Abstract
The field of evo-devo studies what, how, and why developmental patterning processes have evolved. While comparative approaches based in experimental data are essential for answering the first two types of questions, evo-devo simulations studies are critical to answer why questions. By simulating evo-devo processes, the evolutionary tape can be replayed both under the same and different conditions, enabling us to answer questions on contingency, convergence, and constraints and their roles in determining evolutionary outcomes.
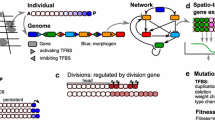
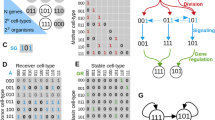
In this chapter, we describe the basic ingredients of computational models simulating evo-devo processes: gene expression regulation; cell and tissue behavior; and mutation-selection driven evolution. We describe for each of these model ingredients the choices that need to be made, e.g., whether the model simulates a one, two, or three-dimensional tissue, and how these affect computational efficiency as well as modeling outcomes. We focus on the importance of incorporating a realistic, nonlinear, and evolvable genotype-phenotype map in evo-devo simulation models.
We end with an illustration of how evo-devo models have helped answer why questions in the field of animal body plan segmentation.
Similar content being viewed by others
References
Chipman AD (2010) Parallel evolution of segmentation by co-option of ancestral gene regulatory networks. BioEssays 32(1):60–70
Cotterell J, Sharpe J (2010) An atlas of gene regulatory networks reveals multiple three-gene mechanisms for interpreting morphogen gradients. Mol Syst Biol 6(1)
Cotterell J, Robert-Moreno A, Sharpe J (2015) A local, self-organizing reaction-diffusion model can explain somite patterning in embryos. Cell Syst 1(4):257–269
François P, Hakim V, Siggia ED (2007) Deriving structure from evolution: metazoan segmentation. Mol Syst Biol 3(1)
Fujimoto K, Ishihara S, Kaneko K (2008) Network evolution of body plans. PLoS One 3(7):e2772
Graner F m c, Glazier JA (1992) Simulation of biological cell sorting using a two-dimensional extended potts model. Phys Rev Lett 69:2013–2016
Hogeweg P (2000) Evolving mechanisms of morphogenesis: on the interplay between differential adhesion and cell differentiation. J Theor Biol 203(4):317–333
Jiménez A, Munteanu A, Sharpe J (2015) Dynamics of gene circuits shapes evolvability. PNAS 112(7):2103–2108
Kohsokabe T, Kaneko K (2016) Evolution-development congruence in pattern formation dynamics: bifurcations in gene expression and regulation of networks structures. J Exp Zool B Mol Dev Evol 326(1):61–84
Marin-Riera M, Brun-Usan M, Zimm R, Välikangas T, Salazar-Ciudad I (2016) Computational modeling of development by epithelia, mesenchyme and their interactions: a unified model. Bioinformatics 32(2):219–225
Salazar-ciudad I, Jernvall J (2004) How different types of pattern formation mechanisms affect the evolution of form and development. Evol Dev 6(1):6–16
Salazar-Ciudad I, Newman SA, Solé RV (2001) Phenotypic and dynamical transitions in model genetic networks I. Emergence of patterns and genotype-phenotype relationships. Evol Dev 3(2):84–94
Sánchez L, Thieffry D (2003) Segmenting the fly embryo: a logical analysis of the pair-rule cross-regulatory module. J Theor Biol 224(4):517–537
Sánchez L, Chaouiya C, Thieffry D (2008) Segmenting the fly embryo: logical analysis of the role of the segment polarity cross-regulatory module. Int J Dev Biol 52:1059–1075
Solé RV, Salazar-Ciudad I, Garcia-Fernández J (2002) Common pattern formation, modularity and phase transitions in a gene network model of morphogenesis. Physica A 305(34):640–654
Spirov A, Holloway D (2013) Using evolutionary computations to understand the design and evolution of gene and cell regulatory networks. Methods 62(1):39–55
ten Tusscher K (2013) Mechanisms and constraints shaping the evolution of body plan segmentation. Eur Phys J E 36(5):1–12
ten Tusscher KH, Hogeweg P (2009) The role of genome and gene regulatory network canalization in the evolution of multi-trait polymorphisms and sympatric speciation. BMC Evol Biol 9(1):159
ten Tusscher KH, Hogeweg P (2011) Evolution of networks for body plan patterning; interplay of modularity, robustness and evolvability. PLoS Comput Biol 7(10):e1002208
Vroomans RMA, Hogeweg P, ten Tusscher KHWJ (2015) Segment-specific adhesion as a driver of convergent extension. PLoS Comput Biol 11(2):1–24
Vroomans RMA, Hogeweg P, ten Tusscher KHWJ (2016) In silico evo-devo: reconstructing stages in the evolution of animal segmentation. EvoDevo 7(1):14
Wagner A (1996) Does evolutionary plasticity evolve? Evolution 50(3):1008–1023
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Section Editor information
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG
About this entry
Cite this entry
Vroomans, R.M.A., ten Tusscher, K.H.W.J. (2018). Modeling Evolution of Developmental Gene Regulatory Networks. In: Nuno de la Rosa, L., Müller, G. (eds) Evolutionary Developmental Biology. Springer, Cham. https://doi.org/10.1007/978-3-319-33038-9_118-1
Download citation
DOI: https://doi.org/10.1007/978-3-319-33038-9_118-1
Received:
Accepted:
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-33038-9
Online ISBN: 978-3-319-33038-9
eBook Packages: Springer Reference Biomedicine and Life SciencesReference Module Biomedical and Life Sciences