Abstract
Epigenetics is an area of research that has recently gained much attention from scientists. Epigenetic processes can induce changes within an organism without altering its genetic makeup. More interestingly, epigenetic mechanisms have the strong ability to modulate gene expression without directly altering the sequences of DNA bases. Dietary compounds consist of several bioactive constituents, which actively regulate different molecular targets involved in tumorigenesis. Keeping these facts in view, we provide evidence that these dietary components (e.g. resveratrol (RES), curcumin, genistein, polyphenols and sulforaphane) might interact with various epigenetic targets in cancer therapeutics. These bioactive compounds can modulate normal DNA methylation and histone acetylation patterns, which are essential for the activation of cancer fighting genes. Compounds, such as the ones listed above, induce epigenetic changes associated with the expression of tumor suppressor genes, such as p53, and inhibition of tumor promoting genes such as telomerase reverse transcriptase during tumor progression. Therefore, in this chapter, we present considerable evidence that bioactive compounds and their epigenetic targets are linked with cancer therapeutics, which may open the door to novel drug discovery and development. Remarkable improvements in our understanding of basic epigenetic mechanisms coupled with the rapid progress in the development of powerful new technologies hold great promise for the advancement of cancer treatment.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
3.1 Introduction
Natural dietary compounds isolated from fruits, vegetables, and spices have shown great potential in the prevention and treatment of various diseases such as cancer [1–12]. These compounds contain several bioactive properties that are ubiquitous in plants, many of which have been used in ancient traditional medicines. Herbs, fruits, and veggies are not only a good source of fiber, vitamins and minerals, but also consist of constituents like resveratrol (RES), curcumin, genistein, polyphenols, alkaloids, phenolics and sulforaphane. Evidence indicates that these compounds may serve more than a basic nutritional function; thereby, effectively mediating the regression of multiple debilitating diseases including cancer. In addition to the compounds listed above, other polyphenols such as isothiocynates, silymarin, dialyl sulfide, lycopene, rosmarinic acid, apigenin and gingerol have demonstrated their potency against cancer [1–12]. Interestingly, these compounds have shown the ability to inhibit cancer via the facilitation of various epigenetic processes. Therefore, this chapter will focus on the epigenetic targets of these compounds, which are heavily involved in cancer prevention and therapy.
The study of epigenetics is comprehensive and includes all intracellular and extracellular interactions that may affect the expression of specific genes without directly altering nucleotide sequences [11–25]. Epigenetics can best be defined as the study of the mechanisms affecting temporal and spatial control of gene activity during the development of complex organisms [26]. Perhaps one of the best examples of this is the epigenetic modification of chromatin during embryonic development after the fertilization of eukaryotic eggs. In fact, epigenetic changes are so wide-ranging that they can be used as molecular tools in the screening and treatment of various diseases including cancer. Cancer is the result of genetic mutations and/or epigenetic modifications stemming from the exposure to various adverse environmental factors [27–29]. Studies have shown that exposure to environmental toxins, the quality of nutrition and other factors including physical and chemical pollutants can alter gene expression and modulate individual genetic susceptibility to changes within the epigenome [17, 30]. To this end, there are several known mechanisms that are capable of altering the epigenome, which include DNA methylation , histone acetylation, chromatin remodeling and RNA-interference/interaction.
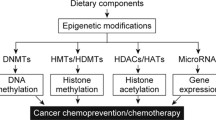
Epigenetic mechanisms often regulate the transcription of genes that facilitate cellular proliferation, differentiation, and survival. These mechanisms have also been linked with tumorigenesis. Aberrant chromatin modifications such as DNA methylation and histone acetylation are the main processes studied in cancer epigenetics [17, 31, 32]. Recent studies have demonstrated that during cancer development, approximately 50 % of all tumor suppressor genes are most likely inactivated by epigenetic rather than genetic, mechanisms [33]. Reports also suggest that bioactive dietary compounds can often restore the function of tumor suppressor genes, increase survival, and under certain circumstances induce apoptosis in many kinds of cancers [34, 35]. In addition to the transcriptional silencing of tumor suppressor genes, non-coding micro-RNAs (miRNAs) can be used to affect mRNA stability and subsequent translation by epigenetic processes during cancer progression [29, 32]. More interestingly, these miRNAs can regulate the expression of various epigenetic modifying enzymes such as methyltransferases (DNMTs), histone methyltransfereases (HMTs), and histone deacetylases (HDACs), which historically have been documented to participate in tumorigenesis [36, 37]. Recent studies also suggest that bioactive dietary compounds may target different tumor suppressor miRNAs to change the function(s) of genes that are being used to classify human cancers [38, 39]. Furthermore, miRNAs either directly or indirectly regulate cancer progression by acting as a tumor suppressor or epigenetically modifying enzyme. In a recent study, miRNA-221 and miRNA-222 inhibit the oncogene KIT, and therefore functions as a tumor suppressor in erythroblastic cells and other solid tumors of human origin [40]. Conversely, the miRNA-29 family can directly control the expression of DNMTs and enhance the expression of both DNMT-3a and DNMT-3b causing genomic hypermethylation and the silencing of sensitive tumor suppressor genes: FHIT and WWOX [41].
