Abstract
Spiders have developed specialized silks with outstanding biophysical properties over millions of years of evolution. As biopolymers composed by highly repetitive amino acid motifs, spider silks have been the focus of research for years. Due to recent advances in genetic engineering, recombinant spider silks have been produced, revealing the relationships between their protein structure and their mechanical properties. Each amino acid motif present in the silk adopts a particular secondary structure responsible for conferring a specific mechanical property to it. This feature has opened up the possibility to produce recombinant silks with controlled properties for various biotechnological applications. Moreover, spider silks are biocompatible and biodegradable biomaterials, which also allow their application in medicine. Accordingly, the relationship between molecular composition, secondary structure, and mechanical properties of spider silks is described in this chapter, along with a discussion of the current strategies for the production of recombinant spider silks, their importance in new biotechnological applications, and the current status of the field.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
9.1 Introduction
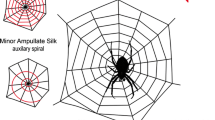
Spiders have developed specialized silks with outstanding biophysical properties over millions of years of evolution [1–3]. They are capable of producing the most architecturally complex spider webs possible, which are able to stop the tremendous kinetic energy of flying insect prey. Orb-weaving spiders (Araneomorphae: Orbiculariae) spin silks that are mechanically superior to most synthetic or natural high-performance fibers [4]. Spiders can possess up to seven morphologically distinct pairs of spinning glands; this allows them to produce fibers from an aqueous silk dope solution, composed basically of high molecular weight proteins, at ambient temperature and pressure [5–7]. With the exception of the aggregate gland, which is responsible for the production of the aqueous coat present on the spiral threads in orb webs, the major and minor ampullate, flagelliform, tubuliform, pyriform, and aciniform glands are responsible for the production of a single kind of silk perfectly adapted for the specific requirements demanded by the spider’s lifestyle and habitat (Fig. 9.1) [8].
At the molecular level, spider silk proteins (spidroins) consist of the repetition of different structural amino acid motifs, which are ultimately responsible for the mechanical properties of the silk fibers [2, 9–11]. These motifs vary according to the type of the silk and are normally present at the central core of the protein flanked by non-repetitive amino (N–) and carboxyl (C–) termini regions. This feature, together with biocompatibility [12, 13], low inflammatory potential, antimicrobial activity [14], and a slow rate of degradation [15], has opened up the possibility to produce recombinant silks with controlled properties for various biotechnological applications, from ballistic vests and sporting goods to regenerative medicine [16–18].
Advances in genetic engineering have led several laboratories to successfully produce synthetic spider silks from different heterologous host systems [19–24]. However, the mechanical properties of natural spider silk are still unmatched and continue to be the most significant challenge. Accordingly, in this chapter we aim to describe the relationship between molecular composition, secondary structure, and mechanical properties of spider silks, along with a discussion of the current strategies for the production of recombinant spider silks and their importance in new biotechnological applications.
9.2 Molecular Properties of Spider Silks
Spiders are outstanding silk producers. One spider can produce up to six different types of silk, mainly composed of four amino acid motifs: polyalanine (A n ), alternating glycine and alanine (GA)n, amino acid triplets composed of two glycines and a third variable amino acid (GGX)n, and glycine-proline-glycine modules (GPGXX)n. Each motif is responsible for the adoption of a specific secondary structure in the protein, and the different combinations will determine the complex array of the mechanical properties exhibited by each silk [9, 10].
Among the different silks produced by spiders, dragline and flagelliform silks are the best characterized because of their remarkable mechanical properties. Being so, this chapter will focus on the characterization of these two spidroins. The function and structure of other silk types are reviewed elsewhere [1, 25, 26].
9.2.1 Dragline Silks
Dragline silk forms the frame and radii of the web and serves as a lifeline for the spider while moving through the environment. Produced by the major ampullate gland, dragline silks are known for their high tensile strength and toughness, with strength values in the range of 1–2 GPa [10, 27, 28]. Table 9.1 compares the material properties of some dragline silks from different spider species. The dragline silk produced by Darwin’s bark spider (Caerostris darwini), which is about five times higher than Kevlar, is the toughest silk fiber measured to date (250 MJ/m3) [29].
Dragline is composed of two different spidroins with molecular size of more than 300 kDa – major ampullate spidroin 1 and 2 (MaSp1 and MaSp2) [30]. The repetitive core region of MaSp1 is formed by the repetition of An, (GA)n, and (GGX)n (X = Y, L, and Q) motifs, flanked by non-repetitive domains at the N– and C– terminal ends. Although MaSp1 and MaSp2 share common features, the latter also has a high proline content, reflecting the fact that (GGX)n motifs are alternated with (GPGXX)n repeats (X = Q, G,Y) (Fig. 9.2). Each of these motifs is responsible for the three-dimensional structure of the protein, which determines the mechanical properties exhibited by dragline silks. The An and (GA)n motifs are responsible for fiber toughness due to the adoption of β-sheet structures which assemble into crystalline substructures in the thread [9]. Surrounding these crystalline structures is an amorphous matrix composed of (GGX)n and the (GPGXX)n motifs from MaSp2 which is responsible for the elasticity of dragline silk [31]. These regions, which are rich in glycine, include helix-like structures and β-turns, whereby the GGX helices act as cross-links between the crystalline β-sheets in adjacent protein molecules, facilitating fiber alignment [10]. Additionally, dragline silk has the ability to undergo supercontraction in contact with water or a relative humidity greater than 60 %. This feature is probably controlled by the proline content, which tunes the mechanical behavior by changing the crystal cross-link density and hydrogen-bonding network [32, 33].
