Abstract
Oncogenic transcription factors represent unique and potentially high value targets for cancer therapy. Proteins like STAT3 and STAT5 are generally not mutated themselves. However, oncogenic signals arising from a wide array of upstream mutations and signaling events converge on a small number of these transcription factors to regulate expression of key genes involved in critical processes including proliferation, survival and invasion. While cancer cells frequently show a high dependency on continued activation of these proteins, normal cells are largely tolerant to interruption of these pathways due to redundancies in transcriptional regulators. Consequently, inhibition of STATs holds the potential to have a very high therapeutic index. The challenge has been to develop strategies to inhibit these proteins that lack domains that are easily amenable to antagonism by small molecules. In recent years, a number of promising strategies have emerged, and now clinical trials of approaches to directly inhibit activated STATs have been developed. The success of these studies, both in terms of clinical efficacy and understanding the molecular effects of STAT inhibitors in humans, may open a new front in the rational, targeted eradication of cancer.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Cancer
- Drug discovery
- Gene expression
- Signal transduction
- STAT transcription factors
- Targeted therapy
- Clinical trials
3.1 Introduction
Cancer therapy has evolved greatly since its advent in the 1940s, progressing from non-specifically cytotoxic anti-metabolites and alkylating agents to the targeted agents, like kinase inhibitors, that are now available. With the introduction of imatinib (Gleevec), an inhibitor of the Bcr/Abl1 fusion kinase found essentially universally in chronic myeloid leukemia (CML), the treatment of CML was revolutionized. However, for most other cancers it has proven difficult to identify activated kinases that provide the same therapeutic opportunity. This has raised the question of whether there are common downstream mediators of cancer-driving mutations, which may not be mutated themselves, but which are critical convergence points of oncogenic signaling. In particular, transcription factors, which tightly choreograph the expression of genes under physiologic conditions, can become activated inappropriately in most cancers. While a single transcription factor can often be deleted from normal cells without deleterious consequences to an organism, typically due to redundancies in physiologic signaling, the same transcription factor may represent a critical dependency to a cancer cell. Transcription factors are difficult targets from a medicinal chemistry standpoint. However, the opportunity presented by the fact that they may provide a high therapeutic index has attracted increased attention. The key questions that emerge are whether they are truly critical targets, and whether they can successfully be inhibited in human clinical trials.
3.2 STAT Activation in Cancer
Signal transducers and activators of transcription (STATs) are a family of transcription factors that play important roles in a range of cellular functions. STATs reside in the cytoplasm under basal conditions. Upon activation by tyrosine phosphorylation, STATs form active dimers, translocate to the nucleus, bind to DNA, and regulate transcription of target genes [1]. Under physiological conditions, STATs are activated only transiently. By contrast, in many forms of cancer, STAT family members are activated constitutively and drive the expression of genes underlying malignant cellular behavior. Two family members in particular, STAT3 and STAT5, are activated most commonly in a range of human cancers. Constitutive activation of these transcription factors can directly lead to cancer pathogenesis [2].
3.2.1 Hematologic Malignancies
The transcription factor STAT5 encompasses two highly homologous proteins, STAT5A and STAT5B. In hematological malignancies in particular, inappropriate activation of STAT5 is a common event that leads to increased expression of genes regulating cell cycle progression and survival [3–5]. STAT5 is constitutively active in chronic myelogenous leukemia (CML) [6, 7], acute myelogenous leukemia (AML), acute lymphocytic leukemia (ALL) [3, 8], and Hodgkin lymphoma [9]. STAT5 phosphorylation can be mediated by mutated tyrosine kinases (TKs, such as BCR-ABL1 and JAK2V617F [10, 11]), or autocrine secretion of cytokines that signal through Janus kinase (JAK) [9]. STAT5 plays a crucial role in mediating survival signals emanating from these upstream oncogenic kinases, since disruption of STAT5 abrogate tumorigenesis induced by the oncogenic kinases.
STAT3 is activated in leukemia and lymphoma often through Janus kinases (JAKs). Recently it was found that in anaplastic large cell lymphoma, multiple driver genetic alterations leads to oncogenic STAT3 activation. JAK/STAT3 pathway inhibition consistently impaired lymphoma cell growth in vitro and in vivo [12]. STAT3 also mediates oncogenic addiction to TEL-AML1 in t(12;21) ALL. Consequently, human leukemic cell lines carrying this translocation are highly sensitive to treatment with STAT3 inhibitor [13].
3.2.2 Solid Tumors
STAT activation in solid tumors often occurs through autocrine or paracrine secretion of cytokines and this is also mediated through JAKs. Reflecting how oncogenic pathways often subvert physiologic signaling events, STAT5 plays an important role in normal mammary gland development, and it frequently becomes constitutively activated in breast cancer. The activation of STAT5 in breast cancer may be due to the autocrine, paracrine, or endocrine secretion of prolactin. In the mammary gland, STAT5 is activated late in pregnancy in response to prolactin to promote terminal differentiation and milk production [14–16]. In breast cancer, constitutively activated STAT5 enhances both survival and anchorage-independent growth of human mammary carcinoma cells [17]. Mice that express a constitutively activated form of STAT5 develop mammary carcinomas, whereas mice that lack STAT5A are protected against mammary tumors induced by transforming growth factor α [15, 18, 19].
Using immunohistochemistry to tyrosine phosphorylated STAT3, and similar techniques, it has been found that STAT3 is constitutively activated in an even wider range of solid tumors compared to STAT5, including breast cancer [20], ovarian cancer [21, 22], gastric cancer [23], colorectal cancer [24], lung cancer [25], glioblastoma [26], and pancreatic cancer [27]. Other methods have also revealed a critical role for STAT3 in human cancers. For example, by combining genome-wide RNAi screens with regulatory network analysis, STAT3 has been identified as a critically activated master regulator of HER2(+) breast cancers [28]. In these systems, STAT3 is frequently activated through an IL-6-dependent JAK2-calprotectin axis and inhibition of this axis alone or in combination with HER2 inhibitors reduces tumorigenicity of hormone receptor (−)/HER2(+) breast cancers.
3.3 Critical Role of STATs in the Survival of Cancer Stem-Like Cells
Persistence of cancer stem cells may promote resistance and recurrence of cancer after treatment. Therefore, therapies that target stem and progenitor cells may be particularly important in achieving long-term remissions of cancer. STAT activation has been suggested to play a critical role in cancer stem cell survival in both hematopoietic and solid cancers. Constitutive STAT activation has been implicated in leukemia stem cell self-renewal [29–31]. For example, STAT signaling is enriched in and critical for leukemia stem cell self-renewal in MN1- and HOXA9-co-expressing leukemias, types that harbor a particularly poor prognosis [29]. STAT5 can confer long-term expansion exclusively on human HSCs, by directly modulating Hypoxia-induced factor 2α (HIF2α) expression [31]. In comparing JAK/STAT signaling between leukemia stem cells (LSCs) and normal stem cells from clinical samples, it was found that JAK/STAT signaling is significantly increased in LSCs, particularly from high-risk AML patients. JAK2 inhibition using small molecule inhibitors or RNA interference reduced the growth of AML LSCs while sparing normal stem cells both in vitro and in vivo [32]. Recently it was found that CML stem cell survival is not dependent on the BCR-ABL1 protein kinase, but rather JAK/STAT5 signaling. Thus, while treatment with an ABL tyrosine kinase inhibitor alone may not cure CML patients, dual inhibition of both JAK and BCR-ABL1 may be critical for eradicating primitive quiescent CML stem cells [33]. This observation not only further supports the importance of inhibiting STAT5 in eliminating leukemia stem cells, but also highlights the fact that STAT5 can be activated by multiple aberrant kinases within leukemic stem cells to maintain their survival. Inhibiting STAT5 directly, as the convergence point of multiple upstream oncogenic kinases, may be crucial in achieving a durable therapeutic response.
