Abstract
Sphingolipids, a once overlooked class of lipids in plants, are now recognized as abundant and essential components of plasma membrane and other endomembranes of plant cells. In addition to providing structural integrity to plant membranes, sphingolipids contribute to Golgi trafficking and protein organizational domains in the plasma membrane. Sphingolipid metabolites have also been linked to the regulation of cellular processes, including programmed cell death. Advances in mass spectrometry-based sphingolipid profiling and analyses of Arabidopsis mutants have enabled fundamental discoveries in sphingolipid structural diversity, metabolism, and function that are reviewed here. These discoveries are laying the groundwork for the tailoring of sphingolipid biosynthesis and catabolism for improved tolerance of plants to biotic and abiotic stresses.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
Sphingolipids were originally identified in the late nineteenth century by Johann Thudichum as an “enigmatic” major lipidic component of the brain (Thudichum 1884). Since this discovery, sphingolipids have been recognized as essential components of eukaryotic cells and have been extensively studied in humans due to their association with a number of lipid storage disorders, including Tay-Sachs disease and Niemann-Pick disease (Sandhoff 2013). Sphingolipids, however, were not identified in plants until the late 1950s (Carter et al. 1958) and for nearly four decades following this discovery, sphingolipid research in plants was limited mainly to structural and compositional analyses, including studies of sphingolipid compositional changes in response to abiotic stresses. Since the late 1990s, plant sphingolipids have become an increasing research focus. Driving this heightened interest is the realization that sphingolipids are among the most abundant endomembrane lipids in plant cells and that they contribute not only to membrane structure and function that underlies abiotic and biotic stress resistance but also to the regulation of cellular processes. Recent advances in plant sphingolipid research have been spurred by development and application of advanced mass spectrometry methods that enable the rapid and quantitative measurement of molecular species of specific sphingolipid classes (Markham and Jaworski 2007). Coupling of these methods with the characterization of Arabidopsis mutants has resulted in advances in our fundamental understanding of plant sphingolipid metabolism and its regulation. The availability of Arabidopsis mutants has also led to discoveries of the involvement of sphingolipids in plant growth and responses to environmental stimuli. In addition, unexpected connections between sphingolipids and physiological processes such as programmed cell death and the related hypersensitive response that is important for plant resistance to bacterial and fungal pathogens have arisen from forward genetic studies of Arabidopsis. Recent research has also been directed at understanding the specialized functions of sphingolipids in plasma membrane microdomains that are believed to contribute to cell surface processes such as cell wall metabolism and ion and auxin transport. This chapter provides an overview of our recent understanding of plant sphingolipid structure, metabolism, and function and highlights unanswered questions in plant sphingolipid biology.
Sphingolipid Structure
Sphingolipids consist of hydrophobic ceramide backbones that are typically linked to polar sugar residues to form amphipathic lipid components of membranes (Lynch and Dunn 2004; Chen et al. 2010). The ceramide backbone contains a long chain amino alcohol referred to as a sphingoid long-chain base (LCB) linked through an amide bond to a fatty acid. LCBs are unique to sphingolipids. In plants, LCBs typically have chain lengths of 18 carbon atoms and can contain double bonds in the Δ4 or Δ8 positions (Fig. 11.1a). The Δ4 double bond is found only in the trans configuration, while the Δ8 double bond can be found in either the trans or cis configurations. Following its initial synthesis, a LCB has two hydroxyl groups at the C-1 and C-3 carbons (Lynch and Dunn 2004; Chen et al. 2010). These LCBs are referred to as dihydroxy LCBs. A third hydroxyl group can be enzymatically added at the C-4 carbon to form a trihydroxy LCB. In the short-hand nomenclature, a dihydroxy LCB with 18 carbons and 1 double bond is referred to as “d18:1”, and a trihydroxy LCB with 18 carbons and 1 double bond is referred to as “t18:1”. LCBs can be phosphorylated at the C-1 position to form LCB-phosphates (LCB-P). Free LCBs and their phosphorylated forms are typically in low abundance in plant cells (Markham and Jaworski 2007; Markham et al. 2006). Instead, the majority of LCBs are found linked to fatty acids in ceramides (Fig. 11.1b). The chain-lengths of plant ceramide fatty acids range from 16 to 26 carbon atoms, the majority of which contain an enzymatically added hydroxyl group at the C-2 or α-position (Lynch and Dunn 2004; Chen et al. 2010). Analogous to the diacylglycerol backbone of glycerolipids, ceramides serve as the hydrophobic component of complex sphingolipids. The polar head group of ceramides is attached at its C-1 position and can be a phosphate residue or a variety of sugar residues (Chen et al. 2010). The latter are referred to as glycosphingolipids. The simplest glycosphingolipid in plants is the glucosylceramide (GlcCer) with a single glucose residue and comprises approximately one-third of the glycosphingolipids of Arabidopsis leaves (Markham and Jaworski 2007; Markham et al. 2006) (Fig. 11.1c). The most abundant glycosphingolipid in plants contains an inositol phosphate bound to the ceramide with up to seven additional hexose and pentose residues (Fig. 11.1c) (Cacas et al. 2013). These molecules are referred to as glycosyl inositolphosphoceramides or GIPCs and comprise approximately two-thirds of the glycosphingolipids of Arabidopsis leaves (Markham and Jaworski 2007; Markham et al. 2006). The quantitative significance of GIPCs in plants was overlooked for many years due to the difficulty in their extraction using standard lipid analytical protocols because of the high polarity of their glycosylated head groups. Between the different carbon chain-lengths and hydroxylation and unsaturation states of LCBs and fatty acids and the array of polar head groups, hundreds of potentially different sphingolipid species can occur in plants, the individual significance of which are only beginning to be elucidated (Markham et al. 2013; Bure et al. 2011).
Examples of long-chain bases (LCB) and sphingolipids found in plants. (a) Examples of LCB modifications found in plants. Shown are examples of dihydroxy and trihydroxy LCBs The nomenclature “d18:0” indicates that the LCB has two hydroxyl groups (d) and 18 carbon atoms and no double bonds, and the nomenclature “t18:0” indicates that the LCB has three hydroxyl groups (t) and 18 carbon atoms and no double bonds. (b) Hydoxyceramide composed of the LCB t18:1 Δ8trans and the fatty acid 24:1 ω9cis that is hydroxylated at the C-2 position. (c) Most abundant glycosyl inositolphosphoceramide (GIPC) found in Arabidopsis leaves. (d) Glucosylceramide
Biosynthesis of Long-Chain Bases
Serine Palmitoyltransferase Complex
The biosynthesis of LCBs is initiated through an endoplasmic reticulum-localized reaction catalyzed by serine palmitoyltransferase (SPT) that condenses serine and palmitoyl-CoA to form the 18 carbon intermediate 3-ketosphinganine (Figs. 11.2 and 11.3) (Chen et al. 2006; Dietrich et al. 2008; Teng et al. 2008). The product of this reaction is then reduced by 3-ketosphinganine reductase (KSR) to form sphinganine or d18:0, the simplest long-chain base in plants and other eukaryotes (Chao et al. 2011). SPT is a member of the α-oxoamine synthase subfamily and is generally regarded as the main regulated step in sphingolipid biosynthesis (Hanada 2003). Similar to other eukaryotes, the Arabidopsis SPT functions as a heterodimer comprised of LCB1 and LCB2 subunits (Table 11.1) (Tamura et al. 2001; Chen et al. 2006; Dietrich et al. 2008; Teng et al. 2008). Although both LCB1 and LCB2 show similarity with α-oxoamine synthases, the catalytic lysine residue that forms a Schiff base with pyridoxal phosphate is found in the LCB2 subunit (Tamura et al. 2001; Hanada 2003). A third smaller subunit, termed the small subunit of SPT or ssSPT, also interacts with the LCB1/LCB2 subunits (Table 11.1) (Han et al. 2004; Kimberlin et al. 2013). ssSPT polypeptides in Arabidopsis contain only 56 amino acids that lack any predicted enzymatic activity but contain a single transmembrane domain (Kimberlin et al. 2013). It is believed that the active site of SPT occurs at the interface of LCB1 and LCB2 with LCB1 and ssSPT acting to stabilize the complex (Gable et al. 2000, 2002). Although SPT can function as a heterodimer (LCB1 and LCB2) with minimal enzymatic activity, ssSPT enhances SPT activity to levels that produce LCBs in amounts that are sufficient to support cell viability in Arabidopsis (Kimberlin et al. 2013).
Schematic of the serine palmitoyltransferase (SPT) complex. In plants, SPT exists as an ER-localized heterodimer of LCB1 and LCB2 subunits that interacts with several regulatory proteins. A stimulatory small subunit of SPT (ssSPT) binds to the core SPT heterodimer to increase SPT activity. Sac1 phosphatase and Ypk1 have been shown in Saccharomyces cerevisiae to catalyze reversible phosphorylation of the inhibitory regulator ORM in response to intracellular sphingolipid levels. The regulatory mechanism of ORM in plants remains to be elucidated. Long-chain bases (LCBs) produced by SPT and 3-ketosphinganine reductase (KSR) are used by ceramide synthase (CS) along with fatty acid acyl-CoAs to produce ceramides. Fumonisin B1 (FB 1) is a mycotoxin that inhibits CS and triggers programmed cell death (Figure is derived from Kimberlin et al. 2013)
Studies of LCB1, LCB2, and ssSPT mutants have demonstrated that SPT activity is essential, and consequently, sphingolipids are required for the viability of plant cells (Kimberlin et al. 2013; Dietrich et al. 2008; Teng et al. 2008). In this regard, the fbr11-2 mutant of the single Arabidopsis LCB1 gene (At4g36480) displays male gametophytic lethality (Teng et al. 2008). Loss of pollen viability is also observed in double mutants of the two redundant LCB2 genes LCB2a (At5g23670) and LCB2b (At3g48780) (Dietrich et al. 2008) as well as in null mutants of ssSPTa (At1g06515), the more highly expressed of the two ssSPT genes in Arabidopsis (Kimberlin et al. 2013). Pollen deficient in sphingolipid synthesis in lcb2a −/− lcb2b −/+ mutants lack Golgi stacks and surrounding intine layer and have vesiculated ER, all of which are consistent with the contributions of sphingolipids to the structural and functional integrity of the endomembrane system (Dietrich et al. 2008).
The regulation of SPT is thought to occur primarily through ssSPT and a second class of interacting proteins termed orosomucoid or ORM polypeptides, rather than transcriptional mechanisms (Fig. 11.2 and Table 11.1) (Markham et al. 2013). The ssSPTs appear to be limiting and so modulation of their expression alters SPT activity. This has been shown in Arabidopsis with ssSPT over-expression leading to increased SPT activity and ssSPT RNAi suppression resulting in reduced SPT activity (Kimberlin et al. 2013). Whether alteration of ssSPT levels occurs naturally in response to intracellular cues to mediate sphingolipid homeostasis is unclear. In addition, no evidence currently exists for regulation of SPT activity through post-translational modifications of ssSPT polypeptides.
It is also notable that ssSPTs can dictate the acyl-CoA specificity of SPT (Han et al. 2009). In this regard, human ssSPTa and ssSPTb polypeptides have been shown to confer different acyl-CoA specificity when bound to SPT leading to the production of either C18 LCBs using palmitoyl (16:0)-CoA substrates or C20 LCBs using stearoyl (18:0)-CoA substrates (Han et al. 2009). These differences in acyl-CoA substrate specificities were shown to result from a single amino acid residue that is a Met in the human ssSPTa and Val in human ssSPTb (Han et al. 2009). The Arabidopsis ssSPTa and ssSPTb polypeptides both contain Met at the analogous position, but mutation of Met to Val results in the aberrant production of C20 LCBs when expressed in transgenic Arabidopsis (Kimberlin et al. 2013).
In yeast, ORM proteins have been shown to act as homeostatic negative regulators of SPT in response to intracellular sphingolipid levels (Roelants et al. 2011; Han et al. 2010; Breslow et al. 2010). The regulation involves TORC2-dependent phosphorylation of ORM to gradually relieve ORM suppression of SPT to enhance LCB synthesis in response to sub-optimal intracellular sphingolipid levels and reversible dephosphorylation of ORM by SAC1 phosphatase to engage ORM suppression of SPT activity in response to excess intracellular sphingolipid levels (Muir et al. 2014; Roelants et al. 2011). Arabidopsis has two homologs of the yeast ORM genes, ORM1 (At1g01230) and ORM2 (At5g42000). Although these proteins have not yet been characterized, RNAi suppression of ORM genes in rice results in temperature sensitivity and pollen abnormalities (Chueasiri et al. 2014).