The molecular structure of orb-weaver spider spidroins. (a) Alignment of the consensus repetitive sequences of MaSp1 and Flag spidroins. (b) Structural motifs found in spider silk proteins. Each box represents a motif module in the spider silk. The empty box marked with “?” indicates that the secondary structure is unknown. MaSp1 and 2, major ampullate spidroin; MiSp, Minor ampullate spidroin; Flag, flagelliform spidroin (Reproduced with permission from Teulé et al. [56])
Once the silk amino acid content has been directly associated with the mechanical properties of the thread, the performance of the dragline silk depends on the ratio between the two protein components [34, 35]. By calculating the number of proline residues, it is possible to estimate the fiber content of MaSp1 and 2, which varies among spider species. The dragline silk from Nephila clavipes, for instance, is composed of approximately 80 % MaSp1 and 20 % MaSp2. However, the dragline silk of the orb-weaver spider, Argiope aurantia, contains 41 % MaSp1 and 59 % MaSp2. Stress–strain curves determined by polynomial regression show significant differences between major ampullate silk fibers from N. clavipes and A. aurantia. The N. clavipes stress–strain curve is characteristic of an elastic material; in contrast, A. aurantia dragline silk has the behavior of a viscoelastic composite [36].
In addition to the repetitive domains, dragline silks contain non-repetitive N- and C-terminal ends. The C-terminal region is highly conserved between species and is essential for controlled switching between the storage and assembly forms of silk proteins [37]. However, while the C-terminal domain is a prerequisite for the formation of continuous silk fibers as opposed to amorphous molecular aggregates, it is the N-terminal domain that senses the pH differences in the glandular lumen and thereby appears to regulate the assembly process in vivo [38].
9.2.2 Flagelliform Silk
Among the different silks produced by spiders, the flagelliform silk (Flag) is extremely elastic and has a very high hysteresis, being three times tougher than Kevlar [2, 3, 39]. It is responsible for the formation of the capture spiral in the orb web, an effective energy-absorbing trap for flying prey. Single molecules of the capture silk, Flag spidroins, are very large in size, measuring approximately 360 kDa. Like other elastomeric proteins, such as elastin and abductin, Flag is also characterized by a high content of glycine residues [40]. Its amino acid content is organized into a large ensemble of repeats composed of the individual repetitive motifs, (GPGGX)n (X represents A, V, S, and Y) and GGX (X represents A, S, and T), and a “spacer” sequence containing charged and hydrophilic amino acids [39, 41, 42] (Fig. 9.2). The glycine-rich central Flag core is also flanked by conserved non-repetitive N- and C-termini. Like in dragline silk, these regions are suggested to control solubility and fiber formation in native spider silk.
The elastic nature of flagelliform silk is conferred by the amino acid motif GPGGX. This motif has been suggested to conform to a β-spiral secondary structure similar to a spring, which could easily contribute to the elastic mechanism of the fiber, whereby the proline bonds generate force for the retraction of the silk after stretching [9]. For instance, dragline silk has 35 % elasticity, with an average of five β-turns in a row, whereas flagelliform silk, with >200 % elasticity, has this same module repeated ∼50 times [43]. The glycoprotein “glue” coating the capture silk also contributes to the extensibility of the thread. Like MaSp2, Flag has the ability to contract from its original length in contact with water. The hygroscopic components present in the glycoprotein glue attract water from the environment, keeping the web hydrated and thus playing a critical role in the mechanical properties of flagelliform silk fibers [44, 45].
Finally, the spacer regions found in Flag contain amino acids with charged groups and separate the peptide motifs into clusters. Although these may provide organizational areas within silks or surface regions for interactions when a mature fiber is formed, their real function is currently undetermined [9]. Recently, Adrianos and colleagues [11] investigated the contribution of various motifs to the mechanical properties of Flag spidroin using recombinant silk gene expression. Their results indicate that the spacer motif is likely a primary contributor to silk strength, with the GGX motif supplying mobility to the protein network of native flagelliform silk fibers.
9.3 Recombinant Spider Silk Proteins and Their Application
Since ancient times, spider silks have fascinated humankind because of their great mechanical characteristics of strength, toughness, and elasticity. In addition, spider silks are biocompatible and biodegradable, which make them excellent material for biomedical applications. Due to these characteristics spider silks can be used to make such diverse products as parachute cords, bulletproof vests, composite material for aircrafts, products used in cosmetics, bandages to promote wound healing, drug delivery systems, and scaffolds for tissue engineering [46–49].
Despite the biotechnological potential of spider silks, there are limitations in harvesting large quantities of silk from spiders. They cannot be farmed as for the silkworm (Bombyx mori) owing to their aggressive behavior and territorial nature. In addition, silk must be harvested manually, directly from a spider or collected from the web, both extremely time-consuming procedures. Recently, a golden color spider silk cape was made in Madagascar from 1.2 million golden orb spiders (genus Nephila) which took 8 years to collect [50]. Thus, new strategies based on genetic engineering have been developed for the production of new biomaterials based on the molecular structure of spider silks.
