Abstract
This chapter describes a workflow for measuring a barcode’s accuracy when identifying species. First, assemble a database of specimens with their marker sequences and their species binomials. The species binomials provide a “taxonomic gold standard” for species identification and should be as accurate as possible, to avoid penalizing correct species assignment. Second, select a computer algorithm for assigning species to barcode sequences. Only one algorithm (BLAST+P) has improved notably on the simple strategy of assigning specimens to the species of the database sequence(s) nearest under p-distance. Global sequence alignments (e.g., with the Needleman-Wunsch algorithm, or with multiple sequence alignment algorithms) align entire barcode sequences, using all available information, so they sometimes produce more accurate species identifications than local sequence alignments (e.g., with BLAST), particularly when BLAST produces barcode alignments of small subsequences within the sequences. Finally, consensus has settled on “the probability of correct identification” (PCI) as the appropriate measurement of species identification accuracy. The overall PCI for a data set is the average of the species PCIs, taken over all species in the data set. The chapter discusses some variant PCIs, their calculation and the estimation of their statistical sampling errors. It also discusses good practice in incorporating PCR failure and species with singleton representatives into data summaries. For software relevant to this chapter, see http://tinyurl.com/spouge-barcode.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
The Anthropocene era is the geologic era when human activities started to have a significant effect on the Earth’s ecosystems (Zalasiewicz et al. 2000). The Anthropocene is extinguishing species, making the conservation of biodiversity a major challenge. The first step to debate the benefits and costs of preserving biodiversity is to assess the facts, namely, to catalog species throughout the world, and to document changes in their populations, a staggering task requiring more taxonomists than presently exist. Fortunately, the technology of DNA barcodes provides an alternative strategy to field assessment by trained taxonomists, because barcodes require DNA samples and not the immediate identification of specimens. Barcodes have many uses other than taxonomic identification (recognition of novel species, taxonomic classification, and the construction of phylogenetic trees, to name but a few), but applications in biodiversity justify restricting this chapter to the measurement of accuracy in species identification (Hebert et al. 2003a). The ability to measure barcode accuracy when identifying species within a taxon is critical, for it measures the success of the barcode enterprise within that taxon.
In its essence, a barcode is any standardized subset of DNA collected from taxonomic specimens (Floyd et al. 2002). To fix the terminology in the chapter, “marker” connotes any contiguous region of DNA (coding or non-coding), whereas “barcode” is a collective noun connoting the one or more markers comprising the standardized subset of DNA. Presently, all official barcode markers are genes, e.g., CO1 (historically, the first official barcode), matK and rbcL (the barcode markers for plants), and ITS (the barcode marker for fungi). Initial investigations into DNA barcodes, particularly in animals, indicated that when used as a barcode, the cytochrome c oxidase 1 (CO1) gene identified many species correctly (Hebert et al. 2003b), so selection of CO1 as the primary barcode followed naturally (Hajibabaei et al. 2006; Hogg and Hebert 2004; Lorenz et al. 2005; Meyer and Paulay 2005; Saunders 2005; Smith et al. 2005, 2006).
The extension of DNA barcodes to plants (Chase et al. 2005; Cowan et al. 2006; Kress and Erickson 2008; CBOL Plant Working Group 2009) and fungi (Schoch et al. 2012) became problematic with the recognition that the CO1 gene evolved too slowly to identify the corresponding species accurately (Kress et al. 2005). Likewise, CO1 often does not identify insect species accurately (Meier et al. 2006; Huang et al. 2008). The lack of a clear consensus for a barcode in these taxa stimulated interest in an objective, quantitative measurement of barcode accuracy in identifying species. Although consensus about barcodes themselves remains elusive in some taxa, notably plants (Pang et al. 2012; Chen et al. 2015; Ferri et al. 2015), a clear consensus on measuring barcode accuracy has emerged (CBOL Plant Working Group 2009; Meier et al. 2006; Erickson et al. 2008). As a byproduct, the consensus provides a clear standard for measuring barcode accuracy in specific taxa, as well.
Section 2 describes the rationale behind and the methods of evaluating identification accuracy, while Sect. 7 provides a practical chapter summary. The web page http://tinyurl.com/spouge-barcode also provides some practical tools for measuring barcode identification accuracy.