Regulation of SPT by ORM proteins in plants and other eukaryotes appears to be more complex than ORM phosphorylation/dephosphorylation in yeast. As described above, the primary regulatory mechanism of ORM in yeast occurs through TORC2 dependent YPK1 phosphorylation of ORM that relieves inhibition of SPT that can be reversed by SAC1 phosphatase activity that restores inhibition of SPT (Han et al. 2010; Muir et al. 2014). This mechanism is adjustable and dependent on intracellular sphingolipid levels and has been shown to be coordinated with ceramide synthase activity (Muir et al. 2014). An N-terminal extension of approximately 80 amino acids in yeast ORM was found to contain several Ser residues that are responsible for this phosphorylation mechanism (Roelants et al. 2011). This phosphorylation domain, however, is absent in plant and mammalian ORM homologs (Roelants et al. 2011). As a result, it is unclear if ORM phosphorylation/dephosphorylation also regulates SPT activity in plants and mammals. Overall, homeostatic regulation of SPT in plants and mammals remains an open and active area of research.
3-Ketosphinganine Reductase
In the second step of LCB synthesis, the SPT product 3-ketosphinganine is reduced by the enzyme 3-ketosphinganine reductase (KSR ) to form sphinganine (d18:0), the simplest LCB found in plants (Fig. 11.3 and Table 11.1). KSR is encoded by two genes in Arabidopsis thaliana, KSR-1 (At3g06060) and KSR-2 (At5g19200). Both genes are essential and contribute to the reductase activity (Chao et al. 2011), although KSR-1 is more highly expressed throughout the plant (Chao et al. 2011). KSR-1 and KSR-2 are functionally redundant, but KSR-1 is the primary contributor to the reductase activity (Chao et al. 2011). Loss-of-function mutants of KSR-1 are viable but display greatly reduced reductase activity (Chao et al. 2011). These mutants also display an altered leaf ionome that is associated with increased root suberization, altered root morphology, and altered root iron homeostasis (Chao et al. 2011). The sphinganine (d18:0) produced from the combined activities of SPT and KSR can be used directly by ceramide synthase or modified by hydroxylation or desaturation at the C-4 position prior to use for ceramide synthesis.
LCB Modifications
LCB C-4 Hydroxylation
The d18:0 LCB resulting from the sequential activities of SPT and KSR can undergo combinations of three modification reactions to generate trihydroxylated and unsaturated LCBs (Figs. 11.1a, 11.3, and Table 11.1). In Arabidopsis leaves, ~90 % of the total LCBs contain three hydroxyl groups and Δ8 unsaturation. The third hydroxyl group of these LCBs occurs at the C-4 position and is introduced by a LCB C-4 hydroxylase (Chen et al. 2008; Sperling et al. 2001). This enzyme is a di-iron oxo protein with homology to desaturases and hydroxylases (Sperling et al. 2001). The two genes that encode the LCB C-4 hydroxylase in Arabidopsis are designated SPHINGOID BASE HYDROXYLASE (SBH)1 (At1g69640) and 2 (At1g14290). Expression of these genes in mutants of the Saccharomyces cerevisiae SUR2 gene (Haak et al. 1997) that encodes a related LCB C-4 hydroxylase restores trihydroxy LCB synthesis (Chen et al. 2008; Sperling et al. 2001). It is presumed that the Arabidopsis LCB C-4 hydroxylase uses a free dihydroxy LCB as its substrate, in part, because of the prevalence of trihydroxy LCBs in the free LCB pool (Markham and Jaworski 2007).
LCB Δ8 Desaturation
LCBs with Δ8 unsaturation , either in the dihydroxy or trihydroxy form, are also abundant in sphingolipids of most plant species (Lynch and Dunn 2004) (Figs. 11.1a and 11.3). Like the LCB C-4 hydroxylase, LCB Δ8 desaturases are di-iron oxo enzymes (Shanklin and Cahoon 1998). The plant Δ8 LCB desaturase was originally identified in sunflower as a desaturase-like enzyme that also contains an N-terminal cytochrome b5 domain and shown to confer production of Δ8 unsaturated LCBs when expressed in Saccharomyces cerevisiae (Sperling et al. 1995). Notably, the LCB Δ8 desaturase is not found in mammals and Saccharomyces cerevisiae, but is present in plants and filamentous or dimorphic fungi such as Pichia patoris and Yarrowia lipolytica. Two homologs, SLD1 (At3g61580) and SLD2 (At2g46210), were identified in Arabidopsis and confirmed to be Δ8 desaturases through yeast and in planta studies (Sperling et al. 1998; Chen et al. 2012). To further add to the structural diversity found in LCBs, the Δ8 double bond can be introduced in either the cis or trans configuration (Markham et al. 2006), which likely results from presentation of LCB substrates in alternative conformations relative to the di-iron oxo atoms in the active site of these enzymes (Beckmann et al. 2002). Though evidence to date cannot preclude that at least a portion of LCB Δ8 desaturation uses free LCBs as substrates, it is presumed that these enzymes largely use LCBs bound in ceramides as substrates (Beckmann et al. 2002; Sperling et al. 1998).
LCB Δ4 Desaturation
Long-chain bases (LCBs) with Δ4 unsaturation are also prevalent in sphingolipids in many plant species. LCB Δ4 unsaturation occurs almost entirely in combination with LCB Δ8 unsaturation in dihydroxy LCBs. These di-unsaturated, dihydroxy LCBs (d18:2) also are found almost exclusively in ceramides of GlcCer, but absent from ceramides of GIPCs (Markham and Jaworski 2007; Markham et al. 2006; Sperling et al. 2005) (Figs. 11.1a, c, 11.3, and Table 11.1). Arabidopsis contains one Δ4 desaturase gene (At4g04930) that was identified by homology to analogous genes in filamentous fungi and mammals (Ternes et al. 2002). In contrast to the LCB Δ8 desaturase, the Δ4 desaturase introduces double bonds exclusively in the trans configuration, most likely using free LCBs as substrates (Ternes et al. 2002). As a result, two d18:2 isomers occur in plants: d18:2-transΔ4, transΔ8 and d18:2-transΔ4, cisΔ8. It is notable that LCB C-4 hydroxylases and LCB Δ4 desaturase can both use d18:0 as substrates. As a result, C-4 hydroxylation precludes Δ4 desaturation, and conversely, Δ4 desaturation prevents C-4 hydroxylation. In Arabidopsis and likely other Brassicaceae, the LCB Δ4 desaturase gene has little or no expression in leaves (Michaelson et al. 2009). Instead, expression is limited almost entirely to flowers and, specifically, pollen, which is consistent with the occurrence of d18:2 in Arabidopsis organs (Michaelson et al. 2009). In most species outside of the Brassicaceae family, LCB Δ4 desaturation, as evidenced by d18:2 production, occurs throughout the plant, and in species such as tomato and soybean, d18:2 is the most abundant LCB in GlcCer (Markham et al. 2006; Sperling et al. 2005).
Influence of Long-Chain Base Hydroxylation and Desaturation on Metabolic Routing
Evidence that has emerged from sphingolipid compositional profiling of Arabidopsis mutants indicates that LCB hydroxylation and desaturation affect metabolic outcomes in sphingolipid biosynthesis. LCB C-4 hydroxylase mutants, for example, accumulate high levels of sphingolipids with ceramide backbones containing C16 fatty acids and dihydroxy LCBs, rather than the more typical ceramides with very long-chain fatty acids and trihydroxy LCBs (Chen et al. 2008). As discussed below, this metabolic phenotype arises from the substrate preferences of ceramide synthases. In addition, Arabidopsis sld1sld2 double mutants lacking LCB Δ8 unsaturation have a 50 % reduction of GlcCer, perhaps due to the substrate specificity of GlcCer synthase (Chen et al. 2012). Similarly, Arabidopsis mutants for the LCB Δ4 desaturase, have a 50 % reduction in GlcCer in pollen (Michaelson et al. 2009). This phenotype is more extreme in LCB Δ4 mutants of the yeast Pichia pastoris (Michaelson et al. 2009). Disruption of this LCB Δ4 desaturase in P. pastoris results in nearly complete loss of GlcCer, likely due to a ceramide synthase that exclusively uses LCBs with Δ4 unsaturation and is dedicated to GlcCer synthesis (Michaelson et al. 2009).
In addition to metabolic alterations, loss of LCB C-4 hydroxylation and Δ8 desaturation affects plant performance. LCB C-4 hydroxylase mutants have impaired growth and constitutive up-regulation of PCD (Chen et al. 2008). In addition, Arabidopsis mutants lacking LCB Δ8 unsaturation are more sensitive to low temperature (Chen et al. 2012), and the relative ratio of cis-trans Δ8 unsaturation affects resistance of Arabidopsis to aluminum (Ryan et al. 2007). However, Arabidopsis LCB Δ4 desaturase mutants have no detectable impairment of pollen viability or germination or other growth phenotypes, despite the reduction of GlcCer levels (Michaelson et al. 2009). These findings bring into question the quantitative significance of GlcCer for plant performance.
Sphingolipid Fatty Acid Synthesis and Structural Modifications
Carbon chain-length, unsaturation, and hydroxylation of fatty acids also contribute to the structural diversity of the ceramide backbone of sphingolipids (Table 11.1). In plants, the fatty acid component ranges from 16 to 26 carbon atoms (Markham and Jaworski 2007), including small amounts of odd-chain fatty acids with 21, 23, and 25 carbon atoms (Cahoon and Lynch 1991). In Arabidopsis leaves, C16, C24, and C26 fatty acids predominate (Markham and Jaworski 2007; Markham et al. 2006). The C16 fatty acids of ceramides arise from palmitic acid formed by de novo fatty acid synthesis, whereas the very long-chain fatty acids or VLCFAs (i.e., fatty acids with ≥C20) of sphingolipids arise from the ER-localized reactions involving the two-carbon sequential elongation of fatty acids produced de novo in plastids (Smith et al. 2013). Each two carbon elongation cycle involves the four successive reactions catalyzed by 3-ketoacyl-CoA synthase (KCS), 3-ketoacyl-CoA reductase (KCR), hydroxyacyl-CoA dehydratase (HACD), and enoyl-CoA reductase (ECR) (Smith et al. 2013). Interestingly, KCS enzymes do not occur in Saccharomyces cerevisiae (Paul et al. 2006). Instead, fatty acid elongation is initiated by ELO enzymes that are structurally unrelated, but functionally equivalent to KCS (Paul et al. 2006). Four ELO homologs occur in Arabidopsis, but no findings to date link these enzymes to the synthesis of sphingolipid VLCFAs (Haslam and Kunst 2013). Arabidopsis mutants of the PAS2 gene (At5g10480) encoding HACD are defective in VLCFA synthesis and have demonstrated the importance of sphingolipid VLCFAs for cellular function. Partial PAS2 mutants are defective in growth and phragmoplast (or cell plate) formation resulting in impaired cell division, and null PAS2 mutants display embryo lethality (Bach et al. 2008, 2011).
Sphingolipid VLCFAs are typically saturated in the plant kingdom, but monounsaturated VLCFAs occur in sphingolipids of Brassicaceae and some Poaceae species as well as selected species from other families (Cahoon and Lynch 1991; Lynch and Dunn 2004; Markham et al. 2006; Sperling et al. 2005). The double bond in sphingolipid VLCFAs of these species is at the ω-9 position (Imai et al. 2000). In Arabidopsis, this double bond is introduced by an enzyme encoded by ADS2 (At2g31360) that has homology to acyl-CoA desaturases (Smith et al. 2013). The ADS2 gene is induced by low temperatures and ADS2 null mutants display chilling sensitivity, indicating a link between sphingolipid structure and low temperature performance, as also shown for the LCB Δ8 desaturase (Chen and Thelen 2013). Notably, ADS2-encoded enzymes do not occur in Poaceae and other monocots (unpublished observation), suggesting that a distinct pathway has evolved for biosynthesis of monounsaturated VLCFA synthesis in Poaceae.
Fatty acids in ceramides of glycosphingolipids occur almost entirely with C-2 or α-hydroxylation (Lynch and Dunn 2004). The C-2 hydroxyl group is introduced by a di-iron-oxo enzyme related to the Saccharomyces cerevisiae fatty acid C-2 hydroxylase encoded by the FAH1 or SCS7 gene (Haak et al. 1997; Mitchell and Martin 1997). The Arabidopsis homologs AtFAH1 (encoded by At2g34770) and AtFAH2 (encoded by At4g20870) notably lack the N-terminal cytochrome b5 domain that is found in the Saccharomyces cerevisiae enzyme (Konig et al. 2012; Mitchell and Martin 1997; Nagano et al. 2012). Based on phenotypes in T-DNA insertion mutants and RNAi suppression lines, AtFAH1 appears to be primarily associated with hydroxylation of VLCFAs, and AtFAH2 appears to be primarily associated with hydroxylation of C16 fatty acids in planta (Nagano et al. 2012). It is presumed that AtFAH1 and AtFAH2 use fatty acids in ceramides rather than free or CoA esters of fatty acids as substrates, given that a substantial portion of fatty acids in the free ceramide pool lack C-2 hydroxylation, even though hydroxylated fatty acids predominate in glycosphingolipid ceramide backbones (Markham and Jaworski 2007). Double mutants of the AtFAH1 and AtFAH2 genes have elevated levels of ceramides but ~25 % reduction in glucosylceramide level (Konig et al. 2012). These results suggest that ceramides with C-2 hydroxylated fatty acids are important for metabolic channeling of ceramides to form glycosphingolipids, due possibly to the substrate preference of enzymes such as glucosylceramide synthase (see below). Suppression of PCD by ER-associated Bax inhibitor-1 protein in Arabidopsis has been shown to be dependent on functional fatty acid C-2 hydroxylases, and overexpression of the Bax inhibitor 1 gene increases fatty acid C-2 hydroxylation of ceramides through direct interaction with cytochrome b5 (Nagano et al. 2009, 2012). From these findings, it has been speculated that accumulation of ceramides with fatty acids lacking the C-2 hydroxyl group initiates PCD, whereas this response is reduced when the fatty acids of these ceramides are hydroxylated (Nagano et al. 2012).