3.2 Mechanism of Epigenetic: DNA Methylation
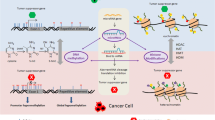
DNA methylation has been observed in many different types of organisms including mammals, plants and bacteria [42, 43]. DNA methylation occurs during DNA replication and is considered a stable gene-silencing mechanism. During this process DNMTs add methyl groups to the 5′ end of the DNA molecule, thus inactivating the affected gene by directly interfering with the assembly of transcription factors essential for gene expression . These enzymes use S-adneylmethionine (SAMs) to transfer methyl groups to cytosine-phosphate-guanine (CpG) sites along the DNA. However, CpG sites are not randomly distributed in the genome, but are concentrated in short CpG-rich DNA fragments commonly referred to as CpG islands [33, 44–46]. Additionally, the majority of CpG sites (except the nucleotide cytosine) are methylated, during development and differentiation in normal cells. Certain subsets of CpG islands at promoter regions may be methylated leading to long term inactivation of target genes, which can be seen in the CpG islands of tumor suppressor genes [47–51]. DNA methylation patterns are formed during cell proliferation, and can disrupt cellular division. DNA methylation is tissue specific, and distinct methylation patterns have been observed across various tissue types. Evidence indicates that the hypermethylation of genes often facilitates conditions that are conducive to carcinogenesis (Fig. 3.1) [33, 44–48, 54–59].
3.3 Histone Modification
The basic structure of the nucleosome consists of the histone octamer, which includes two molecules of each H2A, H2B, H3 and H4 proteins. The N-terminal of these proteins extends from the nucleosome core and the exposed amino acids undergo a series of covalent modifications including methylation, acetylation, phosphorylation, ubiquitinization and sumolization [11, 18, 32, 60]. Singular occurrence or a combination of these modifying events are believed to cause inheritable epigenetic programs that facilitate different nucleosome functions such as gene transcription, the inactivation of the X-chromosome, formation of heterochromatin, mitosis and DNA repair and replication [10, 36, 57, 61, 62]. Direct interaction between the chromodomain of Tip60 and histone H3 trimethylized on lysine 9 (H3K9me3) at double-strand breaks (DSBs) activate acetyltransferase. H3K9me3 deletion inhibits acetyltransferase activation of Tip60, resulting in defective ATM activation that leads to defective DSB repair. These functions are induced either by altered nucleosome interactions with chromatin or by recruiting effector proteins that possess modules that recognize specific histone modifications in a sequence specific manner. The epigenetic codes reside in the substrate specificity of the enzymes that catalyzes the various covalent modifications as well as the enzyme that reverses these modifications.
Chromatin is the template for DNA mediated processes; therefore, it might be worthy to note that histone modifications are an important component in controlling the structure and/or function of the chromatin, which often produces functional consequences. Previous reports suggest that site-specific histone modification can be linked with gene transcription [33, 63, 64]. For instance, histone H3, lysine 9 acetylation (H3K9ac), H3 serine 10 (H3S10) phosphorylation and H3 lysine 4 trimethylation (H3K4me3) are found to be associated with transcriptional activation [33, 64–67]. However, hypomethylation of H3 and H4 have shown to suppress transcription. In brief, the importance of histone modification is highlighted after the revelation that transcription apparatuses often recognize and respond to histone modifying activity [44, 58, 68]. Studies have also shown that histone H3S10 phosphorylation is catalyzed by mitogen and stress activated protein kinase 1 (MSK1). H3S10 phosphorylation is also recognized by a 14-3-3e/14-3-3y heterodimer through its interaction with H3K4 trimethyltransferase (SMYD3) and the p52 subunit of FIIH (Fig. 3.2) [64].
3.4 microRNAs Interaction
MicroRNAs are evolutionarily conserved endogenous non-coding RNAs. MiRNAs are typically 19–25 nucleotides long, which partially or completely match the 3′ untranslated regions (3′UTR) of target RNAs. The hybridization of miRNAs to target RNAs controls gene expression by post-translational modification, silencing, and degradation mechanisms [21, 38, 40, 41, 68, 71–73]. Previous reports suggests that more than 30 % of human genes are controlled by miRNAs which suggests that these small non-coding RNAs play important roles in many biological processes including cell cycle regulation, cell growth, apoptosis , cell differentiation and stress reactions [42, 43, 74–78].
In recent studies, increased detection of miRNA among clinical samples clearly suggests that regulatory functions involve miRNAs [12, 16, 18, 21, 73, 79, 80]. According to data retrieved from the Sanger miRNA Registry in 2013, more than 800 or 1000 human miRNAs have been recorded however; many more miRNAs are expected to be discovered in the future [81]. miRNA control is very similar to the regulation of tightly controlled protein encoding genes. However, during cases of cancer proliferation miRNAs have been found to be greatly deregulated [42, 43, 74–78, 82–85].
