Abstract
The chlorarachniophytes are a group of single-celled phototrophic, mixotrophic eukaryotes in marine environments. They are most common in tropical and temperate seas. The group is primarily studied due to their evolutionary history. Chlorarachniophytes acquired photosynthesis by secondary endosymbiosis, where an amoeboflagellate host took up a green algal symbiont and retained it. The symbiont is distinguished by having retained a relict nucleus, or nucleomorph, which has been intensively studied to help elucidate the process of organelle origins by endosymbiosis. Historically, work on the nucleomorph was an important clue suggesting that secondary endosymbiosis played a role in the distribution of photosynthesis and plastids in eukaryotes. More recently, a number of genomic and cell biological studies, in particular focusing on gene flow within the cell and protein targeting, have further contributed to our understanding of organelle integration during endosymbiosis. The host component is now known to be a member of the Cercozoa and can include amoeboid, flagellate, and cyst stages, various species having any combination of one or more stages in the life cycle.
Access provided by CONRICYT-eBooks. Download reference work entry PDF
Similar content being viewed by others
Keywords
Summary Classification
-
●Chlorarachniophyceae
-
●●Amorphochlora
-
●●Bigelowiella
-
●●Chlorarachnion
-
●●Cryptochlora
-
●●Gymnochlora
-
●●Lotharella
-
●●Norrisiella
-
●●Partenskyella
-
Introduction
The chlorarachniophytes are a small group of tropical to temperate marine amoeboflagellates with chlorophyll a- and b-containing chloroplasts. They have attracted the attention of biologists primarily due to their complex cell biology and evolutionary history, stemming from the fact that they acquired their green chloroplasts by secondary endosymbiosis and have retained a vestigial nucleus of the engulfed alga, now called a nucleomorph (McFadden et al. 1994; Archibald 2007). All known chlorarachniophytes are phototrophic and can possess from one to several chloroplasts, each associated with a nucleomorph. The host cells may be found as amoebae, some plasmodial with individual cells linked by a network of reticulopodia, as thick-walled coccoid cells, or as highly motile uniflagellated zoospore. In some genera, all three cell types are found, although one type is the dominant trophic stage, whereas in other genera, only two or one of the cell types have been observed (Ishida et al. 2007). The endosymbiont is known to be derived from a green alga, and the host is a member of the Cercozoa (Cavalier-Smith 1999; Rogers et al. 2007).
The type species for the group is Chlorarachnion reptans, originally described by Geitler (Geitler 1930) and also the first species to be investigated in detail by light microscopy (LM), electron microscopy (EM), and pigment composition analysis (Hibberd and Norris 1984). Bigelowiella natans has since emerged as the best-studied species, and now most of our information comes from this organism.
History of Study and Literature
For many years after the discovery of C. reptans, there remained little literature on chlorarachniophytes (Hibberd 1990). The suggestion that they originated by secondary endosymbiosis led to new interest in the group, and early work proving this and preliminary characterization of the nucleomorph both led to a surge in reports on the group, in particular on B. natans. Recent work has focused on genomics (McFadden et al. 1997a; Williams et al. 2005; Gilson et al. 2006; Rogers et al. 2007; Curtis et al. 2012), molecular evolution (Archibald et al. 2002, 2003; Takishita et al. 2005; Burki et al. 2007), protein trafficking (Rogers et al. 2004; Gile and Keeling 2008; Hirakawa and Ishida 2010; Hirakawa et al. 2009, 2010, 2011a, 2012a, b), and on the description of new species. Many genomes are now available, especially from B. natans, but the emerging model for cell biology is Amorphochlora amoebiformis, due to the creation of a transient transfection system that has been used with green fluorescent protein (GFP) markers (Hirakawa et al. 2009). Most information on the cell structure, life history, and habitat is found in the formal descriptions of the 14 species described to date (Hibberd and Norris 1984; Calderon-Saenz and Schnetter 1987; Ishida and Hara 1994; Ishida et al. 1996, 2000, 2011b; Moestrup and Sengco 2001; Dietz et al. 2003; Ota et al. 2005, 2007a, b, 2009a, b, 2011, 2012). There is a large number of review articles on chlorarachniophytes, almost all focusing on molecular biology and endosymbiosis, due to the presence of the nucleomorph and its importance to understanding genome reduction and the endosymbiotic history of plastids (McFadden and Gilson 1995; Gilson et al. 1997; Gilson and McFadden 1997, 2002; McFadden et al. 1997a; Gilson 2001; Archibald and Keeling 2002; Cavalier-Smith 2002; Archibald 2007; Ishida et al. 2007).
Characterization and Recognition
General Characteristics
Although there is much variation between members of the group, there are three common life history stages in chlorarachniophytes: amoeboid, coccoid, and zoospore. In some species all three stages have been observed, whereas in others only one or two stages have been observed, and where more than one is known, the dominant trophic stage can vary. Characteristics common to different stages are discussed here, and stage-specific characteristics will be discussed in turn.
Cells contain a single nucleus (with the exception of a giant amoeboid/coccoid stage of Gymnochlora dimorpha/Lotharella reticulosa (Ota et al. 2011, 2012)), which divides by open or semi-open mitosis. Mitochondria display typical tubular cristae and are dispersed throughout the cell. Extrusomes have been observed in several species. Cells contain numerous Golgi bodies, some associated with the pyrenoid and the pyrenoid-capping vesicle. The pyrenoid-capping vesicle is bounded by a single membrane and contains a homogeneous material that has been shown to react with antibodies specific to β-1,3-glucans, which are the primary carbohydrate storage product (McFadden et al. 1997b). The chloroplast lacks starch (Hibberd and Norris 1984), so all carbohydrate storage seems to be carried out in this form by the host. The lipid composition of chlorarachniophytes has also been examined and found to be unlike that of other algae (Leblond et al. 2005).
