Abstract
Gram-positive bacteria have a more ancient and primitive membrane structure than their Gram-negative counterparts which generally results in higher levels of intrinsic susceptibility to various lipophilic and amphiphilic antimicrobial drugs. Nonetheless, these bacteria encode similar numbers of efflux pumps in their respective genomes. In this chapter, we provide a historical overview of the identification and current understanding of such systems in Gram-positive genera of practical and industrial significance – including some clinically relevant organisms not covered elsewhere in this book. In general, these systems have been less thoroughly investigated than their Gram-negative counterparts with respect to transporter and substrate identification and their associated regulation. However, some key findings in the progression of the bacterial drug efflux field were first identified in less clinically relevant organisms such as Bacillus subtilis and Lactococcus lactis. Given this framework, the physiological relevance of efflux has become increasingly significant with concepts involving the innate immune response, metabolites, and bactericidal host-derived resistance and “natural” substrates.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Gram-positive bacteria
- Bacillus
- Clostridium
- Enterococcus
- Lactococcus
- Listeria
- Streptococcus
- Antimicrobial resistance
- Multidrug transporters
- P-glycoprotein
- ABC superfamily
- Major facilitator superfamily
1 Introduction
Model Gram-negative bacteria such as Escherichia coli and Pseudomonas aeruginosa encode a number of membrane efflux pumps that are responsible for significant levels of resistance to a variety of noxious compounds. In the E. coli genome alone, there are approximately 37 efflux pumps that belong to five different phylogenetic families and represent approximately 9 % of the encoded transporters [1–3]. The main constitutive pump, AcrB, covered in-depth elsewhere in this book, is a member of the resistance-nodulation-cell division (RND) superfamily and is a part of a system of three proteins that spans the inner membrane, periplasm, and outer membrane to coordinate efflux of amphiphiles simultaneously across both membranes into the extracellular milieu. This archetype complex alone is responsible for significant levels of efflux of several classes of antibiotics (β-lactams, macrolides, fluoroquinolones, etc.), dyes, natural and synthetic detergents (including bile acids), organic solvents, and even steroid hormones [4–6]. Furthermore, E. coli also encodes five other systems in this phylogenetic family that are poorly expressed except under specialized conditions, usually as a result of upregulation via two-component sensor kinases. Such systems have been extensively characterized in P. aeruginosa, a more clinically significant microbe, but are slightly more complex. In fact, several Gram-negative bacteria of various genera have been shown to chromosomally encode AcrB homologs [7].
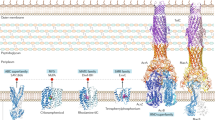
On the contrary, Gram-positive efflux-based resistance is less well reported in the literature. Nevertheless, genomic analysis suggests that Gram-positive bacteria encode as many putative multidrug transporters as Gram-negative. Thus, the Enterococcus genome reveals 34 potential drug efflux-related genes [8]. Likewise, further comparative genomics of 11 Gram-positive bacteria of importance to the health and food industry is generally similar to E. coli in genomic prevalence ranging around 10 %, the exception being Bacillus subtilis in which 17 % of its transporter cadre is putatively dedicated to drug and toxic compound extrusion [1]. Gram-positive bacteria have a cell envelope that consists of a single phospholipid bilayer, surrounded by a thick layer of peptidoglycan. Consequently, unlike multidrug transporters of Gram-negative bacteria which often assemble into multicomponent complexes designed to span both membranes, multidrug transporters of Gram-positive bacteria have only a single transmembrane component. Since the archetypal RND superfamily transporter, the major clinically relevant superfamily of multidrug transporters in Gram-negative bacteria, consists of three components spanning both inner and outer membrane, it was believed for a long time that there are no RND-type multidrug transporters in Gram-positive bacteria. In 2001, this view was modified when YerP, a transporter of the RND superfamily, was identified in B. subtilis [9]. However, the number of multidrug efflux pumps belonging to RND superfamily in Gram-positive bacteria (except mycobacteria) is generally very limited [10]. Four other efflux families or superfamilies found in Gram-negative bacteria are well represented in Gram-positive bacteria: the major facilitator superfamily (MFS), the small multidrug resistance (SMR) family, the multidrug and toxic compound extrusion (MATE) family, and the ATP-binding cassette (ABC) superfamily [11]. Considering the vast difference in membrane structure between Gram-negatives and Gram-positives [12, 13], it is important to determine whether such proteins can contribute to similar levels of intrinsic resistance in this, a more ancient division of bacteria.
Identification of multidrug transporters remains quite a challenge even today, in a post-genomic era. Multidrug transporters lack definitive signatures for substrate specificity, so while it is possible to identify putative multidrug transporters by analysis of the genomic DNA sequence, their ability to transport multiple unrelated compounds has to be confirmed experimentally. Thus, putative SMR family proteins PsmrAB were cloned in E. coli from metagenomic DNA from a halophilic environment and were found to function as a two-component Na+/H+ antiporter, rather than involved in resistance to drugs [14]. Also, often, multidrug transporters are not expressed under physiological conditions, and whereas some of them are activated by their substrates, for many, the activator(s) are still not known. These transporters need to be overexpressed in order to confirm their identity, but for many bacteria, the difficulties in culturing or genetic manipulation and the availability of genetic tools make overexpression or disruption of the gene very difficult. In this chapter, we review the presently characterized multidrug efflux pumps in Gram-positive bacteria.
2 Bacillus subtilis
B. subtilis, while of little clinical importance, is an excellent model organism, easily cultured and with a lot of genetic tools available. In addition, B. subtilis has significant genomic abundance of multidrug transporters relative to other Gram-positives [1]. For these reasons, these B. subtilis transporters have been extensively studied. In fact, the phenomenon of bacterial multidrug resistance was first discovered in B. subtilis [15]. Authors hypothesized the presence of a mechanism analogous to the mammalian multidrug transporter, P-glycoprotein. Indeed, multidrug-resistant cells were obtained after selection with increasing concentrations of one of the substrates of P-glycoprotein, rhodamine 6G. These cells exhibited resistance to some other known substrates of P-glycoprotein, such as ethidium bromide, chloramphenicol, and puromycin, as well as to tetraphenylphosphonium and cetyltrimethylammonium bromide, which are not transported by P-glycoprotein [15]. The mechanism of resistance was shown to be efflux based and was sensitive to the same inhibitors, reserpine and verapamil, as mammalian P-glycoprotein. These cells were used to clone the first bacterial multidrug transporter, Bmr, whose gene was found to be amplified in resistant cells. Analysis of the Bmr sequence, however, showed little similarity with P-glycoprotein. Indeed, Bmr is a multidrug transporter of the MFS and is very different from an ABC transporter P-glycoprotein. It was shown to use a different energy source – secondary-active transport with the transmembrane pH gradient – whereas P-glycoprotein couples the transport of substrates with primary-active ATP hydrolysis [16, 17]. Later, a second multidrug transporter of the MFS, Blt, was identified in B. subtilis [18].
Blt is 51 % identical to Bmr and transports a similar set of compounds; however, the pattern of their expression is quite different. Bmr is expressed under standard cultivating conditions and is further regulated by BmrR, a member of the family of MerR-like transcriptional activators [19]. BmrR activates the expression of Bmr after binding its substrates. In contrast, Blt expression is normally not detectable. It is regulated by BltR [18], which is related to BmrR, but has a different inducer-binding domain, and its substrates are not yet known. Blt is cotranscribed with a downstream gene encoding spermine-spermidine acetyltransferase, indicating physiological function(s) apart from synthetic drug resistance per se – a theme central to efflux systems in Gram-positive and Gram-negative systems alike [11, 20]. Another layer of regulation was reported later for Bmr and Blt [21]. Their expression was found to be further controlled by a MerR-type regulator Mta. Apo-Mta acted as a repressor of the bmr and blt gene transcription. Although Mta inducer was not identified in this report, Mta was converted into transcriptional activator by the removal of the C-terminal inducer-binding domain. The authors proposed that this removal mimics the binding of inducer to Mta [21].
Several more multidrug transporters were identified in B. subtilis. In 1996, a stunning discovery was made of a first bacterial multidrug transporter of the ABC family, LmrA from Lactococcus lactis [22]. LmrA was homologous to both halves of the mammalian P-glycoprotein which is arranged in a 6 + 6 transmembrane motif [23] common to many multidrug transporters of different families [24]. Subsequently, two multidrug transporters of ABC family, BmrA and BmrC/BmrD, which functions as heterodimer, were identified in B. subtilis. BmrA was first identified from genome sequencing of B. subtilis [25] and demonstrated to transport Hoechst33342 (a fluorescent dye used to stain DNA), doxorubicin, and 7-aminoactinomycin D in highly enriched inverted membrane vesicles from E. coli [26]. BmrC/BmrD was likewise shown in the same system to transport Hoechst 33342, doxorubicin, BCECF (2′,7′-bis-(2-carboxyethyl)-5-(and-6)-carboxyfluorescein, fluorescent compound) and mitoxantrone [27]. BmrC/BmrD expression is regulated, first, by the main transcription phase regulator AbrB, and second, via a dedicated ribosome-mediated transcriptional attenuation mechanism that requires the bmrB-encoded leader peptide [28]. Another substrate-specific ABC transporter deserves mentioning here because of its clinical relevance. BceAB is positively regulated by BceRS two-component regulatory system and contributes to intrinsic resistance to bacitracin [29, 30].
