Abstract
Pseudomonas aeruginosa is a major opportunistic pathogen that exhibits high-level intrinsic and acquired multiple antimicrobial resistance. In addition to the accumulation of individual drug-specific resistance mechanisms, such resistance phenotypes are attributed to the interplay between the polyspecific multidrug efflux pumps and the low outer membrane permeability, and this reflects evolution of P. aeruginosa in exposure to diverse hostile environments. A dozen drug efflux pumps, which belong to the resistance-nodulation-cell division (RND) superfamily, have been characterized in P. aeruginosa. Several RND pumps, as represented by MexAB-OprM and MexXY, play important roles in clinically relevant resistance, stress responses, and virulence. Regulation of these pumps is often under the control of local regulators (repressors or activators), global regulators, two-component regulatory systems, and modulators, whose mutations produce elevated antimicrobial resistance in many clinical isolates. This chapter provides an up-to-date overview of antimicrobial drug efflux pumps in P. aeruginosa with a focus on their substrates, regulation, inhibition, and clinical significance.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Pseudomonas aeruginosa
- Multidrug resistance
- Efflux pumps
- Outer membrane
- Efflux pump inhibitor
- RND
- MexAB-OprM
- MexXY
1 Introduction
Pseudomonas aeruginosa is a non-fermentative Gram-negative rod thriving in aquatic environments impacted by human activities. It is a notorious cause of severe healthcare-associated infections in immunocompromised patients, as well as pro-inflammatory chronic lung colonization in cystic fibrosis patients [1, 2]. This species is also a pathogen for animals such as dogs, cats, and bovines [3], whose virulence holds on production of multiple cell wall-associated or secreted factors (e.g., alginate, pili, lipopolysaccharide, toxins, and proteases) [4–6] and formation of biofilms [7, 8]. Because of the relatively high-level intrinsic resistance of this species to a wide range of structurally diverse antimicrobials, chemotherapy of P. aeruginosa infections relies on a limited number of antipseudomonal antimicrobials [1]. However, clinically significant resistance to these agents is commonly developed by clinical strains via various adaptive or acquired mechanisms [9, 10]. In the USA, it was estimated that 13 % of severe healthcare-associated P. aeruginosa infections are due to multidrug-resistant strains [11]. A more recent US study showed non-susceptible rates of up to 25 % toward major antipseudomonal drugs (except colistin) among 1,743 P. aeruginosa isolates [12]. In addition to the accumulation of individual drug-specific resistance mechanisms, multidrug resistance (MDR) may be achieved through the synergistic interplay between the low permeability outer membrane (OM) barrier and a number of multidrug efflux pumps belonging to the resistance-nodulation-cell division (RND) superfamily of transporters [13, 14]. Initially discovered in the early 1990s with the predominant role of the MexAB-OprM efflux system in both intrinsic and acquired resistance [15–18], multidrug transporters of P. aeruginosa have been further characterized for their roles in drug resistance and other functions [14, 19–21]. This chapter provides an up-to-date overview of efflux pump-mediated drug resistance in P. aeruginosa with an emphasis on the substrates, regulation, inhibition, and clinical relevance of these export systems. The roles of MDR efflux pumps beyond drug resistance such as in biofilm formation, stress responses, and pathogenicity of P. aeruginosa are described elsewhere (see Chaps. 25, 26, and 27).
2 Historical Perspectives on P. aeruginosa Chromosomal MDR Efflux Pumps
During the early studies on P. aeruginosa in the 1960s, MDR phenotypes characterized by a simultaneous resistance to aminoglycosides, chloramphenicol, penicillins, sulfonamides, and tetracyclines were observed [22, 23]. While at that time resistant P. aeruginosa strains were known to produce drug-inactivating enzymes (e.g., β-lactamases and aminoglycoside-modifying enzymes) [23, 24], these drug-specific enzymatic mechanisms offered no satisfactory explanation of resistance to structurally distinct antimicrobials. Because of the barrier function of the OM, most Gram-negative bacteria are less susceptible than Gram-positive bacteria to amphiphilic or bulky drug molecules [25, 26]. Breakthrough studies also specifically demonstrated that P. aeruginosa is a species with exceptionally low OM permeability [27–31], which is due to its major porin OprF mainly existing as closed channels [32, 33]. (Of note, P. aeruginosa and Escherichia coli have similar low permeable asymmetric lipid bilayer domains [34, 35].) Indeed, an antimicrobial-hypersusceptible P. aeruginosa mutant had OM lipopolysaccharide deficiency with easy drug access [29, 36, 37]. (This mutant was later found to be also deficient in drug efflux activity [16].) Moreover, drug uptake in P. aeruginosa may be further reduced in isolates resistant to aminoglycosides (e.g., streptomycin) or carbapenems (e.g., imipenem) by quantitative or qualitative changes in the lipopolysaccharide or porin (OprD) content of the OM [38, 39].
In the 1980s, the use of advanced broad-spectrum β-lactamase-stable β-lactams and fluoroquinolones was accompanied with increased isolation of multidrug-resistant isolates in vivo during drug administration [40–44]. These agents were also found to readily select MDR in vitro under laboratory conditions. While investigating the biochemical mechanisms of MDR or fluoroquinolone resistance, the OM protein profiles of P. aeruginosa isolates were assessed in numerous studies, which showed overproduction of ca. 50 kDa OM proteins that were associated with several gene loci named as nalB, nfxB, and nfxC [45–51]. One of these reports by Masuda and Ohya [51] designated the MDR-associated OM protein as OprM. Importantly, quinolone-resistant isolates also showed reduced uptake of ciprofloxacin [47] and active extrusion of ofloxacin [52]. Regardless of these studies, it became clear that the OM permeability barrier and periplasmic β-lactamase activity [53, 54] cannot fully explain MDR phenotypes (including β-lactam resistance in multidrug-resistant isolates/impermeability-type carbenicillin-resistant isolates) [55], which led to our initiative to investigate intrinsic and acquired MDR of P. aeruginosa [16, 17, 56]. In 1993, Poole et al. [15] reported the identification of the mexAB-oprK (i.e., mexAB-oprM) operon from P. aeruginosa which encodes a three-component efflux system involved in MDR. Together, these studies demonstrated a predominant role of drug efflux mechanism in intrinsic and acquired MDR (including β-lactam resistance) and expression of multiple drug efflux pumps in P. aeruginosa [15–18]. Subsequently, three MexAB-OprM homologues, MexCD-OprJ [57], MexEF-OprN [58], and MexXY (initially referred to as MexGH or AmrAB) [59–61], were also reported to be involved in P. aeruginosa MDR before the availability of the first whole genome annotated sequence for P. aeruginosa strain PAO1 [62]. All these Mex pumps belong to the RND superfamily of secondary active transporters [63], which typically require multiple components to form an energy-dependent functional extrusion complex across the entire cytoplasmic (inner) and outer membranes of Gram-negative bacteria [14].
3 Antimicrobial Drug Efflux Pumps and Their Clinical Significance in P. aeruginosa
P. aeruginosa genome sequences show the presence of a larger number of primary and secondary active transporters (TransportDB at http://www.membranetransport.org; accessed on February 15, 2016) [62, 64]. Both the widely studied strain PAO1 and more virulent strain UCBPP-PA14 contain 17 RND-type transporters. To date, 12 RND efflux pumps have been characterized for their substrate profiles as shown in Table 14.1. The rest of the RND members include homologues to protein export components such as SecD and SecF, but their role in drug resistance, if any, remains unknown. These RND pumps are generally encoded by operons and are each composed of three components that include a cytoplasmic membrane transporter (e.g., MexB), a cytoplasmic membrane-associated periplasmic adaptor protein (membrane fusion protein) (e.g., MexA), and an OM channel protein (e.g., OprM) (see Chaps. 1 and 5). These multicomponent pumps reflect the complex structures of P. aeruginosa cell envelopes and provide the structural and functional basis to directly extrude substrates out of the cell. In vitro transport activity of an assembled MexAB-OprM in proteoliposomes was recently demonstrated to show energy-dependent substrate translocation in a system mimicking Gram-negative dual-membrane envelope architecture [129]. Additionally, members of other transporter superfamilies or families have been identified, including the members of the major facilitator superfamily (MFS), the multidrug and toxic compound extrusion (MATE) family, the small multidrug resistance (SMR) family, and the ATP-binding cassette (ABC) superfamily (Table 14.1). For instance, of five proteins of the SMR family present in P. aeruginosa, one pump with the highest identity to the EmrE homologue of E. coli was shown to contribute to intrinsic resistance to aminoglycosides and dyes in low ionic strength media [120]. An ABC exporter was recently noted to be regulated by the PhoPQ two-component regulatory system and to contribute to tetracycline resistance [125]. However, the roles of the non-RND pumps in drug resistance remain largely unclear. Hence, we limit the descriptions below to RND efflux pumps.
3.1 MexAB-OprM
This efflux system, which has a constitutive though growth phase-dependent expression in wild-type strains, significantly contributes to intrinsic drug resistance of P. aeruginosa [130]. Inactivation of any component of MexAB-OprM renders the wild-type strains extremely hypersusceptible with ≥8-fold reduction in the values of the minimal inhibitory concentrations (MICs) for diverse antimicrobial agents (e.g., carbenicillin MIC was reduced by ≥128-fold) [18, 65–67]. MexAB-OprM overproduction contributes to the acquired MDR and is observed in clinical isolates of several mutant types including mexR (nalB), nalC, and nalD mutants [18, 51, 131–144]. An investigation of 12 multidrug-resistant MexAB-OprM-overproducing strains showed an equivalent distribution of mexR, nalC, and nalD mutants [136], which was supported by similar findings from independent studies [138, 145]. MexAB-OprM overproducers have also been selected in vitro after exposure to β-lactams, quinolones, chloramphenicol, macrolides, tetracyclines, as well as biocides and organic solvents [16, 45, 51, 146–148]. In vitro studies on reference strains have shown that any mutations inactivating genes mexR, nalC, or nalD or impairing the activity of their respective products, MexR, ArmR, or NalD (see Sect. 14.4 below), can result in overexpression (≥threefold) of mexAB-oprM with concomitant increase in resistance (2- to 16-fold MIC increases) to the pump substrates compared to baseline levels, with nalC mutants being in general twofold more susceptible than the nalB and nalD mutants [68–70, 82, 132, 149–151].
Similar to E. coli AcrAB-TolC (see Chap. 9), the MexAB-OprM efflux system shows the broadest substrate profile among the known multidrug efflux pumps of P. aeruginosa (Table 14.1) [15–18, 51, 65–67, 71, 72, 120, 152–156]. Those antimicrobial agents that have been confirmed as substrates are comprised of β-lactams (including β-lactamase inhibitors), chloramphenicol, quinolones/fluoroquinolones, macrolides, novobiocin, sulfonamides, trimethoprim, tetracyclines, cerulenin, pacidamycin, and thiolactomycin [16–18, 65, 73, 74, 157]. Moreover, the substrates also extend to nonantibiotics, such as dyes (acridine orange, acriflavine, crystal violet, and ethidium bromide), detergents, triclosan, organic solvents, tea tree oils, and quorum-sensing molecules/inhibitors [66, 72, 73, 75, 120, 148, 155, 158]. MexAB-OprM is also involved in reduced aminoglycoside susceptibility in low ionic strength medium [120]. Intriguingly, antipseudomonal activity of imipenem, a carbapenem β-lactam, appears not a substrate of the MexAB-OprM pump since MexAB-OprM overexpression has no impact on imipenem MIC in an OprD-deficient mutant vs. wild-type OprD strain [159]. (The OprD channel protein functions as a specific pathway for active basic amino acid uptake and also permits rapid penetration of imipenem [39], thus potentially masking the role of an efflux pump.) Nevertheless, other carbapenems such as doripenem, panipenem, and meropenem are substrates for MexAB-OprM [51, 160–162].
Reminiscent of E. coli TolC protein, OprM serves as a universal OM efflux protein and functions in multiple efflux systems (Table 14.1) [163–165]. OprM contributes to MDR, not only in conjunction with MexAB [18] but also independent of MexAB [166]. To date, OprM is known to work with other RND transporters (whose encoding operons often lack a linked gene for an OM component) including MexXY [59–61], MexJK [107], MexMN [109], MexVW [110], and TriABC [114], although other OM proteins can function with some of these transporters such as OpmH [114, 167] and OprA [168] (Table 14.1). Moreover, OprM can functionally replace the role of either OprJ of MexCD-OprJ or OprN of MexEF-OprN without affecting substrate profiles of these systems [163, 169].
Relevant to its clinical significance, MexAB-OprM when overproduced decreases the susceptibility of clinical isolates to antipseudomonal antimicrobials by a two- to eightfold in MIC values in comparison with the baseline levels in the absence of non-efflux resistance mechanisms such as enzymatic drug inactivation and drug target alterations [133, 142, 154, 170]. Based on the clinical susceptibility breakpoints from the Clinical and Laboratory Standards Institute (CLSI) [171], a maximal effect from the elevated MexAB-OprM efflux mechanism (eightfold MIC increase) would change strain categorization for a small number of antipseudomonal drugs such as aztreonam and ticarcillin (from drug susceptible [S] to intermediate [I] or resistant [R]) and meropenem, ciprofloxacin, and levofloxacin (from S to I). Another study showed that MexAB-OprM overproduction (via measuring mexA expression) was linked to median MIC values above the clinical resistance breakpoints (from the European Committee on Antimicrobial Susceptibility Testing [EUCAST]) for ciprofloxacin, cefepime, and meropenem [142]. Although further investigations are required to assess the therapeutic impact of MexAB-OprM in vivo [172], a recent study demonstrated that isolates with overproduction of either MexAB-OprM, MexCD-OprJ, or MexEF-OprN negatively affected antimicrobial efficacy in a Galleria mellonella in vivo infection model [173]. Higher drug dosages or antimicrobial-efflux pump inhibitor combinations are expected to be required in the treatment of infections associated with MexAB-OprM overproducers [173, 174]. Additionally, elevated MexAB-OprM expression also facilitates the emergence of other resistance mechanisms [147, 175]. Simultaneous expression of MexAB-OprM and other Mex pumps (e.g., MexXY or MexEF-OprN) have been reported, and this can produce additive effects in raising drug MIC levels as evident with fluoroquinolones [136, 141, 142, 176–180].
As a key mechanism responsible for high-level intrinsic resistance, the role of MexAB-OprM is also tightly linked to the low OM permeability barrier. Thus, the OM barrier and MexAB-OprM interplay to limit the access of antimicrobials to their cellular targets. The differential MIC values shown in Table 14.2 clearly demonstrate such synergistic interplay between the membrane barrier and the major efflux system in P. aeruginosa [181, 182]. Membrane disorganizers, such as chelating agent ethylenediaminetetraacetate (EDTA), potentiate antimicrobial activity of amphiphilic agents (which are expected to cross the OM through the lipidic domains), especially in the absence of MexAB-OprM (Table 14.2) [181]. This is also supported by an observation on the association of the deficiency in both MexAB-OprM and lipopolysaccharide with the hypersusceptible phenotype of strain Z61 [16, 37, 183]. Together, these data support a strategy to reverse antimicrobial resistance through the inhibition of drug efflux pumps and disruption of the OM barrier.
3.2 MexXY-OprM/MexXY-OprA
Encoded by a two-gene operon that lacks a gene for an OM protein, the MexXY system utilizes OprM to form a functional efflux pump in most P. aeruginosa strains [59, 60]. However, in the phylogenetically distinct isolate, PA7, and related strains, the mexXY genes are linked to a downstream gene encoding an OM protein dubbed OprA [168]. MexXY can function with either OprM or OprA in PA7 [168]. MexXY can also operate with another OM protein, OpmB, under still unclear conditions [83]. Inducibly expressed in P. aeruginosa, MexXY pump provides intrinsic resistance to aminoglycosides, a class of highly hydrophilic antimicrobial drugs, and to other agents that can, at subinhibitory levels, induce MexXY expression [59, 184]. Intriguingly, all of these inducers target ribosomes and this feature is related to regulation of MexXY expression (see Sect. 14.4 below) [184, 185]. Inactivation of MeXY in wild-type strains leads to a four- to eightfold reduction in MIC values of aminoglycosides (e.g., amikacin, gentamicin, isepamicin, netilmicin, and tobramycin), erythromycin, and tetracycline [59]. Aminoglycoside resistance in so-called “impermeability-type” clinical isolates is caused by MexXY overproduction [61]. Amino acid residues important for aminoglycoside recognition in MexY have been identified recently [186]. Elevated MexXY expression confers a 2- to 16-fold higher resistance to its pump substrates. When overexpressed from plasmid vectors in P. aeruginosa or E. coli, MexXY also mediates resistance to fluoroquinolones [59, 60]. Interestingly, induction of MexXY expression by spectinomycin is correlated with an increased susceptibility to polymyxins (up to a fourfold MIC reduction), due to the reduced expression of polymyxin resistance-promoting lipopolysaccharide modification locus [187].
MexXY-overproducing mutants can be easily selected in vitro and in vivo in the presence of substrate antimicrobial agents including peptide deformylase inhibitors [82, 84–86, 142, 188] [85]. Indeed, MexXY overproducers are highly prevalent in clinical isolates from cystic fibrosis [61, 189–194] and non-cystic fibrosis patients worldwide [137, 140, 178, 179, 195–205]. Abundance of reactive oxygen species in the cystic fibrosis lung environment may offer an explanation for such high rates of resistance [206]. Consistent with this, prolonged exposure of P. aeruginosa to hydrogen peroxide was shown to facilitate the emergence of MexXY overproducers in vitro [207].
Based on the locations of mutations, MexXY-overproducing mutants can be divided into three types: agrZ, agrW1, and agrW2 mutants. With agrZ mutants, various mutations occur in gene mexZ that encodes a repressor of MexXY [84, 138, 168, 188–191, 193, 197, 200, 205, 208, 209]. With agrW1 mutants, mutations affect ribosomal proteins such as L1 [61], L25 [210], L21, and L27 [211] or components of the methionyl-tRNAfmet formylation bypass [85]. Actually, lines of evidence suggest that whatever its origin (e.g., mutations, ribosome targeting drugs) impairment of protein synthesis is a stimulus for MexXY expression. For agrW2 mutants, mutational activation of the two-component regulatory system ParRS results in constitutive expression of MexXY [86, 203]. The presence of these three types of mutants in clinical isolates was confirmed among non-cystic fibrosis isolates that exhibited a moderate, nonenzymatic resistance to aminoglycosides [205]. However, the agrZ type predominates over the two others in cystic fibrosis isolates [189–191].