The chloroplast is the best-studied structure of chlorarachniophytes. All cells contain one or more bilobed, peripheral, chlorophyll a- and b-containing chloroplasts, typically with a central, inwardly projecting pyrenoid that is closely surrounded by a capping vesicle. Chloroplast lamellae are usually composed of one to three thylakoids, and a girdle lamella is absent. Each chloroplast is bounded by four membranes that may appear closely appressed or as two pairs separated by a space. The outermost membrane is derived from the endomembrane system of the Cercozoan host but is smooth and is not directly connected to the rough endoplasmic reticulum (ER), as is the case in several other algal groups with secondary plastids. The second membrane is derived from the plasma membrane of the green algal endosymbiont. The space between the outer pair and inner pair (which corresponds to the cytoplasm of the green alga) is sometimes referred to as the periplastid space or periplastidial compartment. The inner pair of membranes is derived from the chloroplast envelope. Protein targeting to the plastid has been examined in some detail and is mediated by a bipartite leader consisting of a signal peptide followed by a transit peptide-like (TPL) sequence. The signal peptide directs the protein to the endomembrane system, and the TPL directs it across the remaining three membranes (Hirakawa et al. 2009, 2010, 2012a). Proteins cross the last two membranes using a fairly conventional plastid translocon complex (TOC and TIC systems: Hirakawa et al. 2012a), but how proteins cross the membrane derived from the endosymbiont plasma membrane remains mysterious. The characteristics of the leader that mediate this process are understood, but the mechanism is unknown: currently it seems unlikely that chlorarachniophytes use symbiont ERAD-like machinery (SELMA) including Der1 proteins (Hirakawa et al. 2012a), which is used by red algal secondary plastids (Hempel et al. 2009).
The periplastid space contains a dense homogeneous matrix including many visible ribosomes equivalent in size to eukaryotic cytosolic ribosomes (Hibberd and Norris 1984; McFadden et al. 1994). The periplastid space is generally only a thin layer around most of the chloroplast, but around the base of the pyrenoid or within a wedge-shaped invagination in the pyrenoid, the space is enlarged and contains the nucleomorph, the relict nucleus of the green algal endosymbiont (Fig. 1i). The nucleomorph is small, bounded by a double membrane with pores (Hibberd and Norris 1984; Ludwig and Gibbs 1989; McFadden et al. 1994), and has been shown in all examined species to contain three small, linear chromosomes amounting to 330–1133 kbp of DNA (Gilson and McFadden 1999; Silver et al. 2007; Ishida et al. 2011a). Like the plastid, the periplastidal space also lacks sufficient nucleomorph-encoded genes for function (Gilson et al. 2006), and now a number of nucleus-encoded periplastid-targeted proteins have been identified. Direct evidence for targeting is only available for three such proteins, histones H2A and H2B (which are targeted to the nucleomorph: Hirakawa et al. 2011a) and EFL (Gile and Keeling 2008). Targeting of these proteins is mediated by a bipartite leader resembling the plastid-targeting leader, except that the TPL portion is distinguished by a net neutral/negative charge, and some proteins have a hydrophilic tail enriched with lysine and aspartic acid residues (Hirakawa et al. 2009, 2010). A large group of potentially periplastidal compartment (PPC)-targeted proteins were identified in the nuclear genome with similar characteristics (Curtis et al. 2012).
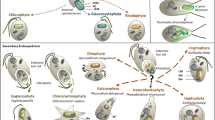
Morphology and diversity of chlorarachniophytes. (a) Transmission electron micrograph of B. natans showing the nucleomorph (nm), periplastid space with eukaryotic size ribosomes (pps), two lobes of the chloroplast (cp), and bulbous pyrenoid (p) with pyrenoid-capping vesicle (pcv). (b) Schematic diagram of chlorarachniophyte plastid. Labeled to the right are the three genome-containing compartments (host and endosymbiont nucleus and plastid – the mitochondrion is not shown), the pyrenoid (the pyrenoid-capping vesicle is not shown), and the various membranes and compartments of the complex plastid, including their evolutionary origin (abbreviations are: EM endomembrane, pps periplastid space, OM outer membrane, IM inner membrane). (c–f) Examples of the amoeboid stage: a scanning electron micrograph of G. stellata showing cell body and emerging filopodia is shown in (c), and light micrographs of amoeboid stages are shown for G. stellata (d), Lotharella amoeboformis (e), and Chlorarachnion reptans (f). (g–h) Examples of the coccoid stage from Lotharella vacuolata (g) and Chlorarachnion reptans (h) where the thickened wall is apparent. (i–j) Examples of the zoospore stage from unnamed strain (CCMP622) of the Lotharella clade (i), in which the helically coiled flagellum can be seen, and Bigelowiella natans (j). Scale bars on all LM and SEM parts are 10 μm. Scale bar on TEM scale bar is 500 nm (TEM is courtesy of Ø. Moestrup, SEM is courtesy of S. Bowser, and all differential interference contrast light micrographs are by P. Keeling)
Amoeboid Stage
Amoeboid cells (Fig. 1c, e–h) are roughly isodiametric, angular cells found in benthic environments, ranging from 8 to 20 μm (not including the filopodia). From each cell radiate several filose pseudopodia, in C. reptans, L. polymorpha, L. vaculolata, and L. reticulosa apparently fusing to form a network of reticulopodia (Hibberd and Norris 1984; Dietz et al. 2003; Ota et al. 2005, 2012) but in others remaining distinct and unconnected (Calderon-Saenz and Schnetter 1989; Ishida et al. 1996, 2000; Ota et al. 2007b). Movement on surfaces is very slow and in some species nonexistent (Ota et al. 2007a), although cytoplasmic streaming can readily be seen in the pseudopodia. Contents of filopodia are mostly restricted to microtubules and vesicular and granular material; mitochondria are the only organelles occurring in the reticulopodia. The amoeboid cells are phagotrophic, engulfing a variety of motile and nonmotile eukaryotes and prokaryotes in the pseudopodia, which may develop large ingestion vesicles. Uptake of prey species has been described for C. reptans as differentiated, and some species are taken up preferentially and digested quickly whereas others are resistant to digestion (Hibberd and Norris 1984). Most digestion takes place in pseudopodia (Hibberd and Norris 1984) but has infrequently been observed in the cell body (Ishida et al. 1996). Amoeboid cells may divide or give rise directly to zoospores or coccoid cells, depending on the species.