Additional work identified Bmr3 [31], a member of the MFS. Bmr3 was shown to transport puromycin, norfloxacin, and tosufloxacin which are also substrates of the Bmr and Blt. Bmr3 expression is growth phase dependent and is drastically reduced as the cells enter late log phase. A spontaneous multidrug-resistant mutant selected by puromycin exhibited an increased stability of bmr3 transcripts [32]. In addition, MdtP is a multidrug transporter of the MFS that contributes to resistance to actinomycin, fusidic acid, novobiocin, and streptomycin [33]. MdtP expression is induced by its substrate fusidic acid, which binds to repressor MdtR (whose encoding gene is cotranscribed with mdtP) and causes its dissociation from the mdtP promoter. YerP is the sole RND-type multidrug transporter characterized in Gram-positive bacteria (except mycobacteria) and has been shown to be involved in resistance to acriflavine, ethidium bromide, and surfactin, a cyclic lipopeptide biosurfactant synthesized by some species of B. subtilis [9]. Finally, EbrAB is a paired multidrug transporter which belongs to the SMR family and functions as heterooligomer. Its overexpression in B. subtilis confers resistance against acriflavine, ethidium bromide, pyronine Y, and safranin O [34].
3 Clostridium difficile
C. difficile is a major cause of nosocomial diarrhea. It is also implicated in 95 % of pseudomembranous colitis cases [35]. This species is intrinsically less susceptible to antibiotics, in particular β-lactams, fluoroquinolones, chloramphenicol, and lincosamides, than the other clostridia. Active efflux was long thought to be responsible for this resistance; however, the first description of multidrug transporter from C. difficile dates to 2004, when Dridi et al. [36] characterized CdeA, an MATE family transporter. When overexpressed in hypersensitive strain of E. coli, CdeA caused resistance to acriflavine, ethidium bromide, ciprofloxacin, and norfloxacin. CdeA was shown to cause energy-dependent efflux of ethidium bromide in E. coli cells, and quantitative reverse transcription-PCR assay showed that cdeA expression in C. difficile was significantly increased by exposure to ethidium bromide, but not to ciprofloxacin. The authors could not test the effect of CdeA inactivation in C. difficile, due to inability to genetically transform this species. The same year, Lebel et al. [37] identified four genes in C. difficile encoding putative proteins homologous to NorA from Staphylococcus aureus, a multidrug transporter of the MFS. Of these sequences, only one, designated cme, conferred resistance to ethidium bromide, safranin O, and erythromycin when expressed in Enterococcus faecalis. It was not known whether the lack of effect of the other three open reading frames was due to inefficient expression in a different species, to inappropriate substrates, or to inactivity of these open reading frames as multidrug transporters.
4 Listeria monocytogenes
L. monocytogenes is an important food-borne pathogen that can cause such severe diseases as septicemia, meningitis, stillbirth, and abortion. High-risk groups include immunocompromised patients, neonates, and pregnant women [38]. Infection with L. monocytogenes causes significant mortality and morbidity in these groups. L. monocytogenes invades host cells and replicates within their cytoplasm [39]. Acquired antimicrobial resistance in L. monocytogenes is a very rare event. However, this pathogen is intrinsically resistant to several antimicrobial agents and also to the bile which has bactericidal properties and to which it is exposed during several stages of its lifecycle in the human gastrointestinal tract. In addition, multidrug transporters play a fascinating role in interaction of Listeria with mammalian innate immunity during its infection cycle. For these reasons, information about multidrug transporters in Listeria is highly clinically significant.
The first multidrug transporter in Listeria spp., MdrL, was identified serendipitously while sequencing genomic region around a gene encoding a putative histone-like protein, flaR, in an effort to find genes implicated in its regulation [40]. The authors identified a gene similar to a number of multidrug transporters of the MFS. Disruption of the allele in the wild-type strain of L. monocytogenes resulted in a small but reproducible increase in susceptibility to erythromycin, josamycin, clindamycin, and heavy metals and about a tenfold increase in susceptibility to cefotaxime. Functional characterization of MdrL as an efflux pump was confirmed by observing reserpine-dependent inhibition of ethidium bromide efflux, which was virtually eliminated in the MdrL disruption mutant.
Two other multidrug transporters of MFS family present in L. monocytogenes, MdrM and MdrT, were identified based on predicted protein sequence similarity [41]. To date, no experimental work addressing directly and conclusively their function as multidrug transporters has been published, but there is a significant indirect supporting evidence. The expression of these genes, as well as of mdrL, is induced by common multidrug transporter substrates. MdrL, MdrM, and MdrT were shown to be regulated by repressors LadR, MarR, and BrtA (previously TetR), respectively [42–44]. These repressors are encoded adjacently to the corresponding multidrug transporter genes. The expression of MdrL is induced by rhodamine 6G in the LadR-dependent fashion [43]. The transcription of mdrM and mdrT is also upregulated in response to rhodamine 6G and tetraphenylphosphonium, although the involvement of the aforementioned repressors in their activation by these compounds has not been demonstrated [42]. Cholic acid, another common multidrug transporter substrate, was shown to bind BrtA and cause its dissociation from the mdrT promoter, resulting in the induction of mdrT transcription [44]. Moreover, MdrT was shown to transport cholic acid out of the cells [44]. There is significant cross-regulation among these genes. A ladR mutant upregulates not only mdrL but also mdrM [42]. In response to the bile acid and cholic acid, BrtA upregulates not only mdrT but mdrM as well [44].
The most fascinating function of MdrM and MdrT was described in the Portnoy laboratory [42, 45]. The authors showed that these multidrug transporters control the magnitude of the host cytosolic innate immune response to L. monocytogenes. On the entry into the host cytosol, L. monocytogenes activates host response that leads to transcription of dozens of genes, including robust expression of interferon beta (IFN-β) [46, 47]. MdrM and MdrT expression was shown to affect the induction of IFN-β in infected macrophages [42, 48, 49]. Disruption of mdrM [42] or mdrT in the strain with mutated brtA [49] decreased IFN-β production, while overexpression of either MdrM or MdrT resulted in increased induction of IFN-β in infected macrophages [42]. The molecule that triggers the cytosolic host response was shown to be the cyclic dinucleotide c-di-AMP [45]. This molecule is produced by many bacteria and is a second messenger that is implicated in a variety of functions including cell wall metabolism, potassium homeostasis, DNA repair, and control of gene expression [50]. C-di-AMP in L. monocytogenes is secreted by MdrM and MdrT [45]. It is sensed by the cytosolic innate immune receptor, STING [51]. Stimulation of this pathway results in the activation of the interferon regulatory factor-3 and nuclear factor-kB transcription factors and, ultimately, to host transcriptional activation of IFN-β [46, 51]. While innate immune system is indispensable for defense against microbial pathogens, paradoxically, the production of IFN-β increases the bacterial burden and lethality of L. monocytogenes infection in mouse models [52–54], through mechanisms that are not well understood, but may involve the enhanced susceptibility of lymphocytes to apoptosis in response to a pore-forming toxin and a major virulence factor of L. monocytogenes, listeriolysin O [53, 54].
Finally, AnrAB is an ABC-type multidrug transporter that was isolated by screening for nisin-sensitive mutants of L. monocytogenes [55]. A mutant strain exhibited enhanced susceptibility to nisin, gallidermin, cefuroxime, cefotaxime, ampicillin, penicillin G, and bacitracin.
5 Lactococcus lactis
L. lactis is broadly used for food manufacturing. Despite a few case reports of L. lactis being an opportunistic pathogen [56], it is a generally regarded as safe organism. However, the “resistance gene reservoir” hypothesis suggests that beneficial and commensal bacterial populations in gastrointestinal tract may play a role in horizontal transfer of antimicrobial resistance to pathogenic microorganisms [57].
