Abstract
An overview over the role and past evolution of High Throughput Screening (HTS) in early drug discovery is given and the different screening phases which are sequentially executed to progressively filter out the samples with undesired activities and properties and identify the ones of interest are outlined. The goal of a complete HTS campaign is to identify a validated set of chemical probes from a larger library of small molecules, antibodies, siRNA, etc. which lead to a desired specific modulating effect on a biological target or pathway. The main focus of this chapter is on the description and illustration of practical assay and screening data quality assurance steps and on the diverse statistical data analysis aspects which need to be considered in every screening campaign to ensure best possible data quality and best quality of extracted information in the hit selection process. The most important data processing steps in this respect are the elimination of systematic response errors (pattern detection, categorization and correction), the detailed analysis of the assay response distribution (mixture distribution modeling) in order to limit the number of false negatives and false discoveries (false discovery rate and p-value analysis), as well as selecting appropriate models and efficient estimation methods for concentration-response curve analysis.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Compound and RNAi screening processes
- Data quality control
- Data normalization
- Correction of systematic response errors
- Hit identification and ranking
- Dose-response curve analysis
1 Introduction
1.1 HTS in Drug Discovery
The beginnings of High-Throughput Screening (HTS) in the pharmaceutical and biotech industry go back to the early 1990s when more and more compounds needed to be tested for a broader series of targets in an increasing number of biological assay systems. In some companies the investigation of larger compound series for activity in a biochemical or cell-based assay systems had its origins in natural product screening, but was extended to look for modulating effects of compounds from the historical and growing in-house collections and added libraries of compounds from combinatorial synthesis. The goal of HTS is the identification of a subset of molecules (small molecular compounds, siRNA, antibodies, antibody conjugates, etc.) from a larger library which have a modulating effect on a given biological target. A large part of HTS work was, and still is, centered on investigating the effects of small molecules against various intra- and extracellular molecular targets and identifying compounds and compound series with a desired mode of action. In the past 20 years these compound collections have been strongly and continually enhanced by complementing the initially available sets with further sets from in-house syntheses, carefully selected additions from commercial sources, various classes of natural products, and known drugs. Stored libraries of compounds covering a broad chemical space are thus available for repeated screening or picking for special purpose investigations and have reached several 100,000 in many biotech and screening service companies, as well as academic facilities, and >1 million (up to 2 million) in most major pharmaceutical companies. Automated compound stores and retrieval systems are in use in most of the companies and allow fast replication of compound library copies into assay plates for regular screening and fast picking of sub-libraries of smaller sets of compounds for more focused screens and follow-up confirmation and verification of any activity found in the broader initial (primary) screening rounds (Fox et al. 2006; Pereira and Williams 2007; Mayr and Fuerst 2008; Mayr and Bojanic 2009; Macarron et al. 2011).
On the experimental side essentially all High-Throughput Screening experiments are performed in a highly automated, standardized and controlled fashion using microtiter plates with 96, 384 or 1536 wells, i.e. grids of ‘reaction containers’ embedded in rectangular plastic plates following an industry standard format with typically between 50–300 μL (96), 10–100 μL (384) and 1–10 μL (1536) working volumes containing the biological material (proteins, cells, cell fragments), assay buffer, reagents and the compound (or other) sample solutions whose activities will be determined. Nowadays the industrial HTS labs essentially only use 384- and 1536 well plates, whereas lower throughput labs may still perform a part of their experiments and measurements in 96-well plates. Most of the quality control diagnostics and data analysis aspects discussed later in this chapter can—and should—be applied irrespective of actual plate formats and throughput (ultrahigh, high, mid or low).
In essentially the same time period the sequencing of the human (and other) genomes has allowed to identify several thousand potential molecular targets for pharmaceutical intervention, some (but by far not all) of them coming with an understanding of the function and, thus, allowing the pursuit of drug discovery efforts. Large efforts in functional genomics, i.e. the determination of the function of genes, RNA transcripts and the resulting protein products as well as their regulation are needed on top of generating the pure sequence information to identify potentially druggable targets (Sakharkar et al. 2007; Bakheet and Doig 2009). The methods of identification and validation of disease-relevant molecular targets are wide and diverse (Kramer and Cohen 2004; Hughes et al. 2011). Information from DNA, RNA and protein expression profiling, proteomics experiments, phenotypic observations, RNAi (RNA interference) screens and extensive data mining and bioinformatics approaches is used to link information on diseases, gene functions and biological interaction networks with molecular properties. In RNAi screening larger libraries of siRNA or shRNA samples are used to investigate the modulation of gene function, the subsequent modification of protein expression levels and resulting loss of function in large scale high-throughput experiments (genome-scale RNAi research, genome-wide screens seeking to identify all possible regulators of a biological process, or screens limited to a subset of target genes related to a specific biological pathway) using phenotypic readouts (Matson 2004; Root et al. 2006). Several of the target identification and validation approaches thus share a series of technological elements (automation, measurement equipment and data analysis methods) with the ‘classical’ plate-based small molecule HTS, and most of the data quality assessment methods and hit selection steps described below can be directly applied in these areas. Some of the differences in specific analysis steps for particular types of probes, e.g. for RNAi screens will be mentioned.
A high degree of automation and standardized assay execution processes are key ingredients for effective HTS and quite large investments into the development and deployment of robotics and automated laboratory equipment have been made by the industry in the past two decades. Many vendors have also developed smaller automation workstations or work cells which can be installed and used in smaller laboratories. As with most initially specialized technologies we are also seeing here a movement from the centralized industry labs into low- and mid-throughput laboratories which are more distributed in drug discovery organizations, and also into academic institutions (Kaiser 2008; Baker 2010). The large body of experience gained over more than two decades in the pharmaceutical industry and larger government laboratories about optimized sample management and screening processes, possible screening artifacts and suitable data analysis techniques can beneficially be applied in the institutions which have more recently adopted these technologies. On the other hand, some of the statistical error correction methods described further below were elaborated within the last decade in academic institutions and are now benefitting the whole screening community.