Walled Coccoid Stage
Coccoid cells (Fig. 1d–e) are also found in benthic environments and are spherical cells 5–15 μm in diameter with a firm wall of variable thickness composed of multiple layers (Ishida and Hara 1994; Ota et al. 2005). They are often found in older cultures where they probably act as a resting stage or cyst (Hibberd and Norris 1984; Ota et al. 2007b), but in some species, the main vegetative cell type is walled or walled amoebae with short pseudopodia can extending from the wall (Ishida and Hara 1994; Dietz et al. 2003; Ota et al. 2005, 2007a). These have more irregularly shaped chloroplasts than in the amoeboid stage, a laterally positioned nucleus, and contain a large number of vesicles with contents similar in appearance to those of the pyrenoid-capping vesicles. Coccoid cells may divide or give rise directly to the amoeboid stage and also to zoospores via a tetrad division, depending on the species.
Zoospore Stage
Zoospores (Fig. 1a–b, i) are small, planktonic, pyriform, ellipsoid, or ovoid cells ranging from 4 to 24 μm long × 3–7 μm wide. The cytoplasm often has a distinct granular appearance at the anterior end. Zoospores are uniflagellate, although ultrastructural investigation of B. natans has revealed a vestigial second basal body (Moestrup and Sengco 2001). The flagellum typically has a hair point and fine lateral hairs and is anchored by a root system consisting of a microtubular component and a second root (Hibberd and Norris 1984; Moestrup and Sengco 2001). During swimming, the flagellum is wrapped helically around the cell body within a concavity. Swimming is rapid – about 100 μm per second for C. reptans and faster for smaller cells. In some species the zoospore may become temporarily amoeboid, with the anterior end forming one or more blunt pseudopodia (Hibberd and Norris 1984; Moestrup and Sengco 2001). Zoospores may divide or give rise directly to the amoeboid or coccoid stages, depending on the species.
Reproduction and Life Cycle
The division of the nucleus, nucleomorph, and plastid has all been examined, as has the order of events in B. natans where the order of division is pyrenoid, nucleomorph, chloroplast, and finally the nucleus (Moestrup and Sengco 2001). In nuclear division, the envelope breaks down but fragments of it remain, and mitosis is otherwise not unusual (Moestrup and Sengco 2001). Separation of the daughter cells, on the other hand, can be by very unusual means, including a variation on cytoplasmic streaming in B. longifolia and L. vacuolata (Ota et al. 2005, 2007b). The nucleomorph divides amitotically: no chromosomal condensation or microtubules have been observed. Rather, the inner membrane invaginates and joins to form a barrier, after which the outer membrane invaginates and the two daughter nucleomorphs are separated (Ludwig and Gibbs 1989). Sexual reproduction is poorly understood in chlorarachniophytes, but gametes and sexual reproduction have been reported in zoospores of C. reptans (Grell 1990) and amoebae of Cryptochlora perforans (Calderon-Saenz and Schnetter 1989).
As described above, there are three main life history stages, and in some species all three are known whereas in others one or two are absent. Life history stages have been documented from members of all eight genera, although direct observation of transformations between the various stages is lacking for some species. Available data on which life history stages are present in which genera are summarized in Fig. 2 and Table 1, and see Ishida et al. (2007) for review.
Life history stages common to chlorarachniophytes. Amoeboid, coccoid, and zoospore states are illustrated, with arrows representing transitions (between stages) or vegetative division of a stage. Initials adjacent to stages or within vegetative division loops represent the genus and species where that stage or that vegetative division has been observed (see Classification section for all full genus and species names). G. stellata is also known to form a plasmodial form that divides through synchronous cytokinesis, shown parallel to the transition between amoeboid and coccoid stages. All transitions have not been observed in all species, but each transition is known in at least one species
Classification
Chlorarachniophytes are currently classified as members of the Cercozoa, as described below. Currently only 14 species and one variety have been formally described, distributed in 8 genera (Table 1). Classification schemes have relied on pyrenoid shape, location of the nucleomorph, presence or absence of different life history stages, and molecular phylogeny (see Ishida et al. 2007 for review).