Initially, only secondary, proton motive force-driven multidrug transporters were described in bacteria, when, in 1994, Bolhius et al. [58] reported isolation of three mutants of L. lactis, selected for resistance to high concentrations of ethidium bromide, daunomycin, or rhodamine 6G. These mutants were found to be cross-resistant to a number of structurally and functionally unrelated drugs, such as quinine, actinomycin D, and gramicidin D. The drug resistance of these strains was due to energy-dependent efflux and was inhibited by reserpine, a multidrug efflux pump inhibitor. Efflux was also inhibited by orthovanadate (an inhibitor of ATP-dependent efflux activity characteristic of ABC transporters) in one of the strains, and in two others, it was partially inhibited both by orthovanadate and by nigericin (an ionophore). This observation suggested that a proton motive force-dependent and ATP-dependent systems were involved in drug efflux. A year later, a lactococcal proton motive force-dependent multidrug efflux pump, LmrP, was characterized in the same laboratory [59]. LmrP was cloned in E. coli and was shown to belong to the MFS. In E. coli, its substrates included ethidium bromide, daunomycin, and tetraphenylphosphonium, which were transported in a proton gradient-dependent manner. Overexpression of lmrP in L. lactis resulted in elevated resistance to ethidium bromide; however, an lmrP deletion mutant was only slightly more susceptible to ethidium bromide than the wild-type strain. The resistance of the lmrP-deficient strain to ethidium bromide could be significantly decreased by treating the cells with orthovanadate. This observation confirmed that an ATP-dependent multidrug transporter was functional in L. lactis.
In 1996, LmrA, the first bacterial ATP-dependent multidrug transporter, was characterized in the same laboratory [22]. LmrA was homologous to the human mdr1, which encoded the P-glycoprotein and, moreover, complemented MDR1 in human lung fibroblast cells [60]. LmrA was targeted to the plasma membrane and conferred typical multidrug resistance in these human cells. Blockers of P-glycoprotein-mediated multidrug resistance also inhibited LmrA-dependent drug resistance. Like P-glycoprotein, LmrA removed drugs from the inner leaflet of the cytoplasmic membrane [61]. The expression of lmrA in a hypersensitive E. coli strain increased resistance to the very wide variety of drugs, including aminoglycosides, chloramphenicol, β-lactams, lincosamides, macrolides, quinolones, streptogramins, and tetracyclines [62].
LmrA is equivalent to half of the P-glycoprotein and functions as homodimer. Later, however, a functional heterodimeric ABC-type multidrug transporter LmrCD was described in L. lactis [63]. LmrC and LmrD were copurified as a heterodimer, and overexpression of both LmrC and LmrD in LmrA-negative strain of L. lactis demonstrated ATP-dependent efflux of ethidium bromide, BCECF-acetoxymethyl ester, daunomycin, and Hoechst 33342. As a corollary, the cells did not show drug extrusion when either gene was overexpressed singly. LmrCD is also responsible for bile resistance [64]. The expression of lmrCD is controlled by transcriptional repressor LmrR, encoded upstream of the lmrCD [65, 66]. LmrR also autoregulates its own expression. LmrR binds the LmrCD substrates: Hoechst 33342, daunomycin, and rhodamine 6G [65, 67]. Drug binding to LmrR relieves the LmrR-dependent repression of the lmrCD genes [68]. Interestingly, when four mutant multidrug-resistant strains of L. lactis selected by challenging with increasing concentrations of daunomycin, ethidium bromide, rhodamine 6G, or cholate were analyzed, only lmrCD multidrug transporter genes were significantly and strongly upregulated in all four strains [69]. These data suggested that LmrCD was a major determinant of multidrug resistance in L. lactis. This study, however, did not address the expression of other putative multidrug transporters in mutant strains. Finally, in 2013, CmbT was characterized as an MFS-type multidrug transporter [70]. Overexpression of cmbT in L. lactis resulted in marginally increased resistance to cholate, ethidium bromide, Hoechst 33342, lincomycin, puromycin, rifampicin, streptomycin, sulbactam, sulfadiazine, and sulfamethoxazole (IC50 increased approximately 1.2–3 times). Overexpressed CmbT mediated extrusion of ethidium bromide and Hoechst 33342, and ionophores inhibited the CmbT-mediated transport of Hoechst 33342. Based on the increased level of thiol groups in supernatant of strain overproducing CmbT, the authors hypothesized possible involvement of CmbT in sulfur metabolism [70]. However, this observation was dependent on methionine and cysteine content of the medium and was not further investigated in this report. In addition to LmrP, LmrA, LmrCD, and CmbT, the genome of L. lactis contains 36 putative multidrug transporters; however, they are still to be characterized experimentally. Additionally, a multidrug transporter of the ABC superfamily, LmrB, was identified on a plasmid carried by a natural isolate of L. lactis [71]. LmrB was shown to be an active multidrug transporter capable of the extrusion from the cell of ethidium bromide and Hoechst 3342. Interestingly, two genes encoding polypeptidic bacteriocins LsbA and LsbB are located on the same plasmid as LmrB, in the immediate vicinity of the multidrug transporter gene. LmrB was shown to render the cells immune to both bacitracins, and to mediate their secretion into the medium. In this function, LmrB could be complemented by LmrA but not LmrP [71]. The location of the lmrB gene on a plasmid may facilitate transfer of this multidrug transporter from L. lactis to pathogenic bacteria and may deserve further investigation.
6 Enterococcus spp.
The enterococci are commensal bacteria that normally populate the human intestine. Over the last two decades, enterococci were identified as causative agents of nosocomial infections with increasing frequency. Infections caused by enterococci include urinary tract infections, nosocomial bacteremia, intra-abdominal infections, and endocarditis. Most enterococci have intrinsic resistance to various antimicrobial agents. However, increasingly frequent isolation of enterococci with acquired resistance to most commonly used drugs has been observed in recent years. As early as 1997, Lynch et al. [72] hypothesized that intrinsic resistance of enterococci to various antimicrobial agents, in the absence of outer membrane, is due, at least in part, to active efflux system(s). They examined four wild-type strains of E. faecalis and a strain of Enterococcus faecium and found that all strains showed energy-driven efflux of chloramphenicol, and all but one strain of E. faecalis extruded norfloxacin. In contrast, active efflux did not play a role in resistance to β-lactams. In this work, genetic determinants of these efflux pumps were not identified. Four years later, enterococcal genome-scanning identified a potential multidrug transporter EmeA [73] due to its homology to NorA from S. aureus, MFS-type multidrug transporter covered in-depth in Chap. 7 in this book. Deletion of this gene in E. faecalis resulted in an approximately twofold increase in susceptibility to acriflavine, ethidium bromide, clindamycin, erythromycin, novobiocin, ciprofloxacin, and norfloxacin compared to the wild-type strain. Functional complementation with wild-type plasmid-expressed emeA restored the resistance to ethidium bromide and resulted in the resistance to norfloxacin fourfold higher than in the wild-type strain. This resistance was due to energy-dependent efflux. Incubation with reserpine (competitive multidrug transporter blocker), verapamil (a calcium channel blocker), or lansoprazole (a H+ and K+-ATPase pump inhibitor) decreased resistance of both wild-type and complemented strains. The resistance of the mutant strain was unaffected by these agents, except for resistance to ethidium bromide which was lowered twofold by reserpine. These data allowed the authors to conclude that EmeA was the main enterococcal pump for these agents.
Later, Lee et al. [74] cloned EfrAB, an ABC multidrug transporter from E. faecalis, by using a drug-hypersusceptible mutant of E. coli host. When expressed in E. coli, EfrAB conferred resistance to norfloxacin, ciprofloxacin, doxycycline, arbekacin, novobiocin, daunorubicin, doxorubicin, acriflavine, 4′,6-diamidino-2-phenylindole, ethidium bromide, safranin O, and tetraphenylphosphonium. Furthermore, EfrAB demonstrated energy-dependent efflux of acriflavine. This efflux was inhibited by verapamil, reserpine, and sodium orthovanadate (an ATPase inhibitor). Similar to other two-component ABC multidrug transporters, both EfrA and EfrB were required for resistance. The expression of EfrAB is induced by subinhibitory concentrations of chloramphenicol, gentamicin, and streptomycin [75]. In the same laboratory, E. faecium multidrug transporter belonging to the MFS, EfmA, was cloned in a similar fashion [76]. E. coli harboring EfmA showed energy-dependent efflux of 4′,6-diamidino-2-phenylindole and tetraphenylphosphonium, as well as norfloxacin/H+ antiport. EfmA was found to be constitutively expressed by E. faecium. Overall, 34 putative multidrug transporters in E. faecalis have been identified from genome sequencing [8]. However, the majority of them are still experimentally unexplored.
7 Streptococcus spp.
7.1 Streptococcus pneumoniae
S. pneumoniae is the main bacterial cause of community-acquired pneumonia and represents a major disease burden worldwide [77]. Despite the recent introduction of the heptavalent pneumococcal conjugate vaccine, antimicrobial resistance is an increasing problem in this organism due to the spread of multidrug-resistant clones and increases in antimicrobial resistance among nonvaccine serotypes [78].
The initial report of the multidrug transporter in S. pneumoniae did not identify the transport protein associated with the phenotype [79]. A few years later, PmrA, a multidrug efflux pump of the MFS family, was identified using S. pneumoniae genomic sequence as homologous to norA of S. aureus [80]. The gene was overexpressed in S. pneumoniae and found to confer resistance to norfloxacin, ciprofloxacin, acriflavine, and ethidium bromide [80]. In later reports, a knock-out of pmrA [81, 82] did not result in increased susceptibility to drugs, indicating that PmrA is not intrinsically active in S. pneumoniae.
