TMS1, -4, -7, -10 residues (blue helices on Fig. 18.2a) and, in particular, residues of the proton-dependent oligopeptide transporter (POT) family-defining motif Ex2ERFxYY of TMS1, in substrate selectivity, proton binding and oligopeptide symport was revealed by the structural analysis of the Streptococcus thermophilus POT oligopeptide symporter PepTSt (Solcan et al. 2012). These structural and mechanistic principles appear to hold true for most, if not all, MFS transporters, and, in the absence of an Mdr1 structure, they provide a model for how C. albicans multidrug efflux pump Mdr1 works.
18.5.2 ABC Transporter Cdr1
18.5.2.1 TMDs and ELs
Early random mutagenesis studies with S. cerevisiae Pdr5 identified several residues that were critical for drug transport. It was not clear, however, how these residues, which were distributed throughout the molecule, caused reduced efflux of some or all drugs and/or affected the ATPase activity of Pdr5 (Egner et al. 2000; Egner et al. 1998; Tutulan-Cunita et al. 2005). Both MDR and PDR ABC exporters appear to have multiple overlapping substrate-binding sites encompassing a large substrate-binding cavity within their TMDs, with Pdr5 and Cdr1 possessing at least three separate substrate-binding sites (Golin et al. 2003; Golin et al. 2007; Puri et al. 2009; Shukla et al. 2003). There are, however, differences in the preferred substrates of ABC proteins. Pgp for instance prefers hydrophobic substrates with hydrogen-bond acceptor groups (Seelig and Landwojtowicz 2000), whereas size (optimum ~ 200–225 Å3), but not hydrophobicity, seemed the major determinant for a good Pdr5 substrate (Golin et al. 2003; Golin et al. 2007). Cdr1, on the other hand, seemed to prefer hydrophobic substrates and molecular-branching (Puri et al. 2010).
In the absence of a high-resolution structure, bioinformatics and biochemistry are the best ways to identify residues involved in drug binding and/or transport. Heterologous expression of efflux pumps in S. cerevisiae provides many advantages for studying structure–function relationships. S. cerevisiae can be easily genetically manipulated and a vast mutant collection is available. Cdr1/Pdr5 variants can be rapidly generated and expressed at high levels at the correct cellular location (Lamping and Cannon 2010; Niimi et al. 2012; Tanabe et al. 2011). In addition, one can quickly select for naturally occurring suppressor mutants of transport defective variants by isolating resistant colonies that appear after prolonged incubation at 30 °C on plates containing high (Niimi et al. 2012) concentrations of toxic efflux pump substrates (Golin and Ambudkar 2015). SDM of all 252 Cdr1 TMS residues indicated that almost half of them (~42%) affected substrate transport and/or ATPase activity (Fig. 18.4a) (Rawal et al. 2013), making interpretation of the effects of mutations of individual residues very challenging. Interestingly, most residues affecting substrate transport (grey circles; Fig. 18.4a) and/or ATPase activity (black circles; Fig. 18.4a) were located in TMS1, -2, -5, -8 and -11, whereas TMS4, -6, -10 and -12 contributed least to substrate transport and/or ATPase activity (Fig. 18.4a) (Rawal et al. 2013). Given this result, it was perhaps a little surprising that only ~ 18% of all IL mutants affected Cdr1 drug transport and none of them affected the Cdr1 ATPase activity (Fig. 18.4a) (Shah et al. 2015). The observed phenotypes of TMS5 mutant residues affecting Cdr1 pump function did not relate to which face of the helix they belonged to, the degree of conservation of the residues or the nature of their side chains (Puri et al. 2009; Rawal et al. 2013). In contrast, TMS11 mutations of both Cdr1 and Pdr5 that affected drug efflux were mainly clustered on the hydrophilic face of the helix, suggesting a direct contribution of TMS11 to a centrally located drug-binding cavity (Egner et al. 2000; Kueppers et al. 2013; Rawal et al. 2013; Saini et al. 2005; Shukla et al. 2004).
The role of EL residues in protein assembly, cell surface localisation and substrate specificity has been highlighted in a random mutagenesis study of Pdr5 (Egner et al. 1998). A single substitution of one of the six conserved EL3/EL6 cysteines (yellow residues in Fig. 18.4a) of fungal PDR transporters (Lamping et al. 2010), C1427Y, caused mislocalization of the mutant Pdr5 to the endoplasmic reticulum. The equivalent residue in Cdr1, C1418, was found to be essential for pump function (Shukla et al. 2003). These studies highlighted the importance of the conserved extracellular cysteines that possibly form three extracellular disulfide bonds that stabilise the structure of the highly conserved extracellular domains of PDR transporters (Lamping et al. 2010). Molecular mapping and biochemical characterization of more than 50 Pdr5/Cdr1 suppressor mutants that had lost their ability to respond to efflux pump inhibitors RC21v3, a Cdr1 specific D-octapeptide inhibitor (Niimi et al. 2012), macrolides such as FK506 (Egner et al. 1998; Kralli and Yamamoto 1996; Lamping et al. 2007; Tanabe et al. 2011) and milbemycins (Lamping et al. 2007; Lee et al. 2001; Niimi et al. 2012; Silva et al. 2013), depsipeptides enniatin (Hiraga et al. 2005; Holmes et al. 2008; Lamping et al. 2007; Lee et al. 2001; Tanabe et al. 2011) and beauvericin (Lee et al. 2001; Tanabe et al. 2011) further highlighted the importance of the conserved EL3 and EL6 in Cdr1/Pdr5 efflux pump function ((Lamping et al. 2014) and unpublished results). How these extracellular domains contribute to type II exporter function is an important question that needs to be answered in order to understand how PDR multidrug efflux pumps work.