Robotic screening systems come either as fully integrated setups where all assay steps (plate movement, reagent addition, compound addition, incubation, readouts, discarding of used plates) are scheduled and executed in automated fashion, or as a series of separate independent workstation cells which are used sequentially, often with a ‘manual’ transfer of plate batches between them. The large fully integrated HTS systems can process and measure up to 100,000 compounds or even more per day (ultra-high throughput screening, uHTS), depending on assay technology, details of the assay protocol and processing times for individual steps. Workstation based mid-throughput screening (MTS) typically reaches throughputs of 10,000–20,000 samples/day (e.g. in batches of 20–50 384 well plates per day). Throughput will naturally be smaller if lengthy process steps like e.g. incubation phases are needed or if the measurement time is prolonged, either because of limitations of the measurement instrument, or because the course of a biochemical reaction needs to be followed for a certain amount of time. The aspect of performing the complete set of experiments of a particular screen in separate batches of individual plates is clearly very important and will have an effect on the necessary data analysis steps.
The optimization of the high throughput screening processes has over time gone through different phases which have initially focused on obtaining higher throughput and larger numbers of screened entities, then on higher sophistication in assay optimization, standardization of processes, miniaturization, added pilot-, counter- and orthogonal screening experiments, as well as improved analysis techniques, i.e. a focus on higher efficiency and better data quality, and in the last 8–10 years the screening and decision making processes were set up in much more flexible ways to better allow diversified and case-by-case handling for each campaign—either full deck screens or adapted focused screens including best possible validation of results with parallel specificity and selectivity screens to obtain higher quality and better characterized hits than resulting from the ‘raw’ hit list of the main screen (Mayr and Fuerst 2008; Macarron et al. 2011).
Counter screens are used to identify compound samples which don’t show an activity directed towards the intended biological target, but which nonetheless give positive readouts (‘false positive responses’ in an assay) by interfering with the readout mechanism or act otherwise nonspecifically. Some compounds will even do this in concentration dependent manner, thus mimicking a desired activity. This can occur due to aggregation, colored components of assay ‘cocktail’, fluorescence, inhibition of detection enzymes (reporter mechanism), cytotoxicity, etc. (Thorne et al. 2010; Hughes et al. 2012).
An orthogonal screen is based on an assay which uses a different format or a different readout mechanism to measure the same phenomenon and confirm that the activity is really directed towards the target of interest. Compounds which are active in both the original and orthogonal assay are usually prioritized for follow-up work.
Selectivity screens are used to determine whether a particular compound is acting solely on the target of interest or also on other targets of the same family (e.g. enzymes, protease inhibitors, related ion channels, receptor families, etc.).
Counter-, orthogonal and selectivity screens are thus used to stratify the hit list of the putative actives on the target of interest. The selection of the main screening assay and the setup of suitable filter-experiments coupled with optimal data analysis approaches to extract the cleanest and most complete information possible are important ingredients for the success of HTS-based hit discovery projects. Secondary follow-up assays for further confirmation and quantification of the desired modulation of the target and possibly determination of mechanisms of action will need similar care in processing of the underlying plate-based data and best possible characterization of concentration-dependent responses. Hit compounds or compound series possessing suitable pharmacological or biological and physicochemical properties and characterized in such a complete fashion, including a final structural verification, can then be considered possible starting points for further chemical optimization, i.e. become a ‘lead compound’ or a ‘lead series’ in a given drug discovery project (Hughes et al. 2011).
High-Throughput Screening methods and tools in their diverse forms (using biochemical, cell- or gene-based assay systems) with small molecular compounds, siRNA, shRNA, antibodies, antibody drug conjugates, or other probes to modulate the intended target or biological process of interest using a wide variety of assay and readout technologies have thus become an essential research tool for drug discovery, i.e. screening for hits and leads, functional genomics (target discovery), biomarker detection and identification in proteomics using mass spectrometry readouts , automated large scale characterization of samples (e.g. for sample quality assessment), as well as the detailed characterization of hit series with biophysical measurements, often using label-free assay technologies, and ADMET (absorption, distribution, metabolism, excretion, toxicity) screens (Macarron et al. 2011). In all cases the choice of the most suitable statistical methods for the different data analysis steps forms an important part of the usefulness and success of this toolset.
1.2 HTS Campaign Phases
Compound Screening
Typically several different rounds of screens are run for each project in compound based drug discovery. An initial primary screen is applied to assess the activity of a collection of compounds or other samples and to identify hits against a biological target of interest, usually employing single measurements (n = 1) due to the sheer number of samples which need to be processed. The primary screen identifies actives from a large diverse library of chemical probes, or alternatively, from a more focused library depending on pre-existing knowledge on a particular target. Selected hits and putative actives are then processed in second stage confirmation screen, either employing replicates at a particular single concentration, or a concentration response curve with just a few concentration points. In Table 5.1 we show the typical experimental characteristics (numbers of replicates, numbers of concentrations) of various screening phases for a large compound based HTS campaign in the author’s organization as an illustration for the overall experimental effort needed and the related data volumes to be expected. The numbers are also representative for other larger screening organizations. The addition of counter- or selectivity measurements will of course lead to a correspondingly higher effort in a particular phase.
