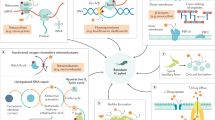
Abstract
Helicobacter spp. play important etiological roles in the pathogenesis of gastroenteric diseases such as in the case of Helicobacter pylori. Despite wild-type strains of H. pylori being generally susceptible to multiple antimicrobial agents, increasing prevalence of antimicrobial resistance in this species constitutes a key risk factor that affects the effective therapy of H. pylori infections. Resistance to anti-H. pylori agents is mainly mediated by multiple drug-specific mechanisms. However, drug efflux systems, represented by the Hef pumps of the resistance-nodulation-cell division superfamily, are implicated in both intrinsic and acquired multidrug resistance as well as in bile salt/nitrosative stress response and gastric colonization of these pathogens. This chapter provides an overview of antimicrobial resistance and mechanisms in Helicobacter with an emphasis on drug efflux systems.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Helicobacter pylori
- Antimicrobial resistance
- Efflux
- RND pumps
- Outer membrane
- Stress response
- Amoxicillin
- Clarithromycin
- Metronidazole
- Bile salts
- HefABC
- HefDEF
- HefGHI
1 Introduction
Helicobacter spp. are Gram-negative bacteria belonging to the Epsilonproteobacteria class. The representative species, Helicobacter pylori, is believed to infect at least 50 % of the world’s human population [1]. Although most individuals with H. pylori infection do not experience any clinical complications, these infections are often implicated in the development of chronic gastritis, peptic and duodenal ulcers, as well as gastric cancers [1, 2]. Indeed, H. pylori has been identified as a carcinogen [3]. H. pylori, along with additional non-H. pylori Helicobacter species, can be divided into three groups (i.e., gastric, enterohepatic, and unsheathed flagella) based on 16S rRNA sequence similarity [4]. Although many of these species have primary animal hosts, some are also known to be associated with gastroenteric and/or hepatic diseases in humans and include, for example, Helicobacter hepaticus, Helicobacter bilis, and Helicobacter cinaedi [4–8]. Furthermore, the clinical significance of Helicobacter spp. in the development of gastrointestinal diseases has been supported by microbiome data generated for gut microbiota [9, 10]. Helicobacter species can persist and cause chronic inflammation in human gut, thus contributing to the pathogenesis of various gastroenteric diseases [7, 8, 10, 11]. Antimicrobial therapy is needed for the eradication of these bacterial infections; however, the antimicrobials available for the treatment of these infections are quite limited, and a combination therapy is required to achieve optimal clinical effectiveness. Moreover, increasing prevalence of antimicrobial resistance has been observed in H. pylori against agents used in H. pylori treatment emerges as a crucial issue when tackling H. pylori infections [12]. Drug efflux pumps are one of many mechanisms increasingly recognized to play an important role in the emerging resistance in H. pylori and other species. Drug efflux pumps of various known transporter families are inherently encoded in Helicobacter genomes. In this chapter, we examine the current status of antimicrobial resistance in this genus with an emphasis on drug efflux pumps.
2 Antimicrobial Susceptibility, Therapeutic Options, and Resistance Prevalence
Antimicrobial susceptibility studies of Helicobacter spp. have been mostly limited to H. pylori, which displays significant in vitro susceptibility to a number of antimicrobial agents including β-lactams, macrolides, fluoroquinolones, nitroimidazoles, nitrofurans, and tetracyclines [1, 13, 14]. Wild-type strains of H. pylori generally have greater susceptibility to antimicrobial agents compared to Escherichia coli and Pseudomonas aeruginosa, with certain exceptions such as polymyxins, glycopeptides, nalidixic acid, sulfonamides, trimethoprim, and streptogramins (Table 19.1) [13, 17]. The lowered pH within the gastrointestinal tract, the habitat of H. pylori, has a negative impact on antimicrobial activity of β-lactams, macrolides, tetracycline, and fluoroquinolones, with values of minimal inhibitory concentrations (MICs) decreased by 4- to 130-fold with pH changes from 7.2 to 5.5 [13]. Susceptibility data for non-H. pylori Helicobacter species are very limited, though several isolates of H. hepaticus show significant intrinsic resistance to amoxicillin with MIC values of 8–64 μg/ml (cf. with values of ≤0.5 μg/ml for H. pylori) [28].
Despite the high in vitro susceptibility of H. pylori to numerous agents (Table 19.1), in vivo therapy of H. pylori infections may not correlate well with expectations based on in vitro data [13]. The harsh environment within the stomach poses a challenge for drug selection among orally administered antimicrobial agents. Indeed, therapy has been limited to certain individual agents of various classes which include amoxicillin, clarithromycin, furazolidone, fluoroquinolones, metronidazole, rifabutin, and tetracycline [13, 29]. Therapeutic regimens require combination therapy or sequential therapy with the abovementioned antimicrobials [13, 29–31]. The recommended first-line therapy for the treatment of H. pylori infections consists of a standard triple-drug therapy with any two of three antibiotics (amoxicillin, clarithromycin, and metronidazole) and either a proton pump inhibitor (e.g., esomeprazole) or ranitidine bismuth citrate for a duration of 7–14 days [13]. Proton pump inhibitors and bismuth salts possess anti-H. pylori activity at high concentration levels [13]. A second-line therapy consists of a quadruple regimen of tetracycline, metronidazole, a bismuth salt, and a proton pump inhibitor [13, 29–31]. Third-line treatment regimens and other rescue therapies are based on the antimicrobial susceptibility profile of the specific strain in question and may include fluoroquinolones, tetracyclines, rifabutin, and furazolidone [32–35].
Antimicrobial resistance is increasingly being recognized as a risk factor affecting treatment efficacy against Helicobacter infections [31, 36–39]. A review from 20 years ago has documented a variable but overall high prevalence of 10–70 % metronidazole resistance [1]. Antimicrobial treatment failure has been linked to increased prevalence of resistant isolates [40]. In a recent study that tested around 340 isolates (including those from patients with up to three treatment failures), the MIC values of amoxicillin varied from <0.015 to 4 μg/ml, with higher prevalence of amoxicillin resistance in isolates from the treatment failures [41]. The rates of resistance to clarithromycin, a major agent for first-line therapies, have increased from 9 % to 18 % in 1998–2008 in Europe and from 7 % to 28 % in 2000–2006 in Japan (reviewed in reference [42]). A recent report has described the rates of resistance to clarithromycin (18 %), levofloxacin (14 %), and metronidazole (35 %) in Europe, and the increased prevalence of resistance was attributable to the increased use of fluoroquinolones and macrolides in clinic [43]. Similarly, a study conducted in China showed increased rates of resistance to clarithromycin (9 % in 2000 to 21 % in 2009) and levofloxacin (10 % in 2000 to 33 % in 2009) with stable rates of about 40–50 % for resistance to metronidazole within a 10-year period [44]. Yet resistance to amoxicillin, furazolidone, or tetracycline was not detectable and/or rarely occurred [44]. A surveillance of nearly 18,000 isolates that were sampled in China between 2009 and 2012 revealed resistance rates of ca. 21 % for clarithromycin and levofloxacin and 94 % for metronidazole with only 0.1 % for amoxicillin, furazolidone, and gentamicin [45]. Resistance to rifabutin remains generally low with the rates of 1.4 % in Germany and 0.24 % in Japan [24, 46]. A German study identified simultaneous resistance to three or four agents in 15 % of isolates contributing to unsuccessful antimicrobial treatment [47]. Furthermore, a Canadian study has also suggested a general increase in resistance to clarithromycin, ciprofloxacin, levofloxacin, and metronidazole beginning from the early 2000s. Together, these data also suggest variable prevalence of resistance in different regions and countries [31]. Newer agents such as finafloxacin and linezolid have been tested for their activity against H. pylori, but their implications for therapy require clinical trials [48, 49]. Lastly, heteroresistance, a circumstance in which subpopulations of isogenic strains develop varying antimicrobial susceptibilities [50], was also observed in isolates from the same patients (even before antimicrobial treatment) [51, 52]. Resistance identification can be hindered by heteroresistance with an undesired consequence of selecting more resistant isolates via antimicrobial therapy [50, 51, 53].
It is also noteworthy that the combinatory use of antimicrobials for treating H. pylori infections can have an adverse long-term in vivo impact on resistance development and persistence in the gut microbiota. For instance, a short-term clarithromycin-metronidazole combination regimen dramatically reduced the diversity of gut microbiota and resulted in a 1,000-fold increase in the ermB gene (encoding the macrolide target-modifying RNA methylase), which then persisted in the gut microbiota for at least 4 years [54]. This observation is consistent with an earlier study showing the persistence of ermB-mediated resistant enterococci for 1–3 years following an anti-H. pylori treatment regimen [55]. Additionally, by modifing lipid A and biofilm formation, H. pylori can adapt in vivo to resist the antimicrobial activity of calprotein, which is a component of the host innate immune system and is present during the inflammatory response [56].
3 Mechanisms of Antimicrobial Resistance
H. pylori displays intrinsic resistance to multiple-unrelated antimicrobials including glycopeptides and polymyxins (Table 19.1) [13], suggesting that access to drug targets likely contributes to resistance manifestation. Acquired resistance can be further developed. One early study from 1990 showed the in vitro selection of resistant mutants by antimicrobials at the levels of 4× or 8× MIC, with spontaneous resistance frequencies in the range of 10−8–10−6 for ciprofloxacin, erythromycin, metronidazole, and tobramycin [22]. Another study in 2001 reported the frequencies of the in vitro spontaneous mutants resistant to clarithromycin, ciprofloxacin, metronidazole, and rifampin being 3 × 10−9 to 7 × 10−8, while no mutants were recovered for amoxicillin [57]. Development of increasing resistance in H. pylori has prompted the investigation of resistance mechanisms. Table 19.2 lists the identified mechanisms of resistance to the major antimicrobials used in the treatment of H. pylori infection. Although antimicrobial target changes are a major form of resistance for H. pylori, the role of drug efflux systems should not be underestimated.
