Abstract
Cytokines serve essential roles in cell to cell communication during development, maintenance of homeostasis, and responses to external stimuli. The expression of cytokines and the signals transmitted by cytokines to target cells need to be tightly regulated at several levels, including transcriptional and post-transcriptional levels. We briefly summarize the role of post-transcriptional mechanisms in coordinating the regulation of cytokine signaling during inflammatory responses, with an emphasis on AU-rich elements (AREs) and GU-rich elements (GREs). AREs and GREs work in concert to coordinate networks of cytokine signaling. The precise regulation of cytokine signaling is particularly important because its dysregulation can be a contributing factor for human diseases, including autoimmune diseases and cancer.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- AU-rich element: ARE
- GU-rich element: GRE
- mRNA degradation
- mRNA stabilization
- Endoribonuclease
- CELF1, ELAVL1, ZFP36
- Combinatorial control
- Competitive binding
- mRNA protein complex: mRNP
1 Introduction
Cytokines are important mediators of cell to cell communication, controlling a variety of cellular activities such as inflammation, growth, differentiation, cell proliferation and apoptosis. In the immune system, cytokines play essential roles in native immunity, proper immune effector function and maintenance of peripheral tolerance. In addition, cytokines can act as hormones, regulating proper growth, cellular differentiation, and appropriate programmed cell death to help coordinate the gene expression pathways during development and maturation of multicellular organisms. Given the complexity and crosstalk between pathways involved in cytokine signaling, dysregulated cytokine networks are often associated with human diseases. For example, autoimmune diseases such as diabetes involve abnormal production of inflammatory cytokines, which lead to destruction of beta cells in the pancreas. Consequently, many of the medications used to treat autoimmune and inflammatory diseases target cytokines or kinase signaling downstream of cytokine receptors (reviewed in Beyaert et al. 2013; Kontzias et al. 2012; Nepom et al. 2013). Cytokine and growth factor dysregulation leading to uncontrolled cell growth is an important mechanism of cancer pathogenesis (Saharinen et al. 2011). Thus, it is crucial that cytokine signaling pathways are tightly regulated through multiple mechanisms, including transcriptional and post-transcriptional mechanisms (reviewed in Duan et al. 2013; Friedel et al. 2009; Ivanov and Anderson 2013; Seko et al. 2006).
Here, we review post-transcriptional regulation of cytokines and cytokine signaling, focusing on the role of AU-rich elements (AREs) and GU-rich elements (GREs) in coordinating signals transmitted by cytokines during inflammatory responses. AREs and GREs are conserved sequences in the 3′ untranslated region (UTR ) of short-lived transcripts that encode a variety of proteins involved in cellular activation, growth and apoptosis, including components of cytokine signaling pathways. AREs function as post-transcriptional regulators of a variety of cytokines, and the mechanisms by which AREs mediate mRNA degradation have been studied extensively (reviewed in Anderson 2010). Current understanding of AREs has helped to explain the molecular mechanisms of post-transcriptional regulation of cytokine production (Anderson 2008) and has helped us to understand how networks of genes are coordinately regulated by conserved sequences in mRNA. In contrast, our knowledge about the role of GREs in coordinating gene expression is not as well understood. We discuss the role of GREs in regulating cytokine signaling networks, and suggest that AREs and GREs regulate distinct sets of transcripts but function in concert to coordinate cytokine expression with cytokine signaling.
2 Regulation by AREs
During inflammatory responses, cytokines and other early response gene proteins are induced transiently for a defined period of time through a preset program of gene expression events that include transcriptional induction, transcriptional repression, and regulated mRNA stability . For example, T cell receptor stimulation of human T lymphocytes induces ARE -containing cytokine gene expression, such as IL2, IFN-gamma, IL4, TNF-alpha, etc., through transcriptional pulses (Hao and Baltimore 2009; Raghavan et al. 2004). This transcriptional induction is followed by transcript degradation mediated by AREs found in the 3′UTRs of these transcripts (reviewed in Al-Haj and Khabar 2012; Khabar 2007; Schott and Stoecklin 2010). Figure 3.1 shows the intricate network of cytokines that are regulated by AREs in a variety of immune cell types. AREs function as instability elements that mediate cytokine transcript degradation by interacting with ARE-binding proteins that recruit the cellular mRNA degradation machinery to the transcript. Along with regulating mRNA decay , ARE-binding proteins control additional post-transcriptional events such as pre-mRNA processing, transport, intracellular localization, and translation (reviewed in Beisang and Bohjanen 2012; Raghavan and Bohjanen 2004; Stumpo et al. 2010).
The role of cytokines in mediating communication between immune cells. Transcripts in grey are cytokine transcripts that contain AREs in their 3′UTRs . Transcripts labeled with an asterisk (*) contain GREs in their 3′UTRs. Arrows indicate direct interactions and/or activations. Blunt-ended lines indicate inhibitory effects. Reproduced from Vlasova-St. Louis and Bohjanen (2014)
A variety of ARE -containing transcripts interact with many different ARE-binding proteins, resulting in complex patterns of post-transcriptional regulation. The destabilizing functions of different AREs, and their ability to interact with different ARE-binding proteins are not equivalent. The sequence characteristics and decay patterns of different AREs allowed them to be categorized into classes based on their sequence features and decay kinetics (Chen and Shyu 1995). A more extensive database of ARE-containing transcripts from mouse, rat and man has been generated using bioinformatics (Bakheet et al. 2003, 2006; Halees et al. 2008). This database groups AREs into five clusters depending on the number of overlapping AUUUA pentamers within the 3′UTR of a transcript with clusters 1–5 having five to one overlapping AUUUA pentamers, respectively. Cluster I AREs are enriched in secreted proteins, such as cytokines, and are involved in the growth of hematopoietic and immune cells. The other ARE clusters are found within a diverse set of transcripts. In total, ARE-containing transcripts compose approximately 5–8 % of the human transcriptome (Bakheet et al. 2001). AREs play decisive roles in regulating the effects of cytokines on inflammatory responses since mutation of the ARE in cytokine genes such as TNF-alpha or IFN-gamma resulted in profound autoimmune-like inflammatory syndrome (Hodge et al. 2014; Jacob et al. 1991).