If primary hit rates are very high and cannot be reduced by other means then counter-screens may need to be run in parallel to the primary screen (i.e. one has to run two large screens in parallel!) in order to reduce the number of candidates in the follow phase to a reasonable level and to be able to more quickly focus on the more promising hit compounds. If primary hit rates are manageable in number then such filter experiments (whether counter- or selectivity-screens) can also be run in the confirmation phase. Selectivity measurements are often delayed to the validation screening phase in order to be able to compare sufficient details of the concentration-response characteristics of the compounds which were progressed to this stage on the different targets, and not just have to rely on single-point or only very restricted concentration dependent measurements in the previous phases. Thus, the actual details of the makeup of the screening stages and the progression criteria are highly project dependent and need to be adapted to the specific requirements.
A validation screen, i.e. a detailed measurement of full concentration-response curves with replicate data points and adequately extended concentration range, is then finally done for the confirmed actives, as mentioned, possibly in parallel to selectivity measurements.
The successful identification of interesting small-molecule compounds or other type of samples exhibiting genuine activity on the biological target of interest is dependent on selecting suitable assay setups, but often also on adequate ‘filter’ technologies and measurements, and definitely, as we will see, also on employing the most appropriate statistical methods for data analysis. These also often need to be adapted to the characteristics of the types of experiments and types of responses observed. All these aspects together play an important role for the success of a given HTS project (Sittampalam et al. 2004).
Besides the large full deck screening methods to explore the influence of the of the particular accessible chemical space on the target of interest in a largely unbiased fashion several other more ‘knowledge based’ approaches are also used in a project specific manner: (a) focused screening with smaller sample sets known to be active on particular target classes (Sun et al. 2010), (b) using sets of drug-like structures or structures based on pharmacophore matching and in-silico docking experiments when structural information on the target of interest is available (Davies et al. 2006), (c) iterative screening approaches which use repeated cycles of subset screening and predictive modeling to classify the available remaining sample set into ‘likely active’ and ‘likely inactive’ subsets and including compounds predicted to likely show activity in the next round of screening (Sun et al. 2010), and (d) fragment based screening employing small chemical fragments which may bind weakly to a biological target and then iteratively combining such fragments into larger molecules with potentially higher binding affinities (Murray and Rees 2008).
In some instances the HTS campaign is not executed in a purely sequential fashion where the hit analysis and the confirmation screen is only done after the complete primary run has been finished, but several alternate processes are possible and also being used in practice: Depending on established practices in a given organization or depending on preferences of a particular drug discovery project group one or more intermediate hit analysis steps can be made before the primary screening step has been completed. The two main reasons for such a partitioned campaign are: (a) To get early insight into the compound classes of the hit population and possible partial refocusing of the primary screening runs by sequentially modifying the screening sample set or the sequence of screened plates, a process which is brought to an ‘extreme’ in the previously mentioned iterative screening approach, (b) to get a head start on the preparation of the screening plates for the confirmation run so that it can start immediately after finishing primary screening, without any ‘loss’ of time. There is an obvious advantage in keeping a given configuration of the automated screening equipment, including specifically programmed execution sequences of the robotics and liquid handling equipment, reader configurations and assay reagent reservoirs completely intact for an immediately following screening step. The careful analysis of the primary screening hit candidates, including possible predictive modeling, structure-based chemo-informatics and false-discovery rate analyses can take several days, also depending on assay quality and specificity. The preparation of the newly assembled compound plates for the confirmation screen will also take several days or weeks, depending on the existing compound handling processes, equipment, degree of automation and overall priorities. Thus, interleaving of the mentioned hit finding investigations with the continued actual physical primary screening will allow the completion of a screen in a shorter elapsed time.
RNAi Screening
RNAi ‘gene silencing’ screens employing small ribonucleic acid (RNA) molecules which can interfere with the messenger RNA (mRNA) production in cells and the subsequent expression of gene products and modulation of cellular signaling come in different possible forms. The related experiments and progression steps are varying accordingly. Because of such differences in experimental design aspects and corresponding screening setups also the related data analysis stages and some of the statistical methods will then differ somewhat from the main methods typically used for (plate-based) single-compound small molecule screens. The latter are described in much more depth in the following sections on statistical analysis methods.
Various types of samples are used in RNAi (siRNA, shRNA) screening: (a) non-pooled standard siRNA duplexes and siRNAs with synthetically modified structures, (b) low-complexity pools (3–6 siRNAs with non-overlapping sequences) targeting the same gene, (c) larger pools of structurally similar silencing molecules, for which measurements of the loss of function is assessed through a variety of possible phenotypic readouts made in plate array format in essentially the same way as for small molecule compounds, and also—very different—(d) using large scale pools of shRNA molecules where an entire library is delivered to a single population of cells and identification of the ‘interesting’ molecules is based on a selection process.
RNAi molecule structure (sequence) is essentially known when using approaches (a) to (c) and, thus, the coding sequence of the gene target (or targets) responsible for a certain phenotypic response change can be inferred in a relatively straightforward way (Echeverri and Perrimon 2006; Sharma and Rao 2009). RNAi hit finding progresses then in similar way as for compound based screening: An initial primary screening run with singles or pools of related RNAi molecule samples, followed by a confirmation screen using (at least 3) replicates of single or pooled samples, and if possible also the individual single constituents of the initial pools. In both stages of screening statistical scoring models to rank distinct RNAi molecules targeting the same gene can be employed when using pools and replicates of related RNAi samples (redundant siRNA activity analysis, RSA, König et al. 2007), thus minimizing the influence of strong off-target activities on the hit-selection process. In any case candidate genes which cannot be confirmed with more than one distinct silencing molecule will be eliminated from further investigation (false positives). Birmingham et al. (2009) give an overview over statistical methods used for RNAi screening analysis. A final set of validation experiments will be based on assays measuring the sought phenotypic effect and other secondary assays to verify the effect on the biological process of interest on one hand, and a direct parallel observation of the target gene silencing by DNA quantitation, e.g. by using PCR (polymerase chain reaction) amplification techniques). siRNA samples and libraries are available for individual or small sets of target genes and can also be produced for large ‘genome scale’ screens targeting typically between 20,000 and 30,000 genes.