2 Measurement of Species Identification Accuracy
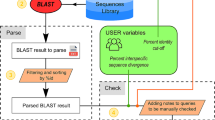
As its ultimate aim, any measurement of the accuracy of species identification should reflect the practical performance of the corresponding barcode database. A barcode database has the following workflow. Users sample a DNA barcode retrieved from a specimen and then query the database with the sample barcode sequence(s); in response, the database software returns a putative species identification. In principle, a species apparently not in the database might evoke the output of “unknown species”. Sometimes species boundaries are said to correlate with DNA differences like 2 % (i.e., p-distance of 0.98), but in practice the boundary difference varies with the species. Incomplete lineage sorting can even make the difference vanish. Different attempts to redefine species boundaries with threshold differences (Hebert et al. 2003a; Floyd et al. 2002; Blaxter et al. 2005; Lambert et al. 2005) conflicted heavily with traditional taxonomy (Meier et al. 2006), so no consensus exists on criteria for defining species boundaries with DNA barcodes. Thus, the database output “unknown species” (i.e., “outside every known species boundary”) becomes problematic, so for pedagogical clarity, this chapter restricts itself to discussing the identification of known species, i.e., it assumes that every sample for querying the barcode database represents a species already in the database.
3 The Barcode Database
The first step in estimating barcode accuracy is to assemble a database of specimens with their marker sequences and their species binomials. In particular, the species binomials provide a “taxonomic gold standard” for species identification, i.e., they represent a standard for measuring the accuracy of algorithms implemented on a computer. The standard is often imperfect, but it should be the best available.
An investigator should recognize potential defects in a dataset (Spouge and Mariño-Ramírez 2012). To exemplify possible imperfections, the taxonomy in GenBank sequences (unless annotated by the “barcode” keyword) can be noticeably inferior to the taxonomy in databases curated by taxonomists. Imperfect taxonomy degrades measurements of barcode accuracy by penalizing correct species assignments. Moreover, GenBank usually does not provide the original specimen with its entries, rendering it unsuitable for studies of barcodes with more than one marker (e.g., botanic barcode studies using both rbcL and matK), because sequences from different markers are not associated with the single originating specimen. GenBank sequences may also contain hidden sampling biases, particularly if they lack the barcode keyword (Spouge and Mariño-Ramírez 2012). Few methods exist to assess imperfections in a taxonomic gold standard, but investigators can partition available sequences into classes expected to correlate with the accuracy of taxonomic classification (e.g., taxonomically curated and non-curated sequences). Taxonomic inaccuracies can then be inferred from statistical differences between the classes (Suwannasai et al. 2013).
If the aim of the measurement of the identification accuracy is to reflect the performance of the corresponding barcode database (as it should be), the aim constrains the number of specimens sampled within a species and the number of species sampled within genera. Section 5 discusses some nuances of sampling.
The evaluation of barcode accuracy should also account for the PCR failures occurring while obtaining relevant markers from a specimen: clearly, if PCR fails to amplify a marker from a specimen, the marker can contribute little to species identification. Measures of the identification accuracy exist to incorporate the effect of PCR failure directly (Spouge and Mariño-Ramírez 2012). PCR failure rates may diminish rapidly with technological advances, however, so current practice distinguishes PCR failure from identification accuracy by stating the (present) PCR failure rate and then restricting the barcode database to samples with successful PCR amplification (Schoch et al. 2012; Hollingsworth et al. 2009).
4 Algorithms to Assign Species
With an appropriate barcode database in hand, the computer must assign a species to each query (or declare “failure to assign”). The next step, therefore, is to develop a computer method for assigning a species to each specimen and its marker sequence(s). Unless designed for uncommonly narrow purposes, the method needs to handle datasets whose species might have either few or many representative specimens. For future reference, note that like any classification method, species assignment should consider all available information, at least in principle. The discourse below focuses on sequence information, but other types of information (e.g., morphological, geographical, etc.) might be available to influence species assignment.
Species assignment algorithms examining more than a specimen’s nearest neighbors (e.g., algorithms building phlylogenetic trees, e.g., parsimony (Farris 1972), neighbor-joining (Saitou and Nei 1987), and Bayesian inference on trees (Munch et al. 2008) have not been noticeably more accurate than the simple strategy of assigning specimens to the species of their nearest neighbor(s) within the barcode database (Austerlitz et al. 2009; Austerlitz 2007; Little and Stevenson 2007). Moreover, many algorithms are too slow for the high-throughput species identifications large barcode databases require (e.g., most tree-building and probabilistic algorithms (Felsenstein 1981, 1988).
Another class of species assignment algorithms (“diagnostic methods”) treat differences between aligned sequences as potential taxonomic characters (e.g., DNA-BAR (DasGupa et al. 2005), BLOG (Weitschek et al. 2013), CAOS (Sarkar et al. 2008), BRONX (Little 2011), PTIGS-DIdIt (Liu et al. 2011), and Linker (Albu et al. 2011). Diagnostic methods (when properly formalized) are essentially machine-learning methods, which generally require 4 samples per species to be effective (Weitschek et al. 2014). In datasets with noticeably fewer than 4 specimens per species, diagnostic methods may over-fit, reducing their identification accuracy on sparsely represented species.