18.5.2.2 NBDs and ILs
The highly conserved NBDs are defined by the Walker A and the Walker B motifs (Walker et al. 1982) and the hallmark ABC signature motif which are critical for ATP-binding and hydrolysis (Fig. 18.4b; (Prasad and Goffeau 2012)). Typically, ABC transporters have two canonical ATP-binding sites, also referred to as composite nucleotide-binding domains (CNBD), comprising Walker A and Walker B motifs of one NBD and the ABC signature motif of the other. CNBD1 and CNBD2 of most ABC proteins are believed to alternatingly bind and hydrolyse ATP (see Sect. 18.4.2). However, PDR transporters have a high, non-inducible, basal ATPase activity (Ernst et al. 2008) and they are characterised by one canonical CNBD2 (i.e. typical Walker A2, Walker B2 and ABC1 signature motifs; Fig. 18.4b) and one non-canonical CNBD1 with its Walker A1 K residue replaced by a C (C193 in Cdr1), its Walker B1 catalytic E replaced by an N (N328 in Cdr1), the conserved H-loop H of NBD1 replaced by a Y (Y361 in Cdr1) and the C-loop G of ABC2 replaced by a E (E1004 in Cdr1; red residues in Fig. 18.4b).
SDM of C199 of Walker A1 and K911 of Walker A2 (Fig. 18.4b) indicated that the Walker A2 K911, but not the Walker A1 C199, was critical for Pdr5 ATPase activity and drug transport (Ernst et al. 2008). SDM of Cdr1 Walker B1 (D327, N328), Walker B2 (D1026, E1027) as well as all five ABC1 (VSGGE) and ABC2 (LNVEQ; Fig. 18.4b) residues found that all four Walker B residues (Rai et al. 2006) and ABC residues S304, G306 and E307 of ABC1 and the equivalent ABC2 residues N1002 and E1004, but not Q1005 (Kumar et al. 2010), were critical for both the ATPase activity as well as drug efflux, indicating the importance of both CNBDs in ATP-binding and/or hydrolysis (Kumar et al. 2010). In contrast, alanine substitution of the Pdr5 D-loop residues D340A and D1042A (red residues; Fig. 18.4b) ‘uncoupled’ the ATPase activity from drug transport by affecting drug efflux but not the ATPase activity of Pdr5, indicating that D-loop residues are important for the crosstalk between the NBDs and the TMDs of PDR transporters (Furman et al. 2013). Surprisingly, mutation of the conserved H-loop H1068A had no effect on Pdr5 ATPase activity, instead it selectively abolished R6G transport without affecting the transport of any other substrates (Ernst et al. 2008). This led to the Kinetic Substrate Selection model for Pdr5 transport, whereby the authors proposed that the H1068A mutation changed the transport kinetics in such a way that only the transport of ‘slow’ substrates such as R6G was impaired because the average time it took ‘slow’ substrates to enter or diffuse into the transporter far exceeded the time the mutant transporter was open for substrates to enter (Ernst et al. 2010).
A study of the Pdr5 S558Y (Cdr1 A548; Fig. 18.4a) mutation at the top of TMS2 provided important clues about how TMS2 may be critical for the crosstalk between the NBDs and the TMDs (Sauna et al. 2008). The phenotype of yeast with this mutation was drug hypersensitivity despite wild-type ATPase activity or, in other words, this mutant had a pump defect because its ATPase activity could not be translated into the conformational changes needed for drug efflux. This defect could be recovered by second-site suppressor mutations in Q-loop1 (N242 K and E244G; Fig. 18.4b) and the centre of IL1 (S597I, S597T (Cdr1 S587)) (Ananthaswamy et al. 2010). Similarly, second-site Pdr5 N242 K suppressor mutants with improved CHX transport were found in the IL2-TMS5 (M649I, V656L, A666G) region and just C-terminal of ABC2 (K1016I) (Ananthaswamy et al. 2010). The same V656L second-site suppressor mutation was also found in a search for improved CHX transport (i.e. increased CHX resistance) of yeast over-expressing wild-type Pdr5 (Downes et al. 2013). Alanine substitution of Pdr5 V656, however, caused the Pdr5 ATPase activity to become uncoupled from drug efflux and led to drug hypersensitivity of the strain over-expressing the mutant protein. Taken together, these data support a model whereby Pdr5 IL2 may possibly be part of a cis-transmission interface between TMD1 and NBD1, unlike MmABCB1 where IL2 is a critical part of a trans-transmission interface by being in direct contact with Q-loop2 of NBD2 (Fig. 18.2b). The critical importance of the TMS2-IL1 region for the crosstalk between the NBDs and TMDs of PDR transporters was further supported by the Cdr1 SDM studies described in Sect. 18.5.2.1 with 7 of 24 functionally critical Cdr1 TMD residues (black residues; Fig. 18.4a) located in TMS2 and almost all remaining TMS2 residues affecting efflux, and more than 50% of all IL residues affecting Cdr1 drug efflux were located in IL1 (Fig. 18.4a) (Rawal et al. 2013; Shah et al. 2015).