When using a large-scale pooled screening approach, as in d) above, the selection of the molecules of interest and identification of the related RNA sequences progresses in a very different fashion: In some instances the first step necessary is to sort and collect the cells which exhibit the relevant phenotypic effect (e.g. the changed expression of a protein). This can for example be done by fluorescence activated cell sorting (FACS), a well-established flow-cytometric technology. DNA is extracted from the collected cells and enrichment or depletion of shRNA related sequences is quantified by PCR amplification and microarray analysis (Ngo et al. 2006). Identification of genes which are associated with changed expression levels using ‘barcoded’ sets of shRNAs which are cultured in cells both under neutral reference conditions and test conditions where an effect-inducing agent, e.g. a known pathway activating molecule is added (Brummelkamp et al. 2006) can also be done through PCR quantification, or alternatively through massive parallel sequencing (Sims et al. 2011) and comparison of the results of samples cultured under different conditions. These are completely ‘non-plate based’ screening methods and will not be further detailed here.
2 Statistical Methods in HTS Data Analysis
2.1 General Aspects
In the next sections we will look at the typical process steps of preparing and running a complete HTS campaign, and we will see that statistical considerations play and important role in essentially all of them because optimal use of resources and optimal use of information from the experimental data are key for a successful overall process. In the subsequent sections of this chapter we will not touch on all of these statistical analysis aspects with the same depth and breadth, not least because some of these topics are actually treated in more detail elsewhere in this book, or because they are not very specific to HTS data analysis. We will instead concentrate more on the topics which are more directly related to plate-based high-throughput bioassay experiments and related efficient large scale screening data analysis.
2.2 Basic Bioassay Design and Validation
Given a particular relevant biological target the scientists will need to select the assay method, detection technology and the biochemical parameters. This is sometimes done independently from HTS groups, or even outside the particular organization. Primarily the sensitivity (ability to detect and accurately distinguish and rank order the potency of active samples over a wide range of potencies, e.g. based on measurements with available reference compounds) and reproducibility questions need to be investigated in this stage. Aspects of specificity in terms of the potential of a particular assay design and readout technology to produce signals from compounds acting through unwanted mechanisms as compared to the pharmacologically relevant mechanism also need to be assessed.
The exploration and optimization of assay conditions is usually done in various iterations with different series of experiments employing (fractional) factorial design, replication and response surface optimization methods to determine robust response regions. The potentially large series of assay parameters (selection of reagents, concentrations, pH, times of addition, reaction time courses, incubation times, cell numbers, etc.) need to be investigated at multiple levels in order to be able to detect possible nonlinearities in the responses (Macarron and Hertzberg 2011). The practical experimentation involving the exploration of many experimental factors and including replication often already takes advantage of the existing screening automation and laboratory robotics setups to control the various liquid handling steps, which otherwise would be rather cumbersome and error-prone when performed manually (Taylor et al. 2000). Standard methods for the analysis of designed experiments are used to optimize dynamic range and stability of the assay readout while maintaining sensitivity (Box et al. 1978; Dean and Lewis 2006). Most often the signal-to-noise ratio SNR, signal window SW and Z′-factor are used as optimization criteria. See the section on assay quality measures and Table 5.2 below for the definition of these quantities.
2.3 Assay Adaptation to HTS Requirements and Pilot Screening
The adaptation of assay parameters and plate designs with respect to the positioning and numbers of control samples, types of plates, plate densities and thus, liquid volumes in the individual plate wells, to fulfill possible constraints of the automated HTS robotics systems needs to follow, if this was not considered in the initial assay design, e.g. when the original bioassay was designed independently of the consideration to run it as a HTS. The initially selected measurement technology often has an influence on the obtainable screening throughput. If these design steps were not done in initial collaboration with the HTS groups, then some of the assays parameters may have to be further changed and optimized with respect to relevant assay readout quality measures (see Table 5.2) for the assay to be able to run under screening conditions. Again, experimental design techniques will be employed, albeit with a now much reduced set of factors. These assay quality metrics are all based on the average response levels \( \overline{x} \) of different types of controls, corresponding to readout for an inactive probe (N, neutral control) or the readout for a ‘fully active’ probe (P, positive control) and the respective estimates of their variability (standard deviation) s.
The quality control estimators in Table 5.2 are shown as being based on the mean \( \overline{x} \) and standard deviation s, but in practice the corresponding outlier resistant ‘plug-in’ equivalents using the median \( {\tilde {x}} \) and median absolute deviation (mad) \( {\tilde {s}} \) estimators are often used, where \( {\tilde {s}} \) includes the factor 1.4826 to ensure consistency with s, so that \( E\left({\tilde {s}}(x)\right)=E\left(s(x)\right)=\sigma \) if \( x\sim N(\mu, {\sigma}^2 \)) (Rousseeuw and Croux 1993).
The quality measures which incorporate information on variability of the controls often much more useful than simple ratios of average values because probability based decision making in the later hit selection process needs to take into account the distributions of the activity values. In order to gain a quick overview and also see aspects of the distribution of the measured values (e.g. presence of outliers, skewness) which are not represented and detected by the simple statistical summary values the data should always be visualized using e.g. categorized scatterplots, boxplots, strip plots and normal quantile-quantile plots.