Isolates with overexpression of MexXY (via measuring mexX expression) also showed median MIC values higher than the EUCAST resistance breakpoints for amikacin, ciprofloxacin, cefepime, and meropenem [142]. However, only a few studies have assessed the potential role of MexXY in clinical therapeutic outcomes. In a rabbit experimental model of pneumonia treated with intravenous administration of tobramycin, a modest influence from MexXY overexpression on animal survival and post-treatment bacterial loads was observed [212]. Elevated efflux activity due to mexXY derepression is likely one of the multiple means P. aeruginosa can accumulate gradually to increase its resistance toward potent antimicrobials [205, 210]. As mentioned earlier, simultaneous overexpression of multiple efflux pumps (e.g., MexAB-OprM, MexXY, and MexEF-OprN) in conjunction with other resistance mechanisms is common in hospital strains [16, 136, 140, 178, 179, 213].
3.3 MexCD-OprJ
This efflux system is apparently quiescent in wild-type strains under normal laboratory growth conditions, and thus, chromosomal disruption of the mexCD-oprJ operon does not alter antimicrobial susceptibility of wild-type cells [57, 158]. MexCD-OprJ expression is inducible by various membrane-damaging nonantibiotic toxicants, including acriflavine, ethidium bromide, rhodamine 6G, chlorhexidine, and tetraphenylphosphonium, which are also the substrates of MexCD-OprJ [92, 93]. Exposure of P. aeruginosa to waste water was found to lead to MexCD-OprJ overexpression [214]. Mutation-mediated overexpression of this operon in nfxB-type mutants significantly contributes to resistance to fourth-generation cephalosporins (cefepime and cefpirome), quinolones/fluoroquinolones, chloramphenicol, cerulenin, pacidamycin, tetracycline, and novel inhibitors of lipid A synthesis such as CHIR-90 [57, 67, 74, 76, 94, 215]. Similar to MexAB-OprM, the substrates for MexCD-OprJ also include other cytotoxic compounds, such as acriflavine, ethidium bromide, quaternary ammonium compounds, rhodamine 6G, triclosan, and organic solvents [66, 92, 93, 95]. Based on the variability in drug resistance levels, MexCD-OprJ-overproducing nfxB mutants can be grouped into two types [215]. Type A mutants are resistant to erythromycin, ofloxacin, and zwitterionic cephems (cefclidin, cefozopran, cefoselis, and cefpirome), while type B mutants are resistant not only to these aforementioned agents but also to chloramphenicol and tetracycline [215]. Type B mutants are, however, four- to eightfold more susceptible to many conventional penicillins (e.g., carbenicillin), atypical β-lactams (aztreonam and moxalactam), carbapenems (biapenem and imipenem), and aminoglycosides (gentamicin and kanamycin) than the wild-type PAO1 strain [215]. This hypersusceptibility to conventional β-lactams and aminoglycosides [215, 216] is possibly attributable to the downregulation of MexAB-OprM [183, 216], MexXY [94], and the AmpC β-lactamase [217] in the MexCD-OprJ-overproducing mutants, although other mechanism(s) may exist [218].
MexCD-OprJ overproducers are associated with fluoroquinolone resistance, although fluoroquinolone-resistant isolates may also overexpress other efflux pumps (e.g., MexAB-OprM, MexXY, or MexEF-OprN) and/or carry quinolone-target mutations [94, 178, 219]. Indeed, overexpression of MexCD-OprJ, as with that of MexAB-OprM or MexXY, was reported to occur in a large proportion of fluoroquinolone- and/or carbapenem-resistant clinical isolates [220, 221]. (However, this MexCD-OprJ overexpression issue remains controversial and requires further investigations since nfxB-type MexCD-OprJ-overexpressing mutants are strongly deficient in fitness. The reverse-transcription-qPCR thresholds used to arbitrarily define mexCD-oprJ overexpression may have a strong impact on the mutant rates found in the clinical setting such as non-cystic fibrosis patients.) An in vitro study suggested newer fluoroquinolone agents were in favor of the selection of MexCD-OprJ producers [147]. Supporting this notion, elevated MexCD-OprJ expression is linked to levofloxacin resistance in P. aeruginosa isolates from urinary tract infections [222]. A patient treated by two substrates of MexCD-OprJ, ciprofloxacin and cefepime, produced over the treatment period nfxB mutants which had a change of bacteria from S to I or R as regards their susceptibility to fluoroquinolones based on the CLSI resistance breakpoints [223]. nfxB mutations causing MexCD-OprJ overproduction can be the first-step mutations in addition to further mutations in other resistance determinants as evident by a selection with a novel bacterial type II topoisomerase inhibitors [96, 224]. Here, it is worth mentioning that the genotypic alterations in MexCD-OprJ overproducers may not correlate with the phenotype [213, 225], likely attributed at least partly to global changes in the physiology and metabolism caused by nfxB mutations [111, 226]. MexCD-OprJ overexpression produces an increased susceptibility to complement-mediated killing and consequently results in reduced virulence [227]. In this regard, a recent study showed that quaternary compounds were substrates of MexCD-OprJ but were unable to select resistant mutants including MexCD-OprJ-overproducing mutants with these biocides [95]. Nevertheless, the negative resistance selection results warrant further investigation. Another study revealed that P. aeruginosa adapted to 2-phenoxyethanol displayed reduced susceptibility to different biocides but increased susceptibility to several antipseudomonal antibiotics including amikacin, tobramycin, ceftazidime, and ciprofloxacin [228]. Another biocide, triclosan, can select MexCD-OprJ overproducers [229, 230]. High-level resistance to this biocide was speculated to play a role in antibiotic resistance in an epidemic isolate [231].
3.4 MexEF-OprN
This efflux system is also not well expressed in wild-type cells of P. aeruginosa, and thus its inactivation leads to no or little change in antimicrobial susceptibility [58, 183]. MexEF-OprN is highly expressed in nfxC mutants to confer an increased resistance to chloramphenicol, quinolones/fluoroquinolones, tetracycline, and trimethoprim [58, 169, 178]. Decrease in susceptibility to carbapenems, a phenotype characteristic of some nfxC mutants [58], is partly attributable to the downregulation of OprD expression [232, 233]. nfxC mutants are readily selected by chloramphenicol and fluoroquinolones, but not by carbapenems [147, 234, 235]. They have been observed in clinical isolates from cystic fibrosis and other patients [221, 225], but their prevalence varies from one study to another. While many studies apparently suggest low frequencies or even no detection of nfxC mutants among clinical isolates [178, 195, 199, 204, 233], recent studies revealed more prevalence of MexEF-OprF overproducers [221, 236]; for example, about 30 % of 62 isolates (mostly obtained from intensive care unit patients and with reduced carbapenem susceptibility) had an increased production of MexEF-OprN (from >4- to 19-fold in mexF mRNA transcripts in comparing with a wild-type reference isolate) [236]. MexEF-OprN overproducers can likely also be among the first-step mutants, which may further acquire higher resistance [178]. Unexpectedly, tobramycin-hypersusceptible mutants (eightfold MIC reduction) were associated with insertional inactivation of mexF that generated the aberrant hybrid MexF-alkaline phosphatase proteins. These hybrid proteins were interpreted to cause cytoplasmic membrane stress for gain-of-function changes with increased aminoglycoside susceptibility [87].
3.5 MexJK-OprM/OpmH
This efflux system is expressed at low levels in wild-type cells [107, 112]. Despite the lack of a gene for OM protein in its encoding operon, MexJK requires an OM channel protein for drug efflux. While OprM is involved in the extrusion by MexJK of ciprofloxacin, erythromycin, and tetracycline, MexJK is dependent on another OM protein, OpmH, for providing resistance to triclosan [107, 167]. Clinical significance of this pump remains unknown. Nevertheless, MexJK overproduction was observed in two MexXY-hyperexpressing cefepime-resistant isolates [197] as well as in a MexXY-/MexVW-overproducing isolate [179].
3.6 MexGHI-OpmD
Encoded by a four-gene operon, PA4205-PA4208 genes [62], MexGHI-OpmD is operative in wild-type cells and mediates intrinsic resistance to vanadium [237]. While MexH and MexI are, respectively, the cytoplasmic membrane exporter and the accessory membrane fusion protein, MexG is a protein with unknown function. This system is involved in PQS (pseudomonas quinolone signal) homeostasis and is associated with quorum sensing in P. aeruginosa [112]. Its inactivation results in reduced production of several virulence factors, thus linking antimicrobial susceptibility and pathogenicity [104, 237]. Intriguingly, vanadium hypersusceptibility of the mexGHI-opmD null mutants is accompanied by increased resistance to netilmicin, tetracycline, and ticarcillin-clavulanic acid [237], likely due to the compensating overexpression of other MDR pumps [183].
3.7 Other RND Efflux Pumps
Several additional RND efflux systems, when expressed from vectors, were able to confer resistance in P. aeruginosa or E. coli host deficient in major RND pumps (Table 14.1): MexMN-OprM for resistance to fluoroquinolones and macrolides [109]; MexPQ-OpmE for resistance to amphenicols [109]; MexVW-OprM for resistance to chloramphenicol, fluoroquinolones, macrolides, and tetracycline [110]; MuxABC-OpmB for resistance to aztreonam, macrolides, novobiocin, and tetracyclines [112, 113]; and TriABC-OpmH for triclosan resistance [114]. While the MuxABC-OpmB system possesses two RND components, MuxBC [114], TriABC-OpmH requires two periplasmic accessory membrane fusion proteins, TriA and TriB [114], which play different roles in the assembly and function of TriABC pump [238]. MuxABC-OpmB is expressed in wild-type strains, and interestingly, its inactivation results in elevated β-lactamase production with increased β-lactam resistance [112]. MuxABC-OpmB is one of the RND efflux systems that affect the development of colistin-tolerant subpopulations in P. aeruginosa biofilms [111]. Co-overexpression of MexVW and MexXY was also reported [179]. Finally, one RND efflux system, CzcCBA (also called CzrCBA), contributes to resistance to cadmium, cobalt, and zinc salts [115, 116]. Phenotypically, CzcCBA-mediated heavy metal resistance is also linked to imipenem resistance as a result of the downregulated OprD expression and the elevated expression of quorum-sensing autoinducer molecules, due to a shared two-component regulatory system, CzcRS [117, 239]. We have observed one imipenem-insusceptible isolate with overexpressed CzcCBA and reduced OprD production from an intensive care unit [203].
4 Regulation of P. aeruginosa RND Efflux Pumps
Regulation of P. aeruginosa RND efflux pumps has been studied intensively over the last two decades, which shows the complexity of RND pump regulatory network with involvement of various transcriptional regulators and modulators at multiple levels [14]. Changes in natural or host environments of P. aeruginosa such as antimicrobial exposures and nutrient, oxidative, and nitrosative stresses can affect the expression of drug efflux pumps and subsequently contribute to phenotypic adaptations such as the development of MDR [20, 240]. The expressional status of one drug efflux pump may also be linked to the production of other pumps through complex regulatory networks [94, 183]. Together, different regulatory pathways interconnect antimicrobial susceptibility, stress responses, pathogenicity, and even biofilm formation.
4.1 MexAB-OprM
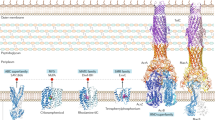
Although constitutively expressed, the mexAB-oprM operon is subject to a complex and finely tuned regulation. Multiple gene products MexR, NalD, ArmR (via NalC), and a two-component regulatory system, RocS1/S2-RocA2, are involved in the regulation of mexAB-oprM expression (Fig. 14.1).
Regulation of the expression of four RND multidrug efflux systems of P. aeruginosa. These pumps are controlled by a local regulator (mostly by a repressor [MexR, NfxB, EsrC, or MexZ] or by an activator [MexT]). Positive and negative regulations of the relevant gene transcriptions are, respectively, denoted by red and green lines. Local repressors are controlled by anti-repressor proteins (ArmR and ArmZ) and can also bind to ligands (e.g., antimicrobial agents) or be induced under various conditions (nitrosative, oxidative, or cell envelope stress). Mutational changes can also lead to inactivation of these regulators. See text for details
MexR, a MarR-family repressor, is encoded by a self-regulated gene (mexR/nalB) that is divergently transcribed upstream of the mexAB-oprM operon [68]. Binding of MexR as a dimer to the intergenic divergent overlapping promoters of mexR and mexAB-oprM produces a balanced transcription of both mexR and mexAB-oprM, which provides P. aeruginosa with a protective baseline level of wide-spectrum efflux activity [149, 241]. Mutations in mexR (nalB mutants) are typically associated with MexAB-OprM overexpression [68, 135, 144, 148]. The crystal structure of MexR suggests an effector-induced conformational change for inhibiting DNA binding [242]. Based on in vitro data that MexR dimerization through the formation of intermonomer disulfide bonds between two redox-active cysteines prevents MexR from interacting with its cognate DNA binding sites, redox modulation of MexR was proposed to occur in vivo under stressful conditions such as the presence of oxidative agents (e.g., hydrogen peroxide) or antibiotics (meropenem and nalidixic acid) [243, 244]. However, several agents including hydrogen peroxide, colistin, and tobramycin apparently do not significantly induce mexAB-oprM transcription [245–248].
The expression of mexAB-oprM is positively modulated by ArmR, a 53-residue peptide, whose encoding gene is located in a two-gene operon, PA3720-armR [69]. By an allosteric polypeptide-protein interaction of high affinity, ArmR function as an anti-repressor to sequester MexR, consequently reducing the MexR repressor activity [249, 250]. Unless mutations inactivate gene nalC (PA3721) which encodes a TetR-family repressor to strongly downregulate the adjacent PA3720-armR operon, basal amounts of ArmR do not affect MexAB-OprM production in wild-type cells [69]. In fact, nalC mutants only show modestly elevated expression of mexAB-oprM, and disruption of ArmR in these nalC mutants reduces MexAB-OprM expression to wild-type levels and compromises MDR [69]. By reversible, non-covalent binding to NalC, various chlorinated phenols including pentachlorophenol at relatively high levels can induce the expression of operons PA3720-armR and mexAB-oprM [77, 251]. Although pentachlorophenol affects expression of armR, MexAB-OprM induction by pentachlorophenol can also be ArmR-independent, yet MexR-dependent [252]. This suggests that in vivo generated catabolite effectors may mimic more specific phenolic antimicrobial compounds than pentachlorophenol that P. aeruginosa encounters in its natural habitat [252].
NalD, a TetR-family repressor, is another regulator of mexAB-oprM that binds to the proximal promoter upstream of the efflux operon [70, 78], resulting in mexAB-oprM being expressed essentially from its distal promoter [78]. A recent study demonstrated direct binding of novobiocin to NalD to result in dissociation of NalD from the promoter with subsequent derepression of mexAB-oprM expression [253]. The combinational mutations in mexR, nalC, and nalD have been observed in clinical isolates including epidemic strains which are MexAB-OprM overproducers [136, 138, 145, 254].
The mexAB-oprM expression is also growth-phase regulated and reaches a maximum level at the onset of the stationary phase, independently of MexR and of LasR, a transcriptional regulator controlling the production of quorum-sensing cell-to-cell signal N-3-oxo-dodecanoyl-L-homoserine lactone (3-oxo-C12-HSL) [130, 255, 256]. P. aeruginosa has several intertwined quorum-sensing systems, such as Las, Pqs, and Rhl, that control virulence gene expression [257, 258]. The Rhl quorum-sensing signal N-butanoyl-L-homoserine lactone (C4-HSL) can induce mexAB-oprM expression [152, 259], possibly via its role in the growth-phase-dependent regulation of MexAB-OprM [72, 79] with MexR being not required in this control [260]. This regulation of MexAB-OprM can be canceled by MexT, the activator of mexEF-oprN operon in nfxC mutants [79]. Additionally, AmpR, a LysR-family global transcriptional regulator implicated in AmpC β-lactamase expression and other genes of the AmpR regulon, was reported to potentially derepress the MexAB-OprM expression by negatively modulating the mexR expression [261]. Several 14- and 15-membered macrolides including azithromycin at subinhibitory levels can repress the cell density-dependent expression of MexAB-OprM in a MexR-dependent manner through yet-unidentified mechanism [262].
MexAB-OprM expression in biofilm cells is further regulated by additional mechanisms. MexAB-OprM pump contributes to tolerance to colistin in a metabolically active subpopulation of biofilm cells [263]. The histidine kinase sensors RocS1 and RocS2 act through their cognate response regulator RocA2 to repress mexAB-oprM expression in biofilms [80]. BrlR, a biofilm-specific MerR-family regulator, functions as an activator and is required to sustain expression of mexAB-oprM (and mexEF-oprN) during an early stage of biofilm development through its binding to the promoter regions of the two operons [81, 264]. Intriguingly, BrlR is responsive to the secondary messenger, cyclic dinucleotide c-di-GMP, which is required for BrlR production and function [265]. During the early developmental stage of biofilms, the two-component hybrid histidine kinase SagS is also produced, and it positively affects the production of c-di-GMP and BrlR, which contribute to increased expression of MexAB-OprM and MexEF-OprN and high-level biofilm-specific resistance to antimicrobial agents [266, 267]. Therefore, mexAB-oprM expression in biofilms is likely affected by at least two distinct signal transducing systems (i.e., RocS1/RocS2-RocA2 and SagS-BrlR). However, contribution of the MexAB-OprM pump to antimicrobial resistance in P. aeruginosa biofilms remains controversial and might depend upon the experimental conditions used or stage of biofilm development [268].
4.2 MexXY
The MexXY efflux system is also subject to a multi-level regulation (Fig. 14.1). MexZ is the local transcriptional repressor of the mexXY or mexXY-oprA operon [59, 168]. Binding of dimerized MexZ to the overlapping promoters of mexXY and mexZ allows very low baseline production of MexXY [208, 269, 270]. Unlike many other TetR-family regulators [271], MexZ’s DNA binding is not relieved by antimicrobials through a direct ligand-regulator interaction but seemingly via indirect protein-protein sequestration, which is dependent on the anti-repressor of mexZ, ArmZ (PA5471) [270, 272, 273]. Induction of mexXY expression occurs through an ArmZ-dependent manner [88, 272] in response to the exposure of P. aeruginosa to a number of ribosome-targeting antimicrobials (such as aminoglycosides, chloramphenicol, macrolides, and tetracyclines) [185] or oxidative stress conditions [207]. Expression of armZ itself is induced by ribosome-targeting agents through a transcriptional attenuation, ribosome stalling mechanism that involves a short 13-amino acid leader peptide, PA5471.1 [88]. Ribosome stalling at this leader peptide mRNA yields armZ transcription to subsequently upregulate mexXY expression [88]. In this regard, another protein, SuhB, was found to interact with the ribosome [89]. The suhB gene was first identified as an entragenic suppressor of a component of the type II secretion system in E. coli [274] and was also revealed to be involved as a regulator of multiple virulence genes implying types III and VI secretion systems and biofilm formation in P. aeruginosa [275]. A suhB mutant exhibited higher level of PA5471.1 mRNA with elevated mexXY expression, which was consistent with the reduced susceptibility of the suhB mutant to aminoglycosides [89]. Additionally, in pan-aminoglycoside-resistant mutants, reduced expression of the rplU-rpmA operon is attributable to mutations in the promoter region of the operon, which encodes ribosomal proteins L21 and L27 [211]. This change is also linked to an ArmZ-dependent MexXY overproduction. Hence, the ribosome-perturbing mutations act in a way reminiscent of mexXY induction by ribosome-targeting antimicrobials [89, 211]. However, mexXY expression still remains inducible to some extent in mexZ and mexZ-armZ null mutants [185, 273], suggesting the presence of additional contributors in induction of mexXY.