Sequence distance methods (or related similarity methods, e.g., BLAST and PSI-BLAST (Altschul et al. 1997), BLAST+P (Pang et al. 2012), NN (Austerlitz et al. 2009), and TaxonDNA (Meier et al. 2006) can identify species despite sparse representation. A distance method essentially brings prior knowledge to a species with sparse representation, aiding in its identification.
Most distance methods begin by aligning marker sequences. In contrast, alignment-free distances do not require sequence alignment (Kuksa and Pavlovic 2009), making them simple and fast (Little and Stevenson 2007, Kuksa and Pavlovic 2009). They provide competitive identification accuracy in fish, butterflies, and birds (Kuksa and Pavlovic 2009), but they remain untested in problematic taxa like plants or fungi. So far, therefore, they have not been widely adopted for species identification.
Other distance methods necessitate sequence alignment. Evolutionary distances require a global sequence alignment, i.e., alignment over the full length of the sequences examined. Similarly, sequence distance methods use the full sequence lengths and are in fact equivalent to global sequence similarity methods (Smith et al. 1981), which assess similarity over full sequence lengths. In contrast, local sequence similarity methods assess only the two most similar subsequences in the sequences. (See Fig. 2 in Ref. Spouge and Mariño-Ramírez 2012, for diagrams of local and global alignments.) Local sequence alignment programs (e.g., BLAST) therefore might declare a statistically significant similarity based only on small subsequences displaying convergent evolution (homoplasy) (Wouters and Husain 2001). Local sequence alignment can therefore make distant species appear spuriously close, whereas global alignment always highlights contrasting dissimilarities across the whole sequence. In principle (and therefore if feasible), barcode studies should prefer global alignment (e.g., they should use one of the many tools performing the Needleman-Wunsch Algorithm (Needleman and Wunsch 1970) over local alignment (e.g., they should avoid the Smith-Waterman Algorithm (Smith and Waterman 1981) or BLAST, if possible). Alignments types other than global and local exist (e.g., semi-global alignment), but they did not assign species noticeably better than global alignment, at least in fungi (Schoch et al. 2012; Suwannasai et al. 2013).
In practice, however, nearest neighbor species are critical to species identification. When aligning markers from nearest neighbors, local alignment programs like BLAST often align their full sequence lengths, because local alignment then fuses subsequence alignments by bridging the short gaps between them in closely related sequences. Thus, in the alignments critical to species identification, local alignment programs often perform global alignment anyway. Investigators should be note, however, that local alignment unnecessarily complicates the interpretation of their results, possibly to the point of invalidating them, if the local alignments are over small subsequences of the original sequences. Large gaps in global alignments signal the possibility of local alignment of small subsequences and are a particularly troubling possibility in the intergenic spacers often used as adjunct markers in botanic studies, e.g., trnH-psbA.
The specific alignment or evolutionary distance chosen for a barcode analysis does not influence the nearest neighbor(s) much, so contrary statements notwithstanding (Hebert et al. 2003a), it does not affect the accuracy of most species assignments materially (Suwannasai et al. 2013; Kwong et al. 2012; Fregin et al. 2012; Collins 2012). In a pairwise alignment, therefore, the proportion of nucleotide pairs consisting of different nucleotides (“p-distance”) recommends itself as a particularly simple and effective distance (Little and Stevenson 2007).
The Barcode of Life Database (BOLD http://www.boldsystems.org; Ratnasingham and Hebert 2007) stores sequences in global multiple sequence alignments (MSAs). Many publicly available computer programs (e.g., MUSCLE Edgar 2004), MAFFT (Katoh et al. 2002), or HMMer (Eddy 1995) create MSAs; BOLD uses HMMer to align marker sequences before comparing them. Large barcode databases use MSAs, because MSAs are computationally much faster than pairwise sequence alignments. (For a database of \( N \) barcodes, the time for pairwise alignment is approximately proportional to \( N^{2} \).)
On the other hand, studies with fewer sequences (say, less than 500) have the luxury of more computationally intensive alignment methods. Global alignment performs inconsistently but slightly better overall than MSA, although not statistically beyond sampling errors when used with databases of 500 or fewer sequences (Schoch et al. 2012; Suwannasai et al. 2013; Hollingsworth et al. 2009; Erickson et al. 2008). The BLAST+P method identifies the species of a query by forming an MSA from its 100 top BLAST hits and then assigning species in accord with the closest MSA neighbor under sequence distance. Its identification accuracy noticeably improved on BLAST alone, at least some cases (Pang et al. 2012).