Epigenetic manipulation of miRNAs is believed to be highly complex [4, 18, 21, 22, 68, 73]. Additionally, tissue specific expression of miRNAs is tightly regulated by epigenetic mechanisms such as DNA methylation and histone modification ; however, miRNAs themselves can also affect epigenetic mechanisms and regulate gene transcription via post-translational gene silencing [16, 37, 41, 73]. In addition to these important biochemical pathways miRNAs can also be regulated by dietary supplements such as RES. Research shows that oncomirs such as miR-21 are upregulated during the manifestation of various types of cancers. RES is an effective regulator of these [86–89].
3.5 Epigenetic and Carcinogenesis
Epigenetic mechanisms help to maintain cellular homeostasis during normal physiological conditions [5, 10, 13, 20, 21, 23, 24, 30, 48]. However, alterations in epigenetic regulation may lead to aberrant gene expression , which can result in the development of cancer. Cancer development is typically associated with genetic mutation and the subsequent improper unregulated functioning of genes [9, 15, 40, 44, 54, 90–94]. However, our understanding shows that carcinogenesis cannot be the result of genetic alterations alone, but also involve epigenetic changes such as DNA methylation , histone modifications and microRNAs (Fig. 3.2). The level of lysine methylation varies and depends upon cell type. Data suggests that these molecular changes are associated with different types of cancers (Table 3.1).
Additionally, the deregulation of lysine methyltransferase and demethylases has been found in a variety of cancers as shown in Tables 3.2 and 3.3.
These changes lead to stable alterations in the pattern of gene expression that control the neoplastic phenotype, such as cellular growth and invasiveness. At this point, we focused on epigenetic targets of the bioactive compound resveratrol (RES) and its role in cancer prevention and therapy.
RES is a dietary polyphenol obtained from grapes, berries, peanuts, and other plant sources. RES shows a wide range of anti-cancer benefits such as modulating signal transduction pathways that regulate growth, differentiation, apoptosis , inflammation, angiogenesis , and metastasis [117–122]. Studies also suggest that treatment with RES inhibits the proliferation of various human cancers such as skin, breast, prostate, lung and colon [123–127]. The success of RES has led to the development of preclinical animal studies in an effort to determine the potential of this agent for cancer chemotherapeutics . Furthermore, RES has shown remarkable effects against cancer cells at both the biochemical and molecular levels [128].
RES has weaker DNMT inhibitory activity as compared to other bioactive compounds such as epigallocatechin-3-gallate (EGCG). In addition, RES inhibits epigenetic silencing of BRCA-1 induced by aromatic hydrogen receptor (AhR) in MCF-7 cells [129]. Studies show that treatment with RES results in AhR-mediated enrichment of mono-methylated-H3K9, DNMT1, and methyl-binding domain protein-2 at the BRCA-1 promoter, which was associated with BRCA-1 reactivation in MCF-7 cells [129]. Conversely, it has also been reported that RES induces retinoic acid receptor beta 2 (RARbeta2) expressions by blocking RARbeta2 promoter methylation in MCF-7 cells as compared to other adenosine analogs [130]. Furthermore, RES induced activation of the type III HDAC inhibitors, sitrin 1 (SIRT1) and p300, in several in vitro and in vivo models [131]. However, activated SIRT1 negatively down regulated the expression of survivin by deacetylase activity [132–135]. Human BRCA-1 breast cancer cells showed decrease expression of SIRT1 [132–135]. RES has been shown to induce the activation of SIRT1 by altering H3 acetylation. This proved to be a useful approach for target therapy for BRCA-1 mediated breast cancer [136]. Furthermore, SIRT1 associated BRCA1 signaling is important for targeting tumorigenesis by activating oncoproteins in human breast cancer [136]. It has been shown that SIRT1-encoded proteins are needed for RES-induced chemotherapy in APC/+ and APC/− mice [137]. SIRT1 also play an important role in aging, since SIRT1 null mice are unable to tolerate caloric restriction and fail to extend their life duration [137]. This demonstrates RES’s ability to modulate epigenetic processes via the activation of expressed HDAC inhibitors [138].
3.6 Conclusion and Future Prospects
The emerging field that involves nutritional genomics to target nutrient related genetic and epigenetic alterations for cancer therapeutics is unique and timely. The bioactive dietary compound (RES) holds great potential not only in the prevention, but also in the therapy of a wide range of cancers by inducing epigenetic modifications . Cancer is a highly resistant disease and uses several survival pathways to prevail over normal cells. RES can act at several levels to inhibit multiple cellular pathways (for instance the induction of SIRT1 and the inhibition of NFkB) and can be developed as a potential therapeutic agent. Many bioactive dietary compounds have shown great promise in targeting many cellular pathways involved in carcinogenesis as compared to other traditional therapies. However, further research is needed to assess organ specificity, bioavailability and general safety of these dietary compounds for any prudent conclusions. Empirical evidence of the healing powers of ancient medicines strongly supports the use of RES for cancer therapy.
References
Bobrowska-Korczak B, Skrajnowska D, Tokarz A. Effect of zinc and copper supplementation on the prognostic value of urinary 5-methyl-2′-deoxycytidine in DMBA-induced carcinogenesis in rats. Cancer Biomark. 2013;13(6):403–10.