By 2006, the overexpression of the ABC superfamily efflux proteins PatA and PatB was found to be responsible for the multidrug-resistant phenotype of a mutant of S. pneumoniae selected after exposure to ciprofloxacin [83]. Disruption of patA and patB resulted in increased sensitivity to acriflavine, ethidium bromide, berberine, erythromycin, oxolinic acid, norfloxacin, ciprofloxacin, and novobiocin [81, 82] thus demonstrating that PatAB is normally expressed by S. pneumoniae. Each subunit consists of a nucleotide-binding domain and a membrane spanning domain, and heterodimerization of PatA and PatB is required to form a functional transporter [84]. Expression of patAB is induced by subinhibitory concentrations of fluoroquinolones [85, 86]. In clinical fluoroquinolone-resistant isolates of S. pneumoniae, whose resistance is ascribable to the overexpression of multidrug transporters, either PmrA [87] or PatA/PatB [88], were found to be responsible for the phenotype. Similar to other Gram-positive examples covered in this chapter, additional putative multidrug transporters exist in the genome of S. pneumoniae, but so far, no phenotype was associated with them [82].
7.2 Streptococcus agalactiae
S. agalactiae causes neonatal sepsis, pneumonia, meningitis, as well as infections of the bovine udder. S. agalactiae produces α-hemolysin, which is an important virulence factor. It is capable of damaging erythrocytes, lung epithelial cells [89], and brain microvascular endothelial cells [90], which is regarded as an initial step in invasive disease. cylA and cylB were identified as genes essential for the production of the S. agalactiae hemolysin [91] and encode an ABC-type transporter. These genes are part of the 12-gene cyl operon, which contains, in addition to cylA and cylB, 5 genes similar to fatty acid biosynthesis enzymes (cylD, cylG, acpC, cylZ, and cylI), one similar to an aminomethyltransferase (cylF), one carrying the conserved domain of a glycosyltransferase (cylJ), a gene predicted to encode an acetyl coenzyme A carboxylase (cylX), a putative phosphopantetheinyl transferase (cylK), and a putative acyl-coA acyltransferase (cylE) [92]. cylA and cylB deletion mutants resulted in a nonhemolytic phenotype [93]. cylA mutant was shown to still harbor intracellular hemolytic activity, which was released by sonication. Since CylAB contained the signature sequence of a multidrug resistance transporter, wild-type and nonhemolytic cylA mutant were exposed to known substrates of multidrug transporters. Deletion of cylA resulted in significant increase in susceptibility to daunorubicin, doxorubicin, and rhodamine 6G. Furthermore, the cylA-negative strain displayed a markedly reduced capacity to export doxorubicin. Growth in the presence of reserpine resulted in a dose-dependent decrease of extractable hemolytic activity, supporting the hypothesis that hemolysin is transported out of the cell by a multidrug transporter. At the time, the nature of the S. agalactiae hemolysin was unknown, and it was believed to be a pore-forming protein toxin. However, Gottschalk et al. [93] raised doubts in the protein nature of hemolysin based on the described work. Multidrug transporters were known to transport small molecules, rather than proteins. Indeed, in 2013, Whidbey et al. [94] showed that an ornithine rhamnolipid pigment known as granadaene [95] is responsible for the hemolytic activity of the bacterium. This is perhaps one of the few cases where the nature of the natural substrate of the multidrug transporters is proven very strongly.
7.3 Streptococcus mutans
S. mutans is a major causative agent in human caries and forms biofilm known as dental plaque [96, 97]. Although S. mutans strains are generally susceptible to antimicrobial agents [98], prolonged antimicrobial exposure can select antimicrobial resistance [99, 100]. Involvement of multidrug transporters in drug resistance has been reported [101, 102]. Genome of S. mutans UAB159 [103] shows the presence of 71 putative ABC transporters and 10 putative MFS transporters (TransportDB at http://www.membranetransport.org).
Increased susceptibility to methyl viologen (paraquat), benzyl and ethyl viologens, and quaternary ammonium compounds was observed in mutant strains that were deficient in a function of ABC transporter complex, VltAB, which is encoded by an operon (SMU.905-906) [104]. The same laboratory also reported another putative ABC transporter complex, SmbFT, which is not present in strain UAB159 [105] and is encoded by genes located in the same locus and provides protection against lantibiotics Smb and haloduracin, but not against other lantibiotics (e.g., nisin) and several peptide antibiotics such as bacitracin, polymyxin B, and vancomycin [106]. A newer study also described two ABC transporter systems, SMU.654-655-656-657 and LctFEG (SMU.1148-1149-1150), which are, respectively, encoded by the genes linked to the two-component regulatory genes nsrRS (located downstream of SMU.654-655-656-657) and lcrRS (located upstream of lctFEG). Inactivation of nsrRS or nsrS (but not SMU.654-655-656-657) rendered the mutant strains more susceptible to nisin A (16-fold MIC reduction), while disruption of lcrRS, lcrS, or lcrFEG increased the susceptibility of the mutants to nukacin (eightfold MIC decrease) [101], suggesting involvement of LctFEG transporter in nukacin resistance. In S. mutans and Streptococcus gordonii (a commensal species), the gene locus rcrRPQ encodes an MarR-like transcriptional regulator (RcrR) and an ABC efflux complex (RcrPQ), which are linked to stress tolerance [107–109]. Inactivation of rcrP in S. mutans or rcrR in S. gordonii rendered the mutant cells more susceptible to lower pH or oxidative stress agents such as H2O2 and methyl viologen [107, 109]. Either overproduction of or deficiency in RcrPQ impaired the biofilm formation in S. mutans [107], suggesting an optimal status of this ABC exporters is essential for biofilm formation. In this regard, another exporter, the NrgA ammonium transporter, is also essential for biofilm formation in S. mutans [110]. Similar to that observed in B. subtilis [29], an ABC transporter named BceAB is encoded by part of the four-gene operon bceABRS that also encodes the BceRS two-component regulatory system, and BceABRS contributes in response to bacitracin-induced cell envelope stress [111]. Recently, the copYAZ operon (SMU.424-426-427) that encodes CopA ABC copper exporter and CopYZ regulators was demonstrated to play an important role in copper homeostasis, stress tolerance, and biofilm formation [112].
Additionally, a multidrug transporter, MdeA of the MFS predicted with 12 transmembrane domains, conferred resistance to ampicillin, oxacillin, nalidixic acid, ciprofloxacin, kanamycin, tetracycline, acriflavine, and rhodamine 6G (4- to 32-fold increase in MIC values) when expressed on a plasmid in a hypersusceptible E. coli host [102]. It is also noted that several regulatory systems such as LytST and ScnRK have been reported to contribute to tolerance to oxidative stress [113, 114]. Whether these regulatory systems are linked to any multidrug transporters remains to be investigated (Table 8.1).
8 Concluding Remarks
Gram-positive organisms are generally more susceptible to antimicrobial drugs than their Gram-negative counterparts. Typical model archetypes for drug efflux have been more well developed genetically, biochemically, and structurally in Gram-negative organisms. This research bias may be generally attributable to the relative contributions of tripartite efflux systems to clinically significant drug resistance phenotypes as targets for inhibitory compounds with potential dramatic modulation of drug resistance phenotypes. From a historical perspective, however, it is significant that some of the first bacterial drug efflux systems were identified and characterized in Gram-positive organisms (with little clinical significance) but noteworthy genetic and functional conservation with the major mammalian multidrug transporter, P-glycoprotein. However, counterintuitive to membrane evolution and drug kinetics, Gram-positive organisms encode a similar, if not greater, genomic investment (in the case of Bacillus subtilis) to efflux-based transport mechanisms. This observation suggests efflux may be physiologically and functionally more essential and, hence, more intrinsically active in Gram-positive organisms to accommodate their respective environments and survival. Further research into pump regulation and expression-level comparisons may be useful to determine relative functional balance between efflux and cell energetics in context with cell physiology and metabolism. In this regard, Gram-positive organisms described herein may be instructive in identifying physiologically relevant substrates and roles for respective efflux systems in the host-bacterial interface. The increasing availability of genetic tools in these organisms will facilitate genetic manipulation required to conduct such studies and in vivo modeling.