Chlorarachnion reptans (Hibberd and Norris 1984) is the type species and only described species of Chlorarachnion. It includes all three cell types in its life history and is also distinguished by the location of its nucleomorph within the pyrenoid slit. The genus Bigelowiella includes B. natans (Moestrup and Sengco 2001), the model species for most chlorarachniophyte cell and molecular biology, and B. longifolia (Ota et al. 2007b). B. natans has only been observed as zoospores (although they can form pseudopodia (Moestrup and Sengco 2001)), whereas a true amoeboid stage is known in B. longifolia (Ota et al. 2007b). There are currently six known species of Lotharella: L. vacuolata (Ota et al. 2005), L. globosa (Ishida and Hara 1994), L. globosa var. fortis (Hirakawa et al. 2011b), L. polymorpha (Dietz et al. 2003), L. oceanica (Ota et al. 2009b), and L. reticulosa (Ota and Vaulot 2012), some of which are primarily coccoid and some primarily amoeboid, but all share a pyrenoid with a deep slit and a nucleomorph positioned at the base of the pyrenoid. Amorphochlora amoebiformis was originally described as L. amoeboformis (Ishida et al. 2000) but was transferred to a new genus based on molecular phylogenetic evidence (Ishida et al. 2011b). Gymnochlora stellata and G. dimorpha are the described species of Gymnochlora (Ishida et al. 1996; Ota et al. 2011), and Norrisiella sphaerica is the only described species of Norrisiella (Ota et al. 2007a). Both are distinguished from other genera by pyrenoid and nucleomorph characters. Partenskyella glossopodia is the only species of Partenskyella and is distinguished by a complete absence of a pyrenoid (Ota et al. 2009a). A last genus comprises the enigmatic and poorly described species Cryptochlora perforans, which is a mixotrophic species that is attracted to damaged algal thalli that it can physically penetrate and feed upon. It has been classified as a chlorarachniophyte based on similarities in life history complexity and plastid characteristics (Calderon-Saenz and Schnetter 1987, 1989). Unfortunately, it has not been described at the ultrastructural or molecular levels, so its exact relationship to chlorarachniophytes is not completely clear.
The relationships between chlorarachniophyte genera are not at all clear from morphological characters, but molecular phylogenies based on genes from the nucleus, nucleomorph, and mitochondria all support the currently analyzable species as being distinct, as well as genera-level distinctions. Phylogenies generally support an overall picture shown in Fig. 3. There is a consistent and well-supported close relationship between the genera Bigelowiella and Norrisiella, both of which are in turn related to Chlorarachnion, with Partenskyella, Lotharella, and Gymnochlora forming an unresolved radiation at the base of the group (Gilson and McFadden 1999; Ishida and Cavalier-Smith 1999; Silver et al. 2007; Ota et al. 2009a; Ota and Vaulot 2012).
Molecular bar codes have been established and tested for all chlorarachniophyte species available in culture (Gile et al. 2010). Nucleomorph ribosomal RNA intergenic spacer (ITS) sequence was found to provide good resolution at the species level, at least for the few genera with multiple species, and was subsequently used to characterize new isolates (Hirakawa et al. 2011b; Ota and Vaulot 2012).
Maintenance and Cultivation
Many species grow easily but slowly in a variety of marine media (see Hibberd and Norris 1984; Ishida et al. 2000; Moestrup and Sengco 2001; Ota et al. 2007a, b). Primarily amoeboid and coccoid species mostly accumulate in masses on surfaces, while primarily flagellated forms can be grown to high densities by shaking or aeration. Currently over 30 strains are available from several culture collections, the largest collection of strains being at the Provasoli-Guillard National Center for Culture of Marine Phytoplankton.
Genomics
The unique evolutionary history and current complexity of chlorarachniophytes has led to several genomic and comparative genomic projects. Currently complete genomes for all four compartments, plastid, mitochondrion, nucleomorph, and nucleus, have been sequenced for the model species B. natans (Gilson et al. 2006; Rogers et al. 2007; Curtis et al. 2012). Organelle genomes, proteomics, and surveys of gene expression have also been carried out for a number of species (Williams et al. 2005; Slamovits and Keeling 2009; Hopkins et al. 2012; Tanifuji et al. 2014; Suzuki et al. 2015), all revealing a model for the effects of endosymbiotic integration and nuclear genome compaction. The nucleomorph genome is severely reduced with only about 300 tightly packed genes but still retains over 800 introns that are all compacted, nearly all to 18–21 bp in length (Gilson et al. 2006; Slamovits and Keeling 2009). Nuclear genome organization is conventional but revealed a large number of genes derived by horizontal gene transfer, as well as genes originating by endosymbiotic gene transfer from the endosymbiont (Archibald et al. 2003; Gile et al. 2008; Hirakawa et al. 2011a; Curtis et al. 2012). The nucleus is haploid and nucleomorph diploid (Hirakawa and Ishida 2014).
Evolutionary History
The unique combination of characters found in chlorarachniophytes led to much speculation and confusion about their possible evolutionary origin in early studies (Geitler 1930; Hibberd and Norris 1984; Grell 1990; Hibberd 1990), particularly before it was understood that they are a symbiotic fusion of a colorless amoeboflagellate and a green alga (Whatley and Whatley 1981; Cavalier-Smith 1982). Now this endosymbiotic origin of chlorarachniophytes has been demonstrated beyond any doubt by ultrastructural studies and characterization of the nucleomorph genome (Ludwig and Gibbs 1989; McFadden et al. 1994; Gilson et al. 2006). While the nucleomorph and its genome have been retained, many endosymbiont genes were moved to the host nucleus (Deane et al. 2000; Archibald et al. 2003; Gile and Keeling 2008; Hirakawa et al. 2011a; Curtis et al. 2012) and many other features simply lost, for example, Golgi bodies, mitochondria, locomotary organelles, and carbohydrate storage, during the integration of chlorarachniophyte plastids.