5 Probability of Correct Identification (PCI)
Given a dataset and species assignment algorithm, we now want to measure identification accuracy. In most cases, the measure should scale, so that it estimates the performance of a comparable high-throughput database in identifying a specimen’s species correctly. With this aim in mind, consensus has focused on “the probability of correct identification” (PCI) as an appropriate measurement (CBOL Plant Working Group 2009; Meier et al. 2006; Erickson et al. 2008). The definition of PCI is broad enough to accommodate legitimate scientific disagreement about species identification, so in fact the concept generates a class of measures capable of accommodating various scientific needs.
Consider any particular dataset, and for the moment, assume that each species within it generates a known PCI (specified later). The overall PCI for the dataset is the species PCI for each species, averaged over all species in the dataset. If a few data subsets are particularly important (e.g., non-basal angiosperms, basal angiosperms, and gymnosperms within plant taxa (Hollingsworth et al. 2009); or Pezizomycotina, Saccharomycotina, Basidiomycota, and early diverging lineages within fungi (Schoch et al. 2012), the PCIs can be reported separately for each subset. In principle, a weight within the average could reflect a species’ under- or over-representation (Pang et al. 2012) or its intrinsic importance within the dataset. Most scientists do not weight averages when calculating overall PCI, however, because usually the weights just represent ephemeral sampling biases. In any case, we need only calculate a species PCI, a probability quantifying identification accuracy in each individual species, to calculate the overall PCI for a dataset.
A leave-one-out procedure (“the jackknife” in statistical language (Efron and Stein 1981) provides the species PCI. Consider a particular species and its representative specimens within the dataset. Remove each representative in turn from the dataset, and use the representative as a query specimen for the dataset. Removed from the dataset, the representative mimics a newly sampled specimen from the species, and the species is “known” if other representatives of the species exist in the dataset. Thus, the leave-one-out procedure is possible only for a species with more than one representative in the dataset. Although a singleton species (i.e., a species with a unique representative) cannot contribute to a species PCI under the leave-one-out, singletons do provide realistic “decoys” within the dataset when considering queries from other species.
Singletons with unique sequences are also a useful ancillary statistic in a barcode study, but the singletons have little relation to the correct identification of the corresponding species in realistic barcode databases (Hollingsworth et al. 2009). For similar reasons, datasets should try to include several species in each genus sampled. In any case, optimistically conflating singleton uniqueness with perfect identification in a heavily sampled species (which is much more demanding) is just plain misleading.
In a non-singleton species, scientists might legitimately disagree over the definition of successful identification of the species. Some scientists might define “success” as a monophyly, where every specimen in the species is closer to all specimens in the species than to any other specimen (CBOL Plant Working Group 2009) Success then is a binary decision, with the species PCI being either 1 or 0. Other scientists might define success pragmatically, by analogy to correct assignment of the species as in a database, where each specimen from the species has as its nearest neighbor(s) only specimens in the species (Meier et al. 2006). Again, species PCI is then either 1 or 0. Additional definitions of success are possible, depending on ties outside the species for a nearest neighbor, assignment of specimens from other species to the species in question, etc.
Some authors have used loose criteria for success (e.g., for some \( k > 1 \), a specimen’s nearest \( k \) neighbors must contain at least one other specimen from the same species (Kuksa and Pavlovic 2009). Other authors have experimented with placing additional conditions on “success” as defined above, e.g., the presence of barcode gaps (i.e., p-distance differences between intra- and inter-species comparisons) such as 2 or 3 % (18). Although detection of unknown species with p-distance thresholds can be an artificial constraint (Ferguson 2002), any specific choice might be appropriate in different circumstances, depending on the scientific aim.
The following PCI definition relies mostly on averages over individuals. Define the PCI of a query specimen as the fraction of nearest neighbors with the same species as the query, then define the species PCI as the average of the specimen PCI over specimens within a species (Erickson et al. 2008). The definition has little dependence on sudden 0 to 1 changes, as might occur with PCI definitions using barcode gaps. Consequently, its statistical error (discussed briefly in Sect. 6) is smaller than many other definitions of PCI, making the value of the PCI when scaled to large databases more predictable.