Fernandez AF, Fraga MF. The effects of the dietary polyphenol resveratrol on human healthy aging and lifespan. Epigenetics. 2011;6(7):870–4.
Mai A. Small-molecule chromatin-modifying agents: therapeutic applications. Epigenomics. 2010;2(2):307–24.
Martin SL, Hardy TM, Tollefsbol TO. Medicinal chemistry of the epigenetic diet and caloric restriction. Curr Med Chem. 2013;20(32):4050–9.
Meeran SM, Ahmed A, Tollefsbol TO. Epigenetic targets of bioactive dietary components for cancer prevention and therapy. Clin Epigenetics. 2010;1(3–4):101–16.
Wu ML, et al. Short-term resveratrol exposure causes in vitro and in vivo growth inhibition and apoptosis of bladder cancer cells. PLoS One. 2014;9(2), e89806.
Zhang P, et al. Biological significance and therapeutic implication of resveratrol-inhibited Wnt, Notch and STAT3 signaling in cervical cancer cells. Genes Cancer. 2014;5(5–6):154–64.
Zhong LX, et al. Inhibition of STAT3 signaling as critical molecular event in resveratrol-suppressed ovarian cancer cells. J Ovarian Res. 2015;8(1):25.
Arumuggam N, Bhowmick NA, Rupasinghe HP. A review: phytochemicals targeting JAK/STAT signaling and IDO expression in cancer. Phytother Res. 2015;29(6):805–17.
De Fabiani E, et al. When food meets man: the contribution of epigenetics to health. Nutrients. 2010;2(5):551–71.
Gao Y, Tollefsbol TO. Impact of epigenetic dietary components on cancer through histone modifications. Curr Med Chem. 2015;22(17):2051–64.
Hardy TM, Tollefsbol TO. Epigenetic diet: impact on the epigenome and cancer. Epigenomics. 2011;3(4):503–18.
Barber BA, Rastegar M. Epigenetic control of Hox genes during neurogenesis, development, and disease. Ann Anat. 2010;192(5):261–74.
Berdasco M, et al. Epigenetic inactivation of the Sotos overgrowth syndrome gene histone methyltransferase NSD1 in human neuroblastoma and glioma. Proc Natl Acad Sci U S A. 2009;106(51):21830–5.
Choudhuri S, Cui Y, Klaassen CD. Molecular targets of epigenetic regulation and effectors of environmental influences. Toxicol Appl Pharmacol. 2010;245(3):378–93.
Gerhauser C. Cancer chemoprevention and nutriepigenetics: state of the art and future challenges. Top Curr Chem. 2013;329:73–132.
Herceg Z. Epigenetics and cancer: towards an evaluation of the impact of environmental and dietary factors. Mutagenesis. 2007;22(2):91–103.
Huang J, Plass C, Gerhauser C. Cancer chemoprevention by targeting the epigenome. Curr Drug Targets. 2011;12(13):1925–56.
Liu S. Epigenetics advancing personalized nanomedicine in cancer therapy. Adv Drug Deliv Rev. 2012;64(13):1532–43.
Park LK, Friso S, Choi SW. Nutritional influences on epigenetics and age-related disease. Proc Nutr Soc. 2012;71(1):75–83.
Rouhi A, et al. MiRNAs, epigenetics, and cancer. Mamm Genome. 2008;19(7–8):517–25.
Sharma S, Kelly TK, Jones PA. Epigenetics in cancer. Carcinogenesis. 2010;31(1):27–36.
Shu XH, et al. Metabolic patterns and biotransformation activities of resveratrol in human glioblastoma cells: relevance with therapeutic efficacies. PLoS One. 2011;6(11), e27484.
Stefanska B, et al. Epigenetic mechanisms in anti-cancer actions of bioactive food components—the implications in cancer prevention. Br J Pharmacol. 2012;167(2):279–97.
Wolff GL, et al. Maternal epigenetics and methyl supplements affect agouti gene expression in Avy/a mice. FASEB J. 1998;12(11):949–57.
Holliday R. DNA methylation and epigenetic inheritance. Philos Trans R Soc Lond B Biol Sci. 1990;326(1235):329–38.
Ellis L, Atadja PW, Johnstone RW. Epigenetics in cancer: targeting chromatin modifications. Mol Cancer Ther. 2009;8(6):1409–20.
Esteller M. Epigenetics in cancer. N Engl J Med. 2008;358(11):1148–59.
Ducasse M, Brown MA. Epigenetic aberrations and cancer. Mol Cancer. 2006;5:60.
Suter MA, Aagaard-Tillery KM. Environmental influences on epigenetic profiles. Semin Reprod Med. 2009;27(5):380–90.
Herceg Z, Hainaut P. Genetic and epigenetic alterations as biomarkers for cancer detection, diagnosis and prognosis. Mol Oncol. 2007;1(1):26–41.
Herranz M, Esteller M. DNA methylation and histone modifications in patients with cancer: potential prognostic and therapeutic targets. Methods Mol Biol. 2007;361:25–62.
