Abstract
Progress in high-throughput sequencing and development of computational tools for identification of SECIS elements, selenoprotein genes and selenocysteine machinery allows recognition of organisms that are dependent, or not dependent, on selenium (Se) and identification of selenoproteins responsible for this trait. Full sets of selenoproteins in organisms, designated selenoproteomes, have been characterized for humans, which have 25 selenoprotein genes, as well as for most other organisms with sequenced genomes. This chapter offers an overview of eukaryotic selenoproteins at the level of individual proteins, protein families and entire selenoproteomes. Comparative genomic and ionomic analyses offer exciting avenues for studying selenoprotein function and evolution, provide insights into the biological functions of the trace element, Se, and allow addressing other important biological questions.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Selenium (Se) is utilized by organisms in three different ways. First, this element is present in the form of 2-selenouridine in the wobble position of certain bacterial tRNAs [1]. Second, it occurs in some bacterial Se-containing molybdoproteins as a labile cofactor that contains a Se-Mo bond that is directly involved in catalysis [2–4]. Third, Se is present in proteins in the form of selenocysteine (Sec) , the 21st amino acid in the genetic code [5, 6]. Sec is encoded by UGA and has been found in each of the three domains of life (i.e., bacteria, archaea and eukaryotes). It is now clear that the essential roles of Se in biology, as well as its beneficial functions in human health, are due to its presence in proteins in the form of Sec. Interestingly, unlike 20 common amino acids in proteins (which can be used throughout the protein sequence), Sec is utilized only when it is essential for protein function. Accordingly, it is often a key functional group in proteins. Moreover, Sec is specifically used for redox catalysis. As such, information on identity and functions of selenoproteins is often a key to biological and biomedical roles of Se, and it can be used to address a variety of other biomedical issues .
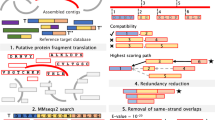
Selenoprotein families . Known selenoprotein families are shown. Selenoproteins that occur in vertebrates and unicellular eukaryotes are indicated and shown in bold and red, respectively. Additional selenoproteins that only occur in prokaryotes are shown below eukaryotic selenoproteins. On the right, relative sizes of selenoproteins are shown (relative to a 100 amino acid scale) and the location of Sec within protein sequence is shown by a red mark
2 Computational Tools for Selenoprotein Identification
All selenoprotein genes have two characteristic features : (1) an UGA codon that designates Sec; and (2) a Sec insertion sequence (SECIS) element. The UGA codon does not provide sufficient information to identify Sec positions. However, SECIS elements can be used for this purpose as these structures are highly specific for selenoprotein genes, have conserved sequence elements and possess a sufficiently complex secondary structure. Therefore, previous bioinformatics analyses mainly focused on SECIS elements , and selenoprotein discovery was based on the following general strategy: (1) finding candidate SECIS elements; (2) analyzing upstream regions to identify coding regions of selenoproteins; and (3) testing these candidates for insertion of Sec by metabolically labeling cells with 75Se. The first selenoproteins identified using this approach were mammalian selenoproteins R (now known as methionine-R-sulfoxide reductase 1 or MsrB1), N and T [9, 10]. These searches were initially restricted to small nucleotide sequence databases and later adapted to entire genomes [11–13]. To aid in these analyses, groups of closely related genomes are analyzed in order to identify conserved SECIS elements that belong to selenoprotein orthologs in these organisms [14]. Over the years, the technique was refined, and currently an arsenal of tools is available for identification of SECIS elements and selenoprotein genes, most notably, SECISearch3 [7], Selenoprofiles [8] and Seblastian [7].