AREs regulate mRNA decay by interacting with ARE -binding proteins that can either function to stabilize or destabilize the bound transcript. Numerous ARE-binding proteins involved in the regulation of mRNA turnover have been described in a variety of cell types (reviewed in Gratacos and Brewer 2010; Li et al. 2012; Von Roretz et al. 2011). Typically, mRNA molecules move through different cellular compartments within messenger ribonucleoprotein (mRNP ) complexes, dynamically associating with RNA-binding proteins (RBPs) that bind to conserved cis-elements found in subsets of transcripts (Turner and Hodson 2012a, b). The association of specific RBPs with subsets of transcripts containing conserved regulatory cis-elements coordinates the fate of these bound transcripts through post-transcriptional processes such as translation, intracellular localization, storage or mRNA decay (reviewed in Blackinton and Keene 2014; Keene 2007). Following cellular activation, the set of transcripts bound by a given RBP can be used to define post-transcriptional regulatory networks. RNA-immunoprecipitation (RNA-IP) experiments, in which RNA-binding proteins are immunoprecipitated from cellular lysates using an antibody against a given RBP, and co-purified mRNAs are identified, provided evidence that such networks of coordinately regulated RNAs exist. Target transcripts of several ARE-binding proteins, including HNRNPD (Wu et al. 2013), ZFP36 (Emmons et al. 2008; Stoecklin et al. 2008), ELAVL1 (Fan et al. 2011; Lopez De Silanes et al. 2004b; Mukherjee et al. 2009), KSRP (Winzen et al. 2007) and TIA-1 (Lopez De Silanes et al. 2005) were identified using RNA-IP techniques, and these targets represent distinct but overlapping sets of transcripts. ARE-binding protein targets include transcripts encoding cytokines and cytokine signaling components. Integration of the effects of multiple ARE-binding proteins likely decides the fate of ARE-containing transcripts (Mansfield and Keene 2009).
Several ARE -binding proteins, including ZFP36 (also known as TTP ), BRF1, BRF2, KSRP , and HNRNPD (AUF1 ) destabilize transcripts by recruiting cellular deadenylases and enzymes involved in both 5′→3′ and 3′→5′ mRNA decay processes, whereas other ARE-binding proteins, such as ELAVL1 (also known as HuR ) and ELAVL4, stabilize target transcripts by preventing deadenylation and decay (Bronicki and Jasmin 2013; Brooks and Blackshear 2013; Khabar 2010; Sarkar et al. 2011; White et al. 2013; Wu et al. 2013). ARE-containing mRNA can also be degraded by ribonucleases, such as ZC3H12A, which initiates decay by cleaving the stem loop mRNA structure in the near proximity of an ARE. Examples of ZC3H12A targets are TNF-alpha, IL1b, IL6 and IL12b, IL2, c-Rel and Ox40 transcripts (Matsushita et al. 2009; Uehata et al. 2013), which are important regulators of lymphocyte activation. These same ARE-containing transcripts are also targets of ELAVL1 (Fan et al. 2011; Lopez De Silanes et al. 2004b; Mukherjee et al. 2009). ELAVL1 has been reported to stabilize transcripts of multiple proinflammatory cytokines including, TNF-alpha, VEGF , IL13, IL17 COX2, GATA3 and a number of chemokines through AREs in their 3′UTRs , often leading to increases in both mRNA and protein levels (Stellato et al. 2011; Winzen et al. 2004). The stabilization of ARE-containing transcripts does not always lead to increased protein expression, however. For example when ELAVL1 is in a complex with TIA-1 protein, TNF-alpha and COX2 mRNAs were stabilized but their translation did not increase (Katsanou et al. 2005).
ARE -containing transcripts can interact with different ARE-binding proteins during the course of an inflammatory response through competitive binding . For example, the ARE-binding proteins ELAVL1 and ZFP36 compete with one another for certain ARE-containing transcripts (Raghavan et al. 2001). More than 50 % of ZFP36 target sites in 3′UTRs also function as ELAVL1 target sites (Mukherjee et al. 2014). This potential for ZFP36 and ELAVL1 to compete for the same binding site on a single mRNA has implications for dynamic changes in binding during the course of adaptive immune responses. For example, shortly following T lymphocyte activation, cytoplasmic ELAVL1 levels increase transiently, stabilizing a network of ARE-containing transcripts and allowing their increased expression. Later in T cell activation, ZFP36 induction promotes the displacement of ELAVL1 from ARE-containing transcripts, allowing ZFP36 to mediate their rapid decay (Raghavan et al. 2001). This process allows the transient expression and subsequent degradation of ARE-containing transcripts during the course of immune responses. Cellular signals can also alter the functions of ARE-binding proteins over the course of innate immune responses. ZFP36 activity is regulated through its phosphorylation by p38 MAPK-activated protein kinase 2 (p38/MK2), following lipopolysaccharide (LPS) stimulation of macrophages. This phosphorylation of ZFP36 promotes its association with 14-3-3 protein and prevents ZFP36 from recruiting 3′ deadenylases to the bound transcript (Sun et al. 2007). Thus, through LPS-mediated phosphorylation of ZFP36, ARE-containing transcripts encoding inflammatory mediators such as CCL3, IL1, IL6, IL17, and COX2 are induced by virtue of transcript stabilization and thereby promote inflammation (Kang et al. 2011; Lee et al. 2012; Ronkina et al. 2010). As the immune response resolves, ZFP36 is dephosphorylated by the phosphatase PP2A, allowing ZFP36 to return to its baseline function, promoting the degradation of transcripts encoding these pro-inflammatory mediators (Frasca et al. 2010; Sun et al. 2007).