Ceramide Synthesis
Ceramides are synthesized by the condensation of a long-chain base and fatty acyl-CoA through an acyltransferase-type reaction catalyzed by ceramide synthase (Fig. 11.3 and Table 11.1). Three ceramide synthases have been identified in Arabidopsis through homology with the yeast ceramide synthase encoded by LAG1 (LONGEVITY ASSURANCE GENE1). These enzymes are designated Lag One Homolog (LOH)-1, -2, and -3 and correspond to genes encoded by LOH1, At3g25440; LOH2, At3g19260; and LOH3, At1g13580, respectively (Ternes et al. 2011; Markham et al. 2011). Homologs of these three enzymes are found throughout the plant kingdom and appear to form two distinct evolutionary branches, LOH1/LOH3-related isoforms and LOH2-related isoforms (Markham et al. 2011; Ternes et al. 2011). Arabidopsis LOH1 and LOH3 share approximately 90 % amino acid sequence identity, while LOH2 shares approximately 60 % identity with LOH1 and LOH3 (Ternes et al. 2011; Markham et al. 2011). In other mammals, multiple ceramide synthases occur that have distinct specificity for fatty acyl-CoAs and/or long-chain bases (Venkataraman et al. 2002; Laviad et al. 2008; Mizutani et al. 2005, 2006; Riebeling et al. 2003). Studies of Arabidopsis LCB C-4 hydroxylase mutants initially pointed to the likelihood that two functional classes of ceramide synthases occur in plants (Chen et al. 2008). Loss of or reduced LCB C-4 hydroxylation has been shown to result in the aberrant accumulation of high levels of sphingolipids with ceramides containing C16 fatty acids bound to dihydroxy LCBs (Chen et al. 2008). Based on this observation, it was proposed that Arabidopsis has one class of ceramide synthase that links C16 fatty acyl-CoAs with dihydoxy LCBs (termed “Class I”), and a second class (“Class II”) that primarily links very long-chain fatty acyl CoAs with trihydroxy LCBs (Chen et al. 2008) (Fig. 11.4). This prediction was supported by the identification, biochemical and genetic characterization of LOH1, LOH2, and LOH3 in Arabidopsis. Studies using yeast complementation showed that LOH2 prefers C16 acyl-CoAs, similar to the predicted Class I ceramide synthase (Ternes et al. 2011). Similarly, Arabidopsis LOH2 mutants were found to be deficient in sphingolipids with ceramide backbones containing C16 fatty acids and dihydroxy fatty acids (Markham et al. 2011). Consistent with the substrate properties of Class II ceramide synthase, partial knock-out mutants of LOH1 and LOH3 contained reduced amounts of ceramides with very long-chain fatty acids and trihydroxy LCBs (Markham et al. 2011). It is notable that under ideal growth conditions, null mutants of LOH2 are viable, suggesting that the Class I ceramide synthase and hence ceramides with C16 fatty acids and dihydroxy LCBs are not essential in Arabidopsis (Markham et al. 2011). Conversely, double null mutants of LOH1 and LOH3 were not recoverable, indicating that the Class II ceramide synthase and ceramides with very long-chain fatty acids and trihydroxy LCBs are essential (Markham et al. 2011).
Model of ceramide synthase mediated long-chain base (LCB) and fatty acid routing. The Arabidopsis gene names are shown as reference. As indicated, Class I ceramide synthase (CSI) encoded by LOH2 displays strict substrate specificity of C16 fatty acid acyl-CoAs and dihydroxy LCBs, and Class II ceramide synthase (CSII) encoded by LOH1 or LOH3 display strict substrate specificity for very long-chain fatty acyl-CoAs and trihydroxy LCBs. One or more products of the CSII pathway appear to negatively regulate serine palmitoyltransferase (SPT) activity. In addition, sphingolipids with ceramides from the CSI pathway do not support growth, while those from the CSII pathway are essential for plant growth. The mycotoxin fumonisin B1 (FB 1) appears to preferentially inhibit CSII enzymes. KSR, 3-ketosphinganine reductase; SBH, LCB C-4 hydroxylase
Ceramide synthases are known targets for competitive inhibition by sphinganine analog mycotoxins (SAMs) such as fumonisin B1 or FB1 produced by a variety of Fusarium species and AAL toxin produced by Alternaria alternata f. sp. lycopersici (Abbas et al. 1994). These compounds, particularly FB1, have been widely used as tools for induction of programmed cell death (PCD) in plants, presumably due to the accumulation of cytotoxic LCBs from their inhibition of ceramide synthases (Stone et al. 2000). Recent evidence using FB1 treatment of Arabidopsis ceramide synthase mutants has suggested that FB1 is a more potent inhibitor of Class II ceramide synthases (i.e. LOH1 and LOH3 ceramide synthases) (Markham et al. 2011). Interestingly, in addition to accumulation of free LCBs, elevated levels of ceramides with C16 fatty acids and dihydroxy LCBs formed by Class I ceramide synthases (i.e. LOH2 ceramide synthase) are detectable following treatment of Arabidopsis with FB1 (Markham et al. 2011). These results suggest that FB1 cytotoxicity and PCD induction may be triggered by accumulated ceramides rather than or in addition to accumulated LCBs. FB1 has also been used as a tool to study sphingolipid homeostasis in plants based on the observation that down-regulation of serine palmitoyltransferase (SPT) activity reduces FB1 cytotoxicity and up-regulation of SPT activity enhances sensitivity of plants to FB1 (Kimberlin et al. 2013; Shi et al. 2007).
Glucosylceramide Synthesis
Following its synthesis by Class I or Class II ceramide synthases, the ceramide backbone can be glycosylated at its C-1 OH to form either of two classes of glycosphingolipids: glucosylceramides (GlcCer) or glycosyl inositolphosphoceramides (GIPCs) (Fig. 11.3 and Table 11.1). GlcCer are the simplest glycosphingolipid and occur broadly in eukaryotes, with the notable exception of Saccharomyces cerevisiae (Lynch and Dunn 2004). GlcCer consist of a glucose bound to the ceramide backbone by a 1,4-glycosidic linkage and are formed by the condensation of a ceramide substrate with UDP-glucose (Leipelt et al. 2001). This reaction is catalyzed by GlcCer synthase, an ER-localized enzyme in Arabidopsis that is encoded by At2g19980 (Melser et al. 2010). Compared to GIPCs, GlcCer are more enriched in ceramides with C16 fatty acids and dihydroxy LCBs (Markham et al. 2006; Sperling et al. 2005). In plants such as tomato and soybean, ceramides with C16 fatty acids and the LCB d18:2 predominate (Markham et al. 2006; Sperling et al. 2005). Based on this composition, it appears that a large portion of the GlcCer ceramide backbone is channeled from Class I-type ceramide synthases that have substrate preference for C16 fatty acids and dihydroxy LCBs (Markham et al. 2011). Although it is an abundant glycosphingolipid in plants, null mutants of the LCB Δ4 deasaturase in Arabidopsis have 30 % reductions in GlcCer levels in flowers (Michaelson et al. 2009) and 50 % reduction in GlcCer levels in pollen (Luttgeharm et al. 2015) without any apparent effect on flower and pollen physiology and function (Michaelson et al. 2009). GlcCer synthase is potently inhibited by d,l-threo-1-phenyl-2-decanoylamino-3-morpholino-1-propanol (PDMP) (Melser et al. 2010). Treatment of Arabidopsis roots with PDMP results in altered Golgi morphology, including reduced numbers of Golgi stacks, and defects in endomembrane trafficking (Melser et al. 2010, 2011). PDMP application to Arabidopsis root cells has also been shown to result in rapid vacuolar fusion and altered vacuole morphology including the appearance of vacuolar invaginations (Kruger et al. 2013). Arabidopsis GlcCer synthase mutants devoid of GlcCer have yet to be described. Such mutants will clarify whether GlcCer are essential in plants, which is an open question because viable fungal cells can be recovered that lack GlcCer (Michaelson et al. 2009; Rittenour et al. 2011).
Inositolphosphoceramide Synthesis
As an alternative fate to GlcCer synthesis, ceramides can be used for the production of GIPCs. GIPCs, which are approximately twofold more abundant in Arabidopsis leaves than GlcCer, are typically enriched in ceramides with VLCFAs and trihydroxy LCBs that arise from Class II ceramide synthases (Markham et al. 2006). The first step in GIPC synthesis occurs by the transfer of the inositolphosphoryl head group of phosphatidylinositol (PI) onto ceramide to form inositolphosphoceramides (IPCs) (Mina et al. 2010; Wang et al. 2008) (Fig. 11.3 and Table 11.1). This activity is catalyzed by IPC synthase, a phosphatidic acid phosphatase-2 (PAP2)-related enzyme, that is encoded by three genes in Arabidopsis: IPCS1 (or ERH1), At2g37940; IPCS2, At2g37940; IPCS3, At2g29525 (Mina et al. 2010; Wang et al. 2008). In contrast to the ER localization of GlcCer synthase, IPC synthases predominantly occur in Golgi bodies of Arabidopsis. Plant IPC synthases are most closely related to analogous enzymes in the protozoa Leishmania major and Trypanosoma brucei than to the Saccharomyces cerevisiae IPC synthase (encoded by the AUR1 gene) (Mina et al. 2010; Wang et al. 2008). Despite this, the three Arabidopsis IPC synthase genes are able to rescue lethality associated with the loss of IPC production in the Saccharomyces cerevisiae AUR1 mutant (Mina et al. 2010; Wang et al. 2008). Although triple mutants of the three Arabidopsis IPC synthase genes have not been reported, it is presumed that IPC biosynthesis is essential, as the three genes are likely partially redundant.
Following the synthesis of IPC, up to seven additional sugar residues can be added to the inositolphosphoryl head group to form an array of different GIPCs (Bure et al. 2011; Cacas et al. 2013). The first residue added to the inositophosphoryl head group is a glucuronic acid moiety (Rennie et al. 2014). This reaction, which uses a UDP-glucuronic acid substrate, was recently shown to be catalyzed by a glycosyltransferase encoded by IPUT1 (At5g18480) in Arabidopsis (Rennie et al. 2014). T-DNA null mutants of IPUT1 are not transmitted through pollen, indicating that this gene is essential in Arabidopsis (Rennie et al. 2014). The remaining glycosyltransferases associated with GIPC synthesis have yet to be identified. Interestingly, a Golgi lumen-localized GDP-mannose transporter encoded by GOLGI-LOCALIZED NUCLEOTIDE SUGAR TRANSPORTER 1 (GONST1, At2g13650) in Arabidopsis was found to be deficient in GIPCs with glycosylation beyond the glucuronic acid introduced by the IPUT1-encoded glycosyltransferase (Mortimer et al. 2013). This suggests that like Saccharomyces cerevisiae, Arabidopsis GIPCs contain mannose, presumably bound to the glucuronic acid moiety. Interestingly, GONST1 mutants display a dwarfed phenotype, constitutive induction of the hypersensitive response, and elevated salicylic acid levels, consistent with a connection between GIPC synthesis and plant pathogen defense (Mortimer et al. 2013).
LCB and Ceramide Phosphorylation/Dephosphorylation
In addition to their occurrence in free form and in ceramides, LCBs are also detectable in low levels as phosphate derivatives that have been attributed to triggers of physiological responses, such as ABA-dependent guard cell closure (Coursol et al. 2003; Ng et al. 2001) (Figs. 11.5, 11.6 and Table 11.1). Phosphorylation of LCBs at their C-1 hydroxyl group is catalyzed by LCB kinases (often referred to sphingosine kinases or SPHKs). To date three LCB kinases have been identified in Arabidopsis: SPHK1 (At5g23450), SPHK2 (At2g46090), and AtLCBK1 (At5g23450) (Imai and Nishiura 2005; Worrall et al. 2008; Guo et al. 2012). Release of the phosphate group from LCB-P molecules is catalyzed by the enzyme LCB-P phosphatase, which are encoded by two genes in Arabidopsis (At3g58490 and At5g03080) (Nakagawa et al. 2012; Worrall et al. 2008). As described below, the interplay between LCB kinases and LCB-P phosphatases are believed to be important for signaling pathways in plants (Nakagawa et al. 2012; Worrall et al. 2008).