Taken together, these studies suggest a critical role for the deviant CNBD1 in intra-domain signalling of ATP-binding at the canonical CNBD2 (Ananthaswamy et al. 2010; Furman et al. 2013) which causes the NBDs to move closer together and, in turn, leads to large conformational changes at the TMDs followed by drug release in the closed conformation of the transporter. In addition, TMS2-IL1, and possibly IL2, seem to play a critical role in the cis-transmission crosstalk between the NBDs and the TMDs. Despite all these findings, however, it is still not clear whether CNBP1 can also bind and/or hydrolyze ATP, or whether it has evolved an alternative ‘regulatory’ function and is only able to bind—possibly at all times—but not hydrolyze ATP (Furman et al. 2013; Gupta et al. 2011; Prasad and Goffeau 2012).
18.6 Model of Fungal PDR Transporters
It is evident that there has been a plethora of studies dissecting Cdr1/Pdr5 structure and function. The vast amount of biochemical data accumulated over the past two decades combined with PDR transporter phylogeny and the first type II ABC exporter structure, HsABCG5-G8, provide evidence for the following tentative model of how PDR transporters function. Cdr1/Pdr5 TMS1, -2, -5, -7, 8, -11 (yellow to red; Fig. 18.5c) form a large central substrate-binding cavity with TMS3, -4, -6, -9, -10, -12 (green to blue; Fig. 18.5c) stabilising the protein during the transport cycle. Salt-bridges help stabilise critical contacts between the positively charged Ch near TMS1 and IL1, Q-loop1 and E-helix1 of NBD1. The homologous parts of the C-terminal half provide similar cis interactions between NBD2 and TMD2. CNBD1 is possibly bound to ATP at all times while CNBD2 provides the energy required for the large conformational changes associated with drug efflux by binding and hydrolysing ATP. Openings between TMS1 and TMS11 (gate 1; Fig. 18.5c) and TMS5 and TMS7 (gate 2; Fig. 18.5c) may provide gates for substrates to enter, in contrast to type I exporters where substrates are thought to enter between TMS4 and TMS6 and TMS10 and TMS12 (Fig. 18.5a). The extracellular domain also plays a critical part in this process, as evidenced by phylogeny and the HsABCG5-G8 structure. How these extracellular domains contribute to drug efflux, however, remains to be explored.
Fungal PDR transporters are predicted to fold like HsABCG5-G8. The TMD folds for MmABCB1 (a) and HsABCG5-G8 (b) are the same as shown in Fig. 18.2b, c. This figure shows how MDR transporters (a) have TMS4, -5, -6 and TMS10, -11, -12 lining the substrate-binding pocket, whereas the PDR transporter homolog HsABCG5-G8 (b) has TMS1, -2, and -5 of ABCG5 (left half) and ABCG8 (right half) lining the substrate-binding pocket. Annotations are the same as in Fig. 18.2 c. The predicted TMS topology for Cdr1 and Pdr5 (Lamping et al. 2014) (c). This topology model was created by careful consideration of all published and unpublished data relating to the structure and function of Cdr1 and Pdr5. The colours of TMSs: red, orange, yellow, green, turquois and blue indicate homologous N- (lighter colours) and C-terminal (darker colours) TMS pairs from the most to the least ‘important’ (i.e. the most important TMSs had the most number of residues affecting Cdr1/Pdr5 ATPase activity and/or drug efflux; also see Fig. 18.4a). Proposed (a) and possible gates (b; c) where pump substrates may enter the transporter from the inner leaflet of the lipid bilayer are indicated with black (a) and grey (b; c) arrows. Note how the predicted PDR TMS topology closely resembles that of ABCG5-G8 (b) but has little semblance to type I exporters (a)
18.7 Concluding Remarks
C. albicans possesses many membrane transport proteins which perform vital cellular functions. ABC and, to a lesser extent, MFS transporters have broad substrate specificities that can include antifungal drugs and xenobiotics. Increased expression of pumps, particularly Cdr1, is selected for by growth in the presence of drugs and confers drug resistance. Because of their promiscuity, the normal biological function of multidrug efflux pumps is an unanswered question. In vitro and in vivo studies have demonstrated that inhibition of Cdr1 can sensitise azole-resistant C. albicans clinical isolates to FLC and ITC. Further advances in the determination of the 3D structures of multidrug transporters will help design more potent specific and broad spectrum pump inhibitors and may also elucidate the reaction cycle for an important class of membrane transporters. Molecular modelling and site-directed mutagenesis is beginning to reveal that the arrangement of external loops, transmembrane helices and interactions between these helices and the NBDs are important for pump function and are potential drug targets.