In any case, two PCIs are not necessarily comparable just because they both lie between 0 and 1! PCIs are comparable only if their underlying definitions are similar. For example, the definition above using nearest \( k \) neighbors obviously produces larger PCIs than the more stringent definition using only the nearest neighbor(s); singleton species if included in a PCI inflate its value unrealistically, etc. Referees should insist on uninflated measures of identification success, to encourage comparability of the resulting PCIs.
6 Statistical Sampling Error
The overall PCI is the (usually unweighted) average of the species PCIs. As a reasonable approximation, assume that species PCIs are independent across species. Every database is a sample of all possible species, so the overall PCI \( \hat{p} \) from the database is an estimate of the “true” overall PCI \( p \) (i.e., the overall PCI \( p \) in a hypothetical database including all organisms on Earth). As such, \( \hat{p} \) has a (known) sampling error. The corresponding confidence intervals are often surprisingly broad, making them extremely useful in resolving scientific disagreements.
The binomial distribution provides several possible (but approximately concordant) confidence intervals for \( \hat{p} \), of which two at least have appeared in barcode studies. The Wilson score interval (Suwannasai et al. 2013; Little 2011) is described in detail elsewhere (Wilson 1927), whereas the normal approximation was described in a previous review (Spouge and Mariño-Ramírez 2012). In most circumstances, the Wilson score interval is probably preferable, although (as in a t-test) the normal approximation yields the probability that the difference between two PCIs is different from 0.
7 Summary
A consensus has settled on the probability of correct identification (PCI) for the measurement of species identification accuracy with a barcode. The measurement involves several steps.
First, assemble databases corresponding to the markers. The choice of database should receive careful consideration, because it profoundly influences conclusions. Because GenBank taxonomy is often undependable, and because most GenBank sequences do not specify the originating specimen, studies based on GenBank sequences lacking the barcode keyword may have less reproducible conclusions than a carefully controlled taxonomic study. The database should try to include several samples per species and several species per genus.
Second, select a computer algorithm for assigning species to a query specimen’s barcode sequences. Only one algorithm has improved noticeably on the following: compute the p-distance between the query and each database marker, and then assign to the query the species of its nearest neighbor(s) in the database. The superior BLAST+P algorithm identifies the species of a query by forming a multiple sequence alignment (MSA) from the query’s 100 top BLAST hits in the database and then assigning the query to the species of the closest MSA neighbor under a sequence distance. The essential improvement in BLAST+P might be restricting the MSA to the query’s 100 closest neighbors to improve the alignment quality, but definitive conclusions must await further investigation.
Global alignment (e.g., with Needleman-Wunsch algorithm, or with any MSA algorithm) uses all the information in barcode sequences. By contrast, local alignment programs like BLAST might match only small subsequences within two sequences. Thus, barcode investigations using BLAST run the risk of producing an artifact, particularly if the resulting alignments do not extend over entire sequence lengths, and particularly when concluding the inferiority of intergenic markers. As long as sequence alignments use the entire lengths of sequences, algorithms using p-distance (which requires only base-pair counts) identify species just as well as algorithms using other, more complicated distances (e.g., alignment distance and similarity scores, and evolutionary distances like Kimura 2-Parameter Distance, etc.).
Consensus has converged on “the probability of correct identification” (PCI) as a measurement of species identification accuracy in barcode studies. The overall PCI for a dataset is the average of the species PCIs, taken over all species in the dataset. If a dataset contains some distinguished subsets, the investigator can report PCIs for those subsets.
To calculate a species PCI, remove in turn each representative of the species from the database, and consider its distance (e.g., p-distance) from the remaining representatives. Section 5 gives several possible definitions of successful identification within a species. Some were more stringent than others, because scientific purpose makes different definitions of “successful assignment” appropriate to different circumstances. Singletons with unique sequences provide a useful ancillary statistic in barcode investigations, but they bear little on the correct identification of the corresponding species in a realistic barcode database. Referees should discourage the optimistic conflation of singleton uniqueness with perfect identification of a heavily sampled species.
The evaluation of identification accuracy should also assess PCR failure rates. Because the rates may diminish rapidly with technological advances, current practice distinguishes PCR failure from intrinsic identification accuracy by stating the (present) PCR failure rate and then restricting the barcode database for measuring identification accuracy to samples with successful PCR amplification (Schoch et al. 2012; Hollingsworth et al. 2009).
Finally, a dataset provides a statistical sample exemplifying a class of possible datasets. The PCI estimated from a dataset therefore estimates a true overall PCI, and as such, it has a statistical error. The errors are sometimes surprisingly large, and therefore well worth calculating.
For software relevant to this chapter, see http://tinyurl.com/spouge-barcode.
