Abstract
The vast majority of the human genome is transcribed into RNA molecules that do not code for proteins, which could be small ones approximately 20 nucleotide in length, known as microRNAs, or transcripts longer than 200 bp, defined as long noncoding RNAs. The prevalent deregulation of microRNAs in human cancers prompted immediate interest on the therapeutic value of microRNAs as drugs and drug targets. Many features of microRNAs such as well-defined mechanisms, and straightforward oligonucleotide design further make them attractive candidates for therapeutic development. The intensive efforts of exploring microRNA therapeutics are reflected by the large body of preclinical studies using oligonucleotide-based mimicking and blocking, culminated by the recent entry of microRNA therapeutics in clinical trial for several human diseases including cancer. Meanwhile, microRNA therapeutics faces the challenge of effective and safe delivery of nucleic acid therapeutics into the target site. Various chemical modifications of nucleic acids and delivery systems have been developed to increase targeting specificity and efficacy, and reduce the associated side effects including activation of immune response. Recently, long noncoding RNAs become attractive targets for therapeutic intervention because of their association with complex and delicate phenotypes, and their unconventional pharmaceutical activities such as capacity of increasing output of proteins. Here I discuss the general therapeutic strategies targeting noncoding RNAs, review delivery systems developed to maximize noncoding RNA therapeutic efficacy, and offer perspectives on the future development of noncoding RNA targeting agents for colorectal cancer.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Noncoding RNAs are useful targets for therapeutic interventions of human cancer. Here we focus on the potential of microRNAs and long noncoding RNAs as therapeutic targets and tools in treatment of colorectal cancer. MicroRNAs have the advantages of targeting multiple protein-coding genes at once. In addition, microRNAs have their function in the physiological conditions, and thus restoring or reducing microRNA to their normal levels may lead to favorable consequences. Furthermore, microRNAs may be used to target proteins that are difficult to design small molecular chemical inhibitors. Another advantages of microRNAs are their relative simple structures, and their predictable mechanisms. These features made the design of mimics or anti-miRs easier than that of the conventional chemical drugs.
The levels of long noncoding RNAs are usually lower than those of the protein coding genes [1]. However, long noncoding RNAs tend to have more tissue specific expression pattern than the protein coding genes, and are thus possibly associated with certain cancer subtypes [1]. The fact that many long noncoding RNAs are identified from important cancer associated genomic locus suggests that they should be functionally important and relevant. Disruption of these transcripts, as demonstrated by previous studies, could lead to significant consequences in the biological activities and disease status. The theoretically specificity and efficacy of small interference RNA and antisense oligonucleotides in reducing levels of a long noncoding RNA, made readily available the means for manipulating such transcripts. Additionally, since long noncoding RNA could interact with proteins such as transcription factors and histone modifiers, targeting long noncoding RNA will lead to specific and delicate changes, which may be desirable in the cancer treatment [2].
2 Therapeutic Strategies Targeting Noncoding RNAs
Generally, there are two strategies for therapeutic targeting of noncoding RNAs in colorectal cancer (Fig. 12.1). The first is to restore the function of noncoding RNAs with tumor suppressor activities that are lost in colorectal cancer. The second is to block the actions of noncoding RNAs with oncogenic function that are aberrantly overexpressed in colorectal cancer. Both strategies could be applied to microRNAs . Since microRNAs are with small size, and often localized in the cytosolic subcellular fraction, it is possible to restore microRNA function with synthetic microRNA mimics . The function of microRNAs could also be blocked by a variety of strategies interfering with microRNA activities. For long noncoding RNAs , blocking their function is more plausible than restoring biological activities of such transcripts, because of several reasons. First, unlike microRNAs, long noncoding RNAs could fold into secondary and higher order structures, and its function is hard to predict with the sequence information [3]. This makes it difficult to synthesize long noncoding RNA to replace their original function. Second, many long noncoding RNAs are localized in the cell nucleus, thus the restoration of their function requires one more step of synthesized molecules entering the nucleus. Third, similar as mRNAs, long noncoding RNAs are more easily degraded than microRNAs , creating one more barrier for replacement strategy for long noncoding RNAs. Lastly, the mechanisms underlying long noncoding RNAs are not well elucidated. If a long noncoding RNA functions via cis regulatory mechanisms, it is impossible to restore its function without expressing at the specific genomic locus. However, blocking of the function of long noncoding RNAs could be easily achieved by several strategies, with the most straightforward approach of downregulation with RNA interference. The approaches discussed below were summarized in Table 12.1.