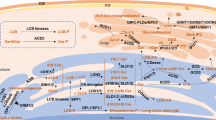
Sphingolipid catabolic and ceramide and long-chain base phosphorylation/dephosphorylation pathways. Dashed arrows represent enzymatic steps involved in catabolism. Abbreviations: LCB long-chain base, LCB-P long-chain base-1-phosphate, Glc glucose, PI phosphatidylinositol, DAG diacylglycerol, IP inositolphosphate, IPCase inositolphosphoceramidase
Phosphorylated/dephosphorylated long-chain bases (LCBs) and ceramides serve as mediators of physiological processes in plants. The interplay between LCBs and ceramides and their phosphorylated forms regulates cellular process and responses to environmental stimuli. Abbreviations: LCB long-chain base, LCB-P long-chain base-1-phosphate, ABA Abscisic acid, ROS Reactive oxygen species, NO nitrous oxide
Similar to LCBs, ceramides can also be found in phosphorylated forms (Figs. 11.5, 11.6. and Table 11.1). Although ceramide-1-phosphates are believed to be of low abundance in plants, they have proven difficult to measure by recently developed mass spectrometry-based protocols. Mutants of the proposed ceramide kinase (encoded by At5g51290), termed accelerated death 5 or acd5 display spontaneous onset of programmed cell death or PCD in late development (Greenberg et al. 2000; Liang et al. 2003). This is accompanied by enhanced accumulation of ceramides (Greenberg et al. 2000; Liang et al. 2003). This observation led to the hypothesis, now accepted as dogma, that elevation of ceramide levels triggers PCD in plants (Greenberg et al. 2000; Liang et al. 2003) via accumulation of mitochondrial-derived hydrogen peroxide (Bi et al. 2014). A ceramide-1-phosphate phosphatase that would convert ceramide-1-phosphates to their free form has yet to be identified in plants.
Sphingolipid Turnover
The net content and composition of sphingolipids in membranes are determined by rates of synthesis and turnover. Little is currently known about rates of sphingolipid turnover, and the contributions of sphingolipid catabolism to membrane function and plant responses to altered environmental conditions. Also unexplored to date in plants are enzymes associated with removal of glycosphingolipid head groups, although candidate genes have been proposed (Chen et al. 2010). More is known about ceramide turnover. Enzymes referred to as ceramidases convert ceramides to free LCBs and fatty acids. Ceramidases are classified into three distinct forms based upon their optimal pH preferences in in vitro assays: acid , neutral , and alkaline ceramidases (Mao and Obeid 2008). Three predicted neutral ceramidase genes and one predicted alkaline ceramidase have been identified by homology with human ceramidase genes (Chen et al. 2010; Wu et al. 2015). The Arabidopsis alkaline ceramidase homolog, AtACER (At4g22330), has been shown to function as a ceramidase with mutant, and RNAi suppression lines for this gene had elevated ceramide levels and increased salt sensitivity and enhanced susceptibility to a bacterial pathogen (Wu et al. 2015). A second gene TOD1 corresponding to At5g46220 was recently shown to encode a polypeptide with alkaline ceramidase activity, but notably, lacked close homology to known alkaline ceramidases (Chen et al. 2015). Based on mutant phenotypes, this enzyme was linked to control of turgor pressure in pollen tubes and silique guard cells (Chen et al. 2015). A neutral ceramidase has also been cloned from rice and confirmed in vitro to be a member of the neutral ceramidase subclass (Pata et al. 2008) (Fig. 11.5 and Table 11.1).
LCBs, including those released by ceramidase activity, can be degraded following phosphorylation by LCB kinases. This process is catalyzed by LCB-P lyase (often referred to as DPL1, based on homology to the yeast enzyme), which generates C16-fatty aldehyde and phosphoethanolamine from a C18 LCB-P. Arabidopsis contains only a single DPL1 gene (At1g27980) (Nishikawa et al. 2008; Tsegaye et al. 2007; Worrall et al. 2008) that is constitutively expressed and strongly upregulated by senescence (Tsegaye et al. 2007). Null mutants of DPL1 displayed small increases in accumulation of the LCB-P t18:1-P, but surprisingly no obvious growth phenotypes (Tsegaye et al. 2007) (Fig. 11.5 and Table 11.1).
Sphingolipids and Membrane Structure-Function
Membrane Function
Sphingolipids compose an estimated ~40 % of the total lipids in plasma membrane of plants (Sperling et al. 2005), where they are enriched in the outer leaflet (van Genderen et al. 1991; Burger et al. 1996; Tjellstrom et al. 2010). Sphingolipids are also abundant lipid components of other endomembranes in the plant, including ER, Golgi, and tonoplast (Bayer et al. 2014; Mongrand et al. 2004; Sperling et al. 2005; Verhoek et al. 1983). GlcCer was first identified in the plasma membrane and tonoplast of plant cells in a number of studies conducted in the 1980s (Cahoon and Lynch 1991; Lynch and Steponkus 1987; Verhoek et al. 1983; Yoshida et al. 1986). In these membranes, GlcCer was reported to compose between 7 and 30 % of plasma membrane and tonoplast, depending on the plant species and tissue type analyzed (Cahoon and Lynch 1991; Lynch and Steponkus 1987; Uemura et al. 1995; Uemura and Steponkus 1994; Verhoek et al. 1983; Yoshida et al. 1986). More recently, it has been shown that GIPCs, rather than GlcCer, are the more abundant glycosphingolipid in plants (Markham and Jaworski 2007; Markham et al. 2006). The quantitative importance of GIPCs was largely overlooked until very recently because their highly glycosylated head groups offer challenges for extraction using typical organic solvents, such as mixtures of chloroform and methanol (Markham et al. 2006). Initially, by quantification of long-chain bases of GIPCs and GlcCer and later by LC-MS/MS analysis, GIPCs were found to be nearly twofold more abundant than GlcCer in Arabidopsis leaves, whereas amounts of GIPCs and GlcCer were nearly the same in tomato leaves (Markham and Jaworski 2007; Markham et al. 2006). GIPCs have subsequently been identified as one of the most abundant lipids of plant plasma membrane and are also enriched in detergent resistant membranes (DRMs) derived from isolated plasma membrane and in plasmodesmata (Grison et al. 2015; Cacas et al. 2012). Given their abundance in plasma membrane and tonoplast, it is likely that the content and composition of sphingolipids affect the ability of plants to respond to abiotic stress, particularly osmotic stresses such as freezing, drought, and salinity. For example, GlcCer concentrations were shown to decrease by nearly half in plasma membrane during cold acclimation of rye and Arabidopsis (Lynch and Steponkus 1987; Uemura et al. 1995; Uemura and Steponkus 1994). More recently, it was reported that GIPCs increase and GlcCer decrease in response to chilling of Arabidopsis (Nagano et al. 2014). Although this is likely an adaptive response to low temperatures, the impact of such adjustments in relative amounts of GIPCs and GlcCer on plant performance has not yet been established. In addition, the fatty acid and long-chain base composition of sphingolipids also affects plant resistance to abiotic stress. For example, Arabidopsis mutants lacking LCB Δ8 and ceramide fatty acid unsaturation display sensitivity to low temperature growth and alterations in the relative amounts of LCB cis-trans Δ8 unsaturation affects resistance of plants to aluminum (Chen et al. 2012; Chen and Thelen 2013; Ryan et al. 2007).
The unique structural components of sphingolipid hydrophobic ceramide backbones include VLCFAs and an abundance of hydroxyl groups distributed between the LCB and fatty acid moieties. Through these structural features, sphingolipids confer rigidity to membranes. In addition, the hydroxyl groups enable the formation of extensive hydrogen bonding networks that result in elevated phase transition temperatures and reduced ion permeability (Lunden et al. 1977; Pascher 1976). The rigidity and high phase transition temperatures of sphingolipid micelles is moderated by interactions with other lipids, including sterols (Curatolo 1987). Sphingolipids have been shown to cluster with sterol in membrane microdomains or lipid rafts (Cacas et al. 2012). Lipid microdomains have long been hypothesized to be present in membranes (Karnovsky et al. 1982) with sphingolipids potentially aiding in the sorting of membrane proteins, such as GPI-anchored proteins, by forming lipid domains that slow lateral protein diffusion (Simons and van Meer 1988; van Meer and Simons 1988; Brown and Rose 1992; Simons and Ikonen 1997; de Almeida et al. 2003). Indeed, pure sphingolipid membranes form a ‘solid gel’ phase with little lateral movement that is fluidized by the presence of sterols (van Meer et al. 2008; Estep et al. 1980; Roche et al. 2008; Grosjean et al. 2015). The co-localization of sterols and sphingolipids in the membrane may be due to sphingolipids complex sugar head group’s ability to shield the non-polar sterol from the bulk solvent, much like an umbrella (Huang and Feigenson 1999; Ali et al. 2006). Raft formation is also dependent on the various sphingolipid modifications, including fatty acid chain length and fatty acid and LCB hydroxylation (Klose et al. 2010).
Proteomic analysis of DRMs prepared from plant plasma membrane has revealed an enrichment of proteins in lipid rafts related to signaling, responses to biotic and abiotic stress, cellular trafficking, auxin transport, and cell wall synthesis and degradation (Brodersen et al. 2002; Lefebvre et al. 2007; Lin et al. 2008; Morel et al. 2006), suggesting that raft regions contribute to these cellular functions. Similar to DRMs, plasmodesmata have also recently been shown to be enriched in sphingolipids and sterols and contain specific GPI-anchored proteins (Bayer et al. 2014; Grison et al. 2015).
Endomembrane Trafficking
Given their abundance in the endomembrane system, sphingolipids are presumed to play a major role in ER export and Golgi-mediated trafficking of proteins through the secretory system. Consistent with this, Arabidopsis pollen deficient in sphingolipids have been shown to have vesiculated ER and lack Golgi stacks (Dietrich et al. 2008). Consistent with defects in Golgi-trafficking to the plasma membrane, the sphingolipid-deficient pollen lacked a surrounding intine layer (Dietrich et al. 2008). Similarly, chemical inhibition of GlcCer synthesis has been shown to alter Golgi morphology and impair Golgi-mediated trafficking of secretory proteins to the plasma membrane (Melser et al. 2010, 2011). Recent studies using an Arabidiopsis KSR mutant and GlcCer synthase and ceramide synthase inhibitors demonstrated the importance of sphingolipids in trafficking of ATP-binding cassette B19 (ABCB19) auxin transporter to the Golgi, trans-Golgi network, and plasma membrane (Yang et al. 2012). Similar studies targeting sterols indicated that sterols have greater significance for post-Golgi transport of ABCB19 from trans-Golgi to the plasma membrane (Yang et al. 2012). Of particular importance for trafficking of proteins through the secretory system is the presence of the very long-fatty acid (VLCFA) component of sphingolipids, which are enriched in GIPCs. Mutants defective in VLCFA synthesis or that have reduced activity of Class II ceramide synthases that incorporate VLCFAs in ceramides have impaired trafficking of secretory proteins, including PIN1 and AUX1 that are required for auxin transport and plant growth (Bach et al. 2008, 2011; Markham et al. 2011; Zheng et al. 2005).
Sphingolipids as Physiological Mediators
ABA-Dependent Guard Cell Closure
In addition to the contributions of the glycosphingolipids GIPCs and GlcCer to membrane structure and function, the less abundant sphingolipid biosynthetic intermediates LCBs, LCBPs, ceramides, and ceramide-1-phosphates have been linked to the mediation of numerous, diverse physiological processes in plant cells. An important contributor to the formation of these physiological mediators are kinase and phosphatase reactions that convert LCBs and ceramides between their phosphorylated and free forms, as described above. The phosphorylation status of LCBs and ceramides are key to the particular physiological process that they regulate (Fig. 11.6).