Regulation of MexXY is also mediated by the two-component regulatory system ParRS. Either mutations or exposure to subinhibitory levels of polycationic compounds such as polymyxins can activate ParRS [276], which upregulates both mexXY and lipopolysaccharide modification operon arnBCADTEF-ugd and downregulates the oprD expression, yielding an MDR phenotype by activation of three distinct mechanisms (efflux, lipopolysaccharide modification, and OprD reduction) [86, 205]. Analysis of tobramycin-hypersusceptible mutants revealed mutations in more than a dozen genes that included mexXY, oprM, and the two-component regulatory system genes amgRS with amgRS mutants showing 8- to 16-fold reduction of tobramycin MIC values [87]. AmgRS was shown to be required for tobramycin induction of several genes, including three genes, htpX, PA5528, and yccA (which encode, respectively, a cytoplasmic membrane-associated protease, a modulator of the FtsH protease, and a protease-associated factor) involved in positively stimulating mexXY expression [87, 90, 91]. A recent study showed AmgRS-dependent potentiation of the activity of 4,5-linked aminoglycosides (such as neomycin, paromomycin, and ribostamycin) by rifampicin [277]. The latter targets AmgRS and repressed expression of AmgRS-dependent genes including htpX, yccA, and mexXY. Rifampicin also potentiated the activity of two 4,6-linked aminoglycosides such as amikacin and gentamicin in two clinical isolates [277].
Inactivation of either gene PA2572 (for a non-canonical response regulator) or PA2573 (for a probable methyl-accepting chemotaxis protein) strongly increased mexXY expression by >10-fold when measuring mexX or mexY transcripts, and this explains the reduced susceptibility of these mutants to two aminoglycosides, amikacin and tobramycin (10- to 40-fold MIC increase) [278]. However, the detailed cascade affecting mexXY expression remains unknown.
4.3 MexCD-OprJ
The MexCD-OprJ pump is negatively controlled by NfxB and EsrC repressors, whose encoded genes are located, respectively, upstream and downstream of the mexCD-oprJ operon (Fig. 14.1) [57, 97, 279]. NfxB acts as a multimer (dimer of dimers) with C-termini required for multimerization and N-termini in DNA binding [97, 280]. nfxB mutations can occur over the entire nfxB gene with the deletion-generated frameshifts frequently observed in clinical strains [94, 281]. Inactivation of DNA oxidative repair system also increases frequencies of nfxB mutations [282]. Intriguingly, VqsM, an AraC-family master transcriptional regulator involved in the regulation of virulence factors and quorum-sensing compounds, can bind to the promoter of nfxB to likely increase nfxB expression, although vqsM mutants derived from wild-type PAO1 strain show higher resistance to kanamycin and tetracycline (16- and 32-fold MIC increase, respectively) with no changes in susceptibility to ceftazidime, ciprofloxacin, polymyxin B, and tobramycin [98]. Given the low-level expression of mexCD-oprJ in wild-type cells, it would be interesting to know whether VqsM influences MexCD-OprJ production in nfxB mutants.
Another regulator of MexCD-OprJ, EsrC, is functionally dependent on NfxB for repressing mexCD-oprJ expression when cells are under envelope stress [279]. Expression of mexCD-oprJ is induced by a number of biocides (e.g., benzalkonium chloride and chlorhexidine), dyes (ethidium bromide), and other membrane-damaging agents (detergents, solvents, polymyxin B, and antimicrobial peptides including human host defense peptide LL-37) [92, 93, 283]. Exposure to chlorhexidine diacetate produces a significant transcriptomic response [284]. These membrane-damaging agents apparently generate membrane lipid derivatives to stimulate the membrane-associated Muc proteins and to eventually activate the stress response sigma factor, AlgU, for upregulating MexCD-OprJ expression. nfxB mutation-related mexCD-oprJ hyperexpression is also dependent on AlgU [93]. Finally, disruption of the aforementioned gene PA2572 that codes for a putative response regulator was also found to modestly increase mexCD-oprJ activity (a fourfold increase in mexC transcripts) [278].
4.4 MexEF-OprN
Expression of MexEF-OprN is also controlled by several regulators (Fig. 14.1). MexT, a LysR-family global regulator, controls expression of multiple genes in nfxC mutants including mexEF-oprN, oprD, and genes for virulence factors [232, 285–288]. Inactive and active forms of MexT exist, respectively, in wild-type strains and nfxC mutants. One gene of the MexT regulon, mexS (encoding an oxidoreductase of unknown function [285]), because of its alteration in nfxC mutants, promotes mexEF-oprN expression with concomitant development of MDR [289]. This induction occurs as a result of MexS-MexT interplays through presumed intracellular accumulation of toxic metabolites recognized by MexT as co-inducers [289]. Indeed, exposure of P. aeruginosa to nitrosative stressors such as S-nitrosoglutathione activates mexEF-oprN transcription via MexT [290]. Disulfide stress response and the type III secretion system are affected by MexS-MexT interaction [100, 291], thus providing another example for the linked regulation among drug efflux pumps, redox stress response, and virulence factor production. But, MexS-independent mexEF-oprN overexpression has also been observed [292]. Similar upregulation of mexS and mexEF-oprN was noted when P. aeruginosa was exposed to human airway epithelial cells releasing unknown efflux-inducing signals [293]. Expression of mexEF-oprN was also found to be abolished by the downregulation of MexS through mutations in the ParRS two-component regulatory system [101]. The latter is also involved in the regulation of MexXY, OprD, and lipopolysaccharide modifications [86, 276]. A recent study showed single amino acid substitutions in MexS in a good proportion of clinical nfxC mutants, which had an association with moderate effects on drug resistance and virulence factor production, supporting the notion of in vivo selection of partially defective mexS mutants retaining some degree of pathogenicity [294]. Additionally, the global regulator MvaT influences expression of hundreds of genes including mexEF-oprN and others involved in biofilm formation, quorum sensing, and virulence [295–297]. Independent of mexT or mexS, inactivation of mvaT results in MexEF-OprN hyperexpression and marginal OprD reduction (associated with increased susceptibility to imipenem) [102], suggesting the complexity in MexEF-OprN expression. Consistently, despite the observed mutations in mexS, mexT, and mvaT in MexEF-OprN-overproducing clinical isolates [178], a good proportion of nfxC mutants do not show any mutations in these genes [294], revealing involvement of additional regulatory mechanisms. In this regard, AmpC β-lactamase regulator AmpR affects expression of >500 genes, and its inactivation increases MexEF-OprN production with an MDR phenotype [261]. The abovementioned BrlR also positively affects MexEF-OprN expression in biofilm cells [81]. The reduced virulence of nfxC mutants has been attributed to MexEF-OprN-dependent extrusion of 4-hydroxy-2-heptylquinoline [103] and/or kynurenine [298, 299], to two precursors of quorum-sensing molecule PQS, and to MexT-dependent downregulation of type III secretion system and pyocyanin production [286, 291].
4.5 Other RND Pumps
The mexGHI-ompD operon is positively regulated by SoxR transcriptional regulator as part of an oxidative stress response to the presence of methyl viologen [105], the phenazine pyocyanin (a heterocyclic, redox-active agent) [106], and oxidative compounds such as 7-hydroxyindole involved in anti-virulence [300]. A human host defense peptide, LL-37, is also able to induce expression of MexGHI-OmpD [283]. Expression of MexJK is negatively regulated by MexL repressor, which is encoded by a gene transcribed divergently from the adjacent mexJK operon [107]. CzcCBA metal exporter is upregulated by at least two two-component regulatory systems, CzcRS (CzrRS) [115] and CopRS [117]. Subinhibitory concentrations of zinc or copper salts can induce expression of czcCBA, czcRS, and copRS. CzcRS and CopRS are also involved in the downregulation of OprD expression with concomitant resistance to carbapenems [116]. CzcR further affects various genes involved in virulence including gene expression of quorum-sensing 3-oxo-C12-HSL and C4-HSL autoinducers [239]. A mvaT mutant also shows a decreased expression of the two-component regulator gene (PA2570) located immediately downstream of the czcABC efflux operon [296].
5 Overcoming P. aeruginosa Drug Efflux Activities
The characterization of RND pumps shows the scientific challenge of finding antimicrobial drugs that can bypass the efflux mechanisms (see Chap. 28). Numerous newer antimicrobial agents are substrates of RND pumps, such as ceftobiprole, doripenem, and tigecycline [14, 82, 301, 302]. In fact, recent success in clinical use of a new β-lactam-β-lactamase inhibitor combination product, ceftazidime-avibactam, has faced an unexpected challenge from archived P. aeruginosa isolates, i.e., drug efflux and membrane permeability barrier to reduce activity of this product [303]. To combat the efflux impact, rational drug design can be exploited to minimize or to avoid efflux. This approach is becoming increasingly feasible due to the in-depth structural and biochemical understanding of RND efflux pumps [129, 165, 304–309].
The following examples show that despite the multi-specificity and multiplicity of RND transporters in P. aeruginosa, novel antimicrobials can be developed to escape efflux mechanism. The activity of a novel parenteral aminopyrazolium cephalosporin, FR264205, is unlikely affected by the expression of MexAB-OprM, MexCD-OprJ, MexEF-OprN, or MexXY [310, 311]. A methylcarbapenem, tomopenem, displays broad-spectrum activity against Gram-positive and Gram-negative pathogens including P. aeruginosa, and this is at most minimally impacted by overexpression of Mex pumps [312, 313]. The latter may, however, be partly attributable to the high affinity of tomopenem to the major lethal targets, penicillin-binding proteins 2 and 3 [314]. Antimicrobial polypeptides generally do not appear to be impacted by efflux systems including RND pumps [315]. Polymyxins are often active against multidrug-resistant P. aeruginosa despite reports suggesting that MexAB-OprM, MexCD-OprJ, and MuxABC-OpmB pumps contribute to nonspecific adaptive resistance to polymyxins in biofilms [111, 263]. In comparing with several fluoroquinolones such as ciprofloxacin, activity of clinafloxacin is less compromised by Mex pumps [71]. Overall, multiple factors such as efflux pump effect, affinity to the drug targets, and membrane permeation contribute to collectively the antipseudomonal activity of drug molecules.
The role of clinically relevant efflux pumps also highlights a needed strategy to look for agents that can function as efflux pump inhibitors either to restore susceptibility of multidrug-resistant strains or to prevent the emergence of mutation-driven resistance mechanisms, when combined with conventional antibiotics. Since the discovery of RND pumps, efforts have also been undertaken to identify pump inhibitors and P. aeruginosa RND pumps have particularly been a major target (see Chaps. 29 and 30) [240, 316]. Phenylalanine-arginine β-naphthylamide is one of the earliest efflux pump inhibitors identified and is accepted as a typical efflux pump inhibitor of RND pumps [316, 317]. It potentiates in vitro activity of a number of antipseudomonal agents against multidrug-resistant strains [316, 318], but its clinical applications have been challenged by various factors including unfavorable pharmacokinetics and toxicity [14]. Compounds of synthetic pyridopyrimidine series have also been investigated for MexAB-OprM-specific inhibition, and these include a potential preclinical candidate, quaternary analogue D13-9001 [319, 320], which potentiates the activity of aztreonam and levofloxacin and reduces in vitro invasiveness of P. aeruginosa into mammalian cells [319, 321]. Molecular modes of action of these inhibitors including their interaction with RND pumps were reviewed recently [14, 320, 322]. Similar to the effect from genetic inactivation of PvdRT-OpmQ efflux pump [126, 127], reserpine was found to inhibit this exporter to synergize both in vitro and in vivo activities of a siderophore-monobactam conjugate [128].
Certain existing drug agents have also been assessed for their potential to be used as efflux pump inhibitors such as sertraline and trimethoprim [173]. (However, further studies are required since only wild-type strains, not efflux-upregulated mutants, were affected.) Various natural extracts have been assessed for combinational use with conventional antibiotics against P. aeruginosa [240, 323–326]. The compound 3,4-dibromopyrrole-2,5-dione isolated from a Pseudoalteromonas spp. was shown to potentiate activity of multiple antimicrobials against Mex pump overproducers [327]. However, more investigations are needed to rule out any non-efflux inhibitory effects of these compounds on cell growth. Transcriptional inhibition of the RND pumps has been shown to reduce efflux-mediated resistance although clinical implications of this approach remain unknown. Andrographolide, isolated from an herb, appears to reduce MexAB-OprM expression via transcriptional inhibition and to increase drug susceptibility [328]. The use of a deoxyribozyme (i.e., DNA molecules with catalytic action in gene replication) against the mRNA of a probable ATP-binding component of an ABC transporter (which is likely PA2812, homologous to CcmA involving in cytochrome c maturation) seems to be able to decrease ciprofloxacin resistance in vitro [124]. The antisense phosphorothioate oligodeoxynucleotides which targeted the oprM gene and were encapsulated in anionic liposomes were shown to reduce oprM expression and to increase antimicrobial susceptibility of multidrug-resistant isolates [329].
6 Concluding Remarks
Over the last two decades, huge advances have been achieved in our in-depth understanding of multidrug efflux systems of P. aeruginosa. These efflux pumps play a predominant role in clinically relevant MDR, which demonstrates a remarkable ability of P. aeruginosa to develop sophisticated defense mechanisms against a variety of old and new antimicrobial agents. Actually, very few existing drugs appear to escape the multiple and complementary efflux pumps in this microorganism. Efflux phenomenon can not only serve as the initial mechanism of resistance to acquire other means of resistance but also interplay synergistically with them to raise resistance levels. The high percentages of efflux mutants from clinical settings around the globe further highlight the significance of these drug efflux systems as a major in vivo mechanism of resistance, which also link resistance selection and cross-resistance between conventional antibiotics and biocides. Minimizing exposure of P. aeruginosa to multiple structurally unrelated efflux selecting antimicrobial agents would limit the development of resistance, including multidrug-resistant efflux mutants, thus providing another compelling argument for antimicrobial stewardship in any environment that includes prudent antimicrobial use in both clinical settings and community hygiene practice. Efflux mechanisms can also be taken into consideration in pharmacokinetic-pharmacodynamics of individual antimicrobial agents to guide clinical drug use in minimizing resistance emergence [330]. Evidently, therapeutic approaches to intervene in efflux mechanisms are attractive for antimicrobial research and development, in particular because drug efflux systems also contribute to stress responses and virulence factor production. The increasing structural and biochemical understanding of drug efflux pumps such as drug recognition or binding sites and transport kinetics should facilitate such an effort. However, despite the progress made in the field of drug efflux research to date, challenges continue to be faced in the development of novel antimicrobial agents or efflux pump inhibitors that can be applied to combat infections associated with multidrug-resistant P. aeruginosa.
7 Addendum in Proof
A lytic bacteriophage of the Myoviridae family was recently shown to utilize OprM as a receptor-binding site and consequently to compromise the function of MexAB-OprM and MexXY-OprM efflux systems, leading to restore antimicrobial susceptibility in multidrug-resistant isolates [331]. A new study has reported the transport, via MexGHI-OpmD pump, of 5-methylphenazine-1-carboxylate, an intermediate involved in phenazine biosynthesis in the conversion of phenazine-1-carboxylic acid to pyocyanin [332]. Expression of MexGHI-OmpD is sufficiently induced by 5-methylphenazine-1-carboxylate and this induction is required for biofilm development. Finally, a recent study has revealed that MexR with Arg21Trp mutation displays a mutation-induced allosteric coupling of contact networks that are independent of the wild-type MexR protein in the regulation of MexAB-OprM expression, suggesting a novel mechanism for MarR family derepression that mimics derepression by small-molecule binding to MarR proteins [333].