9.3.1 Production of Recombinant Silk Proteins
Several heterologous host systems have been used for the production of different spider silk proteins [50, 51]. The most widely used host is the gram-negative enterobacterium, Escherichia coli, which offers a well-controlled, cost-efficient system suitable for large-scale production. However, spider silk proteins are rather difficult to produce, mainly because of the highly repetitive nature of the nucleotide sequence and the high content of specific amino acids, mainly alanine and glycine, which require a large pool of the respective tRNAs [47, 50]. For this reason, attempts to produce major ampullate silks based on the native DNA sequence in E. coli resulted in low protein yields and instability of the spider silk gene, which led to the production of limited size proteins [19]. As an alternative, the methylotrophic yeast, Pichia pastoris, was used and was able to produce and secrete recombinant silk proteins up to 65 kDa to the extracellular medium [20]. Mammalian cell lines have also been investigated as a potential host system [21]. Although they were capable of producing spider silk proteins with molecular size ranging from 60 to 140 kDa, as the size of the recombinant silk gene increased, translational pausing occurred, resulting in the expression of heterogeneous size proteins and a decrease in the protein yield.
In order to optimize protein yield and size, strategies using synthetic genes designed with host-specific codons were developed. Additionally, completely novel silk proteins were created by hybridizing motifs from different silk types. Several groups have used these approaches to produce a variety of engineered silk proteins, most of which are based on the sequences of major and flagelliform spidroins [52–55]. Teulé and colleagues [56] developed an iterative polymerization strategy to produce very large and repetitive spider silk genes in a precisely controlled manner (Fig. 9.3). This technique is extremely powerful since it allows the construction of basic repeats that exactly reflect the native silk protein sequences or represent new engineered variants of native sequences by mixing and matching structural motifs in different ratios and combinations [57]. This approach is also extremely flexible since two different recombinant plasmids containing silk multimer inserts at different stages of assembly can be used simultaneously as templates to generate a higher size multimer [56]. Utilizing a similar strategy, Xia et al [58] were able to produce a native-sized recombinant spider silk protein in a metabolically engineered E. coli with an elevated glycyl-tRNA pool. The 284.9 kDa recombinant protein was based on the MaSp1 motif from N. clavipes, which is extremely rich in glycine (43–45 %), and was produced by fed-batch culture reaching a protein yield of 2.7 g/L. In addition, the mechanical properties of the fibers spun with this ultrahigh molecular weight recombinant spider silk were comparable to those of the native one. Thus, the high molecular weight of native spider silk proteins indeed plays an important role in determining its mechanical properties.
Strategy for construction of a tandem spider silk synthetic gene based on a monomer with compatible but nonregenerable sites. ⋆ denotes DNA cut by a restriction enzyme. N × indicates that this strategy can be repeated as many times as one wishes (Reproduced with permission from Teulé et al. [56])
Another approach to producing recombinant spider silks utilizes transgenic animals and plants that carry a segment of silk DNA in their genome. Mice and goats have been transgenically modified to produce spider silk proteins in the female’s mammary gland [59, 60]. However, the maximum protein yields observed for recombinant silk production in transgenic animals are low when compared with bacteria (117 mg/l and up to 300 mg/l, respectively) [19, 61, 62]. Scheller et al. [63] reported the expression of spider silk proteins in tobacco and potatoes for the first time. Using the Cauliflower mosaic virus (CaMV) 35S promoter, stable transgenic tobacco and potato lines were engineered to express different sizes of the MaSp1 gene derivatives from N. clavipes in the endoplasmic reticulum (ER) of leaves. This technique has proven to be effective for the expression of MaSp1 and MaSp2 derivatives in greenhouse and field trials [23]. Additionally, recombinant MaSp1-like proteins were also produced in Arabidopsis leaves and seeds, as well as in somatic soybean embryos [22]. These results demonstrated that recombinant spider silk proteins had higher accumulation levels in seeds than in the leaves. Recently, the production of synthetic, high molecular weight spider silk proteins larger than 250 kDa based on the assembly of Flag protein monomers via intein-mediated trans-splicing was achieved in plants [64]. This method avoids the need for highly repetitive transgenes, which may result in higher genetic and transcriptional stability. The resulting multimeric structures formed microfibers with up to 500 μm in length and diameters ranging from 1 to 2 μm, thereby demonstrating their great potential as a biomaterial.
Because of their high capacity to produce natural silk, silkworms (B. mori) have also been studied as a host for the production of recombinant spider silk proteins. In a recent publication [65], transgenic silkworms were developed using a piggyBac vector composed of the B. mori fibroin heavy chain promoter and enhancer, a genetic sequence encoding a 78 kDa synthetic spider silk, and an enhanced green fluorescent protein (EGFP) tag (Fig. 9.4). The chimeric silkworm/spider silk fibers produced by the transgenic B. mori were significantly tougher than silkworm silk and as tough as dragline spider silk native fibers. These results showed that silkworms can be engineered to manufacture composite fibers containing stably integrated spider silk protein sequences with improved mechanical properties.