In addition to the approach that identifies SECIS elements, a method was developed that searches for UGA codons flanked by conserved coding sequences [13–16]. This approach is based on the finding that the majority of selenoprotein genes have orthologs, in which Sec is replaced with cysteine (Cys). For example, an ortholog of mammalian selenoprotein MsrB1 is a Cys-containing MsrB in yeast and plants. This method is used as follows: protein databases, e.g., protein sequences from NCBI, are searched against NCBI nucleotide sequences with TBLASTN to identify nucleotide sequences which, when translated in one of six reading frames, align with Cys-containing protein sequences from the protein database, such that Cys residues correspond to UGA in the nucleotide sequences and the resulting Sec/Cys pairs are flanked by conserved sequences. Such a Sec/Cys homology approach is completely independent of the searches for SECIS elements. As both Sec/Cys and SECIS-based algorithms identify very similar sets of selenoprotein genes in organisms, both methods show excellent performance and can identify the majority, and often all, selenoprotein genes in sequenced genomes.
Figure 11.1 summarizes known eukaryotic selenoprotein families. We further provide a brief overview of selenoproteins that have been functionally characterized. Readers are referred to individual chapters within the book for additional information on various selenoproteins.
3 Mammalian Selenoproteins
3.1 Glutathione Peroxidases
There are eight glutathione peroxidases (GPX1-GPX8) in mammals, five of which are selenoproteins (GPX1, GPX2, GPX3, GPX4 and GPX6). GPX1-GPX4 are selenoproteins in all mammals, whereas GPX6 is a selenoprotein in many species, but in some it is a Cys-containing protein [14]. GPX1 was the first animal selenoprotein identified [17]. It is also the most abundant mammalian selenoprotein that exhibits a particularly high expression in the liver and kidney. This antioxidant enzyme catalyzes glutathione-dependent hydroperoxide reduction. GPX4 has received much attention recently due to its essential function (e.g., its knockout in mice leads to early embryonic lethality), and its role in ferroptosis and regulation of phospholipid hydroperoxide levels [18]. Interestingly, a mitochondrial form of this protein serves a structural role in mature sperm and participates in disulfide bond formation during spermiogenesis [19]. Whereas GPX1 and GPX4 are expressed in all cells, GPX2 expression is largely restricted to the gastrointestinal tract, and GPX3 is primarily made in the kidney and secreted into the blood stream. GPX3 localizes to the basement membrane of the proximal tubules in kidney [20]. However, it remains unclear how it can function in the extracellular milieu in the absence of sufficient levels of thiol reductants. Besides mammals, selenoprotein GPX homologs were found in most animals as well as in various single-celled eukaryotes and even in bacteria. However, the ancestral form of these enzymes is a Cys-containing protein, and it is thought that Cys was replaced with Sec during evolution, making these enzymes better catalysts.
3.2 Thyroid Hormone Deiodinases
Mammals have three deiodinase genes (DIO1, DIO2 and DIO3), which activate and/or inactivate thyroid hormones by reductive deiodination. Deiodinases also occur in other vertebrates, and their homologs (some of them are Cys-containing proteins) were even detected in unicellular eukaryotes and bacteria, although their function must be different in these organisms. Like GPXs and the majority of other selenoproteins, deiodinases are thioredoxin-fold proteins.
3.3 Thioredoxin Reductases
All three mammalian thioredoxin reductases (TXNRDs ) are selenoproteins, hence the entire thioredoxin (TXN) system in mammals is dependent on Se. In these proteins, Sec is present in the C-terminal penultimate position, preceded by Cys and followed by glycine. These enzymes evolved from glutathione reductases by including a C-terminal Sec-containing extension that serves as an intraprotein substrate for the N-terminal active center of pyridine nucleotide disulfide oxidoreductase family members [21–24]. TXNRD1 is a cytosolic and nuclear protein. Its main function is to maintain TXN1 in the reduced state by reducing a disulfide in this protein in an NADPH-dependent manner. It exhibits broad substrate specificity, especially with regard to low molecular weight compounds [25] and occurs in the form of multiple isoforms generated by alternative transcription initiation and splicing [26–28]. Another member of the mammalian TXNRD family is thioredoxin/glutathione reductase (TGR, TXNRD3). This enzyme has an additional N-terminal glutaredoxin domain [22], which is implicated in the formation/isomerization of disulfide bonds during sperm maturation [29]. TGR can catalyze reactions specific for both TXN and glutathione systems. A third member of the family, TXNRD2, is a mitochondrial protein, which maintains mitochondrial TXN2 as well as glutaredoxin 2 in the reduced state. TXNRD1 and TXNRD2 are essential for embryonic development in mice [30, 31].