References
Lorca GL, Barabote RD, Zlotopolski V, Tran C, Winnen B, Hvorup RN, Stonestrom AJ, Nguyen E et al (2007) Transport capabilities of eleven Gram-positive bacteria: comparative genomic analyses. Biochim Biophys Acta 1768:1342–1366. doi:10.1016/j.bbamem.2007.02.007
Nishino K, Yamaguchi A (2001) Analysis of a complete library of putative drug transporter genes in Escherichia coli. J Bacteriol 183:5803–5812. doi:10.1128/JB.183.20.5803-5812.2001
Sulavik MC, Houseweart C, Cramer C, Jiwani N, Murgolo N, Greene J, DiDomenico B, Shaw KJ et al (2001) Antibiotic susceptibility profiles of Escherichia coli strains lacking multidrug efflux pump genes. Antimicrob Agents Chemother 45:1126–1136. doi:10.1128/AAC.45.4.1126-1136.2001
Ma D, Cook DN, Alberti M, Pon NG, Nikaido H, Hearst JE (1995) Genes acrA and acrB encode a stress-induced efflux system of Escherichia coli. Mol Microbiol 16:45–55. doi:10.1111/j.1365-2958.1995.tb02390.x
Elkins CA, Nikaido H (2002) Substrate specificity of the RND-type multidrug efflux pumps AcrB and AcrD of Escherichia coli is determined predominantly by two large periplasmic loops. J Bacteriol 184:6490–6498. doi:10.1128/JB.184.23.6490-6499.2002
Elkins CA, Mullis LB (2006) Mammalian steroid hormones are substrates for the major RND- and MFS-type tripartite multidrug efflux pumps of Escherichia coli. J Bacteriol 188:1191–1195. doi:10.1128/JB.188.3.1191-1195.2006
Li X-Z, Plésiat P, Nikaido H (2015) The challenge of efflux-mediated antibiotic resistance in Gram-negative bacteria. Clin Microbiol Rev 28:337–418. doi:10.1128/CMR.00117-14
Davis DR, McAlpine JB, Pazoles CJ, Talbot MK, Alder EA, White C, Jonas BM, Murray BE et al (2001) Enterococcus faecalis multi-drug resistance transporters: application for antibiotic discovery. J Mol Microbiol Biotechnol 3:179–184
Tsuge K, Ohata Y, Shoda M (2001) Gene yerP, involved in surfactin self-resistance in Bacillus subtilis. Antimicrob Agents Chemother 45:3566–3573. doi:10.1128/AAC.45.12.3566-3573.2001
Serizawa M, Sekiguchi J (2005) The Bacillus subtilis YdfHI two-component system regulates the transcription of ydfJ, a member of the RND superfamily. Microbiology 151:1769–1778. doi:10.1099/mic.0.27619-0
Li X-Z, Nikaido H (2009) Efflux-mediated drug resistance in bacteria: an update. Drugs 69:1555–1623. doi:10.2165/11317030-000000000-00000
Navarre WW, Schneewind O (1999) Surface proteins of Gram-positive bacteria and mechanisms of their targeting to the cell wall envelope. Microbiol Mol Biol Rev 63:174–229
Nikaido H (2003) Molecular basis of bacterial outer membrane permeability revisited. Microbiol Mol Biol Rev 67:593–656. doi:10.1128/MMBR.67.4.593-656.2003
Jiang J, Wang L, Zhang H, Wu H, Huang H, Yang L (2013) Putative paired smallmultidrug resistance family proteins PsmrAB, the homolog of YvdSR, actually function as a novel two-component Na+/H+ antiporter. FEMS Microbiol Lett 338:31–38. doi:10.1111/1574-6968.12008
Neyfakh AA, Bidnenko VE, Chen LB (1991) Efflux-mediated multidrug resistance in Bacillus subtilis: similarities and dissimilarities with the mammalian system. Proc Natl Acad Sci U S A 88:4781–4785
Ambudkar SV, Lelong IH, Zhang J, Cardarelli CO, Gottesman MM, Pastan I (1992) Partial purification and reconstitution of the human multidrug-resistance pump: characterization of the drug-stimulatable ATP hydrolysis. Proc Natl Acad Sci U S A 89:8472–8476. doi:10.1073/pnas.89.18.8472
Hamada H, Tsuruo T (1988) Characterization of the ATPase activity of the Mr 170,000 to 180,000 membrane glycoprotein (P-glycoprotein) associated with multidrug resistance in K562/ADM cells. Cancer Res 48:4926–4932
Ahmed M, Lyass L, Markham PN, Taylor SS, Vazquez-Laslop N, Neyfakh AA (1995) Two highly similar multidrug transporters of Bacillus subtilis whose expression is differentially regulated. J Bacteriol 177:3904–3910
Ahmed M, Borsch CM, Taylor SS, Vazquez-Laslop N, Neyfakh AA (1994) A protein that activates expression of a multidrug efflux transporter upon binding the transporter substrates. J Biol Chem 269:28506–28513
Piddock LJ (2006) Multidrug-resistance efflux pumps – not just for resistance. Nat Rev Microbiol 4:629–636. doi:10.1038/nrmicro1464
Baranova NN, Danchin A, Neyfakh AA (1999) Mta, a global MerR-type regulator of the Bacillus subtilis multidrug-efflux transporters. Mol Microbiol 31:1549–1559. doi:10.1046/j.1365-2958.1999.01301.x
van Veen HW, Venema K, Bolhuis H, Oussenko I, Kok J, Poolman B, Driessen AJ, Konings WN (1996) Multidrug resistance mediated by a bacterial homolog of the human multidrug transporter MDR1. Proc Natl Acad Sci U S A 93:10668–10672
Chen CJ, Chin JE, Ueda K, Clark DP, Pastan I, Gottesman MM, Roninson IB (1986) Internal duplication and homology with bacterial transport proteins in the mdr1 (P-glycoprotein) gene from multidrug-resistant human cells. Cell 47:381–389. doi:10.1016/0092-8674(86)90595-7
Saier MH Jr (2003) Tracing pathways of transport protein evolution. Mol Microbiol 48:1145–1156. doi:10.1046/j.1365-2958.2003.03499.x
Kunst F, Ogasawara N, Moszer I, Albertini AM, Alloni G, Azevedo V, Bertero MG, Bessieres P et al (1997) The complete genome sequence of the Gram-positive bacterium Bacillus subtilis. Nature 390:249–256. doi:10.1038/36786
Steinfels E, Orelle C, Fantino JR, Dalmas O, Rigaud JL, Denizot F, Di Pietro A, Jault JM (2004) Characterization of YvcC (BmrA), a multidrug ABC transporter constitutively expressed in Bacillus subtilis. Biochemistry 43:7491–7502. doi:10.1021/bi0362018
Torres C, Galian C, Freiberg C, Fantino JR, Jault JM (2009) The YheI/YheH heterodimer from Bacillus subtilis is a multidrug ABC transporter. Biochim Biophys Acta 1788:615–622. doi:10.1016/j.bbamem.2008
Reilman E, Mars RA, van Dijl JM, Denham EL (2014) The multidrug ABC transporter BmrC/BmrD of Bacillus subtilis is regulated via a ribosome-mediated transcriptional attenuation mechanism. Nucleic Acids Res 42:11393–11407. doi:10.1093/nar/gku832
Ohki R, Giyanto, Tateno K, Masuyama W, Moriya S, Kobayashi K, Ogasawara N (2003) The BceRS two-component regulatory system induces expression of the bacitracin transporter, BceAB, in Bacillus subtilis. Mol Microbiol 49:1135–1144. doi:10.1046/j.1365-2958.2003.03653.x
Ohki R, Tateno K, Okada Y, Okajima H, Asai K, Sadaie Y, Murata M, Aiso T (2003) A bacitracin-resistant Bacillus subtilis gene encodes a homologue of the membrane-spanning subunit of the Bacillus licheniformis ABC transporter. J Bacteriol 185:51–59. doi:10.1128/JB.185.1.51-59.2003
Ohki R, Murata M (1997) bmr3, a third multidrug transporter gene of Bacillus subtilis. J Bacteriol 179:1423–1427
Ohki R, Tateno K (2004) Increased stability of bmr3 mRNA results in a multidrug-resistant phenotype in Bacillus subtilis. J Bacteriol 186:7450–7455. doi:10.1128/JB.186.21.7450-7455.2004, doi:10.1128/JB.185.1.51-59.2003
Kim J-Y, Inaoka T, Hirooka K, Matsuoka H, Murata M, Ohki R, Adachi Y, Fujita Y et al (2009) Identification and characterization of a novel multidrug resistance operon mdtRP (yusOP) of Bacillus subtilis. J Bacteriol 191:3273–3281. doi:10.1128/JB.00151-09
Masaoka Y, Ueno Y, Morita Y, Kuroda T, Mizushima T, Tsuchiya T (2000) A two-component multidrug efflux pump, EbrAB, in Bacillus subtilis. J Bacteriol 182:2307–2310. doi:10.1128/JB.182.8.2307-2310.2000
Kelly CP, Pothoulakis C, LaMont JT (1994) Clostridium difficile colitis. N Engl J Med 330:257–262. doi:10.1056/NEJM199401273300406