Understanding the role of AREs in balancing the expression of pro-inflammatory cytokines during inflammatory responses has led to the development of new anti-inflammatory therapies. For example, the novel drug, aganirsen, functions as an anti-inflammatory agent by impairing 14-3-3b-ZFP36 complex formation, preventing the stabilization and upregulation of ARE -containing cytokines, including IL8, TNF-alpha, IL21-beta, IL12, and IL22, during inflammatory responses (Colin et al. 2014). Thus, understanding ARE-mediated regulation of cytokine gene expression may lead to the further development of anti-inflammatory agents. In summary, the coordinate regulation of networks of ARE-containing transcripts, including cytokine networks, is a dynamic process following immune cell activation, which involves activation-induced expression of ARE-binding proteins, competition between ARE-binding proteins with opposing activities, and multiple phosphorylation events mediated through signaling pathways.
3 Regulation by GREs
Studies regarding AREs and ARE -binding proteins have led to identification of other cis-elements and proteins that coordinately regulate mRNA decay (Vlasova and Bohjanen 2008). For example, bioinformatic analysis of short-lived transcripts expressed in primary human T cells led to the identification of the sequence, UGUUUGUUUGU (known as a GRE), to be highly enriched in the 3′UTR of transcripts that exhibited rapid degradation (Vlasova et al. 2008). Introduction of the GRE into the 3′UTR of a beta-globin reporter transcript induced rapid decay of the otherwise stable beta-globin transcript, demonstrating that the GRE functions as a mediator of mRNA degradation . The GRE binds to the protein CELF1, leading to the decay of GRE-containing transcripts. Knockdown of CELF1 in HeLa cells led to stabilization of GRE-containing reporter transcripts, further implicating CELF1 as a mediator of GRE-dependent mRNA degradation (Rattenbacher et al. 2010; Vlasova et al. 2008). A database of GRE-containing transcripts (Halees et al. 2011) was generated by categorizing GREs into five clusters based on the number of overlapping GUUUG pentamers found in the 3′UTR of a transcript, with clusters 1–5 having five to one overlapping GUUUG pentamers, respectively. These GRE-containing transcripts encode a variety of proteins with diverse biological functions including cellular signaling, growth, development and apoptosis regulation. Our recent analysis of CELF1 revealed that targets contained not only the GRE sequence UGUUUGUUUGU but also a GU-repeat sequence which could bind CELF1 and mediate mRNA decay. Based on these findings, the GRE sequence definition was revised to UGU[G/U]UGU[G/U]UGU (Rattenbacher et al. 2010).
CELF1 binds numerous target transcripts involved in cytokine signaling pathways (Vlasova-St. Louis and Bohjanen 2011, 2014; Vlasova-St. Louis et al. 2013). Genome-wide RNA-IP experiments in HeLa cells (Rattenbacher et al. 2010), primary human T cells (Beisang et al. 2012), and mouse myoblasts (Lee et al. 2010), identified hundreds of CELF1 target transcripts. Comparison of molecular functions of the CELF1 bound transcripts among the different cell types revealed enrichment of mRNAs encoding regulators of transcription, post-transcriptional control and cell cycle. In primary human resting T cells, CELF1 binds to and mediates the degradation of numerous transcripts involved in cellular activation and proliferation, presumably to maintain the cell in a quiescent state. Rapid changes in the expression of these GRE-containing transcripts occur immediately following activation of T lymphocytes. T cell receptor-mediated activation promotes the phosphorylation of CELF1, inhibiting its ability to bind to GRE-containing transcripts (Beisang et al. 2012). The lack of CELF1 binding to mRNA within first 24 h following T cell stimulation correlates with a transient increase in expression of GRE-containing transcripts involved in cellular activation and proliferation. Thus, CELF1 functions in resting T cells to mediate the degradation of GRE-containing transcripts that promote cell proliferation, and subsequent CELF1 phosphorylation following T cell activation prevents CELF1 binding to mRNA, leading to the stabilization and upregulation of GRE-containing transcripts (Vlasova-St. Louis and Bohjanen 2014).
Post-transcriptional networks are instrumental for cytokines to rapidly transmit intercellular signals to maintain homeostasis and direct the normal course of inflammatory responses. Post-transcriptional regulation through AREs impacts cytokine production by affecting the stability of cytokine transcripts. Numerous components of cytokine signaling, however, are regulated through GREs. Figures 3.2, 3.3, and 3.4 depict transcripts encoding signaling components downstream of various cytokine receptors whose steady state mRNA levels changed following immune activation. As can be seen, GRE-containing transcripts (grey transcripts) encode components of signaling pathways downstream from these receptors. Overall, these receptors transduce a broad range of intracellular signals that shift cell activation states, alter the rate of proliferation and production of other cytokines. These pathway analyses indicate that the GRE/CELF1 network plays important roles in regulating the dynamic expression of cytokine signaling components through mRNA degradation and thereby modulates the strength and duration of inflammatory responses.