3.1 Amoxicillin Resistance
Resistance to amoxicillin in H. pylori appears to occur less frequently [36, 44, 57]. This phenomenon is attributable to mutations in the genes encoding penicillin-binding proteins (PBPs) [41]. H. pylori possess three to four major PBPs [58]. Amoxicillin-resistant mutants show significant reduction in the affinity of PBP1 to amoxicillin [59]. Furthermore, amino acid substitutions were observed in PBP1 of resistant isolates [41]. Although mutations in the pbp2 gene were also noted with those in the pbp1, they did not affect amoxicillin resistance [85]. In addition to mutations in PBP1, other unidentified mechanism(s) are likely also needed for high-level amoxicillin resistance [85]. A cysteine-rich protein named HcpA (encoded by HP0211) was earlier suggested to not only be a PBP but also a β-lactamase of H. pylori that slowly hydrolyzes penicillin derivatives [86]. However, more recent studies only demonstrated HcpA as a bacterial virulence factor triggering the release of a concerted set of cytokines [87, 88]. No further studies support HcpA as a typical β-lactamase. Indeed, typical β-lactamase activity is not detectable in H. pylori [89], although it is well known that PBPs generally may have certain β-lactamase activity. This is consistent with the observation that the H. pylori genome does not contain genes encoding typical β-lactamases, whose production constitutes the predominant mechanism of β-lactam resistance in Gram-negative bacteria. However, given that many β-lactamase genes are located on plasmids and that H. pylori has a strong natural transformation capability, it is not surprising to see the report of a high-level amoxicillin-resistant isolate (≥256 μg/ml amoxicillin) carrying the bla TEM gene [60]; it remains unclear whether this gene was plasmid borne or chromosome encoded.
Interestingly, two studies have revealed that amoxicillin-resistant/multidrug-resistant mutants accumulate less penicillin, chloramphenicol, and/or tetracycline than susceptible strains [59, 89]. Yet, the accumulation of penicillin and tetracycline by resistant strains was not affected by the ionophore proton conductor, carbonyl cyanide m-chlorophenyl hydrazone (CCCP) [59, 89]). Thus, a reduced accumulation of drugs was explained by investigators as due to reduced uptake and not active efflux. However, the involvement of drug efflux systems needs to be carefully assessed before a solid conclusion is made regarding additional mechanisms of amoxicillin resistance. The outer membrane permeability barrier alone cannot sufficiently explain drug accumulation differences in the steady state (e.g., within 30–60 min of accumulation). These drug molecules are expected to cross the outer membrane barrier in less than a second [90]. Based on various drug accumulation assays, the steady state of drug levels in intact cells should generally be reached within 30 min [59, 89, 90]. Indeed, the contribution from porin alterations and efflux pumps to amoxicillin resistance in Helicobacter spp. has been observed [16, 28] as discussed in the next section. This finding explains a phenotypic relationship between high β-lactam resistance and low- to moderate-level multidrug resistance [89].
3.2 Clarithromycin Resistance
Macrolides inhibit bacterial protein synthesis by targeting 23S rRNA. Mutations in the 23S rRNA genes reduce the binding of macrolides to the 23S rRNA [62–64, 91] and are the major mechanism of resistance to macrolides (and particularly clarithromycin for H. pylori) [42]. Major mutations include A2142G and A2143G transitions and an A2142C transversion [62, 92] with additional mutations in the 23S rRNA genes reported in the literature [19, 36]. Mutations within other genes, including those in infB (encoding translation initiation factor IF-2) and rpl22 (ribosomal protein L22), were also found to cooperate with 23S rRNA gene mutations in raising resistance level [20]. Lastly, macrolides are often the substrates of multidrug resistance or macrolide-specific efflux pumps in various bacteria [93, 94], and indeed efflux pumps also mediate resistance to clarithromycin, as described in the next section.
3.3 Metronidazole Resistance
As a nitroimidazole agent, metronidazole requires reduction by oxygen-insensitive NADPH nitroreductase (RdxA), NADPH-flavin oxidoreductase (FrxA), and ferredoxin-like enzymes (FrxB) to be activated from its prodrug form. Thus, mutations in relevant encoding genes (rdxA, frxA, and frxB) are responsible for metronidazole resistance [66, 85, 91, 95, 96]. Annotated mutations in rdxA consist of frame shift mutations, missense mutations, deletions, and insertions [36, 72]. The rdxA mutations are also better correlated to clinically relevant resistance (metronidazole MIC >8 μg/ml) than those in frxA [73]. Mutations in frxA alone may not be sufficient in generating metronidazole resistance [53, 96]. In addition to confirming the role of rdxA and frxA mutations, a recent study also identified, via whole genome sequencing and natural transformation approaches, mutations in another gene, rpsU [97]. The rpsU gene encodes ribosomal protein S21; mutations rpsU alone do not produce sufficient resistance levels but instead cooperate with rdxA mutations to achieve high resistance [97]. Inhibition of superoxide dismutase production in strains with mutations in the ferric uptake regulator is also known to be involved in metronidazole resistance [74]. The contribution of efflux mechanism to metronidazole resistance is discussed in the next section. Interestingly, the loss of metronidazole resistance occurs under low oxygen conditions (that mimic in vivo microaerophilic situation) or in the presence of chloramphenicol [98], suggesting multiple factors affecting metronidazole susceptibility.
3.4 Fluoroquinolone Resistance
Fluoroquinolones act on DNA gyrase (A2B2 complex) and topoisomerase IV. However, H. pylori strains lack the gene encoding topoisomerase IV [42]. Thus, resistance occurs mainly as a result of mutations in the quinolone resistance determining region of the gyrase A gene (e.g., Asn87Lys; Asn87Tyr; Asp91Gly, Asp91Asn, or Asp91Tyr) [42, 70, 91]. Mutations in the gyrase B gene were have also been noted [70]. Furthermore, although fluoroquinolone resistance frequently occurs as a result of efflux pump overproduction in Gram-negative bacteria [94], no efflux pumps affecting quinolone susceptibility have been identified in H. pylori to date.
3.5 Furazolidone Resistance
The broad-spectrum furazolidone inhibits DNA biosynthesis by crossing-linking to DNA molecules [67]. Mechanisms for furazolidone resistance in Helicobacter spp. are not well understood. However, resistance to nitrofurans in E. coli has primarily been linked to mutations in genes encoding nitroreductases such as nfsA and nfsB [99]. A recent study further demonstrated involvement of these mutations in furazolidone-resistant E. coli [68]. Pyruvate/flavodoxin oxidoreductase (PorCDAB) and 2-oxoglutarate oxidoreductase (OorDABC) act as nitrofuran nitroreductases in H. pylori [66], and mutations in the porD and oorD genes have been noted in all furazolidone-resistant (>2 μg/ml furazolidone) clinical isolates of H. pylori that were obtained from patients previously treated with metronidazole [69].
3.6 Rifabutin Resistance
Rifabutin acts on the β-subunit of the DNA-dependent RNA polymerase encoded by the rpoB gene. Amino acid substitutions resulting from point mutations in rpoB confer high-level resistance to rifampicin and rifabutin (with >128-fold MIC increases) [23, 24]. These resistance levels are dependent on the amino acid substitutions with four distinct regions identified in rpoB [76]. Similar to many other species, rifamycin resistance in H. pylori occurs more frequently than resistance to other agents [57]. The history of rifamycin use has been linked to the emergence of rifabutin-resistant isolates including those from cases with treatment failure [24, 46, 77].
3.7 Tetracycline Resistance
Tetracyclines inhibit protein synthesis by binding to the 30S subunit of the ribosome and preventing association between aminoacyl-tRNAs with the ribosome [78, 79]. Mutations in the 16S rRNA rrnA/rrnB genes reduce drug binding to the ribosome [100] and yield high-level tetracycline resistance (>40-fold MIC increases for tetracycline, doxycycline, and minocycline [25, 80, 81]). However, other types of tetracycline-resistant isolates were found to lack any mutations in the 16S rRNA genes and instead rely on the altered uptake or efflux [82, 83]. One study has shown proton motive force-dependent efflux of tetracycline in clinical isolates without identifying specific pump(s) [84]. The requirement for multiple mutations in the development of tetracycline resistance may explain its low prevalence in clinical resistance [101, 102]. The involvement of efflux pumps (HP1165) in tetracycline resistance [83] will be discussed in the next section.
3.8 Molecular Methods for Resistance Detection
Molecular methods have been developed to detect resistance caused by the specific gene mutations, and have been facilitated by advances in technology such as whole genome sequencing [92]. Commercially available molecular methods for detection of antimicrobial resistance in H. pylori also exist as reviewed in the reference [42]. The GenoType HelicoDR test is able to identify point mutations in the rrn and gyrA genes that are linked to clarithromycin and levofloxacin resistance, respectively [103], but this application has limited in its sensitivity and specificity, apparently making it infeasible for clinical applications [104]. Overall, genetic molecular methods are only applied to known resistance mechanisms for certain genes, as mutations can also be independent of the resistance phenotype [13]. Molecular approaches for examining genetic mutations alone will not be sufficient in characterizing resistance attributable to efflux mechanisms, particularly because multiple regulatory genes can impact efflux gene expression and ultimately the resistance phenotype.