2.1 Restoring Noncoding RNA Function
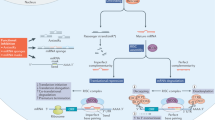
To regain the function of a microRNA that is lost or downregulated in colorectal cancer, the simplest method is to supply with synthetic microRNA molecules with same function. This could be achieved with microRNA mimics or with microRNAs expression vectors [2]. microRNA mimics are chemically modified double stranded RNAs that mimic endogenous microRNAs [4]. When transfected into cells, microRNA mimics could be processed into single-strand microRNA molecule to target coding genes similar as the endogenous microRNA . An alternative way of replacing a microRNA function is to produce it in an expression vector. With the microRNA production machinery engineered, the designed vector could produce continuously the intended microRNA molecules for replacement. In addition, the microRNA expression vectors can be engineered with promoters to specifically express microRNA in a tumor- and tissue- specific manner, giving this expression vector method an advantage over microRNA mimics.
The loss of microRNA expression could be due to genomic deletion or epigenetic silencing. In the latter scenario, it is possible to recover a microRNA expression by reversing the epigenetic quenching. Decitabine and 5-azacytidine, two hypomethylating agents that have been approved for treatment of myelodysplastic syndromes, were shown to re-induce the expression of several miRNAs including miR-124a [5]. However, this regulation is non-specific to one microRNA . In addition, the spectrum of induced microRNAs is context dependent [5–7]. The antibacterial compound enoxacin has also been shown to boost the expression of a subset of microRNAs in colon cancer cell lines in vitro and in vivo by acting on TARBP2, a protein regulating microRNA processing [8].
Among these replacement strategies, microRNA mimics gain popularity in the development into therapeutic agents by biopharmaceutical companies. This can be reflected by the first miRNA replacement therapy entering clinical trial for treatment of human cancer - formulated miR-34a mimics for treatment of patients with advanced hepatocellular carcinoma [9].
2.2 Antagonizing Noncoding RNA Function
Current strategies to antagonize microRNA function include locked nucleic acids (LNA anti-miRs ), tiny LNA anti-miRs, antagomirs , and miRNA sponges [2]. LNA anti-miRs are antisense oligonucleotides with several nucleotides substituted by bicyclic RNA analogues in a ‘locked’ conformation [10]. This LNA modification renders high affinity for the binding of the targeted microRNA by generating an ideal conformation for Watson–Crick binding, and allows for effective blockade of microRNA function with short sequences (13–22 nucleotides). Additionally, LNA anti-miRs are resistant to degradation, and efficient in uptake by many tissues. These features of LNA anti-miRs eliminate the need for sophisticated formulation and delivery, which is indispensible for most other antagonizing strategies. The exploration of LNA anti-miRs for clinical usages culminated with the entering of miravirsen (SPC3649; Santaris Pharma), an LNA anti-miR against miR-122 , in Phase I and Phase IIa clinical trials for the treatment of hepatitis C virus (HCV) [11, 12]. Since 5′-seed region at positions 2–7 of a microRNA is essential for the binding of microRNA to its mRNA targets, tiny LNA anti-miRs specifically designed to target the microRNA seed region were explored [13]. Tiny LNA anti-miRs have the advantage of targeting multiple microRNAs within the same family; however, the specificity of microRNA targeting was compromised [13].