References
Albu M, Nikbakht H, Hajibabaei M, Hickey DA (2011) The DNA barcode linker. Mol Ecol Resour 11:84–88
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Austerlitz F (2007) Comparing phylogenetic and statistical classification methods for DNA barcoding. In: The second international barcode of life conference, Taipei
Austerlitz F, David O, Schaeffer B, Bleakley K, Olteanu M, Leblois R, Veuille M, Laredo C (2009) DNA barcode analysis: a comparison of phylogenetic and statistical classification methods. BMC Bioinform 10:1
Blaxter M, Mann J, Chapman T, Thomas F, Whitton C, Floyd R, Abebe E (2005) Defining operational taxonomic units using DNA barcode data. Philos Trans R Soc B Biol Sci 360:1935–1943
CBOL Plant Working Group (2009) A DNA barcode for land plants. Proc Natl Acad Sci USA 106:12794–12797
Chase MW, Salamin N, Wilkinson M, Dunwell JM, Kesanakurthi RP, Haidar N, Savolainen V (2005) Land plants and DNA barcodes: short-term and long-term goals. Philos Trans R Soc Lond B Biol Sci 360:1889–1895
Chen J, Zhao JT, Erickson DL, Xia NH, Kress WJ (2015) Testing DNA barcodes in closely related species of Curcuma (Zingiberaceae) from Myanmar and China. Mol Ecol Resour 15:337–348
Collins RA (2012) Barcoding’s next top model: an evaluation of nucleotide substitution models for specimen identification. Methods Ecol Evol 3:457–465
Cowan RS, Chase MW, Kress JW, Savolainen V (2006) 300,000 species to identify: problems, progress, and prospects in DNA barcoding of land plants. Taxon 55:611–616
DasGupa B, Konwar KM, Mandoiu II, Shvartsman AA (2005) DNA-BAR: distinguisher selection for DNA barcoding. Bioinformatics 21:3424–3426
Eddy SR (1995) Multiple alignment using hidden Markov models. Proc Int Conf Intell Syst Mol Biol 3:114–120
Edgar RC (2004) MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinform 5:113
Efron B, Stein C (1981) The Jackknife estimate of variance. Ann Stat 9:586–596
Erickson DL, Spouge JL, Resch A, Weight LA, Kress JW (2008) DNA barcoding in land plants: developing standards to quantify and maximize success. Taxon 13:1304–1316
Farris JS (1972) Estimating phylogenetic trees from distance matrices. Am Nat 106:645–668
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1988) Phylogenies from molecular sequences—inference and reliability. Annu Rev Genet 22:521–565
Ferguson JWH (2002) On the use of genetic divergence for identifying species. Biol J Linn Soc 75:509–516
Ferri G, Corradini B, Ferrari F, Santunione AL, Palazzoli F, Alu M (2015) Forensic botany II, DNA barcode for land plants: which markers after the international agreement? Forensic Sci Int Genet 15:131–136
Floyd R, Abebe E, Papert A, Blaxter M (2002) Molecular barcodes for soil nematode identification. Mol Ecol 11:839–850
Fregin S, Haase M, Olsson U, Alstrom P (2012) Pitfalls in comparisons of genetic distances: a case study of the avian family Acrocephalidae. Mol Phylogenet Evol 62:319–328
Hajibabaei M, Janzen DH, Burns JM, Hallwachs W, Hebert PDN (2006) DNA barcodes distinguish species of tropical Lepidoptera. Proc Natl Acad Sci USA 103:968–971
Hebert PD, Cywinska A, Ball SL, deWaard JR (2003a) Biological identifications through DNA barcodes. Proc Biol Sci 270:313–321
Hebert PD, Ratnasingham S, deWaard JR (2003b) Barcoding animal life: cytochrome c oxidase subunit 1 divergences among closely related species. Proc Biol Sci 270(Suppl 1):S96–S99
Hogg ID, Hebert PDN (2004) Biological identification of springtails (Hexapoda: Collembola) from the Canadian Arctic, using mitochondrial DNA barcodes. Can J Zool (Revue Canadienne De Zoologie) 82:749–754