Kondo Y, et al. Gene silencing in cancer by histone H3 lysine 27 trimethylation independent of promoter DNA methylation. Nat Genet. 2008;40(6):741–50.
Majid S, et al. Genistein induces the p21WAF1/CIP1 and p16INK4a tumor suppressor genes in prostate cancer cells by epigenetic mechanisms involving active chromatin modification. Cancer Res. 2008;68(8):2736–44.
Landis-Piwowar KR, Dou QP. Polyphenols: biological activities, molecular targets, and the effect of methylation. Curr Mol Pharmacol. 2008;1(3):233–43.
Nakagawa Y, et al. Class II HDACs mediate CaMK-dependent signaling to NRSF in ventricular myocytes. J Mol Cell Cardiol. 2006;41(6):1010–22.
Guil S, Esteller M. DNA methylomes, histone codes and miRNAs: tying it all together. Int J Biochem Cell Biol. 2009;41(1):87–95.
Calin GA, et al. MicroRNA profiling reveals distinct signatures in B cell chronic lymphocytic leukemias. Proc Natl Acad Sci U S A. 2004;101(32):11755–60.
Calin GA, et al. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci U S A. 2004;101(9):2999–3004.
Croce CM. Causes and consequences of microRNA dysregulation in cancer. Nat Rev Genet. 2009;10(10):704–14.
Fabbri M, et al. MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc Natl Acad Sci U S A. 2007;104(40):15805–10.
Migicovsky Z, Kovalchuk I. Changes to DNA methylation and homologous recombination frequency in the progeny of stressed plants. Biochem Cell Biol. 2013;91(1):1–5.
Sanchez-Romero MA, Cota I, Casadesus J. DNA methylation in bacteria: from the methyl group to the methylome. Curr Opin Microbiol. 2015;25:9–16.
Doi A, et al. Differential methylation of tissue- and cancer-specific CpG island shores distinguishes human induced pluripotent stem cells, embryonic stem cells and fibroblasts. Nat Genet. 2009;41(12):1350–3.
Feng W, et al. Imprinted tumor suppressor genes ARHI and PEG3 are the most frequently down-regulated in human ovarian cancers by loss of heterozygosity and promoter methylation. Cancer. 2008;112(7):1489–502.
Fujii S, et al. Enhancer of zeste homologue 2 (EZH2) down-regulates RUNX3 by increasing histone H3 methylation. J Biol Chem. 2008;283(25):17324–32.
Khan SI, et al. Epigenetic events associated with breast cancer and their prevention by dietary components targeting the epigenome. Chem Res Toxicol. 2012;25(1):61–73.
Papoutsis AJ, et al. Gestational exposure to the AhR agonist 2,3,7,8-tetrachlorodibenzo-p-dioxin induces BRCA-1 promoter hypermethylation and reduces BRCA-1 expression in mammary tissue of rat offspring: preventive effects of resveratrol. Mol Carcinog. 2015;54(4):261–9.
Schlesinger Y, et al. Polycomb-mediated methylation on Lys27 of histone H3 pre-marks genes for de novo methylation in cancer. Nat Genet. 2007;39(2):232–6.
Vanden Berghe W. Epigenetic impact of dietary polyphenols in cancer chemoprevention: lifelong remodeling of our epigenomes. Pharmacol Res. 2012;65(6):565–76.
Xi Y, et al. Validation of biomarkers associated with 5-fluorouracil and thymidylate synthase in colorectal cancer. Oncol Rep. 2008;19(1):257–62.
Chen QW, et al. Epigenetic regulation and cancer (review). Oncol Rep. 2014;31(2):523–32.
Chen Y, et al. Differential methylation of the micro-RNA 7b gene targets postnatal maturation of murine neuronal Mecp2 gene expression. Dev Neurobiol. 2014;74(4):407–25.
Esteller M, et al. Detection of aberrant promoter hypermethylation of tumor suppressor genes in serum DNA from non-small cell lung cancer patients. Cancer Res. 1999;59(1):67–70.
Grady WM, et al. Detection of aberrantly methylated hMLH1 promoter DNA in the serum of patients with microsatellite unstable colon cancer. Cancer Res. 2001;61(3):900–2.
Lee TL, et al. Detection of gene promoter hypermethylation in the tumor and serum of patients with gastric carcinoma. Clin Cancer Res. 2002;8(6):1761–6.
Sanchez-Cespedes M, et al. Gene promoter hypermethylation in tumors and serum of head and neck cancer patients. Cancer Res. 2000;60(4):892–5.
Singh V, Sharma P, Capalash N. DNA methyltransferase-1 inhibitors as epigenetic therapy for cancer. Curr Cancer Drug Targets. 2013;13(4):379–99.
Widschwendter M, et al. Epigenetic stem cell signature in cancer. Nat Genet. 2007;39(2):157–8.
Agresti A, Bianchi ME. HMGB proteins and gene expression. Curr Opin Genet Dev. 2003;13(2):170–8.
Mirza S, et al. Expression of DNA methyltransferases in breast cancer patients and to analyze the effect of natural compounds on DNA methyltransferases and associated proteins. J Breast Cancer. 2013;16(1):23–31.