References
McCarthy K (2015) Pseudomonas aeruginosa: evolution of antimicrobial resistance and implications for therapy. Semin Respir Crit Care Med 36:44–55. doi:10.1055/s-0034-1396907
Sousa AM, Pereira MO (2014) Pseudomonas aeruginosa diversification during infection development in cystic fibrosis lungs-a review. Pathogens 3:680–703. doi:10.3390/pathogens3030680
Haenni M, Hocquet D, Ponsin C, Cholley P, Guyeux C, Madec JY, Bertrand X (2015) Population structure and antimicrobial susceptibility of Pseudomonas aeruginosa from animal infections in France. BMC Vet Res 11:9. doi:10.1186/s12917-015-0324-x
King JD, Kocincova D, Westman EL, Lam JS (2009) Review: lipopolysaccharide biosynthesis in Pseudomonas aeruginosa. Innate Immunol 15:261–312. doi:10.1177/1753425909106436
Burrows LL (2012) Pseudomonas aeruginosa twitching motility: type IV pili in action. Annu Rev Microbiol 66:493–520. doi:10.1146/annurev-micro-092611-150055
Kazmierczak BI, Schniederberend M, Jain R (2015) Cross-regulation of Pseudomonas motility systems: the intimate relationship between flagella, pili and virulence. Curr Opin Microbiol 28:78–82. doi:10.1016/j.mib.2015.07.017
Mah TF, Pitts B, Pellock B, Walker GC, Stewart PS, O’Toole GA (2003) A genetic basis for Pseudomonas aeruginosa biofilm antibiotic resistance. Nature 426:306–310. doi:10.1038/nature02122
Høiby N, Ciofu O, Bjarnsholt T (2010) Pseudomonas aeruginosa biofilms in cystic fibrosis. Future Microbiol 5:1663–1674. doi:10.2217/fmb.10.125
Livermore DM (2001) Of Pseudomonas, porins, pumps and carbapenems. J Antimicrob Chemother 47:247–250. doi:10.1093/jac/47.3.247
Breidenstein EB, de la Fuente-Nunez C, Hancock RE (2011) Pseudomonas aeruginosa: all roads lead to resistance. Trends Microbiol 19:419–426. doi:10.1016/j.tim.2011.04.005
U. S. Centers for Disease Control and Prevention (2013) Antibiotic resistance threats in the United States, 2013. CDC, Atlanta
Huband MD, Castanheira M, Flamm RK, Farrell DJ, Jones RN, Sader HS (2016) In vitro activity of ceftazidime-avibactam against contemporary Pseudomonas aeruginosa isolates from United States medical centers by Census region (2014). Antimicrob Agents Chemother 60:2537–2541. doi:10.1128/AAC.03056-15
Nikaido H (2003) Molecular basis of bacterial outer membrane permeability revisited. Microbiol Mol Biol Rev 67:593–656. doi:10.1128/MMBR.67.4.593-656.2003
Li X-Z, Plésiat P, Nikaido H (2015) The challenge of efflux-mediated antibiotic resistance in Gram-negative bacteria. Clin Microbiol Rev 28:337–418. doi:10.1128/CMR.00117-14
Poole K, Krebes K, McNally C, Neshat S (1993) Multiple antibiotic resistance in Pseudomonas aeruginosa: evidence for involvement of an efflux operon. J Bacteriol 175:7363–7372
Li X-Z, Livermore DM, Nikaido H (1994) Role of efflux pump(s) in intrinsic resistance of Pseudomonas aeruginosa: resistance to tetracycline, chloramphenicol, and norfloxacin. Antimicrob Agents Chemother 38:1732–1741. doi:10.1128/AAC.38.8.1732
Li X-Z, Ma D, Livermore DM, Nikaido H (1994) Role of efflux pump(s) in intrinsic resistance of Pseudomonas aeruginosa: active efflux as a contributing factor to β-lactam resistance. Antimicrob Agents Chemother 38:1742–1752. doi:10.1128/AAC.38.8.1742
Li X-Z, Nikaido H, Poole K (1995) Role of MexA-MexB-OprM in antibiotic efflux in Pseudomonas aeruginosa. Antimicrob Agents Chemother 39:1948–1953. doi:10.1128/AAC.39.9.1948
Poole K (2001) Multidrug efflux pumps and antimicrobial resistance in Pseudomonas aeruginosa and related organisms. J Mol Microbiol Biotechnol 3:255–264
Li X-Z (2012) Multidrug resistance efflux pumps of Pseudomonas aeruginosa: a 10-year update. Chin J Antibiot 37:481–500. doi:10.13461/j.cnki.cja.005039
Poole K (2013) Pseudomonas aeruginosa efflux pumps. In: Yu EW, Zhang Q, Brown MH (eds) Microbial efflux pumps: current research. Caister Academic Press, Norfolk, pp 175–206
Holloway BW (1969) Genetics of Pseudomonas. Bacteriol Rev 33:419–443
Rolinson GN (1971) Bacterial resistance to penicillins and cephalosporins. Proc R Soc Lond B Biol Sci 179:403–410. doi:10.1098/rspb.1971.0105
Sykes RB (1975) Resistance of Pseudomonas aeruginosa to antimicrobial drugs. Prog Med Chem 12:333–393. doi:10.1016/S0079-6468(08)70180-2
Leive L (1974) The barrier function of the Gram-negative envelope. Ann N Y Acad Sci 235:109–129. doi:10.1111/j.1749-6632.1974.tb43261.x
Decad GM, Nikaido H (1976) Outer membrane of Gram-negative bacteria. XII. Molecular-sieving function of cell wall. J Bacteriol 128:325–336
Hancock RE, Decad GM, Nikaido H (1979) Identification of the protein producing transmembrane diffusion pores in the outer membrane of Pseudomonas aeruginosa PAO1. Biochim Biophys Acta 554:323–331. doi:10.1016/0005-2736(79)90373-0
Benz R, Hancock RE (1981) Properties of the large ion-permeable pores formed from protein F of Pseudomonas aeruginosa in lipid bilayer membranes. Biochim Biophys Acta 646:298–308. doi:10.1016/0005-2736(81)90336-9
Angus BL, Carey AM, Caron DA, Kropinski AM, Hancock RE (1982) Outer membrane permeability in Pseudomonas aeruginosa: comparison of a wild-type with an antibiotic-supersusceptible mutant. Antimicrob Agents Chemother 21:299–309. doi:10.1128/AAC.21.2.299
Yoshimura F, Nikaido H (1982) Permeability of Pseudomonas aeruginosa outer membrane to hydrophilic solutes. J Bacteriol 152:636–642
Nikaido H, Nikaido K, Harayama S (1991) Identification and characterization of porins in Pseudomonas aeruginosa. J Biol Chem 266:770–779
Sugawara E, Nestorovich EM, Bezrukov SM, Nikaido H (2006) Pseudomonas aeruginosa porin OprF exists in two different conformations. J Biol Chem 281:16220–16229. doi:10.1074/jbc.M600680200
Sugawara E, Nagano K, Nikaido H (2010) Factors affecting the folding of Pseudomonas aeruginosa OprF porin into the one-domain open conformer. mBio 1:e00228–10. doi:10.1128/mBio.00228-10
Plésiat P, Nikaido H (1992) Outer membranes of Gram-negative bacteria are permeable to steroid probes. Mol Microbiol 6:1323–1333. doi:10.1111/j.1365-2958.1992.tb00853.x
Plésiat P, Aires JR, Godard C, Kohler T (1997) Use of steroids to monitor alterations in the outer membrane of Pseudomonas aeruginosa. J Bacteriol 179:7004–7010
Zimmermann W (1980) Penetration of β-lactam antibiotics into their target enzymes in Pseudomonas aeruginosa: comparison of a highly sensitive mutant with its parent strain. Antimicrob Agents Chemother 18:94–100. doi:10.1128/AAC.18.1.94
Kropinski AM, Kuzio J, Angus BL, Hancock RE (1982) Chemical and chromatographic analysis of lipopolysaccharide from an antibiotic-supersusceptible mutant of Pseudomonas aeruginosa. Antimicrob Agents Chemother 21:310–319. doi:10.1128/AAC.21.2.310
Bryan LE, O’Hara K, Wong S (1984) Lipopolysaccharide changes in impermeability-type aminoglycoside resistance in Pseudomonas aeruginosa. Antimicrob Agents Chemother 26:250–255. doi:10.1128/AAC.26.2.250
Trias J, Nikaido H (1990) Outer membrane protein D2 catalyzes facilitated diffusion of carbapenems and penems through the outer membrane of Pseudomonas aeruginosa. Antimicrob Agents Chemother 34:52–57. doi:10.1128/AAC.34.1.52
Preheim LC, Penn RG, Sanders CC, Goering RV, Giger DK (1982) Emergence of resistance to β-lactam and aminoglycoside antibiotics during moxalactam therapy of Pseudomonas aeruginosa infections. Antimicrob Agents Chemother 22:1037–1041. doi:10.1128/AAC.22.6.1037
Sanders CC, Sanders WE Jr, Goering RV, Werner V (1984) Selection of multiple antibiotic resistance by quinolones, β-lactams, and aminoglycosides with special reference to cross-resistance between unrelated drug classes. Antimicrob Agents Chemother 26:797–801. doi:10.1128/AAC.26.6.797
Bragman S, Sage R, Booth L, Noone P (1986) Ceftazidime in the treatment of serious Pseudomonas aeruginosa sepsis. Scand J Infect Dis 18:425–429. doi:10.3109/00365548609032359
Chow AW, Wong J, Bartlett KH, Shafran SD, Stiver HG (1989) Cross-resistance of Pseudomonas aeruginosa to ciprofloxacin, extended-spectrum β-lactams, and aminoglycosides and susceptibility to antibiotic combinations. Antimicrob Agents Chemother 33:1368–1372. doi:10.1128/AAC.33.8.1368
Piddock LJ, Hall MC, Bellido F, Bains M, Hancock RE (1992) A pleiotropic, posttherapy, enoxacin-resistant mutant of Pseudomonas aeruginosa. Antimicrob Agents Chemother 36:1057–1061. doi:10.1128/AAC.36.5.1057
Rella M, Haas D (1982) Resistance of Pseudomonas aeruginosa PAO to nalidixic acid and low levels of β-lactam antibiotics: mapping of chromosomal genes. Antimicrob Agents Chemother 22:242–249. doi:10.1128/AAC.33.1.124
Robillard NJ, Scarpa AL (1988) Genetic and physiological characterization of ciprofloxacin resistance in Pseudomonas aeruginosa PAO. Antimicrob Agents Chemother 32:535–539. doi:10.1128/AAC.33.1.124
Legakis NJ, Tzouvelekis LS, Makris A, Kotsifaki H (1989) Outer membrane alterations in multiresistant mutants of Pseudomonas aeruginosa selected by ciprofloxacin. Antimicrob Agents Chemother 33:124–127. doi:10.1128/AAC.33.1.124
Celesk RA, Robillard NJ (1989) Factors influencing the accumulation of ciprofloxacin in Pseudomonas aeruginosa. Antimicrob Agents Chemother 33:1921–1926. doi:10.1128/AAC.33.11.1921
Fukuda H, Hosaka M, Hirai K, Iyobe S (1990) New norfloxacin resistance gene in Pseudomonas aeruginosa PAO. Antimicrob Agents Chemother 34:1757–1761. doi:10.1128/AAC.34.9.1757
Hashmi ZS, Smith JM (1991) Outer membrane changes in quinolone resistant Pseudomonas aeruginosa. J Antimicrob Chemother 28:465–470. doi:10.1093/jac/28.3.465
Masuda N, Ohya S (1992) Cross-resistance to meropenem, cephems, and quinolones in Pseudomonas aeruginosa. Antimicrob Agents Chemother 36:1847–1851. doi:10.1128/AAC.36.9.1847
Lei Y, Sato K, Nakae T (1991) Ofloxacin-resistant Pseudomonas aeruginosa mutants with elevated drug extrusion across the inner membrane. Biochem Biophys Res Commun 178:1043–1048. doi:10.1016/0006-291X(91)90997-L
Zimmermann W, Rosselet A (1977) Function of the outer membrane of Escherichia coli as a permeability barrier to β-lactam antibiotics. Antimicrob Agents Chemother 12:368–372. doi:10.1128/AAC.12.3.368
Nikaido H, Normark S (1987) Sensitivity of Escherichia coli to various β-lactams is determined by the interplay of outer membrane permeability and degradation by periplasmic β-lactamases: a quantitative predictive treatment. Mol Microbiol 1:29–36. doi:10.1111/j.1365-2958.1987.tb00523.x
Livermore DM, Davy KW (1991) Invalidity for Pseudomonas aeruginosa of an accepted model of bacterial permeability to β-lactam antibiotics. Antimicrob Agents Chemother 35:916–921. doi:10.1128/AAC.35.5.916
Nikaido H (2011) To the happy few. Ann Rev Microbiol 65:1–18. doi:10.1146/annurev-micro-090110-102920
Poole K, Gotoh N, Tsujimoto H, Zhao Q, Wada A, Yamasaki T, Neshat S, Yamagishi J et al (1996) Overexpression of the mexC-mexD-oprJ efflux operon in nfxB-type multidrug-resistant strains of Pseudomonas aeruginosa. Mol Microbiol 21:713–724. doi:10.1046/j.1365-2958.1996.281397.x
Köhler T, Michea-Hamzehpour M, Henze U, Gotoh N, Curty LK, Pechère JC (1997) Characterization of MexE-MexF-OprN, a positively regulated multidrug efflux system of Pseudomonas aeruginosa. Mol Microbiol 23:345–354. doi:10.1046/j.1365-2958.1997.2281594.x
Aires JR, Köhler T, Nikaido H, Plésiat P (1999) Involvement of an active efflux system in the natural resistance of Pseudomonas aeruginosa to aminoglycosides. Antimicrob Agents Chemother 43:2624–2628
Mine T, Morita Y, Kataoka A, Mizushima T, Tsuchiya T (1999) Expression in Escherichia coli of a new multidrug efflux pump, MexXY, from Pseudomonas aeruginosa. Antimicrob Agents Chemother 43:415–417
Westbrock-Wadman S, Sherman DR, Hickey MJ, Coulter SN, Zhu YQ, Warrener P, Nguyen LY, Shawar RM et al (1999) Characterization of a Pseudomonas aeruginosa efflux pump contributing to aminoglycoside impermeability. Antimicrob Agents Chemother 43:2975–2983
Stover CK, Pham XQ, Erwin AL, Mizoguchi SD, Warrener P, Hickey MJ, Brinkman FS, Hufnagle WO et al (2000) Complete genome sequence of Pseudomonas aeruginosa PAO1, an opportunistic pathogen. Nature 406:959–964. doi:10.1038/35023079
Tseng TT, Gratwick KS, Kollman J, Park D, Nies DH, Goffeau A, Saier MH Jr (1999) The RND permease superfamily: an ancient, ubiquitous and diverse family that includes human disease and development proteins. J Mol Microbiol Biotechnol 1:107–125. doi:10.1007/s13205-013-0155-z
Lee DG, Urbach JM, Wu G, Liberati NT, Feinbaum RL, Miyata S, Diggins LT, He J et al (2006) Genomic analysis reveals that Pseudomonas aeruginosa virulence is combinatorial. Genome Biol 7:R90. doi:10.1186/gb-2006-7-10-r90
Li X-Z, Zhang L, Srikumar R, Poole K (1998) β-Lactamase inhibitors are substrates for the multidrug efflux pumps of Pseudomonas aeruginosa. Antimicrob Agents Chemother 42:399–403
Li X-Z, Zhang L, Poole K (1998) Role of the multidrug efflux systems of Pseudomonas aeruginosa in organic solvent tolerance. J Bacteriol 180:2987–2991
Masuda N, Sakagawa E, Ohya S, Gotoh N, Tsujimoto H, Nishino T (2000) Substrate specificities of MexAB-OprM, MexCD-OprJ, and MexXY-OprM efflux pumps in Pseudomonas aeruginosa. Antimicrob Agents Chemother 44:3322–3327. doi:10.1128/AAC.44.12.3322-3327.2000
Poole K, Tetro K, Zhao Q, Neshat S, Heinrichs DE, Bianco N (1996) Expression of the multidrug resistance operon mexA-mexB-oprM in Pseudomonas aeruginosa: mexR encodes a regulator of operon expression. Antimicrob Agents Chemother 40:2021–2028
Cao L, Srikumar R, Poole K (2004) MexAB-OprM hyperexpression in NalC-type multidrug-resistant Pseudomonas aeruginosa: identification and characterization of the nalC gene encoding a repressor of PA3720–PA3719. Mol Microbiol 53:1423–1436. doi:10.1111/j.1365-2958.2004.04210.x
Sobel ML, Hocquet D, Cao L, Plesiat P, Poole K (2005) Mutations in PA3574 (nalD) lead to increased MexAB-OprM expression and multidrug resistance in laboratory and clinical isolates of Pseudomonas aeruginosa. Antimicrob Agents Chemother 49:1782–1786. doi:10.1128/AAC.49.5.1782-1786.2005
Zhang L, Li X-Z, Poole K (2001) Fluoroquinolone susceptibilities of efflux-mediated multidrug-resistant Pseudomonas aeruginosa, Stenotrophomonas maltophilia and Burkholderia cepacia. J Antimicrob Chemother 48:549–552. doi:10.1093/jac/48.4.549
Minagawa S, Inami H, Kato T, Sawada S, Yasuki T, Miyairi S, Horikawa M, Okuda J et al (2012) RND type efflux pump system MexAB-OprM of Pseudomonas aeruginosa selects bacterial languages, 3-oxo-acyl-homoserine lactones, for cell-to-cell communication. BMC Microbiol 12:70. doi:10.1186/1471-2180-12-70
Schweizer HP (1998) Intrinsic resistance to inhibitors of fatty acid biosynthesis in Pseudomonas aeruginosa is due to efflux: application of a novel technique for generation of unmarked chromosomal mutations for the study of efflux systems. Antimicrob Agents Chemother 42:394–398
Mistry A, Warren MS, Cusick JK, Karkhoff-Schweizer RR, Lomovskaya O, Schweizer HP (2013) High-level pacidamycin resistance in Pseudomonas aeruginosa is mediated by an opp oligopeptide permease encoded by the opp-fabI operon. Antimicrob Agents Chemother 57:5565–5571. doi:10.1128/AAC.01198-13
Moore JD, Gerdt JP, Eibergen NR, Blackwell HE (2014) Active efflux influences the potency of quorum sensing inhibitors in Pseudomonas aeruginosa. Chembiochem 15:435–442. doi:10.1002/cbic.201300701
Caughlan RE, Jones AK, Delucia AM, Woods AL, Xie L, Ma B, Barnes SW, Walker JR et al (2012) Mechanisms decreasing in vitro susceptibility to the LpxC inhibitor CHIR-090 in the Gram-negative pathogen Pseudomonas aeruginosa. Antimicrob Agents Chemother 56:17–27. doi:10.1128/AAC.05417-11
Muller JF, Stevens AM, Craig J, Love NG (2007) Transcriptome analysis reveals that multidrug efflux genes are upregulated to protect Pseudomonas aeruginosa from pentachlorophenol stress. Appl Environ Microbiol 73:4550–4558. doi:10.1128/AEM.00169-07
Morita Y, Cao L, Gould G, Avison MB, Poole K (2006) nalD encodes a second repressor of the mexAB-oprM multidrug efflux operon of Pseudomonas aeruginosa. J Bacteriol 188:8649–8654. doi:10.1128/JB.01342-06