Even after the endosymbiotic origin of chlorarachniophytes was well established, however, the origin of both the host and the endosymbiont continued to be controversial. The presence of chlorophyll b immediately suggested a link to green algae (Hibberd and Norris 1984), but numerous theories about which kind of green algae were put forward (Sasa et al. 1882; Cavalier-Smith et al. 1994; Van de Peer et al. 1996; Ishida et al. 1997; Ishida and Cavalier-Smith 1999). Current data only suggest it is a member of the ulvophyte-trebouxiophyte-chlorophyte complex (Rogers et al. 2007; Turmel et al. 2009). The evolutionary history of the host was considerably more obscure since chlorarachniophytes do not share any obvious defining morphological feature with any other group. Molecular data have shed considerable light on this, however, and consistently show the chlorarachniophyte host to be part of a large and diverse group of flagellate s, amoeba e, and amoeboflagellates, called Cercozoa (Cavalier-Smith 1999). There are currently no structural characteristics that uniquely unite all Cercozoa, but phylogenies based on all genes that have been examined individually or as large concatenates (Bhattacharya et al. 1995; Cavalier-Smith and Chao 1997; Keeling et al. 1998; Keeling 2001; Longet et al. 2003; Nikolaev et al. 2004; Takishita et al. 2005; Burki et al. 2007, 2012), as well as the presence of unique insertion/deletions in polyubiquitins and rRNA (Cavalier-Smith and Chao 1997; Archibald et al. 2002), all consistently show Cercozoa to be monophyletic group that includes chlorarachniophytes, likely as an early-branching subgroup. Cercozoa, in turn has been shown to be part of an even larger group called Rhizaria, which also includes a number of mostly amoeboid lineages such as foraminiferans, acantharians, and polycystines (Sierra et al. 2012). Altogether, Rhizaria consistently branch as sisters to the alveolates and stramenopiles in large multigene phylogenetic analyses (Burki et al. 2007, 2012). The closest cercozoan sister group to the chlorarachniophytes in current phylogenies appears to be the pico-heterotroph, Minorisa, one of the smallest known eukaryotes (del Campo et al. 2013).
References
Archibald, J. M. (2007). Nucleomorph genomes: Structure, function, origin and evolution. Bioessays, 29, 392–402.
Archibald, J. M., & Keeling, P. J. (2002). Recycled plastids: A green movement in eukaryotic evolution. Trends in Genetics, 18, 577–584.
Archibald, J. M., Longet, D., Pawlowski, J., & Keeling, P. J. (2002). A novel polyubiquitin structure in Cercozoa and Foraminifera: Evidence for a new eukaryotic supergroup. Molecular Biology and Evolution, 20, 62–66.
Archibald, J. M., Rogers, M. B., Toop, M., Ishida, K., & Keeling, P. J. (2003). Lateral gene transfer and the evolution of plastid-targeted proteins in the secondary plastid-containing alga Bigelowiella natans. Proceedings of the National Academy of Sciences of the United States of America, 100, 7678–7683.
Bhattacharya, D., Helmchen, T., & Melkonian, M. (1995). Molecular evolutionary analyses of nuclear-encoded small subunit ribosomal RNA identify an independent rhizopod lineage containing the Euglyphidae and the Chlorarachniophyta. The Journal of Eukaryotic Microbiology, 42, 64–68.
Burki, F., Shalchian-Tabrizi, K., Minge, M., Skjaeveland, A., Nikolaev, S. I., Jakobsen, K. S., & Pawlowski, J. (2007). Phylogenomics reshuffles the eukaryotic supergroups. PLoS One, 2, e790.
Burki, F., Okamoto, N., Pombert, J.-F., & Keeling, P. J. (2012). Phylogenomic evidence for a polyphyletic origin of the cryptophytes, haptophytes, and associated heterotrophic lineages. Proceedings of the Royal Society B, 279, 2246–2254.
Calderon-Saenz, E., & Schnetter, R. (1987). Cryptochlora perforans, a new genus and species of alga (Chlorarachniophyta), capable of penetrating dead algal filaments. Plant Systematics and Evolution, 158, 69–71.
Calderon-Saenz, E., & Schnetter, R. (1989). Morphology, biology and systematics of Cryptochlora perforans (Chlorarachniophyta), a phagotropic marine alga. Plant Systematics and Evolution, 163, 165–176.
Cavalier-Smith, T. (1982). The origins of plastids. Biological Journal of the Linnean Society, 17, 289–306.
Cavalier-Smith, T. (1999). Principles of protein and lipid targeting in secondary symbiogenesis: Euglenoid, dinoflagellate, and sporozoan plastid origins and the eukaryote family tree. The Journal of Eukaryotic Microbiology, 46, 347–366.
Cavalier-Smith, T. (2002). Nucleomorphs: Enslaved algal nuclei. Current Opinion in Microbiology, 5, 612–619.
Cavalier-Smith, T., & Chao, E. E. (1997). Sarcomonad ribosomal RNA sequences, rhizopod phylogeny, and the origin of euglyphid amoebae. Archiv für Protistenkunde, 147, 227–236.
Cavalier-Smith, T., Allsopp, M. T., & Chao, E. E. (1994). Chimeric conundra: Are nucleomorphs and chromists monophyletic or polyphyletic? Proceedings of the National Academy of Sciences of the United States of America, 91, 11368–11372.
Curtis, B. A., Tanifuji, G., Burki, F., Gruber, A., Irimia, M., Maruyama, S., Arias, M. C., Ball, S. G., Gile, G. H., Hirakawa, Y., Hopkins, J. F., Kuo, A., Rensing, S. A., Schmutz, J., Symeonidi, A., Elias, M., Eveleigh, R. J., Herman, E. K., Klute, M. J., Nakayama, T., Oborník, M., Reyes-Prieto, A., Armbrust, E. V., Aves, S. J., Beiko, R. G., Coutinho, P., Dacks, J. B., Durnford, D. G., Fast, N. M., Green, B. R., Grisdale, C. J., Hempel, F., Henrissat, B., Höppner, M. P., Ishida, K., Kim, E., Kořený, L., Kroth, P. G., Liu, Y., Malik, S. B., Maier, U. G., McRose, D., Mock, T., Neilson, J. A., Onodera, N. T., Poole, A. M., Pritham, E. J., Richards, T. A., Rocap, G., Roy, S. W., Sarai, C., Schaack, S., Shirato, S., Slamovits, C. H., Spencer, D. F., Suzuki, S., Worden, A. Z., Zauner, S., Barry, K., Bell, C., Bharti, A. K., Crow, J. A., Grimwood, J., Kramer, R., Lindquist, E., Lucas, S., Salamov, A., McFadden, G. I., Lane, C. E., Keeling, P. J., Gray, M. W., Grigoriev, I. V., & Archibald, J. M. (2012). Algal genomes reveal evolutionary mosaicism and the fate of nucleomorphs. Nature, 492, 59–65.