The role of GREs and CELF1 in TNF, TLR or IL1 receptor signaling. Transcripts shown exhibited changes in steady-state levels following T cell receptor stimulation of primary human T cells (Raghavan et al. 2004). Transcripts in grey contain GREs. Transcripts with underlined text were identified as CELF1 target transcripts in human T cells by RNA-immunoprecipitation (Beisang et al. 2012). This figure is modified from (Vlasova-St Louis and Bohjanen 2014)
The role of GREs and CELF1 in interferon and interleukin receptor signaling. Transcripts shown exhibited changes in steady-state levels following T cell receptor stimulation of primary human T cells (Raghavan et al. 2004). Transcripts in grey contain GREs. Transcripts with underlined text were identified as CELF1 target transcripts in human T cells by RNA-immunoprecipitation (Beisang et al. 2012). Transcripts marked with text in italics are also bound to ELAVL1 in activated cells (Mukherjee et al. 2009; Fan et al. 2011). This figure is modified from Vlasova-St. Louis and Bohjanen (2014)
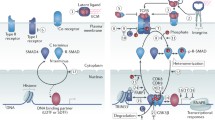
The role of GREs in growth factor signaling. Signals transmitted through tumor growth factor receptors (TGFR), vascular endothelial, epidermal, platelet-derived or insulin growth factor receptors (VEGFR, EGFR, IGFR, PTDGFR) are directed through several major pathways, such as SMAD, mTOR, RAS, PLC, or JAK/STAT. Proper expressions of these transcripts cooperatively contribute to the overall cellular signaling outcome (such as cell proliferation, cell survival and cell growth). Transcripts in grey contain GREs. Transcripts with underlined text were identified as CELF1 target transcripts in resting human T cells by RNA-immunoprecipitation (Beisang et al. 2012). Transcripts marked with text in italics are also bind ELAVL1 in activated cells (Mukherjee et al. 2009; Fan et al. 2011). Arrows indicate direct interactions and/or activations. Blunt-ended lines indicate inhibitory effects. This network diagram was built using Ingenuity Pathway Assistant Software
4 Cross-Talk between Networks
The network of ARE -containing transcripts encodes various cytokines (Fig. 3.1) and cytokine signaling molecules that depends on circuits of GRE-containing transcripts to function. In particular, GRE-containing transcripts encode diverse components of cytokine receptor signaling (Figs. 3.2, 3.3, and 3.4). Moreover, numerous GRE-containing CELF1 target transcripts are also targets of the ARE-binding protein ELAVL1, indicating a functional interaction and potential competition between these RNA-binding proteins. RNA recognition sequences for CELF1 require precise GU repeats or GUUU repeats, whereas the recognition sequence for ELAVL1 is less precise, and ELAVL1 binds to a variety of U-rich sequences, including GU-rich sequences or a poly U sequence. Thus, CELF1 and ELAVL1 may compete for binding to a subset of target transcripts. In contrast to ELAVL1, other ARE-binding proteins that recognize precise AUUU repeats, such as ZFP36, do not bind to GRE sequences and do not compete for binding sites. The comparison of CELF1 targets that were immunoprecipitated from normal T cells (Beisang et al. 2012) with ELAVL1 targets from HeLa (Lopez De Silanes et al. 2004a) or Jurkat cells (Mukherjee et al. 2009), showed a subset of transcripts involved in cytokine signaling that are targets of both proteins (Vlasova-St. Louis and Bohjanen 2014). Depending on which protein competitively binds to the GRE in the target transcript, dichotomous biochemical effects may result in either CELF1 mediating degradation or ELAVL1 mediating stabilization. We hypothesize that in resting cells these transcripts are bound to CELF1 and targeted for degradation, but in activated immune cells, CELF1 becomes phosphorylated and is rendered unable to bind to GRE sequences. ELAVL1 then binds to the GRE, displacing CELF1, stabilizing the transcripts, and allowing them to be translated. Later in the immune response, CELF1 becomes dephosphorylated and then displaces ELAVL1 to mediate transcript degradation. Although further evidence is needed, this model could explain the transient expression of certain GRE-containing transcripts following T cell activation.
The dynamic interplay of GRE and ARE networks appears to regulate nearly all known cytokine signaling pathways. For example, signals downstream of TNF-alpha receptors interact with signals downstream of FAS (also TNFRSF6) to regulate apoptosis. CELF1 regulates expression of a number of apoptotic proteins downstream of FAS receptor through destabilization of pro-apoptotic mRNAs, simultaneously affecting cell death rate and proliferation. CELF1 has been shown to co-immunoprecipitate with transcripts encoding TNF receptor super family members and other molecules downstream of TNF receptors (underlined transcripts in Fig. 3.2) (Beisang et al. 2012; Lee et al. 2010; Rattenbacher et al. 2010). Interestingly, CELF1 and ELAVL1 both bind to and affect the stability of TNF mRNA (Dean et al. 2001; Zhang et al. 2008) and other regulators including effector caspases and cytochrome C mRNA, which are activated downstream of TNF receptors involved in the regulation of apoptosis. Both CELF1 and ELAVL1 proteins regulate stability of transcripts encoding members of the BCL2 superfamily such as BCL2, BNIP3, BCL2L2, BAD and BAX, which modulate programmed cell death (Abdelmohsen et al. 2007; Beisang et al. 2012; Talwar et al. 2013). Thus, signaling through the TNF receptors is controlled by GRE-containing transcripts (also CELF1 targets), which cooperate with ELAVL1 target transcripts to regulate apoptosis.
As shown in Fig. 3.2, multiple GRE-containing transcripts encode signaling components downstream of the IL1 receptor. Proper signaling through this cytokine receptor requires coordination of ARE and GRE networks. Several IL1 receptor signaling components, including STK4, CDC42, MYD88, TIFA, MEF2, RSK, cyclin D have been shown to be ELAVL1 targets in RNA-IP experiments (Mukherjee et al. 2009). Many target transcripts, shared by CELF1 and ELAVL1, are transcription factors or early response genes such as RUNX1, JUN, myc or CREB that are transiently up-regulated following cellular activation and then down-regulated through rapid mRNA decay . Chronic imbalances in the turnover rates of these transcripts could promote onset and progression in diseases such as autoimmunity or cancer through aberrant immune cell activation, excess inflammation and inappropriate cell death (Candido and Hagemann 2013; Moudgil and Choubey 2011; Vlasova et al. 2005).