3.4 Methionine-R-Sulfoxide Reductase 1 (MSRB1)
This protein was initially designated as selenoprotein R [9] and selenoprotein X [10], but after it was found to catalyze stereospecific reduction of methionine-R-sulfoxide residues in proteins, it was renamed MSRB1 [32]. Mammals have two additional MSRBs (MSRB2 and MSRB3), which contain catalytic Cys in place of Sec and reside in mitochondria and endoplasmic reticulum, respectively [33]. At least in the liver and kidney of mammals, MSRB1 has the highest activity of all MSRBs, so the protein reductive repair function is dependent on Se in mammals. MSRB1 is located in the cytosol and nucleus [33]. Msrb1 knockout mice are viable, but are characterized by oxidative stress [34]. An important recent discovery is the role of MSRB1 in regulation of actin polymerization by reversible methionine-R-sulfoxidation [35].
3.5 15 kDa Selenoprotein (SEP15)
SEP15 is a conserved eukaryotic selenoprotein that occurs in most animals as well as in some unicellular eukaryotes, such as algae. It resides in the endoplasmic reticulum where it binds UDP-glucose:glycoprotein glucosyltransferase, a sensor of protein folding [36]. SEP15 is composed of an N-terminal ER signal peptide, a Cys-rich domain responsible for binding UDP-glucose:glycoprotein glucosyltransferase, and a C-terminal domain characterized by the thioredoxin-like fold. SEP15 may in part be responsible for the effect of Se in cancer [37, 38]. Sep15 knockout mice are viable, but develop cataracts [39].
3.6 Selenophosphate Synthetase 2 (SEPHS2)
By analogy to bacterial selenophosphate synthetase SelD [40], SEPHS2 (also known as SPS2) was thought to synthesize selenophosphate, a Se donor compound. It is essential for selenoprotein biosynthesis in mammals. Mammals and some other metazoans have an SEPHS2 paralog, SEPHS1, whose function remains unknown [41], but it is not related to Sec biosynthesis [42].
3.7 Selenoprotein P (SEPP1)
SEPP1 is the only selenoprotein with multiple Sec residues in mammals [43], e.g., there are ten Sec residues in human and mouse SEPP1 , and seven in the naked mole rat protein [44]. However, the number of Sec residues in SEPP1 homologs varies greatly (7–16 in mammals) [44]. SEPP1 is the major plasma selenoprotein, which is synthesized primarily in the liver and delivers Se to certain other organs and tissues [45, 46]. The Sepp1 knockout mouse has been a particularly useful model in examining Se metabolism in mammals [47].
3.8 Selenoproteins W (SelW, SEPW1) and V (SELV)
SelW is the smallest mammalian selenoprotein [48]. Although it was one of the first identified, its function remains unknown. SEPW1 homologs were identified in lower eukaryotes and even bacteria, but these findings did not help identify the function of SEPW1 [16]. A SEPW1 paralog, SELV , is a larger protein due to an additional N-terminal sequence of unknown function [14]. This protein is expressed exclusively in testes. Its function is also not known.
3.9 Selenoproteins T (SELT), M (SELM) and H (SELH)
Functions of these mammalian selenoproteins are not known. They are listed here together because they belong to a group of thioredoxin-like fold proteins (together with SEP15, SEPW1 and SELV). SELT is among the first selenoproteins identified through bioinformatics [9]. SELM is a distant homolog of SEP15 and, like SEP15, it resides in the endoplasmic reticulum [49]. SELH was first identified as BthD in fruit flies [12, 14]. It resides in the nucleus. Several studies have found that knockdown of these proteins leads to oxidative stress suggesting roles, at least partially, as antioxidants.