One of the first links between sphingolipids and control of cellular processes was the observation that the LCB-P sphingosine-1-phosphate or S1P participates in the ABA-mediated signaling pathway that controls stomatal aperture by elevating cytosolic Ca2+ levels which, in turn, activates ion channels in guard cell membranes, with the resulting K+ efflux causing loss of guard cell turgor pressure and stomatal closing (Kim et al. 2010). After drought treatment of the plant Commelina communis, S1P levels were found to increase in leaves, and when applied exogenously, S1P resulted in a Ca2+ spike followed by stomatal closure (Ng et al. 2001). Phytosphingosine-1-phosphate elicits the same response, although sphinganine-1-phosphate does not, indicating some level of LCB specificity in the mediation of guard cell closure (Coursol et al. 2003). Treatment of Arabidopsis plants with ABA was found to activate LCB kinase, and this activity was sensitive to the mammalian LCB kinase inhibitor, N,N-dimethylsphingosine (Coursol et al. 2003). As in mammals, the target of S1P in plants is presumed to be a G-protein coupled receptor, and Arabidopsis mutants lacking the G-protein α-subunit (GPA1) did not respond upon exogenous S1P application (Coursol et al. 2005; Strub et al. 2010). More recently, a connection between phospholipids and sphingolipids in the signaling pathway for ABA-dependent guard cell closure has been proposed. In this regard, ABA and phosphatidic acid (PA) produced by phospholipase Dα1 (PLDα1) have been shown to activate sphingosine kinase (SPK) to promote production of LCB-P (Guo et al. 2012; Guo and Wang 2012; Worrall et al. 2008). PA enhancement of SPK activity was found to occur by direct interaction of PA with this enzyme (Guo et al. 2012; Guo and Wang 2012). Given that LCB-P induction of guard cell closure requires a functional PLDα1, it was proposed that LCB-P functions upstream of PLDα1 in this signaling pathway (Guo et al. 2012; Guo and Wang 2012).
Programmed Cell Death
Sphingolipids, primarily in the form of ceramides and LCBs, have also been strongly implicated in mediation of programmed cell death (PCD ) in plants. As described above, an initial indication of the role of ceramides as PCD triggers was obtained from the Arabidopsis acd5 mutant that is defective in a proposed ceramide kinase (Greenberg et al. 2000; Liang et al. 2003). This mutant accumulates enhanced levels of free ceramides and displays early onset of PCD relative to wild-type controls (Greenberg et al. 2000; Liang et al. 2003), resulting in part to the enhanced release of mitochondrial reactive oxygen species (Bi et al. 2014). PCD induction in the acd11 mutant has also been linked to ceramide accumulation associated with defects in ceramide-1-phosphate transport in this mutant (Simanshu et al. 2014). Similar findings have been obtained by treatment of Arabidopsis cell cultures with C2 ceramide at a concentration of 50 μM (Townley et al. 2005). This treatment induces a transient increase in cytosolic Ca2+and hydrogen peroxide production, followed by cell death, which was reversed by inhibition of Ca2+ release (Townley et al. 2005). These findings implicate Ca2+ as an essential component of ceramide induction of PCD. Notably, C2 ceramides containing 2- or α-hydroxylated fatty acids were not effective in PCD induction in Arabidopsis cell cultures (Townley et al. 2005). Consistent with this observation, the ability of Bax inhibitor-1 (BI-1) to suppress cell death in Arabidopsis is dependent on 2-hydroxylation of ceramide VLCFAs (Nagano et al. 2012).
Similar to results with ceramides, application of the free LCBs d18:1, d18:0, and t18:0 to Arabidopsis leaves also induces PCD, albeit at concentrations lower than that observed with ceramides (Shi et al. 2007). This induction of PCD was also dependent on ROS generation, but was suppressed by application of LCB-P along with free LCBs (Alden et al. 2011; Shi et al. 2007). These findings suggest that the ratio of free LCB to LCB-P, mediated by LCB kinases and LCB-P phosphatases, is an important “rheostat” for regulation of PCD (Fig. 11.6) (Alden et al. 2011; Shi et al. 2007). This is analogous to the dependence of PCD induction on relative levels of ceramides and ceramide-1-phosphates (Greenberg et al. 2000; Liang et al. 2003). The transduction pathway for elicitation of PCD by free LCBs has been shown to be dependent in Arabidopsis on mitogen-activated protein kinase 6 (MPK6) (Saucedo-García et al. 2011) as well as 14-3-3 protein phosphorylation by calcium-dependent kinase 3 (CPK3) that is activated by LCB-triggered release of cytosolic Ca2+ (Lachaud et al. 2013).
Fungal-derived sphingosine-analog mycotoxins (SAMs) including fumonisin B1 produced by Fusarium species and AAL toxin produced by Alternaria alternata lycopersici are also potent triggers of PCD in plants. These molecules competitively inhibit ceramide synthases leading to the accumulation of free LCBs that, in turn, elicit PCD (Abbas et al. 1994). Consistent with this, reduction of LCB synthesis by chemical inhibition of serine palmitoyltransferase (SPT) activity enhances resistance of plants to SAMs (Spassieva et al. 2002). Increased resistance to FB1-triggered PCD induction has also been observed in an Arabidopsis LCB1 mutant and in small subunit of SPT RNAi suppression lines that have reduced SPT activity (Kimberlin et al. 2013; Shi et al. 2007). Recent evidence has also emerged that FB1 not only increases levels of free LCBs in plant cells but also elevates levels of ceramides containing C16 fatty acids (Markham et al. 2011; Ternes et al. 2011). This finding suggests that FB1 most effectively inhibits Class II ceramide synthases (i.e., LOH1, LOH3) that produce ceramides with VLCFAs and are less effective inhibitors of Class I ceramide synthase (i.e., LOH2) that produces ceramides with C16 fatty acids (Markham et al. 2011; Ternes et al. 2011). These findings suggest that the potency of SAMs for PCD-induction is due to their ability to enhance accumulation of LCBs and ceramides.
Pathogen Resistance
The hypersensitive response (HR) is an important process for resistance to bacterial and fungal pathogens that is characterized by localized induction of PCD that reduces or prevents the spread of pathogens in plants. Given the importance of LCBs and ceramides to PCD induction, a considerable body of research has emerged linking sphingolipids to bacterial and fungal pathogen resistance as described in a recent review (Berkey et al. 2012). Notably, ceramide accumulation in acd5 and acd11 mutants has been shown to be associated with salicylic acid (SA)-dependent upregulation of HR-type PCD and pathogen-resistance genes, including genes for PR1, ERD11, and chitinase (Brodersen et al. 2002; Greenberg et al. 2000). More recently, Arabidopsis mutants defective in 2-hydroxylation of ceramide fatty acids were found to have elevated LCB and ceramide levels, as well as, increased levels of free and glycosylated SA and constitutive induction of PR1 and PR2 genes (Konig et al. 2012). These mutants also displayed enhanced resistance to the biotrophic fungal pathogen Golovinomyces cichoracearum (Konig et al. 2012). In addition, infection of Arabidopsis with the bacterial pathogen Pseudomonas syringae was accompanied by transient increases in the LCB phytosphingosine (t18:0) and induction of ROS and cell death (Fig. 11.6) (Peer et al. 2010; Bach et al. 2011). Furthermore, resistance to the bacterial pathogen Pseudomonas cichorii was compromised in tobacco upon chemical inhibition of SPT and an accompanying reduction in LCB synthesis (Takahashi et al. 2009). This resistance appears to be mediated by MPK6, as FB1-elicited Arabidopsis mpk6 mutants displayed reduced resistance to the bacterial pathogen Pseudomonas syringae pv. tomato avrRpm1 due to compromised induction of PCD in this mutant (Saucedo-García et al. 2011).
Cold Stress Signaling
Sphingolipids as abundant components of plasma membrane and tonoplast contribute to the ability of plants to resist chilling and freezing stresses. As evidence of this, Arabidopsis mutants lacking LCB Δ8 unsaturation have increased sensitivity to prolonged exposure to low, non-freezing temperatures (Chen et al. 2012). In addition to their roles as membrane components, recent studies have implicated sphingolipids in cold stress signaling pathways (Cantrel et al. 2011; Guillas et al. 2011, 2013). Exposure of Arabidopsis plants to 4 °C resulted in accumulation of PA and nitrous oxide (NO). In addition, within 5 min of this cold treatment amounts of the LCB-P phytosphingosine phosphate and ceramide-1-phosphate increased by ~50 % (Fig. 11.6) (Cantrel et al. 2011). This increase was negatively regulated by NO, as chemical inhibition of NO production enhanced the accumulation of these molecules but chemically-induced enhancement of NO levels reduced accumulation of the phosphorylated LCB and ceramides (Cantrel et al. 2011). From these findings, it was suggested that NO may regulate the relative levels of phosphorylated and dephosphorylated LCBs and ceramides as part of a rapid signaling response pathway to low, non-freezing temperatures (Cantrel et al. 2011; Guillas et al. 2011). The mechanistic details of this potential signaling pathway remain uncharacterized.
Conclusions
As outlined here, significant advances have been made in plant sphingolipid biology during the past 15 years. Nearly all of the genes involved in sphingolipid synthesis have now been identified and numerous insights have been made in understanding the role of sphingolipids in membrane function and physiological mediation pathways. In addition, the combination of mass spectrometry-based sphingolipid compositional profiling or sphingolipidomics and the characterization of Arabidopsis mutants have revealed the unexpected importance of LCB modifications in regulating flux into glycosphingolipids due, in part, to the narrow substrate specificities of the two functional classes of ceramide synthases. Furthermore, the biological importance of sphingolipids in processes such as programmed cell death and pathogen resistance have been fortuitously discovered by forward genetic screens. Despite the large amount of progress to date, many open questions and challenges remain for plant sphingolipid research. For example, sphingolipid biosynthesis and its regulation have been mostly examined in isolation, and metabolic networks that supply the serine and fatty acid precursors to sphingolipid synthesis have not been examined. We also have yet to understand the interconnections of sphingolipid metabolic and phospholipid and sterol metabolic pathways, particularly in response to environmental stimuli, such as altered growth temperature, that together lead to adjustments in membrane properties. In addition, details of sphingolipid homeostatic regulation in plants have yet to be fully resolved. Also missing in our knowledge is the contributions of sphingolipid turnover and the identities of many of the catabolic enzymes that contribute to mediation of sphingolipid levels in plants. Furthermore, the exact sugar residues, their linkages, and the enzymes that introduce these residues in GIPC head groups have yet to be fully identified. Moreover, it is not currently understood if the distinctive ceramide compositions in GlcCer and GIPCs arise from substrate preferences of GlcCer and IPC synthases or from differential partitioning of specific ceramides to the GlcCer synthase in the ER and to IPC synthase in the Golgi. Similarly, the distinct roles of GlcCer and GIPCs in plants have yet to be elucidated. Finally, an understanding of sphingolipid-associated signaling pathways and their possible interconnections await further resolution. Ultimately, with this information, it will be possible to more precisely genetically tailor sphingolipid metabolism for improved plant performance, particularly in response to environmental perturbations.