Maseda H, Sawada I, Saito K, Uchiyama H, Nakae T, Nomura N (2004) Enhancement of the mexAB-oprM efflux pump expression by a quorum-sensing autoinducer and its cancellation by a regulator, MexT, of the mexEF-oprN efflux pump operon in Pseudomonas aeruginosa. Antimicrob Agents Chemother 48:1320–1328. doi:10.1128/AAC.48.4.1320-1328.2004
Sivaneson M, Mikkelsen H, Ventre I, Bordi C, Filloux A (2011) Two-component regulatory systems in Pseudomonas aeruginosa: an intricate network mediating fimbrial and efflux pump gene expression. Mol Microbiol 79:1353–1366. doi:10.1111/j.1365-2958.2010.07527.x
Liao J, Schurr MJ, Sauer K (2013) The MerR-like regulator BrlR confers biofilm tolerance by activating multidrug efflux pumps in Pseudomonas aeruginosa biofilms. J Bacteriol 195:3352–3363. doi:10.1128/JB.00318-13
Dean CR, Visalli MA, Projan SJ, Sum PE, Bradford PA (2003) Efflux-mediated resistance to tigecycline (GAR-936) in Pseudomonas aeruginosa PAO1. Antimicrob Agents Chemother 47:972–978. doi:10.1128/AAC.47.3.972-978.2003
Murata T, Gotoh N, Nishino T (2002) Characterization of outer membrane efflux proteins OpmE, OpmD and OpmB of Pseudomonas aeruginosa: molecular cloning and development of specific antisera. FEMS Microbiol Lett 217:57–63. doi:10.1111/j.1574-6968.2002.tb11456.x
Baum EZ, Crespo-Carbone SM, Morrow BJ, Davies TA, Foleno BD, He W, Queenan AM, Bush K (2009) Effect of MexXY overexpression on ceftobiprole susceptibility in Pseudomonas aeruginosa. Antimicrob Agents Chemother 53:2785–2790. doi:10.1128/AAC.00018-09
Caughlan RE, Sriram S, Daigle DM, Woods AL, Buco J, Peterson RL, Dzink-Fox J, Walker S et al (2009) Fmt bypass in Pseudomonas aeruginosa causes induction of MexXY efflux pump expression. Antimicrob Agents Chemother 53:5015–5021. doi:10.1128/AAC.00253-09
Muller C, Plésiat P, Jeannot K (2011) A two-component regulatory system interconnects resistance to polymyxins, aminoglycosides, fluoroquinolones, and β-lactams in Pseudomonas aeruginosa. Antimicrob Agents Chemother 55:1211–1221. doi:10.1128/AAC.01252-10
Lee S, Hinz A, Bauerle E, Angermeyer A, Juhaszova K, Kaneko Y, Singh PK, Manoil C (2009) Targeting a bacterial stress response to enhance antibiotic action. Proc Natl Acad Sci U S A 106:14570–14575. doi:10.1073/pnas.0903619106
Morita Y, Gilmour C, Metcalf D, Poole K (2009) Translational control of the antibiotic inducibility of the PA5471 gene required for mexXY multidrug efflux gene expression in Pseudomonas aeruginosa. J Bacteriol 191:4966–4975. doi:10.1128/JB.00073-09
Shi J, Jin Y, Bian T, Li K, Sun Z, Cheng Z, Jin S, Wu W (2015) SuhB is a novel ribosome associated protein that regulates expression of MexXY by modulating ribosome stalling in Pseudomonas aeruginosa. Mol Microbiol 98:370–383. doi:10.1111/mmi.13126
Lau CH, Fraud S, Jones M, Peterson SN, Poole K (2013) Mutational activation of the AmgRS two-component system in aminoglycoside-resistant Pseudomonas aeruginosa. Antimicrob Agents Chemother 57:2243–2251. doi:10.1128/AAC.00170-13
Lau CH, Krahn T, Gilmour C, Mullen E, Poole K (2014) AmgRS-mediated envelope stress-inducible expression of the mexXY multidrug efflux operon of Pseudomonas aeruginosa. Microbiol Open 4:121–135. doi:10.1002/mbo3.226
Morita Y, Murata T, Mima T, Shiota S, Kuroda T, Mizushima T, Gotoh N, Nishino T et al (2003) Induction of mexCD-oprJ operon for a multidrug efflux pump by disinfectants in wild-type Pseudomonas aeruginosa PAO1. J Antimicrob Chemother 51:991–994. doi:10.1093/jac/dkg173
Fraud S, Campigotto AJ, Chen Z, Poole K (2008) MexCD-OprJ multidrug efflux system of Pseudomonas aeruginosa: involvement in chlorhexidine resistance and induction by membrane-damaging agents dependent upon the AlgU stress response sigma factor. Antimicrob Agents Chemother 52:4478–4482. doi:10.1128/AAC.01072-08
Jeannot K, Elsen S, Köhler T, Attree I, van Delden C, Plésiat P (2008) Resistance and virulence of Pseudomonas aeruginosa clinical strains overproducing the MexCD-OprJ efflux pump. Antimicrob Agents Chemother 52:2455–2462. doi:10.1128/AAC.01107-07
De Silva M, Ning C, Ghanbar S, Zhanel G, Logsetty S, Liu S, Kumar A (2015) Evidence that a novel quaternary compound and its organic N-chloramine derivative do not select for resistant mutants of Pseudomonas aeruginosa. J Hosp Infect 91:53–58. doi:10.1016/j.jhin.2015.05.009
Nayar AS, Dougherty TJ, Reck F, Thresher J, Gao N, Shapiro AB, Ehmann DE (2015) Target-based resistance in Pseudomonas aeruginosa and Escherichia coli to NBTI 5463, a novel bacterial type II topoisomerase inhibitor. Antimicrob Agents Chemother 59:331–337. doi:10.1128/AAC.04077-14
Purssell A, Poole K (2013) Functional characterization of the NfxB repressor of the mexCD-oprJ multidrug efflux operon of Pseudomonas aeruginosa. Microbiology 159:2058–2073. doi:10.1099/mic.0.069286-0
Liang H, Deng X, Li X, Ye Y, Wu M (2014) Molecular mechanisms of master regulator VqsM mediating quorum-sensing and antibiotic resistance in Pseudomonas aeruginosa. Nucleic Acids Res 42:10307–10320. doi:10.1093/nar/gku586
Gillis RJ, White KG, Choi K-H, Wagner VE, Schweizer HP, Iglewski BH (2005) Molecular basis of azithromycin-resistant Pseudomonas aeruginosa biofilms. Antimicrob Agents Chemother 49:3858–3867. doi:10.1128/AAC.49.9.3858-3867.2005
Fargier E, Mac Aogain M, Mooij MJ, Woods DF, Morrissey JP, Dobson AD, Adams C, O’Gara F (2012) MexT functions as a redox-responsive regulator modulating disulfide stress resistance in Pseudomonas aeruginosa. J Bacteriol 194:3502–3511. doi:10.1128/JB.06632-11
Wang D, Seeve C, Pierson LS 3rd, Pierson EA (2013) Transcriptome profiling reveals links between ParS/ParR, MexEF-OprN, and quorum sensing in the regulation of adaptation and virulence in Pseudomonas aeruginosa. BMC Genomics 14:618. doi:10.1186/1471-2164-14-618
Westfall LW, Carty NL, Layland N, Kuan P, Colmer-Hamood JA, Hamood AN (2006) mvaT mutation modifies the expression of the Pseudomonas aeruginosa multidrug efflux operon mexEF-oprN. FEMS Microbiol Lett 255:247–254. doi:10.1111/j.1574-6968.2005.00075.x
Lamarche MG, Deziel E (2011) MexEF-OprN efflux pump exports the Pseudomonas quinolone signal (PQS) precursor HHQ (4-hydroxy-2-heptylquinoline). PLoS One 6:e24310. doi:10.1371/journal.pone.0024310
Aendekerk S, Diggle SP, Song Z, Hoiby N, Cornelis P, Williams P, Camara M (2005) The MexGHI-OpmD multidrug efflux pump controls growth, antibiotic susceptibility and virulence in Pseudomonas aeruginosa via 4-quinolone-dependent cell-to-cell communication. Microbiology 151:1113–1125. doi:10.1099/mic.0.27631-0
Palma M, Zurita J, Ferreras JA, Worgall S, Larone DH, Shi L, Campagne F, Quadri LE (2005) Pseudomonas aeruginosa SoxR does not conform to the archetypal paradigm for SoxR-dependent regulation of the bacterial oxidative stress adaptive response. Infect Immun 73:2958–2966. doi:10.1128/IAI.73.5.2958-2966.2005
Dietrich LE, Price-Whelan A, Petersen A, Whiteley M, Newman DK (2006) The phenazine pyocyanin is a terminal signalling factor in the quorum sensing network of Pseudomonas aeruginosa. Mol Microbiol 61:1308–1321. doi:10.1111/j.1365-2958.2006.05306.x
Chuanchuen R, Narasaki CT, Schweizer HP (2002) The MexJK efflux pump of Pseudomonas aeruginosa requires OprM for antibiotic efflux but not for efflux of triclosan. J Bacteriol 184:5036–5044. doi:10.1128/JB.184.18.5036-5044.2002
Chuanchuen R, Gaynor JB, Karkhoff-Schweizer R, Schweizer HP (2005) Molecular characterization of MexL, the transcriptional repressor of the mexJK multidrug efflux operon in Pseudomonas aeruginosa. Antimicrob Agents Chemother 49:1844–1851. doi:10.1128/AAC.49.5.1844-1851.2005
Mima T, Sekiya H, Mizushima T, Kuroda T, Tsuchiya T (2005) Gene cloning and properties of the RND-type multidrug efflux pumps MexPQ-OpmE and MexMN-OprM from Pseudomonas aeruginosa. Microbiol Immunol 49:999–1002. doi:10.1111/j.1348-0421.2005.tb03696.x
Li Y, Mima T, Komori Y, Morita Y, Kuroda T, Mizushima T, Tsuchiya T (2003) A new member of the tripartite multidrug efflux pumps, MexVW-OprM, in Pseudomonas aeruginosa. J Antimicrob Chemother 52:572–575. doi:10.1093/jac/dkg390
Chiang WC, Pamp SJ, Nilsson M, Givskov M, Tolker-Nielsen T (2012) The metabolically active subpopulation in Pseudomonas aeruginosa biofilms survives exposure to membrane-targeting antimicrobials via distinct molecular mechanisms. FEMS Immunol Med Microbiol 65:245–256. doi:10.1111/j.1574-695X.2012.00929.x
Yang L, Chen L, Shen L, Surette M, Duan K (2011) Inactivation of MuxABC-OpmB transporter system in Pseudomonas aeruginosa leads to increased ampicillin and carbenicillin resistance and decreased virulence. J Microbiol 49:107–114. doi:10.1007/s12275-011-0186-2
Mima T, Kohira N, Li Y, Sekiya H, Ogawa W, Kuroda T, Tsuchiya T (2009) Gene cloning and characteristics of the RND-type multidrug efflux pump MuxABC-OpmB possessing two RND components in Pseudomonas aeruginosa. Microbiology 155:3509–3517. doi:10.1099/mic.0.031260-0
Mima T, Joshi S, Gomez-Escalada M, Schweizer HP (2007) Identification and characterization of TriABC-OpmH, a triclosan efflux pump of Pseudomonas aeruginosa requiring two membrane fusion proteins. J Bacteriol 189:7600–7609. doi:10.1128/JB.00850-07
Hassan MT, van der Lelie D, Springael D, Romling U, Ahmed N, Mergeay M (1999) Identification of a gene cluster, czr, involved in cadmium and zinc resistance in Pseudomonas aeruginosa. Gene 238:417–425. doi:10.1016/S0378-1119(99)00349-2
Perron K, Caille O, Rossier C, Van Delden C, Dumas JL, Köhler T (2004) CzcR-CzcS, a two-component system involved in heavy metal and carbapenem resistance in Pseudomonas aeruginosa. J Biol Chem 279:8761–8768. doi:10.1074/jbc.M312080200
Caille O, Rossier C, Perron K (2007) A copper-activated two-component system interacts with zinc and imipenem resistance in Pseudomonas aeruginosa. J Bacteriol 189:4561–4568. doi:10.1128/JB.00095-07
He GX, Kuroda T, Mima T, Morita Y, Mizushima T, Tsuchiya T (2004) An H+-coupled multidrug efflux pump, PmpM, a member of the MATE family of transporters, from Pseudomonas aeruginosa. J Bacteriol 186:262–265. doi:10.1128/JB.186.1.262-265.2004
Colinon C, Jocktane D, Brothier E, Rossolini GM, Cournoyer B, Nazaret S (2010) Genetic analyses of Pseudomonas aeruginosa isolated from healthy captive snakes: evidence of high inter- and intrasite dissemination and occurrence of antibiotic resistance genes. Environ Microbiol 12:716–729. doi:10.1111/j.1462-2920.2009.02115.x
Li X-Z, Poole K, Nikaido H (2003) Contributions of MexAB-OprM and an EmrE homolog to intrinsic resistance of Pseudomonas aeruginosa to aminoglycosides and dyes. Antimicrob Agents Chemother 47:27–33. doi:10.1128/AAC.47.1.27-33.2003
Kucken D, Feucht H, Kaulfers P (2000) Association of qacE and qacEΔ1 with multiple resistance to antibiotics and antiseptics in clinical isolates of Gram-negative bacteria. FEMS Microbiol Lett 183:95–98. doi:10.1111/j.1574-6968.2000.tb08939.x
Jeong JH, Shin KS, Lee JW, Park EJ, Son SY (2009) Analysis of a novel class 1 integron containing metallo-β-lactamase gene VIM-2 in Pseudomonas aeruginosa. J Microbiol 47:753–759. doi:10.1007/s12275-008-0272-2
Zhang L, Mah TF (2008) Involvement of a novel efflux system in biofilm-specific resistance to antibiotics. J Bacteriol 190:4447–4452. doi:10.1128/JB.01655-07
Zhou J, Hao D, Wang X, Liu T, He C, Xie F, Sun Y, Zhang J (2006) An important role of a “probable ATP-binding component of ABC transporter” during the process of Pseudomonas aeruginosa resistance to fluoroquinolone. Proteomics 6:2495–2503
Chen L, Duan K (2016) A PhoPQ-regulated ABC transporter system exports tetracycline in Pseudomonas aeruginosa. Antimicrob Agents Chemother 60:3016–3024. doi:10.1128/AAC.02986-15
Hannauer M, Yeterian E, Martin LW, Lamont IL, Schalk IJ (2010) An efflux pump is involved in secretion of newly synthesized siderophore by Pseudomonas aeruginosa. FEBS Lett 584:4751–4755. doi:10.1016/j.febslet.2010.10.051
Hannauer M, Braud A, Hoegy F, Ronot P, Boos A, Schalk IJ (2012) The PvdRT-OpmQ efflux pump controls the metal selectivity of the iron uptake pathway mediated by the siderophore pyoverdine in Pseudomonas aeruginosa. Environ Microbiol 14:1696–1708. doi:10.1111/j.1462-2920.2011.02674.x
Tomaras AP, Crandon JL, McPherson CJ, Nicolau DP (2015) Potentiation of antibacterial activity of the MB-1 siderophore-monobactam conjugate using an efflux pump inhibitor. Antimicrob Agents Chemother 59:2439–2442. doi:10.1128/AAC.04172-14
Verchère A, Dezi M, Adrien V, Broutin I, Picard M (2015) In vitro transport activity of the fully assembled MexAB-OprM efflux pump from Pseudomonas aeruginosa. Nat Commun 6:6890. doi:10.1038/ncomms7890
Evans K, Poole K (1999) The MexA-MexB-OprM multidrug efflux system of Pseudomonas aeruginosa is growth-phase regulated. FEMS Microbiol Lett 173:35–39. doi:10.1111/j.1574-6968.1999.tb13481.x
Masuda N, Sakagawa E, Ohya S (1995) Outer membrane proteins responsible for multiple drug resistance in Pseudomonas aeruginosa. Antimicrob Agents Chemother 39:645–649. doi:10.1128/AAC.35.5.916
Srikumar R, Paul CJ, Poole K (2000) Influence of mutations in the mexR repressor gene on expression of the MexA-MexB-OprM multidrug efflux system of Pseudomonas aeruginosa. J Bacteriol 182:1410–1414. doi:10.1128/JB.182.5.1410-1414.2000
Ziha-Zarifi I, Llanes C, Köhler T, Pechere JC, Plésiat P (1999) In vivo emergence of multidrug-resistant mutants of Pseudomonas aeruginosa overexpressing the active efflux system MexA-MexB-OprM. Antimicrob Agents Chemother 43:287–291
Saito K, Yoneyama H, Nakae T (1999) nalB-type mutations causing the overexpression of the MexAB-OprM efflux pump are located in the mexR gene of the Pseudomonas aeruginosa chromosome. FEMS Microbiol Lett 179:67–72. doi:10.1111/j.1574-6968.1999.tb08709.x
Boutoille D, Corvec S, Caroff N, Giraudeau C, Espaze E, Caillon J, Plesiat P, Reynaud A (2004) Detection of an IS21 insertion sequence in the mexR gene of Pseudomonas aeruginosa increasing β-lactam resistance. FEMS Microbiol Lett 230:143–146. doi:10.1016/S0378-1097(03)00882-6
Llanes C, Hocquet D, Vogne C, Benali-Baitich D, Neuwirth C, Plésiat P (2004) Clinical strains of Pseudomonas aeruginosa overproducing MexAB-OprM and MexXY efflux pumps simultaneously. Antimicrob Agents Chemother 48:1797–1802. doi:10.1128/AAC.48.5.1797-1802.2004
Hocquet D, Roussel-Delvallez M, Cavallo JD, Plesiat P (2007) MexAB-OprM- and MexXY-overproducing mutants are very prevalent among clinical strains of Pseudomonas aeruginosa with reduced susceptibility to ticarcillin. Antimicrob Agents Chemother 51:1582–1583. doi:10.1128/AAC.01334-06
Campo Esquisabel AB, Rodriguez MC, Campo-Sosa AO, Rodriguez C, Martinez-Martinez L (2011) Mechanisms of resistance in clinical isolates of Pseudomonas aeruginosa less susceptible to cefepime than to ceftazidime. Clin Microbiol Infect 17:1817–1822. doi:10.1111/j.1469-0691.2011.03530.x
Sacha P, Wieczorek P, Ojdana D, Hauschild T, Milewski R, Czaban S, Poniatowski B, Tryniszewska E (2014) Expression of MexAB-OprM efflux pump system and susceptibility to antibiotics of different Pseudomonas aeruginosa clones isolated from patients hospitalized in two intensive care units at University Hospital in Bialystok (northeastern Poland) between January 2002 and December 2009. APMIS 122:931–940. doi:10.1111/apm.12236
Aghazadeh M, Hojabri Z, Mahdian R, Nahaei MR, Rahmati M, Hojabri T, Pirzadeh T, Pajand O (2014) Role of efflux pumps: MexAB-OprM and MexXY(-OprA), AmpC cephalosporinase and OprD porin in non-metallo-β-lactamase producing Pseudomonas aeruginosa isolated from cystic fibrosis and burn patients. Infect Genet Evol 24:187–192. doi:10.1016/j.meegid.2014.03.018
Castanheira M, Mills JC, Farrell DJ, Jones RN (2014) Mutation-driven β-lactam resistance mechanisms among contemporary ceftazidime-nonsusceptible Pseudomonas aeruginosa isolates from U.S. hospitals. Antimicrob Agents Chemother 58:6844–6850. doi:10.1128/AAC.03681-14