Expression of a chimeric silkworm/spider silk/EGFP protein in Bombyx mori (a) cocoons, (b, c) silk glands, and (d) silk fibers (Reproduced with permission from Teulé et al. [65])
9.3.2 New Biomaterials from Recombinant Spider Silk Proteins
Silk proteins have been shown to solubilize in water, organic solvents, and ionic liquids, indicating the versatile options available [66]. In addition, depending on the processing route, phase transition of spider silk proteins is controlled by parameters of chemical and physical processes leading to different material morphologies, including fibers, films, gels, porous sponges, and other related systems (Fig. 9.5). For instance, a film can be formed by simple evaporation of the solvent, and if porogens are introduced into the silk protein solution prior to evaporation of the solvent, porous structures or foams can be produced [46].
Processing of silk into new biomaterials. Native and recombinant spider silks can be solubilized and processed into a range of different structures (Reproduced from Kluge et al. [66] with permission of Elsevier Limited)
Transparent films, ranging in thickness from 0.5 to 1.5 μm, have been produced using a hexafluoroisopropanol (HFIP) solution containing recombinant dragline silk proteins. Treating the films with potassium phosphate or methanol could convert the proteins’ secondary structure from α-helix to β-sheet, making them water insoluble [67]. The portion of induced β-sheet structure can be regulated by the incubation time or by treatment of the film with methanol/water solutions mixed at different ratios. Different contents of β-sheet structure will not only determine the stability of silk films but will also influence their mechanical properties. Generally, a higher amount of β-sheet increases the elastic modulus and strength of the film but reduces its elasticity [46].
Microparticles are also one of the manifold morphologies spider silk proteins can adopt. These particles are formed by precipitating aqueous silk protein solutions with kosmotropic salts such as potassium phosphate or ammonium sulfate [68, 69]. The diameter of the spherical colloidal assemblies is variable (100 nm to 10 μm) and depends on the initial protein concentration and the chosen kosmotropic salt. For example, the addition of potassium phosphate (>300 mM) results in increased hydrophobic interactions accompanied by a dense packing of the spider silk proteins. Such tight packing leads to the formation of chemically stable spheres, with a high content of β-sheet secondary structure and extremely smooth surfaces [70].
Despite the wide range of biomaterials that can be assembled from recombinant spider silk proteins, the mechanical properties exhibited by drawn synthetic fibers are still below the measured properties of native spider silks. Solvent extrusion [56], electrospinning [71], and microfluidics [72] are some of the techniques that have already been used to process recombinant spider silk proteins into fibers (Fig. 9.6). However, it is still difficult to manufacture long lengths of material in sufficient quantity and with superior quality. Moreover, most of the fibers created by spinning processes are so brittle that their mechanical properties cannot be properly measured [71, 73]. Even so, these studies are important to provide some understanding about the main keys to spinning recombinant silk proteins. For instance, drawing the newly formed fiber for strain elongation improves the mechanical properties of synthetic fibers by the induction of β-sheet formation [61, 74]. Future challenges will include the optimization of the spinning processes in order to scale up production and maximize the properties of the processed material.
Schematic representation of current silk fiber-forming techniques. (a) Solvent extrusion is performed by drawing the fiber through a coagulation bath in a controlled manner. Microfluidic devices use a contracting channel and multiple solvent inputs to regulate the geometry and chemistry of the resulting fiber. Electrospinning processes combine strong voltage gradients and syringe pump extrusion and result in either random or aligned fiber deposition. (b) Characteristics of the different fiber-forming processes. To date, electrospinning has proved to be the most useful approach, mainly because of the low amounts of spider silk materials needed and the utility of the resulting electrospun fiber mats for cell and tissue culture studies (Reproduced from Kluge et al. [66] with permission of Elsevier Limited)
9.3.3 Biotechnological Applications of Synthetic Spider Silk
Due to their outstanding mechanical properties, different biotechnological applications have been foreseen for spider silk proteins. They have desirable self-assembly characteristics, which allow their utilization as molecular building blocks for the production of different biomaterials. Furthermore, they can be genetically modified to allow the covalent coupling of peptides, enzymes, or particles before and after silk processing into biomaterials, therefore expanding the range of potential applications [75–78].
Recently, the synthesis of novel spider silk uranyl-binding proteins was demonstrated through bioengineering of varying repeats of a uranyl-binding sequence adopted from a mutated calmodulin sequence in combination with spider silk sequences [79]. This new chimeric protein system can be used in environmental engineering as a substrate for uranium collection and removal. Once selective metal binding can be exploited, the development of such silk-based material might also find applications in microelectronics, biosensors, solar energy, and electrical batteries [79].
Spider silks also possess several desired characteristics for biomedical applications, such as biocompatibility, low inflammatory potential, antimicrobial activity, and a slow rate of degradation. The specific biomedical application of synthetic spider silks will not only rely on its molecular structure but also on the morphology adopted by the silk proteins. Silk fibroin films, for instance, can be used for different medical applications such as lenses, drug-loading films, and, mainly, cell-supporting scaffolds for tissue engineering [80–83]. Genes and drug delivery platforms can be developed using spider silk microcapsules [48, 84, 85]. Spider silk proteins can also be processed into coatings that can be used to improve the biocompatibility and surface properties of biomaterials, such as silicone implants [86].
Overall, the macroscopic structure of silk-based materials will be responsible for their interface with the environment, and the advantages and limitations of each system will determine its use in a specific application or for commercial exploitation [66].