References
Aller SG, Yu J, Ward A, Weng Y, Chittaboina S, Zhuo R, Harrell PM, Trinh YT, Zhang Q, Urbatsch IL, Chang G (2009) Structure of P-glycoprotein reveals a molecular basis for poly-specific drug binding. Science 323:1718–1722
Ananthaswamy N, Rutledge R, Sauna ZE, Ambudkar SV, Dine E, Nelson E, Xia D, Golin J (2010) The signaling interface of the yeast multidrug transporter Pdr5 adopts a cis conformation, and there are functional overlap and equivalence of the deviant and canonical Q-loop residues. Biochemistry 49:4440–4449
Braun BR, van Het Hoog M, d’Enfert C, Martchenko M, Dungan J, Kuo A, Inglis DO, Uhl MA, Hogues H, Berriman M, Lorenz M, Levitin A, Oberholzer U, Bachewich C, Harcus D, Marcil A, Dignard D, Iouk T, Zito R, Frangeul L, Tekaia F, Rutherford K, Wang E, Munro CA, Bates S, Gow NA, Hoyer LL, Kohler G, Morschhauser J, Newport G, Znaidi S, Raymond M, Turcotte B, Sherlock G, Costanzo M, Ihmels J, Berman J, Sanglard D, Agabian N, Mitchell AP, Johnson AD, Whiteway M, Nantel A (2005) A human-curated annotation of the Candida albicans genome. PLoS Genet 1:36–57
Calabrese D, Bille J, Sanglard D (2000) A novel multidrug efflux transporter gene of the major facilitator superfamily from Candida albicans (FLU1) conferring resistance to fluconazole. Microbiology 146(Pt 11):2743–2754
Cannon RD, Lamping E, Holmes AR, Niimi K, Baret PV, Keniya MV, Tanabe K, Niimi M, Goffeau A, Monk BC (2009) Efflux-mediated antifungal drug resistance. Clin Microbiol Rev 22:291–321
Cheng Y, Grigorieff N, Penczek PA, Walz T (2015) A primer to single-particle cryo-electron microscopy. Cell 161:438–449
Coleman JJ, Mylonakis E (2009) Efflux in fungi: la piece de resistance. PLoS Pathog 5:e1000486
Costa C, Dias PJ, Sa-Correia I, Teixeira MC (2014) MFS multidrug transporters in pathogenic fungi: do they have real clinical impact? Front Physiol 5:197
Coste A, Selmecki A, Forche A, Diogo D, Bougnoux ME, d’Enfert C, Berman J, Sanglard D (2007) Genotypic evolution of azole resistance mechanisms in sequential Candida albicans isolates. Eukaryot Cell 6:1889–1904
Coste AT, Karababa M, Ischer F, Bille J, Sanglard D (2004) TAC1, transcriptional activator of CDR genes, is a new transcription factor involved in the regulation of Candida albicans ABC transporters CDR1 and CDR2. Eukaryot Cell 3:1639–1652
Crouzet J, Trombik T, Fraysse AS, Boutry M (2006) Organization and function of the plant pleiotropic drug resistance ABC transporter family. FEBS Lett 580:1123–1130
Dawson RJ, Locher KP (2006) Structure of a bacterial multidrug ABC transporter. Nature 443:180–185
Dawson RJ, Locher KP (2007) Structure of the multidrug ABC transporter Sav 1866 from Staphylococcus aureus in complex with AMP-PNP. FEBS Lett 581:935–938
de Micheli M, Bille J, Schueller C, Sanglard D (2002) A common drug-responsive element mediates the upregulation of the Candida albicans ABC transporters CDR1 and CDR2, two genes involved in antifungal drug resistance. Mol Microbiol 43:1197–1214
Delmar JA, Bolla JR, Su CC, Yu EW (2015) Crystallization of membrane proteins by vapor diffusion. Methods Enzymol 557:363–392
Downes MT, Mehla J, Ananthaswamy N, Wakschlag A, Lamonde M, Dine E, Ambudkar SV, Golin J (2013) The transmission interface of the Saccharomyces cerevisiae multidrug transporter Pdr5: Val-656 located in intracellular loop 2 plays a major role in drug resistance. Antimicrob Agents Chemother 57:1025–1034
Dunkel N, Blass J, Rogers PD, Morschhauser J (2008) Mutations in the multi-drug resistance regulator MRR1, followed by loss of heterozygosity, are the main cause of MDR1 overexpression in fluconazole-resistant Candida albicans strains. Mol Microbiol 69:827–840
Egner R, Bauer BE, Kuchler K (2000) The transmembrane domain 10 of the yeast Pdr5p ABC antifungal efflux pump determines both substrate specificity and inhibitor susceptibility. Mol Microbiol 35:1255–1263
Egner R, Rosenthal FE, Kralli A, Sanglard D, Kuchler K (1998) Genetic separation of FK506 susceptibility and drug transport in the yeast Pdr5 ATP-binding cassette multidrug resistance transporter. Mol Biol Cell 9:523–543
Ernst R, Kueppers P, Klein CM, Schwarzmueller T, Kuchler K, Schmitt L (2008) A mutation of the H-loop selectively affects rhodamine transport by the yeast multidrug ABC transporter Pdr5. Proc Natl Acad Sci U S A 105:5069–5074
Ernst R, Kueppers P, Stindt J, Kuchler K, Schmitt L (2010) Multidrug efflux pumps: substrate selection in ATP-binding cassette multidrug efflux pumps–first come, first served? FEBS J 277:540–549
Fling ME, Kopf J, Tamarkin A, Gorman JA, Smith HA, Koltin Y (1991) Analysis of a Candida albicans gene that encodes a novel mechanism for resistance to benomyl and methotrexate. Mol Gen Genet 227:318–329
Ford RC, Kamis AB, Kerr ID, Callaghan R (2009) The ABC transporters: structural insights into drug transport. Transporters as Drug Carriers: Struct Funct Substrates 44:3