Possible modifications of the original assay design may entail gaining higher stability of response values over longer time scales or in smaller volumes to be able to run larger batches of plates, shortening some biochemical reaction or incubation phases to gain higher throughput, or to have smaller influence of temperature variations on responses, etc. Such modifications may sometimes even have to be done at the cost of reduced readout response levels. As long as the assay quality as measured by a suitable metric stays above the acceptance criteria defined by the project group such ‘compromises’ can usually be done without large consequences for the scientific objectives of the HTS project.
Besides the optimization of the assay signal range and signal stability an important part of the quality determination and validation in the assay adaptation phase is the initial determination of assay reproducibility for samples with varying degrees of activity using correlation- and analysis of agreement measures, as well providing rough estimates on expected false positive and false negative rates which are based on the evaluation of single point %-activity data at the planned screening concentration and the determination of the corresponding concentration-response data as an activity reference for the complete standard ‘pilot screening library’ of diverse compounds with known mechanisms of action and a wide range of biological and physicochemical properties for use in all HTS assays in the pilot screening phase (Coma et al. 2009a, b). See Table 5.3 for an overview of typical analyses performed in this phase.
These investigations with the standard pilot screening library will sometimes also allow one to gain early information on possible selectivity (to what extent are compounds acting at the target vs. other targets of the same family) and specificity (is the compound acting through the expected mechanism or through an unwanted one) of the assay for an already broader class of samples available in the pilot screening library than are usually investigated in the initial bioassay design.
While the detailed evaluation of factors and response optimization using experimental design approaches with replicated data are feasible (and necessary) at the assay development and adaptation stages, this is no longer possible in the actual large primary HTS runs. They will usually need to be executed with n = 1 simply because of the large scale of these experiments with respect to reagent consumption, overall cost and time considerations. Because of this bulk- and batch execution nature of the set of measurements collected over many days or weeks it is not possible to control all the factors affecting the assay response. Given these practical experimental (and resource) restrictions, the observed random and systematic variations of the measured responses need to be accommodated and accounted for in the data analysis steps in order to extract the highest quality activity- and activity-rank order information possible. The use of optimal data normalization and hit selection approaches are key in this respect.
2.4 Assay Readouts, Raw and Initial Derived Values
Raw readouts from plate readers are generated in some instrument-specific units (often on an ‘arbitrary’ scale) which can be directly used for data quality and activity assessment, but in other assay setups there may be a need for an initial data transformation step. This can be needed when performing time- or temperature-dependent measurements which need a regression analysis step to deliver derived readouts in meaningful and interpretable physical units (assay readout endpoints), e.g. using inverse estimation on data of a calibration curve, determining kinetic parameters of a time-course measurement or protein melting transition temperatures in a thermal shift assay, or extracting the key time course signal characteristics in a kinetic fluorescence intensity measurement. Sometimes such derived information can be obtained directly from the instrument software as an alternate or additional ‘readout’, but when developing new assays and measurement methods it is sometimes necessary to be able to add different, more sophisticated or more robust types of analyses as ad-hoc data preprocessing steps. It is a definite advantage if a standard software framework is available where such additional analysis steps and related methods can be easily explored, developed and readily plugged into the automated assay data processing path, e.g. by using R scripts (R Development Core Team 2013) in the Pipeline Pilot® (http://accelrys.com/products/pipeline-pilot/), Knime (https://www.knime.org/) or similar analytics platform included upstream of an organization’s standard screening data processing system, where data thus transformed can then easily be processed using the available standard HTS data analysis methods in the same way as any other type of screening reader output.
Assay and readout technologies have gone through many changes and advancements in the past years. Whereas initially in HTS the measurement of only one or very few (2–3) readout parameters per well (e.g. fluorescence intensities at two different wavelengths) was customary—and still is for many practical applications—the advent of automated microscopy and cellular imaging coupled with automated image analysis (image based High Content Analysis or Screening, HCA, HCS) which can detect changes in the morphology of cells or of separately labeled cell compartments (nucleus, membrane, organelles, etc.), thus resulting in a large number of parameters for a given well or even for each individual cell, has led to the need for the exploration and evaluation of suitable multivariate statistical data analysis methods (Hill et al. 2007). Intensities, textures, morphological and other parameters from the segmented images are captured at several different wavelengths and corresponding feature vectors are associated with each identified object or well (Abraham et al. 2004; Carpenter 2007; Duerr et al. 2007; Nichols 2007). Cell level analysis enables the analysis of the various cell-cycles and the separation of the effects of the probes on cells in a particular state (Loo et al. 2007; Singh et al. 2014). Besides the now quite broadly used image based HCS approaches there are several other assay technologies which produce multivariate readouts of high dimensions, Cytof (Qiu et al. 2011), Luminex gene expression profiling (Wunderlich et al. 2011), RPA (van Oostrum et al. 2009), laser cytometry (Perlman et al. 2004 ), and others, with medium throughput . For most of these technologies the most suitable optimal data analysis methods are still being explored. Questions of normalization, correction of systematic errors, discrimination and classification are under active investigation in many labs (Reisen et al. 2013; Kümmel et al. 2012; Abraham et al. 2014; Singh et al. 2014; Smith and Horvath 2014; Haney 2014). It is clear that all these different types of assay technologies can benefit from a common informatics infrastructure for large scale multivariate data analysis, which includes a large set of dimension reduction, feature selection, clustering, classification and other statistical data analysis methods, as well as a standardized informatics systems for data storage and metadata handling, coupled to high performance computing resources (compute clusters) and large volume file stores and databases (Millard et al. 2011).