The critical role of STATs was also found in solid tumor stem-like cells. In glioblastoma, STAT3 was only found to be activated in stem-like cells where it promoted tumorigenicity, but not in more differentiated cells populations [34]. In breast cancer, the JAK2/STAT3 signaling pathway is preferentially active in and required for growth of CD44+CD24− stem cell-like cancer cells in human tumors [35]. In endometrial cancer, IL-6/JAK1/STAT3 signaling is essential for maintenance of an ALDHhi/CD126+ stem-like component [36]. Furthermore, a small molecule inhibitor targeting STAT3 is effective in inhibiting expression of “stemness” genes and suppresses cancer relapse and metastasis [37]. All of these observations suggest that targeting STATs has the potential to reduce a stem-like cancer cell compartment, which may be the cause of resistance to therapy and tumor recurrence.
3.4 STAT Activation as an Important Mechanism of Resistance to Cancer Therapy
While oncogenic kinase targeted therapy has been extremely successful in the treatment of CML, resistance to targeted therapies develops rapidly in most forms of cancer. One of the important common pathways that mediate this resistance is alternative activation of STATs. It was found that AML cells quickly developed resistance to multi-targeted tyrosine kinase inhibitors through activated JAK2/STAT5 signaling [38]. Even in CML, patients who initially responded well to TKIs could acquire resistance leading to progression of their disease. Indeed, increased activation of STAT5 has been associated with leukemia progression and TKI resistance in CML [39, 40]. In addition, it has been suggested that an increased level of STAT5 triggers BCR-ABL1 mutation, leading to an increase in inhibitor-resistant BCR-ABL1 mutations [41].
STAT activation has also been found to be an important resistance mechanism in solid tumors. It was found that JAK-mediated STAT5 signaling closely interacts with the PI3K/AKT pathway and mediates resistance to PI3K/AKT inhibition in breast cancer [42]. In melanoma, activation of STAT3 can be induced by MEK or BRAF inhibitors, leading to melanoma cells that are not only resistant to those inhibitors, but also acquire a more invasive phenotype [43, 44]. Thus BRAF inhibitors may need to be combined with STAT3 inhibition to achieve a clinically sustainable response in melanoma [45]. Similarly, in ovarian cancer, resistance to anti-VEGF therapy was found to be mediated through autocrine IL-6/STAT3 signaling [46].
STAT3 activation not only accounts for resistance to targeted therapy, but also plays an important role in resistance to traditional cytotoxic cancer therapies. For example, activated STAT3 can upregulate BCL2 in metastatic breast cancer to promote resistance to chemotherapy [47]. In addition, inhibiting STAT3 activation by blocking IL-6 signaling has been shown to sensitize multiple tumor types to chemotherapy [48]. Tumor permeability is a critical determinant of drug delivery and sensitivity. Using three-dimensional (3D) culture condition, JAK/STAT3 signaling pathway was identified as an essential regulator of tumor permeability barrier function, and STAT3 inhibition increased drug sensitivity. The combination of STAT3 inhibition and 5-FU chemotherapy markedly reduced tumor growth compared to monotherapy. STAT3 activation was also found to be associated with proneural-to-mesenchymal transition observed in gliomas upon radiation therapy [49]. Thus, STAT3 inhibition could be helpful in preventing emergence of therapy-resistant mesenchymal glioma at relapse.
3.5 STAT-Mediated Modulation of the Tumor Microenvironment
STAT3 activation not only directly regulates genes that mediate anti-apoptotic signals and promote malignant cells survival within cells, STAT3 also modulates genes that modify the tumor microenvironment to promote tumor cell survival. For example, not only does activated STAT3 promote angiogenesis, activation of STAT3 also contributes to tumor immune evasion [50]. In STAT3-deficient mice, hyperplastic and early adenoma-like lesions initially formed, but they later completely regressed. This tumor regression correlated with massive immune infiltration into the STAT3-deficient lesions, leading to their elimination [51]. In head and neck squamous cells carcinoma, STAT3 inhibition by siRNA knock-down resulted in enhanced expression and secretion of both pro-inflammatory cytokines and chemokines, and led to the activation of dendritic cells and lymphocytes [52]. STAT3 inhibition was also found to enhance the therapeutic efficacy of immunogenic chemotherapeutic drugs, such as anthracyclines, by stimulating type 1 interferon production by cancer cells [53]. The important immune checkpoint pathway mediated through programmed death-1 (PD-1) is activated by STAT3 in classic Hodgkin’s lymphoma, with JAK-STAT signaling found to promote the induction and increase the abundance of PD-1 ligands expressed on Reed-Sternberg cells [54], which upon binding with PD-1 on tumor infiltrating lymphocytes (TILs), leads to TIL dysfunction.
3.6 Potential Disadvantages of Targeting STAT Transcription Factors in Cancer
Despite the convincing evidence that inappropriate activation of STAT3 and STAT5 can promote oncogenesis, there is evidence showing that in certain cellular context, STAT3 or STAT5 may exert tumor suppressive activities. Identifying and characterizing these cellular contexts is equally essential in designing effective targeted therapies for these proteins. For example, it has been found that mouse models with STAT5-deficiency in hematopoietic cells are permissive for Myc-induced B-cell leukemogenesis [55]. In JAK2V617F-driven myeloproliferative neoplasms in mouse models, deletion of STAT3 enhances myeloid cell expansion and increases the severity of myeloproliferative diseases [56]. In a Pten-deficient prostate cancer mouse model, genetic inactivation of STAT3 or IL-6 signaling accelerates cancer progression leading to metastasis. In addition, loss of STAT3 signaling was found to disrupt the ARF-Mdm2-p53 tumor suppressor axis through bypassing senescence [57].
The role of STAT3 in KRAS-induced malignancy is more complicated, with different mouse models showing distinct roles of STAT3 in KRAS-driven malignancy. In mouse pancreatic cancer models, STAT3 was shown to be essential for pancreatic ductal adenocarcinoma initiation and progression driven by KRAS [58, 59]. On the other hand, in lung adenocarcinoma models also driven by KRAS, two different groups demonstrated that depletion of STAT3 accelerates RAS-induced lung cancer [60–62]. Since the mitochondrial role of STAT3 in supporting KRAS-induced transformation has been well established [63], it is possible that the conflicting effects of STAT3 in KRAS-dependent malignancy may be related to its role as a transcription factor. STAT3 may drive different sets of target genes expression that either support or antagonize KRAS-induced transformation in different cellular contexts.
In breast cancer, both molecular and epidemiological evidence suggests that the co-activation of STAT5 with STAT3 leads to a less aggressive tumor. This may be mediated, at least in part, by modulation of expression of the oncogenic transcriptional regulator BCL6. Whereas expression of this gene is induced by STAT3, it is repressed by STAT5, even in the presence of activated STAT3 [64, 65]. Thus, it remains unclear as to whether inhibition of STAT5 will be of therapeutic value in the large fraction of breast cancers in which both STAT3 and STAT5 are activated.