Dridi L, Tankovic J, Petit JC (2004) CdeA of Clostridium difficile, a new multidrug efflux transporter of the MATE family. Microb Drug Resist 10:191–196. doi:10.1089/mdr.2004.10.191
Lebel S, Bouttier S, Lambert T (2004) The cme gene of Clostridium difficile confers multidrug resistance in Enterococcus faecalis. FEMS Microbiol Lett 238:93–100. doi:10.1111/j.1574-6968.2004.tb09742.x
Vazquez-Boland JA, Dominguez-Bernal G, Gonzalez-Zorn B, Kreft J, Goebel W (2001) Pathogenicity islands and virulence evolution in Listeria. Microbes Infect 3:571–584. doi:10.1016/S1286-4579(01)01413-7
Portnoy DA, Chakraborty T, Goebel W, Cossart P (1992) Molecular determinants of Listeria monocytogenes pathogenesis. Infect Immun 60:1263–1267
Mata MT, Baquero F, Perez-Diaz JC (2000) A multidrug efflux transporter in Listeria monocytogenes. FEMS Microbiol Lett 187:185–188. doi:10.1111/j.1574-6968.2000.tb09158.x
Glaser P, Frangeul L, Buchrieser C, Rusniok C, Amend A, Baquero F, Berche P, Bloecker H et al (2001) Comparative genomics of Listeria species. Science 294:849–852. doi:10.1126/science.1063447
Crimmins GT, Herskovits AA, Rehder K, Sivick KE, Lauer P, Dubensky TW Jr, Portnoy DA (2008) Listeria monocytogenes multidrug resistance transporters activate a cytosolic surveillance pathway of innate immunity. Proc Natl Acad Sci U S A 105:10191–10196. doi:10.1073/pnas.0804170105
Huillet E, Velge P, Vallaeys T, Pardon P (2006) LadR, a new PadR-related transcriptional regulator from Listeria monocytogenes, negatively regulates the expression of the multidrug efflux pump MdrL. FEMS Microbiol Lett 254:87–94. doi:10.1111/j.1574-6968.2005.00014.x
Quillin SJ, Schwartz KT, Leber JH (2011) The novel Listeria monocytogenes bile sensor BrtA controls expression of the cholic acid efflux pump MdrT. Mol Microbiol 81:129–142. doi:10.1111/j.1365-2958.2011.07683.x
Woodward JJ, Iavarone AT, Portnoy DA (2010) c-di-AMP secreted by intracellular Listeria monocytogenes activates a host type I interferon response. Science 328:1703–1705. doi:10.1126/science.1189801
O’Riordan M, Yi CH, Gonzales R, Lee KD, Portnoy DA (2002) Innate recognition of bacteria by a macrophage cytosolic surveillance pathway. Proc Natl Acad Sci U S A 99:13861–13866. doi:10.1073/pnas.202476699
Stockinger S, Materna T, Stoiber D, Bayr L, Steinborn R, Kolbe T, Unger H, Chakraborty T et al (2002) Production of type I IFN sensitizes macrophages to cell death induced by Listeria monocytogenes. J Immunol 169:6522–6529. doi:10.4049/jimmunol.169.11.6522
Schwartz KT, Carleton JD, Quillin SJ, Rollins SD, Portnoy DA, Leber JH (2012) Hyperinduction of host beta interferon by a Listeria monocytogenes strain naturally overexpressing the multidrug efflux pump MdrT. Infect Immun 80:1537–1545. doi:10.1128/IAI.06286-11
Yamamoto T, Hara H, Tsuchiya K, Sakai S, Fang R, Matsuura M, Nomura T, Sato F et al (2012) Listeria monocytogenes strain-specific impairment of the TetR regulator underlies the drastic increase in cyclic di-AMP secretion and beta interferon-inducing ability. Infect Immun 80:2323–2332. doi:10.1128/IAI.06162-11
Commichau FM, Dickmanns A, Gundlach J, Ficner R, Stulke J (2015) A jack of all trades: the multiple roles of the unique essential second messenger cyclic di-AMP. Mol Microbiol 97:189–204. doi:10.1111/mmi.13026
Sauer JD, Sotelo-Troha K, von Moltke J, Monroe KM, Rae CS, Brubaker SW, Hyodo M, Hayakawa Y et al (2011) The N-ethyl-N-nitrosourea-induced Goldenticket mouse mutant reveals an essential function of Sting in the in vivo interferon response to Listeria monocytogenes and cyclic dinucleotides. Infect Immun 79:688–694. doi:10.1128/IAI.00999-10
Auerbuch V, Brockstedt DG, Meyer-Morse N, O’Riordan M, Portnoy DA (2004) Mice lacking the type I interferon receptor are resistant to Listeria monocytogenes. J Exp Med 200:527–533. doi:10.1084/jem.20040976
Carrero JA, Calderon B, Unanue ER (2004) Type I interferon sensitizes lymphocytes to apoptosis and reduces resistance to Listeria infection. J Exp Med 200:535–540. doi:10.1084/jem.20040769
O’Connell RM, Saha SK, Vaidya SA, Bruhn KW, Miranda GA, Zarnegar B, Perry AK, Nguyen BO et al (2004) Type I interferon production enhances susceptibility to Listeria monocytogenes infection. J Exp Med 200:437–445. doi:10.1084/jem.20040712
Collins B, Curtis N, Cotter PD, Hill C, Ross RP (2010) The ABC transporter AnrAB contributes to the innate resistance of Listeria monocytogenes to nisin, bacitracin, and various β-lactam antibiotics. Antimicrob Agents Chemother 54:4416–4423. doi:10.1128/AAC.00503-10
Mannion PT, Rothburn MM (1990) Diagnosis of bacterial endocarditis caused by Streptococcus lactis and assisted by immunoblotting of serum antibodies. J Infect 21:317–318. doi:10.1016/0163-4453(90)94149-T
Salyers AA, Gupta A, Wang Y (2004) Human intestinal bacteria as reservoirs for antibiotic resistance genes. Trends Microbiol 12:412–416. doi:10.1016/j.tim.2004.07.004
Bolhuis H, Molenaar D, Poelarends G, van Veen HW, Poolman B, Driessen AJ, Konings WN (1994) Proton motive force-driven and ATP-dependent drug extrusion systems in multidrug-resistant Lactococcus lactis. J Bacteriol 176:6957–6964
Bolhuis H, Poelarends G, van Veen HW, Poolman B, Driessen AJ, Konings WN (1995) The lactococcal lmrP gene encodes a proton motive force-dependent drug transporter. J Biol Chem 270:26092–26098. doi:10.1074/jbc.270.44.26092
van Veen HW, Callaghan R, Soceneantu L, Sardini A, Konings WN, Higgins CF (1998) A bacterial antibiotic-resistance gene that complements the human multidrug-resistance P-glycoprotein gene. Nature 391:291–295. doi:10.1038/34669
Bolhuis H, van Veen HW, Molenaar D, Poolman B, Driessen AJ, Konings WN (1996) Multidrug resistance in Lactococcus lactis: evidence for ATP-dependent drug extrusion from the inner leaflet of the cytoplasmic membrane. EMBO J 15:4239–4245
Poelarends GJ, Mazurkiewicz P, Putman M, Cool RH, Veen HW, Konings WN (2000) An ABC-type multidrug transporter of Lactococcus lactis possesses an exceptionally broad substrate specificity. Drug Resist Updat 3:330–334. doi:10.1054/drup.2000.0173
Lubelski J, Mazurkiewicz P, van Merkerk R, Konings WN, Driessen AJ (2004) ydaG and ydbA of Lactococcus lactis encode a heterodimeric ATP-binding cassette-type multidrug transporter. J Biol Chem 279:34449–34455. doi:10.1074/jbc.M404072200
Zaidi AH, Bakkes PJ, Lubelski J, Agustiandari H, Kuipers OP, Driessen AJ (2008) The ABC-type multidrug resistance transporter LmrCD is responsible for an extrusion-based mechanism of bile acid resistance in Lactococcus lactis. J Bacteriol 190:7357–7366. doi:10.1128/JB.00485-08
Agustiandari H, Lubelski J, van den Berg van Saparoea HB, Kuipers OP, Driessen AJ (2008) LmrR is a transcriptional repressor of expression of the multidrug ABC transporter LmrCD in Lactococcus lactis. J Bacteriol 190:759–763. doi:10.1128/JB.01151-07
Agustiandari H, Peeters E, de Wit JG, Charlier D, Driessen AJ (2011) LmrR-mediated gene regulation of multidrug resistance in Lactococcus lactis. Microbiology 157:1519–1530. doi:10.1099/mic.0.048025-0
van der Berg JP, Madoori PK, Komarudin AG, Thunnissen AM, Driessen AJ (2015) Binding of the lactococcal drug dependent transcriptional regulator LmrR to its ligands and responsive promoter regions. PLoS One 10:e0135467. doi:10.1371/journal.pone.0135467
Madoori PK, Agustiandari H, Driessen AJ, Thunnissen AM (2009) Structure of the transcriptional regulator LmrR and its mechanism of multidrug recognition. EMBO J 28:156–166. doi:10.1038/emboj.2008.263