References
Abbas HK, Tanaka T, Duke SO, Porter JK, Wray EM, Hodges L, Sessions AE, Wang E, Merrill AH Jr, Riley RT (1994) Fumonisin- and AAL-toxin-induced disruption of sphingolipid metabolism with accumulation of free sphingoid bases. Plant Physiol 106(3):1085–1093
Alden KP, Dhondt-Cordelier S, McDonald KL, Reape TJ, Ng CK, McCabe PF, Leaver CJ (2011) Sphingolipid long chain base phosphates can regulate apoptotic-like programmed cell death in plants. Biochem Biophys Res Commun 410(3):574–580
Ali MR, Cheng KH, Huang J (2006) Ceramide drives cholesterol out of the ordered lipid bilayer phase into the crystal phase in 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine/cholesterol/ceramide ternary mixtures. Biochemistry 45(41):12629–12638
Bach L, Michaelson LV, Haslam R, Bellec Y, Gissot L, Marion J, Da Costa M, Boutin JP, Miquel M, Tellier F, Domergue F, Markham JE, Beaudoin F, Napier JA, Faure JD (2008) The very-long-chain hydroxy fatty acyl-CoA dehydratase PASTICCINO2 is essential and limiting for plant development. Proc Natl Acad Sci U S A 105(38):14727–14731
Bach L, Gissot L, Marion J, Tellier F, Moreau P, Satiat-Jeunemaitre B, Palauqui JC, Napier JA, Faure JD (2011) Very-long-chain fatty acids are required for cell plate formation during cytokinesis in Arabidopsis thaliana. J Cell Sci 124(19):3223–3234
Bayer EM, Mongrand S, Tilsner J (2014) Specialised membrane domains of plasmodesmata, plant intercellular nanopores. Front Plant Sci 5:507
Beaudoin F, Gable K, Sayanova O, Dunn T, Napier JA (2002) A Saccharomyces cerevisiae gene required for heterologous fatty acid elongase activity encodes a microsomal β-keto-reductase. J Biol Chem 277(13):11481–11488
Beckmann C, Rattke J, Oldham NJ, Sperling P, Heinz E, Boland W (2002) Characterization of a Δ8-sphingolipid desaturase from higher plants: a stereochemical and mechanistic study on the origin of E, Z isomers. Angew Chem Int Ed 41(13):2298–2300
Beeler T, Bacikova D, Gable K, Hopkins L, Johnson C, Slife H, Dunn T (1998) The Saccharomyces cerevisiae TSC10/YBR265w gene encoding 3-ketosphinganine reductase is identified in a screen for temperature-sensitive suppressors of the Ca2+-sensitive csg2Δ mutant. J Biol Chem 273(46):30688–30694
Berkey R, Bendigeri D, Xiao S (2012) Sphingolipids and plant defense/disease: the “death” connection and beyond. Front Plant Sci 3:68
Bi FC, Liu Z, Wu JX, Liang H, Xi XL, Fang C, Sun TJ, Yin J, Dai GY, Rong C, Greenberg JT, Su WW, Yao N (2014) Loss of ceramide kinase in Arabidopsis impairs defenses and promotes ceramide accumulation and mitochondrial H2O2 bursts. Plant Cell 26(8):3449–3467
Blacklock BJ, Jaworski JG (2006) Substrate specificity of Arabidopsis 3-ketoacyl-CoA synthases. Biochem Biophys Res Commun 346(2):583–590
Boot RG, Verhoek M, Donker-Koopman W, Strijland A, van Marle J, Overkleeft HS, Wennekes T, Aerts JMFG (2007) Identification of the non-lysosomal glucosylceramidase as β-glucosidase 2. J Biol Chem 282(2):1305–1312
Brandwagt BF, Mesbah LA, Takken FLW, Laurent PL, Kneppers TJA, Hille J, Nijkamp HJJ (2000) A longevity assurance gene homolog of tomato mediates resistance to Alternaria alternata f. sp lycopersici toxins and fumonisin B1. Proc Natl Acad Sci U S A 97(9):4961–4966
Breslow DK, Collins SR, Bodenmiller B, Aebersold R, Simons K, Shevchenko A, Ejsing CS, Weissman JS (2010) Orm family proteins mediate sphingolipid homeostasis. Nature 463(7284):1048–1053
Brodersen P, Petersen M, Pike H, Olszak B, Skov S, Odum N, Jørgensen L, Brown R, Mundy J (2002) Knockout of Arabidopsis accelerated-cell-death11 encoding a sphingosine transfer protein causes activation of programmed cell death and defense. Genes Dev 16(4):490–502
Brown DA, Rose JK (1992) Sorting of GPI-anchored proteins to glycolipid-enriched membrane subdomains during transport to the apical cell surface. Cell 68(3):533–544
Bure C, Cacas JL, Wang F, Gaudin K, Domergue F, Mongrand S, Schmitter JM (2011) Fast screening of highly glycosylated plant sphingolipids by tandem mass spectrometry. Rapid Commun Mass Spectrom 25(20):3131–3145
Burger KN, van der Bijl P, van Meer G (1996) Topology of sphingolipid galactosyl transferases in ER and Golgi: transbilayer movement of monohexosyl sphingolipids is required for higher glycosphingolipid biosynthesis. J Cell Biol 133(1):15–28
Cacas JL, Furt F, Le Guedard M, Schmitter JM, Bure C, Gerbeau-Pissot P, Moreau P, Bessoule JJ, Simon-Plas F, Mongrand S (2012) Lipids of plant membrane rafts. Prog Lipid Res 51(3):272–299
Cacas JL, Bure C, Furt F, Maalouf JP, Badoc A, Cluzet S, Schmitter JM, Antajan E, Mongrand S (2013) Biochemical survey of the polar head of plant glycosylinositolphosphoceramides unravels broad diversity. Phytochemistry 96:191–200
Cahoon EB, Lynch DV (1991) Analysis of glucocerebrosides of rye (Secale cereale L. cv Puma) leaf and plasma membrane. Plant Physiol 95(1):58–68
Cantrel C, Vazquez T, Puyaubert J, Reze N, Lesch M, Kaiser WM, Dutilleul C, Guillas I, Zachowski A, Baudouin E (2011) Nitric oxide participates in cold-responsive phosphosphingolipid formation and gene expression in Arabidopsis thaliana. New Phytol 189(2):415–427
Carter HE, Gigg RH, Law JH, Nakayama T, Weber E (1958) Biochemistry of the sphingolipides: structure of phytoglycolipide. J Biol Chem 233(6):1309–1314
Chao DY, Gable K, Chen M, Baxter I, Dietrich CR, Cahoon EB, Guerinot ML, Lahner B, Lu S, Markham JE, Morrissey J, Han G, Gupta SD, Harmon JM, Jaworski JG, Dunn TM, Salt DE (2011) Sphingolipids in the root play an important role in regulating the leaf ionome in Arabidopsis thaliana. Plant Cell 23(3):1061–1081
Chen M, Thelen JJ (2013) ACYL-LIPID DESATURASE2 is required for chilling and freezing tolerance in Arabidopsis. Plant Cell 25(4):1430–1444
Chen M, Han G, Dietrich CR, Dunn TM, Cahoon EB (2006) The essential nature of sphingolipids in plants as revealed by the functional identification and characterization of the Arabidopsis LCB1 subunit of serine palmitoyltransferase. Plant Cell 18(12):3576–3593
Chen M, Markham JE, Dietrich CR, Jaworski JG, Cahoon EB (2008) Sphingolipid long-chain base hydroxylation is important for growth and regulation of sphingolipid content and composition in Arabidopsis. Plant Cell 20(7):1862–1878
Chen M, Cahoon E, Saucedo-García M, Plasencia J, Gavilanes-Ruíz M (2010) Plant sphingolipids: structure, synthesis and function. In: Wada H, Murata N (eds) Lipids in photosynthesis, vol 30, Advances in Photosynthesis and Respiration. Springer, Dordrecht, p 77
Chen M, Markham JE, Cahoon EB (2012) Sphingolipid Δ8 unsaturation is important for glucosylceramide biosynthesis and low-temperature performance in Arabidopsis. Plant J 69(5):769–781
Chen LY, Shi DQ, Zhang WJ, Tang ZS, Liu J, Yang WC (2015) The Arabidopsis alkaline ceramidase TOD1 is a key turgor pressure regulator in plant cells. Nat Commun 6:6030
Chueasiri C, Chunthong K, Pitnjam K, Chakhonkaen S, Sangarwut N, Sangsawang K, Suksangpanomrung M, Michaelson LV, Napier JA, Muangprom A (2014) Rice ORMDL controls sphingolipid homeostasis affecting fertility resulting from abnormal pollen development. PLoS One 9(9):e106386
Costaglioli P, Joubes K, Garcia C, Stef M, Arveiler B, Lessire R, Garbay B (2005) Profiling candidate genes involved in wax biosynthesis in Arabidopsis thaliana by microarray analysis. BBA-Mol Cell Biol Lipids 1734(3):247–258
Coursol S, Fan LM, Le Stunff H, Spiegel S, Gilroy S, Assmann SM (2003) Sphingolipid signalling in Arabidopsis guard cells involves heterotrimeric G proteins. Nature 423(6940):651–654
Coursol S, Le Stunff H, Lynch D, Gilroy S, Assmann S, Spiegel S (2005) Arabidopsis sphingosine kinase and the effects of phytosphingosine-1-phosphate on stomatal aperture. Plant Physiol 137(2):724–737
Curatolo W (1987) The physical properties of glycolipids. Biochim Biophys Acta 906(2):111–136
de Almeida RF, Fedorov A, Prieto M (2003) Sphingomyelin/phosphatidylcholine/cholesterol phase diagram: boundaries and composition of lipid rafts. Biophys J 85(4):2406–2416
Denny PW, Shams-Eldin H, Price HP, Smith DF, Schwarz RT (2006) The protozoan inositol phosphorylceramide synthase – a novel drug target that defines a new class of sphingolipid synthase. J Biol Chem 281(38):28200–28209
Dietrich CR, Han G, Chen M, Berg RH, Dunn TM, Cahoon EB (2008) Loss-of-function mutations and inducible RNAi suppression of Arabidopsis LCB2 genes reveal the critical role of sphingolipids in gametophytic and sporophytic cell viability. Plant J 54(2):284–298
Duan RD, Bergman T, Xu N, Wu J, Cheng Y, Duan JX, Nelander S, Palmberg C, Nilsson A (2003) Identification of human intestinal alkaline sphingomyelinase as a novel ecto-enzyme related to the nucleotide phosphodiesterase family. J Biol Chem 278(40):38528–38536
Dunn TM, Lynch DV, Michaelson LV, Napier JA (2004) A post-genomic approach to understanding sphingolipid metabolism in Arabidopsis thaliana. Ann Bot 93(5):483–497
Estep TN, Calhoun WI, Barenholz Y, Biltonen RL, Shipley GG, Thompson TE (1980) Evidence for metastability in stearoylsphingomyelin bilayers. Biochemistry 19(1):20–24
Fiebig A, Mayfield JA, Miley NL, Chau S, Fischer RL, Preuss D (2000) Alterations in CER6, a gene identical to CUT1, differentially affect long-chain lipid content on the surface of pollen and stems. Plant Cell 12(10):2001–2008
Gable K, Slife H, Bacikova D, Monaghan E, Dunn TM (2000) Tsc3p is an 80-amino acid protein associated with serine palmitoyltransferase and required for optimal enzyme activity. J Biol Chem 275(11):7597–7603
Gable K, Han G, Monaghan E, Bacikova D, Natarajan M, Williams R, Dunn TM (2002) Mutations in the yeast LCB1 and LCB2 genes, including those corresponding to the hereditary sensory neuropathy type I mutations, dominantly inactivate serine palmitoyltransferase. J Biol Chem 277(12):10194–10200
Gable K, Garton S, Napier JA, Dunn TM (2004) Functional characterization of the Arabidopsis thaliana orthologue of Tsc13p, the enoyl reductase of the yeast microsomal fatty acid elongating system. J Exp Bot 55(396):543–545
Gray JE, Holroyd GH, van der Lee FM, Bahrami AR, Sijmons PC, Woodward FI, Schuch W, Heterington AM (2000) The HIC signalling pathway links CO2 perception to stomatal development. Nature 408(6813):713–716
Greenberg JT, Silverman FP, Liang H (2000) Uncoupling salicylic acid-dependent cell death and defense-related responses from disease resistance in the Arabidopsis mutant acd5. Genetics 156(1):341–350
Grilley MM, Stock SD, Dickson RC, Lester RL, Takemoto JY (1998) Syringomycin action gene SYR2 is essential for sphingolipid 4-hydroxylation in Saccharomyces cerevisiae. J Biol Chem 273(18):11062–11068
Grison MS, Brocard L, Fouillen L, Nicolas W, Wewer V, Dormann P, Nacir H, Benitez-Alfonso Y, Claverol S, Germain V, Boutte Y, Mongrand S, Bayer EM (2015) Specific membrane lipid composition is important for plasmodesmata function in Arabidopsis. Plant Cell 27(4):1228–1250
Grosjean K, Mongrand S, Beney L, Simon-Plas F, Gerbeau-Pissot P (2015) Differential effect of plant lipids on membrane organization: specificities of phytosphingolipids and phytosterols. J Biol Chem 290(9):5810–5825
Guillas I, Zachowski A, Baudouin E (2011) A matter of fat: interaction between nitric oxide and sphingolipid signaling in plant cold response. Plant Signal Behav 6(1):140–142
Guillas I, Guellim A, Reze N, Baudouin E (2013) Long chain base changes triggered by a short exposure of Arabidopsis to low temperature are altered by AHb1 non-symbiotic haemoglobin overexpression. Plant Physiol Biochem 63:191–195