Riou M, Avrain L, Carbonnelle S, El Garch F, Pirnay JP, De Vos D, Plésiat P, Tulkens PM et al (2016) Increase of efflux-mediated resistance in Pseudomonas aeruginosa during antibiotic treatment in patients suffering from nosocomial pneumonia. Int J Antimicrob Agents 47:77–83. doi:10.1016/j.ijantimicag.2015.11.004
Vestergaard M, Paulander W, Marvig RL, Clasen J, Jochumsen N, Molin S, Jelsbak L, Ingmer H et al (2016) Antibiotic combination therapy can select for broad-spectrum multidrug resistance in Pseudomonas aeruginosa. Int J Antimicrob Agents 47:48–55. doi:10.1016/j.ijantimicag.2015.09.014
Choudhury D, Ghosh A, Dhar Chanda D, Das Talukdar A, Dutta Choudhury M, Paul D, Maurya AP, Chakravorty A et al (2016) Premature termination of MexR leads to overexpression of MexAB-OprM efflux pump in Pseudomonas aeruginosa in a tertiary referral hospital in India. PLoS One 11:e0149156. doi:10.1371/journal.pone.0149156
Quale J, Bratu S, Gupta J, Landman D (2006) Interplay of efflux system, ampC, and oprD expression in carbapenem resistance of Pseudomonas aeruginosa clinical isolates. Antimicrob Agents Chemother 50:1633–1641. doi:10.1128/aac.50.5.1633-1641.2006
Hamzehpour MM, Pechere JC, Plésiat P, Köhler T (1995) OprK and OprM define two genetically distinct multidrug efflux systems in Pseudomonas aeruginosa. Antimicrob Agents Chemother 39:2392–2396. doi:10.1128/AAC.39.11.2392
Köhler T, Michea-Hamzehpour M, Plésiat P, Kahr AL, Pechere JC (1997) Differential selection of multidrug efflux systems by quinolones in Pseudomonas aeruginosa. Antimicrob Agents Chemother 41:2540–2543
Li X-Z, Poole K (1999) Organic solvent-tolerant mutants of Pseudomonas aeruginosa display multiple antibiotic resistance. Can J Microbiol 45:18–22. doi:10.1139/w98-127
Adewoye L, Sutherland A, Srikumar R, Poole K (2002) The mexR repressor of the mexAB-oprM multidrug efflux operon in Pseudomonas aeruginosa: characterization of mutations compromising activity. J Bacteriol 184:4308–4312. doi:10.1128/JB.184.15.4308-4312.2002
Saito K, Akama H, Yoshihara E, Nakae T (2003) Mutations affecting DNA-binding activity of the MexR repressor of mexR-mexA-mexB-oprM operon expression. J Bacteriol 185:6195–6198. doi:10.1128/JB.185.20.6195-6198.2003
Andrésen C, Jalal S, Aili D, Wang Y, Islam S, Jarl A, Liedberg B, Wretlind B et al (2010) Critical biophysical properties in the Pseudomonas aeruginosa efflux gene regulator MexR are targeted by mutations conferring multidrug resistance. Protein Sci 19:680–692. doi:10.1002/pro.343
Pearson JP, Van Delden C, Iglewski BH (1999) Active efflux and diffusion are involved in transport of Pseudomonas aeruginosa cell-to-cell signals. J Bacteriol 181:1203–1210
Nakae T, Saito K, Nakajima A (2000) Effect of sulbactam on anti-pseudomonal activity of β-lactam antibiotics in cells producing various levels of the MexAB-OprM efflux pump and β-lactamase. Microbiol Immunol 44:997–1001. doi:10.1111/j.1348-0421.2000.tb02595.x
Dupont P, Hocquet D, Jeannot K, Chavanet P, Plésiat P (2005) Bacteriostatic and bactericidal activities of eight fluoroquinolones against MexAB-OprM-overproducing clinical strains of Pseudomonas aeruginosa. J Antimicrob Chemother 55:518–522. doi:10.1093/jac/dki030
Papadopoulos CJ, Carson CF, Chang BJ, Riley TV (2008) Role of the MexAB-OprM efflux pump of Pseudomonas aeruginosa in tolerance to tea tree (Melaleuca alternifolia) oil and its monoterpene components terpinen-4-ol, 1,8-cineole, and alpha-terpineol. Appl Environ Microbiol 74:1932–1935. doi:10.1128/AEM.02334-07
Robertson GT, Doyle TB, Du Q, Duncan L, Mdluli KE, Lynch AS (2007) A novel indole compound that inhibits Pseudomonas aeruginosa growth by targeting MreB is a substrate for MexAB-OprM. J Bacteriol 189:6870–6881. doi:10.1128/jb.00805-07
Köhler T, Kok M, Michea-Hamzehpour M, Plesiat P, Gotoh N, Nishino T, Curty LK, Pechere JC (1996) Multidrug efflux in intrinsic resistance to trimethoprim and sulfamethoxazole in Pseudomonas aeruginosa. Antimicrob Agents Chemother 40:2288–2290
Srikumar R, Li X-Z, Poole K (1997) Inner membrane efflux components are responsible for β-lactam specificity of multidrug efflux pumps in Pseudomonas aeruginosa. J Bacteriol 179:7875–7881
Köhler T, Michea-Hamzehpour M, Epp SF, Pechere JC (1999) Carbapenem activities against Pseudomonas aeruginosa: respective contributions of OprD and efflux systems. Antimicrob Agents Chemother 43:424–427
Okamoto K, Gotoh N, Nishino T (2001) Pseudomonas aeruginosa reveals high intrinsic resistance to penem antibiotics: penem resistance mechanisms and their interplay. Antimicrob Agents Chemother 45:1964–1971. doi:10.1128/AAC.45.7.1964-1971.2001
Okamoto K, Gotoh N, Nishino T (2002) Extrusion of penem antibiotics by multicomponent efflux systems MexAB-OprM, MexCD-OprJ, and MexXY-OprM of Pseudomonas aeruginosa. Antimicrob Agents Chemother 46:2696–2699. doi:10.1128/AAC.46.8.2696-2699.2002
Riera E, Cabot G, Mulet X, Garcia-Castillo M, del Campo R, Juan C, Canton R, Oliver A (2011) Pseudomonas aeruginosa carbapenem resistance mechanisms in Spain: impact on the activity of imipenem, meropenem and doripenem. J Antimicrob Chemother 66:2022–2027. doi:10.1093/jac/dkr232
Gotoh N, Tsujimoto H, Nomura A, Okamoto K, Tsuda M, Nishino T (1998) Functional replacement of OprJ by OprM in the MexCD-OprJ multidrug efflux system of Pseudomonas aeruginosa. FEMS Microbiol Lett 165:21–27. doi:10.1111/j.1574-6968.1998.tb13122.x
Li X-Z, Poole K (2001) Mutational analysis of the OprM outer membrane component of the MexA-MexB-OprM multidrug efflux system of Pseudomonas aeruginosa. J Bacteriol 183:12–27. doi:10.1128/JB.183.1.12-27.2001
Akama H, Kanemaki M, Yoshimura M, Tsukihara T, Kashiwagi T, Yoneyama H, Narita S, Nakagawa A et al (2004) Crystal structure of the drug discharge outer membrane protein, OprM, of Pseudomonas aeruginosa: dual modes of membrane anchoring and occluded cavity end. J Biol Chem 279:52816–52819. doi:10.1074/jbc.C400445200
Zhao Q, Li X-Z, Srikumar R, Poole K (1998) Contribution of outer membrane efflux protein OprM to antibiotic resistance in Pseudomonas aeruginosa independent of MexAB. Antimicrob Agents Chemother 42:1682–1688
Chuanchuen R, Murata T, Gotoh N, Schweizer HP (2005) Substrate-dependent utilization of OprM or OpmH by the Pseudomonas aeruginosa MexJK efflux pump. Antimicrob Agents Chemother 49:2133–2136. doi:10.1128/AAC.49.5.2133-2136.2005
Morita Y, Tomida J, Kawamura Y (2012) Primary mechanisms mediating aminoglycoside resistance in the multidrug-resistant Pseudomonas aeruginosa clinical isolate PA7. Microbiology 158:1071–1083. doi:10.1099/mic.0.054320-0
Maseda H, Yoneyama H, Nakae T (2000) Assignment of the substrate-selective subunits of the MexEF-OprN multidrug efflux pump of Pseudomonas aeruginosa. Antimicrob Agents Chemother 44:658–664. doi:10.1128/AAC.44.3.658-664.2000
Cavallo JD, Hocquet D, Plesiat P, Fabre R, Roussel-Delvallez M, Gerpa (2007) Susceptibility of Pseudomonas aeruginosa to antimicrobials: a 2004 French multicentre hospital study. J Antimicrob Chemother 59:1021–1024. doi:10.1093/jac/dkm076
CLSI (2015) Performance standards for antimicrobial susceptibility testing; twenty-fifth informational supplement M100-S25. Clinical and Laboratory Standards Institute, Wayne
Ong CT, Tessier PR, Li C, Nightingale CH, Nicolau DP (2007) Comparative in vivo efficacy of meropenem, imipenem, and cefepime against Pseudomonas aeruginosa expressing MexA-MexB-OprM efflux pumps. Diagn Microbiol Infect Dis 57:153–161. doi:10.1016/j.diagmicrobio.2006.06.014
Adamson DH, Krikstopaityte V, Coote PJ (2015) Enhanced efficacy of putative efflux pump inhibitor/antibiotic combination treatments versus MDR strains of Pseudomonas aeruginosa in a Galleria mellonella in vivo infection model. J Antimicrob Chemother 70:2271–2278. doi:10.1093/jac/dkv111
Boutoille D, Jacqueline C, Le Mabecque V, Potel G, Caillon J (2009) In vivo impact of the MexAB-OprM efflux system on β-lactam efficacy in an experimental model of Pseudomonas aeruginosa infection. Int J Antimicrob Agents 33:417–420. doi:10.1016/j.ijantimicag.2008.10.029
Lomovskaya O, Lee A, Hoshino K, Ishida H, Mistry A, Warren MS, Boyer E, Chamberland S et al (1999) Use of a genetic approach to evaluate the consequences of inhibition of efflux pumps in Pseudomonas aeruginosa. Antimicrob Agents Chemother 43:1340–1346
Lee A, Mao W, Warren MS, Mistry A, Hoshino K, Okumura R, Ishida H, Lomovskaya O (2000) Interplay between efflux pumps may provide either additive or multiplicative effects on drug resistance. J Bacteriol 182:3142–3150. doi:10.1128/JB.182.11.3142-3150.2000
Pumbwe L, Piddock LJ (2000) Two efflux systems expressed simultaneously in multidrug-resistant Pseudomonas aeruginosa. Antimicrob Agents Chemother 44:2861–2864. doi:10.1128/AAC.44.10.2861-2864.2000
Llanes C, Köhler T, Patry I, Dehecq B, van Delden C, Plésiat P (2011) Role of the MexEF-OprN efflux system in low-level resistance of Pseudomonas aeruginosa to ciprofloxacin. Antimicrob Agents Chemother 55:5676–5684. doi:10.1128/AAC.00101-11
Poonsuk K, Tribuddharat C, Chuanchuen R (2014) Simultaneous overexpression of multidrug efflux pumps in Pseudomonas aeruginosa non-cystic fibrosis clinical isolates. Can J Microbiol 60:437–443. doi:10.1139/cjm-2014-0239
Kanchana P, Rungtip C (2014) The multidrug-resistant Pseudomonas aeruginosa clinical isolates from dogs and cats expressed three multidrug efflux systems simultaneously. Thai J Vet Med 44:453–459
Li X-Z, Zhang L, Poole K (2000) Interplay between the MexA-MexB-OprM multidrug efflux system and the outer membrane barrier in the multiple antibiotic resistance of Pseudomonas aeruginosa. J Antimicrob Chemother 45:433–436. doi:10.1093/jac/45.4.433
Li X-Z (2003) Efflux-mediated multiple antibiotic resistance in Pseudomonas aeruginosa. Chin J Antibiot 28:577–596. doi:10.13461/j.cnki.cja.003185
Li X-Z, Barré N, Poole K (2000) Influence of the MexA-MexB-OprM multidrug efflux system on expression of the MexC-MexD-OprJ and MexE-MexF-OprN multidrug efflux systems in Pseudomonas aeruginosa. J Antimicrob Chemother 46:885–893. doi:10.1093/jac/46.6.885
Masuda N, Sakagawa E, Ohya S, Gotoh N, Tsujimoto H, Nishino T (2000) Contribution of the MexX-MexY-OprM efflux system to intrinsic resistance in Pseudomonas aeruginosa. Antimicrob Agents Chemother 44:2242–2246. doi:10.1128/AAC.44.9.2242-2246.2000
Jeannot K, Sobel ML, El Garch F, Poole K, Plesiat P (2005) Induction of the MexXY efflux pump in Pseudomonas aeruginosa is dependent on drug-ribosome interaction. J Bacteriol 187:5341–5346. doi:10.1128/JB.187.15.5341-5346.2005
Lau CH, Hughes D, Poole K (2014) MexY-promoted aminoglycoside resistance in Pseudomonas aeruginosa: involvement of a putative proximal binding pocket in aminoglycoside recognition. mBio 5:e01068–14. doi:10.1128/mBio.01068-14
Poole K, Lau CH, Gilmour C, Hao Y, Lam JS (2015) Polymyxin susceptibility in Pseudomonas aeruginosa linked to the MexXY-OprM multidrug efflux system. Antimicrob Agents Chemother 59:7276–7289. doi:10.1128/AAC.01785-15
Hocquet D, Muller A, Blanc K, Plésiat P, Talon D, Monnet DL, Bertrand X (2008) Relationship between antibiotic use and incidence of MexXY-OprM overproducers among clinical isolates of Pseudomonas aeruginosa. Antimicrob Agents Chemother 52:1173–1175. doi:10.1128/AAC.01212-07
Vogne C, Aires JR, Bailly C, Hocquet D, Plésiat P (2004) Role of the multidrug efflux system MexXY in the emergence of moderate resistance to aminoglycosides among Pseudomonas aeruginosa isolates from patients with cystic fibrosis. Antimicrob Agents Chemother 48:1676–1680. doi:10.1128/AAC.48.5.1676-1680.2004
Smith EE, Buckley DG, Wu Z, Saenphimmachak C, Hoffman LR, D’Argenio DA, Miller SI, Ramsey BW et al (2006) Genetic adaptation by Pseudomonas aeruginosa to the airways of cystic fibrosis patients. Proc Natl Acad Sci U S A 103:8487–8492. doi:10.1073/pnas.0602138103
Feliziani S, Lujan AM, Moyano AJ, Sola C, Bocco JL, Montanaro P, Canigia LF, Argarana CE et al (2010) Mucoidy, quorum sensing, mismatch repair and antibiotic resistance in Pseudomonas aeruginosa from cystic fibrosis chronic airways infections. PLoS One 5: e12669. doi:10.1371/journal.pone.0012669
Mulcahy LR, Burns JL, Lory S, Lewis K (2010) Emergence of Pseudomonas aeruginosa strains producing high levels of persister cells in patients with cystic fibrosis. J Bacteriol 192:6191–6199. doi:10.1128/JB.01651-09
Qin X, Zerr DM, McNutt MA, Berry JE, Burns JL, Kapur RP (2012) Pseudomonas aeruginosa syntrophy in chronically colonized airways of cystic fibrosis patients. Antimicrob Agents Chemother 56:5971–5981. doi:10.1128/AAC.01371-12
Llanes C, Pourcel C, Richardot C, Plésiat P, Fichant G, Cavallo JD, Merens A, Group GS (2013) Diversity of β-lactam resistance mechanisms in cystic fibrosis isolates of Pseudomonas aeruginosa: a French multicentre study. J Antimicrob Chemother 68:1763–1771. doi:10.1093/jac/dkt115
Henrichfreise B, Wiegand I, Pfister W, Wiedemann B (2007) Resistance mechanisms of multiresistant Pseudomonas aeruginosa strains from Germany and correlation with hypermutation. Antimicrob Agents Chemother 51:4062–4070. doi:10.1128/AAC.00148-07
Pournaras S, Maniati M, Spanakis N, Ikonomidis A, Tassios PT, Tsakris A, Legakis NJ, Maniatis AN (2005) Spread of efflux pump-overexpressing, non-metallo-β-lactamase-producing, meropenem-resistant but ceftazidime-susceptible Pseudomonas aeruginosa in a region with blaVIM endemicity. J Antimicrob Chemother 56:761–764. doi:10.1093/jac/dki296
Hocquet D, Nordmann P, El Garch F, Cabanne L, Plesiat P (2006) Involvement of the MexXY-OprM efflux system in emergence of cefepime resistance in clinical strains of Pseudomonas aeruginosa. Antimicrob Agents Chemother 50:1347–1351. doi:10.1128/AAC.50.4.1347-1351.2006
Hocquet D, Berthelot P, Roussel-Delvallez M, Favre R, Jeannot K, Bajolet O, Marty N, Grattard F et al (2007) Pseudomonas aeruginosa may accumulate drug resistance mechanisms without losing its ability to cause bloodstream infections. Antimicrob Agents Chemother 51:3531–3536. doi:10.1128/AAC.00503-07
Xavier DE, Picao RC, Girardello R, Fehlberg LC, Gales AC (2010) Efflux pumps expression and its association with porin down-regulation and β-lactamase production among Pseudomonas aeruginosa causing bloodstream infections in Brazil. BMC Microbiol 10:217. doi:10.1186/1471-2180-10-217
Cabot G, Ocampo-Sosa AA, Dominguez MA, Gago JF, Juan C, Tubau F, Rodriguez C, Moya B et al (2012) Genetic markers of widespread extensively drug-resistant Pseudomonas aeruginosa high-risk clones. Antimicrob Agents Chemother 56:6349–6357. doi:10.1128/AAC.01388-12
Moya B, Beceiro A, Cabot G, Juan C, Zamorano L, Alberti S, Oliver A (2012) Pan-β-lactam resistance development in Pseudomonas aeruginosa clinical strains: molecular mechanisms, penicillin-binding protein profiles, and binding affinities. Antimicrob Agents Chemother 56:4771–4778. doi:10.1128/AAC.00680-12
Fehlberg LC, Xavier DE, Peraro PP, Marra AR, Edmond MB, Gales AC (2012) β-Lactam resistance mechanisms in Pseudomonas aeruginosa strains causing bloodstream infections: comparative results between Brazilian and American isolates. Microb Drug Resist 18:402–407. doi:10.1089/mdr.2011.0174
Fournier D, Richardot C, Muller E, Robert-Nicoud M, Llanes C, Plésiat P, Jeannot K (2013) Complexity of resistance mechanisms to imipenem in intensive care unit strains of Pseudomonas aeruginosa. J Antimicrob Chemother 68:1772–1780. doi:10.1093/jac/dkt098
Castanheira M, Deshpande LM, Costello A, Davies TA, Jones RN (2014) Epidemiology and carbapenem resistance mechanisms of carbapenem-non-susceptible Pseudomonas aeruginosa collected during 2009–11 in 14 European and Mediterranean countries. J Antimicrob Chemother 69:1804–1814. doi:10.1093/jac/dku048