The high numbers of readout parameters (300–600) (Yin et al. 2008; Reisen et al. 2013) which must be simultaneously analyzed and the much higher data volumes which need to be processed introduce new aspects into high-throughput screening data analysis which are usually not covered by the available features in the established standard screening informatics systems (Heyse 2002; Gunter et al. 2003; Kevorkov and Makarenkov 2005; Gubler 2006; Boutros et al. 2006; Zhang and Zhang 2013). This makes these data much more challenging to analyze from the point of view of methodology, complexity of assay signals and the sheer amounts of data. But it is clear that these types of screening technologies and efficient methods to analyze the large data volumes will become even more important and widespread in future. While one can say that the analysis methods for standard HTS data have been largely settled—at least from the point of view of the main recommended data processing and quality assurance steps as outlined in this chapter—this is definitely not yet the case for the high dimensional multivariate screening data analysis, especially when going to the single cell level. Note that the screening literature occasionally refers to multi-parametric analysis in this context. Systematic investigations on advantages and disadvantages of particular methods and the preferred approaches for determining assay and screening quality metrics, correction of systematic response errors, classification of actives, etc. with such types of data are ongoing and are naturally more complex than for the cases where just a few readout parameters can be processed in a largely independent manner up to the point where the final values need to be correlated to each other (Kümmel et al. 2012).
2.5 Assay Quality Measures
The overall error which accumulates over the many different chemical, biological and instrumental processing steps to obtain the final readout in a screening assay needs to be kept as small as possible so that there is high confidence in the set of compounds identified as active in a screening campaign. The assay quality metrics to measure and monitor this error are based on simple location and scale estimates derived from raw readout data from the different types of wells on a microtiter plate (zero-effect and full inhibition of full activation controls for normalization of the data, reference controls exhibiting responses in the middle of the expected response range, background wells, and test sample wells). Different quality indicators have been proposed to measure the degree of separability between positive and zero-effect (neutral) assay controls: Signal to background ratio or high-low ratio, coefficient of variation, signal to noise ratio, Z- and Z′-factor (not to be confused with a Z-score) (Zhang et al. 1999), strictly standardized mean difference (SSMD) (Zhang et al. 2007) and others are in routine use to optimize and measure assay response quality (see Table 5.2).
The Z′-factor has become an accepted and widely used quality metric to assess the discriminatory power of a screening assay. It is a relative measure and quantifies the ‘usable window’ for responses between the upper and lower controls outside of their respective 3s limits. Z′ can be between \( -\infty \) (if the control averages which define the response limits are identical), 0 when the two 3s limits ‘touch’ each other, and 1 if the standard deviation of the controls becomes vanishingly small. Z′ is an empirical point measure and the derivation of its large sample interval estimator was only recently published (Majumdar and Stock 2011). The sampling uncertainty of Z′ should be considered when setting acceptance thresholds, especially for lower density plates with small numbers of control wells. Small sample intervals can be estimated by bootstrap resampling (Iversen et al. 2006). Other quality indicators than those listed were proposed and described in the literature (e.g. assay variability ratio, signal window and others), but are not so widely used in standard practice because they are related to the Z′-factor and don’t represent independent information (Sui and Wu 2007; Iversen et al. 2006). The V-factor is a generalization of the Z′-factor to multiple response values between \( {\overline{x}}_N \) and \( {\overline{x}}_P \) (Ravkin 2004).
Some screening quality problems can occur for actual sample wells which are not captured by control well data and the measures listed in Table 5.2, e.g. higher variability for sample wells than for control wells, additional liquid handling errors due to additional process steps for sample pipetting, non-uniform responses across the plates, etc. Such effects and tools for their diagnosis are described in more detail further below in the screening data quality and process monitoring section.
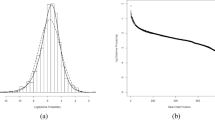
In Fig. 5.1 we show an example of the behavior of the High/Low control ratio (HLR) and the Z′ factor for a set of 692 1536-well plates from a biochemical screen exhibiting several peculiarities: (a) clear batch boundary effects in the ratio of the HLR values for batch sizes varying between 80 and 200 plates, (b) ‘smooth’ time dependence of the HLR (fluorescence intensity) ratio due to the use of continuous assay product formation reaction and related detection method, (c) no ‘strongly visible’ influence of the varying HLR on the Z′-factor, i.e. a negligible influence of the varying HLR on the relative ‘assay window’, (d) an interleaved staggering pattern of HLR which is due to the use of a robotic system with two separate processing lanes with different liquid handling and reader instruments. This latter aspect may be important to take into account when analyzing the data because any systematic response errors, if they occur at a detectable and significant level, are likely to be different between the two subsets of plates, hence a partially separate analysis may need to be envisaged. We also see that for assays of this nature setting a tight range limit on HLR will not make sense; only a lower threshold could be useful as a potential measurement failure criterion.
(a) Screening quality control metrics: High/Low ratio HLR for complete screening run showing batch start and end effects, signal changes over time and alternating robot lane staggering effects (inset). (b) Z′-factor (assay window coefficient) for the same plate set with occasional low values for this metric, indicating larger variance of control values for individual plates, but generally negligible influence of robot lane alternation on assay data quality (inset, saw tooth pattern barely visible)
2.6 Screening Data Quality and Process Monitoring
Automated screening is executed in largely unattended mode and suitable procedures to ensure that relevant quality measures are staying within adequate acceptance limits need to be set up. Some aspects of statistical process control (SPC) methodology (Shewhart 1931; Oakland 2002) can directly be transferred to HTS as an ‘industrial’ data production process (Coma et al. 2009b; Shun et al. 2011).