4 Outer Membrane Permeability Barrier and Drug Efflux Systems
In Gram-negative bacteria, the outer membrane permeability barrier and drug exporters affect the influx and efflux of antimicrobial agents, respectively, and thus play a role in determining the susceptibility phenotype [94]. A large number of outer membrane and efflux proteins are encoded in the Helicobacter genomes [105–107], although the sizes of several known Helicobacter genomes are relatively small (only about 1.7–2.0 Mbp) [105, 108–110]. For example, at least 32 outer membrane proteins [111] and 27 proven or putative drug transporters [112] have been identified in H. pylori. These transporters belong to one of the following superfamilies or families [94]: (i) resistance-nodulation-cell division (RND) superfamily, (ii) the major facilitator superfamily (MFS), (iii) the multidrug and toxic compound extrusion (MATE) family, (iv) the small multidrug resistance (SMR) family, and (v) the ATP-binding cassette (ABC) superfamily (Table 19.3).
4.1 Outer Membrane Permeability Barrier
The outer membrane consists of a lipopolysaccharide-containing lipid bilayer with water-filled porins and serves as an effective barrier in limiting the influx of antimicrobial molecules [117]. Small hydrophilic agents, such as amoxicillin, cross the outer membrane via the porin channels, while large or hydrophobic agents require penetration of the outer membrane lipid bilayer [94]. Many outer membrane proteins of H. pylori have been studied for their role in infection pathogenesis [107, 118, 119], with five proteins (HopA to E) investigated for their channel-forming activity [120, 121]. The HopA to HopD porins form similar pores with relatively small channel size [120], while the less abundant HopE protein forms a larger nonspecific channel in vitro [121]. The presence of these porins explains the high susceptibility of H. pylori to small hydrophilic antimicrobials such as amoxicillin, which is expected to enter the periplasm through the porin channels. Indeed, mutations in HopB and HopC proteins render cells less susceptible to β-lactams (two- to eightfold reductions in the MIC values) and cooperate with PBP1 mutations to raise levels of β-lactam resistance (16- to 64-fold amoxicillin MIC reductions) [16]. Alterations in outer membrane protein profiles were observed in high-level amoxicillin-resistant isolates [89]. The outer membrane permeability of H. pylori to the small hydrophobic agent, 1-N-phenylnaphthylamine, was found to be higher than that of E. coli [17], an observation consistent with the low MIC values of many hydrophobic agents (Table 19.1). An increased susceptibility to metronidazole occurred in the presence of aspirin, which enhanced intracellular concentrations of tetracycline, but no significant changes in the transcriptional expression of the genes encoding the HopA, HopB, HopC, HopD, and HopE porins and HefABC efflux system were observed [122]. Hypersusceptibility to several hydrophobic agents (e.g., erythromycin, novobiocin, and rifampicin) was reported for mutants carrying null mutations in the ostA (also called imp) and/or msbA genes [27], which encode an organic solvent tolerance outer membrane protein and a lipopolysaccharide lipid precursor exporter, respectively – both of which are involved in the biogenesis of lipopolysaccharide [107, 123].
4.2 RND Pumps
Multiple putative RND pumps have been identified, based on protein homology, in several Helicobacter spp. (as presented in Table 19.3). The total numbers are fewer than those found in E. coli (which contains six RND pumps) or P. aeruginosa (>12 RND pumps) [94]; these differences are likely due to the relatively small genome sizes of Helicobacter spp. The four RND systems of H. pylori are each encoded by a putative three-gene operon [17], which produces the typical three components of RND tripartite efflux complexes; these components include an efflux transporter located in the cytoplasmic membrane, an accessory membrane fusion protein, and an outer membrane channel protein [94]. For two non-H. pylori species, putative RND pumps are instead each encoded by a two-gene operon [106] and likely requires an outer membrane channel protein encoded elsewhere in the genome for proper functioning (Table 19.3). Interestingly, unlike in E. coli or P. aeruginosa [94], no regulatory genes have been identified adjacent to the structural genes of these RND systems. One exception is HefGHI (also known as CrdB-CzcB-CzcA), where the encoded HP1326 (CrdA) is required for induction of HefGHI by copper [114]. CrdA expression is further controlled by a two-component regulatory system CrdRS (HP1364-1365) [124]. A recent study has demonstrated the importance of CrdRS in nitrosative response of H. pylori and its influence on the transcriptional expression of about 100 genes (including the upregulation of crdA) [125]. Overall, regulation of Helicobacter RND pump expression remains a mystery.
Phylogenetic analysis of the RND pumps of H. pylori suggest that HefC is closer to the RND pumps involved in drug efflux while HefF and HefC are related to RND pumps involved in the extrusion of divalent cations [17]. Further studies have been conducted to analyze their expression and functional roles [17, 113]. Despite their expression in wild-type cells, an early study used a genetic inactivation approach to suggest only a minimal role of HefABC, HefDEF, and HefGHI in the antimicrobial resistance in H. pylori [17]. Indeed, pretreatment of H. pylori cells with CCCP did not result in increased accumulation of either chloramphenicol or tetracycline (on the contrary, reduced drug accumulation was observed), arguing against involvement of a proton motive force-dependent drug efflux pump in intrinsic resistance to chloramphenicol or tetracycline [17].
Two other studies indicate a strong contribution of the HefABC efflux system to intrinsic and acquired multidrug resistance [15, 126]. The expression of hefABC in one study was generally the strongest among the four RND systems in clarithromycin-resistant (≥1.0 μg/ml clarithromycin) isolates [65]. Inactivation of the HefC pump gene in a wild-type strain rendered the mutant hypersusceptible to β-lactams (aztreonam, cefotaxime, ceftriaxone, penicillin, and piperacillin but not amoxicillin), clindamycin, erythromycin, ethidium bromide, novobiocin, and tetracycline with four- to 330-fold MIC reduction (Table 19.4) [15]. These antimicrobials are known substrates for RND pumps. Furthermore, CCCP treatment of wild-type cells increased the accumulation of ethidium bromide [15]. Chloramphenicol accumulation was increased slightly in CCCP-treated resistant cells [89]. However, susceptibility to quinolones was not affected by genetic disruption of the hefA or hefC gene [15, 126]. In another study, disruption of hefA (but not hefD, hefG, or HP1489) made the cells more susceptible to deoxycholate and novobiocin and the simultaneous inactivation of hefA and hefD sensitized cells to metronidazole [112]. Together, these results support that the HefABC pump plays an important role in the intrinsic drug resistance of H. pylori. This conclusion is further supported by the involvement of HefABC (not HefDEF or HefGHI) in resistance to bile salts and their derivatives, ceragenins [127]. Elevated hefA expression was noted in multidrug-resistant chloramphenicol-selected mutants [126] as well as in multidrug-resistant isolates from another study [128].
The contribution of HefC pumps to amoxicillin resistance was noted in certain resistant isolates, and the combination of 1-(1-naphthylmethyl)-piperazine (NMP; an RND pump inhibitor) at 100 μg/ml reduced the amoxicillin MIC by 16-fold in HefC overproducers [61]. In this case, given the overall hydrophilic nature of amoxicillin, one would expect that the efflux process alone may have a limited role in amoxicillin resistance, but this process may still be possible if the influx of amoxicillin is also affected by reduced porin expression (as already reported in the reference [16]; see above in the outer membrane permeability barrier section). Thus, it would be ideal to assess the porins for the isolates of this study [61]. Mutations in multiple genes were analyzed in this study [61], and surprisingly mutations in HefC (Asp131Glu and Leu378Phe) were identified in several resistant strains; these mutations appeared to yield a gain of function. In another paper, the inclusion of RND pump inhibitor phenylalanine-arginine β-naphthylamide (PAβN) reduced clarithromycin MIC values by four to eightfolds (from 4–32 to 1–8 μg/ml) for 15 clinical clarithromycin-resistant isolates [65] Lastly, HefABC overproduction was revealed to be the first step in the development of acquired resistance [72] to metronidazole. All of these results jointly suggest that HefABC plays a major role in acquired multidrug resistance.
Although the regulation of HefABC expression remains unknown, this efflux system is clearly inducible. The exposure of five clinical isolates to metronidazole at 8 or 16 μg/ml revealed a concentration-dependent increase in hefA expression, even though hefA was already constitutively expressed in these isolates [129]. The presence of cholesterol also induces hefABC expression and thus contributes to resistance to bile salts [127]. It was also found that the expression of all four RND systems was elevated in biofilm cells in comparison with that of planktonic cells [130].
Another RND pump of H. pylori, CznABC (i.e., HefDEF), is a metal efflux pump involved in cadmium, nickel, and zinc resistance. Only minimal growth occurred for pump-deficient mutants in the presence of cadmium (10 μM), nickel (1.2 mM), and zinc (0.8 mM) [113]. Furthermore, CznABC is also critical for gastric colonization and modulation of urease activity [113], providing a possible physiological role of RND pumps in H. pylori. (Urease activity plays an essential role in acid tolerance of H. pylori [131].) The third RND pump, HefGHI, confers resistance to copper, and its inactivation renders cells more susceptible to copper – with only minimal growth in the presence of 0.1 mM copper [114]. Based on the CrdRS system’s involvement in regulation of crdA (located immediately upstream of the hefGHI genes in the same transcriptional direction) [125], CrdRS could also influence hefGHI expression, and thus HefGHI may possibly contribute to nitrosative stress response. Indeed, the role of RND pumps in nitrosative stress response has been demonstrated in E. coli, Klebsiella pneumoniae, and P. aeruginosa [132–134]. Additionally, in contrast to the HefC pump, both HefF and HefH are not involved in cholesterol-dependent resistance to bile salts [127]. Inactivation of either the HefF or HefI pump did not alter the drug susceptibility of cells (with 20 tested agents) [15].