Pallier C, et al. Association of chromatin proteins high mobility group box (HMGB) 1 and HMGB2 with mitotic chromosomes. Mol Biol Cell. 2003;14(8):3414–26.
Li B, Carey M, Workman JL. The role of chromatin during transcription. Cell. 2007;128(4):707–19.
Li Y, et al. The histone modifications governing TFF1 transcription mediated by estrogen receptor. J Biol Chem. 2011;286(16):13925–36.
Casanova M, et al. Polycomblike 2 facilitates the recruitment of PRC2 Polycomb group complexes to the inactive X chromosome and to target loci in embryonic stem cells. Development. 2011;138(8):1471–82.
Hassan YI, Zempleni J. Epigenetic regulation of chromatin structure and gene function by biotin. J Nutr. 2006;136(7):1763–5.
Magerl C, et al. H3K4 dimethylation in hepatocellular carcinoma is rare compared with other hepatobiliary and gastrointestinal carcinomas and correlates with expression of the methylase Ash2 and the demethylase LSD1. Hum Pathol. 2010;41(2):181–9.
Pan W, et al. MicroRNA-21 and microRNA-148a contribute to DNA hypomethylation in lupus CD4+ T cells by directly and indirectly targeting DNA methyltransferase 1. J Immunol. 2010;184(12):6773–81.
Baynam G, et al. Intersections of epigenetics, twinning and developmental asymmetries: insights into monogenic and complex diseases and a role for 3D facial analysis. Twin Res Hum Genet. 2011;14(4):305–15.
Godfrey KM, et al. Non-imprinted epigenetics in fetal and postnatal development and growth. Nestle Nutr Inst Workshop Ser. 2013;71:57–63.
Fabbri M, et al. Whole genome analysis and microRNAs regulation in HepG2 cells exposed to cadmium. ALTEX. 2012;29(2):173–82.
Friedman JM, et al. The putative tumor suppressor microRNA-101 modulates the cancer epigenome by repressing the polycomb group protein EZH2. Cancer Res. 2009;69(6):2623–9.
Iorio MV, Croce CM. microRNA involvement in human cancer. Carcinogenesis. 2012;33(6):1126–33.
Adamsen BL, et al. Apoptosis, cell cycle progression and gene expression in TP53-depleted HCT116 colon cancer cells in response to short-term 5-fluorouracil treatment. Int J Oncol. 2007;31(6):1491–500.
Mobarra N, et al. Overexpression of microRNA-16 declines cellular growth, proliferation and induces apoptosis in human breast cancer cells. In Vitro Cell Dev Biol Anim. 2015;51(6):604–11.
Wang Z, et al. MicroRNA-378-5p suppresses cell proliferation and induces apoptosis in colorectal cancer cells by targeting BRAF. Cancer Cell Int. 2015;15:40.
Yang T, et al. MicroRNA-15a induces cell apoptosis and inhibits metastasis by targeting BCL2L2 in non-small cell lung cancer. Tumour Biol. 2015;36(6):4357–65.
Zhao X, et al. RNA silencing of integrin-linked kinase increases the sensitivity of the A549 lung cancer cell line to cisplatin and promotes its apoptosis. Mol Med Rep. 2015;12(1):960–6.
Lai EC, et al. Computational identification of Drosophila microRNA genes. Genome Biol. 2003;4(7):R42.
Wang Y, et al. Genetic and epigenetic studies for determining molecular targets of natural product anticancer agents. Curr Cancer Drug Targets. 2013;13(5):506–18.
Griffiths-Jones S. The microRNA registry. Nucleic Acids Res. 2004;32(Database issue):D109–11.
Kupczyk M, Kuna P. MicroRNAs—new biomarkers of respiratory tract diseases. Pneumonol Alergol Pol. 2014;82(2):183–90.
Li X, et al. MicroRNA expression profiles in differentiated thyroid cancer, a review. Int J Clin Exp Med. 2013;6(1):74–80.
Sanchez-Espiridion B, et al. MicroRNA signatures and treatment response in patients with advanced classical Hodgkin lymphoma. Br J Haematol. 2013;162(3):336–47.
Yang H, et al. Up-regulation of microRNA-138 induce radiosensitization in lung cancer cells. Tumour Biol. 2014;35(7):6557–65.
Kaminski J, et al. Resveratrol initiates differentiation of mouse skeletal muscle-derived C2C12 myoblasts. Biochem Pharmacol. 2012;84(10):1251–9.
Lancon A, et al. Control of MicroRNA expression as a new way for resveratrol to deliver its beneficial effects. J Agric Food Chem. 2012;60(36):8783–9.
Liu P, et al. Resveratrol induces apoptosis of pancreatic cancers cells by inhibiting miR-21 regulation of BCL-2 expression. Clin Transl Oncol. 2013;15(9):741–6.
Sheth S, et al. Resveratrol reduces prostate cancer growth and metastasis by inhibiting the Akt/MicroRNA-21 pathway. PLoS One. 2012;7(12), e51655.
Bachmann IM, et al. EZH2 expression is associated with high proliferation rate and aggressive tumor subgroups in cutaneous melanoma and cancers of the endometrium, prostate, and breast. J Clin Oncol. 2006;24(2):268–73.