Antagomirs are synthetic, cholesterol-conjugated RNAs complementary to the targeted microRNA sequence, featured by a 2′-O-methyl linkage and phosphorothioate modification [14]. These added features help to increase cellular uptake and protect from degradation by nucleases [14]. While antagomirs have been shown to block microRNA function in mouse models, their uses are currently limited to experimental tools, probably due to high effective dosages associated with antagomirs [10].
Another strategy of blocking microRNA function is to generate microRNA sponges to competitively inhibit microRNA function [15]. These microRNA sponges contain multiple tandem bindings that are complementary to the microRNA seed sequence . By sequestering aimed microRNAs from their endogenous mRNA targets, this microRNA sponge method effectively blocks the microRNA function [15]. To achieve enough concentration of sponge RNAs, expression vectors with strong promoter are usually used to maintain high level of transcription. Several studies showed that microRNA sponges tend to have long-lasting effect [16]; however, because sponges are RNAs without chemical modification, the concentrations for effective inhibition of microRNA function may be much higher than other anti-miRs. Furthermore, whether the excess of sponge transcripts produce undesired effects remains to be determined by further studies.
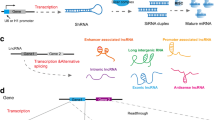
The therapeutic exploration of long noncoding RNAs lags far behind the microRNA therapeutics. The function of long noncoding RNAs could be blocked by several strategies. First, the level of long noncoding RNA could be regulated by specifically designed siRNAs . The length of long noncoding RNA also makes the design of specific siRNAs not a difficult task. Previous studies have shown that siRNAs could successfully achieve knockdown of long noncoding RNAs, irrespective of their subcellular localization [17]. Considering the fact that many long noncoding RNA are upregulated in colorectal cancer, the use of siRNAs targeting such oncogenic long noncoding RNA could probably reverse the cancer malignancies. On the other hand, many protein-coding genes have corresponding natural antisense transcripts, which could negatively regulate expression of these protein-coding genes [18]. Therefore, targeting of natural antisense transcripts by single-stranded oligonucleotides represents a unique opportunity for therapeutic upregulation of tumor suppressor genes, which is difficult to realize with the conventional drug design of chemical compounds.
3 Delivery Systems
In almost all of the strategies of noncoding RNA therapeutics, safe and effective delivery of the oligonucleotides into the cancer tissue without causing deleterious side effects remains the premier challenges. Unmodified oligonucleotides are not stable in the circulation, can be attacked by immune system, and hardly penetrate into cells. Although modifications as discussed above could increase affinity to targets, and increase the stability, most of the oligonucleotide therapies need additional optimal delivery system to achieve the desired biological effects. Several aspects need to be considered when selecting a delivery system: stability against serum nucleases, evasion of the innate immune system, avoidance of non-specific interactions with serum proteins and non-target cells, prevention of renal clearance, release from blood vessels to target tissues, cell entry, incorporation into the RNA interference or other machinery [19].
Shielding the exterior of delivery vehicles with polyethylene glycol (PEG) is a common strategy to increase the circulation time for therapeutic oligonucleotides [20]. This strategy could prevent non-specific interaction of formulated particles with serum proteins, immune cells and other non-target tissues [20]. Particles with size of 8 nm to 20 nm in the circulation are subject to renal clearance, with the exception of dynamic polyconjugates (DPCs) and triantennary N-acetylgalactosamine (GalNAc) conjugates [19]. These two conjugates therefore could offer advantage of avoiding elimination of formulated particles by urine. To take effect on the target site, formulated particles need to release from the circulation into the aimed cancer tissues. Many solid tumors including colorectal cancer have discontinuous endothelia, and thus are more prone to permeation than the normal tissues [21]. Together with impaired lymphatic drainage in cancer, tumor tissues could accumulate more circulating particles.
Once reaching the tumor site, the delivery particles usually enter the cells via endocytosis. To facilitate such process, the delivery system could be engineered with targeting ligands that specifically recognize receptors on target cells. Alternatively, cell-penetrating peptides could increase the cellular uptake [22]. Tumors are characterized by acidic environment partially because of lack of nutrition and metabolic changes induced by Warburg effect [23]. The acidic environment of tumors offers opportunity to incorporate materials that can be released in the low pH environment. A recent study by the Slack group has developed a delivery system attaching antimiRs to a peptide with low pH-inducible transmembrane structure, and demonstrated the success of this system in blocking miR-155 function in a mouse model of lymphoma [23].