Several RND systems exist in H. hepaticus (Table 19.3). Intriguingly, H. hepaticus strains appear to be much less susceptible to amoxicillin than H. pylori, and they do not have PBP alterations nor produce β-lactamases [28]. Inactivation of hefA rendered a mutant strain hypersusceptible to amoxicillin (256-fold MIC reduction but not to another tested β-lactam tested, cefotaxime), rifampicin (ninefold MIC reduction), ofloxacin (fourfold MIC reduction), ethidium bromide (>fourfold MIC reduction), and bile salts (2.5- to 10-fold MIC decreases) [28]. Thus, HefABC likely contributes to intrinsic resistance in H. hepaticus to multiple agents including amoxicillin. Moreover, the expression of hefA is inducible by bile salts (but not by amoxicillin) [28]. This fact suggests that HefABC may be involved in the survival of H. hepaticus in the gastrointestinal tract where it would be exposed to high bile salt concentrations, similar to the role of the E. coli AcrAB-TolC system and the Campylobacter jejuni CmeABC system [135, 136].
4.3 Non-RND Pumps
This group includes ABC, MFS, MATE, and SMR pumps that remain to be characterized (Table 19.3) [105, 112, 115]. Inactivation of the ABC-type MsbA transporter rendered the mutant strain more susceptible to several agents including erythromycin and glutaraldehyde. The impact of CCCP treatment on the accumulation of ethidium bromide supported an efflux process contributed by MsbA [27]. This pump also cooperates synergistically with another lipopolysaccharide biogenesis protein OstA to enhance hydrophobic drug resistance [27]. However, since MsbA is a lipopolysaccharide flippase [107], mutants with MsbA deficiency (and/or with OstA defect) have a reduced lipopolysaccharide production [27]. A few ABC transporters such as CadA and CopA are involved in heavy metal resistance [114, 116].
The HP1165 protein is a homolog of the TetA(P) efflux pump of Clostridium perfringens belonging to the MFS family [83]. Its gene is constitutively expressed in any growth phase of a wild-type tetracycline-susceptible strain and its inactivation renders the mutant strain more susceptible to tetracycline (tenfold MIC reduction). While the overproduction of HP1165 is has been linked to tetracycline resistance following tetracycline exposure, its absence abolishes the ability of tetracycline to induce tetracycline resistance [83]. Additionally, the HP1181 protein is another putative MFS exporter, a homolog of the NorA pump of Staphylococcus aureus, but its functional properties remain to be characterized [115].
4.4 Effect of Efflux Pump Inhibitors and Methodological Considerations
As described above, many studies have employed efflux pump inhibitors in characterizing drug efflux contribution to resistance in Helicobacter. PAβN and NMP are known inhibitors of RND pumps [94]. We have been unable to find data on the activities of these two inhibitors alone against Helicobacter spp. (such as MIC values), but these values can be informative in assessing the effect of these agents themselves on Helicobacter [94]. In one study [65], PAβN was used at 10, 20, 40, 60, and 120 μg/ml, and it appeared that this agent alone at levels of up to 40 μg/ml did not affect the growth of a particular H. pylori strain [65]. Thus, the observations with effect of 40 μg/ml PAβN on the reduction of the MIC values of clarithromycin [65] and metronidazole [72] are interpreted as the involvement of an efflux mechanism. Similarly, NMP can be used at 100 μg/ml without detectable adverse impact on Helicobacter, and thus the effect of NMP on HefC is also considered to be related to efflux inhibition [61]. In this regard, it is worth mentioning that the inclusion of a plant extract (baicalin, berberine, emodin, or schizandrin) enhanced antibacterial activity of amoxicillin and tetracycline [128].
Multiple studies have used the proton conductor CCCP, which abolishes proton motive force across the cytoplasmic membrane and therefore is not an efflux pump inhibitor per se. In two independent studies, CCCP at 40 and 100 μM reduced (instead of increased) the accumulation of chloramphenicol and tetracycline [17, 122], suggesting an impact on uptake processes [17]. However, two other studies showed an increase in accumulation of ethidium bromide or chloramphenicol in the presence of 10 or 100 μM CCCP [27, 89]. No impact of 40 or 100 μM CCCP on penicillin and tetracycline resistance was also reported [61, 89]. CCCP at 100 μM produced more effects in drug-CCCP combination susceptibility testing on chloramphenicol-selected multidrug-resistant isolates than the parental strains for multiple drugs [137]. There are also studies that used CCCP at a high level of 200 μM, which increased accumulation of ethidium bromide and tetracycline [84, 138]. Given the apparently inconsistent results on the effect of CCCP on drug accumulation in H. pylori, additional investigations are needed to carefully reassess the use of CCCP including its appropriate concentrations.
H. pylori infection also requires the treatment with the proton pump inhibitor acid-inhibitory drugs as part of a drug combination regimen. Proton pump inhibitors themselves exhibit anti-H. pylori activities at the levels which are not achievable in vivo [13]. Interestingly, studies have examined the effect of proton pump inhibitors on the multidrug resistance phenotype of either bacterial isolates or mammalian tumor cells [139–141]. Specific to H. pylori, esomeprazole, lansoprazole, omeprazole, pantoprazole, and rabeprazole (each at 10 μg/ml) were found to exhibit certain reduction of MICs of amoxicillin and metronidazole (particularly with pantoprazole and rabeprazole) [137]. Yet, clinical relevance of this observation remains unknown.
5 Concluding Remarks
The treatment of Helicobacter infections is adversely affected by the increasing emergence of acquired resistance in H. pylori. Even though drug-specific mechanisms are predominantly responsible for the clinically relevant resistance phenotypes that compromise H. pylori infection therapy effectiveness, the contribution of drug efflux systems (particularly the RND-type pumps) to intrinsic and acquired multidrug resistance in Helicobacter spp. is being recognized. Furthermore, as already observed in other bacteria of the same class as Helicobacter spp., these efflux systems likely also function beyond drug resistance and are involved in its pathogenesis. However, the data regarding the role of the HefABC pump and its substrate profile vary within the literature. These discrepancies may be due to methodological challenges in conducting antimicrobial susceptibility testing with H. pylori as well as the high susceptibility of the wild-type H. pylori to many agents in vitro. A better understanding of Helicobacter drug efflux pumps should be pursued to characterize better strategies for therapeutic interventions. Agents that inhibit the efflux pumps could serve as antimicrobial adjuvants to improve the activities of the existing anti-Helicobacter drugs. Furthermore, an open-ended question remains on how expression of these efflux systems is regulated. Physiological roles of the Helicobacter drug efflux systems are also likely linked to stress response or colonization [113, 125, 127]. The current knowledge clearly warrants further investigations of Helicobacter drug efflux pumps and, in particular, their regulation and functional roles.