Baptista T, et al. Regulation of histone H2A.Z expression is mediated by sirtuin 1 in prostate cancer. Oncotarget. 2013;4(10):1673–85.
Chen WD, et al. Detection in fecal DNA of colon cancer-specific methylation of the nonexpressed vimentin gene. J Natl Cancer Inst. 2005;97(15):1124–32.
Cooney CA. Are somatic cells inherently deficient in methylation metabolism? A proposed mechanism for DNA methylation loss, senescence and aging. Growth Dev Aging. 1993;57(4):261–73.
Feng ZJ, et al. Lung cancer cell migration is regulated via repressing growth factor PTN/RPTP beta/zeta signaling by menin. Oncogene. 2010;29(39):5416–26.
Ellinger J, et al. Global levels of histone modifications predict prostate cancer recurrence. Prostate. 2010;70(1):61–9.
Elsheikh SE, et al. Global histone modifications in breast cancer correlate with tumor phenotypes, prognostic factors, and patient outcome. Cancer Res. 2009;69(9):3802–9.
Li Q, et al. Polycomb CBX7 directly controls trimethylation of histone H3 at lysine 9 at the p16 locus. PLoS One. 2010;5(10), e13732.
Canaani E, et al. ALL-1/MLL1, a homologue of Drosophila TRITHORAX, modifies chromatin and is directly involved in infant acute leukaemia. Br J Cancer. 2004;90(4):756–60.
Liu H, et al. Phosphorylation of MLL by ATR is required for execution of mammalian S-phase checkpoint. Nature. 2010;467(7313):343–6.
Scacheri PC, et al. Genome-wide analysis of menin binding provides insights into MEN1 tumorigenesis. PLoS Genet. 2006;2(4), e51.
Kobayashi Y, et al. DNA methylation profiling reveals novel biomarkers and important roles for DNA methyltransferases in prostate cancer. Genome Res. 2011;21(7):1017–27.
Velichutina I, et al. EZH2-mediated epigenetic silencing in germinal center B cells contributes to proliferation and lymphomagenesis. Blood. 2010;116(24):5247–55.
Tell R, et al. Gastrin-releasing peptide signaling alters colon cancer invasiveness via heterochromatin protein 1Hsbeta. Am J Pathol. 2011;178(2):672–8.
Wang XQ, et al. SMYD3 tandem repeats polymorphism is not associated with the occurrence and metastasis of hepatocellular carcinoma in a Chinese population. Exp Oncol. 2007;29(1):71–3.
Fang W, et al. Preferential loss of a polymorphic RIZ allele in human hepatocellular carcinoma. Br J Cancer. 2001;84(6):743–7.
Lucio-Eterovic AK, et al. Role for the nuclear receptor-binding SET domain protein 1 (NSD1) methyltransferase in coordinating lysine 36 methylation at histone 3 with RNA polymerase II function. Proc Natl Acad Sci U S A. 2010;107(39):16952–7.
Nimura K, et al. A histone H3 lysine 36 trimethyltransferase links Nkx2-5 to Wolf-Hirschhorn syndrome. Nature. 2009;460(7252):287–91.
Taketani T, et al. NUP98-NSD3 fusion gene in radiation-associated myelodysplastic syndrome with t(8;11)(p11;p15) and expression pattern of NSD family genes. Cancer Genet Cytogenet. 2009;190(2):108–12.
Watanabe H, et al. Deregulation of histone lysine methyltransferases contributes to oncogenic transformation of human bronchoepithelial cells. Cancer Cell Int. 2008;8:15.
Suikki HE, et al. Genetic alterations and changes in expression of histone demethylases in prostate cancer. Prostate. 2010;70(8):889–98.
Fukuda T, et al. Fbxl10/Kdm2b deficiency accelerates neural progenitor cell death and leads to exencephaly. Mol Cell Neurosci. 2011;46(3):614–24.
Vinatzer U, et al. Mucosa-associated lymphoid tissue lymphoma: novel translocations including rearrangements of ODZ2, JMJD2C, and CNN3. Clin Cancer Res. 2008;14(20):6426–31.
Yang ZQ, et al. Identification of a novel gene, GASC1, within an amplicon at 9p23-24 frequently detected in esophageal cancer cell lines. Cancer Res. 2000;60(17):4735–9.
Zeng J, et al. The histone demethylase RBP2 Is overexpressed in gastric cancer and its inhibition triggers senescence of cancer cells. Gastroenterology. 2010;138(3):981–92.
Rao M, et al. Inhibition of histone lysine methylation enhances cancer-testis antigen expression in lung cancer cells: implications for adoptive immunotherapy of cancer. Cancer Res. 2011;71(12):4192–204.
Xiang Y, et al. JMJD3 is a histone H3K27 demethylase. Cell Res. 2007;17(10):850–7.
Bishayee A. Cancer prevention and treatment with resveratrol: from rodent studies to clinical trials. Cancer Prev Res (Phila). 2009;2(5):409–18.