Guénard S, Muller C, Monlezun L, Benas P, Broutin I, Jeannot K, Plésiat P (2014) Multiple mutations lead to MexXY-OprM-dependent aminoglycoside resistance in clinical strains of Pseudomonas aeruginosa. Antimicrob Agents Chemother 58:221–228. doi:10.1128/AAC.01252-13
Galli F, Battistoni A, Gambari R, Pompella A, Bragonzi A, Pilolli F, Iuliano L, Piroddi M et al (2012) Oxidative stress and antioxidant therapy in cystic fibrosis. Biochim Biophys Acta 1822:690–713. doi:10.1016/j.bbadis.2011.12.012
Fraud S, Poole K (2011) Oxidative stress induction of the MexXY multidrug efflux genes and promotion of aminoglycoside resistance development in Pseudomonas aeruginosa. Antimicrob Agents Chemother 55:1068–1074. doi:10.1128/AAC.01495-10
Alguel Y, Lu D, Quade N, Sauter S, Zhang X (2010) Crystal structure of MexZ, a key repressor responsible for antibiotic resistance in Pseudomonas aeruginosa. J Struct Biol 172:305–310. doi:10.1016/j.jsb.2010.07.012
Jahandideh S (2013) Diversity in structural consequences of MexZ mutations in Pseudomonas aeruginosa. Chem Biol Drug Des 81:600–606. doi:10.1111/cbdd.12104
El’Garch F, Jeannot K, Hocquet D, Llanes-Barakat C, Plésiat P (2007) Cumulative effects of several nonenzymatic mechanisms on the resistance of Pseudomonas aeruginosa to aminoglycosides. Antimicrob Agents Chemother 51:1016–1021. doi:10.1128/AAC.00704-06
Lau CH, Fraud S, Jones M, Peterson SN, Poole K (2012) Reduced expression of the rplU-rpmA ribosomal protein operon in mexXY-expressing pan-aminoglycoside-resistant mutants of Pseudomonas aeruginosa. Antimicrob Agents Chemother 56:5171–5179. doi:10.1128/AAC.00846-12
Martha B, Croisier D, Durand D, Hocquet D, Plésiat P, Piroth L, Portier H, Chavanet P (2006) In-vivo impact of the MexXY efflux system on aminoglycoside efficacy in an experimental model of Pseudomonas aeruginosa pneumonia treated with tobramycin. Clin Microbiol Infect 12:426–432. doi:10.1111/j.1469-0691.2006.01371.x
Bruchmann S, Dotsch A, Nouri B, Chaberny IF, Haussler S (2013) Quantitative contributions of target alteration and decreased drug accumulation to Pseudomonas aeruginosa fluoroquinolone resistance. Antimicrob Agents Chemother 57:1361–1368. doi:10.1128/AAC.01581-12
Schwartz T, Armant O, Bretschneider N, Hahn A, Kirchen S, Seifert M, Dotsch A (2015) Whole genome and transcriptome analyses of environmental antibiotic sensitive and multi-resistant Pseudomonas aeruginosa isolates exposed to waste water and tap water. Microb Biotechnol 8:116–130. doi:10.1111/1751-7915.12156
Masuda N, Gotoh N, Ohya S, Nishino T (1996) Quantitative correlation between susceptibility and OprJ production in NfxB mutants of Pseudomonas aeruginosa. Antimicrob Agents Chemother 40:909–913
Gotoh N, Tsujimoto H, Tsuda M, Okamoto K, Nomura A, Wada T, Nakahashi M, Nishino T (1998) Characterization of the MexC-MexD-OprJ multidrug efflux system in ΔmexA-mexB-oprM mutants of Pseudomonas aeruginosa. Antimicrob Agents Chemother 42:1938–1943
Masuda N, Sakagawa E, Ohya S, Gotoh N, Nishino T (2001) Hypersusceptibility of the Pseudomonas aeruginosa nfxB mutant to β-lactams due to reduced expression of the AmpC β-lactamase. Antimicrob Agents Chemother 45:1284–1286. doi:10.1128/AAC.45.4.1284-1286.2001
Wolter DJ, Hanson ND, Lister PD (2005) AmpC and OprD are not involved in the mechanism of imipenem hypersusceptibility among Pseudomonas aeruginosa isolates overexpressing the MexCD-OprJ efflux pump. Antimicrob Agents Chemother 49:4763–4766. doi:10.1128/AAC.49.11.4763-4766.2005
Li X-Z, Nikaido H (2004) Efflux-mediated drug resistance in bacteria. Drugs 64:159–204. doi:10.2165/00003495-200464020-00004
Rodriguez-Martinez JM, Poirel L, Nordmann P (2009) Molecular epidemiology and mechanisms of carbapenem resistance in Pseudomonas aeruginosa. Antimicrob Agents Chemother 53:4783–4788. doi:10.1128/AAC.00574-09
Terzi HA, Kulah C, Ciftci IH (2014) The effects of active efflux pumps on antibiotic resistance in Pseudomonas aeruginosa. World J Microbiol Biotechnol 30:2681–2687. doi:10.1007/s11274-014-1692-2
Shigemura K, Osawa K, Kato A, Tokimatsu I, Arakawa S, Shirakawa T, Fujisawa M (2015) Association of overexpression of efflux pump genes with antibiotic resistance in Pseudomonas aeruginosa strains clinically isolated from urinary tract infection patients. J Antibiot (Tokyo) 68:568–572. doi:10.1038/ja.2015.34
Reinhardt A, Kohler T, Wood P, Rohner P, Dumas JL, Ricou B, van Delden C (2007) Development and persistence of antimicrobial resistance in Pseudomonas aeruginosa: a longitudinal observation in mechanically ventilated patients. Antimicrob Agents Chemother 51:1341–1350. doi:10.1128/AAC.01278-06
Dougherty TJ, Nayar A, Newman JV, Hopkins S, Stone GG, Johnstone M, Shapiro AB, Cronin M et al (2014) NBTI 5463 is a novel bacterial type II topoisomerase inhibitor with activity against Gram-negative bacteria and in vivo efficacy. Antimicrob Agents Chemother 58:2657–2664. doi:10.1128/AAC.02778-13
Wolter DJ, Black JA, Lister PD, Hanson ND (2009) Multiple genotypic changes in hypersusceptible strains of Pseudomonas aeruginosa isolated from cystic fibrosis patients do not always correlate with the phenotype. J Antimicrob Chemother 64:294–300. doi:10.1093/jac/dkp185
Stickland HG, Davenport PW, Lilley KS, Griffin JL, Welch M (2010) Mutation of nfxB causes global changes in the physiology and metabolism of Pseudomonas aeruginosa. J Proteome Res 9:2957–2967. doi:10.1021/pr9011415
Martinez-Ramos I, Mulet X, Moya B, Barbier M, Oliver A, Alberti S (2014) Overexpression of MexCD-OprJ reduces Pseudomonas aeruginosa virulence by increasing its susceptibility to complement-mediated killing. Antimicrob Agents Chemother 58:2426–2429. doi:10.1128/AAC.02012-13
Abdel Malek SM, Badran YR (2010) Pseudomonas aeruginosa PAO1 adapted to 2-phenoxyethanol shows cross-resistance to dissimilar biocides and increased susceptibility to antibiotics. Folia Microbiol 55:588–592. doi:10.1007/s12223-010-0094-6
Chuanchuen R, Beinlich K, Hoang TT, Becher A, Karkhoff-Schweizer RR, Schweizer HP (2001) Cross-resistance between triclosan and antibiotics in Pseudomonas aeruginosa is mediated by multidrug efflux pumps: exposure of a susceptible mutant strain to triclosan selects nfxB mutants overexpressing MexCD-OprJ. Antimicrob Agents Chemother 45:428–432. doi:10.1128/AAC.45.2.428-432.2001
Chuanchuen R, Karkhoff-Schweizer RR, Schweizer HP (2003) High-level triclosan resistance in Pseudomonas aeruginosa is solely a result of efflux. Am J Infect Control 31:124–127. doi:10.1067/mic.2003.11
D’Arezzo S, Lanini S, Puro V, Ippolito G, Visca P (2012) High-level tolerance to triclosan may play a role in Pseudomonas aeruginosa antibiotic resistance in immunocompromised hosts: evidence from outbreak investigation. BMC Res Notes 5:43. doi:10.1186/1756-0500-5-43
Ochs MM, McCusker MP, Bains M, Hancock RE (1999) Negative regulation of the Pseudomonas aeruginosa outer membrane porin OprD selective for imipenem and basic amino acids. Antimicrob Agents Chemother 43:1085–1090
Fukuda H, Hosaka M, Iyobe S, Gotoh N, Nishino T, Hirai K (1995) nfxC-type quinolone resistance in a clinical isolate of Pseudomonas aeruginosa. Antimicrob Agents Chemother 39:790–792. doi:10.1128/AAC.39.3.790
Rådberg G, Nilsson LE, Svensson S (1990) Development of quinolone-imipenem cross resistance in Pseudomonas aeruginosa during exposure to ciprofloxacin. Antimicrob Agents Chemother 34:2142–2147. doi:10.1128/AAC.34.11.2142
Aubert G, Pozzetto B, Dorche G (1992) Emergence of quinolone-imipenem cross-resistance in Pseudomonas aeruginosa after fluoroquinolone therapy. J Antimicrob Chemother 29:307–312. doi:10.1093/jac/29.3.307
Bubonja-Sonje M, Matovina M, Skrobonja I, Bedenic B, Abram M (2015) Mechanisms of carbapenem resistance in multidrug-resistant clinical isolates of Pseudomonas aeruginosa from a Croatian Hospital. Microb Drug Resist 21:261–269. doi:10.1089/mdr.2014.0172
Aendekerk S, Ghysels B, Cornelis P, Baysse C (2002) Characterization of a new efflux pump, MexGHI-OpmD, from Pseudomonas aeruginosa that confers resistance to vanadium. Microbiology 148:2371–2381. doi:10.1099/00221287-148-8-2371
Weeks JW, Nickels LM, Ntreh AT, Zgurskaya HI (2015) Non-equivalent roles of two periplasmic subunits in the function and assembly of triclosan pump TriABC from Pseudomonas aeruginosa. Mol Microbiol 98:343–356. doi:10.1111/mmi.13124
Dieppois G, Ducret V, Caille O, Perron K (2012) The transcriptional regulator CzcR modulates antibiotic resistance and quorum sensing in Pseudomonas aeruginosa. PLoS One 7: e38148. doi:10.1371/journal.pone.0038148
Li X-Z, Nikaido H (2009) Efflux-mediated drug resistance in bacteria: an update. Drugs 69:1555–1623. doi:10.2165/11317030-000000000-00000
Evans K, Adewoye L, Poole K (2001) MexR repressor of the mexAB-oprM multidrug efflux operon of Pseudomonas aeruginosa: identification of MexR binding sites in the mexA-mexR intergenic region. J Bacteriol 183:807–812. doi:10.1128/JB.183.3.807-812.2001
Lim D, Poole K, Strynadka NC (2002) Crystal structure of the MexR repressor of the mexRAB-oprM multidrug efflux operon of Pseudomonas aeruginosa. J Biol Chem 277:29253–29259
Chen H, Hu J, Chen PR, Lan L, Li Z, Hicks LM, Dinner AR, He C (2008) The Pseudomonas aeruginosa multidrug efflux regulator MexR uses an oxidation-sensing mechanism. Proc Natl Acad Sci U S A 105:13586–13591. doi:10.1073/pnas.0803391105
Chen H, Yi C, Zhang J, Zhang W, Ge Z, Yang CG, He C (2010) Structural insight into the oxidation-sensing mechanism of the antibiotic resistance of regulator MexR. EMBO Rep 11:685–690. doi:10.1038/embor.2010.96
Chang W, Small DA, Toghrol F, Bentley WE (2005) Microarray analysis of Pseudomonas aeruginosa reveals induction of pyocin genes in response to hydrogen peroxide. BMC Genomics 6:115. doi:10.1186/1471-2164-6-115
Salunkhe P, Topfer T, Buer J, Tummler B (2005) Genome-wide transcriptional profiling of the steady-state response of Pseudomonas aeruginosa to hydrogen peroxide. J Bacteriol 187:2565–2572. doi:10.1128/JB.187.8.2565-2572.2005
Cummins J, Reen FJ, Baysse C, Mooij MJ, O’Gara F (2009) Subinhibitory concentrations of the cationic antimicrobial peptide colistin induce the pseudomonas quinolone signal in Pseudomonas aeruginosa. Microbiology 155:2826–2837. doi:10.1099/mic.0.025643-0
Whiteley M, Bangera MG, Bumgarner RE, Parsek MR, Teitzel GM, Lory S, Greenberg EP (2001) Gene expression in Pseudomonas aeruginosa biofilms. Nature 413:860–864. doi:10.1038/35101627
Daigle DM, Cao L, Fraud S, Wilke MS, Pacey A, Klinoski R, Strynadka NC, Dean CR et al (2007) Protein modulator of multidrug efflux gene expression in Pseudomonas aeruginosa. J Bacteriol 189:5441–5451. doi:10.1128/JB.00543-07
Wilke MS, Heller M, Creagh AL, Haynes CA, McIntosh LP, Poole K, Strynadka NC (2008) The crystal structure of MexR from Pseudomonas aeruginosa in complex with its antirepressor ArmR. Proc Natl Acad Sci U S A 105:14832–14837. doi:10.1073/pnas.0805489105
Ghosh S, Cremers CM, Jakob U, Love NG (2011) Chlorinated phenols control the expression of the multidrug resistance efflux pump MexAB-OprM in Pseudomonas aeruginosa by interacting with NalC. Mol Microbiol 79:1547–1556. doi:10.1111/j.1365-2958.2011.07544.x
Starr LM, Fruci M, Poole K (2012) Pentachlorophenol induction of the Pseudomonas aeruginosa mexAB-oprM efflux operon: involvement of repressors NalC and MexR and the antirepressor ArmR. PLoS One 7: e32684. doi:10.1371/journal.pone.0032684
Chen W, Wang D, Zhou W, Sang H, Liu X, Ge Z, Zhang J, Lan L et al (2016) Novobiocin binding to NalD induces the expression of the MexAB-OprM pump in Pseudomonas aeruginosa. Mol Microbiol 100:749–758. doi:10.1111/mmi.13346
Tomás M, Doumith M, Warner M, Turton JF, Beceiro A, Bou G, Livermore DM, Woodford N (2010) Efflux pumps, OprD porin, AmpC β-lactamase, and multiresistance in Pseudomonas aeruginosa isolates from cystic fibrosis patients. Antimicrob Agents Chemother 54:2219–2224. doi:10.1128/AAC.00816-09
Sánchez P, Rojo F, Martínez JL (2002) Transcriptional regulation of mexR, the repressor of Pseudomonas aeruginosa mexAB-oprM multidrug efflux pump. FEMS Microbiol Lett 207:63–68. doi:10.1111/j.1574-6968.2002.tb11029.x
Jimenez PN, Koch G, Thompson JA, Xavier KB, Cool RH, Quax WJ (2012) The multiple signaling systems regulating virulence in Pseudomonas aeruginosa. Microbiol Mol Biol Rev 76:46–65. doi:10.1128/MMBR.05007-11
Pearson JP, Pesci EC, Iglewski BH (1997) Roles of Pseudomonas aeruginosa las and rhl quorum-sensing systems in control of elastase and rhamnolipid biosynthesis genes. J Bacteriol 179:5756–5767
Lee J, Wu J, Deng Y, Wang J, Wang C, Wang J, Chang C, Dong Y et al (2013) A cell-cell communication signal integrates quorum sensing and stress response. Nat Chem Biol 9:339–343. doi:10.1038/nchembio.1225
Evans K, Passador L, Srikumar R, Tsang E, Nezezon J, Poole K (1998) Influence of the MexAB-OprM multidrug efflux system on quorum sensing in Pseudomonas aeruginosa. J Bacteriol 180:5443–5447
Sawada I, Maseda H, Nakae T, Uchiyama H, Nomura N (2004) A quorum-sensing autoinducer enhances the mexAB-oprM efflux-pump expression without the MexR-mediated regulation in Pseudomonas aeruginosa. Microbiol Immunol 48:435–439. doi:10.1111/j.1348-0421.2004.tb03533.x
Balasubramanian D, Schneper L, Merighi M, Smith R, Narasimhan G, Lory S, Mathee K (2012) The regulatory repertoire of Pseudomonas aeruginosa AmpC β-lactamase regulator AmpR includes virulence genes. PLoS One 7: e34067. doi:10.1371/journal.pone.0034067
Sugimura M, Maseda H, Hanaki H, Nakae T (2008) Macrolide antibiotic-mediated downregulation of MexAB-OprM efflux pump expression in Pseudomonas aeruginosa. Antimicrob Agents Chemother 52:4141–4144. doi:10.1128/aac.00511-08
Pamp SJ, Gjermansen M, Johansen HK, Tolker-Nielsen T (2008) Tolerance to the antimicrobial peptide colistin in Pseudomonas aeruginosa biofilms is linked to metabolically active cells, and depends on the pmr and mexAB-oprM genes. Mol Microbiol 68:223–240. doi:10.1111/j.1365-2958.2008.06152.x
Liao J, Sauer K (2012) The MerR-like transcriptional regulator BrlR contributes to Pseudomonas aeruginosa biofilm tolerance. J Bacteriol 194:4823–4836. doi:10.1128/JB.00765-12
Chambers JR, Liao J, Schurr MJ, Sauer K (2014) BrlR from Pseudomonas aeruginosa is a c-di-GMP-responsive transcription factor. Mol Microbiol 92:471–487. doi:10.1111/mmi.12562
Gupta K, Marques CN, Petrova OE, Sauer K (2013) Antimicrobial tolerance of Pseudomonas aeruginosa biofilms is activated during an early developmental stage and requires the two-component hybrid SagS. J Bacteriol 195:4975–4987. doi:10.1128/JB.00732-13
Gupta K, Liao J, Petrova OE, Cherny KE, Sauer K (2014) Elevated levels of the second messenger c-di-GMP contribute to antimicrobial resistance of Pseudomonas aeruginosa. Mol Microbiol 92:488–506. doi:10.1111/mmi.12587
De Kievit TR, Parkins MD, Gillis RJ, Srikumar R, Ceri H, Poole K, Iglewski BH, Storey DG (2001) Multidrug efflux pumps: expression patterns and contribution to antibiotic resistance in Pseudomonas aeruginosa biofilms. Antimicrob Agents Chemother 45:1761–1770. doi:10.1128/AAC.45.6.1761-1770.2001
Matsuo Y, Eda S, Gotoh N, Yoshihara E, Nakae T (2004) MexZ-mediated regulation of mexXY multidrug efflux pump expression in Pseudomonas aeruginosa by binding on the mexZ-mexX intergenic DNA. FEMS Microbiol Lett 238:23–28. doi:10.1111/j.1574-6968.2004.tb09732.x
Yamamoto M, Ueda A, Kudo M, Matsuo Y, Fukushima J, Nakae T, Kaneko T, Ishigatsubo Y (2009) Role of MexZ and PA5471 in transcriptional regulation of mexXY in Pseudomonas aeruginosa. Microbiology 155:3312–3321. doi:10.1099/mic.0.028993-0
Cuthbertson L, Nodwell JR (2013) The TetR family of regulators. Microbiol Mol Biol Rev 77:440–475. doi:10.1128/MMBR.00018-13
Morita Y, Sobel ML, Poole K (2006) Antibiotic inducibility of the MexXY multidrug efflux system of Pseudomonas aeruginosa: involvement of the antibiotic-inducible PA5471 gene product. J Bacteriol 188:1847–1855. doi:10.1128/JB.188.5.1847-1855.2006