Deane, J. A., Fraunholz, M., Su, V., Maier, U.-G., Martin, W., Durnford, D. G., & McFadden, G. (2000). Evidence for nucleomorph to host nucleus gene transfer: Light-harvesting complex proteins from cryptomonads and chlorarachniophytes. Protist, 151, 239–252.
del Campo, J., Not, F., Forn, I., Sieracki, M. E., & Massana, R. (2013). Taming the smallest predators of the oceans. The ISME Journal, 7, 351–8.
Dietz, C., Ehlers, K., Wilhelm, C., Gil-Rodriguez, M. C., & Schnetter, R. (2003). Lotharella polymorpha sp. nov. (Chlorarachniophyta) from the coast of Portugal. Phycologia, 42, 582–593.
Geitler, L. (1930). Ein grünes Filarplamodium und andere neue Protisten. Archiv für Protistenkunde, 69, 221–230.
Gile, G. H., & Keeling, P. J. (2008). Nucleus-encoded periplastid-targeted EFL in chlorarachniophytes. Molecular Biology and Evolution, 25(9), 1967–1977.
Gile, G. H., Stern, R. F., James, E. R., & Keeling, P. J. (2010). DNA barcoding of chlorarachniophytes using nucleomorph ITS. Journal of Phycology, 46, 743–750.
Gilson, P. R. (2001). Nucleomorph genomes: Much ado about practically nothing. Genome Biol, 2, R1022.
Gilson, P. R., & McFadden, G. I. (1997). Good things in small packages: The tiny genomes of chlorarachniophyte endosymbionts. Bioessays, 19, 167–173.
Gilson, P., & McFadden, G. (1999). Molecular, morphological and phylogenetic characterization of six chlorarachniophyte strains. Phycological Research, 47, 7–19.
Gilson, P. R., & McFadden, G. I. (2002). Jam packed genomes – A preliminary, comparative analysis of nucleomorphs. Genetica, 115, 13–28.
Gilson, P. R., Maier, U. G., & McFadden, G. I. (1997). Size isn’t everything: Lessons in genetic miniaturisation from nucleomorphs. Current Opinion in Genetics and Development, 7, 800–806.
Gilson, P. R., Su, V., Slamovits, C. H., Reith, M. E., Keeling, P. J., & McFadden, G. I. (2006). Complete nucleotide sequence of the chlorarachniophyte nucleomorph: Nature’s smallest nucleus. Proceedings of the National Academy of Sciences of the United States of America, 103, 9566–9571.
Grell, K. G. (1990). Some light microscope observations on Chlorarachnion reptans Geitler. Archiv für Protistenkunde, 138, 271–290.
Hempel, F., Bullmann, L., Lau, J., Zauner, S., & Maier, U. G. (2009). ERAD-derived preprotein transport across the second outermost plastid membrane of diatoms. Molecular Biology and Evolution, 26, 1781–1790.
Hibberd, D. J. (1990). Phylum Chlorarachnida. In L. Margulis, J. O. Corliss, M. Melkonian, & D. J. Chapman (Eds.), Handbook of protoctista (pp. 288–292). Boston: Jones and Bartlett Publishers.
Hibberd, D. J., & Norris, R. E. (1984). Cytology and ultrastructure of Chlorarachnion reptans (Chlorarachniophyta divisio nova, Chlorarachniophyceae classis nova). Journal of Phycology, 20, 310–330.
Hirakawa, Y., & Ishida, K. (2010). Internal plastid-targeting signal found in a RubisCO small subunit protein of a chlorarachniophyte alga. The Plant Journal, 64, 402–410.
Hirakawa, Y., & Ishida, K. (2014). Polyploidy of endosymbiotically derived genomes in complex algae. Genome Biology and Evolution, 6, 974–80.
Hirakawa, Y., Nagamune, K., & Ishida, K. (2009). Protein targeting into secondary plastids of chlorarachniophytes. Proceedings of the National Academy of Sciences of the United States of America, 106, 12820–12825.
Hirakawa, Y., Gile, G. H., Ota, S., Keeling, P. J., & Ishida, K. I. (2010). Characterization of periplastidal compartment targeting signals in chlorarachniophytes. Molecular Biology and Evolution, 27, 1538–1545.
Hirakawa, Y., Burki, F., & Keeling, P. J. (2011a). Nucleus- and nucleomorph-targeted histone proteins in a chlorarachniophyte alga. Molecular Microbiology, 80, 1339–1449.
Hirakawa, Y., Howe, A., James, E. R., & Keeling, P. J. (2011b). Morphological diversity between culture strains of a chlorarachniophyte, Lotharella globosa. PLoS One, 6, e23193.
Hirakawa, Y., Burki, F., & Keeling, P. J. (2012a). Genome-based reconstruction of the protein import machinery in the secondary plastid of a chlorarachniophyte alga. Eukaryotic Cell, 11, 324–333.