References
Dunn BE, Cohen H, Blaser MJ (1997) Helicobacter pylori. Clin Microbiol Rev 10:720–741
Polk DB, Peek RM Jr (2010) Helicobacter pylori: gastric cancer and beyond. Nat Rev Cancer 10:403–414. doi:10.1038/nrc2857
Correa P, Houghton J (2007) Carcinogenesis of Helicobacter pylori. Gastroenterology 133:659–672. doi:10.1053/j.gastro.2007.06.026
Lawson AJ (2011) Chapter 54. Helicobacter. In: Versalovic J (ed) Manual of clinical microbiology, vol 1, 10th edn. ASM Press, Washington, DC, pp 900–915
Okoli AS, Menard A, Mendz GL (2009) Helicobacter spp. other than Helicobacter pylori. Helicobacter 14(Suppl 1):69–74. doi:10.1111/j.1523-5378.2009.00701.x
Rossi M, Hanninen ML (2012) Helicobacter spp. other than H. pylori. Helicobacter 17(Suppl 1):56–61. doi:10.1111/j.1523-5378.2012.00984.x
Segura-Lopez FK, Guitron-Cantu A, Torres J (2015) Association between Helicobacter spp. infections and hepatobiliary malignancies: a review. World J Gastroenterol 21:1414–1423. doi:10.3748/wjg.v21.i5.1414
Goldman CG, Mitchell HM (2010) Helicobacter spp. other than Helicobacter pylori. Helicobacter 15(Suppl 1):69–75. doi:10.1111/j.1523-5378.2010.00780.x
Schulz C, Koch N, Schutte K, Pieper DH, Malfertheiner P (2015) H. pylori and its modulation of gastrointestinal microbiota. J Dig Dis 16:109–117. doi:10.1111/1751-2980.12233
Khosravi Y, Seow SW, Amoyo AA, Chiow KH, Tan TL, Wong WY, Poh QH, Sentosa IM et al (2015) Helicobacter pylori infection can affect energy modulating hormones and body weight in germ free mice. Sci Rep 5:8731. doi:10.1038/srep08731
Li J, Butcher J, Mack D, Stintzi A (2015) Functional impacts of the intestinal microbiome in the pathogenesis of inflammatory bowel disease. Inflamm Bowel Dis 21:139–153. doi:10.1097/MIB.0000000000000215
Fock KM, Graham DY, Malfertheiner P (2013) Helicobacter pylori research: historical insights and future directions. Nat Rev Gastroenterol Hepatol 10:495–500. doi:10.1038/nrgastro.2013.96
Megraud F, Lehours P (2007) Helicobacter pylori detection and antimicrobial susceptibility testing. Clin Microbiol Rev 20:280–322. doi:10.1128/CMR.00033-06
Gisbert JP, Calvet X (2012) Review article: rifabutin in the treatment of refractory Helicobacter pylori infection. Aliment Pharmacol Ther 35:209–221. doi:10.1111/j.1365-2036.2011.04937.x
Kutschke A, de Jonge BL (2005) Compound efflux in Helicobacter pylori. Antimicrob Agents Chemother 49:3009–3010. doi:10.1128/AAC.49.7.3009-3010.2005
Co EM, Schiller NL (2006) Resistance mechanisms in an in vitro-selected amoxicillin-resistant strain of Helicobacter pylori. Antimicrob Agents Chemother 50:4174–4176. doi:10.1128/AAC.00759-06
Bina JE, Alm RA, Uria-Nickelsen M, Thomas SR, Trust TJ, Hancock RE (2000) Helicobacter pylori uptake and efflux: basis for intrinsic susceptibility to antibiotics in vitro. Antimicrob Agents Chemother 44:248–254. doi:10.1128/AAC.44.2.248-254.2000
Yang JC, Lee PI, Hsueh PR (2010) In vitro activity of nemonoxacin, tigecycline, and other antimicrobial agents against Helicobacter pylori isolates in Taiwan, 1998–2007. Eur J Clin Microbiol Infect Dis 29:1369–1375. doi:10.1007/s10096-010-1009-9
Francesco VD, Zullo A, Hassan C, Giorgio F, Rosania R, Ierardi E (2011) Mechanisms of Helicobacter pylori antibiotic resistance: an updated appraisal. World J Gastrointest Pathophysiol 2:35–41. doi:10.4291/wjgp.v2.i3.35
Binh TT, Shiota S, Suzuki R, Matsuda M, Trang TT, Kwon DH, Iwatani S, Yamaoka Y (2014) Discovery of novel mutations for clarithromycin resistance in Helicobacter pylori by using next-generation sequencing. J Antimicrob Chemother 69:1796–1803. doi:10.1093/jac/dku050
Taylor DE (2000) Pathophysiology of antibiotic resistance: clarithromycin. Can J Gastroenterol 14:891–894
Haas CE, Nix DE, Schentag JJ (1990) In vitro selection of resistant Helicobacter pylori. Antimicrob Agents Chemother 34:1637–1641. doi:10.1128/AAC.34.9.1637
Heep M, Beck D, Bayerdorffer E, Lehn N (1999) Rifampin and rifabutin resistance mechanism in Helicobacter pylori. Antimicrob Agents Chemother 43:1497–1499
Nishizawa T, Suzuki H, Matsuzaki J, Muraoka H, Tsugawa H, Hirata K, Hibi T (2011) Helicobacter pylori resistance to rifabutin in the last 7 years. Antimicrob Agents Chemother 55:5374–5375. doi:10.1128/AAC.05437-11
Gerrits MM, de Zoete MR, Arents NL, Kuipers EJ, Kusters JG (2002) 16S rRNA mutation-mediated tetracycline resistance in Helicobacter pylori. Antimicrob Agents Chemother 46:2996–3000. doi:10.1128/AAC.46.9.2996-3000.2002
Hoffman PS, Bruce AM, Olekhnovich I, Warren CA, Burgess SL, Hontecillas R, Viladomiu M, Bassaganya-Riera J et al (2014) Preclinical studies of amixicile, a systemic therapeutic developed for treatment of Clostridium difficile infections that also shows efficacy against Helicobacter pylori. Antimicrob Agents Chemother 58:4703–4712. doi:10.1128/AAC.03112-14
Chiu HC, Lin TL, Yang JC, Wang JT (2009) Synergistic effect of imp/ostA and msbA in hydrophobic drug resistance of Helicobacter pylori. BMC Microbiol 9:136. doi:10.1186/1471-2180-9-136
Belzer C, Stoof J, Breijer S, Kusters JG, Kuipers EJ, van Vliet AH (2009) The Helicobacter hepaticus hefA gene is involved in resistance to amoxicillin. Helicobacter 14:72–79. doi:10.1111/j.1523-5378.2009.00661.x
Megraud F, Marshall BJ (2000) How to treat Helicobacter pylori. First-line, second-line, and future therapies. Gastroenterol Clin North Am 29:759–773. doi:10.1016/S0889-8553(05)70145-X
Wannmacher L (2011) Review of the evidence for H. pylori treatment regimens. (The 18th Expert Committee on the Selection and Use of Essential Medicines. March 21–25, 2011) http://www.who.int/selection_medicines/committees/expert/18/applications/Review_171.pdf. Accessed 12 Mar 2016
Xie C, Lu NH (2015) Review: clinical management of Helicobacter pylori infection in China. Helicobacter 20:1–10. doi:10.1111/hel.12178
Matsuzaki J, Suzuki H, Nishizawa T, Hirata K, Tsugawa H, Saito Y, Okada S, Fukuhara S et al (2012) Efficacy of sitafloxacin-based rescue therapy for Helicobacter pylori after failures of first- and second-line therapies. Antimicrob Agents Chemother 56:1643–1645. doi:10.1128/AAC.05941-11
Hirata Y, Ohmae T, Yanai A, Sakitani K, Hayakawa Y, Yoshida S, Sugimoto T, Mitsuno Y et al (2012) Sitafloxacin resistance in Helicobacter pylori isolates and sitafloxacin-based triple therapy as a third-line regimen in Japan. Int J Antimicrob Agents 39:352–355. doi:10.1016/j.ijantimicag.2011.12.002
Liu X, Wang H, Lv Z, Wang Y, Wang B, Xie Y, Zhou X, Lv N (2015) Rescue therapy with a proton pump inhibitor plus amoxicillin and rifabutin for Helicobacter pylori infection: a systematic review and meta-analysis. Gastroenterol Res Pract 2015:415648. doi:10.1155/2015/415648
Zhang Y, Gao W, Cheng H, Zhang X, Hu F (2014) Tetracycline- and furazolidone-containing quadruple regimen as rescue treatment for Helicobacter pylori infection: a single center retrospective study. Helicobacter 19:382–386. doi:10.1111/hel.12143
Megraud F (2001) Resistance of Helicobacter pylori to antibiotics and its impact on treatment options. Drug Resist Updat 4:178–186. doi:10.1054/drup.2001.0203
Alfizah H, Norazah A, Hamizah R, Ramelah M (2014) Resistotype of Helicobacter pylori isolates: the impact on eradication outcome. J Med Microbiol 63:703–709. doi:10.1099/jmm.0.069781-0
Draeger S, Wuppenhorst N, Kist M, Glocker EO (2015) Outcome of second- and third-line Helicobacter pylori eradication therapies based on antimicrobial susceptibility testing. J Antimicrob Chemother 70:3141–3145. doi:10.1093/jac/dkv223
Wu IT, Chuah SK, Lee CH, Liang CM, Lu LS, Kuo YH, Yen YH, Hu ML et al (2015) Five-year sequential changes in secondary antibiotic resistance of Helicobacter pylori in Taiwan. World J Gastroenterol 21:10669–10674. doi:10.3748/wjg.v21.i37.10669
Romano M, Iovene MR, Russo MI, Rocco A, Salerno R, Cozzolino D, Pilloni AP, Tufano MA et al (2008) Failure of first-line eradication treatment significantly increases prevalence of antimicrobial-resistant Helicobacter pylori clinical isolates. J Clin Pathol 61:1112–1115. doi:10.1136/jcp.2008.060392