Many other interleukin and interferon receptors also utilize signaling proteins encoded by GRE-containing transcripts (Fig. 3.3), and rely on interplay between ARE and GRE networks to maintain homeostasis. Cytokines that bind to receptors containing the common gamma chain, such as IL2, IL4, IL7, IL9, IL15 and IL21, are pivotal for immune responses (Baker et al. 2007; Liao et al. 2011). These receptors activate Janus kinases (JAK1 and JAK2) to phosphorylate members of the STAT (Signal transducers and activators of transcription) family of transcription factors (Jenkins 2014). STAT proteins form hetero- or homodimers, and upon translocation into nucleus they induce the transcription of genes including cytokine, chemokine and adhesion molecule genes. One of the induced gene family, SOCS, plays a role in inhibiting JAK kinase activity and consequently, down-regulates immune activation (Linossi et al. 2013). Interestingly, SOCS3 mRNA is also a CELF1 and ELAVL1 target, perhaps to allow its expression only at the appropriate time during the resolution phase of the immune response. Engagement of interferons by interferon receptors also activates JAK-STAT, PI3K and MAPK signaling pathways (Khabar and Young 2007). Interferon family members of type I, II, and III contain functional AREs that allow precise post-transcriptional control by ARE-binding proteins. Interferon receptor mRNAs contain GREs and likely function to maintain the balance in cytokine-receptor signaling and ultimately the course of immune responses (Fig. 3.3). Thus, the induction and control of interferon responses requires coordination of ARE and GRE networks.
Growth factor-induced cell proliferation, adhesion, and migration often depend on ARE and GRE post-transcriptional regulatory networks. Transforming growth factor-beta (TGF-beta) signals are transduced by trans-membrane type I and type II serine/threonine kinase receptors (TGFR1/2), which are encoded by GRE- and ARE-containing mRNAs (Fig. 3.4). TGF-beta maintains tissue homeostasis by regulating the cellular proliferation rate, differentiation and survival. The activated TGF receptor complex induces oligomerization of several downstream SMAD protein family members. Activated SMADs regulate transcription of a number of GRE-containing mRNAs (ex. CITED2, CXCL2, NKX, CDKN1A, VEGF , IL3RA, TGFBR1, etc.), which in turn, become a subject of post-transcriptional regulation by CELF1 (or ELAVL1), once they leave the nucleus. In pathological conditions, TGF-beta signaling, by activating RAS oncogenic pathway, upregulates a large set of ARE-containing genes and specifically VEGF mRNA (Kanies et al. 2008), perhaps due in part to dysfunctional mRNA decay pathways. The RAS–PI3K–Akt kinase–mTOR pathway is a well-established signaling axis that modulates the proliferation and survival of many cell types through growth factor receptor signaling (reviewed elsewhere Bitterman and Polunovsky 2012; Cao et al. 2008; Weichhart and Saemann 2008). Growth factors are beneficial for tissue growth and regeneration, but abnormal growth factor signaling can fuel inflammation and metastases (Carmeliet and Jain 2011). Since ARE and GRE-containing transcripts encode important regulators of apoptosis and cell cycle downstream of growth factor receptors, transformed cells may usurp these post-transcriptional networks to their advantage as they become malignant (Fuxe and Karlsson 2012).
Numerous studies describe an important role of ELAVL1 in the pathogenesis of inflammatory diseases that involve over-production of inflammatory cytokines or diseases characterized by cytokine deficiency. Most studies support a role for ELAVL1 malfunction in the promotion of inflammation and proliferation in human diseases such as autoimmune diseases and cancer (Khabar 2010; Srikantan and Gorospe 2012). Furthermore, genetic deletion of ELAVL1 in thymocytes (Papadaki et al. 2009) and myelocytes (Yiakouvaki et al. 2012) predisposed mice to exaggerated inflammatory responses and inflammation-driven oncogenesis. In contrast, CELF1 functions as an inhibitor of a network of transcripts that promote cellular activation and proliferation (Beisang and Bohjanen 2012). The opposing effects of CELF1 and ELAVL1 may have important implications for new therapies for proliferative diseases such as cancer or autoimmunity. Thus, understanding the crosstalk of CELF1, ELAVL1 and other RBPs may uncover novel therapeutic strategies to fine-tune the balance in proliferative or proinflammatory pathways involved in human diseases. In the future, more work needs to be done to develop interventions targeting post-transcriptional mechanisms involved in inflammatory responses such as the severe reaction that occurs in sepsis, which involves self-amplifying networks of inflammatory mediators often referred to as ‘cytokine storm’.
5 Summary and Perspective
Post-transcriptional networks defined by AREs and GREs regulate cytokine production and signaling. The ARE and GRE networks represent distinct subsets of transcripts that work in concert to coordinate the function of cytokines over the course of an immune response. AREs are critical regulators of cytokine production and play important roles in cytokine signaling, but a distinct subset of cytokine signaling components, regulated by GREs, is needed for precise physiological function of cytokines. Thus, an effective immune response requires crosstalk between ARE and GRE pathways to appropriately regulate cytokine responses over time. Perhaps, integrative studies of interactions between GREs, AREs and RBPs as modulators of coordinated gene expression could be translated to novel treatment strategies.