Fribourg PF, Chami M, Sorzano CO, Gubellini F, Marabini R, Marco S, Jault JM, Levy D (2014) 3D cryo-electron reconstruction of BmrA, a bacterial multidrug ABC transporter in an inward-facing conformation and in a lipidic environment. J Mol Biol 426:2059–2069
Furman C, Mehla J, Ananthaswamy N, Arya N, Kulesh B, Kovach I, Ambudkar SV, Golin J (2013) The deviant ATP-binding site of the multidrug efflux pump Pdr5 plays an active role in the transport cycle. J Biol Chem 288:30420–30431
Gaur M, Choudhury D, Prasad R (2005) Complete inventory of ABC proteins in human pathogenic yeast, Candida albicans. J Mol Microbiol Biotechnol 9:3–15
Gaur M, Puri N, Manoharlal R, Rai V, Mukhopadhayay G, Choudhury D, Prasad R (2008) MFS transportome of the human pathogenic yeast Candida albicans. BMC Genom 9:579
George AM, Jones PM (2012) Perspectives on the structure-function of ABC transporters: the switch and constant contact models. Prog Biophys Mol Biol 109:95–107
Golin J, Ambudkar SV (2015) The multidrug transporter Pdr5 on the 25th anniversary of its discovery: an important model for the study of asymmetric ABC transporters. Biochem J 467:353–363
Golin J, Ambudkar SV, Gottesman MM, Habib AD, Sczepanski J, Ziccardi W, May L (2003) Studies with novel Pdr5p substrates demonstrate a strong size dependence for xenobiotic efflux. J Biol Chem 278:5963–5969
Golin J, Ambudkar SV, May L (2007) The yeast Pdr5p multidrug transporter: how does it recognize so many substrates? Biochem Biophys Res Commun 356:1–5
Gupta RP, Kueppers P, Schmitt L, Ernst R (2011) The multidrug transporter Pdr5: a molecular diode? Biol Chem 392:53–60
Gutmann DA, Ward A, Urbatsch IL, Chang G, van Veen HW (2010) Understanding polyspecificity of multidrug ABC transporters: closing in on the gaps in ABCB1. Trends Biochem Sci 35:36–42
Henderson R (2015) Overview and future of single particle electron cryomicroscopy. Arch Biochem Biophys 581:19–24
Higgins CF, Linton KJ (2004) The ATP switch model for ABC transporters. Nat Struct Mol Biol 11:918–926
Hiraga K, Yamamoto S, Fukuda H, Hamanaka N, Oda K (2005) Enniatin has a new function as an inhibitor of Pdr5p, one of the ABC transporters in Saccharomyces cerevisiae. Biochem Biophys Res Commun 328:1119–1125
Hollenstein K, Dawson RJ, Locher KP (2007) Structure and mechanism of ABC transporter proteins. Curr Opin Struct Biol 17:412–418
Hollenstein K, Frei DC, Locher KP (2007) Structure of an ABC transporter in complex with its binding protein. Nature 446:213–216
Holmes AR, Lin YH, Niimi K, Lamping E, Keniya M, Niimi M, Tanabe K, Monk BC, Cannon RD (2008) ABC transporter Cdr1p contributes more than Cdr2p does to fluconazole efflux in fluconazole-resistant Candida albicans clinical isolates. Antimicrob Agents Chemother 52:3851–3862
Hvorup RN, Goetz BA, Niederer M, Hollenstein K, Perozo E, Locher KP (2007) Asymmetry in the structure of the ABC transporter-binding protein complex BtuCD-BtuF. Science 317:1387–1390
Janbon G, Ormerod KL, Paulet D, Byrnes EJ 3rd, Yadav V, Chatterjee G, Mullapudi N, Hon CC, Billmyre RB, Brunel F, Bahn YS, Chen W, Chen Y, Chow EW, Coppee JY, Floyd-Averette A, Gaillardin C, Gerik KJ, Goldberg J, Gonzalez-Hilarion S, Gujja S, Hamlin JL, Hsueh YP, Ianiri G, Jones S, Kodira CD, Kozubowski L, Lam W, Marra M, Mesner LD, Mieczkowski PA, Moyrand F, Nielsen K, Proux C, Rossignol T, Schein JE, Sun S, Wollschlaeger C, Wood IA, Zeng Q, Neuveglise C, Newlon CS, Perfect JR, Lodge JK, Idnurm A, Stajich JE, Kronstad JW, Sanyal K, Heitman J, Fraser JA, Cuomo CA, Dietrich FS (2014) Analysis of the genome and transcriptome of Cryptococcus neoformans var. grubii reveals complex RNA expression and microevolution leading to virulence attenuation. PLoS Genet 10:e1004261
Jin MS, Oldham ML, Zhang Q, Chen J (2012) Crystal structure of the multidrug transporter P-glycoprotein from Caenorhabditis elegans. Nature 490:566–569
Jones PM, George AM (2009) Opening of the ADP-bound active site in the ABC transporter ATPase dimer: evidence for a constant contact, alternating sites model for the catalytic cycle. Proteins 75:387–396
Jones PM, O’Mara ML, George AM (2009) ABC transporters: a riddle wrapped in a mystery inside an enigma. Trends Biochem Sci 34:520–531
Kerr ID, Jones PM, George AM (2010) Multidrug efflux pumps: the structures of prokaryotic ATP-binding cassette transporter efflux pumps and implications for our understanding of eukaryotic P-glycoproteins and homologues. FEBS J 277:550–563
Kolaczkowski M, Sroda-Pomianek K, Kolaczkowska A, Michalak K (2013) A conserved interdomain communication pathway of pseudosymmetrically distributed residues affects substrate specificity of the fungal multidrug transporter Cdr1p. Biochim Biophys Acta 1828:479–490