9.4 Concluding Remarks
In the last few decades, several prokaryotic and eukaryotic hosts have been tested for the production of recombinant spider silk proteins. So far, there has been considerable progress in understanding the genetic organization of spider silk proteins and the self-assembly and processing of synthetic spider silks into many material formats. Recently, the production of native-sized (285 kDa) spider silk protein and the production of chimeric silkworm/spider silk fibers with improved mechanical properties have been reported [58, 65].
For any commercial application of synthetic spider silks, it is still necessary to develop new strategies for the production of homogeneous and reproducible fibers with defined mechanical properties. Moreover, applications of spider silks in commodity materials, such as textiles, high-performance composite materials, and durable and tough materials, in general, will require the success of robust and low-cost expression systems coupled with environmentally friendly assembly processes. Further exploitation of the aqueous processing and assembly of highly hydrophobic silk proteins into material structures together with a better comprehension of the natural events associated with the storage and spinning of the silk in spiders may be the keys to the development of a green improved biomaterial production and design.
Abbreviations
- A :
-
Amino acid alanine
- CaMV:
-
Cauliflower mosaic virus
- EGFP:
-
Enhanced green fluorescent protein
- ER:
-
Endoplasmic reticulum
- Flag:
-
Flagelliform spidroin
- G:
-
Amino acid glycine
- HFIP:
-
Hexafluoroisopropanol
- MaSp1:
-
Major ampullate spidroin 1
- MaSp2:
-
Major ampullate spidroin 2
- MiSp:
-
Minor ampullate spidroin
- P:
-
Amino acid proline
- T-RNA:
-
Transfer ribonucleic acid
References
Lewis, R. V. (2006). Spider silk: ancient ideas for new biomaterials. Chemical Reviews, 106, 37262–3774.
Gosline, J. M., Guerette, P. A., Ortlepp, C. S., & Savage, K. N. (1999). The mechanical design of spider silks: From fibroin sequence to mechanical function. The Journal of Experimental Biology, 202, 3295–3303.
Gosline, J. M., Denny, M. W., & DeMont, M. E. (1984). Spider silk as rubber. Nature, 309, 551–552.
Blackledge, T. A., & Hayashi, C. Y. (2006). Silken toolkits: Biomechanics of silk fibers spun by the orb web spider Argiope argentata (Fabricius 1775). The Journal of Experimental Biology, 209, 2452–2461.
Vollrath, F., & Knight, D. P. (2001). Liquid crystalline spinning of spider silk. Nature, 410, 541–548.
Knight, D. P., & Vollrath, F. (2001). Changes in element composition along the spinning duct in a Nephila spider. Naturwissenschaften, 4, 179–182.
Sponner, A., Schlott, B., Vollrath, F., Unger, E., et al. (2005). Characterization of the protein components of Nephila clavipes dragline silk. Biochemistry, 44, 4727–4736.
Gatesy, J., Hayashi, C., Motriuk, D., Woods, J., & Lewis, R. (2001). Extreme diversity, conservation, and convergence of spider silk fibroin sequences. Science, 291, 2603–2605.
Hayashi, C. Y., Shipley, N. H., & Lewis, R. V. (1999). Hypotheses that correlate the sequence, structure, and mechanical properties of spider silk proteins. International Journal of Biological Macromolecules, 24, 271–275.
Savage, K. N., & Gosline, J. M. (2008). The role of proline in the elastic mechanism of hydrated spider silks. The Journal of Experimental Biology, 211, 1948–1957.
Adrianos, S. L., Teulé, F., Hinman, M. B., Jones, J. A., Weber, W. S., Yarger, J. L., & Lewis, R. V. (2013). Nephila clavipes Flagelliform silk-like GGX motifs contribute to extensibility and spacer motifs contribute to strength in synthetic spider silk fibers. Biomacromolecules, 14, 1751–1760.
Vollrath, F., Barth, P., Basedow, A., Engstrom, W., & List, H. (2002). Local tolerance to spider silks and protein polymers in vivo. In Vivo, 16, 229–234.
Altman, G. H., Diaz, F., Jakuba, C., Calabro, T., et al. (2003). Silk-based biomaterials. Biomaterials, 24, 401–416.
Wright, S., & Goodacre, S. L. (2012). Evidence for antimicrobial activity associated with common house spider silk. BMC Research Notes, 5, 326.
Horan, R. L., Antle, K., Collette, A. L., Wang, Y., et al. (2005). In vitro degradation of silk fibroin. Biomaterials, 26, 3385–3393.
Sponner, A. (2007). Spider silk as a resource for future biotechnologies. Entomological Research, 37, 238–250.
Allmeling, C., Jokuszies, A., Reimers, K., Kall, S., et al. (2008). Spider silk fibres in artificial nerve constructs promote peripheral nerve regeneration. Cell Proliferation, 41, 408–420.
Vollrath, F., & Porter, D. (2009). Silks as ancient models for modern polymers. Polymer, 50, 5623–5632.
Fahnestock, S. R., & Irwin, S. L. (1997). Synthetic spider dragline silk proteins and their production in Escherichia coli. Applied Microbiology and Biotechnology, 47, 23–32.