3.7 Unbiased Approaches to Identify STAT Inhibitors
Based on our understanding of the mechanism of STAT activation in cancer, various strategies to inhibit STAT transcriptional function have been designed. One approach is to use structure-based design, targeting specific STAT domains or critical steps in STAT function [4]. Such approaches include cytokine receptor-directed monoclonal antibodies, tyrosine kinase inhibitors, SH2 domain inhibitors [66], and antisense oligonucleotides or small molecules [67] that target the STAT DNA binding domain [4]. An alternate approach is to use screening strategies to identify compounds that inhibit STAT-based transcription. One way to do this is to use a chemical biology approach in which a cell-based system is developed that allows the quantitative high-throughput measurement of STAT-dependent gene expression. Another screening strategy makes use of a computational approach using databases that catalog the effect of thousands of drugs on gene expression [68] and gene expression signatures that reflect the activation of STATs in human cancers [69] to identify drugs that lead to gene expression signatures that are the opposite of the STAT signature. These unbiased approaches greatly expand the range of potential STAT inhibitors that can be identified. Compounds identified by these strategies also serve as biological probes that provide insight into the physiologic mechanisms of STAT regulation in a cell, and identify new targets for therapeutic inhibition.
3.8 Post-Translational Modifications and STAT Transcriptional Function
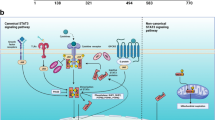
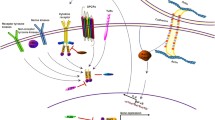
While STATs can be activated by cytokine-induced JAK activation, or receptor or non-receptor tyrosine kinases, there are additional subtleties that regulate their transcriptional function. STAT proteins can be post-translationally modified at different locations, in addition to the canonical tyrosine phosphorylation, and several of those modifications have been shown to modulate STAT transcriptional function (Fig. 3.1). For example, STATs can be phosphorylated, acetylated, methylated or ubiquitinated on several amino acid residues. In many tumor types, phosphorylation of both Tyr-705 (Y705) and Ser-727 (S727) is important for STAT3 transcriptional function. Phosphorylation of S727 was believed to occur after Y705 phosphorylation and binding with the target promoter to further augment the transcriptional function of STATs [70]. In certain cancers such as chronic lymphocytic leukemia (CLL), only S727 phosphorylation of STAT3 is observed [71], though this is sufficient to drive target gene expression [72]. In renal cell carcinoma, STAT3 was found to be phosphorylated by glycogen synthase kinase 3α and -β (GSK-3α/β) at T714 and S727, but not Y705, to drive target gene expression [73]. There is also evidence that acetylation of STAT3 enhances the stability and interaction of STAT3 with P300 bromodomain protein to increase transcription [74].
Inappropriate activation of STAT transcription factors drive the expression of critical target genes in cancer, and so STATs represent targets with a potentially high therapeutic index. STATs can become activated constitutively in cancer cells through phosphorylation by mutated oncogenic tyrosine kinases, or through cytokines that are present in the tumor microenvironment through autocrine or paracrine mechanisms, thereby activating JAKs. Upon tyrosine phosphorylation, STATs form active dimers, translocate to the nucleus, bind to DNA, and regulate transcription of target genes that regulate self-renewal (“stemness”), survival, angiogenesis, and immune evasion. The transcriptional function of STATs is modulated by post-translational modifications including phosphorylation, methylation and acetylation. Co-factors that interact with STATs at the genomic level serve as another level of transcriptional regulation. Understanding these mechanisms of regulating STAT function has led to a number of therapeutic opportunities to target these proteins. (P phosphorylation, Me methylation, Ac acetylation)
STAT5 encompass two isoforms, STAT5A and STAT5B. The canonical activation marker for STAT5A is Y694 and for STAT5B is Y699 [75–78]. STAT5A can also be serine phosphorylated at multiple sites such as S726, S780 and S127/128. At least in the case of ERBB4/HER4 activated STAT5A, S779 phosphorylation seemed dispensable for phosphorylation of STAT5A at Y694 and subsequent DNA binding. However S127/S128 was required for ERBB4-induced phosphorylation of Y694 of STAT5A [79]. STAT5B can be serine phosphorylated at S731 and S193 [75, 80]. Furthermore, although Y699 is absolutely required for transcriptional activation of STAT5B, tyrosines 725, 740, and 743 may be involved in a negative regulation of STAT5B-mediated transcription [81].
Recently, key methylation sites that modulate STAT3 transcriptional activity have been identified, though methylation at different sites on STAT3 may exert completely opposite effects on transcriptional activity. For example, following its tyrosine phosphorylation, STAT3 is methylated on K140 by the histone methyl transferase SET9 and demethylated by LSD1. This methylation of K140 is a negative regulatory event [70]. On the other hand, STAT3 can be methylated at different sites by the same enzyme, enhancer of Zeste homolog 2 (EZH2) to activate its transcriptional function. EZH2 is a lysine methyl transferase and EZH2-containing PRC2 catalyzes trimethylation of histone 3 at lysine 27 (H3K27me3) [82]. It has recently been appreciated that EZH2 also methylates non-histone proteins. Two independent studies have demonstrated that EZH2 modulates STAT3 transcriptional activity by methylating distinct sites of STAT3. In glioblastoma stem cells, EZH2 trimethylates STAT3 on K180. Trimethylation at K180 promoted Y705 phosphorylation of STAT3 and activated STAT3 transcriptional activity [34]. It is still unknown how trimethylation at K180 synergize with Y705 phosphorylation of STAT3 in glioblastoma stem cells. In another cellular system in which STAT3 is activated by IL-6, perturbation of EZH2 function did not inhibit Y705 phosphorylation of STAT3, although it significantly reduced STAT3 transcriptional activity. It was found that in this IL-6 dependent system, dimethylation of K49 of STAT3 by EZH2 was crucial for full activation of STAT3 transcriptional activity. Unlike K180 trimethylation that promoted Y705 phosphorylation, dimethylation of K49 had no effect on Y705 phosphorylation. On the contrary, Y705 phosphorylation was required for K49 dimethylation of STAT3 to occur [83]. The mechanism by which K49 modification altered STAT3-dependent gene expression is unclear. It does not appear that K49 methylation affected the binding of STAT3 to its genomic binding site. It has been suggested that K49 methylation of STAT3 promotes the recruitment of co-regulatory factors to genomic target sites to facilitate maximal transcriptional function of STAT3, although these postulated co-regulators have not yet been identified.
3.9 Identification of Clinically-Translatable STAT Inhibitors
Although different modifications can affect STAT3 transcriptional function, it is clear that Y705 phosphorylation is nearly always essential for transcriptional activity. Thus drug screening and structure-based design of STAT inhibitors have mainly focused on inhibition of this phosphorylation event in STAT3. Many inhibitors of STAT tyrosine phosphorylation have been identified that block the STAT3 SH2 domain, which is required for both recruitment to activated kinase-receptor complexes as well as for activating dimerization. In addition, a number of natural products have been described that inhibit STAT3 phosphorylation. While these molecules have encouraging properties in vitro, and some have shown activity in animal models, progress in advancing STAT-targeted small molecules into clinical trials in cancer patients has been slow.
As noted, cell-based screening systems can be used to identify inhibitors of STAT-dependent transcription. This approach can allow the screening of chemical libraries that contain drugs that are already known to be safe in humans, including those that are approved for human use. This approach has identified several notable compounds, two of which function by blocking STAT3 tyrosine phosphorylation, albeit through different mechanisms. Nifuroxazide, an oral antibiotic that is used in many countries to treat colitis and diarrhea in humans, was found to be an inhibitor of STAT3 transcriptional function with an EC 50 of approximately 3 μM [84]. In analyzing its mechanism of action, it was found that nifuroxazide inhibited Y705 phosphorylation of STAT3 through inhibiting the kinase activity of both TYK2 and JAK2 (but not JAK1). Nifuroxazide was found to induce apoptosis and reduce the viability of multiple myeloma cells that are dependent on activated STAT3 for survival.