Nishizawa T, Suzuki H, Tsugawa H, Muraoka H, Matsuzaki J, Hirata K, Ikeda F, Takahashi M et al (2011) Enhancement of amoxicillin resistance after unsuccessful Helicobacter pylori eradication. Antimicrob Agents Chemother 55:3012–3014. doi:10.1128/AAC.00188-11
Nishizawa T, Suzuki H (2014) Mechanisms of Helicobacter pylori antibiotic resistance and molecular testing. Front Mol Biosci 1:19. doi:10.3389/fmolb.2014.00019
Megraud F, Coenen S, Versporten A, Kist M, Lopez-Brea M, Hirschl AM, Andersen LP, Goossens H et al (2013) Helicobacter pylori resistance to antibiotics in Europe and its relationship to antibiotic consumption. Gut 62:34–42. doi:10.1136/gutjnl-2012-302254
Sun QJ, Liang X, Zheng Q, Gu WQ, Liu WZ, Xiao SD, Lu H (2010) Resistance of Helicobacter pylori to antibiotics from 2000 to 2009 in Shanghai. World J Gastroenterol 16:5118–5121. doi:10.3748/wjg.v16.i40.5118
Su P, Li Y, Li H, Zhang J, Lin L, Wang Q, Guo F, Ji Z et al (2013) Antibiotic resistance of Helicobacter pylori isolated in the southeast coastal region of China. Helicobacter 18:274–279. doi:10.1111/hel.12046
Glocker E, Bogdan C, Kist M (2007) Characterization of rifampicin-resistant clinical Helicobacter pylori isolates from Germany. J Antimicrob Chemother 59:874–879. doi:10.1093/jac/dkm039
Wueppenhorst N, Stueger HP, Kist M, Glocker E (2009) Identification and molecular characterization of triple- and quadruple-resistant Helicobacter pylori clinical isolates in Germany. J Antimicrob Chemother 63:648–653. doi:10.1093/jac/dkp003
Lee JW, Kim N, Nam RH, Kim JM, Park JY, Lee SM, Kim JS, Lee DH et al (2015) High antimicrobial efficacy of finafloxacin on Helicobacter pylori isolates at pH 5.0 compared with those of other fluoroquinolones. Antimicrob Agents Chemother 59:7629–7636. doi:10.1128/AAC.01467-15
Boyanova L, Evstatiev I, Gergova G, Yaneva P, Mitov I (2015) Linezolid susceptibility in Helicobacter pylori, including strains with multidrug resistance. Int J Antimicrob Agents 46:703–706. doi:10.1016/j.ijantimicag.2015.08.010
El-Halfawy OM, Valvano MA (2015) Antimicrobial heteroresistance: an emerging field in need of clarity. Clin Microbiol Rev 28:191–207. doi:10.1128/CMR.00058-14
Matteo MJ, Granados G, Olmos M, Wonaga A, Catalano M (2008) Helicobacter pylori amoxicillin heteroresistance due to point mutations in PBP-1A in isogenic isolates. J Antimicrob Chemother 61:474–477. doi:10.1093/jac/dkm504
Kao CY, Lee AY, Huang AH, Song PY, Yang YJ, Sheu SM, Chang WL, Sheu BS et al (2014) Heteroresistance of Helicobacter pylori from the same patient prior to antibiotic treatment. Infect Genet Evol 23:196–202. doi:10.1016/j.meegid.2014.02.009
Matteo MJ, Perez CV, Domingo MR, Olmos M, Sanchez C, Catalano M (2006) DNA sequence analysis of rdxA and frxA from paired metronidazole-sensitive and -resistant Helicobacter pylori isolates obtained from patients with heteroresistance. Int J Antimicrob Agents 27:152–158. doi:10.1016/j.ijantimicag.2005.09.019
Jakobsson HE, Jernberg C, Andersson AF, Sjolund-Karlsson M, Jansson JK, Engstrand L (2010) Short-term antibiotic treatment has differing long-term impacts on the human throat and gut microbiome. PLoS One 5:e9836. doi:10.1371/journal.pone.0009836
Sjolund M, Wreiber K, Andersson DI, Blaser MJ, Engstrand L (2003) Long-term persistence of resistant Enterococcus species after antibiotics to eradicate Helicobacter pylori. Ann Intern Med 139:483–487. doi:10.7326/0003-4819-139-6-200309160-00011
Gaddy JA, Radin JN, Cullen TW, Chazin WJ, Skaar EP, Trent MS, Algood HM (2015) Helicobacter pylori resists the antimicrobial activity of calprotectin via lipid A modification and associated biofilm formation. mBio 6:e01349–15. doi:10.1128/mBio.01349-15
Wang G, Wilson TJ, Jiang Q, Taylor DE (2001) Spontaneous mutations that confer antibiotic resistance in Helicobacter pylori. Antimicrob Agents Chemother 45:727–733. doi:10.1128/AAC.45.3.727-733.2001
DeLoney CR, Schiller NL (1999) Competition of various β-lactam antibiotics for the major penicillin-binding proteins of Helicobacter pylori: antibacterial activity and effects on bacterial morphology. Antimicrob Agents Chemother 43:2702–2709
DeLoney CR, Schiller NL (2000) Characterization of an in vitro-selected amoxicillin-resistant strain of Helicobacter pylori. Antimicrob Agents Chemother 44:3368–3373. doi:10.1128/AAC.44.12.3368-3373.2000
Tseng YS, Wu DC, Chang CY, Kuo CH, Yang YC, Jan CM, Su YC, Kuo FC et al (2009) Amoxicillin resistance with β-lactamase production in Helicobacter pylori. Eur J Clin Invest 39:807–812. doi:10.1111/j.1365-2362.2009.02166.x
Qureshi NN, Gallaher B, Schiller NL (2014) Evolution of amoxicillin resistance of Helicobacter pylori in vitro: characterization of resistance mechanisms. Microb Drug Resist 20:509–516. doi:10.1089/mdr.2014.0019
Versalovic J, Shortridge D, Kibler K, Griffy MV, Beyer J, Flamm RK, Tanaka SK, Graham DY et al (1996) Mutations in 23S rRNA are associated with clarithromycin resistance in Helicobacter pylori. Antimicrob Agents Chemother 40:477–480
Occhialini A, Urdaci M, Doucet-Populaire F, Bebear CM, Lamouliatte H, Megraud F (1997) Macrolide resistance in Helicobacter pylori: rapid detection of point mutations and assays of macrolide binding to ribosomes. Antimicrob Agents Chemother 41:2724–2728
Agudo S, Perez-Perez G, Alarcon T, Lopez-Brea M (2010) High prevalence of clarithromycin-resistant Helicobacter pylori strains and risk factors associated with resistance in Madrid, Spain. J Clin Microbiol 48:3703–3707. doi:10.1128/JCM.00144-10
Hirata K, Suzuki H, Nishizawa T, Tsugawa H, Muraoka H, Saito Y, Matsuzaki J, Hibi T (2010) Contribution of efflux pumps to clarithromycin resistance in Helicobacter pylori. J Gastroenterol Hepatol 25(Suppl 1):S75–S79. doi:10.1111/j.1440-1746.2009.06220.x
Kwon DH, Lee M, Kim JJ, Kim JG, El-Zaatari FA, Osato MS, Graham DY (2001) Furazolidone- and nitrofurantoin-resistant Helicobacter pylori: prevalence and role of genes involved in metronidazole resistance. Antimicrob Agents Chemother 45:306–308. doi:10.1128/AAC.45.1.306-308.2001
Chatterjee SN, Ghose S, Maiti M (1977) Cross linking of deoxyribonucleic acid in furazolidone treated Vibrio cholerae cell. Biochem Pharmacol 26:1453–1454. doi:10.1016/0006-2952(77)90377-X
Martinez-Puchol S, Gomes C, Pons MJ, Ruiz-Roldan L, Torrents de la Pena A, Ochoa TJ, Ruiz J (2015) Development and analysis of furazolidone-resistant Escherichia coli mutants. APMIS 123:676–681. doi:10.1111/apm.12401
Su Z, Xu H, Zhang C, Shao S, Li L, Wang H, Wang H, Qiu G (2006) Mutations in Helicobacter pylori porD and oorD genes may contribute to furazolidone resistance. Croat Med J 47:410–415
Rimbara E, Noguchi N, Kawai T, Sasatsu M (2012) Fluoroquinolone resistance in Helicobacter pylori: role of mutations at position 87 and 91 of GyrA on the level of resistance and identification of a resistance conferring mutation in GyrB. Helicobacter 17:36–42. doi:10.1111/j.1523-5378.2011.00912.x
Suzuki H, Nishizawa T, Muraoka H, Hibi T (2009) Sitafloxacin and garenoxacin may overcome the antibiotic resistance of Helicobacter pylori with gyrA mutation. Antimicrob Agents Chemother 53:1720–1721. doi:10.1128/AAC.00049-09
Tsugawa H, Suzuki H, Muraoka H, Ikeda F, Hirata K, Matsuzaki J, Saito Y, Hibi T (2011) Enhanced bacterial efflux system is the first step to the development of metronidazole resistance in Helicobacter pylori. Biochem Biophys Res Commun 404:656–660. doi:10.1016/j.bbrc.2010.12.034
Masaoka H, Suzuki H, Kurabayashi K, Nomoto Y, Nishizawa T, Mori M, Hibi T (2006) Could frameshift mutations in the frxA and rdxA genes of Helicobacter pylori be a marker for metronidazole resistance? Aliment Pharmacol Ther 24:81–87. doi:10.1111/j.1746-6342.2006.00029.x
Tsugawa H, Suzuki H, Satoh K, Hirata K, Matsuzaki J, Saito Y, Suematsu M, Hibi T (2011) Two amino acids mutation of ferric uptake regulator determines Helicobacter pylori resistance to metronidazole. Antioxid Redox Signal 14:15–23. doi:10.1089/ars.2010.3146
Hoffman PS, Goodwin A, Johnsen J, Magee K, Veldhuyzen van Zanten SJ (1996) Metabolic activities of metronidazole-sensitive and -resistant strains of Helicobacter pylori: repression of pyruvate oxidoreductase and expression of isocitrate lyase activity correlate with resistance. J Bacteriol 178:4822–4829