3.10 Selenoproteins O (SELO) and I (SELI)
SELO is a widely distributed protein with homologs in animals, bacteria, yeast and plants, but the functions of any member of this protein family are not known [14]. Only vertebrate homologs of SELO have Sec, which is located in the C-terminal penultimate position, and the protein is located in mitochondria [50]. SELI is a recently evolved selenoprotein specific to vertebrates [14]. This membrane selenoprotein has no known function.
3.11 Selenoprotein K (SELK) and S (SELS)
SELK and SELS are unusual among selenoproteins in that they do not have a pronounced secondary structure [14]. These small selenoproteins contain a single transmembrane helix in the N-terminal sequence that targets them to the ER membrane. Studies revealed the role of SELS and SELK in retrotranslocation of misfolded proteins from the ER to the cytosol, where these proteins are further degraded [51]. Both proteins bind Derlins, which are ER membrane-resident proteins, and associate with multiprotein complexes [52, 53]. In addition, SELS was implicated in inflammation and the immune response, and SELK in protein palmitoylation [54]. A Selk knockout mouse model is viable [55].
3.12 Selenoprotein N (SEPN1)
One of the first selenoprotein discovered through bioinformatics approaches [10], SEPN1 remains a selenoprotein of unknown function. This protein was implicated in the role of Se in muscle function through biochemical and genetic analyses, as well as through analyses of knockout mice [56], and was found to serve as a cofactor for the ryanodine receptor [57]. Mutations in SEPN1 are associated with a hereditary muscular dystrophy.
4 Additional Selenoproteins in Eukaryotes
The following selenoproteins that are absent in mammals were identified in eukaryotes: methionine-S-sulfoxide reductase (MSRA ), protein disulfide isomerase (PDI ), selenoproteins U (SELU), L (SELL), J (SELJ), FEP15, MCS, and plasmodial selenoproteins Sel1, Sel2, Sel3 and Sel4, and a selenoprotein SelTryp from Trypanosoma. MSRA is a widely distributed protein family, whose function is to repair methionine residues in proteins. Like MSRB, it catalyzes a stereospecific reduction of methionine sulfoxides, but is specific for methionine-S-sulfoxides. MSRA was initially found in the green algae, Chlamydomonas [58], but later was also identified in other eukaryotes as well as in some bacteria. PDI is also very narrowly distributed in eukaryotes [59], in contrast to Cys-containing PDIs, which are essential for the formation of disulfide bonds in the ER of eukaryotic cells. SELU [60], SELJ [61], FEP15 [62], and SELL [63] were only found in fish and/or invertebrates. The four Plasmodium selenoproteins (Sel1-Sel4) showed no detectable homology to any other protein [64]. However, Sel1 and Sel4 have Sec in the C-terminal regions and may be related to SELK and SELS.
5 Selenoprotein Functions
Eukaryotic selenoproteins for which functions are known are oxidoreductases. In these proteins, Sec is the catalytic residue that is employed because it is superior to Cys in this function [40, 65–68]. In selenoproteins, Sec reversibly changes its redox state during catalysis. Functions of many selenoproteins, particularly those found in vertebrates, are not known. However, by analogy to proteins with known functions, it may be expected that the majority of these uncharacterized selenoproteins are also oxidoreductases.
Eukaryotic selenoproteins may be loosely clustered into two groups based on the location of Sec. The most abundant selenoprotein group includes proteins containing Sec in the N-terminal region or in the middle of the protein. Many of these selenoproteins exhibit thioredoxin or thioredoxin-like folds, but some proteins (e.g., SEPHS2, MSRA) show different folds. In these proteins, Sec is the catalytic group, which often works in concert with a resolving Cys. In the second group, Sec is located in the C-terminal sequences. These proteins include selenoproteins K, S, O, I and TXNRDs. Except for TXNRDs, the function of Sec in selenoproteins in this group is not known.