Lipid nanoparticles (LNPs) such as liposomes have been developed to protect oligonucleotides from nuclease degradation, avoid renal clearance, increase cellular uptake, and promote endosomal escape [24]. Several LNP RNAi drugs have passed the preclinical evaluation and entered clinical trials [25]. One example is the LNP drug ALN-VSP, a lipid delivery system developed by Alnylam Pharmaceuticals, which was recently evaluated in phase-I clinical trial for treatment of advanced solid tumors [26]. This study found that ALN-VSP successfully degraded target mRNA in tissue biopsies to exert antitumor activity at dosages well tolerated by patients [26]. As the first anticancer microRNA drug entering clinical trial, the miR-34 mimic MRX34 developed by Mirna Therapeutics is also liposome-based [9].
It should be noted that the delivery systems showing success in vivo vary largely in size, structure, and chemistry. For each specific case, unique designs of delivery system might be necessary to achieve best efficacy without causing deleterious side effect. LNPs are among the most effective formulations in the delivery of oligonucleotides for noncoding RNA therapy. Conjugate systems, which require minimal amounts of delivery material, have the advantage of defined molecular structures, and wide therapeutic window, also show promise as an effective delivery system [19].
4 Challenges of Noncoding RNA Therapeutics
Noncoding RNA therapeutics is a new concept that differs from the conventional chemical drug design. Numerous challenges exist for the therapeutic use of noncoding RNAs in the treatment of colorectal cancer. For instance, while the fact that microRNAs target multiple mRNAs can be an advantage itself, this also cause ambiguity as to the scope of genes that are exactly controlled by microRNAs. Making this even more complicated, studies show that microRNA functions are fine-tuned and context-dependent [27]. The microRNA targets identified by the cell model system or animal models may not be applicable to the clinical scenarios. To serve as a candidate for clinical evaluations, the functional phenotype and mechanisms of a microRNA need to be well elucidated and validated in the most stringent way. For long noncoding RNAs, the challenges are even bigger. The functioning mechanisms of long noncoding RNAs are not well understood, and general principles governing the functioning mechanisms are missing. In addition, long noncoding RNAs are more tissue-specific than protein-coding genes. This adds further challenges in targeting noncoding RNAs in the specific tissue or subcellular compartments. Detailed understanding of the biology and functioning mechanisms holds the key for translation of such knowledge into clinical usages.
For the noncoding RNAs with well-defined activity and functioning mechanisms, the biggest challenges lie in the delivery system. Even the most advanced formulations do not solve the technical requirement for a clinically useful drug. For instance, the manufacturing production of nanoparticles needs a better controlled mixing processes to achieve consistent quality [28]. In addition, the mechanisms underlying the delivery process are not well elucidated, and the established formulation guidelines may not always lead to expected biological phenomenon. Most of the oligonucleotide delivery systems are for well-perfused tissues such as liver, which physiologically allows for the distribution of therapeutic particles into target tissues. Novel delivery systems need to be developed for targeting colorectal cancer. Considering the importance of cancer stem cells in the initial and progression of colorectal cancer, it can be conceived that conjugated ligand specifically recognizing colon cancer stem cells could be used for delivery of therapeutic materials to destroy cancer stem cells. Recent studies showed that microRNAs could be packaged into multivesicular bodies and released into the extracellular environment as exosomes [29]. This represents a natural delivery system and may offer more advantages than the synthetic delivery systems. The detailed understanding of exosome microRNAs in colorectal cancer progression, metastasis , and drug response might offer novel strategies for cancer treatment, and aid the design of more efficient tumor specific delivery systems.