Lubelski J, de Jong A, van Merkerk R, Agustiandari H, Kuipers OP, Kok J, Driessen AJ (2006) LmrCD is a major multidrug resistance transporter in Lactococcus lactis. Mol Microbiol 61:771–781. doi:10.1111/j.1365-2958.2006.05267.x
Filipic B, Golic N, Jovcic B, Tolinacki M, Bay DC, Turner RJ, Antic-Stankovic J, Kojic M et al (2013) The cmbT gene encodes a novel major facilitator multidrug resistance transporter in Lactococcus lactis. Res Microbiol 164:46–54. doi:10.1016/j.resmic.2012.09.003
Gajic O, Buist G, Kojic M, Topisirovic L, Kuipers OP, Kok J (2003) Novel mechanism of bacteriocin secretion and immunity carried out by lactococcal multidrug resistance proteins. J Biol Chem 278:34291–34298. doi:10.1074/jbc.M211100200
Lynch C, Courvalin P, Nikaido H (1997) Active efflux of antimicrobial agents in wild-type strains of enterococci. Antimicrob Agents Chemother 41:869–871
Jonas BM, Murray BE, Weinstock GM (2001) Characterization of EmeA, a NorA homolog and multidrug resistance efflux pump, in Enterococcus faecalis. Antimicrob Agents Chemother 45:3574–3579. doi:10.1128/AAC.45.12.3574-3579.2001
Lee EW, Huda MN, Kuroda T, Mizushima T, Tsuchiya T (2003) EfrAB, an ABC multidrug efflux pump in Enterococcus faecalis. Antimicrob Agents Chemother 47:3733–3738. doi:10.1128/AAC.47.12.3733-3738.2003
Lavilla Lerma L, Benomar N, Valenzuela AS, Casado Munoz Mdel C, Galvez A, Abriouel H (2014) Role of EfrAB efflux pump in biocide tolerance and antibiotic resistance of Enterococcus faecalis and Enterococcus faecium isolated from traditional fermented foods and the effect of EDTA as EfrAB inhibitor. Food Microbiol 44:249–257. doi:10.1016/j.fm.2014.06.009
Nishioka T, Ogawa W, Kuroda T, Katsu T, Tsuchiya T (2009) Gene cloning and characterization of EfmA, a multidrug efflux pump, from Enterococcus faecium. Biol Pharm Bull 32:483–488. doi:10.1248/bpb.32.483
Isaacman DJ, McIntosh ED, Reinert RR (2010) Burden of invasive pneumococcal disease and serotype distribution among Streptococcus pneumoniae isolates in young children in Europe: impact of the 7-valent pneumococcal conjugate vaccine and considerations for future conjugate vaccines. Int J Infect Dis 14:e197–e209. doi:10.1016/j.ijid.2009.05.010
Cornick JE, Bentley SD (2012) Streptococcus pneumoniae: the evolution of antimicrobial resistance to β-lactams, fluoroquinolones and macrolides. Microbes Infect 14:573–583. doi:10.1016/j.micinf.2012.01.012
Baranova NN, Neyfakh AA (1997) Apparent involvement of a multidrug transporter in the fluoroquinolone resistance of Streptococcus pneumoniae. Antimicrob Agents Chemother 41:1396–1398
Gill MJ, Brenwald NP, Wise R (1999) Identification of an efflux pump gene, pmrA, associated with fluoroquinolone resistance in Streptococcus pneumoniae. Antimicrob Agents Chemother 43:187–189
Robertson GT, Doyle TB, Lynch AS (2005) Use of an efflux-deficient Streptococcus pneumoniae strain panel to identify ABC-class multidrug transporters involved in intrinsic resistance to antimicrobial agents. Antimicrob Agents Chemother 49:4781–4783. doi:10.1128/AAC.49.11.4781-4783.2005
Tocci N, Iannelli F, Bidossi A, Ciusa ML, Decorosi F, Viti C, Pozzi G, Ricci S et al (2013) Functional analysis of pneumococcal drug efflux pumps associates the MATE DinF transporter with quinolone susceptibility. Antimicrob Agents Chemother 57:248–253. doi:10.1128/AAC.01298-12
Marrer E, Schad K, Satoh AT, Page MG, Johnson MM, Piddock LJ (2006) Involvement of the putative ATP-dependent efflux proteins PatA and PatB in fluoroquinolone resistance of a multidrug-resistant mutant of Streptococcus pneumoniae. Antimicrob Agents Chemother 50:685–693. doi:10.1128/AAC.50.2.685-693.2006
Boncoeur E, Durmort C, Bernay B, Ebel C, Di Guilmi AM, Croize J, Vernet T, Jault JM (2012) PatA and PatB form a functional heterodimeric ABC multidrug efflux transporter responsible for the resistance of Streptococcus pneumoniae to fluoroquinolones. Biochemistry 51:7755–7765. doi:10.1021/bi300762p
Marrer E, Satoh AT, Johnson MM, Piddock LJ, Page MG (2006) Global transcriptome analysis of the responses of a fluoroquinolone-resistant Streptococcus pneumoniae mutant and its parent to ciprofloxacin. Antimicrob Agents Chemother 50:269–278. doi:10.1128/aac.50.1.269-278.2006
El Garch F, Lismond A, Piddock LJ, Courvalin P, Tulkens PM, Van Bambeke F (2010) Fluoroquinolones induce the expression of patA and patB, which encode ABC efflux pumps in Streptococcus pneumoniae. J Antimicrob Chemother 65:2076–2082. doi:10.1093/jac/dkq287
Piddock LJ, Johnson MM, Simjee S, Pumbwe L (2002) Expression of efflux pump gene pmrA in fluoroquinolone-resistant and -susceptible clinical isolates of Streptococcus pneumoniae. Antimicrob Agents Chemother 46:808–812. doi:10.1128/AAC.46.3.808-812.2002
Garvey MI, Baylay AJ, Wong RL, Piddock LJ (2011) Overexpression of patA and patB, which encode ABC transporters, is associated with fluoroquinolone resistance in clinical isolates of Streptococcus pneumoniae. Antimicrob Agents Chemother 55:190–196. doi:10.1128/AAC.00672-10
Doran KS, Chang JC, Benoit VM, Eckmann L, Nizet V (2002) Group B streptococcal β-hemolysin/cytolysin promotes invasion of human lung epithelial cells and the release of interleukin-8. J Infect Dis 185:196–203. doi:10.1086/338475
Doran KS, Liu GY, Nizet V (2003) Group B streptococcal β-hemolysin/cytolysin activates neutrophil signaling pathways in brain endothelium and contributes to development of meningitis. J Clin Invest 112:736–744. doi:10.1172/JCI17335
Spellerberg B, Pohl B, Haase G, Martin S, Weber-Heynemann J, Lutticken R (1999) Identification of genetic determinants for the hemolytic activity of Streptococcus agalactiae by ISS1 transposition. J Bacteriol 181:3212–3219
Rosa-Fraile M, Dramsi S, Spellerberg B (2014) Group B streptococcal haemolysin and pigment, a tale of twins. FEMS Microbiol Rev 38:932–946. doi:10.1111/1574-6976.12071
Gottschalk B, Broker G, Kuhn M, Aymanns S, Gleich-Theurer U, Spellerberg B (2006) Transport of multidrug resistance substrates by the Streptococcus agalactiae hemolysin transporter. J Bacteriol 188:5984–5992. doi:10.1128/JB.00768-05
Whidbey C, Harrell MI, Burnside K, Ngo L, Becraft AK, Iyer LM, Aravind L, Hitti J et al (2013) A hemolytic pigment of Group B Streptococcus allows bacterial penetration of human placenta. J Exp Med 210:1265–1281. doi:10.1084/jem.20122753
Rosa-Fraile M, Rodriguez-Granger J, Haidour-Benamin A, Cuerva JM, Sampedro A (2006) Granadaene: proposed structure of the group B Streptococcus polyenic pigment. Appl Environ Microbiol 72:6367–6370. doi:10.1128/AEM.00756-06
Loesche WJ (1986) Role of Streptococcus mutans in human dental decay. Microbiol Rev 50:353–380
Yoshida A, Ansai T, Takehara T, Kuramitsu HK (2005) LuxS-based signaling affects Streptococcus mutans biofilm formation. Appl Environ Microbiol 71:2372–2380. doi:10.1128/AEM.71.5.2372-2380.2005
Jarvinen H, Tenovuo J, Huovinen P (1993) In vitro susceptibility of Streptococcus mutans to chlorhexidine and six other antimicrobial agents. Antimicrob Agents Chemother 37:1158–1159. doi:10.1128/AAC.37.5.1158
William B, Rwenyonyi CM, Swedberg G, Kironde F (2012) Cotrimoxazole prophylaxis specifically selects for cotrimoxazole resistance in Streptococcus mutans and Streptococcus sobrinus with varied polymorphisms in the target genes folA and folP. Int J Microbiol 2012:916129. doi:10.1155/2012/916129