Hay T, Fraud S, Lau CH, Gilmour C, Poole K (2013) Antibiotic inducibility of the mexXY multidrug efflux operon of Pseudomonas aeruginosa: involvement of the MexZ anti-repressor ArmZ. PLoS One 8: e56858. doi:10.1371/journal.pone.0056858
Yano R, Nagai H, Shiba K, Yura T (1990) A mutation that enhances synthesis of σ32 and suppresses temperature-sensitive growth of the rpoH15 mutant of Escherichia coli. J Bacteriol 172:2124–2130
Li K, Xu C, Jin Y, Sun Z, Liu C, Shi J, Chen G, Chen R et al (2013) SuhB is a regulator of multiple virulence genes and essential for pathogenesis of Pseudomonas aeruginosa. mBio 4:e00419–13. doi:10.1128/mBio.00419-13
Fernandez L, Gooderham WJ, Bains M, McPhee JB, Wiegand I, Hancock RE (2010) Adaptive resistance to the “last hope” antibiotics polymyxin B and colistin in Pseudomonas aeruginosa is mediated by the novel two-component regulatory system ParR-ParS. Antimicrob Agents Chemother 54:3372–3382. doi:10.1128/AAC.00242-10
Poole K, Gilmour C, Farha MA, Mullen E, Lau CH, Brown ED (2016) Potentiation of aminoglycoside activity in Pseudomonas aeruginosa by targeting the AmgRS envelope stress-responsive two-component system. Antimicrob Agents Chemother 60:3509–3518. doi:10.1128/AAC.03069-15
McLaughlin HP, Caly DL, McCarthy Y, Ryan RP, Dow JM (2012) An orphan chemotaxis sensor regulates virulence and antibiotic tolerance in the human pathogen Pseudomonas aeruginosa. PLoS One 7:e42205. doi:10.1371/journal.pone.0042205
Purssell A, Fruci M, Mikalauskas A, Gilmour C, Poole K (2015) EsrC, an envelope stress-regulated repressor of the mexCD-oprJ multidrug efflux operon in Pseudomonas aeruginosa. Environ Microbiol 17:186–198. doi:10.1111/1462-2920.12602
Shiba T, Ishiguro K, Takemoto N, Koibuchi H, Sugimoto K (1995) Purification and characterization of the Pseudomonas aeruginosa NfxB protein, the negative regulator of the nfxB gene. J Bacteriol 177:5872–5877
Monti MR, Morero NR, Miguel V, Argarana CE (2013) nfxB as a novel target for analysis of mutation spectra in Pseudomonas aeruginosa. PLoS One 8:e66236. doi:10.1371/journal.pone.0066236
Mandsberg LF, Ciofu O, Kirkby N, Christiansen LE, Poulsen HE, Hoiby N (2009) Antibiotic resistance in Pseudomonas aeruginosa strains with increased mutation frequency due to inactivation of the DNA oxidative repair system. Antimicrob Agents Chemother 53:2483–2491. doi:10.1128/AAC.00428-08
Strempel N, Neidig A, Nusser M, Geffers R, Vieillard J, Lesouhaitier O, Brenner-Weiss G, Overhage J (2013) Human host defense peptide LL-37 stimulates virulence factor production and adaptive resistance in Pseudomonas aeruginosa. PLoS One 8: e82240. doi:10.1371/journal.pone.0082240
Nde CW, Jang HJ, Toghrol F, Bentley WE (2009) Global transcriptomic response of Pseudomonas aeruginosa to chlorhexidine diacetate. Environ Sci Technol 43:8406–8415. doi:10.1021/es9015475
Köhler T, Epp SF, Curty LK, Pechere JC (1999) Characterization of MexT, the regulator of the MexE-MexF-OprN multidrug efflux system of Pseudomonas aeruginosa. J Bacteriol 181:6300–6305
Tian ZX, Mac Aogain M, O’Connor HF, Fargier E, Mooij MJ, Adams C, Wang YP, O’Gara F (2009) MexT modulates virulence determinants in Pseudomonas aeruginosa independent of the MexEF-OprN efflux pump. Microb Pathog 47:237–241. doi:10.1016/j.micpath.2009.08.003
Tian ZX, Fargier E, Mac Aogain M, Adams C, Wang YP, O’Gara F (2009) Transcriptome profiling defines a novel regulon modulated by the LysR-type transcriptional regulator MexT in Pseudomonas aeruginosa. Nucleic Acids Res 37:7546–7559. doi:10.1093/nar/gkp828
Lister PD, Wolter DJ, Hanson ND (2009) Antibacterial-resistant Pseudomonas aeruginosa: clinical impact and complex regulation of chromosomally encoded resistance mechanisms. Clin Microbiol Rev 22:582–610. doi:10.1128/CMR.00040-09
Sobel ML, Neshat S, Poole K (2005) Mutations in PA2491 (mexS) promote MexT-dependent mexEF-oprN expression and multidrug resistance in a clinical strain of Pseudomonas aeruginosa. J Bacteriol 187:1246–1253. doi:10.1128/JB.187.4.1246-1253.2005
Fetar H, Gilmour C, Klinoski R, Daigle DM, Dean CR, Poole K (2011) mexEF-oprN multidrug efflux operon of Pseudomonas aeruginosa: regulation by the MexT activator in response to nitrosative stress and chloramphenicol. Antimicrob Agents Chemother 55:508–514. doi:10.1128/AAC.00830-10
Jin Y, Yang H, Qiao M, Jin S (2011) MexT regulates the type III secretion system through MexS and PtrC in Pseudomonas aeruginosa. J Bacteriol 193:399–410. doi:10.1128/JB.01079-10
Uwate M, Ichise YK, Shirai A, Omasa T, Nakae T, Maseda H (2013) Two routes of MexS-MexT-mediated regulation of MexEF-OprN and MexAB-OprM efflux pump expression in Pseudomonas aeruginosa. Microbiol Immunol 57:263–272. doi:10.1111/1348-0421.12032
Frisk A, Schurr JR, Wang G, Bertucci DC, Marrero L, Hwang SH, Hassett DJ, Schurr MJ (2004) Transcriptome analysis of Pseudomonas aeruginosa after interaction with human airway epithelial cells. Infect Immunol 72:5433–5438. doi:10.1128/IAI.72.9.5433-5438.2004
Richardot C, Juarez P, Jeannot K, Patry I, Plésiat P, Llanes C (2016) Amino acid substitutions account for most MexS alterations in clinical nfxC mutants of Pseudomonas aeruginosa. Antimicrob Agents Chemother 60:2302–2310. doi:10.1128/AAC.02622-15
Vallet I, Diggle SP, Stacey RE, Camara M, Ventre I, Lory S, Lazdunski A, Williams P et al (2004) Biofilm formation in Pseudomonas aeruginosa: fimbrial cup gene clusters are controlled by the transcriptional regulator MvaT. J Bacteriol 186:2880–2890. doi:10.1128/JB.186.9.2880-2890.2004
Castang S, McManus HR, Turner KH, Dove SL (2008) H-NS family members function coordinately in an opportunistic pathogen. Proc Natl Acad Sci U S A 105:18947–18952. doi:10.1073/pnas.0808215105
Castang S, Dove SL (2012) Basis for the essentiality of H-NS family members in Pseudomonas aeruginosa. J Bacteriol 194:5101–5109. doi:10.1128/JB.00932-12
Köhler T, van Delden C, Curty LK, Hamzehpour MM, Pechère JC (2001) Overexpression of the MexEF-OprN multidrug efflux system affects cell-to-cell signaling in Pseudomonas aeruginosa. J Bacteriol 183:5213–5222. doi:10.1128/JB.183.18.5213-5222.2001
Olivares J, Alvarez-Ortega C, Linares JF, Rojo F, Köhler T, Martínez JL (2012) Overproduction of the multidrug efflux pump MexEF-OprN does not impair Pseudomonas aeruginosa fitness in competition tests, but produces specific changes in bacterial regulatory networks. Environ Microbiol 14:1968–1981. doi:10.1111/j.1462-2920.2012.02727.x
Lee J, Attila C, Cirillo SL, Cirillo JD, Wood TK (2009) Indole and 7-hydroxyindole diminish Pseudomonas aeruginosa virulence. Microb Biotechnol 2:75–90. doi:10.1111/j.1751-7915.2008.00061.x
Mushtaq S, Ge Y, Livermore DM (2004) Doripenem versus Pseudomonas aeruginosa in vitro: activity against characterized isolates, mutants, and transconjugants and resistance selection potential. Antimicrob Agents Chemother 48:3086–3092. doi:10.1128/AAC.48.8.3086-3092.2004
Queenan AM, Shang W, Bush K, Flamm RK (2010) Differential selection of single-step AmpC or efflux mutants of Pseudomonas aeruginosa by using cefepime, ceftazidime, or ceftobiprole. Antimicrob Agents Chemother 54:4092–4097. doi:10.1128/AAC.00060-10
Winkler ML, Papp-Wallace KM, Hujer AM, Domitrovic TN, Hujer KM, Hurless KN, Tuohy M, Hall G et al (2015) Unexpected challenges in treating multidrug-resistant Gram-negative bacteria: resistance to ceftazidime-avibactam in archived isolates of Pseudomonas aeruginosa. Antimicrob Agents Chemother 59:1020–1029. doi:10.1128/AAC.04238-14
Akama H, Matsuura T, Kashiwagi S, Yoneyama H, Narita S, Tsukihara T, Nakagawa A, Nakae T (2004) Crystal structure of the membrane fusion protein, MexA, of the multidrug transporter in Pseudomonas aeruginosa. J Biol Chem 279:25939–25942. doi:10.1074/jbc.C400164200
Sennhauser G, Bukowska MA, Briand C, Grutter MG (2009) Crystal structure of the multidrug exporter MexB from Pseudomonas aeruginosa. J Mol Biol 389:134–145. doi:10.1016/j.jmb.2009.04.001
Nakashima R, Sakurai K, Yamasaki S, Hayashi K, Nagata C, Hoshino K, Onodera Y, Nishino K et al (2013) Structural basis for the inhibition of bacterial multidrug exporters. Nature 500:102–106. doi:10.1038/nature12300
Du D, Wang Z, James NR, Voss JE, Klimont E, Ohene-Agyei T, Venter H, Chiu W et al (2014) Structure of the AcrAB-TolC multidrug efflux pump. Nature 509:512–515. doi:10.1038/nature13205
Daury L, Orange F, Taveau JC, Verchere A, Monlezun L, Gounou C, Marreddy RK, Picard M et al (2016) Tripartite assembly of RND multidrug efflux pumps. Nat Commun 7:10731. doi:10.1038/ncomms10731
Yonehara R, Yamashita E, Nakagawa A (2016) Crystal structures of OprN and OprJ, outer membrane factors of multidrug tripartite efflux pumps of Pseudomonas aeruginosa. Proteins 84:759–769. doi:10.1002/prot.25022
Takeda S, Nakai T, Wakai Y, Ikeda F, Hatano K (2007) In vitro and in vivo activities of a new cephalosporin, FR264205, against Pseudomonas aeruginosa. Antimicrob Agents Chemother 51:826–830. doi:10.1128/AAC.00860-06
Hong MC, Hsu DI, Bounthavong M (2013) Ceftolozane/tazobactam: a novel antipseudomonal cephalosporin and β-lactamase-inhibitor combination. Infect Drug Resist 6:215–223. doi:10.2147/IDR.S36140
Eguchi K, Ueda Y, Kanazawa K, Sunagawa M, Gotoh N (2007) The mode of action of 2-(thiazol-2-ylthio)-1β-methylcarbapenems against Pseudomonas aeruginosa: the impact of outer membrane permeability and the contribution of MexAB-OprM efflux system. J Antibiot (Tokyo) 60:129–135. doi:10.1038/ja.2007.12
Koga T, Masuda N, Kakuta M, Namba E, Sugihara C, Fukuoka T (2008) Potent in vitro activity of tomopenem (CS-023) against methicillin-resistant Staphylococcus aureus and Pseudomonas aeruginosa. Antimicrob Agents Chemother 52:2849–2854. doi:10.1128/AAC.00413-08
Koga T, Sugihara C, Kakuta M, Masuda N, Namba E, Fukuoka T (2009) Affinity of tomopenem (CS-023) for penicillin-binding proteins in Staphylococcus aureus, Escherichia coli, and Pseudomonas aeruginosa. Antimicrob Agents Chemother 53:1238–1241. doi:10.1128/AAC.01433-08
Rieg S, Huth A, Kalbacher H, Kern WV (2009) Resistance against antimicrobial peptides is independent of Escherichia coli AcrAB, Pseudomonas aeruginosa MexAB and Staphylococcus aureus NorA efflux pumps. Int J Antimicrob Agents 33:174–176. doi:10.1016/j.ijantimicag.2008.07.032
Lomovskaya O, Warren MS, Lee A, Galazzo J, Fronko R, Lee M, Blais J, Cho D et al (2001) Identification and characterization of inhibitors of multidrug resistance efflux pumps in Pseudomonas aeruginosa: novel agents for combination therapy. Antimicrob Agents Chemother 45:105–116. doi:10.1128/AAC.45.1.105-116.2001
Sonnet P, Izard D, Mullie C (2012) Prevalence of efflux-mediated ciprofloxacin and levofloxacin resistance in recent clinical isolates of Pseudomonas aeruginosa and its reversal by the efflux pump inhibitors 1-(1-naphthylmethyl)-piperazine and phenylalanine-arginine-β-naphthylamide. Int J Antimicrob Agents 39:77–80. doi:10.1016/j.ijantimicag.2011.08.005
Mesaros N, Glupczynski Y, Avrain L, Caceres NE, Tulkens PM, Van Bambeke F (2007) A combined phenotypic and genotypic method for the detection of Mex efflux pumps in Pseudomonas aeruginosa. J Antimicrob Chemother 59:378–386. doi:10.1093/jac/dkl504
Yoshida K, Nakayama K, Ohtsuka M, Kuru N, Yokomizo Y, Sakamoto A, Takemura M, Hoshino K et al (2007) MexAB-OprM specific efflux pump inhibitors in Pseudomonas aeruginosa. Part 7: highly soluble and in vivo active quaternary ammonium analogue D13-9001, a potential preclinical candidate. Bioorg Med Chem 15:7087–7097. doi:10.1016/j.bmc.2007.07.039
Zuo Z, Weng J, Wang W (2016) Insights into the inhibitory mechanism of D13-9001 to the multidrug transporter AcrB through molecular dynamics simulations. J Phys Chem B 120:2145–2154. doi:10.1021/acs.jpcb.5b11942
Hirakata Y, Kondo A, Hoshino K, Yano H, Arai K, Hirotani A, Kunishima H, Yamamoto N et al (2009) Efflux pump inhibitors reduce the invasiveness of Pseudomonas aeruginosa. Int J Antimicrob Agents 34:343–346. doi:10.1016/j.ijantimicag.2009.06.007
Opperman TJ, Nguyen ST (2015) Recent advances toward a molecular mechanism of efflux pump inhibition. Front Microbiol 6:421. doi:10.3389/fmicb.2015.00421
Kuete V, Alibert-Franco S, Eyong KO, Ngameni B, Folefoc GN, Nguemeving JR, Tangmouo JG, Fotso GW et al (2011) Antibacterial activity of some natural products against bacteria expressing a multidrug-resistant phenotype. Int J Antimicrob Agents 37:156–161. doi:10.1016/j.ijantimicag.2010.10.020
Fadli M, Chevalier J, Saad A, Mezrioui NE, Hassani L, Pagès JM (2011) Essential oils from Moroccan plants as potential chemosensitisers restoring antibiotic activity in resistant Gram-negative bacteria. Int J Antimicrob Agents 38:325–330. doi:10.1016/j.ijantimicag.2011.05.005
Aparna V, Dineshkumar K, Mohanalakshmi N, Velmurugan D, Hopper W (2014) Identification of natural compound inhibitors for multidrug efflux pumps of Escherichia coli and Pseudomonas aeruginosa using in silico high-throughput virtual screening and in vitro validation. PLoS One 9: e101840. doi:10.1371/journal.pone.0101840
Negi N, Prakash P, Gupta ML, Mohapatra TM (2014) Possible role of curcumin as an efflux pump inhibitor in multidrug resistant clinical isolates of Pseudomonas aeruginosa. J Clin Diagn Res 8:DC04–DC07. doi:10.7860/JCDR/2014/8329.4965
Whalen KE, Poulson-Ellestad KL, Deering RW, Rowley DC, Mincer TJ (2015) Enhancement of antibiotic activity against multidrug-resistant bacteria by the efflux pump inhibitor 3,4-dibromopyrrole-2,5-dione isolated from a Pseudoalteromonas sp. J Nat Prod 78:402–412. doi:10.1021/np500775e
Wu CM, Cao JL, Zheng MH, Ou Y, Zhang L, Zhu XQ, Song JX (2008) Effect and mechanism of andrographolide on the recovery of Pseudomonas aeruginosa susceptibility to several antibiotics. J Int Med Res 36:178–186
Wang H, Meng J, Jia M, Ma X, He G, Yu J, Wang R, Bai H et al (2010) oprM as a new target for reversion of multidrug resistance in Pseudomonas aeruginosa by antisense phosphorothioate oligodeoxynucleotides. FEMS Immunol Med Microbiol 60:275–282. doi:10.1111/j.1574-695X.2010.00742.x
Rees VE, Bulitta JB, Nation RL, Tsuji BT, Sorgel F, Landersdorfer CB (2015) Shape does matter: short high-concentration exposure minimizes resistance emergence for fluoroquinolones in Pseudomonas aeruginosa. J Antimicrob Chemother 70:818–826. doi:10.1093/jac/dku437
Chan BK, Sistrom M, Wertz JE, Kortright KE, Narayan D, Turner PE (2016) Phage selection restores antibiotic sensitivity in MDR Pseudomonas aeruginosa. Sci Rep 6: 26717. doi:10.1038/srep26717
Sakhtah H, Koyama L, Zhang Y, Morales DK, Fields BL, Price-Whelan A, Hogan DA, Shepard K et al (2016) The Pseudomonas aeruginosa efflux pump MexGHI-OpmD transports a natural phenazine that controls gene expression and biofilm development. Proc Natl Acad Sci U S A 113:E3538–3547. doi:10.1073/pnas.1600424113
Anandapadamanaban M, Pilstål R, Andresen C, Trewhella J, Moche M, Wallner B, Sunnerhagen M (2016) Mutation-induced population shift in the MexR conformational ensemble disengages DNA binding: a novel mechanism for MarR family derepression. Structure 24:1311–1321. doi:10.1016/j.str.2016.06.008
Acknowledgments
The views expressed in this chapter do not necessarily reflect those of the authors’ affiliations, Health Canada or University of Franche-Comté. The authors thank Andrea Leclair for helpful reading of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Li, XZ., Plésiat, P. (2016). Antimicrobial Drug Efflux Pumps in Pseudomonas aeruginosa . In: Li, XZ., Elkins, C., Zgurskaya, H. (eds) Efflux-Mediated Antimicrobial Resistance in Bacteria. Adis, Cham. https://doi.org/10.1007/978-3-319-39658-3_14
Download citation
DOI: https://doi.org/10.1007/978-3-319-39658-3_14
Published:
Publisher Name: Adis, Cham
Print ISBN: 978-3-319-39656-9
Online ISBN: 978-3-319-39658-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)