Bishayee A, Dhir N. Resveratrol-mediated chemoprevention of diethylnitrosamine-initiated hepatocarcinogenesis: inhibition of cell proliferation and induction of apoptosis. Chem Biol Interact. 2009;179(2-3):131–44.
Bishayee A, Politis T, Darvesh AS. Resveratrol in the chemoprevention and treatment of hepatocellular carcinoma. Cancer Treat Rev. 2010;36(1):43–53.
Singh NP, et al. Resveratrol (trans-3,5,4′-trihydroxystilbene) suppresses EL4 tumor growth by induction of apoptosis involving reciprocal regulation of SIRT1 and NF-kappaB. Mol Nutr Food Res. 2011;55(8):1207–18.
Singh UP, et al. Resveratrol (trans-3,5,4′-trihydroxystilbene) induces silent mating type information regulation-1 and down-regulates nuclear transcription factor-kappaB activation to abrogate dextran sulfate sodium-induced colitis. J Pharmacol Exp Ther. 2010;332(3):829–39.
Singh UP, et al. Role of resveratrol-induced CD11b(+) Gr-1(+) myeloid derived suppressor cells (MDSCs) in the reduction of CXCR3(+) T cells and amelioration of chronic colitis in IL-10(−/−) mice. Brain Behav Immun. 2012;26(1):72–82.
Bai Y, et al. Resveratrol induces apoptosis and cell cycle arrest of human T24 bladder cancer cells in vitro and inhibits tumor growth in vivo. Cancer Sci. 2010;101(2):488–93.
Kraft TE, et al. Fighting cancer with red wine? Molecular mechanisms of resveratrol. Crit Rev Food Sci Nutr. 2009;49(9):782–99.
Liu PL, et al. Resveratrol inhibits human lung adenocarcinoma cell metastasis by suppressing heme oxygenase 1-mediated nuclear factor-kappaB pathway and subsequently downregulating expression of matrix metalloproteinases. Mol Nutr Food Res. 2010;54 Suppl 2:S196–204.
Mao QQ, et al. Resveratrol confers resistance against taxol via induction of cell cycle arrest in human cancer cell lines. Mol Nutr Food Res. 2010;54(11):1574–84.
Vanamala J, et al. Resveratrol suppresses IGF-1 induced human colon cancer cell proliferation and elevates apoptosis via suppression of IGF-1R/Wnt and activation of p53 signaling pathways. BMC Cancer. 2010;10:238.
Athar M, et al. Multiple molecular targets of resveratrol: anti-carcinogenic mechanisms. Arch Biochem Biophys. 2009;486(2):95–102.
Papoutsis AJ, et al. Resveratrol prevents epigenetic silencing of BRCA-1 by the aromatic hydrocarbon receptor in human breast cancer cells. J Nutr. 2010;140(9):1607–14.
Stefanska B, et al. Hypomethylation and induction of retinoic acid receptor beta 2 by concurrent action of adenosine analogues and natural compounds in breast cancer cells. Eur J Pharmacol. 2010;638(1-3):47–53.
Kaeberlein M, et al. Substrate-specific activation of sirtuins by resveratrol. J Biol Chem. 2005;280(17):17038–45.
Bouras T, et al. SIRT1 deacetylation and repression of p300 involves lysine residues 1020/1024 within the cell cycle regulatory domain 1. J Biol Chem. 2005;280(11):10264–76.
Das C, et al. CBP/p300-mediated acetylation of histone H3 on lysine 56. Nature. 2009;459(7243):113–7.
Li Y, et al. SIRT2 down-regulation in HeLa can induce p53 accumulation via p38 MAPK activation-dependent p300 decrease, eventually leading to apoptosis. Genes Cells. 2011;16(1):34–45.
Lim JH, et al. Sirtuin 1 modulates cellular responses to hypoxia by deacetylating hypoxia-inducible factor 1alpha. Mol Cell. 2010;38(6):864–78.
Lin YL, et al. Biologically active components and nutraceuticals in the Monascus-fermented rice: a review. Appl Microbiol Biotechnol. 2008;77(5):965–73.
Boily G, et al. SirT1-null mice develop tumors at normal rates but are poorly protected by resveratrol. Oncogene. 2009;28(32):2882–93.
Farghali H, Kutinova Canova N, Lekic N. Resveratrol and related compounds as antioxidants with an allosteric mechanism of action in epigenetic drug targets. Physiol Res. 2013;62(1):1–13.
Acknowledgements
M.K.M. has been supported by National Institutes of Health grant P20CA192976, US Department of Defense grant W911NF-12-1-0073 and W911NF-14-1-0064, and National Science Foundation grant 1154214.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Stokes, J.A., Kumar, S., Scissum-Gunn, K., Singh, U.P., Mishra, M.K. (2016). Epigenetic and Cancer: An Evaluation of the Impact of Dietary Components. In: Mishra, M., Bishnupuri, K. (eds) Epigenetic Advancements in Cancer. Springer, Cham. https://doi.org/10.1007/978-3-319-24951-3_3
Download citation
DOI: https://doi.org/10.1007/978-3-319-24951-3_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-24949-0
Online ISBN: 978-3-319-24951-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)