Kulik EM, Waltimo T, Weiger R, Schweizer I, Lenkeit K, Filipuzzi-Jenny E, Walter C (2015) Development of resistance of mutans streptococci and Porphyromonas gingivalis to chlorhexidine digluconate and amine fluoride/stannous fluoride-containing mouthrinses, in vitro. Clin Oral Invest 19:1547–1553. doi:10.1007/s00784-014-1379-y
Kawada-Matsuo M, Oogai Y, Zendo T, Nagao J, Shibata Y, Yamashita Y, Ogura Y, Hayashi T et al (2013) Involvement of the novel two-component NsrRS and LcrRS systems in distinct resistance pathways against nisin A and nukacin ISK-1 in Streptococcus mutans. Appl Environ Microbiol 79:4751–4755. doi:10.1128/AEM.00780-13
Kim DK, Kim KH, Cho EJ, Joo SJ, Chung JM, Son BY, Yum JH, Kim YM et al (2013) Gene cloning and characterization of MdeA, a novel multidrug efflux pump in Streptococcus mutans. J Microbiol Biotechnol 23:430–435. doi:10.4014/jmb.1301.01028
Ajdic D, McShan WM, McLaughlin RE, Savic G, Chang J, Carson MB, Primeaux C, Tian R et al (2002) Genome sequence of Streptococcus mutans UA159, a cariogenic dental pathogen. Proc Natl Acad Sci U S A 99:14434–14439. doi:10.1073/pnas.172501299
Biswas S, Biswas I (2011) Role of VltAB, an ABC transporter complex, in viologen tolerance in Streptococcus mutans. Antimicrob Agents Chemother 55:1460–1469. doi:10.1128/AAC.01094-10
Yonezawa H, Kuramitsu HK (2005) Genetic analysis of a unique bacteriocin, Smb, produced by Streptococcus mutans GS5. Antimicrob Agents Chemother 49:541–548. doi:10.1128/AAC.49.2.541-548.2005
Biswas S, Biswas I (2013) SmbFT, a putative ABC transporter complex, confers protection against the lantibiotic Smb in streptococci. J Bacteriol 195:5592–5601. doi:10.1128/JB.01060-13
Seaton K, Ahn SJ, Sagstetter AM, Burne RA (2011) A transcriptional regulator and ABC transporters link stress tolerance, (p)ppGpp, and genetic competence in Streptococcus mutans. J Bacteriol 193:862–874. doi:10.1128/JB.01257-10
Ahn SJ, Kaspar J, Kim JN, Seaton K, Burne RA (2014) Discovery of novel peptides regulating competence development in Streptococcus mutans. J Bacteriol 196:3735–3745. doi:10.1128/JB.01942-14
Shields RC, Burne RA (2015) Conserved and divergent functions of RcrRPQ in Streptococcus gordonii and S. mutans. FEMS Microbiol Lett 362. doi:10.1093/femsle/fnv119
Ardin AC, Fujita K, Nagayama K, Takashima Y, Nomura R, Nakano K, Ooshima T, Matsumoto-Nakano M (2014) Identification and functional analysis of an ammonium transporter in Streptococcus mutans. PLoS One 9:e107569. doi:10.1371/journal.pone.0107569
Ouyang J, Tian XL, Versey J, Wishart A, Li YH (2010) The BceABRS four-component system regulates the bacitracin-induced cell envelope stress response in Streptococcus mutans. Antimicrob Agents Chemother 54:3895–3906. doi:10.1128/AAC.01802-09
Singh K, Senadheera DB, Levesque CM, Cvitkovitch DG (2015) The copYAZ operon functions in copper efflux, biofilm formation, genetic transformation, and stress tolerance in Streptococcus mutans. J Bacteriol 197:2545–2557. doi:10.1128/JB.02433-14
Chen PM, Chen HC, Ho CT, Jung CJ, Lien HT, Chen JY, Chia JS (2008) The two-component system ScnRK of Streptococcus mutans affects hydrogen peroxide resistance and murine macrophage killing. Microbes Infect 10:293–301. doi:10.1016/j.micinf.2007.12.006
Ahn SJ, Qu MD, Roberts E, Burne RA, Rice KC (2012) Identification of the Streptococcus mutans LytST two-component regulon reveals its contribution to oxidative stress tolerance. BMC Microbiol 12:187. doi:10.1186/1471-2180-12-187
Bernard R, Joseph P, Guiseppi A, Chippaux M, Denizot F (2003) YtsCD and YwoA, two independent systems that confer bacitracin resistance to Bacillus subtilis. FEMS Microbiol Lett 228:93–97. doi:10.1016/S0378-1097(03)00738-9
Woolridge DP, Vazquez-Laslop N, Markham PN, Chevalier MS, Gerner EW, Neyfakh AA (1997) Efflux of the natural polyamine spermidine facilitated by the Bacillus subtilis multidrug transporter Blt. J Biol Chem 272:8864–8866. doi:10.1074/jbc.272.14.8864
Murata M, Ohno S, Kumano M, Yamane K, Ohki R (2003) Multidrug resistant phenotype of Bacillus subtilis spontaneous mutants isolated in the presence of puromycin and lincomycin. Can J Microbiol 49:71–77. doi:10.1139/w03-014
Yoshida K, Ohki YH, Murata M, Kinehara M, Matsuoka H, Satomura T, Ohki R, Kumano M et al (2004) Bacillus subtilis LmrA is a repressor of the lmrAB and yxaGH operons: identification of its binding site and functional analysis of lmrB and yxaGH. J Bacteriol 186:5640–5648. doi:10.1128/JB.186.17.5640-5648.2004
Singh KV, Weinstock GM, Murray BE (2002) An Enterococcus faecalis ABC homologue (Lsa) is required for the resistance of this species to clindamycin and quinupristin-dalfopristin. Antimicrob Agents Chemother 46:1845–1850. doi:10.1128/AAC.46.6.1845-1850.2002
Perreten V, Schwarz FV, Teuber M, Levy SB (2001) Mdt(A), a new efflux protein conferring multiple antibiotic resistance in Lactococcus lactis and Escherichia coli. Antimicrob Agents Chemother 45:1109–1114. doi:10.1128/AAC.45.4.1109-1114.2001
Romanova NA, Wolffs PF, Brovko LY, Griffiths MW (2006) Role of efflux pumps in adaptation and resistance of Listeria monocytogenes to benzalkonium chloride. Appl Environ Microbiol 72:3498–3503. doi:10.1128/AEM.72.5.3498-3503.2006
Godreuil S, Galimand M, Gerbaud G, Jacquet C, Courvalin P (2003) Efflux pump Lde is associated with fluoroquinolone resistance in Listeria monocytogenes. Antimicrob Agents Chemother 47:704–708. doi:10.1128/AAC.47.2.704-708.2003
Lismond A, Tulkens PM, Mingeot-Leclercq MP, Courvalin P, Van Bambeke F (2008) Cooperation between prokaryotic (Lde) and eukaryotic (MRP) efflux transporters in J774 macrophages infected with Listeria monocytogenes: studies with ciprofloxacin and moxifloxacin. Antimicrob Agents Chemother 52:3040–3046. doi:10.1128/AAC.00105-08
Cai Y, Kong F, Gilbert GL (2007) Three new macrolide efflux (mef) gene variants in Streptococcus agalactiae. J Clin Microbiol 45:2754–2755. doi:10.1128/jcm.00579-07
Clancy J, Dib-Hajj F, Petitpas JW, Yuan W (1997) Cloning and characterization of a novel macrolide efflux gene, mreA, from Streptococcus agalactiae. Antimicrob Agents Chemother 41:2719–2723
Avrain L, Garvey M, Mesaros N, Glupczynski Y, Mingeot-Leclercq MP, Piddock LJ, Tulkens PM, Vanhoof R et al (2007) Selection of quinolone resistance in Streptococcus pneumoniae exposed in vitro to subinhibitory drug concentrations. J Antimicrob Chemother 60:965–972. doi:10.1093/jac/dkm292
Garvey MI, Piddock LJ (2008) The efflux pump inhibitor reserpine selects multidrug-resistant Streptococcus pneumoniae strains that overexpress the ABC transporters PatA and PatB. Antimicrob Agents Chemother 52:1677–1685. doi:10.1128/AAC.01644-07
Tait-Kamradt A, Clancy J, Cronan M, Dib-Hajj F, Wondrack L, Yuan W, Sutcliffe J (1997) mefE is necessary for the erythromycin-resistant M phenotype in Streptococcus pneumoniae. Antimicrob Agents Chemother 41:2251–2255
Acknowledgments
The views expressed in this chapter do not necessarily reflect those of the authors’ affiliation, U.S. Food and Drug Administration.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Baranova, N., Elkins, C.A. (2016). Antimicrobial Drug Efflux Pumps in Other Gram-Positive Bacteria. In: Li, XZ., Elkins, C., Zgurskaya, H. (eds) Efflux-Mediated Antimicrobial Resistance in Bacteria. Adis, Cham. https://doi.org/10.1007/978-3-319-39658-3_8
Download citation
DOI: https://doi.org/10.1007/978-3-319-39658-3_8
Published:
Publisher Name: Adis, Cham
Print ISBN: 978-3-319-39656-9
Online ISBN: 978-3-319-39658-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)