Non-catalytic functions of Sec, while rare, have also been described. Known examples include Sec residues in the C-terminal region of SEPP1, which function in transporting Se from liver to other organs, and recently evolved Sec residues in the Metridium senile MsrB homolog, wherein the function of these Sec residues is not known [69].
6 Selenoproteomes
Availability of tools for efficient identification of selenoprotein genes led to the recognition of all, or almost all, selenoproteins in many organisms [5, 6]. Information on the full sets of selenoproteins in organisms (selenoproteomes) offers an opportunity to address diverse questions relevant to the biology of Se, e.g., by linking individual selenoproteins or selenoprotein groups and specific effects of dietary Se [70]. In this regard, Se differs from other trace elements where new proteins are still discovered biochemically, and often by accident, and where full sets of proteins dependent on a particular element is difficult to ascertain.
Among metazoan selenoproteomes, an interesting case is represented by C. elegans and C. briggsae , which have only a single UGA codon in their genomes that codes for Sec [71]. This codon occurs in the TxnRd1 gene, and phylogenetic analyses suggested that other selenoprotein genes were lost in nematodes during evolution. Thus, the entire Sec machinery is maintained in these organisms to insert a single Sec residue. Selenoproteinless animals have also been identified, most of which are arthropods [72, 73]. Information about such animals (or other organisms that lost selenoproteins, such as yeast and higher plants) helps explain the changing requirements for Se during evolution. Interestingly, selenoproteinless insects lost the entire Sec insertion machinery, but preserved SEPHS1, suggesting that this protein is not involved in Sec biosynthesis [42, 72].
A recent study identified 59 selenoproteins in the harmful alga, Aureococcus anophagefferens [74, 75]. This organism has the largest and the most diverse selenoproteome identified to date, including known eukaryotic selenoproteins, selenoproteins previously only detected in bacteria, and novel selenoproteins. Similar to smaller selenoproteomes, the A. anophagefferens selenoproteome was dominated by the thioredoxin fold proteins, and oxidoreductase functions could be assigned to the majority of detected selenoproteins. Se was required for the growth of A. anophagefferens as cultures grew maximally at nanomolar Se concentrations. Moreover, in a coastal ecosystem, dissolved Se was elevated before and after A. anophagefferens blooms, but reduced by >95 % during the peak of blooms. Consistent with this pattern, enrichment of seawater with selenite before and after a bloom did not affect the growth of A. anophagefferens, but enrichment during the peak of the bloom significantly increased population growth rates. Thus, Se inventories, which can be anthropogenically enriched, can support proliferation of harmful algal blooms through synthesis of a large arsenal of selenoproteins.
Selenoproteome analyses also are capable of uncovering trends in the use of Sec [76], although some limitations of this approach have been described [77]. An analysis of selenoproteomes of several model eukaryotes detected 26–29 selenoprotein genes in two species of Ostreococcus , five in the social amoebae , Dictyostelium discoideum, and 16 in the diatom, Thalassiosira pseudonana, including several new selenoproteins [76]. Further analyses identified massive, independent selenoprotein losses in land plants, fungi, nematodes, insects and some protists. Comparative analyses of selenoprotein-rich and -deficient organisms revealed that aquatic organisms generally have large selenoproteomes, whereas several groups of terrestrial organisms reduced their selenoproteomes through loss of selenoprotein genes and replacement of Sec with Cys. These observations suggested that many selenoproteins originated at the base of the eukaryotic domain and showed that the environment may play a role in selenoproteome evolution. In particular, aquatic organisms apparently retained and sometimes expanded their selenoproteomes, whereas the selenoproteomes of some terrestrial organisms were reduced or completely lost. It is an interesting possibility that aquatic life supports Se utilization , whereas terrestrial habitats lead to the reduced use of this trace element [76].