Hollingsworth PM, Forrest LL, Spouge JL, Hajibabaei M, Ratnasingham S, van der Bank M, Chase MW, Cowan RS, Erickson DL, Fazekas AJ, Graham SW, James KE, Kim K-J, Kress WJ, Schneider H, van AlphenStahl J, Barrett SCH, van den Berg C, Bogarin D, Burgessk KS, Cameron KM, Carine M, Chacón J, Clark A, Clarkson JJ, Conrad F, Devey DS, Ford CS, Hedderson TAJ, Hollingsworth ML, Husband BC, Kellya LJ, Kesanakurti PR, Kim JS, Kim Y-D, Lahaye R, Lee H-L, Long DG, Madriñán S, Maurin O, Meusnier I, Newmaster SG, Park C-W, Percy DM, Petersen G, Richardson JE, Salazar GA, Savolainene V, Seberg O, Wilkinson MJ, Yi D-K, Little DP (2009) A DNA barcode for land plants. Proc Natl Acad Sci USA 106:12794–12797
Huang D, Meier R, Todd PA, Chou LM (2008) Slow mitochondrial COI sequence evolution at the base of the metazoan tree and its implications for DNA barcoding. J Mol Evol 66:167–174
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucl Acids Res 30:3059–3066
Kress WJ, Erickson DL (2008) DNA barcodes: genes, genomics, and bioinformatics. Proc Natl Acad Sci USA 105:2761–2762
Kress WJ, Wurdack KJ, Zimmer EA, Weigt LA, Janzen DH (2005) Use of DNA barcodes to identify flowering plants. Proc Natl Acad Sci USA 102:8369–8374
Kuksa P, Pavlovic V (2009) Efficient alignment-free DNA barcode analytics. BMC Bioinform 10:S9
Kwong S, Srivathsan A, Vaidya G, Meier R (2012) Is the COI barcoding gene involved in speciation through intergenomic conflict? Mol Phylogenet Evol 62:1009–1012
Lambert DM, Baker A, Huynen L, Haddrath O, Hebert PDN, Millar CD (2005) Is a large-scale DNA-based inventory of ancient life possible? J Hered 96:279–284
Little DP (2011) DNA barcode sequence identification incorporating taxonomic hierarchy and within taxon variability. PLoS ONE 6(8):e20552
Little DP, Stevenson DW (2007) A comparison of algorithms for the identification of specimens using DNA barcodes: examples from gymnosperms. Cladistics 23:1–27
Liu C, Liang D, Gao T, Pang X, Song J, Yao H, Han J, Liu Z, Guan X, Jiang K, Li H, Chen S (2011) PTIGS-IdIt, a system for species identification by DNA sequences of the psbA-trnH intergenic spacer region. BMC Bioinform 12:1
Lorenz JG, Jackson WE, Beck JC, Hanner R (2005) The problems and promise of DNA barcodes for species diagnosis of primate biomaterials. Philos Trans R Soc B Biol Sci 360:1869–1877
Meier R, Shiyang K, Vaidya G, Ng PK (2006) DNA barcoding and taxonomy in Diptera: a tale of high intraspecific variability and low identification success. Syst Biol 55:715–728
Meyer CP, Paulay G (2005) DNA barcoding: error rates based on comprehensive sampling. PLoS Biol 3:e422
Munch K, Boomsma W, Huelsenbeck JP, Willerslev E, Nielsen R (2008) Statistical assignment of DNA sequences using Bayesian phylogenetics. Syst Biol 57:750–757
Needleman SB, Wunsch CD (1970) A general method applicable to the search for similarities in the amino acid sequence of two proteins. J Mol Biol 48:443–453
Pang X, Liu C, Shi L, Liu R, Liang D, Li H, Cherny SS, Chen S (2012) Utility of the trnH-psbA intergenic spacer region and its combinations as plant DNA barcodes: a meta-analysis. PLoS ONE 7:e48833
Ratnasingham S, Hebert PD (2007) BOLD: the barcode of life data system, Mol Ecol Notes 7:355–364. http://www.barcodinglife.org
Saitou N, Nei M (1987) The neighbor-joining method—a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sarkar IN, Planet PJ, Desalle R (2008) CAOS software for use in character-based DNA barcoding. Mol Ecol Resour 8:1256–1259
Saunders GW (2005) Applying DNA barcoding to red macroalgae: a preliminary appraisal holds promise for future applications. Philos Trans R Soc B Biol Sci 360:1879–1888
Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque CA, Chen W, Bolchacova E, Voigt K, Crous PW, Miller AN, Wingfield MJ, Aime MC, An KD, Bai FY, Barreto RW, Begerow D, Bergeron MJ, Blackwell M, Boekhout T, Bogale M, Boonyuen N, Burgaz AR, Buyck B, Cai L, Cai Q, Cardinali G, Chaverri P, Coppins BJ, Crespo A, Cubas PP, Cummings C, Damm U, de Beer ZW, de Hoog GS, Del-Prado R, Dentinger B, Dieguez-Uribeondo J, Divakar PK, Douglas B, Duenas M, Duong TA, Eberhardt U, Edwards JE, Elshahed MS, Fliegerova K, Furtado M, Garcia MA, Ge ZW, Griffith GW, Griffiths K, Groenewald JZ, Groenewald M, Grube M, Gryzenhout M, Guo LD, Hagen F, Hambleton S, Hamelin RC, Hansen K, Harrold P, Heller G, Herrera G, Hirayama, K, Hirooka Y, Ho HM, Hoffmann K, Hofstetter V, Hognabba F, Hollingsworth PM, Hong SB, Hosaka K, Houbraken J, Hughes K, Huhtinen S, Hyde KD, James T, Johnson EM, Johnson JE, Johnston PR, Jones EB, Kelly LJ, Kirk PM, Knapp DG, Koljalg U, Kovacs GM, Kurtzman CP, Landvik S, Leavitt SD, Liggenstoffer AS, Liimatainen K, Lombard L, Luangsa-Ard JJ, Lumbsch HT, Maganti H, Maharachchikumbura SS, Martin MP, May TW, McTaggart AR, Methven AS, Meyer W, Moncalvo JM, Mongkolsamrit S, Nagy LG, Nilsson RH, Niskanen T, Nyilasi I, Okada G, Okane I, Olariaga I, Otte J, Papp T, Park D, Petkovits T, Pino-Bodas R, Quaedvlieg W, Raja HA, Redecker D, Rintoul T, Ruibal C, Sarmiento-Ramirez JM, Schmitt I, Schussler A, Shearer C, Sotome K, Stefani FO, Stenroos S, Stielow B, Stockinger H, Suetrong S, Suh SO, Sung GH, Suzuki M, Tanaka K, Tedersoo L, Telleria MT, Tretter E, Untereiner WA, Urbina H, Vagvolgyi C, Vialle A, Vu TD, Walther G, Wang QM, Wang Y, Weir BS, Weiss M, White MM, Xu J, Yahr R, Yang, ZL, Yurkov A, Zamora JC, Zhang N, Zhuang WY, Schindel D, Fungal Barcoding C (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc Natl Acad Sci USA 109:6241–6246
Smith MA, Fisher BL, Hebert PDN (2005) DNA barcoding for effective biodiversity assessment of a hyperdiverse arthropod group: the ants of Madagascar. Philos Trans R Soc B Biol Sci 360:1825–1834
Smith MA, Woodley NE, Janzen DH, Hallwachs W, Hebert PDN (2006) DNA barcodes reveal cryptic host-specificity within the presumed polyphagous members of a genus of parasitoid flies (Diptera: Tachinidae). Proc Natl Acad Sci USA 103:3657–3662
Smith TF, Waterman MS (1981) Identification of common molecular subsequences. J Mol Biol 147:195–197
Smith TF, Waterman MS, Fitch WM (1981) Comparative biosequence metrics. J Mol Evol 18:38–46
Spouge JL, Mariño-Ramírez L (2012) The practical evaluation of DNA barcode efficacy. Methods Mol Biol 858:365–377
Suwannasai N, Martin MP, Phosri C, Sihanonth P, Whalley AJS, Spouge JL (2013) Fungi in Thailand: a case study of the efficacy of an ITS barcode for automatically identifying species within the Annulohypoxylon and Hypoxylon Genera. PLoS ONE 8:e54529
Weitschek E, Fiscon G, Felici G (2014) Supervised DNA barcodes species classification: analysis, comparisons and results. Biodata Min 7:1
Weitschek E, Van Velzen R, Felici G, Bertolazzi P (2013) BLOG 2.0: a software system for character-based species classification with DNA barcode sequences. What it does, how to use it. Mol Ecol Resour 13:1043–1046
Wilson EB (1927) Probable inference, the law of succession, and statistical inference. J Am Stat Assoc 22:209–212
Wouters MA, Husain A (2001) Changes in zinc ligation promote remodeling of the active site in the zinc hydrolase superfamily. J Mol Biol 314:1191–1207
Zalasiewicz J et al (2000) Are we now living in the Anthropocene? GSA Today 18:4–8
Acknowledgments
This research was supported in part by the Intramural Research Program of the NIH, NLM, NCBI.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Spouge, J.L. (2016). Measurement of a Barcode’s Accuracy in Identifying Species. In: Trivedi, S., Ansari, A., Ghosh, S., Rehman, H. (eds) DNA Barcoding in Marine Perspectives. Springer, Cham. https://doi.org/10.1007/978-3-319-41840-7_2
Download citation
DOI: https://doi.org/10.1007/978-3-319-41840-7_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-41838-4
Online ISBN: 978-3-319-41840-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)