5 Conclusions and Summary
Noncoding RNA therapeutics for colorectal cancer is still in its infancy. Nonetheless, the field of noncoding RNA therapeutics is developing fast. Just two decades after the initial discovery of microRNA link with human cancer in 2002, MRX34 entered clinical trials for treating advanced hepatocarcinoma. Both academia and pharmaceutical companies have been enthusiastically pursing the therapeutic value of noncoding RNAs. Companies such as Regulus Therapeutics and Mirna Therapeutics have developed pipelines for microRNA therapeutics in treating diseases including cancer. In addition, companies such as RaNA Therapeutics are exploring the therapeutic potential of long noncoding RNAs . With the experience gained from developing oligonucleotides-based therapeutics, many obstacles that noncoding RNA therapeutics face might be cleared. Colorectal cancer is characterized by genetic alterations; noncoding RNAs including microRNAs and long noncoding RNAs have pivotal role in the regulation of these genetic events. We believe that with improved understanding of noncoding RNA biology and delivery system innovation, we will see in the near future the utility of noncoding RNA in the treatment of patients with colorectal cancer, in combination with chemotherapy and radiotherapy.
References
Rosenbloom KR, Dreszer TR, Long JC, Malladi VS, Sloan CA, Raney BJ, et al. ENCODE whole-genome data in the UCSC Genome Browser: update 2012. Nucleic Acids Res. 2012;40(Database issue):D912–7. Epub 2011/11/15.
Ling H, Fabbri M, Calin GA. MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nat Rev Drug Discov. 2013;12(11):847–65. Epub 2013/11/01.
Wan Y, Qu K, Zhang QC, Flynn RA, Manor O, Ouyang Z, et al. Landscape and variation of RNA secondary structure across the human transcriptome. Nature. 2014;505(7485):706–9. Epub 2014/01/31.
Bader AG, Brown D, Stoudemire J, Lammers P. Developing therapeutic microRNAs for cancer. Gene Ther. 2011;18(12):1121–6. Epub 2011/06/03.
Lujambio A, Ropero S, Ballestar E, Fraga MF, Cerrato C, Setien F, et al. Genetic unmasking of an epigenetically silenced microRNA in human cancer cells. Cancer Res. 2007;67(4):1424–9. Epub 2007/02/20.
Bandres E, Agirre X, Bitarte N, Ramirez N, Zarate R, Roman-Gomez J, et al. Epigenetic regulation of microRNA expression in colorectal cancer. Int J Cancer. 2009;125(11):2737–43. Epub 2009/06/13.
Saito Y, Jones PA. Epigenetic activation of tumor suppressor microRNAs in human cancer cells. Cell Cycle. 2006;5(19):2220–2. Epub 2006/10/03.
Melo S, Villanueva A, Moutinho C, Davalos V, Spizzo R, Ivan C, et al. Small molecule enoxacin is a cancer-specific growth inhibitor that acts by enhancing TAR RNA-binding protein 2-mediated microRNA processing. Proc Natl Acad Sci U S A. 2011;108(11):4394–9. Epub 2011/03/04.
Bouchie A. First microRNA mimic enters clinic. Nat Biotechnol. 2013;31(7):577. Epub 2013/07/11.
Lennox KA, Behlke MA. Chemical modification and design of anti-miRNA oligonucleotides. Gene Ther. 2011;18(12):1111–20. Epub 2011/07/15.
Janssen HL, Reesink HW, Lawitz EJ, Zeuzem S, Rodriguez-Torres M, Patel K, et al. Treatment of HCV infection by targeting microRNA. N Engl J Med. 2013;368(18):1685–94. Epub 2013/03/29.
Lieberman J, Sarnow P. Micromanaging hepatitis C virus. N Engl J Med. 2013;368(18):1741–3. Epub 2013/03/29.
Obad S, dos Santos CO, Petri A, Heidenblad M, Broom O, Ruse C, et al. Silencing of microRNA families by seed-targeting tiny LNAs. Nat Genet. 2011;43(4):371–8. Epub 2011/03/23.