In a separate study involving vertebrates , reconstruction of evolutionary changes in the Se transport domain of SEPP1 revealed a decrease in the Sec content specifically in the mammalian lineage via replacement of Sec with Cys [44]. Compared to mammals, fish showed higher Sec content of SEPP1, larger selenoproteomes, elevated SEPP1 gene expression, and higher levels of tissue Se. In addition, mammals replaced Sec with Cys in several proteins and lost several selenoproteins altogether, whereas such events were not found in fish. These data suggested that evolution from fish to mammals was accompanied by a decreased use of Sec and that analyses of SEPP1, selenoproteomes and Sec/Cys transitions provide a genetic marker of utilization of this trace element in vertebrates. The evolved reduced reliance on Se raises questions regarding the need to maximize selenoprotein expression by Se dietary supplements in situations where pathology is not imminent, which is a currently accepted practice.
A more recent study characterized the selenoproteomes of 44 sequenced vertebrates , and detected 45 selenoprotein subfamilies [78]. Twenty-eight of them were found in mammals, and 41 in bony fishes. The study defined the ancestral vertebrate (28 proteins) and mammalian (25 proteins) selenoproteomes, and described how they evolved along lineages through gene duplication (20 events), gene loss (10 events) and replacement of Sec with Cys (12 events). It was shown that an intronless SEPHS2 gene evolved in early mammals and replaced functionally the original multiexon gene in placental mammals, whereas both genes remain in marsupials. Mammalian TXNRD1 and TGR evolved from an ancestral glutaredoxin-domain containing enzyme, still present in fish. Selenoprotein V and GPX6 evolved specifically in placental mammals from duplications of SEPW1 and GPX3, respectively, and GPX6 lost Sec several times independently. Bony fishes were characterized by duplications of several selenoprotein families (GPX1, GPX3, GPX4, DIO3, MSRB1, SELJ, SELO, SELT, SELlU1, and SEPW2). The study also identified new isoforms for several selenoproteins and described unusually conserved selenoprotein pseudogenes.
7 Applications of Selenoproteome Analyses to Biology
7.1 Genetic Code Supports Targeted Insertion of Two Amino Acids by One Codon
Strict one-to-one correspondence between codons and amino acids was thought to be an essential feature of the genetic code. However, a recent selenoproteome analysis of the ciliate Euplotes crassus revealed that one codon can code for two different amino acids with the choice of the inserted amino acid determined by a specific 3′-untranslated region and location of the dual-function codon within the mRNA [79]. It was found that the codon UGA specified insertion of Sec and Cys in E. crassus, that the dual use of this codon could occur even within the same protein, and that the structural arrangements of Euplotes mRNA preserved the location-dependent dual function of UGA when expressed in mammalian cells. Thus, the genetic code supports the use of one codon to code for multiple amino acids. This finding challenged one of the foundations of the code, i.e., that only one genetic codeword is used for one amino acid in an organism.
7.2 High-Throughput Identification of Catalytic Redox-Active Cysteine Residues
Cys residues often play critical roles in proteins; however, identification of their specific functions has been limited to case-by-case experimental approaches. A recent study developed a procedure for high-throughput identification ofcatalytic redox-active Cys in proteins by searching for sporadic Sec/Cys pairs in sequence databases [80]. This method was independent of protein family, structure, and taxon. It was used to selectively detect the majority of known proteins with redox-active Cys and to make additional predictions, one of which was verified. Rapid accumulation of sequence information from genomic and metagenomic projects, coupled with selenoproteome analyses, should allow detection of many additional oxidoreductase families as well as identification of redox-active Cys in these proteins.