Heep M, Odenbreit S, Beck D, Decker J, Prohaska E, Rieger U, Lehn N (2000) Mutations at four distinct regions of the rpoB gene can reduce the susceptibility of Helicobacter pylori to rifamycins. Antimicrob Agents Chemother 44:1713–1715. doi:10.1128/AAC.44.6.1713-1715.2000
Suzuki S, Suzuki H, Nishizawa T, Kaneko F, Ootani S, Muraoka H, Saito Y, Kobayashi I et al (2009) Past rifampicin dosing determines rifabutin resistance of Helicobacter pylori. Digestion 79:1–4. doi:10.1159/000191204
Chopra I, Roberts M (2001) Tetracycline antibiotics: mode of action, applications, molecular biology, and epidemiology of bacterial resistance. Microbiol Mol Biol Rev 65:232–260. doi:10.1128/MMBR.65.2.232-260.2001
Kwon DH, Kim JJ, Lee M, Yamaoka Y, Kato M, Osato MS, El-Zaatari FA, Graham DY (2000) Isolation and characterization of tetracycline-resistant clinical isolates of Helicobacter pylori. Antimicrob Agents Chemother 44:3203–3205. doi:10.1128/AAC.44.11.3203-3205.2000
Trieber CA, Taylor DE (2002) Mutations in the 16S rRNA genes of Helicobacter pylori mediate resistance to tetracycline. J Bacteriol 184:2131–2140. doi:10.1128/JB.184.8.2131-2140.2002
Toledo H, Lopez-Solis R (2010) Tetracycline resistance in Chilean clinical isolates of Helicobacter pylori. J Antimicrob Chemother 65:470–473. doi:10.1093/jac/dkp457
Khan R, Nahar S, Mukhopadhyay AK, Berg DE, Ahmad MM, Okamoto K, Nair GB, Rahman M (2008) Isolation of tetracycline-resistant clinical Helicobacter pylori without mutations in 16S rRNA gene in Bangladesh. Microbiol Imunol 52:508–511. doi:10.1111/j.1348-0421.2008.00062.x
Li Y, Dannelly HK (2006) Inactivation of the putative tetracycline resistance gene HP1165 in Helicobacter pylori led to loss of inducible tetracycline resistance. Arch Microbiol 185:255–262. doi:10.1007/s00203-006-0093-9
Anoushiravani M, Falsafi T, Niknam V (2009) Proton motive force-dependent efflux of tetracycline in clinical isolates of Helicobacter pylori. J Med Microbiol 58:1309–1313. doi:10.1099/jmm.0.010876-0
Paul R, Postius S, Melchers K, Schafer KP (2001) Mutations of the Helicobacter pylori genes rdxA and pbp1 cause resistance against metronidazole and amoxicillin. Antimicrob Agents Chemother 45:962–965. doi:10.1128/AAC.45.3.962-965.2001
Mittl PR, Luthy L, Hunziker P, Grutter MG (2000) The cysteine-rich protein A from Helicobacter pylori is a β-lactamase. J Biol Chem 275:17693–17699. doi:10.1074/jbc.M001869200
Deml L, Aigner M, Decker J, Eckhardt A, Schutz C, Mittl PR, Barabas S, Denk S et al (2005) Characterization of the Helicobacter pylori cysteine-rich protein A as a T-helper cell type 1 polarizing agent. Infect Immun 73:4732–4742. doi:10.1128/IAI.73.8.4732-4742.2005
Dumrese C, Slomianka L, Ziegler U, Choi SS, Kalia A, Fulurija A, Lu W, Berg DE et al (2009) The secreted Helicobacter cysteine-rich protein A causes adherence of human monocytes and differentiation into a macrophage-like phenotype. FEBS Lett 583:1637–1643. doi:10.1016/j.febslet.2009.04.027
Kwon DH, Dore MP, Kim JJ, Kato M, Lee M, Wu JY, Graham DY (2003) High-level β-lactam resistance associated with acquired multidrug resistance in Helicobacter pylori. Antimicrob Agents Chemother 47:2169–2178. doi:10.1128/AAC.47.7.2169-2178.2003
Nikaido H (1989) Outer membrane barrier as a mechanism of antimicrobial resistance. Antimicrob Agents Chemother 33:1831–1836. doi:10.1128/AAC.33.11.1831
Eng NF, Ybazeta G, Chapman K, Fraleigh NL, Letto R, Altman E, Diaz-Mitoma F (2015) Antimicrobial susceptibility of Canadian isolates of Helicobacter pylori in Northeastern Ontario. Can J Infect Dis Med Microbiol 26:137–144. doi:10.1155/2015/853287
Iwamoto A, Tanahashi T, Okada R, Yoshida Y, Kikuchi K, Keida Y, Murakami Y, Yang L et al (2014) Whole-genome sequencing of clarithromycin resistant Helicobacter pylori characterizes unidentified variants of multidrug resistant efflux pump genes. Gut Pathog 6:27. doi:10.1186/1757-4749-6-27
Li X-Z, Nikaido H (2009) Efflux-mediated drug resistance in bacteria: an update. Drugs 69:1555–1623. doi:10.2165/11317030-000000000-00000
Li X-Z, Plésiat P, Nikaido H (2015) The challenge of efflux-mediated antibiotic resistance in Gram-negative bacteria. Clin Microbiol Rev 28:337–418. doi:10.1128/CMR.00117-14
Kwon DH, Kato M, El-Zaatari FA, Osato MS, Graham DY (2000) Frame-shift mutations in NAD(P)H flavin oxidoreductase encoding gene (frxA) from metronidazole resistant Helicobacter pylori ATCC43504 and its involvement in metronidazole resistance. FEMS Microbiol Lett 188:197–202. doi:10.1111/j.1574-6968.2000.tb09193.x
Jeong JY, Mukhopadhyay AK, Akada JK, Dailidiene D, Hoffman PS, Berg DE (2001) Roles of FrxA and RdxA nitroreductases of Helicobacter pylori in susceptibility and resistance to metronidazole. J Bacteriol 183:5155–5162. doi:10.1128/JB.183.17.5155-5162.2001
Binh TT, Suzuki R, Trang TT, Kwon DH, Yamaoka Y (2015) Search for novel candidate mutations for metronidazole resistance in Helicobacter pylori using next-generation sequencing. Antimicrob Agents Chemother 59:2343–2348. doi:10.1128/AAC.04852-14
Gerrits MM, van der Wouden EJ, Bax DA, van Zwet AA, van Vliet AH, de Jong A, Kusters JG, Thijs JC et al (2004) Role of the rdxA and frxA genes in oxygen-dependent metronidazole resistance of Helicobacter pylori. J Med Microbiol 53:1123–1128. doi:10.1099/jmm.0.45701-0
Whiteway J, Koziarz P, Veall J, Sandhu N, Kumar P, Hoecher B, Lambert IB (1998) Oxygen-insensitive nitroreductases: analysis of the roles of nfsA and nfsB in development of resistance to 5-nitrofuran derivatives in Escherichia coli. J Bacteriol 180:5529–5539
Nonaka L, Connell SR, Taylor DE (2005) 16S rRNA mutations that confer tetracycline resistance in Helicobacter pylori decrease drug binding in Escherichia coli ribosomes. J Bacteriol 187:3708–3712. doi:10.1128/JB.187.11.3708-3712.2005
Dailidiene D, Bertoli MT, Miciuleviciene J, Mukhopadhyay AK, Dailide G, Pascasio MA, Kupcinskas L, Berg DE (2002) Emergence of tetracycline resistance in Helicobacter pylori: multiple mutational changes in 16S ribosomal DNA and other genetic loci. Antimicrob Agents Chemother 46:3940–3946. doi:10.1128/AAC.46.12.3940-3946.2002
Lawson AJ, Elviss NC, Owen RJ (2005) Real-time PCR detection and frequency of 16S rDNA mutations associated with resistance and reduced susceptibility to tetracycline in Helicobacter pylori from England and Wales. J Antimicrob Chemother 56:282–286. doi:10.1093/jac/dki199
Cambau E, Allerheiligen V, Coulon C, Corbel C, Lascols C, Deforges L, Soussy CJ, Delchier JC et al (2009) Evaluation of a new test, genotype HelicoDR, for molecular detection of antibiotic resistance in Helicobacter pylori. J Clin Microbiol 47:3600–3607. doi:10.1128/JCM.00744-09
Lee JW, Kim N, Nam RH, Park JH, Choi YJ, Kim JM, Kim JS, Jung HC (2014) GenoType HelicoDR test in the determination of antimicrobial resistance of Helicobacter pylori in Korea. Scand J Gastroenterol 49:1058–1067. doi:10.3109/00365521.2014.894117
Tomb JF, White O, Kerlavage AR, Clayton RA, Sutton GG, Fleischmann RD, Ketchum KA, Klenk HP et al (1997) The complete genome sequence of the gastric pathogen Helicobacter pylori. Nature 388:539–547. doi:10.1038/41483
Morita Y, Tomida J, Kawamura Y (2012) Multidrug efflux systems in Helicobacter cinaedi. Antibiotics 1:29–43. doi:10.3390/antibiotics1010029
Liechti G, Goldberg JB (2012) Outer membrane biogenesis in Escherichia coli, Neisseria meningitidis, and Helicobacter pylori: paradigm deviations in H. pylori. Front Cell Infect Microbiol 2:29. doi:10.3389/fcimb.2012.00029
Suerbaum S, Josenhans C, Sterzenbach T, Drescher B, Brandt P, Bell M, Droge M, Fartmann B et al (2003) The complete genome sequence of the carcinogenic bacterium Helicobacter hepaticus. Proc Natl Acad Sci U S A 100:7901–7906. doi:10.1073/pnas.1332093100
Arnold IC, Zigova Z, Holden M, Lawley TD, Rad R, Dougan G, Falkow S, Bentley SD et al (2011) Comparative whole genome sequence analysis of the carcinogenic bacterial model pathogen Helicobacter felis. Genome Biol Evol 3:302–308. doi:10.1093/gbe/evr022