Data quality monitoring of larger screens using suitably selected assay quality measures mentioned above and preferably also for some of the additional screening quality measures listed below in Table 5.4, can be done online with special software tools which analyze the data in an automated way directly after the readout is available from the detection instruments (Coma et al. 2009b), or at least very soon after completing the running of a plate batch with the standard data analysis tools which are in use, so that losses, potential waste and the need for unnecessarily repetitions of larger sets of experiments are minimized. As in every large scale process the potential material and time-losses and the related financial aspects cannot be neglected and both plate level and overall batch level quality must be maintained to ensure a meaningful completion of a screening campaign.
Systematic response errors in the data due to uncontrolled (and uncontrollable) factors will most likely also affect some of the general and easily calculated quality measures shown in Table 5.4, and they can thus be indirect indicators of potential problems in the screening or robotic automation setup. When using the standard HTS data analysis software tools to signal the presence of systematic response errors or a general degradation of the readout quality, then the broad series of available diagnostic tools can efficiently flag and ‘annotate’ any such plate for further detailed inspection and assessment by the scientists. This step can also be used to automatically categorize them for inclusion or exclusion in a subsequent response correction step.
A note on indexing in mathematical expressions: In order to simplify notation as far as possible and to avoid overloading the quoted mathematical expressions we will use capital index letters to indicate a particular subset of measured values (e.g. P and N as previously mentioned, C compound samples), and we will use single array indexing of measured or calculated values of particular wells i whenever possible. In those situations where the exact row and column location of the well in the two-dimensional grid is important we will use double array indexing ij. The explicit identification of the subset of values on plate p is almost always required, so this index will appear often.
These additional metrics are relying on data from the sample areas of the plates and will naturally provide additional important insight into the screening performance as compared to the control sample based metrics listed in Table 5.2. As for the previously mentioned control sample based assay quality metrics it is even more important to visualize such additional key screening data quality metrics which are based on the (compound) sample wells in order to get a quick detailed overview on the behavior of the response data on single plates, as well as its variation over time to obtain indications of data quality deteriorations due to instrumental (e.g. liquid handling failures), environmental (e.g. evaporation effects, temperature variations) and biochemical (e.g. reagent aging) factors, or due to experimental batch effects, e.g. when using different reagent or cell batches. Direct displays of the plate data and visualizations of the various assay quality summaries as a function of measurement time or sequence will immediately reveal potentially problematic data and suitable threshold settings can trigger automatic alerts when quality control metrics are calculated online.
Some of the listed screening quality metrics are based on direct estimation of systematic plate and well-location specific experimental response errors S ijp , or are indicators for the presence of spatial autocorrelation due to localized ‘background response’ distortions, e.g. Moran’s I coefficient which also allows the derivation of an associated p-value for the ‘no autocorrelation’ null hypothesis (Moran 1950). Similar visualizations as for the HLR and Z′-factor shown in Fig. 5.1 can also be generated for the listed screening quality metrics, like the Z-factor (screening window), Moran coefficient I, or the VEP measure.
In Fig. 5.2 we show examples of useful data displays for response visualizations of individual plates. In this case both the heatmap and the separate platewise scatterplot of all data, or boxplots of summary row- and column effects of a 384-well plate clearly show previously mentioned systematic deviations of the normalized response values which will need to be analyzed further. In the section on correction of systematic errors further below we also show an illustration of the behavior of the Moran coefficient in presence of systematic response errors, and after their removal (Fig. 5.6).
(a) Example plate data heatmap showing visibly lower signal in border areas due to evaporation effects. (b) Scatterplot of normalized response values of all individual wells with grid lines separating different plate rows. (c) Boxplot of column summaries, and (d) Boxplot of row summaries, all showing location dependent response differences across the plate and offsets of row- and column medians from 0
It is also recommended to regularly intersperse the screening plate batches with sets of quality control plates without compounds (providing a control of liquid handling performance) and plates containing only inhibitor or activator controls and, if relevant, further reference compound samples exhibiting intermediate activity to provide additional data on assay sensitivity. ‘QC plates’ without compounds or other types of samples (just containing assay reagents and solvents) are also very helpful for use in correcting the responses on the compound plates in cases where it is not possible to reliably estimate the systematics errors from an individual plate or a set of sample plates themselves. This is e.g. the case for plates with large numbers of active samples and very high ‘hit rates’ where it is not possible to reliably determine the effective ‘null’ or ‘background’ response, as well as for other cases where a large number of wells on a plate will exhibit nonzero activity, especially also all concentration- response experiments with the previously selected active samples from the primary screens. For these situations we have two possibilities to detect and correct response values: (a) Using the mentioned QC control plates (Murie et al. 2014), and (b) using plate designs with ‘uniform’ or ‘uniform random’ placement of neutral control wells across the whole plate, instead of placing the controls in a particular column or row close to the plate edges (Zhang 2008), as is often done in practice. Such an arrangement of control well locations will allow the estimation of systematic spatial response deviations with the methods which only rely on a small number of free parameters to capture the main characteristics and magnitude of the background response errors, as e.g. polynomial response models.
2.7 Integration of Diagnostic Information from Automated Equipment
Modern liquid handlers (e.g. acoustic dispensers) often have the capability to deliver success/failure information on the liquid transfer for each processed well and this information can be automatically joined with the reader data through plate barcode and well matching procedures, and then considered for automated valid/invalid flagging of the result data, provided suitable data management processes and systems are available. Also other types of equipment may be equally able to signal failures or performance deterioration at the plate or well level which then can also be correlated with reader data and be integrated in suitable alerting and flagging mechanisms for consideration during data analysis.