Kovalchuk A, Driessen AJ (2010) Phylogenetic analysis of fungal ABC transporters. BMC Genomics 11:177
Kralli A, Yamamoto KR (1996) An FK506-sensitive transporter selectively decreases intracellular levels and potency of steroid hormones. J Biol Chem 271:17152–17156
Kueppers P, Gupta RP, Stindt J, Smits SH, Schmitt L (2013) Functional impact of a single mutation within the transmembrane domain of the multidrug ABC transporter Pdr5. Biochemistry 52:2184–2195
Kumar A, Shukla S, Mandal A, Shukla S, Ambudkar SV, Prasad R (2010) Divergent signature motifs of nucleotide binding domains of ABC multidrug transporter, CaCdr1p of pathogenic Candida albicans, are functionally asymmetric and noninterchangeable. Biochim Biophys Acta 1798:1757–1766
Lamping E, Baret PV, Holmes AR, Monk BC, Goffeau A, Cannon RD (2010) Fungal PDR transporters: Phylogeny, topology, motifs and function. Fungal Genet Biol 47:127–142
Lamping E, Cannon RD (2010) Use of a yeast-based membrane protein expression technology to overexpress drug resistance efflux pumps. Methods Mol Biol 666:219–250
Lamping E, Monk BC, Niimi K, Holmes AR, Tsao S, Tanabe K, Niimi M, Uehara Y, Cannon RD (2007) Characterization of three classes of membrane proteins involved in fungal azole resistance by functional hyperexpression in Saccharomyces cerevisiae. Eukaryot Cell 6:1150–1165
Lamping E, Niimi M, Niimi K, Tanabe K, Cannon RD (2014) How do the transmembrane domains of PDR transporters like Cdr1p fold. Paper presented at the conference celebrating 20 years of CDR1 research, New Delhi, India, 4–8 January 2014
Law CJ, Maloney PC, Wang DN (2008) Ins and outs of major facilitator superfamily antiporters. Annu Rev Microbiol 62:289–305
Lee JY, Kinch LN, Borek DM, Wang J, Wang J, Urbatsch IL, Xie XS, Grishin NV, Cohen JC, Otwinowski Z, Hobbs HH, Rosenbaum DM (2016) Crystal structure of the human sterol transporter ABCG5/ABCG8. Nature 533:561–564
Lee MD, Galazzo JL, Staley AL, Lee JC, Warren MS, Fuernkranz H, Chamberland S, Lomovskaya O, Miller GH (2001) Microbial fermentation-derived inhibitors of efflux-pump-mediated drug resistance. Farmaco 56:81–85
Li J, Jaimes KF, Aller SG (2014) Refined structures of mouse P-glycoprotein. Protein Sci 23:34–46
Li R, Kumar R, Tati S, Puri S, Edgerton M (2013) Candida albicans flu1-mediated efflux of salivary histatin 5 reduces its cytosolic concentration and fungicidal activity. Antimicrob Agents Chemother 57:1832–1839
Liu TT, Znaidi S, Barker KS, Xu L, Homayouni R, Saidane S, Morschhauser J, Nantel A, Raymond M, Rogers PD (2007) Genome-wide expression and location analyses of the Candida albicans Tac1p regulon. Eukaryot Cell 6:2122–2138
Locher KP (2004) Structure and mechanism of ABC transporters. Curr Opin Struct Biol 14:426–431
Ma J, Biggin PC (2013) Substrate versus inhibitor dynamics of P-glycoprotein. Proteins 81:1653–1668
Maebashi K, Niimi M, Kudoh M, Fischer FJ, Makimura K, Niimi K, Piper RJ, Uchida K, Arisawa M, Cannon RD, Yamaguchi H (2001) Mechanisms of fluconazole resistance in Candida albicans isolates from Japanese AIDS patients. J Antimicrob Chemother 47:527–536
Mandal A, Kumar A, Singh A, Lynn AM, Kapoor K, Prasad R (2012) A key structural domain of the Candida albicans Mdr1 protein. Biochem J 445:313–322
Moeller A, Lee SC, Tao H, Speir JA, Chang G, Urbatsch IL, Potter CS, Carragher B, Zhang Q (2015) Distinct conformational spectrum of homologous multidrug ABC transporters. Structure 23:450–460
Morschhauser J, Barker KS, Liu TT, Bla BWJ, Homayouni R, Rogers PD (2007) The transcription factor Mrr1p controls expression of the MDR1 efflux pump and mediates multidrug resistance in Candida albicans. PLoS Pathog 3:e164
Nannenga BL, Iadanza MG, Vollmar BS, Gonen T (2013) Overview of electron crystallography of membrane proteins: crystallization and screening strategies using negative stain electron microscopy. In: Coligan John E et al Current protocols in protein science Chapter 17: Unit17 15
Niimi K, Harding DR, Holmes AR, Lamping E, Niimi M, Tyndall JD, Cannon RD, Monk BC (2012) Specific interactions between the Candida albicans ABC transporter Cdr1p ectodomain and a D-octapeptide derivative inhibitor. Mol Microbiol 85:747–767
Nomura N, Verdon G, Kang HJ, Shimamura T, Nomura Y, Sonoda Y, Hussien SA, Qureshi AA, Coincon M, Sato Y (2015) Structure and mechanism of the mammalian fructose transporter GLUT5. Nature 526:397–401
Pasrija R, Banerjee D, Prasad R (2007) Structure and function analysis of CaMdr1p, a major facilitator superfamily antifungal efflux transporter protein of Candida albicans: identification of amino acid residues critical for drug/H+ transport. Eukaryot Cell 6:443–453
Paulsen IT, Brown MH, Littlejohn TG, Mitchell BA, Skurray RA (1996) Multidrug resistance proteins QacA and QacB from Staphylococcus aureus: membrane topology and identification of residues involved in substrate specificity. Proc Natl Acad Sci U S A 93:3630–3635