Another compound identified through this approach is pimozide, which is clinically used as a neuroleptic for the treatment of Tourette syndrome. This drug was found to decrease STAT5 tyrosine phosphorylation. Interestingly, pimozide inhibits STAT5 phosphorylation irrespective of the upstream kinases that activate STAT5. Indeed, pimozide inhibits STAT5 phosphorylation in CML cells in which STAT5 is activated by the BCR-ABL1 fusion kinase [85], AML cells in which STAT5 is activated by FLT3-ITD [86], and myeloproliferative neoplasms in which STAT5 is activated by the mutated kinase JAK2(V617F) [5]. However, pimozide is not a kinase inhibitor. It does not inhibit JAKs, ABL1 or SRC family members in in vitro kinase assays, nor does it inhibit other signaling pathways downstream of those activated kinases. These findings suggested that pimozide inhibits STAT5 phosphorylation using a completely independent mechanism. The exact mechanism by which pimozide mediates this effect is not known, although it may involve modulation of negative regulators of STAT function. However, this non-kinase dependent STAT5 inhibition by pimozide may provide an important therapeutic opportunity. First, kinase mutation or amplification frequently leads to a reduction or loss of efficacy of kinases inhibitors. Therapies that target STAT5 independent of upstream kinases may still be able to achieve therapeutic efficacy. Indeed, hematopoietic cells with the T315I mutation in BCR-ABL are completely resistant to the BCR-ABL1 kinase inhibitor imatinib, but they are still sensitive to STAT5 inhibition by pimozide [85]. Second, even without BCR-ABL mutation, increased amount of STAT5 have been seen in the accelerated stage of CML and can render CML cells more resistant to imatinib [39]. In this situation, it is conceivable that a drug like pimozide that targets STAT5 without depending on upstream kinase inhibition will be valuable in controlling diseases. In addition, two compounds that inhibit different steps of the same oncogenic pathway may have greater efficacy with a lower chance of the emergence of resistance. Consistent with this idea, combining pimozide with kinase inhibition augmented the therapeutic efficacy of a JAK inhibitor in myeloproliferative diseases [5].
3.10 Therapeutic Modulation of Co-Factors of STATs
As with other transcription factors, STATs recruit co-factors to activate transcription, which can include other transcription factors, as well as chromatin remodeling proteins, among others. Cross talk between STATs and members of the nuclear receptor family has been observed in normal breast tissue and breast cancer [87–92]. Progesterone receptor (PR), androgen receptor (AR), and glucocorticoid receptor (GR), have all been shown to synergistically interact with STAT5 and enhance STAT5 target gene expression.
BRG1, the ATPase subunit of a chromatin remodeling complex, is another factor that is essential for STAT3 target gene transcription. Genome-wide STAT3 binding in pluripotent embryonic stem cells (ESCs) is dependent on BRG1, since BRG1 is required to establish chromatin accessibility at STAT3 binding targets [93].
To identify STAT3-interacting proteins that contribute to STAT3 tumorigenesis, one can use mass-spectrometry to profile STAT3-interacting proteins. This approach has allowed the identification of granulin (GRN) as a novel STAT3 interacting protein in triple negative breast cancer cells [94]. GRN can act as an autocrine growth factor [95], and it can bind to and alter the subcellular distribution of positive transcription elongation factor (P-TEFb), leading to the repression of the transcription of tumor suppressor genes [96]. In breast cancer cells, GRN enhances STAT3 DNA binding and increases the time-integrated amount of LIF-induced STAT3 phosphorylation in breast cancer cells. Furthermore, silencing GRN neutralizes STAT3-mediated proliferation and migration of breast cancer cells. The correlation between GRN and STAT3 was also observed in primary breast cancer samples, where GRN mRNA levels were positively correlated with STAT3 gene expression signatures and with reduced patient survival.
Many of the co-regulators of STATs that have been identified may be difficult targets for pharmacological intervention. However, one group of key transcriptional co-factors is the BET (bromodomains and extra-terminal domain) family of bromodomain-containing proteins, which includes BRD2, BRD3, BRD4 and BRDT. Nuclear BET-protein interactome studies have indicated that BET proteins are integral components of a large number of nuclear protein complexes [97, 98]. Consistent with a role for BET proteins as key modulators of STAT signaling, it was found that the bromodomain inhibitor JQ1 inhibits STAT5 transcriptional activity. Further RNA interference-based experiments demonstrated that among the three BET bromodomain proteins expressed in hematological malignancies and targeted by JQ1, only BRD2 is necessary for STAT5 transcriptional function [99]. BRD2 likely participates in the STAT5 transcriptional complex, and acts as a critical co-activator for STAT5 function. The recruitment of STAT5 to its genomic binding sites is not dependent on BRD2, but rather maximal transcriptional initiation of these target genes requires BRD2. Interestingly, although JQ1 significantly reduces the transcriptional function of STAT5, it had essentially no effects on STAT3-dependent gene expression. Given the structural similarity between STAT5 and STAT3, further genomic and structural studies are necessary to elucidate the mechanism of this selectivity. The therapeutic implication of targeting STAT5 by dual BET bromodomain inhibition (JQ1) and tyrosine kinase inhibition (TKIs) was investigated in a clinically aggressive disease, acute T lymphocytic leukemia. Strong synergy in the induction of apoptosis was found in T-ALL cells when JQ1 was combined with TKIs [99]. Over-expression of a constitutively activated STAT5 rescued cell death induced by the combination of JQ1 and TKIs, supporting the notion that the synergistic effect is, at least partially, mediated through STAT5 inhibition. These findings also reaffirm the important role of STAT5 activation in the pathogenesis of T-ALL.
3.11 Limitations of Transcription-Based Drug Discovery for STATs Inhibitors
While most approaches to developing STAT inhibitors are based on inhibition of its transcriptional function, there are some limitations on relying on this approach. Although most of the known oncogenic properties of STATs are attributed to their roles as transcriptional factors, there is evidence that cytoplasmic [77] or mitochondrial STATs [63] can play important roles in malignant cell transformation and survival. It is conceivable that compounds that target these aspects of STAT function may not be discovered from transcription-based drug discovery methods. On the other hand, modifications of STATs that regulate their transcriptional function could also influence their cytoplasmic or mitochondrial localization.
Another potential caveat in transcription-based drug discovery is that STAT activation in these assays is generally induced by exogenous cytokine stimulation. Cytokine-induced STAT activation is transient, generally returning to baseline in 60–90 min. This differs from the continual activation seen in most tumor systems. In addition, the magnitude of the phosphorylation of STATs induced by cytokines, and the induction of transcription, is considerably greater in cytokine-induced systems than that seen with constitutive activation. Thus it is possible that compounds or genetic perturbations that modulate STAT transcriptional activity in a cytokine-induced system may not have the same activity in the setting of constitutively activated STATs as seen in cancer. Finally, it is clear that there are differences in STAT driven gene expression and STAT function that is dependent on the cellular context. Thus, compounds identified in a given system may not have uniform effects in other cells or tissues. Even within a given tumor type, unique aspects related to epigenetic states or the presence or absence of co-regulatory proteins may affect the activity of pharmacological modulators of STAT function. Nonetheless, the large amount of encouraging data generated in pre-clinical systems has generated a great interest in testing the approach of targeting STATs in human cancer.