Goto T, Ogura Y, Hirakawa H, Tomida J, Morita Y, Akaike T, Hayashi T, Kawamura Y (2012) Complete genome sequence of Helicobacter cinaedi strain PAGU611, isolated in a case of human bacteremia. J Bacteriol 194:3744–3745. doi:10.1128/JB.00645-12
Bina J, Bains M, Hancock RE (2000) Functional expression in Escherichia coli and membrane topology of porin HopE, a member of a large family of conserved proteins in Helicobacter pylori. J Bacteriol 182:2370–2375. doi:10.1128/JB.182.9.2370-2375.2000
van Amsterdam K, Bart A, van der Ende A (2005) A Helicobacter pylori TolC efflux pump confers resistance to metronidazole. Antimicrob Agents Chemother 49:1477–1482. doi:10.1128/AAC.49.4.1477-1482.2005
Stahler FN, Odenbreit S, Haas R, Wilrich J, Van Vliet AH, Kusters JG, Kist M, Bereswill S (2006) The novel Helicobacter pylori CznABC metal efflux pump is required for cadmium, zinc, and nickel resistance, urease modulation, and gastric colonization. Infect Immun 74:3845–3852. doi:10.1128/IAI.02025-05
Waidner B, Melchers K, Ivanov I, Loferer H, Bensch KW, Kist M, Bereswill S (2002) Identification by RNA profiling and mutational analysis of the novel copper resistance determinants CrdA (HP1326), CrdB (HP1327), and CzcB (HP1328) in Helicobacter pylori. J Bacteriol 184:6700–6708. doi:10.1128/JB.184.23.6700-6708.2002
Morrison S, Ward A, Hoyle CJ, Henderson PJ (2003) Cloning, expression, purification and properties of a putative multidrug resistance efflux protein from Helicobacter pylori. Int J Antimicrob Agents 22:242–249. doi:10.1016/S0924-8579(03)00222-X
Herrmann L, Schwan D, Garner R, Mobley HL, Haas R, Schafer KP, Melchers K (1999) Helicobacter pylori cadA encodes an essential Cd(II)-Zn(II)-Co(II) resistance factor influencing urease activity. Mol Microbiol 33:524–536. doi:10.1046/j.1365-2958.1999.01496.x
Nikaido H (2003) Molecular basis of bacterial outer membrane permeability revisited. Microbiol Mol Biol Rev 67:593–656. doi:10.1128/MMBR.67.4.593-656.2003
Snelling WJ, Moran AP, Ryan KA, Scully P, McGourty K, Cooney JC, Annuk H, O’Toole PW (2007) HorB (HP0127) is a gastric epithelial cell adhesion. Helicobacter 12:200–209. doi:10.1111/j.1523-5378.2007.00499.x
Odenbreit S, Swoboda K, Barwig I, Ruhl S, Boren T, Koletzko S, Haas R (2009) Outer membrane protein expression profile in Helicobacter pylori clinical isolates. Infect Immun 77:3782–3790. doi:10.1128/IAI.00364-09
Exner MM, Doig P, Trust TJ, Hancock RE (1995) Isolation and characterization of a family of porin proteins from Helicobacter pylori. Infect Immun 63:1567–1572
Doig P, Exner MM, Hancock RE, Trust TJ (1995) Isolation and characterization of a conserved porin protein from Helicobacter pylori. J Bacteriol 177:5447–5452
Zhang XP, Wang WH, Tian Y, Gao W, Li J (2009) Aspirin increases susceptibility of Helicobacter pylori to metronidazole by augmenting endocellular concentrations of antimicrobials. World J Gastroenterol 15:919–926. doi:10.3748/wjg.15.919
Abe S, Okutsu T, Nakajima H, Kakuda N, Ohtsu I, Aono R (2003) n-Hexane sensitivity of Escherichia coli due to low expression of imp/ostA encoding an 87 kDa minor protein associated with the outer membrane. Microbiology 149:1265–1273. doi:10.1099/mic.0.25927-0
Waidner B, Melchers K, Stahler FN, Kist M, Bereswill S (2005) The Helicobacter pylori CrdRS two-component regulation system (HP1364/HP1365) is required for copper-mediated induction of the copper resistance determinant CrdA. J Bacteriol 187:4683–4688. doi:10.1128/JB.187.13.4683-4688.2005
Hung CL, Cheng HH, Hsieh WC, Tsai ZT, Tsai HK, Chu CH, Hsieh WP, Chen YF et al (2015) The CrdRS two-component system in Helicobacter pylori responds to nitrosative stress. Mol Microbiol 97:1128–1141. doi:10.1111/mmi.13089
Liu ZQ, Zheng PY, Yang PC (2008) Efflux pump gene hefA of Helicobacter pylori plays an important role in multidrug resistance. World J Gastroenterol 14:5217–5222. doi:10.3748/wjg.14.5217
Trainor EA, Horton KE, Savage PB, Testerman TL, McGee DJ (2011) Role of the HefC efflux pump in Helicobacter pylori cholesterol-dependent resistance to ceragenins and bile salts. Infect Immun 79:88–97. doi:10.1128/IAI.00974-09
Huang YQ, Huang GR, Wu MH, Tang HY, Huang ZS, Zhou XH, Yu WQ, Su JW et al (2015) Inhibitory effects of emodin, baicalin, schizandrin and berberine on hefA gene: treatment of Helicobacter pylori-induced multidrug resistance. World J Gastroenterol 21:4225–4231. doi:10.3748/wjg.v21.i14.4225
Mehrabadi JF, Sirous M, Daryani NE, Eshraghi S, Akbari B, Shirazi MH (2011) Assessing the role of the RND efflux pump in metronidazole resistance of Helicobacter pylori by RT-PCR assay. J Infect Dev Ctries 5:88–93. doi:10.3855/jidc.1187
Yonezawa H, Osaki T, Hanawa T, Kurata S, Ochiai K, Kamiya S (2013) Impact of Helicobacter pylori biofilm formation on clarithromycin susceptibility and generation of resistance mutations. PLoS One 8:e73301. doi:10.1371/journal.pone.0073301
Weeks DL, Eskandari S, Scott DR, Sachs G (2000) A H+-gated urea channel: the link between Helicobacter pylori urease and gastric colonization. Science 287:482–485. doi:10.1126/science.287.5452.482
Zhang Y, Xiao M, Horiyama T, Zhang Y, Li X, Nishino K, Yan A (2011) The multidrug efflux pump MdtEF protects against nitrosative damage during the anaerobic respiration in Escherichia coli. J Biol Chem 286:26576–26584. doi:10.1074/jbc.M111.243261
Fetar H, Gilmour C, Klinoski R, Daigle DM, Dean CR, Poole K (2011) mexEF-oprN multidrug efflux operon of Pseudomonas aeruginosa: regulation by the MexT activator in response to nitrosative stress and chloramphenicol. Antimicrob Agents Chemother 55:508–514. doi:10.1128/AAC.00830-10
Srinivasan VB, Mondal A, Venkataramaiah M, Chauhan NK, Rajamohan G (2013) Role of OxyRKP, a novel LysR-family transcriptional regulator, in antimicrobial resistance and virulence in Klebsiella pneumoniae. Microbiology 159:1301–1314. doi:10.1099/mic.0.065052-0
Ma D, Cook DN, Alberti M, Pon NG, Nikaido H, Hearst JE (1995) Genes acrA and acrB encode a stress-induced efflux system of Escherichia coli. Mol Microbiol 16:45–55. doi:10.1111/j.1365-2958.1995.tb02390.x
Lin J, Cagliero C, Guo B, Barton YW, Maurel MC, Payot S, Zhang Q (2005) Bile salts modulate expression of the CmeABC multidrug efflux pump in Campylobacter jejuni. J Bacteriol 187:7417–7424. doi:10.1128/JB.187.21.7417-7424.2005
Zhang Z, Liu ZQ, Zheng PY, Tang FA, Yang PC (2010) Influence of efflux pump inhibitors on the multidrug resistance of Helicobacter pylori. World J Gastroenterol 16:1279–1284. doi:10.3748/wjg.v16.i10.1279
Falsafi T, Ehsani A, Niknam V (2009) The role of active efflux in antibiotic – resistance of clinical isolates of Helicobacter pylori. Indian J Med Microbiol 27:335–340. doi:10.4103/0255-0857.55452
Aeschlimann JR, Kaatz GW, Rybak MJ (1999) The effects of NorA inhibition on the activities of levofloxacin, ciprofloxacin and norfloxacin against two genetically related strains of Staphylococcus aureus in an in-vitro infection model. J Antimicrob Chemother 44:343–349. doi:10.1093/jac/44.3.343
Yang Y, Chua KL (2013) Assessment of the effect of efflux pump inhibitors on in vitro antimicrobial susceptibility of multidrug-resistant Acinetobacter baumannii. Int J Antimicrob Agents 42:283–284. doi:10.1016/j.ijantimicag.2013.05.011
De Milito A, Fais S (2005) Tumor acidity, chemoresistance and proton pump inhibitors. Future Oncol 1:779–786. doi:10.2217/14796694.1.6.779
Acknowledgments
The views expressed in this chapter do not necessarily reflect those of Xian-Zhi Li’s affiliation, Health Canada.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Li, J., Li, XZ. (2016). Antimicrobial Resistance and Drug Efflux Pumps in Helicobacter . In: Li, XZ., Elkins, C., Zgurskaya, H. (eds) Efflux-Mediated Antimicrobial Resistance in Bacteria. Adis, Cham. https://doi.org/10.1007/978-3-319-39658-3_19
Download citation
DOI: https://doi.org/10.1007/978-3-319-39658-3_19
Published:
Publisher Name: Adis, Cham
Print ISBN: 978-3-319-39656-9
Online ISBN: 978-3-319-39658-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)