8 Analyses of Ionomes
Mechanisms that regulate Se and other trace elements in human cells were recently characterized by a genome-wide high-throughput siRNA/ionomics screen [81]. Se levels were controlled through the Sec machinery and expression of abundant selenoproteins. On the other hand, copper balance was affected by lipid metabolism and required machinery involved in protein trafficking and post-translational modifications, whereas the iron levels were influenced by iron import and expression of the iron/heme-containing enzymes.
A separate study examined Se and 17 other elements in the brain, heart, kidney, and liver of 26 mammalian species and reported the elemental composition of these organs, the patterns of utilization across the species, and their correlation with body mass and longevity [82]. Across the organs, distinct distribution patterns for abundant elements, transition metals, and toxic elements were observed. In this analysis, Se clustered with cadmium and arsenic, suggesting that Se toxicity is the property that is particularly important for organisms. Some elements showed lineage-specific patterns, including reduced Se utilization in African mole rats Heterocephalus glaber, and positive correlation between the number of Sec residues in SEPP1 and Se levels in liver and kidney across mammals. Interestingly, species lifespan correlated negatively with Se, suggesting that low utilization of Se may help achieve longevity. This study provided insights into the variation of Se levels in mammals according to organ physiology, lineage specialization, and longevity.
9 Concluding Remarks
Largely due to the remarkable progress in genomics, we now know which organisms utilize Se, and which do not, and which selenoproteins and Sec machinery mediate the Se utilization trait. Approximately 100 selenoprotein families are currently known, including 25 selenoprotein genes encoded in the human genome . This information allows researchers to study various aspects of Se biology and selenoprotein functions and address questions, not even imaginable until only recently, such as cross-species, cross-population and geographical distribution of Se and selenoprotein utilization and expansion of the genetic code . In selenoproteins with known functions, Sec is a key functional group that carries out redox catalysis. Further studies on selenoproteins and selenoproteomes should help explain known biological and biomedical effects of Se and identify new biological processes and pathways dependent on this trace element.
References
MD Wolfe et al 2004 J Biol Chem 279:1801
GL Dilworth 1982 Arch Biochem Biophys 219:30
VN Gladyshev et al 1994 Proc Natl Acad Sci USA 91:232
WT Self, TC Stadtman 2000 Proc Natl Acad Sci USA 97:7208
VM Labunskyy et al 2014 Physiol Rev 94:739
DL Hatfield et al 2014 Trends Biochem Sci 39:112
M Mariotti et al 2013 Nucleic Acids Res 41:e149
M Mariotti, R Guigó 2010 Bionformatics 26:2656
GV Kryukov et al 1999 J Biol Chem 274:33888
A Lescure et al 1999 J Biol Chem 274:38147
VN Gladyshev et al 2004 Annu Rev Nutr 24:579
FJ Martin-Romero et al 2001 J Biol Chem 276:29798
S Castellano et al 2001 EMBO Rep 2:697
GV Kryukov et al 2003 Science 300:1439
GV Kryukov, VN Gladyshev 2004 EMBO Rep 5:538
Y Zhang et al 2005 Genome Biol 6:R37
JT Rotruck et al 1973 Science 179:588
A Seiler et al 2008 Cell Metab 8:237