Change history
29 October 2020
The original online version of this chapter was inadvertently published with part of last name of the author “Irina Vlasova-St. Louis” included along with the first name. The correct first and last name of this author is given below:
References
Abdelmohsen K, Pullmann R Jr, Lal A et al (2007) Phosphorylation of HuR by Chk2 regulates SIRT1 expression. Mol Cell 25:543–557
Al-Haj L, Khabar KS (2012) Cloning of cytokine 3′ untranslated regions and posttranscriptional assessment using cell-based GFP assay. Methods Mol Biol 820:91–104
Anderson P (2008) Post-transcriptional control of cytokine production. Nat Immunol 9:353–359
Anderson P (2010) Post-transcriptional regulons coordinate the initiation and resolution of inflammation. Nat Rev Immunol 10:24–35
Baker SJ, Rane SG, Reddy EP (2007) Hematopoietic cytokine receptor signaling. Oncogene 26:6724–6737
Bakheet T, Frevel M, Williams BR et al (2001) ARED: human AU-rich element-containing mRNA database reveals an unexpectedly diverse functional repertoire of encoded proteins. Nucleic Acids Res 29:246–254
Bakheet T, Williams BR, Khabar KS (2003) ARED 2.0: an update of AU-rich element mRNA database. Nucleic Acids Res 31:421–423
Bakheet T, Williams BR, Khabar KS (2006) ARED 3.0: the large and diverse AU-rich transcriptome. Nucleic Acids Res 34:D111–D114
Beisang D, Bohjanen PR (2012) Perspectives on the ARE as it turns 25 years old. Wiley Interdiscip Rev RNA 3:719–731
Beisang D, Rattenbacher B, Vlasova-St. Louis IA et al (2012) Regulation of CUG-binding protein 1 (CUGBP1) binding to target transcripts upon T cell activation. J Biol Chem 287:950–960
Beyaert R, Beaugerie L, Van Assche G et al (2013) Cancer risk in immune-mediated inflammatory diseases (IMID). Mol Cancer 12:98
Bitterman PB, Polunovsky VA (2012) Translational control of cell fate: from integration of environmental signals to breaching anticancer defense. Cell Cycle 11:1097–1107
Blackinton JG, Keene JD (2014) Post-transcriptional RNA regulons affecting cell cycle and proliferation. Semin Cell Dev Biol 34:44–54
Bronicki LM, Jasmin BJ (2013) Emerging complexity of the HuD/ELAVl4 gene; implications for neuronal development, function, and dysfunction. RNA 19:1019–1037
Brooks SA, Blackshear PJ (2013) Tristetraprolin (TTP): interactions with mRNA and proteins, and current thoughts on mechanisms of action. Biochim Biophys Acta 1829:666–679
Candido J, Hagemann T (2013) Cancer-related inflammation. J Clin Immunol 33(Suppl 1):S79–S84
Cao W, Manicassamy S, Tang H et al (2008) Toll-like receptor-mediated induction of type I interferon in plasmacytoid dendritic cells requires the rapamycin-sensitive PI(3)K-mTOR-p70S6K pathway. Nat Immunol 9:1157–1164
Carmeliet P, Jain RK (2011) Molecular mechanisms and clinical applications of angiogenesis. Nature 473:298–307
Chen CY, Shyu AB (1995) AU-rich elements: characterization and importance in mRNA degradation. Trends Biochem Sci 20:465–470
Colin S, Darne B, Kadi A et al (2014) The antiangiogenic insulin receptor substrate-1 antisense oligonucleotide aganirsen impairs AU-rich mRNA stability by reducing 14-3-3beta-tristetraprolin protein complex, reducing inflammation and psoriatic lesion size in patients. J Pharmacol Exp Ther 349:107–117
Dean JL, Wait R, Mahtani KR et al (2001) The 3′ untranslated region of tumor necrosis factor alpha mRNA is a target of the mRNA-stabilizing factor HuR. Mol Cell Biol 21:721–730
Duan J, Shi J, Ge X et al (2013) Genome-wide survey of interindividual differences of RNA stability in human lymphoblastoid cell lines. Sci Rep 3:1318
Emmons J, Townley-Tilson WH, Deleault KM et al (2008) Identification of TTP mRNA targets in human dendritic cells reveals TTP as a critical regulator of dendritic cell maturation. RNA 14:888–902
Fan J, Ishmael FT, Fang X et al (2011) Chemokine transcripts as targets of the RNA-binding protein HuR in human airway epithelium. J Immunol 186:2482–2494
Frasca D, Romero M, Landin AM et al (2010) Protein phosphatase 2A (PP2A) is increased in old murine B cells and mediates p38 MAPK/tristetraprolin dephosphorylation and E47 mRNA instability. Mech Ageing Dev 131:306–314
Friedel CC, Dolken L, Ruzsics Z et al (2009) Conserved principles of mammalian transcriptional regulation revealed by RNA half-life. Nucleic Acids Res 37, e115
Fuxe J, Karlsson MC (2012) TGF-beta-induced epithelial-mesenchymal transition: a link between cancer and inflammation. Semin Cancer Biol 22:455–461
Gratacos FM, Brewer G (2010) The role of AUF1 in regulated mRNA decay. Wiley Interdiscip Rev RNA 1:457–473
Halees AS, El-Badrawi R, Khabar KS (2008) ARED Organism: expansion of ARED reveals AU-rich element cluster variations between human and mouse. Nucleic Acids Res 36:D137–D140
Halees AS, Hitti E, Al-Saif M et al (2011) Global assessment of GU-rich regulatory content and function in the human transcriptome. RNA Biol 8:681–691
Hao S, Baltimore D (2009) The stability of mRNA influences the temporal order of the induction of genes encoding inflammatory molecules. Nat Immunol 10:281–288