Krutzfeldt J, Rajewsky N, Braich R, Rajeev KG, Tuschl T, Manoharan M, et al. Silencing of microRNAs in vivo with ‘antagomirs’. Nature. 2005;438(7068):685–9. Epub 2005/11/01.
Ebert MS, Neilson JR, Sharp PA. MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells. Nat Methods. 2007;4(9):721–6. Epub 2007/08/19.
Xie J, Ameres SL, Friedline R, Hung JH, Zhang Y, Xie Q, et al. Long-term, efficient inhibition of microRNA function in mice using rAAV vectors. Nat Methods. 2012;9(4):403–9. Epub 2012/03/06.
Khalil AM, Guttman M, Huarte M, Garber M, Raj A, Rivea Morales D, et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc Natl Acad Sci U S A. 2009;106(28):11667–72. Epub 2009/07/03.
Wahlestedt C. Targeting long non-coding RNA to therapeutically upregulate gene expression. Nat Rev Drug Discov. 2013;12(6):433–46. Epub 2013/06/01.
Kanasty R, Dorkin JR, Vegas A, Anderson D. Delivery materials for siRNA therapeutics. Nat Mater. 2013;12(11):967–77. Epub 2013/10/24.
Alexis F, Pridgen E, Molnar LK, Farokhzad OC. Factors affecting the clearance and biodistribution of polymeric nanoparticles. Mol Pharm. 2008;5(4):505–15. Epub 2008/08/05.
Kanasty RL, Whitehead KA, Vegas AJ, Anderson DG. Action and reaction: the biological response to siRNA and its delivery vehicles. Mol Ther. 2012;20(3):513–24. Epub 2012/01/19.
Bolhassani A. Potential efficacy of cell-penetrating peptides for nucleic acid and drug delivery in cancer. Biochim Biophys Acta. 2011;1816(2):232–46. Epub 2011/08/16.
Cheng CJ, Bahal R, Babar IA, Pincus Z, Barrera F, Liu C, et al. MicroRNA silencing for cancer therapy targeted to the tumour microenvironment. Nature. 2015;518(7537):107–10. Epub 2014/11/20.
Alabi C, Vegas A, Anderson D. Attacking the genome: emerging siRNA nanocarriers from concept to clinic. Curr Opin Pharmacol. 2012;12(4):427–33. Epub 2012/06/26.
Burnett JC, Rossi JJ, Tiemann K. Current progress of siRNA/shRNA therapeutics in clinical trials. Biotechnol J. 2011;6(9):1130–46. Epub 2011/07/12.
Tabernero J, Shapiro GI, LoRusso PM, Cervantes A, Schwartz GK, Weiss GJ, et al. First-in-humans trial of an RNA interference therapeutic targeting VEGF and KSP in cancer patients with liver involvement. Cancer Discov. 2013;3(4):406–17. Epub 2013/01/30.
Dvinge H, Git A, Graf S, Salmon-Divon M, Curtis C, Sottoriva A, et al. The shaping and functional consequences of the microRNA landscape in breast cancer. Nature. 2013;497(7449):378–82. Epub 2013/05/07.
Gindy ME, Leone AM, Cunningham JJ. Challenges in the pharmaceutical development of lipid-based short interfering ribonucleic acid therapeutics. Expert Opin Drug Deliv. 2012;9(2):171–82. Epub 2012/01/19.
Melo SA, Sugimoto H, O’Connell JT, Kato N, Villanueva A, Vidal A, et al. Cancer exosomes perform cell-independent microRNA biogenesis and promote tumorigenesis. Cancer Cell. 2014;26(5):707–21. Epub 2014/12/03.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Ling, H. (2016). Non-coding RNAs: Therapeutic Strategies and Delivery Systems. In: Slaby, O., Calin, G. (eds) Non-coding RNAs in Colorectal Cancer. Advances in Experimental Medicine and Biology, vol 937. Springer, Cham. https://doi.org/10.1007/978-3-319-42059-2_12
Download citation
DOI: https://doi.org/10.1007/978-3-319-42059-2_12
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-42057-8
Online ISBN: 978-3-319-42059-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)