3.12 Clinical Trials of STAT3 Inhibitors
Despite the large number of papers on developing and testing STAT inhibitors in model systems, relatively few true STAT inhibitors, i.e., compounds designed to specifically inhibit STAT function, have been introduced into clinical trials. This reflects a number of factors, including a relative lack of enthusiasm for targeting transcription factors among many in the field of cancer drug development, due to the pharmacologic challenges in inhibiting these proteins. Thus, for STAT inhibitors being introduced into clinical use, it is essential that appropriate pharmacodynamic markers be followed, to ensure that the target is, in fact, being inhibited. While this should be true for all targeted drug development efforts, it is particularly important for such a novel target as an inhibitor of an oncogenic transcription factor. Particularly in a Phase 1 trial in heavily pre-treated cancer patients, the chance of a large clinical response may be limited. In order to learn as much as possible from every patient who volunteers to participate in such a trial, it is important to first ask the question of whether the designated target is being inhibited. For a compound that blocks the activating tyrosine phosphorylation of STAT3, it can be relatively easy to monitor tyrosine phosphorylation by immunocytochemistry, immunofluorescence, or immunoblots. Where malignant cells and tissue can easily be obtained, as in hematological cancers or superficial lesions, this can be relatively straightforward. For other tumor types, it might be necessary to perform biopsies to obtain the necessary material. To minimize morbidity in patients with advanced cancer, one can also consider approaches such as examining circulating tumor cells to assess functional STAT activation.
For inhibitors that do not alter STAT3 phosphorylation, but inhibit the transcriptional response, it can be even more challenging to measure inhibition of STAT function. In those cases, one can evaluate the mRNA levels of STAT3 target gene signatures. Again, it may be necessary to perform relatively invasive biopsies to obtain adequate tissue, but the use of circulating tumor cells may make this more feasible.
Two clinical trials of true STAT3 inhibitors are particularly illustrative. The first, built on pioneering work from the laboratory of Jennifer Grandis, highlighted several key points [100]. The first is to use an inhibitor that has been tested extensively and rigorously in pre-clinical systems to ensure on target activity. While much work in developing STAT3 inhibitors is focused on inhibitors of the SH2 domain, these investigators used an approach based on blocking DNA binding of activated STAT3 dimers. They used a short double-stranded oligonucleotide that contained a canonical STAT3 binding site. They then were able to show that when this molecule was introduced into cancer cells with activated STAT3, it titrated the active STAT3 dimers away from the endogenous genomic sites to this “decoy”. After validating this approach in cell culture and animal studies, the investigators were then ready to test this approach in human cancer patients. The next key issue, in which physician investigators or collaborators are essential, was to determine the appropriate tumor type in which to test this strategy. These scientists chose squamous cell carcinoma of the head and neck, a disease in which constitutive STAT3 is common, and which is often accessible to direct visualization and injection. They performed a so-called “Phase 0” clinical trial (#NCT00696176), in which patients who were going to have their tumor resected had a single intratumoral injection of either the STAT3 decoy or saline control. No toxicity was noted from this therapy. When the tumor was resected 4–6 h later, assessment of expression of STAT3 regulated cyclin D1 and Bcl-xl were lower in the tumors treated with the STAT3 decoy than in the tumors treated with saline. Although this work is at an early stage, and these genes are regulated by a number of transcription factors, it represented a significant advance in actually translating STAT3 inhibitors from the laboratory to the clinic.
In contrast to this macromolecular approach to STAT3 inhibition, the first small molecule inhibitor of STAT3 to enter a clinical trial was based on a drug, pyrimethamine, that was identified from a chemical library screen for STAT3 inhibition. Pyrimethamine is an anti-microbial drug that is used clinically to treat malaria and toxoplasmosis. Pyrimethamine inhibits the transcriptional function of STAT3, but not that of other STAT family members or unrelated transcription factors like NF-kB [101, 102]. Furthermore, pyrimethamine exerts this effect at low micromolar concentrations, which are known to be readily achieved in human patients, and can safely be sustained for months on end. While pyrimethamine was very desirable from the standpoint of efficacy, specificity, and safety, it had one disadvantage. At the lower range of concentrations at which it inhibits STAT3 transcriptional function, it does not significantly reduce phosphorylation of STAT3. Thus, it seems that this drug acts through a relatively novel mechanism, likely involving disruption of co-activator complexes. However, this property of pyrimethamine would alter the way its activity would have to be monitored in a patient.
In considering a clinical trial with this drug, again it was important to focus on a cancer that was known to be dependent on activated STAT3 in a large majority of patients, to forestall the need to either test tumors prior to study entry or to enroll a large enough cohort so that an adequate number of patients with activated STAT3 were included. In addition, it was necessary to focus on a cancer in which it was easy to obtain sufficient tumor cells to perform pharmacodynamic evaluation of whether STAT3 function was definitively being inhibited. Since the phosphorylation of STAT3 was not affected, this analysis would have to rely on measurements of STAT3-dependent gene expression. The cancer chosen for this trial was chronic lymphocytic leukemia (CLL), and its essentially equivalent counterpart of small lymphocytic lymphoma (SLL). From a logistic standpoint, CLL has the advantage that most patients have a very large number of circulating malignant cells, so that assessment of pharmacodynamic endpoints can easily be achieved with a simple blood draw. CLL is characterized by essentially uniform phosphorylation of STAT3 in leukemic cells [71]. However, although the STAT3 is in the nucleus and transcriptionally active, it is phosphorylated on S727 rather than Y705. Nonetheless, since pyrimethamine could block the transcriptional function of STAT3 in CLL, and could decrease viability of CLL cells in vitro, this disease was chosen for a phase I/II clinical trial (#NCT01066663).
In this study, which is currently ongoing, patients are treated in cohorts of increasing daily doses of oral pyrimethamine. Trough concentrations of pyrimethamine are obtained in both the plasma and the white blood cell fraction (which contains the leukemic cells), so that effects on gene expression can be correlated with drug exposure. Not only are changes in STAT3 target genes determined from the cells taken immediately from the patient, but parallel in vitro experiments are performed on cells obtained from the patient prior to entry on the trial, to determine whether changes in gene expression and survival of the cells treated ex vivo with pyrimethamine match the clinical response.
Should this study show evidence of on-target effects, the integrated pharmacokinetic and pharmacodynamic data can then be used to guide trials in other diseases commonly driven by activated STAT3. If STAT3 inhibition is not occurring, then consideration needs to be given as to whether adequate drug concentrations and exposures over a 24 h time period are being achieved. For example, increased dose levels may need to be considered. If gene expression analyses show that STAT3 is adequately being inhibited, yet there is little clinical benefit, then one could consider combining a STAT3 inhibitor with another modality, including some of the conventional or novel targeted agents in use to treat this disease. For example, by decreasing expression of pro-survival genes like BCL-2 or BCL-xL, a STAT3 inhibitor like pyrimethamine might sensitize CLL cells to conventional cytotoxic drugs like fludarabine and cyclophosphamide, as well as novel kinase inhibitors like ibrutinib or idelalisib.