Hirakawa, Y., Burki, F., & Keeling, P. J. (2012b). Dual targeting of aminoacyl-tRNA synthetases to the mitochondrion and complex plastids of chlorarachniophytes. Journal of Cell Science, 125, 6176–6184.
Hopkins, J. F., Spencer, D. F., Laboissiere, S., Neilson, J. A., Eveleigh, R. J., Durnford, D. G., Gray, M. W., & Archibald, J. M. (2012). Proteomics reveals plastid- and periplastid-targeted proteins in the chlorarachniophyte alga Bigelowiella natans. Genome Biology and Evolution, 4, 1391–406.
Ishida, K., & Cavalier-Smith, T. (1999). Diversification of a chimerica algal group, the chlorarachniophytes: Phylogeny of nuclear and nucleomorph encoded small-subunit rRNA genes. Molecular Biology and Evolution, 16, 321–331.
Ishida, K., & Hara, Y. (1994). Taxonomic studies on the Chlorarachniophyta. I. Chlorarachnion globosum sp. nov. Phycologia, 33, 351–358.
Ishida, K., Nakayama, T., & Hara, Y. (1996). Taxonomic studies on the Chlorarachniophyta. II. Generic delimination of the chlorarachniophytes and description of Gynmochlora stellata gen. et sp. nov. Phycological Research, 44, 37–45.
Ishida, K., Cao, Y., Hasegawa, M., Okada, N., & Hara, Y. (1997). The origin of chlorarachniophyte plastids, as inferred from phylogenetic comparisons of amino acid sequences of EF-Tu. Journal of Molecular Evolution, 45, 682–687.
Ishida, K., Ishida, N., & Hara, Y. (2000). Lotharella amoeboformis sp. nov.: A new species of chlorarachniophyte from Japan. Phycological Research, 48, 221–229.
Ishida, K., Yabuki, A., & Ota, S. (2007). The chlorarachniophytes: Evolution and classification. In J. Brodie & J. Lewis (Eds.), Unravelling the algae – The past, present, and future of algal systematics. Boca Raton: CRC Press.
Ishida, K., Endo, H., & Koike, S. (2011a). Partenskyella glossopodia (Chlorarachniophyceae) possesses a nucleomorph genome of approximately 1Mbp. Phycological Research, 59, 120–122.
Ishida, K., Yabuki, A., & Ota, S. (2011b). Amorphochlora amoebiformis gen. et comb. nov. (Chlorarachniophyceae). Phycological Research, 59, 52–53.
Keeling, P. J. (2001). Foraminifera and Cercozoa are related in actin phylogeny: Two orphans find a home? Molecular Biology and Evolution, 18, 1551–1557.
Keeling, P. J., Deane, J. A., & McFadden, G. I. (1998). The phylogenetic position of alpha- and beta-tubulins from the Chlorarachnion host and Cercomonas (Cercozoa). The Journal of Eukaryotic Microbiology, 45, 561–570.
Leblond, J. D., Dahmen, J. L., Seipelt, R. L., Elrod-Erickson, M. J., Kincaid, R., Howard, J. C., Evens, T. J., & Chapman, P. J. (2005). Lipid composition of chlorarachniophytes (Chlorarachniophyceae) from the genera Bigelowiella, Gymnochlora, and Lotharella. Journal of Phycology, 41, 311–321.
Longet, D., Archibald, J. M., Keeling, P. J., & Pawlowski, J. (2003). Foraminifera and Cercozoa share a common origin according to RNA polymerase II phylogenies. International Journal of Systematic and Evolutionary Microbiology, 53, 1735–1739.
Ludwig, M., & Gibbs, S. P. (1989). Evidence that the nucleomorphs of Chlorarachnion reptans (chlorarachniophyceae) are vestigial nuclei: Morphology, division and DNA-DAPI fluorescence. Journal of Phycology, 25, 385–394.
McFadden, G. I., & Gilson, P. R. (1995). Something borrowed, something green: Lateral transfer of chloroplasts by secondary endosymbiosis. Trends in Ecology & Evolution, 10, 12–17.
McFadden, G. I., Gilson, P. R., Hofmann, C. J., Adcock, G. J., & Maier, U. G. (1994). Evidence that an amoeba acquired a chloroplast by retaining part of an engulfed eukaryotic alga. Proceedings of the National Academy of Sciences of the United States of America, 91, 3690–3694.
McFadden, G. I., Gilson, P. R., Douglas, S. E., Cavalier-Smith, T., Hofmann, C. J., & Maier, U. G. (1997a). Bonsai genomics: Sequencing the smallest eukaryotic genomes. Trends in Genetics, 13, 46–49.
McFadden, G. I., Gilson, P. R., & Sims, I. M. (1997b). Preliminary characterization of carbohydrate stores from chlorarachniophytes (Division: Chlorarachniophyta). Phycological Research, 45, 145–151.
Moestrup, Ø., & Sengco, M. (2001). Ultrastructural studies on Bigelowiella natans, gen. et sp. nov., a chlorarachniophyte flagellate. Journal of Phycology, 37, 624–646.
Nikolaev, S. I., Berney, C., Fahrni, J. F., Bolivar, I., Polet, S., Mylnikov, A. P., Aleshin, V. V., Petrov, N. B., & Pawlowski, J. (2004). The twilight of Heliozoa and rise of Rhizaria, an emerging supergroup of amoeboid eukaryotes. Proceedings of the National Academy of Sciences of the United States of America, 101, 8066–8071.
Ota, S., & Vaulot, D. (2012). Lotharella reticulosa sp. nov.: A highly reticulated network forming chlorarachniophyte from the Mediterranean Sea. Protist, 163, 91–104.