Fahnestock, S. R., & Bedzyk, L. A. (1997). Production of synthetic spider dragline silk protein in Pichia pastoris. Applied Microbiology and Biotechnology, 47, 33–39.
Lazaris, A., Arcidiacono, S., Huang, Y., Zhou, J.-F., Duguay, F., Chretien, N., Welsh, E. A., Soares, J. W., & Karatzas, C. N. (2002). Spider silk fibers spun from soluble recombinant silk produced in mammalian cells. Science, 259, 472–476.
Barr, L. A., Fahnestock, S. R., & Yang, J. J. (2004). Production and purification of recombinant DP1B silk-like protein in plants. Molecular Breeding, 13, 345–356.
Menassa, R., Zhu, H., Karatzas, C. N., Lazaris, A., et al. (2004). Spider dragline silk proteins in transgenic tobacco leaves: Accumulation and field production. Plant Biotechnology Journal, 2, 431–438.
Xu, H. T., Fan, B. L., Yu, S. Y., Huang, Y. H., Zhao, Z. H., Lian, Z. X., Dai, Y. P., Wang, L. L., Liu, Z. L., Fei, J., & Li, N. (2007). Construct synthetic gene encoding artificial spider dragline silk protein and its expression in milk of transgenic mice. Animal Biotechnology, 18, 1–12.
Bittencourt, D., Oliveira, P. F., Prosdocimi, F., & Rech, E. L. (2012). Protein families, natural history and biotechnological aspects of spider silk. Genetics and Molecular Research, 11, 2360–2380.
Tokareva, O., Jacobsen, M., Buehler, M., Wong, J., & Kaplan, D. L. (2014). Structure-function-property-design interplay in biopolymers: Spider silk. Acta Biomaterialia, 10, 1612–1626.
Gosline, J., Lillie, M., Carrington, E., Guerette, P., Ortlepp, C., & Savage, K. (2002). Elastic proteins: Biological roles and mechanical properties. Philosophical Transactions of the Royal Society of London. Series B: Biological Sciences, 357, 121–132.
Blackledge, T. A., & Hayashi, C. Y. (2006). Unraveling the mechanical properties of composite silk threads spun by cribellate orb-weaving spiders. The Journal of Experimental Biology, 209, 3131–3140.
Agnarsson, I., Kuntner, M., & Blackledge, T. A. (2010). Bioprospecting finds the toughest biological material: Extraordinary silk from a giant riverine orb spider. PLoS One, 5, e11234.
Hinman, M. B., & Lewis, R. V. (1992). Isolation of a clone encoding a second dragline silk fibroin. Nephila clavipes dragline silk is a two-protein fiber. Journal of Biological Chemistry, 267, 19320–19324.
Keten, S., & Buehler, M. J. (2010). Nanostructure and molecular mechanics of spider dragline silk protein assemblies. Journal of the Royal Society Interface, 7, 1709–1721.
Termonia, Y. (1994). Molecular modelling of spider silk elasticity. Macromolecules, 27, 7378–7381.
Liu, Y., Shao, Z. Z., & Vollrath, F. (2005). Relationships between supercontraction and mechanical properties of spider silk. Nature Materials, 4, 901–905.
Vollrath, F., & Knight, D. P. (1999). Structure and function of the silk production pathway in the spider Nephila edulis. International Journal of Biological Macromolecules, 24, 243–249.
Marhabaie, M., Leeper, T. C., & Blackledge, T. A. (2014). Protein composition correlates with the mechanical properties of spider (Argiope trifasciata) dragline silk. Biomacromolecules, 15, 20–29.
Brooks, A. E., Steinkraus, H. B., Nelson, S. R., & Lewis, R. V. (2005). An investigation of the divergence of major ampullate silk fibers from Nephila clavipes and Argiope aurantia. Biomacromolecules, 6, 3095–3099.
Hagn, F., Eisoldt, L., Hardy, J. G., Vendrely, C., et al. (2010). A conserved spider silk domain acts as a molecular switch that controls fibre assembly. Nature, 465, 239–242.
Silvers, R., Buhr, F., & Schwalbe, H. (2010). The molecular mechanism of spider-silk formation. Angewandte Chemie International Edition in English, 49, 5410–5412.
Hayashi, C. Y., & Lewis, R. V. J. (1998). Evidence from flagelliform silk cDNA for the structural basis of elasticity and modular nature of spider silks. Molecular Biology, 275, 773–784.
Martino, M., Bavoso, A., Guantieri, V., Coviello, A., & Tamburro, A. M. (2000). On the occurrence of polyproline II structure in elastin. Journal of Molecular Structure, 519, 173–189.
Bittencourt, D., Souto, B. M., Verza, N. C., Vinecky, F., Dittmar, K., Silva, P. I., Jr., Andrade, A. C., da Silva, F. R., Lewis, R. V., & Rech, E. L. (2007). Spidroins from the Brazilian spider Nephilengys cruentata (araneae: Nephilidae). Comparative Biochemistry and Physiology. Part B, Biochemistry and Molecular Biology, 147, 597–606.
Hayashi, C. Y., & Lewis, R. V. (2000). Molecular architecture and evolution of a modular spider silk protein gene. Science, 287, 1477–1479.
Hinman, M. B., Jones, J. A., & Lewis, R. V. (2000). Synthetic spider silk: a modular fiber. Trends in Biotechnology, 18, 374–379.