Guo L, Wang X (2012) Crosstalk between phospholipase D and sphingosine kinase in plant stress signaling. Front Plant Sci 3:51
Guo L, Mishra G, Markham JE, Li M, Tawfall A, Welti R, Wang X (2012) Connections between sphingosine kinase and phospholipase D in the abscisic acid signaling pathway in Arabidopsis. J Biol Chem 287(11):8286–8296
Gupta SD, Gable K, Han GS, Borovitskaya A, Selby L, Dunn TM, Harmon JM (2009) Tsc10p and FVT1: topologically distinct short-chain reductases required for long-chain base synthesis in yeast and mammals. J Lipid Res 50(8):1630–1640
Haak D, Gable K, Beeler T, Dunn T (1997) Hydroxylation of Saccharomyces cerevisiae ceramides requires Sur2p and Scs7p. J Biol Chem 272(47):29704–29710
Han G, Gable K, Yan L, Natarajan M, Krishnamurthy J, Gupta SD, Borovitskaya A, Harmon JM, Dunn TM (2004) The topology of the Lcb1p subunit of yeast serine palmitoyltransferase. J Biol Chem 279(51):53707–53716
Han G, Gupta SD, Gable K, Niranjanakumari S, Moitra P, Eichler F, Brown RH Jr, Harmon JM, Dunn TM (2009) Identification of small subunits of mammalian serine palmitoyltransferase that confer distinct acyl-CoA substrate specificities. Proc Natl Acad Sci U S A 106(20):8186–8191
Han S, Lone MA, Schneiter R, Chang A (2010) Orm1 and Orm2 are conserved endoplasmic reticulum membrane proteins regulating lipid homeostasis and protein quality control. Proc Natl Acad Sci U S A 107(13):5851–5856
Hanada K (2003) Serine palmitoyltransferase, a key enzyme of sphingolipid metabolism. Biochim Biophys Acta 1632(1–3):16–30
Haslam TM, Kunst L (2013) Extending the story of very-long-chain fatty acid elongation. Plant Sci 210:93–107
Huang J, Feigenson GW (1999) A microscopic interaction model of maximum solubility of cholesterol in lipid bilayers. Biophys J 76(4):2142–2157
Imai H, Nishiura H (2005) Phosphorylation of sphingoid long-chain bases in Arabidopsis: functional characterization and expression of the first sphingoid long-chain base Kinase gene in plants. Plant Cell Physiol 46(2):375–380
Imai H, Yamamoto K, Shibahara A, Miyatani S, Nakayama T (2000) Determining double-bond positions in monoenoic 2-hydroxy fatty acids of glucosylceramides by gas chromatography-mass spectrometry. Lipids 35(2):233–236
Joubes J, Raffaele S, Bourdenx B, Garcia C, Laroche-Traineau J, Moreau P, Domergue F, Lessire R (2008) The VLCFA elongase gene family in Arabidopsis thaliana: phylogenetic analysis, 3D modelling and expression profiling. Plant Mol Biol 67(5):547–566
Karnovsky MJ, Kleinfeld AM, Hoover RL, Dawidowicz EA, McIntyre DE, Salzman EA, Klausner RD (1982) Lipid domains in membranes. Ann N Y Acad Sci 401:61–75
Kim T-H, Böhmer M, Hu H, Nishimura N, Schroeder JI (2010) Guard cell signal transduction network: advances in understanding abscisic acid, CO2, and Ca2+ signaling. Annu Rev Plant Biol 61(1):561–591
Kim J, Jung JH, Lee SB, Go YS, Kim HJ, Cahoon R, Markham JE, Cahoon EB, Suh MC (2013) Arabidopsis 3-ketoacyl-coenzyme a synthase9 is involved in the synthesis of tetracosanoic acids as precursors of cuticular waxes, suberins, sphingolipids, and phospholipids. Plant Physiol 162(2):567–580
Kimberlin AN, Majumder S, Han G, Chen M, Cahoon RE, Stone JM, Dunn TM, Cahoon EB (2013) Arabidopsis 56-amino acid serine palmitoyltransferase-interacting proteins stimulate sphingolipid synthesis, are essential, and affect mycotoxin sensitivity. Plant Cell 25(11):4627–4639
Klose C, Ejsing CS, Garcia-Saez AJ, Kaiser HJ, Sampaio JL, Surma MA, Shevchenko A, Schwille P, Simons K (2010) Yeast lipids can phase-separate into micrometer-scale membrane domains. J Biol Chem 285(39):30224–30232
Konig S, Feussner K, Schwarz M, Kaever A, Iven T, Landesfeind M, Ternes P, Karlovsky P, Lipka V, Feussner I (2012) Arabidopsis mutants of sphingolipid fatty acid α-hydroxylases accumulate ceramides and salicylates. New Phytol 196(4):1086–1097
Kruger F, Krebs M, Viotti C, Langhans M, Schumacher K, Robinson DG (2013) PDMP induces rapid changes in vacuole morphology in Arabidopsis root cells. J Exp Bot 64(2):529–540
Lachaud C, Prigent E, Thuleau P, Grat S, Da Silva D, Briere C, Mazars C, Cotelle V (2013) 14-3-3-Regulated Ca2+-dependent protein kinase CPK3 is required for sphingolipid-induced cell death in Arabidopsis. Cell Death Differ 20(2):209–217
Laviad EL, Albee L, Pankova-Kholmyansky I, Epstein S, Park H, Merrill AH Jr, Futerman AH (2008) Characterization of ceramide synthase 2: tissue distribution, substrate specificity, and inhibition by sphingosine 1-phosphate. J Biol Chem 283(9):5677–5684
Lefebvre B, Furt F, Hartmann M-A, Michaelson LV, Carde J-P, Sargueil-Boiron F, Rossignol M, Napier JA, Cullimore J, Bessoule J-J, Mongrand S (2007) Characterization of lipid rafts from Medicago truncatula root plasma membranes: a proteomic study reveals the presence of a raft-associated redox system. Plant Physiol 144(1):402–418
Leipelt M, Warnecke D, Zahringer U, Ott C, Muller F, Hube B, Heinz E (2001) Glucosylceramide synthases, a gene family responsible for the biosynthesis of glucosphingolipids in animals, plants, and fungi. J Biol Chem 276(36):33621–33629
Liang H, Yao N, Song JT, Luo S, Lu H, Greenberg JT (2003) Ceramides modulate programmed cell death in plants. Genes Dev 17(21):2636–2641
Lin S-S, Martin R, Mongrand S, Vandenabeele S, Chen K-C, Jang I-C, Chua N-H (2008) RING1 E3 ligase localizes to plasma membrane lipid rafts to trigger FB1-induced programmed cell death in Arabidopsis. Plant J 56(4):550–561
Lunden BM, Lofgren H, Pascher I (1977) Accommodation of hydroxyl groups and their hydrogen bond system in a hydrocarbon matrix. Chem Phys Lipids 20(4):263–271
Luttgeharm KD, Kimberlin AN, Cahoon RE, Cerny RL, Napier JA, Markham JE, Cahoon EB (2015) Sphingolipid metabolism is strikingly different between pollen and leaf in Arabidopsis as revealed by compositional and gene expression profiling. Phytochemistry 115:121–129
Lynch DV, Dunn TM (2004) An introduction to plant sphingolipids and a review of recent advances in understanding their metabolism and function. New Phytol 161(3):677–702
Lynch DV, Steponkus PL (1987) Plasma membrane lipid alterations associated with cold acclimation of winter rye seedlings (Secale cereale L. cv Puma). Plant Physiol 83(4):761–767
Mao C, Obeid LM (2008) Ceramidases: regulators of cellular responses mediated by ceramide, sphingosine, and sphingosine-1-phosphate. Biochim Biophys Acta 1781(9):424–434
Mao CG, Xu RJ, Bielawska A, Obeid LM (2000) Cloning of an alkaline ceramidase from Saccharomyces cerevisiae – an enzyme with reverse (CoA-independent) ceramide synthase activity. J Biol Chem 275(10):6876–6884
Markham JE, Jaworski JG (2007) Rapid measurement of sphingolipids from Arabidopsis thaliana by reversed-phase high-performance liquid chromatography coupled to electrospray ionization tandem mass spectrometry. Rapid Commun Mass Spectrom 21(7):1304–1314
Markham JE, Li J, Cahoon EB, Jaworski JG (2006) Separation and identification of major plant sphingolipid classes from leaves. J Biol Chem 281(32):22684–22694
Markham JE, Molino D, Gissot L, Bellec Y, Hematy K, Marion J, Belcram K, Palauqui JC, Satiat-Jeunemaitre B, Faure JD (2011) Sphingolipids containing very-long-chain fatty acids define a secretory pathway for specific polar plasma membrane protein targeting in Arabidopsis. Plant Cell 23(6):2362–2378
Markham JE, Lynch DV, Napier JA, Dunn TM, Cahoon EB (2013) Plant sphingolipids: function follows form. Curr Opin Plant Biol 16(3):350–357
Melser S, Batailler B, Peypelut M, Poujol C, Bellec Y, Wattelet-Boyer V, Maneta-Peyret L, Faure JD, Moreau P (2010) Glucosylceramide biosynthesis is involved in Golgi morphology and protein secretion in plant cells. Traffic 11(4):479–490
Melser S, Molino D, Batailler B, Peypelut M, Laloi M, Wattelet-Boyer V, Bellec Y, Faure JD, Moreau P (2011) Links between lipid homeostasis, organelle morphodynamics and protein trafficking in eukaryotic and plant secretory pathways. Plant Cell Rep 30(2):177–193
Michaelson LV, Zauner S, Markham JE, Haslam RP, Desikan R, Mugford S, Albrecht S, Warnecke D, Sperling P, Heinz E, Napier JA (2009) Functional characterization of a higher plant sphingolipid Δ4-desaturase: defining the role of sphingosine and sphingosine-1-phosphate in Arabidopsis. Plant Physiol 149(1):487–498
Mina JG, Okada Y, Wansadhipathi-Kannangara NK, Pratt S, Shams-Eldin H, Schwarz RT, Steel PG, Fawcett T, Denny PW (2010) Functional analyses of differentially expressed isoforms of the Arabidopsis inositol phosphorylceramide synthase. Plant Mol Biol 73(4–5):399–407
Mitchell AG, Martin CE (1997) Fah1p, a Saccharomyces cerevisiae cytochrome b5 fusion protein, and its Arabidopsis thaliana homolog that lacks the cytochrome b5 domain both function in the α-hydroxylation of sphingolipid-associated very long chain fatty acids. J Biol Chem 272(45):28281–28288
Mizutani Y, Kihara A, Igarashi Y (2005) Mammalian Lass6 and its related family members regulate synthesis of specific ceramides. Biochem J 390:263–271
Mizutani Y, Kihara A, Igarashi Y (2006) LASS3 (longevity assurance homologue 3) is a mainly testis-specific (dihydro)ceramide synthase with relatively broad substrate specificity. Biochem J 398:531–538
Mongrand S, Morel J, Laroche J, Claverol S, Carde JP, Hartmann MA, Bonneu M, Simon-Plas F, Lessire R, Bessoule JJ (2004) Lipid rafts in higher plant cells: purification and characterization of Triton X-100-insoluble microdomains from tobacco plasma membrane. J Biol Chem 279(35):36277–36286
Morel J, Claverol S, Mongrand S, Furt F, Fromentin J, Bessoule JJ, Blein JP, Simon-Plas F (2006) Proteomics of plant detergent-resistant membranes. Mol Cell Proteomics 5(8):1396–1411
Mortimer JC, Yu XL, Albrecht S, Sicilia F, Huichalaf M, Ampuero D, Michaelson LV, Murphy AM, Matsunaga T, Kurz S, Stephens E, Baldwin TC, Ishii T, Napier JA, Weber APM, Handford MG, Dupree P (2013) Abnormal glycosphingolipid mannosylation triggers salicylic acid-mediated responses in arabidopsis. Plant Cell 25(5):1881–1894
Muir A, Ramachandran S, Roelants FM, Timmons G, Thorner J (2014) TORC2-dependent protein kinase Ypk1 phosphorylates ceramide synthase to stimulate synthesis of complex sphingolipids. Elife 3:e03779
Nagano M, Ihara-Ohori Y, Imai H, Inada N, Fujimoto M, Tsutsumi N, Uchimiya H, Kawai-Yamada M (2009) Functional association of cell death suppressor, Arabidopsis Bax inhibitor-1, with fatty acid 2-hydroxylation through cytochrome b5. Plant J 58(1):122–134
Nagano M, Takahara K, Fujimoto M, Tsutsumi N, Uchimiya H, Kawai-Yamada M (2012) Arabidopsis sphingolipid fatty acid 2-hydroxylases (AtFAH1 and AtFAH2) are functionally differentiated in fatty acid 2-hydroxylation and stress responses. Plant Physiol 159(3):1138–1148
Nagano M, Ishikawa T, Ogawa Y, Iwabuchi M, Nakasone A, Shimamoto K, Uchimiya H, Kawai-Yamada M (2014) Arabidopsis Bax inhibitor-1 promotes sphingolipid synthesis during cold stress by interacting with ceramide-modifying enzymes. Planta 240(1):77–89
Nakagawa N, Kato M, Takahashi Y, Shimazaki K, Tamura K, Tokuji Y, Kihara A, Imai H (2012) Degradation of long-chain base 1-phosphate (LCBP) in Arabidopsis: functional characterization of LCBP phosphatase involved in the dehydration stress response. J Plant Res 125(3):439–449
Ng CKY, Carr K, McAinsh MR, Powell B, Hetherington AM (2001) Drought-induced guard cell signal transduction involves sphingosine-1-phosphate. Nature 410(6828):596–599