Ota, S., Ueda, K., & Ishida, K. (2005). Lotharella vacuolata sp nov., a new species of chlorarachniophyte algae, and time-lapse video observations on its unique post-cell division behavior. Phycological Research, 53, 275–286.
Ota, S., Ueda, K., & Ishida, K. (2007a). Norrisiella sphaerica gen. et sp. nov., a new coccoid chlorarachniophyte from Baja california, Mexico. Journal of Plant Research, 120, 661–670.
Ota, S., Ueda, K., & Ishida, K. (2007b). Taxonomic study of Bigelowiella longifila sp. nov. (Chlorarachniophyta) and time-lapse video observation of the unique migration of amoeboid cells. Journal of Phycology, 43, 333–343.
Ota, S., Vaulot, D., & Gall, F. L. (2009a). Yabuki, and K. Ishida: Partenskyella glossopodia gen et ap. nov., the first report of a chlorarachniophyte that lacks a pyrenoid. Protist, 160, 137–150.
Ota, S., Sikver, T. D., Archibald, J. M., & Ishida, K. (2009b). Lotharella oceanica sp. nov.-a new planktonic chlorarachniophyte studied by light and electron microscopy. Phycologia, 48, 315–323.
Ota, S., Kudo, A., & Ishida, K. (2011). Gymnochlora dimorpha sp. nov., a chlorarachniophyte with unique daughter cell behavior. Phycologia, 50, 317–326.
Rogers, M. B., Archibald, J. M., Field, M. A., Li, C., Striepen, B., & Keeling, P. J. (2004). Plastid-targeting peptides from the chlorarachniophyte Bigelowiella natans. The Journal of Eukaryotic Microbiology, 51, 529–535.
Rogers, M. B., Gilson, P. R., Su, V., McFadden, G. I., & Keeling, P. J. (2007). The complete chloroplast genome of the chlorarachniophyte Bigelowiella natans: Evidence for independent origins of chlorarachniophyte and euglenid secondary endosymbionts. Molecular Biology and Evolution, 24, 54–62.
Sasa, T., Takaichi, S., Hatakeyama, N., & Watanabe, M. M. (1882). A novel carotenoid ester, lorozathin dodecenoate, from Pryamimonas-parkeae (Prasinophyceae) and a chlorarachniophycean alga. Plant and Cell Physiology, 33, 921–925.
Sierra, R., Matz, M. V., Aglyamova, G., Pillet, L., Decelle, J., Not, F., de Vargas, C., & Pawlowski, J. (2012). Deep relationships of Rhizaria revealed by phylogenomics: A farewell to Haeckel’s Radiolaria. Molecular Phylogenetics and Evolution, 67, 53–59.
Silver, T. D., Koike, S., Yabuki, A., Kofuji, R., Archibald, J. M., & Ishida, K. (2007). Phylogeny and nucleomorph karyotype diversity of chlorarachniophyte algae. The Journal of Eukaryotic Microbiology, 54, 403–410.
Slamovits, C. H., & Keeling, P. J. (2009). Evolution of ultrasmall introns in highly reduced nuclear genomes. Molecular Biology and Evolution, 26, 1699–1705.
Suzuki, S., Shirato, S., Hirakawa, Y., & Ishida, K. (2015). Nucleomorph genome sequences of two chlorarachniophytes, Amorphochlora amoebiformis and Lotharella vacuolata. Genome Biology and Evolution, 7, 1533–45.
Takishita, K., Inagaki, Y., Tsuchiya, M., Sakaguchi, M., & Maruyama, T. (2005). A close relationship between Cercozoa and Foraminifera supported by phylogenetic analyses based on combined amino acid sequences of three cytoskeletal proteins (actin, alpha-tubulin, and beta-tubulin). Gene, 362, 153–160.
Tanifuji, G., Onodera, N. T., Brown, M. W., Curtis, B. A., Roger, A. J., Ka-Shu Wong, G., Melkonian, M., & Archibald, J. M. (2014). Nucleomorph and plastid genome sequences of the chlorarachniophyte Lotharella oceanica: Convergent reductive evolution and frequent recombination in nucleomorph-bearing algae. BMC Genomics, 15, 374.
Turmel, M., Gagnon, M. C., O’Kelly, C. J., Otis, C., & Lemieux, C. (2009). the chloroplast genomes of the green algae Pyramimonas, Monomastix, and Pycnococcus shed new light on the evolutionary history of prasinophytes and the origin of secondary chloroplasts in euglenoids. Molecular Biology and Evolution, 26, 631–648.
Van de Peer, Y., Rensing, S. A., Maier, U. G., & De Wachter, R. (1996). Substitution rate calibration of small subunit ribosomal RNA identifies chlorarachniophyte endosymbionts as remnants of green algae. Proceedings of the National Academy of Sciences of the United States of America, 93, 7732–7736.
Whatley, J. M., & Whatley, F. R. (1981). Chloroplast evolution. The New Phytologist, 87, 233–247.
Williams, B. A., Slamovits, C. H., Patron, N. J., Fast, N. M., & Keeling, P. J. (2005). A high frequency of overlapping gene expression in compacted eukaryotic genomes. Proceedings of the National Academy of Sciences of the United States of America, 102, 10936–10941.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this entry
Cite this entry
Keeling, P.J. (2017). Chlorarachniophytes. In: Archibald, J., Simpson, A., Slamovits, C. (eds) Handbook of the Protists. Springer, Cham. https://doi.org/10.1007/978-3-319-28149-0_34
Download citation
DOI: https://doi.org/10.1007/978-3-319-28149-0_34
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-28147-6
Online ISBN: 978-3-319-28149-0
eBook Packages: Biomedical and Life SciencesReference Module Biomedical and Life Sciences