Hodge DL, Berthet C, Coppola V et al (2014) IFN-gamma AU-rich element removal promotes chronic IFN-gamma expression and autoimmunity in mice. J Autoimmun 53:33–45
Ivanov P, Anderson P (2013) Post-transcriptional regulatory networks in immunity. Immunol Rev 253:253–272
Jacob CO, Hwang F, Lewis GD et al (1991) Tumor necrosis factor alpha in murine systemic lupus erythematosus disease models: implications for genetic predisposition and immune regulation. Cytokine 3:551–561
Jenkins BJ (2014) Transcriptional regulation of pattern recognition receptors by JAK/STAT signaling, and the implications for disease pathogenesis. J Interferon Cytokine Res 34:750–758
Kang JG, Amar MJ, Remaley AT et al (2011) Zinc finger protein tristetraprolin interacts with CCL3 mRNA and regulates tissue inflammation. J Immunol 187:2696–2701
Kanies CL, Smith JJ, Kis C et al (2008) Oncogenic Ras and transforming growth factor-beta synergistically regulate AU-rich element-containing mRNAs during epithelial to mesenchymal transition. Mol Cancer Res 6:1124–1136
Katsanou V, Papadaki O, Milatos S et al (2005) HuR as a negative posttranscriptional modulator in inflammation. Mol Cell 19:777–789
Keene JD (2007) RNA regulons: coordination of post-transcriptional events. Nat Rev Genet 8:533–543
Khabar KS (2007) Rapid transit in the immune cells: the role of mRNA turnover regulation. J Leukoc Biol 81:1335–1344
Khabar KS (2010) Post-transcriptional control during chronic inflammation and cancer: a focus on AU-rich elements. Cell Mol Life Sci 67:2937–2955
Khabar KS, Young HA (2007) Post-transcriptional control of the interferon system. Biochimie 89:761–769
Kontzias A, Kotlyar A, Laurence A et al (2012) Jakinibs: a new class of kinase inhibitors in cancer and autoimmune disease. Curr Opin Pharmacol 12:464–470
Lee JE, Lee JY, Wilusz J et al (2010) Systematic analysis of cis-elements in unstable mRNAs demonstrates that CUGBP1 is a key regulator of mRNA decay in muscle cells. PLoS One 5, e11201
Lee HH, Yoon NA, Vo MT et al (2012) Tristetraprolin down-regulates IL-17 through mRNA destabilization. FEBS Lett 586:41–46
Li X, Lin WJ, Chen CY et al (2012) KSRP: a checkpoint for inflammatory cytokine production in astrocytes. Glia 60:1773–1784
Liao W, Lin JX, Leonard WJ (2011) IL-2 family cytokines: new insights into the complex roles of IL-2 as a broad regulator of T helper cell differentiation. Curr Opin Immunol 23:598–604
Linossi EM, Chandrashekaran IR, Kolesnik TB et al (2013) Suppressor of cytokine signaling (SOCS) 5 utilises distinct domains for regulation of JAK1 and interaction with the adaptor protein Shc-1. PLoS One 8, e70536
Lopez De Silanes I, Fan J, Galban CJ et al (2004a) Global analysis of HuR-regulated gene expression in colon cancer systems of reducing complexity. Gene Expr 12:49–59
Lopez De Silanes I, Zhan M, Lal A et al (2004b) Identification of a target RNA motif for RNA-binding protein HuR. Proc Natl Acad Sci USA 101:2987–2992
Lopez De Silanes I, Galban S, Martindale JL et al (2005) Identification and functional outcome of mRNAs associated with RNA-binding protein TIA-1. Mol Cell Biol 25:9520–9531
Mansfield KD, Keene JD (2009) The ribonome: a dominant force in co-ordinating gene expression. Biol Cell 101:169–181
Matsushita K, Takeuchi O, Standley DM et al (2009) Zc3h12a is an RNase essential for controlling immune responses by regulating mRNA decay. Nature 458:1185–1190
Moudgil KD, Choubey D (2011) Cytokines in autoimmunity: role in induction, regulation, and treatment. J Interferon Cytokine Res 31:695–703
Mukherjee N, Lager PJ, Friedersdorf MB et al (2009) Coordinated posttranscriptional mRNA population dynamics during T-cell activation. Mol Syst Biol 5:288
Mukherjee N, Jacobs NC, Hafner M et al (2014) Global target mRNA specification and regulation by the RNA-binding protein ZFP36. Genome Biol 15:R12
Nepom GT, Ehlers M, Mandrup-Poulsen T (2013) Anti-cytokine therapies in T1D: concepts and strategies. Clin Immunol 149:279–285
Papadaki O, Milatos S, Grammenoudi S et al (2009) Control of thymic T cell maturation, deletion and egress by the RNA-binding protein HuR. J Immunol 182:6779–6788
Raghavan A, Bohjanen PR (2004) Microarray-based analyses of mRNA decay in the regulation of mammalian gene expression. Brief Funct Genomic Proteomic 3:112–124
Raghavan A, Robison RL, Mcnabb J et al (2001) HuA and tristetraprolin are induced following T cell activation and display distinct but overlapping RNA binding specificities. J Biol Chem 276:47958–47965
Raghavan A, Dhalla M, Bakheet T et al (2004) Patterns of coordinate down-regulation of ARE-containing transcripts following immune cell activation. Genomics 84:1002–1013
Rattenbacher B, Beisang D, Wiesner DL et al (2010) Analysis of CUGBP1 targets identifies GU-repeat sequences that mediate rapid mRNA decay. Mol Cell Biol 30:3970–3980
Ronkina N, Menon MB, Schwermann J et al (2010) MAPKAP kinases MK2 and MK3 in inflammation: complex regulation of TNF biosynthesis via expression and phosphorylation of tristetraprolin. Biochem Pharmacol 80:1915–1920
Saharinen P, Eklund L, Pulkki K et al (2011) VEGF and angiopoietin signaling in tumor angiogenesis and metastasis. Trends Mol Med 17:347–362