3.13 Conclusion
Although drug development in oncology had been dominated since its inception by cytotoxic drugs that non-specifically damage DNA or microtubules, or inhibited metabolic pathways, the field is now shifting to a new, more rational approach. Targeted molecular therapies first showed dramatic efficacy when specific kinases, activated by mutation, could be specifically inhibited. However, the targets are now broadening so-that non-mutated kinases that are oncogenic dependencies have become appealing targets. Finally, non-kinase targets, like the pro-survival protein BCL-2, are becoming tractable to pharmacologic intervention. One of the next frontiers in targeted molecular therapy for cancer is oncogenic transcription factors. While usually not directly mutated, these proteins are key convergence points from oncogenic signaling pathways. Since normal cells are generally tolerant of their inhibition, while cancer cells may be completely dependent on their function, transcription factors like STAT3 or STAT5 represent important targets with the potential of having a very high therapeutic index. While somewhat challenging from a medicinal chemistry standpoint, these high value targets can be inhibited using a number of creative strategies, and clinical trials of STAT inhibitors are currently under way. In the coming years, we will gain a better appreciation of the feasibility and potential of targeting STATs and other oncogenic transcription factors for the rational molecular therapy of cancer.
References
Darnell JE Jr (1997) STATs and gene regulation. Science 277:1630–1635
Frank DA (2003) STAT signaling in cancer: insights into pathogenesis and treatment strategies. Cancer Treat Res 115:267–291
Lin TS, Mahajan S, Frank DA (2000) STAT signaling in the pathogenesis and treatment of leukemias. Oncogene 19:2496–2504
Benekli M, Baumann H, Wetzler M (2009) Targeting signal transducer and activator of transcription signaling pathway in leukemias. J Clin Oncol 27:4422–4432
Bar-Natan M, Nelson EA, Walker SR et al (2012) Dual inhibition of Jak2 and STAT5 enhances killing of myeloproliferative neoplasia cells. Leukemia 26:1407–1410
Frank DA, Varticovski L (1996) BCR/abl leads to the constitutive activation of Stat proteins, and shares an epitope with tyrosine phosphorylated Stats. Leukemia 10:1724–1730
Carlesso N, Frank DA, Griffin JD (1996) Tyrosyl phosphorylation and DNA-binding activity of STAT proteins in hematopoietic cell lines transformed by Bcr/Abl. J Exp Med 183:811–820
Sternberg DW, Gilliland DG (2004) The role of signal transducer and activator of transcription factors in leukemogenesis. J Clin Oncol 22:361–371
Scheeren FA, Diehl SA, Smit LA et al (2008) IL-21 is expressed in Hodgkin lymphoma and activates STAT5: evidence that activated STAT5 is required for Hodgkin lymphomagenesis. Blood 111(9):4706–4715
Hoelbl A, Kovacic B, Kerenyi MA et al (2006) Clarifying the role of Stat5 in lymphoid development and Abelson-induced transformation. Blood 107:4898–4906
Walz C, Medrzycki M, Bunting ST et al (2012) Essential role for Stat5a/b in myeloproliferative neoplasms induced by BCR-ABL1 and Jak2V617F in mice. Blood 119:3550–3560
Crescenzo R, Abate F, Lasorsa E et al (2015) Convergent mutations and kinase fusions lead to oncogenic STAT3 activation in anaplastic large cell lymphoma. Cancer Cell 27:516–532
Mangolini M, De Boer J, Walf-Vorderwulbecke V et al (2013) STAT3 mediates oncogenic addiction to TEL-AML1 in t(12;21) acute lymphoblastic leukemia. Blood 122:542–549
Cui Y, Riedlinger G, Miyoshi K et al (2004) Inactivation of Stat5 in mouse mammary epithelium during pregnancy reveals distinct functions in cell proliferation, survival, and differentiation. Mol Cell Biol 24:8037–8047
Iavnilovitch E, Groner B, Barash I (2002) Overexpression and forced activation of stat5 in mammary gland of transgenic mice promotes cellular proliferation, enhances differentiation, and delays postlactational apoptosis. Mol Cancer Res 1:32–47
Liu X, Robinson GW, Gouilleux F, Groner B, Hennighausen L (1995) Cloning and expression of Stat5 and an additional homologue (Stat5b) involved in prolactin signal transduction in mouse mammary tissue. Proc Natl Acad Sci U S A 92:8831–8835
Tang JZ, Zuo ZH, Kong XJ et al (2010) Signal transducer and activator of transcription (STAT)-5A and STAT5B differentially regulate human mammary carcinoma cell behavior. Endocrinology 151:43–55
Ren S, Cai HR, Li M, Furth PA (2002) Loss of Stat5a delays mammary cancer progression in a mouse model. Oncogene 21:4335–4339
Iavnilovitch E, Cardiff RD, Groner B, Barash I (2004) Deregulation of Stat5 expression and activation causes mammary tumors in transgenic mice. Int J Cancer 112:607–619
Walker SR, Nelson EA, Yeh JE et al (2009) Reciprocal effects of STAT5 and STAT3 in breast cancer. Mol Cancer Res 7:966–976
Walker SR, Chaudhury M, Nelson EA, Frank DA (2010) Microtubule-targeted chemotherapeutic agents inhibit signal transducer and activator of transcription 3 (STAT3) signaling. Mol Pharmacol 78:903–908
Burke WM, Jin X, Lin HJ et al (2001) Inhibition of constitutively active Stat3 suppresses growth of human ovarian and breast cancer cells. Oncogene 20:7925–7934
Putoczki TL, Thiem S, Loving A et al (2013) Interleukin-11 is the dominant IL-6 family cytokine during gastrointestinal tumorigenesis and can be targeted therapeutically. Cancer Cell 24:257–271
Rokavec M, Oner MG, Li H et al (2014) IL-6R/STAT3/miR-34a feedback loop promotes EMT-mediated colorectal cancer invasion and metastasis. J Clin Invest 124:1853–1867
Errico A (2015) Lung cancer: driver-mutation-dependent stratification: learning from STAT3. Nat Rev Clin Oncol 12:251
Fan QW, Cheng CK, Gustafson WC et al (2013) EGFR phosphorylates tumor-derived EGFRvIII driving STAT3/5 and progression in glioblastoma. Cancer Cell 24:438–449
Baumgart S, Chen NM, Siveke JT et al (2014) Inflammation-induced NFATc1-STAT3 transcription complex promotes pancreatic cancer initiation by KrasG12D. Cancer Discov 4:688–701
Rodriguez-Barrueco R, Yu J, Saucedo-Cuevas LP et al (2015) Inhibition of the autocrine IL-6-JAK2-STAT3-calprotectin axis as targeted therapy for HR-/HER2+ breast cancers. Genes Dev 29:1631–1648
Heuser M, Sly LM, Argiropoulos B et al (2009) Modeling the functional heterogeneity of leukemia stem cells: role of STAT5 in leukemia stem cell self-renewal. Blood 114:3983–3993
Yoshimoto G, Miyamoto T, Jabbarzadeh-Tabrizi S et al (2009) FLT3-ITD up-regulates MCL-1 to promote survival of stem cells in acute myeloid leukemia via FLT3-ITD-specific STAT5 activation. Blood 114:5034–5043
Fatrai S, Wierenga AT, Daenen SM, Vellenga E, Schuringa JJ (2011) Identification of HIF2alpha as an important STAT5 target gene in human hematopoietic stem cells. Blood 117:3320–3330