Paulsen IT, Skurray RA (1993) Topology, structure and evolution of two families of proteins involved in antibiotic and antiseptic resistance in eukaryotes and prokaryotes–an analysis. Gene 124:1–11
Perlin MH, Andrews J, Toh SS (2014) Essential letters in the fungal alphabet: ABC and MFS transporters and their roles in survival and pathogenicity. Adv Genet 85:201–253
Prajapati R, Sangamwar AT (2014) Translocation mechanism of P-glycoprotein and conformational changes occurring at drug-binding site: Insights from multi-targeted molecular dynamics. Biochim Biophys Acta 1838:2882–2898
Prasad R, Goffeau A (2012) Yeast ATP-binding cassette transporters conferring multidrug resistance. Annu Rev Microbiol 66:39–63
Puri N, Gaur M, Sharma M, Shukla S, Ambudkar SV, Prasad R (2009) The amino acid residues of transmembrane helix 5 of multidrug resistance protein CaCdr1p of Candida albicans are involved in substrate specificity and drug transport. Biochim Biophys Acta 1788:1752–1761
Puri N, Prakash O, Manoharlal R, Sharma M, Ghosh I, Prasad R (2010) Analysis of physico-chemical properties of substrates of ABC and MFS multidrug transporters of pathogenic Candida albicans. Eur J Med Chem 45:4813–4826
Quistgaard EM, Low C, Guettou F, Nordlund P (2016) Understanding transport by the major facilitator superfamily (MFS): structures pave the way. Nat Rev Mol Cell Biol 17:123–132
Rai V, Gaur M, Shukla S, Shukla S, Ambudkar SV, Komath SS, Prasad R (2006) Conserved Asp327 of walker B motif in the N-terminal nucleotide binding domain (NBD-1) of Cdr1p of Candida albicans has acquired a new role in ATP hydrolysis. Biochemistry 45:14726–14739
Ravna AW, Sylte I, Sager G (2009) Binding site of ABC transporter homology models confirmed by ABCB1 crystal structure. Theor Biol Med Model 6:20
Rawal MK, Khan MF, Kapoor K, Goyal N, Sen S, Saxena AK, Lynn AM, Tyndall JD, Monk BC, Cannon RD, Komath SS, Prasad R (2013) Insight into pleiotropic drug resistance ATP-binding cassette pump drug transport through mutagenesis of Cdr1p transmembrane domains. J Biol Chem 288:24480–24493
Reddy VS, Shlykov MA, Castillo R, Sun EI, Saier MH Jr (2012) The major facilitator superfamily (MFS) revisited. FEBS J 279:2022–2035
Rees DC, Johnson E, Lewinson O (2009) ABC transporters: the power to change. Nat Rev Mol Cell Biol 10:218–227
Rigaud J-L, Chami M, Lambert O, Levy D, Ranck J-L (2000) Use of detergents in two-dimensional crystallization of membrane proteins. Biochim Biophys Acta (BBA)-Biomembranes 1508:112–128
Rosenberg MF, Kamis AB, Callaghan R, Higgins CF, Ford RC (2003) Three-dimensional structures of the mammalian multidrug resistance P-glycoprotein demonstrate major conformational changes in the transmembrane domains upon nucleotide binding. J Biol Chem 278:8294–8299
Rutledge RM, Esser L, Ma J, Xia D (2011) Toward understanding the mechanism of action of the yeast multidrug resistance transporter Pdr5p: a molecular modeling study. J Struct Biol 173:333–344
Sa-Correia I, dos Santos SC, Teixeira MC, Cabrito TR, Mira NP (2009) Drug: H+ antiporters in chemical stress response in yeast. Trends Microbiol 17:22–31
Saini P, Prasad T, Gaur NA, Shukla S, Jha S, Komath SS, Khan LA, Haq QM, Prasad R (2005) Alanine scanning of transmembrane helix 11 of Cdr1p ABC antifungal efflux pump of Candida albicans: identification of amino acid residues critical for drug efflux. J Antimicrob Chemother 56:77–86
Sanglard D, Kuchler K, Ischer F, Pagani JL, Monod M, Bille J (1995) Mechanisms of resistance to azole antifungal agents in Candida albicans isolates from AIDS patients involve specific multidrug transporters. Antimicrob Agents Chemother 39:2378–2386
Sauna ZE, Bohn SS, Rutledge R, Dougherty MP, Cronin S, May L, Xia D, Ambudkar SV, Golin J (2008) Mutations define cross-talk between the N-terminal nucleotide-binding domain and transmembrane helix-2 of the yeast multidrug transporter Pdr5: possible conservation of a signaling interface for coupling ATP hydrolysis to drug transport. J Biol Chem 283:35010–35022
Sauna ZE, Kim IW, Nandigama K, Kopp S, Chiba P, Ambudkar SV (2007) Catalytic cycle of ATP hydrolysis by P-glycoprotein: evidence for formation of the E. S reaction intermediate with ATP-gamma-S, a nonhydrolyzable analogue of ATP. Biochemistry 46:13787–13799
Seelig A, Landwojtowicz E (2000) Structure-activity relationship of P-glycoprotein substrates and modifiers. Eur J Pharm Sci 12:31–40
Shah AH, Rawal MK, Dhamgaye S, Komath SS, Saxena AK, Prasad R (2015) Mutational analysis of intracellular loops identify cross talk with nucleotide binding domains of yeast ABC transporter Cdr1p. Sci Rep 5:11211