Nishikawa M, Hosokawa K, Ishiguro M, Minamioka H, Tamura K, Hara-Nishimura I, Takahashi Y, Shimazaki K, Imai H (2008) Degradation of sphingoid long-chain base 1-phosphates (LCB-1Ps): functional characterization and expression of AtDPL1 encoding LCB-1P lyase involved in the dehydration stress response in Arabidopsis. Plant Cell Physiol 49(11):1758–1763
Pascher I (1976) Molecular arrangements in sphingolipids. Conformation and hydrogen bonding of ceramide and their implication on membrane stability and permeability. Biochim Biophys Acta 455(2):433–451
Pata MO, Wu BX, Bielawski J, Xiong TC, Hannun YA, Ng CKY (2008) Molecular cloning and characterization of OsCDase, a ceramidase enzyme from rice. Plant J 55(6):1000–1009
Paul S, Gable K, Beaudoin F, Cahoon E, Jaworski J, Napier JA, Dunn TM (2006) Members of the Arabidopsis FAE1-like 3-ketoacyl-CoA synthase gene family substitute for the Elop proteins of Saccharomyces cerevisiae. J Biol Chem 281(14):9018–9029
Peer M, Stegmann M, Mueller MJ, Waller F (2010) Pseudomonas syringae infection triggers de novo synthesis of phytosphingosine from sphinganine in Arabidopsis thaliana. FEBS Lett 584(18):4053–4056
Rennie EA, Ebert B, Miles GP, Cahoon RE, Christiansen KM, Stonebloom S, Khatab H, Twell D, Petzold CJ, Adams PD, Dupree P, Heazlewood JL, Cahoon EB, Scheller HV (2014) Identification of a sphingolipid α-glucuronosyltransferase that is essential for pollen function in Arabidopsis. Plant Cell 26(8):3314–3325
Riebeling C, Allegood JC, Wang E, Merrill AH, Futerman AH (2003) Two mammalian longevity assurance gene (LAG1) family members, trh1 and trh4, regulate dihydroceramide synthesis using different fatty acyl-CoA donors. J Biol Chem 278(44):43452–43459
Rittenour WR, Chen M, Cahoon EB, Harris SD (2011) Control of glucosylceramide production and morphogenesis by the Bar1 ceramide synthase in Fusarium graminearum. PLoS One 6:e19385
Roche Y, Gerbeau-Pissot P, Buhot B, Thomas D, Bonneau L, Gresti J, Mongrand S, Perrier-Cornet JM, Simon-Plas F (2008) Depletion of phytosterols from the plant plasma membrane provides evidence for disruption of lipid rafts. FASEB J 22(11):3980–3991
Roelants FM, Breslow DK, Muir A, Weissman JS, Thorner J (2011) Protein kinase Ypk1 phosphorylates regulatory proteins Orm1 and Orm2 to control sphingolipid homeostasis in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 108(48):19222–19227
Ryan PR, Liu Q, Sperling P, Dong B, Franke S, Delhaize E (2007) A higher plant Δ8 sphingolipid desaturase with a preference for (Z)-isomer formation confers aluminum tolerance to yeast and plants. Plant Physiol 144(4):1968–1977
Sandhoff K (2013) Metabolic and cellular bases of sphingolipidoses. Biochem Soc Trans 41(6):1562–1568
Saucedo-García M, Guevara-García A, González-Solís A, Cruz-García F, Vázquez-Santana S, Markham J, Lozano-Rosas M, Dietrich C, Ramos-Vega M, Cahoon E, Gavilanes-Ruíz M (2011) MPK6, sphinganine and the LCB2a gene from serine palmitoyltransferase are required in the signaling pathway that mediates cell death induced by long chain bases in Arabidopsis. New Phytol 191(4):943–957
Shanklin J, Cahoon EB (1998) Desaturation and related modifications of fatty acids. Annu Rev Plant Physiol Plant Mol Biol 49:611–641
Shi L, Bielawski J, Mu J, Dong H, Teng C, Zhang J, Yang X, Tomishige N, Hanada K, Hannun YA, Zuo J (2007) Involvement of sphingoid bases in mediating reactive oxygen intermediate production and programmed cell death in Arabidopsis. Cell Res 17(12):1030–1040
Simanshu DK, Zhai X, Munch D, Hofius D, Markham JE, Bielawski J, Bielawska A, Malinina L, Molotkovsky JG, Mundy JW, Patel DJ, Brown RE (2014) Arabidopsis accelerated cell death 11, ACD11, is a ceramide-1-phosphate transfer protein and intermediary regulator of phytoceramide levels. Cell Rep 6(2):388–399
Simons K, van Meer G (1988) Lipid sorting in epithelial cells. Biochemistry 27(17):6197–6202
Simons K, Ikonen E (1997) Functional rafts in cell membranes. Nature 387(6633):569–572
Smith MA, Dauk M, Ramadan H, Yang H, Seamons LE, Haslam RP, Beaudoin F, Ramirez-Erosa I, Forseille L (2013) Involvement of Arabidopsis ACYL-COENZYME A DESATURASE-LIKE2 (At2g31360) in the biosynthesis of the very-long-chain monounsaturated fatty acid components of membrane lipids. Plant Physiol 161(1):81–96
Spassieva SD, Markham JE, Hille J (2002) The plant disease resistance gene Asc-1 prevents disruption of sphingolipid metabolism during AAL-toxin-induced programmed cell death. Plant J 32(4):561–572
Sperling P, Schmidt H, Heinz E (1995) A cytochrome-b containing fusion protein similar to plant acyl lipid desaturases. Eur J Biochem 232(3):798–805
Sperling P, Zahringer U, Heinz E (1998) A sphingolipid desaturase from higher plants. Identification of a new cytochrome b5 fusion protein. J Biol Chem 273(44):28590–28596
Sperling P, Ternes P, Moll H, Franke S, Zahringer U, Heinz E (2001) Functional characterization of sphingolipid C4-hydroxylase genes from Arabidopsis thaliana. FEBS Lett 494(1–2):90–94
Sperling P, Franke S, Luthje S, Heinz E (2005) Are glucocerebrosides the predominant sphingolipids in plant plasma membranes? Plant Physiol Biochem 43(12):1031–1038
Stone JM, Heard JE, Asai T, Ausubel FM (2000) Simulation of fungal-mediated cell death by fumonisin B1 and selection of fumonisin B 1 -resistant (fbr) Arabidopsis mutants. Plant Cell 12(10):1811–1822
Strub GM, Maceyka M, Hait NC, Milstien S, Spiegel S (2010) Extracellular and intracellular actions of sphingosine-1-phosphate. Adv Exp Med Biol 688:141–155
Takahashi Y, Berberich T, Kanzaki H, Matsumura H, Saitoh H, Kusano T, Terauchi R (2009) Serine palmitoyltransferase, the first step enzyme in sphingolipid biosynthesis, is involved in nonhost resistance. Mol Plant Microbe Interact 22(1):31–38
Tamura K, Mitsuhashi N, Hara-Nishimura I, Imai H (2001) Characterization of an Arabidopsis cDNA encoding a subunit of serine palmitoyltransferase, the initial enzyme in sphingolipid biosynthesis. Plant Cell Physiol 42(11):1274–1281
Teng C, Dong H, Shi L, Deng Y, Mu J, Zhang J, Yang X, Zuo J (2008) Serine palmitoyltransferase, a key enzyme for de novo synthesis of sphingolipids, is essential for male gametophyte development in Arabidopsis. Plant Physiol 146(3):1322–1332
Ternes P, Franke S, Zahringer U, Sperling P, Heinz E (2002) Identification and characterization of a sphingolipid Δ4-desaturase family. J Biol Chem 277(28):25512–25518
Ternes P, Feussner K, Werner S, Lerche J, Iven T, Heilmann I, Riezman H, Feussner I (2011) Disruption of the ceramide synthase LOH1 causes spontaneous cell death in Arabidopsis thaliana. New Phytol 192(4):841–854
Thudichum JLW (1884) A treatise on the chemical constitution of the brain. Baillière, Tindall and Cox, London
Tjellstrom H, Hellgren LI, Wieslander A, Sandelius AS (2010) Lipid asymmetry in plant plasma membranes: phosphate deficiency-induced phospholipid replacement is restricted to the cytosolic leaflet. FASEB J 24(4):1128–1138
Todd J, Post-Beittenmiller D, Jaworski Jan G (1999) KCS1 encodes a fatty acid elongase 3-ketoacyl-CoA synthase affecting wax biosynthesis in Arabidopsis thaliana. Plant J 17(2):119–130
Townley HE, McDonald K, Jenkins GI, Knight MR, Leaver CJ (2005) Ceramides induce programmed cell death in Arabidopsis cells in a calcium-dependent manner. Biol Chem 386(2):161–166
Trenkamp S, Martin W, Tietjen K (2004) Specific and differential inhibition of very-long-chain fatty acid elongases from Arabidopsis thaliana by different herbicides. Proc Natl Acad Sci U S A 101(32):11903–11908
Tsegaye Y, Richardson CG, Bravo JE, Mulcahy BJ, Lynch DV, Markham JE, Jaworski JG, Chen M, Cahoon EB, Dunn TM (2007) Arabidopsis mutants lacking long chain base phosphate lyase are fumonisin-sensitive and accumulate trihydroxy-18:1 long chain base phosphate. J Biol Chem 282(38):28195–28206
Uemura M, Steponkus PL (1994) A contrast of the plasma membrane lipid composition of oat and rye leaves in relation to freezing tolerance. Plant Physiol 104(2):479–496
Uemura M, Joseph RA, Steponkus PL (1995) Cold acclimation of Arabidopsis thaliana. effect on plasma membrane lipid composition and freeze-induced lesions. Plant Physiol 109(1):15–30
van Genderen IL, van Meer G, Slot JW, Geuze HJ, Voorhout WF (1991) Subcellular localization of Forssman glycolipid in epithelial MDCK cells by immuno-electron microscopy after freeze-substitution. J Cell Biol 115(4):1009–1019
van Meer G, Simons K (1988) Lipid polarity and sorting in epithelial cells. J Cell Biochem 36(1):51–58
van Meer G, Voelker DR, Feigenson GW (2008) Membrane lipids: where they are and how they behave. Nat Rev Mol Cell Biol 9(2):112–124
Venkataraman K, Riebeling C, Bodennec J, Riezman H, Allegood JC, Sullards MC, Merrill AH Jr, Futerman AH (2002) Upstream of growth and differentiation factor 1 (uog1), a mammalian homolog of the yeast longevity assurance gene 1 (LAG1), regulates N-stearoyl-sphinganine (C18-(dihydro)ceramide) synthesis in a fumonisin B1-independent manner in mammalian cells. J Biol Chem 277(38):35642–35649
Verhoek B, Haas R, Wrage K, Linscheid M, Heinz E (1983) Lipids and enzymatic-activities in vacuolar membranes isolated via protoplasts from oat primary leaves. Z Naturforsch 38(9–10):770–777
Wang W, Yang X, Tangchaiburana S, Ndeh R, Markham JE, Tsegaye Y, Dunn TM, Wang GL, Bellizzi M, Parsons JF, Morrissey D, Bravo JE, Lynch DV, Xiao S (2008) An inositolphosphorylceramide synthase is involved in regulation of plant programmed cell death associated with defense in Arabidopsis. Plant Cell 20(11):3163–3179
Worrall D, Liang YK, Alvarez S, Holroyd GH, Spiegel S, Panagopulos M, Gray JE, Hetherington AM (2008) Involvement of sphingosine kinase in plant cell signalling. Plant J 56(1):64–72
Wu J-X, Li J, Liu Z, Yin J, Chang Z-Y, Rong C, Wu J-L, Bi F-C, Yao N (2015) The Arabidopsis ceramidase AtACER functions in disease resistance and salt tolerance. Plant J 81(5):767–780
Yang Z, Jean-Baptiste G, Khoury C, Greenwood MT (2005) The mouse sphingomyelin synthase 1 (SMS1) gene is alternatively spliced to yield multiple transcripts and proteins. Gene 363:123–132
Yang H, Richter GL, Wang X, Mlodzinska E, Carraro N, Ma G, Jenness M, Chao DY, Peer WA, Murphy AS (2012) Sterols and sphingolipids differentially function in trafficking of the Arabidopsis ABCB19 auxin transporter. Plant J 74(1):37–47
Yephremov A, Wisman E, Huijser P, Huijser C, Wellesen K, Saedler H (1999) Characterization of the FIDDLEHEAD gene of Arabidopsis reveals a link between adhesion response and cell differentiation in the epidermis. Plant Cell 11(11):2187–2201
Yoshida S, Kawata T, Uemura M, Niki T (1986) Isolation and characterization of tonoplast from chilling-sensitive etiolated seedlings of Vigna radiata L. Plant Physiol 80(1):161–166
Zheng H, Rowland O, Kunst L (2005) Disruptions of the Arabidopsis enoyl-CoA reductase gene reveal an essential role for very-long-chain fatty acid synthesis in cell expansion during plant morphogenesis. Plant Cell 17(5):1467–1481
Acknowledgements
Research that contributed to this chapter was supported by the U.S. National Science Foundation (Grant MCB-1158500).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Luttgeharm, K.D., Kimberlin, A.N., Cahoon, E.B. (2016). Plant Sphingolipid Metabolism and Function. In: Nakamura, Y., Li-Beisson, Y. (eds) Lipids in Plant and Algae Development. Subcellular Biochemistry, vol 86. Springer, Cham. https://doi.org/10.1007/978-3-319-25979-6_11
Download citation
DOI: https://doi.org/10.1007/978-3-319-25979-6_11
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-25977-2
Online ISBN: 978-3-319-25979-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)