Cook AM, Li L, Hoy Y et al (2014) Role of altered growth factor receptor-mediated JAK2 signaling in growth and maintenance of human acute myeloid leukemia stem cells. Blood 123:2826–2837
Gallipoli P, Cook A, Rhodes S et al (2014) JAK2/STAT5 inhibition by nilotinib with ruxolitinib contributes to the elimination of CML CD34+ cells in vitro and in vivo. Blood 124:1492–1501
Kim E, Kim M, Woo DH et al (2013) Phosphorylation of EZH2 activates STAT3 signaling via STAT3 methylation and promotes tumorigenicity of glioblastoma stem-like cells. Cancer Cell 23:839–852
Marotta LL, Almendro V, Marusyk A et al (2011) The JAK2/STAT3 signaling pathway is required for growth of CD44(+)CD24(−) stem cell-like breast cancer cells in human tumors. J Clin Invest 121:2723–2735
van der Zee M, Sacchetti A, Cansoy M et al (2015) IL6/JAK1/STAT3 Signaling blockade in endometrial cancer affects the ALDHhi/CD126+ stem-like component and reduces tumor burden. Cancer Res 75:3608–3622
Li Y, Guo G, Li L et al (2015) Suppression of cancer relapse and metastasis by inhibiting cancer stemness. Proc Natl Acad Sci U S A 112:1839–1844
Nishioka C, Ikezoe T, Yang J, Yokoyama A (2009) Multitargeted tyrosine kinase inhibitor stimulates expression of IL-6 and activates JAK2/STAT5 signaling in acute myelogenous leukemia cells. Leukemia 23:2304–2308
Warsch W, Kollmann K, Ecklehart E et al (2011) High STAT5 levels mediate imatinib resistance and indicate disease progression in chronic myeloid leukemia. Blood 117:3409–3420
Nishioka C, Ikezoe T, Yang J, Yokoyama A (2010) Long-term exposure of leukemia cells to multi-targeted tyrosine kinase inhibitor induces activations of AKT, ERK and STAT5 signaling via epigenetic silencing of the PTEN gene. Leukemia 24:1631–1640
Warsch W, Grunschober E, Berger A et al (2012) STAT5 triggers BCR-ABL1 mutation by mediating ROS production in chronic myeloid leukaemia. Oncotarget 3:1669–1687
Britschgi A, Andraos R, Brinkhaus H et al (2012) JAK2/STAT5 inhibition circumvents resistance to PI3K/mTOR blockade: a rationale for cotargeting these pathways in metastatic breast cancer. Cancer Cell 22:796–811
Vultur A, Villanueva J, Krepler C et al (2014) MEK inhibition affects STAT3 signaling and invasion in human melanoma cell lines. Oncogene 33:1850–1861
Liu F, Cao J, Wu J et al (2013) Stat3-targeted therapies overcome the acquired resistance to vemurafenib in melanomas. J Invest Dermatol 133:2041–2049
Girotti MR, Pedersen M, Sanchez-Laorden B et al (2013) Inhibiting EGF receptor or SRC family kinase signaling overcomes BRAF inhibitor resistance in melanoma. Cancer Discov 3:158–167
Eichten A, Su J, Adler A et al (2016) Resistance to anti-VEGF therapy mediated by autocrine IL-6/STAT3 signaling and overcome by IL-6 blockade. Cancer Res 76(8):2327–2339
Real PJ, Sierra A, De Juan A et al (2002) Resistance to chemotherapy via Stat3-dependent overexpression of Bcl-2 in metastatic breast cancer cells. Oncogene 21:7611–7618
Zhong H, Davis A, Ouzounova M et al (2016) A novel IL6 antibody sensitizes multiple tumor types to chemotherapy including trastuzumab-resistant tumors. Cancer Res 76:480–490
Lau J, Ilkhandizadeh S, Wang S et al (2015) STAT3 blockade inhibits radiation-induced malignant progression in glioma. Cancer Res 75:4302–4311
Wang T, Niu G, Kortylewski M et al (2004) Regulation of the innate and adaptive immune responses by Stat-3 signaling in tumor cells. Nat Med 10:48–54
Jones LM, Broz ML, Ranger JJ et al (2016) Stat3 establishes an immunosuppressive microenvironment during the early stages of breast carcinogenesis to promote tumor growth and metastasis. Cancer Res 76:1416–1428
Albesiano E, Davis M, See AP et al (2010) Immunologic consequences of signal transducers and activators of transcription 3 activation in human squamous cell carcinoma. Cancer Res 70:6467–6476
Yang H, Yamazaki T, Pietrocola F et al (2015) STAT3 inhibition enhances the therapeutic efficacy of immunogenic chemotherapy by stimulating type 1 interferon production by cancer cells. Cancer Res 75:3812–3822
Green MR, Monti S, Rodig SJ et al (2010) Integrative analysis reveals selective 9p24.1 amplification, increased PD-1 ligand expression, and further induction via JAK2 in nodular sclerosing Hodgkin lymphoma and primary mediastinal large B-cell lymphoma. Blood 116:3268–3277
Wang Z, Medrzycki M, Bunting ST, Bunting KD (2015) Stat5-deficient hematopoiesis is permissive for Myc-induced B-cell leukemogenesis. Oncotarget 6:28961–28972
Yan D, Jobe F, Hutchison RE, Mohi G (2015) Deletion of Stat3 enhances myeloid cell expansion and increases the severity of myeloproliferative neoplasms in Jak2V617F knock-in mice. Leukemia 29:2050–2061
Pencik J, Schlederer M, Gruber W et al (2015) STAT3 regulated ARF expression suppresses prostate cancer metastasis. Nat Commun 6:7736
Fukuda A, Wang SC, Morris JPT et al (2011) Stat3 and MMP7 contribute to pancreatic ductal adenocarcinoma initiation and progression. Cancer Cell 19:441–455
Lesina M, Kurkowski MU, Ludes K et al (2011) Stat3/Socs3 activation by IL-6 transsignaling promotes progression of pancreatic intraepithelial neoplasia and development of pancreatic cancer. Cancer Cell 19:456–469
Zhou J, Qu Z, Yan S et al (2015) Differential roles of STAT3 in the initiation and growth of lung cancer. Oncogene 34:3804–3814
Qu Z, Sun F, Zhou J et al (2015) Interleukin-6 prevents the initiation but enhances the progression of lung cancer. Cancer Res 75:3209–3215
Grabner B, Schramek D, Mueller KM et al (2015) Disruption of STAT3 signalling promotes KRAS-induced lung tumorigenesis. Nat Commun 6:6285
Gough DJ, Corlett A, Schlessinger K et al (2009) Mitochondrial STAT3 supports Ras-dependent oncogenic transformation. Science 324:1713–1716
Walker SR, Nelson EA, Frank DA (2007) STAT5 represses BCL6 expression by binding to a regulatory region frequently mutated in lymphomas. Oncogene 26:224–233
Walker SR, Nelson EA, Yeh JE et al (2013) STAT5 outcompetes STAT3 to regulate the expression of the oncogenic transcriptional modulator BCL6. Mol Cell Biol 33:2879–2890
Yue P, Lopez-Tapia F, Paladino D et al (2016) Hydroxamic acid and benzoic acid-based Stat3 inhibitors suppress human glioma and breast cancer phenotypes in vitro and in vivo. Cancer Res 76:652–663
Huang W, Dong Z, Chen Y et al (2016) Small-molecule inhibitors targeting the DNA-binding domain of STAT3 suppress tumor growth, metastasis and STAT3 target gene expression in vivo. Oncogene 35:783–792
Lamb J, Crawford ED, Peck D et al (2006) The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313:1929–1935
Alvarez JV, Febbo PG, Ramaswamy S et al (2005) Identification of a genetic signature of activated signal transducer and activator of transcription 3 in human tumors. Cancer Res 65:5054–5062