Sarkar S, Han J, Sinsimer KS et al (2011) RNA-binding protein AUF1 regulates lipopolysaccharide-induced IL10 expression by activating IkappaB kinase complex in monocytes. Mol Cell Biol 31:602–615
Schott J, Stoecklin G (2010) Networks controlling mRNA decay in the immune system. Wiley Interdiscip Rev RNA 1:432–456
Seko Y, Cole S, Kasprzak W et al (2006) The role of cytokine mRNA stability in the pathogenesis of autoimmune disease. Autoimmun Rev 5:299–305
Srikantan S, Gorospe M (2012) HuR function in disease. Front Biosci J Virtual Libr 17:189–205
Stellato C, Gubin MM, Magee JD et al (2011) Coordinate regulation of GATA-3 and Th2 cytokine gene expression by the RNA-binding protein HuR. J Immunol 187:441–449
Stoecklin G, Tenenbaum SA, Mayo T et al (2008) Genome-wide analysis identifies interleukin-10 mRNA as target of tristetraprolin. J Biol Chem 283:11689–11699
Stumpo DJ, Lai WS, Blackshear PJ (2010) Inflammation: cytokines and RNA-based regulation. Wiley Interdiscip Rev RNA 1:60–80
Sun L, Stoecklin G, Van Way S et al (2007) Tristetraprolin (TTP)-14-3-3 complex formation protects TTP from dephosphorylation by protein phosphatase 2a and stabilizes tumor necrosis factor-alpha mRNA. J Biol Chem 282:3766–3777
Talwar S, Balasubramanian S, Sundaramurthy S et al (2013) Overexpression of RNA-binding protein CELF1 prevents apoptosis and destabilizes pro-apoptotic mRNAs in oral cancer cells. RNA Biol 10:277–286
Turner M, Hodson D (2012a) Regulation of lymphocyte development and function by RNA-binding proteins. Curr Opin Immunol 24:160–165
Turner M, Hodson DJ (2012b) An emerging role of RNA-binding proteins as multifunctional regulators of lymphocyte development and function. Adv Immunol 115:161–185
Uehata T, Iwasaki H, Vandenbon A et al (2013) Malt1-induced cleavage of regnase-1 in CD4(+) helper T cells regulates immune activation. Cell 153:1036–1049
Vlasova IA, Bohjanen PR (2008) Posttranscriptional regulation of gene networks by GU-rich elements and CELF proteins. RNA Biol 5:201–207
Vlasova IA, Mcnabb J, Raghavan A et al (2005) Coordinate stabilization of growth-regulatory transcripts in T cell malignancies. Genomics 86:159–171
Vlasova IA, Tahoe NM, Fan D et al (2008) Conserved GU-rich elements mediate mRNA decay by binding to CUG-binding protein 1. Mol Cell 29:263–270
Vlasova-St. Louis I, Bohjanen PR (2011) Coordinate regulation of mRNA decay networks by GU-rich elements and CELF1. Curr Opin Genet Dev 21:444–451
Vlasova-St. Louis I, Bohjanen PR (2014) Post-transcriptional regulation of cytokine signaling by AU-rich and GU-rich elements. J Interferon Cytokine Res 34:233–241
Vlasova-St. Louis I, Dickson AM, Bohjanen PR et al (2013) CELFish ways to modulate mRNA decay. Biochim Biophys Acta 1829:695–707
Von Roretz C, Di Marco S, Mazroui R et al (2011) Turnover of AU-rich-containing mRNAs during stress: a matter of survival. Wiley Interdiscip Rev RNA 2:336–347
Weichhart T, Saemann MD (2008) The PI3K/Akt/mTOR pathway in innate immune cells: emerging therapeutic applications. Ann Rheum Dis 67(Suppl 3):iii70–iii74
White EJ, Brewer G, Wilson GM (2013) Post-transcriptional control of gene expression by AUF1: mechanisms, physiological targets, and regulation. Biochim Biophys Acta 1829:680–688
Winzen R, Gowrishankar G, Bollig F et al (2004) Distinct domains of AU-rich elements exert different functions in mRNA destabilization and stabilization by p38 mitogen-activated protein kinase or HuR. Mol Cell Biol 24:4835–4847
Winzen R, Thakur BK, Dittrich-Breiholz O et al (2007) Functional analysis of KSRP interaction with the AU-rich element of interleukin-8 and identification of inflammatory mRNA targets. Mol Cell Biol 27:8388–8400
Wu X, Chesoni S, Rondeau G et al (2013) Combinatorial mRNA binding by AUF1 and Argonaute 2 controls decay of selected target mRNAs. Nucleic Acids Res 41:2644–2658
Yiakouvaki A, Dimitriou M, Karakasiliotis I et al (2012) Myeloid cell expression of the RNA-binding protein HuR protects mice from pathologic inflammation and colorectal carcinogenesis. J Clin Invest 122:48–61
Zhang L, Lee JE, Wilusz J et al (2008) The RNA-binding protein CUGBP1 regulates stability of tumor necrosis factor mRNA in muscle cells: implications for myotonic dystrophy. J Biol Chem 283:22457–22463
Acknowledgments
This work was supported by National Institutes of Health grants AI057484 and AI072068 to P.R.B. We would like to thank Eric Collins, for critical reading and scientific editing of the manuscript. Our acknowledgements to University of Minnesota Supercomputing Institute for providing the access to Ingenuity Pathway Assistant.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Vlasova-St. Louis, I., Bohjanen, P.R. (2016). Post-transcriptional Regulation of Cytokine Signaling During Inflammatory Responses. In: Menon, PhD, K., Goldstrohm, PhD, A. (eds) Post-transcriptional Mechanisms in Endocrine Regulation. Springer, Cham. https://doi.org/10.1007/978-3-319-25124-0_3
Download citation
DOI: https://doi.org/10.1007/978-3-319-25124-0_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-25122-6
Online ISBN: 978-3-319-25124-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)