F Ursini et al 1999 Science 285:1393
GE Olson et al 2010 Am J Physiol Renal Physiol 298:F1244
S Gromer et al 1998 Biochem J 332:591
QA Sun et al 2001 Proc Natl Acad Sci USA 98:3673
T Sandalova et al 2001 Proc Natl Acad Sci USA 98:9533
QA Sun et al 1999 J Biol Chem 274:24522
ES Arner, A Holmgren 2000 Eur J Biochem 267:6102
QA Sun et al 2001 J Biol Chem 276:3106
AK Rundlof et al 2004 Free Radic Biol Med 36:641
D Su, VN Gladyshev 2004 Biochemistry 43:12177
D Su et al 2005 J Biol Chem 280:26491
M Conrad et al 2004 Mol Cell Biol 24:9414
C Jakupoglu et al 2005 Mol Cell Biol 25:1980
GV Kryukov et al 2002 Proc Natl Acad Sci USA 99:4245
HY Kim, VN Gladyshev 2004 Mol Biol Cell 15:1055
DE Fomenko et al 2009 J Biol Chem 284:5986
BC Lee et al 2013 Mol Cell 51:397
KV Korotkov et al 2001 J Biol Chem 276:15330
E Kumaraswamy et al 2000 J Biol Chem 275:35540
AD Ferguson et al 2005 J Biol Chem 281:3536
MV Kasaikina et al 2011 J Biol Chem 286:33203
TC Stadtman 1996 Annu Rev Biochem 65:83
XM Xu et al 2007 Biochem J 404:115
M Mariotti et al 2015 Genome Res 25:1256
RF Burk, KE Hill 2005 Annu Rev Nutr 25:215
AV Lobanov et al 2008 Genome Biol 9:R62
KE Hill et al 2003 J Biol Chem 278:13640
L Schomburg et al 2003 Biochem J 370:397
RF Burk, KE Hill 2015 Annu Rev Nutr 35:109
SC Vendeland et al 1995 Proc Natl Acad Sci USA 92:8749
KV Korotkov et al 2002 Mol Cell Biol 22:1402
SJ Han et al 2014 PLoS One 9:e95518
Y Ye et al 2004 Nature 429:841
VA Shchedrina et al 2011 J Biol Chem 286:42937
AA Turanov et al 2014 Biochem J 462:555
GJ Fredericks et al 2014 Proc Natl Acad Sci USA 111:16478
S Verma et al 2011 J Immunol 186:2127
B Moghadaszadeh et al 2001 Nat Genet 29:17
MJ Jurynec et al 2008 Proc Natl Acad Sci USA 105:12485
SV Novoselov et al 2002 EMBO J 21:3681
T Obata, Y Shiraiwa 2005 J Biol Chem 280:18462
S Castellano et al 2004 EMBO Rep 5:71
S Castellano et al 2005 Proc Natl Acad Sci USA 102:16188
SV Novoselov et al 2005 Biochem J 394:575
VA Shchedrina et al 2007 Proc Natl Acad Sci USA 104:13919
AV Lobanov et al 2006 Nucl Acids Res 34:496
ES Arnér 2010 Exp Cell Res 316:1296
HJ Reich, RJ Hondal 2016 ACS Chem Biol 11:821
AP Lothrop et al 2014 Biochemistry 53:664
AP Lothrop et al 2014 Biochemistry 53:654
BC Lee et al 2011 J Biol Chem 286:18747
AV Lobanov et al 2009 Biochim Biophys Acta 1790:1424
K Taskov et al 2005 Nucleic Acids Res 33:2227
AV Lobanov et al 2008 Protein Sci 17:176
CE Chapple, R Guigó 2008 PLoS One 3:e2968
CJ Gobler et al 2011 Proc Natl Acad Sci USA 108:4352
CJ Gobler et al 2013 ISME J 7:1333
AV Lobanov et al 2007 Genome Biol 8:R198
S Castellano et al 2009 Mol Biol Evol 26:2031
M Mariotti et al 2012 PLoS One 7:e33066
AA Turanov et al 2009 Science 323:259
DE Fomenko et al 2007 Science 315:387
M Malinouski et al 2014 Nat Commun 5:3301
S Ma et al 2014 Cell Rep 13:1319
Acknowledgements
This work was supported by grants from the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media, LLC
About this chapter
Cite this chapter
Gladyshev, V.N. (2016). Eukaryotic Selenoproteomes. In: Hatfield, D., Schweizer, U., Tsuji, P., Gladyshev, V. (eds) Selenium. Springer, Cham. https://doi.org/10.1007/978-3-319-41283-2_11
Download citation
DOI: https://doi.org/10.1007/978-3-319-41283-2_11
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-41281-8
Online ISBN: 978-3-319-41283-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)