Shukla S, Ambudkar SV, Prasad R (2004) Substitution of threonine-1351 in the multidrug transporter Cdr1p of Candida albicans results in hypersusceptibility to antifungal agents and threonine-1351 is essential for synergic effects of calcineurin inhibitor FK520. J Antimicrob Chemother 54:38–45
Shukla S, Saini P, Smriti Jha S, Ambudkar SV, Prasad R (2003) Functional characterization of Candida albicans ABC transporter Cdr1p. Eukaryot Cell 2:1361–1375
Silva LV, Sanguinetti M, Vandeputte P, Torelli R, Rochat B, Sanglard D (2013) Milbemycins: more than efflux inhibitors for fungal pathogens. Antimicrob Agents Chemother 57:873–886
Solcan N, Kwok J, Fowler PW, Cameron AD, Drew D, Iwata S, Newstead S (2012) Alternating access mechanism in the POT family of oligopeptide transporters. EMBO J 31:3411–3421
Tanabe K, Lamping E, Nagi M, Okawada A, Holmes AR, Miyazaki Y, Cannon RD, Monk BC, Niimi M (2011) Chimeras of Candida albicans Cdr1p and Cdr2p reveal features of pleiotropic drug resistance transporter structure and function. Mol Microbiol 82:416–433
Tsao S, Rahkhoodaee F, Raymond M (2009) Relative contributions of the Candida albicans ABC transporters Cdr1p and Cdr2p to clinical azole resistance. Antimicrob Agents Chemother 53:1344–1352
Tutulan-Cunita AC, Mikoshi M, Mizunuma M, Hirata D, Miyakawa T (2005) Mutational analysis of the yeast multidrug resistance ABC transporter Pdr5p with altered drug specificity. Genes to cells: devoted to molecular & cellular mechanisms 10:409–420
Velamakanni S, Yao Y, Gutmann DA, van Veen HW (2008) Multidrug transport by the ABC transporter Sav 1866 from Staphylococcus aureus. Biochemistry 47:9300–9308
Vinothkumar KR (2015) Membrane protein structures without crystals, by single particle electron cryomicroscopy. Curr Opin Struct Biol 33:103–114
Walker JE, Saraste M, Runswick MJ, Gay NJ (1982) Distantly related sequences in the alpha- and beta-subunits of ATP synthase, myosin, kinases and other ATP-requiring enzymes and a common nucleotide binding fold. EMBO J 1:945–951
Ward A, Reyes CL, Yu J, Roth CB, Chang G (2007) Flexibility in the ABC transporter MsbA: alternating access with a twist. Proc Natl Acad Sci U S A 104:19005–19010
Ward AB, Szewczyk P, Grimard V, Lee CW, Martinez L, Doshi R, Caya A, Villaluz M, Pardon E, Cregger C, Swartz DJ, Falson PG, Urbatsch IL, Govaerts C, Steyaert J, Chang G (2013) Structures of P-glycoprotein reveal its conformational flexibility and an epitope on the nucleotide-binding domain. Proc Natl Acad Sci U S A 110:13386–13391
White TC (1997) Increased mRNA levels of ERG16, CDR, and MDR1 correlate with increases in azole resistance in Candida albicans isolates from a patient infected with human immunodeficiency virus. Antimicrob Agents Chemother 41:1482–1487
White TC, Holleman S, Dy F, Mirels LF, Stevens DA (2002) Resistance mechanisms in clinical isolates of Candida albicans. Antimicrob Agents Chemother 46:1704–1713
Wirsching S, Michel S, Morschhauser J (2000) Targeted gene disruption in Candida albicans wild-type strains: the role of the MDR1 gene in fluconazole resistance of clinical Candida albicans isolates. Mol Microbiol 36:856–865
Xu K, Zhang M, Zhao Q, Yu F, Guo H, Wang C, He F, Ding J, Zhang P (2013) Crystal structure of a folate energy-coupling factor transporter from Lactobacillus brevis. Nature 497:268–271
Yamada-Okabe T, Yamada-Okabe H (2002) Characterization of the CaNAG3, CaNAG4, and CaNAG6 genes of the pathogenic fungus Candida albicans: possible involvement of these genes in the susceptibilities of cytotoxic agents. FEMS Microbiol Lett 212:15–21
Yan N (2013) Structural advances for the major facilitator superfamily (MFS) transporters. Trends Biochem Sci 38:151–159
Yan N (2015) Structural biology of the major facilitator superfamily transporters. Annu Rev Biophys 44:257–283
Yin Y, He X, Szewczyk P, Nguyen T, Chang G (2006) Structure of the multidrug transporter EmrD from Escherichia coli. Science 312:741–744
Znaidi S, Weber S, Al-Abdin OZ, Bomme P, Saidane S, Drouin S, Lemieux S, De Deken X, Robert F, Raymond M (2008) Genomewide location analysis of Candida albicans Upc2p, a regulator of sterol metabolism and azole drug resistance. Eukaryot Cell 7:836–847
Acknowledgements
The authors acknowledge financial support from the Marsden Fund of the Royal Society of New Zealand (UOO1305) and a research grant from Chulalongkorn University, Thailand.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Lamping, E., Madani, G., Lee, H.J., Niimi, M., Cannon, R.D. (2017). Structure–Function Analyses of Multidrug Transporters. In: Prasad, R. (eds) Candida albicans: Cellular and Molecular Biology. Springer, Cham. https://doi.org/10.1007/978-3-319-50409-4_18
Download citation
DOI: https://doi.org/10.1007/978-3-319-50409-4_18
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-50408-7
Online ISBN: 978-3-319-50409-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)