Vollrath, F., Fairbrother, W. J., Williams, R. J. P., Tillinghast, E. K., et al. (1990). Compounds in the droplets of the Orb Spiders Viscid Spiral. Nature, 345, 526–528.
Choresh, O., Bayarmagnai, B., & Lewis, R. V. (2009). Spider web glue: Two proteins expressed from opposite strands of the same DNA sequence. Biomacromolecules, 10, 2852–2856.
Spiess, K., Lammel, A., & Scheibel, T. (2010). Recombinant spider silk proteins for applications in biomaterials. Macromolecular Bioscience, 10, 998–1007.
Rising, A., Widhe, M., Johansson, J., & Hedhammar, M. (2011). Spider silk proteins: Recent advances in recombinant production, structure-function relationships and biomedical applications. Cellular and Molecular Life Sciences, 68, 169–184.
Hofer, M., Winter, G., & Myschik, J. (2012). Recombinant spider silk particles for controlled delivery of protein drugs. Biomaterials, 33, 1554–1562.
Schacht, K., & Scheibel, T. (2014). Processing recombinant spider silk proteins into tailor-made materials for biomaterials applications. Current Opinion in Biotechnology, 29, 62–69.
Chung, H., Kim, T. Y., & Lee, S. Y. (2012). Recent advances in production of recombinant spider silk proteins. Current Opinion in Biotechnology, 23, 957–996.
Tokareva, O., Michalczechen-Lacerda, V. A., Rech, E. L., & Kaplan, D. L. (2013). Recombinant DNA production of spider silk proteins. Microbial Biotechnology, 6, 651–663.
Vendrely, C., & Scheibel, T. (2007). Biotechnological production of spider-silk proteins enables new applications. Macromolecular Bioscience, 7, 401.
Heim, M., Ackerschott, C. B., & Scheibel, T. (2010). Characterization of recombinantly produced spider flagelliform silk domains. Journal of Structural Biology, 170, 420.
Rising, A., Widhe, M., Johansson, J., & Hedhammar, M. (2011). Spider silk proteins: Recent advances in recombinant production, structure-function relationships and biomedical applications. Cellular and Molecular Life Sciences, 68, 169–184.
Teulé, F., Addison, B., Cooper, A. R., Ayon, J., Henning, R. W., Benmore, C. J., et al. (2012). Combining flagelliform and dragline spider silk motifs to produce tunable synthetic biopolymer fibers. Biopolymers, 97, 418–431.
Teulé, F., Cooper, A. R., Furin, W. A., Bittencourt, D., Rech, E. L., Brooks, A., & Lewis, R. V. (2009). A protocol for the production of recombinant spider silk-like proteins for artificial fiber spinning. Nature Protocols, 4, 324–355.
Brooks, A. E., Stricker, S. M., Joshi, S. B., Kamerzell, T. J., Middaugh, C. R., & Lewis, R. V. (2008). Properties of synthetic spider silk fibers based on Argiope aurantia MaSp2. Biomacromolecules, 9, 1506–1510.
Xia, X. X., Qian, Z. G., Ki, C. S., Park, Y. H., Kaplan, D. L., & Lee, S. Y. (2010). Native-sized recombinant spider silk protein produced in metabolically engineered Escherichia coli results in a strong fiber. Proceedings of the National Academy of Sciences of the United States of America, 107, 14059–14063.
Xu, H. T., Fan, B. L., Yu, S. Y., Huang, Y. H., Zhao, Z. H., Lian, Z. X., et al. (2007). Construct synthetic gene encoding artificial spider dragline silk protein and its expression in milk of transgenic mice. Animal Biotechnology, 18, 1–12.
Steinkraus, H. B., Rothfuss, H., Jones, J. A., Dissen, E., Shefferly, E., & Lewis, R. V. (2012). The absence of detectable fetal microchimerism in nontransgenic goats (Capra aegagrus hircus) bearing transgenic offspring. Journal of Animal Science, 90, 481–488.
An, B., Hinman, M. B., Holland, G. P., Yarger, J. L., & Lewis, R. V. (2011). Inducing β-sheets formation in synthetic spider silk fibers by aqueous post-spin stretching. Biomacromolecules, 12, 2375–2381.
Dams-Kozlowska, H., Majer, A., Tomasiewicz, P., Lozinska, J., Kaplan, D. L., & Mackiewicz, A. (2012). Purification and cytotoxicity of tag-free bioengineered spider silk proteins. Journal of Biomedical Materials Research. Part A, 101A, 456–464.
Scheller, J., Guhrs, K. H., Grosse, F., & Conrad, U. (2001). Production of spider silk proteins in tobacco and potato. Nature Biotechnology, 19, 573–577.
Hauptmann, V., Weichert, N., Menzel, M., Knoch, D., et al. (2013). Native sized spider silk proteins synthesized in planta via intein-based multimerization. Transgenic Research, 22, 369–377.
Teulé, F., Miao, Y. G., Sohn, B. H., Kim, Y. S., Hull, J. J., Fraser, M. J., et al. (2012). Silkworms transformed with chimeric silkworm/spider silk genes spin composite silk fibers with improved mechanical properties. Proceedings of the National Academy of Sciences of the United States of America, 109, 923–928.