Yang J, Huang J, Dasgupta M et al (2010) Reversible methylation of promoter-bound STAT3 by histone-modifying enzymes. Proc Natl Acad Sci U S A 107:21499–21504
Frank DA, Mahajan S, Ritz J (1997) B lymphocytes from patients with chronic lymphocytic leukemia contain signal transducer and activator of transcription (STAT) 1 and STAT3 constitutively phosphorylated on serine residues. J Clin Invest 100:3140–3148
Hazan-Halevy I, Harris D, Liu Z et al (2010) STAT3 is constitutively phosphorylated on serine 727 residues, binds DNA, and activates transcription in CLL cells. Blood 115:2852–2863
Waitkus MS, Chandrasekharan UM, Willard B et al (2014) Signal integration and gene induction by a functionally distinct STAT3 phosphoform. Mol Cell Biol 34:1800–1811
Hou T, Ray S, Lee C, Brasier AR (2008) The STAT3 NH2-terminal domain stabilizes enhanceosome assembly by interacting with the p300 bromodomain. J Biol Chem 283:30725–30734
Mitra A, Ross JA, Rodriguez G et al (2012) Signal transducer and activator of transcription 5b (Stat5b) serine 193 is a novel cytokine-induced phospho-regulatory site that is constitutively activated in primary hematopoietic malignancies. J Biol Chem 287:16596–16608
Berger A, Hoelbl-Kovacic A, Bougeais J et al (2014) PAK-dependent STAT5 serine phosphorylation is required for BCR-ABL-induced leukemogenesis. Leukemia 28:629–641
Friedbichler K, Kerenyi MA, Kovacic B et al (2010) Stat5a serine 725 and 779 phosphorylation is a prerequisite for hematopoietic transformation. Blood 116:1548–1558
Gouilleux F, Wakao H, Mundt M, Groner B (1994) Prolactin induces phosphorylation of Tyr694 of Stat5 (MGF), a prerequisite for DNA binding and induction of transcription. EMBO J 13:4361–4369
Clark DE, Williams CC, Duplessis TT et al (2005) ERBB4/HER4 potentiates STAT5A transcriptional activity by regulating novel STAT5A serine phosphorylation events. J Biol Chem 280:24175–24180
Weaver AM, Silva CM (2007) S731 in the transactivation domain modulates STAT5b activity. Biochem Biophys Res Commun 362:1026–1030
Kloth MT, Catling AD, Silva CM (2002) Novel activation of STAT5b in response to epidermal growth factor. J Biol Chem 277:8693–8701
Cao R, Wang L, Wang H et al (2002) Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science 298:1039–1043
Dasgupta M, Dermawan JK, Willard B, Stark GR (2015) STAT3-driven transcription depends upon the dimethylation of K49 by EZH2. Proc Natl Acad Sci U S A 112:3985–3990
Nelson EA, Walker SR, Kepich A et al (2008) Nifuroxazide inhibits survival of multiple myeloma cells by directly inhibiting STAT3. Blood 112:5095–5102
Nelson EA, Walker SR, Weisberg E et al (2011) The STAT5 inhibitor pimozide decreases survival of chronic myelogenous leukemia cells resistant to kinase inhibitors. Blood 117:3421–3429
Nelson EA, Walker SR, Xiang M et al (2012) The STAT5 inhibitor pimozide displays efficacy in models of acute myelogenous leukemia driven by FLT3 mutations. Genes Cancer 3:503–511
Cerliani JP, Guillardoy T, Giulianelli S et al (2011) Interaction between FGFR-2, STAT5, and progesterone receptors in breast cancer. Cancer Res 71:3720–3731
Bertucci PY, Quaglino A, Pozzi AG, Kordon EC, Pecci A (2010) Glucocorticoid-induced impairment of mammary gland involution is associated with STAT5 and STAT3 signaling modulation. Endocrinology 151:5730–5740
Subtil-Rodriguez A, Millan-Arino L, Quiles I et al (2008) Progesterone induction of the 11beta-hydroxysteroid dehydrogenase type 2 promoter in breast cancer cells involves coordinated recruitment of STAT5A and progesterone receptor to a distal enhancer and polymerase tracking. Mol Cell Biol 28:3830–3849
Carsol JL, Gingras S, Simard J (2002) Synergistic action of prolactin (PRL) and androgen on PRL-inducible protein gene expression in human breast cancer cells: a unique model for functional cooperation between signal transducer and activator of transcription-5 and androgen receptor. Mol Endocrinol 16:1696–1710
Richer JK, Lange CA, Manning NG et al (1998) Convergence of progesterone with growth factor and cytokine signaling in breast cancer. Progesterone receptors regulate signal transducers and activators of transcription expression and activity. J Biol Chem 273:31317–31326
Wyszomierski SL, Yeh J, Rosen JM (1999) Glucocorticoid receptor/signal transducer and activator of transcription 5 (STAT5) interactions enhance STAT5 activation by prolonging STAT5 DNA binding and tyrosine phosphorylation. Mol Endocrinol 13:330–343
Ho L, Miller EL, Ronan JL et al (2011) esBAF facilitates pluripotency by conditioning the genome for LIF/STAT3 signalling and by regulating polycomb function. Nat Cell Biol 13:903–913
Yeh JE, Kreimer S, Walker SR et al (2015) Granulin, a novel STAT3-interacting protein, enhances STAT3 transcriptional function and correlates with poorer prognosis in breast cancer. Genes Cancer 6:153–168
Lu R, Serrero G (2001) Mediation of estrogen mitogenic effect in human breast cancer MCF-7 cells by PC-cell-derived growth factor (PCDGF/granulin precursor). Proc Natl Acad Sci U S A 98:142–147
Hoque M, Young TM, Lee CG et al (2003) The growth factor granulin interacts with cyclin T1 and modulates P-TEFb-dependent transcription. Mol Cell Biol 23:1688–1702
Dawson MA, Prinjha RK, Dittmann A et al (2011) Inhibition of BET recruitment to chromatin as an effective treatment for MLL-fusion leukaemia. Nature 478:529–533
Dawson MA, Kouzarides T, Huntly BJ (2012) Targeting epigenetic readers in cancer. N Engl J Med 367:647–657
Liu S, Walker SR, Nelson EA et al (2014) Targeting STAT5 in hematologic malignancies through inhibition of the bromodomain and extra-terminal (BET) bromodomain protein BRD2. Mol Cancer Ther 13:1194–1205
Sen M, Grandis JR (2012) Nucleic acid-based approaches to STAT inhibition. JAKSTAT 1:285–291
Takakura A, Nelson EA, Haque N et al (2011) Pyrimethamine inhibits adult polycystic kidney disease by modulating STAT signaling pathways. Hum Mol Genet 20:4143–4154
Nelson EA, Sharma SV, Settleman J, Frank DA (2011) A chemical biology approach to developing STAT inhibitors: molecular strategies for accelerating clinical translation. Oncotarget 2:518–524
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Liu, S., Frank, D. (2016). Translating STAT Inhibitors from the Lab to the Clinic. In: Ward, A. (eds) STAT Inhibitors in Cancer. Cancer Drug Discovery and Development. Humana Press, Cham. https://doi.org/10.1007/978-3-319-42949-6_3
Download citation
DOI: https://doi.org/10.1007/978-3-319-42949-6_3
Published:
Publisher Name: Humana Press, Cham
Print ISBN: 978-3-319-42947-2
Online ISBN: 978-3-319-42949-6
eBook Packages: MedicineMedicine (R0)