Kluge, J. A., Rabotyagova, O., Leisk, G. G., & Kaplan, D. L. (2008). Spider silks and their applications. Trends in Biotechnology, 26, 244–251.
Huemmerich, D., et al. (2006). Processing and modification of films made from recombinant spider silk proteins. Applied Physics A, 82, 219–222.
Lammel, A., Schwab, M., Slotta, U., Winter, G., & Scheibel, T. (2008). Processing conditions for the formation of spider silk microspheres. ChemSusChem, 1, 413–416.
Slotta, U. K., Rammensee, S., Gorb, S., & Scheibel, T. (2008). An engineered spider silk protein forms microspheres. Angewandte Chemie International Edition in English, 47, 4592–4594.
Rammensee, S., Slotta, U., Scheibel, T., & Bausch, A. R. (2008). Assembly mechanism of recombinant spider silk proteins. Proceedings of the National Academy of Sciences of the United States of America, 105, 6590–6595.
Arcidiacono, S., Mello, C. M., Butler, M., Welsh, E., Soares, J. W., Allen, A., Ziegler, D., Laue, T., & Chase, S. (2002). Aqueous processing and fiber spinning of recombinant spider silks. Macromolecules, 35, 1262–1266.
Scheibel, T., Huemmerich, D., Remmensee, S., Freudiger, C., & Bausch, A. (2007). Microfluidic device for controlled aggregation of spider silk. US patent WO 2007/141131 A1.
Slotta, U., Mougin, N., Romer, L., & Leimer, A. H. (2012). Synthetic spider silk proteins and threads. Chemical Engineering Progress, 108, 43–49.
Albertson, A. E., Teulé, F., Weber, W., Yarger, J. L., & Lewis, R. V. (2014). Effects of different post-spin stretching conditions on the mechanical properties of synthetic spider silk fibers. Journal of the Mechanical Behavior of Biomedical Materials, 29, 225–234.
Lee, S. M., Pippel, E., Gösele, U., Dresbach, C., Qin, Y., Chandran, C. V., Bräuniger, T., Hause, G., & Knez, M. (2009). Greatly increased toughness of infiltrated spider silk. Science, 324, 488–492.
Gomes, S. C., Leonor, I. B., Mano, J. F., Reis, R. L., & Kaplan, D. L. (2011). Antimicrobial functionalized genetically engineered spider silk. Biomaterials, 32, 4255–4266.
Belton, D. J., Mieszawska, A. J., Currie, H. A., Kaplan, D. L., & Perry, C. C. (2012). Silk-silica composites from genetically engineered chimeric proteins: Materials properties correlate with silica condensation rate and colloidal stability of the proteins in aqueous solution. Langmuir, 28, 4373–4381.
Vidal, G., Blanchi, T., Mieszawska, A. J., Calabrese, R., Rossi, C., Vigneron, P., et al. (2013). Enhanced cellular adhesion on titanium by silk functionalized with titanium binding and RGD peptides. Acta Biomaterialia, 9, 4935–4943.
Krishnaji, S. T., & Kaplan, D. L. (2013). Bioengineered chimeric spider silk-uranium binding proteins. Macromolecular Bioscience, 13, 256–264.
Karageorgiou, V., Meinel, L., Hofmann, S., Malhotra, A., et al. (2004). Bone morphogenetic protein-2 decorated silk fibroin films induce osteogenic differentiation of human bone marrow stromal cells. Journal of Biomedial Materials Research Part A, 71, 528–537.
Leal-Egana, A., & Scheibel, T. (2012). Interactions of cells with silk surfaces. Journal of Materials Chemistry, 22, 14330–14336.
Bauer, F., Wohlrab, S., & Scheibel, T. (2013). Controllable cell adhesion, growth and orientation on layered silk protein films. Biomaterials Science, 1, 1244–1249.
Jansson, R., Thatikonda, N., Lindberg, D., Rising, A., Johansson, J., Nygren, P. Å., & Hedhammar, M. (2014). Recombinant spider silk genetically functionalized with affinity domains. Biomacromolecules, 15, 1696–1706.
Lammel, A., Schwab, M., Hofer, M., Winter, G., & Scheibel, T. (2011). Recombinant spider silk particles as drug delivery vehicles. Biomaterials, 32, 2233–2240.
Numata, K., Mieszawska-Czajkowska, A. J., Kvenvold, L. A., & Kaplan, D. L. (2012). Silk-based nanocomplexes with tumor-homing peptides for tumor-specific gene delivery. Macromolecular Bioscience, 12, 75–82.
Zeplin, P. H., Maksimovikj, N. C., Jordan, M. C., Nickel, J., Lang, G., Leimer, A. H., Römer, L., & Scheibel, T. (2014). Spider silk coatings as a bioshield to reduce periprosthetic fibrous capsule formation. Advanced Functional Materials, 24, 2658–2666.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
de C. Bittencourt, D.M. (2016). Spider Silks and Their Biotechnological Applications. In: Raman, C., Goldsmith, M., Agunbiade, T. (eds) Short Views on Insect Genomics and Proteomics. Entomology in Focus, vol 4. Springer, Cham. https://doi.org/10.1007/978-3-319-24244-6_9
Download citation
DOI: https://doi.org/10.1007/978-3-319-24244-6_9
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-24242-2
Online ISBN: 978-3-319-24244-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)