2.8 Plate Data Normalization
Uncontrolled experimental factors will influence the raw assay readouts on each individual plate. This can be likened to the existence of multiplicative and possibly also additive systematic errors between individual plates p which can be represented by the measurement model \( {x}_{ip}={\lambda}_p\left(m+{\kappa}_i\right)+{b}_p \), where x ip is the measured raw value in well i on plate p , m is the ‘intrinsic’ null-effect response value of the assay, κ i is a sample (compound) effect, with κ i = 0 if inactive, and λ p and b p are plate-, instrument- and concentration dependent gain factors and offsets, respectively. x ip can then also equivalently be represented as
when also including an error term \( {\epsilon}_{ip}\sim N\left(0,{\sigma}_p^2\right) \) and setting \( {m}_p = {\lambda}_pm + {b}_p \), \( {\gamma}_{ip}={\lambda}_p{\kappa}_i \). We make \( E\left({\epsilon}_{ip}\right)={\sigma}_p \) explicitly depend on plate index p because this often corresponds to experimental reality, especially between different plate batches. Reagent aging and evaporation as well as time shifts in some process steps will usually lead to smooth trends in plate averages within batches, whereas e.g. the effect of cells plated at very different times may show up as a more discrete batch effect of responses. Plate-level normalization procedures use the response values of specific wells to bring the response values into a standardized numerical range which can be easily interpreted by scientists, usually a 0–100 % scale with respect to the no-effect and the ‘maximal’ effect obtainable with the given assay parameters. In this particular context ‘plate normalization’ is simply understood to adjust the average responses between the different plates: Control based normalization via specifically selected control wells, sample based normalization via the ensemble of all sample wells if the number of active wells is ‘small enough’. The use of robust estimation procedures with high breakdown limits is critical for successful sample based normalization because of the likely presence of ‘outliers’, i.e. active compounds with larger response deviations in usually one and sometimes both possible response directions.
Normalization of Compound Screening Data
In the experiment- and plate design for small molecule compound screening one usually has two types of controls to calibrate and normalize the readout signals to a 0–100 % effect scale: A set of wells (neutral or negative controls) corresponding to the zero effect level (i.e. no compound effect) and a set of wells (positive controls) corresponding to the assay’s signal level for the maximal inhibitory effect, or for activation- or agonist assays, corresponding to the signal level of an effect-inducing reference compound. In the latter case the normalized %-scale has to be understood to be relative to the chosen reference which will vary between different reference compounds, if multiple are known and available, and hence has a ‘less absolute’ meaning compared to inhibition assays whose response scale is naturally bounded by the signal level corresponding to the complete absence of the measured biological effect. In some cases an effect-inducing reference agonist agent may not be known and then the normalization has to be done with respect to neutral controls only, i.e. using Fold- or any of the Z- or R-score normalization variants shown in Table 5.5.
Normalization of RNAi Screening Data
In RNAi screens which use functional readouts any gene silencing event can in principle lead to an inhibition or to an enhancement of the observed phenotypic readout (e.g. a simple cell viability measurement). It is thus advisable to use several different types of controls to be able to assess screen quality and calibrate the response scale. In any case, siRNA controls which have no specific silencing effect on the target gene(s) need to be designed and used as negative controls, and siRNAs which target some genes having a previously known association with the biological process under study and leading to a modulation of the desired readout can be used as positive controls. In a particular screen it is desirable to employ positive controls of different strengths (e.g. weak inhibition, strong inhibition) to compare the often strongly varying observed siRNA effects to the effect sizes of known influencers of a complex biological pathway, and also to use controls exhibiting different effect directions (inhibitors, enhancers) to be able to assess the reliability of the assay response scale in either direction. Besides the natural use of the positive controls as screening QC indicators to assess screen stability, derive assay window quality Z′-factors, etc. the effect sizes of the positive controls need to be considered as almost ‘arbitrary’ reference response levels allowing to classify individual siRNA responses as ‘weak’ to ‘very strong’, which is similar to the use of different types of agonist assay reference wells in compound screening providing different relative normalization scales as described in the last paragraph. Plate-based RNAi screening data are usually normalized to the Fold scale based on the neutral controls (see Table 5.5) and actual relative effect ‘sizes’ of individual single siRNA samples in either direction are either assessed by using R-scores, and in the case of replicates by using a t-statistic, or preferably the SSMD-statistic for unequal sample sizes and unequal variances between siRNA and neutral controls (Zhang 2011a, b). This type of SSMD- based normalization, and in a similar way the consideration of magnitudes of R-score values, already touches on the hit selection process as described further below.
Table 5.5 lists the most frequently used data normalization expressions and it can easily be seen that the values from the various expressions are centered at 0, 1, or 100 and linearly related to each other. The normalized values are proportional to the biological modulation effect κ i , provided the assay and readout system response is linear as assumed in Eq. (5.1), and when disregarding ϵ ip and systematic response errors. For nonlinear (but monotone) system responses the rank orders of the normalized activity values are maintained. With the presence of experimental random and systematic measurement errors this strict proportionality and/or concordance in rank ordering of the sample activities κ i within and across plates is of course no longer guaranteed. Especially in cell-based assay system the assay response variability can be relatively large and also the relative rank ordering of sample activities from primary screening and the follow-up IC 50 potency determinations can differ considerably, mostly due to concentration errors in the different experiments (Gubler et al. 2013).
Besides these average plate effects (‘gain factor differences’) other response differences at the well, row or column level occur frequently in actual experimental practice. They often occur in a similar fashion on multiple plates within a batch and it is important to take these location- and batch dependent errors into account to be able to identify hits in an efficient way. We deal with the detection and correction of these effects separately in the next section, but of course they belong to the same general topic of assay response normalization.
If we observe plate-to-plate or batch-to-batch variations of σ p then the use of Z- orR-score-based normalization is advised to allow the application of a meaningful experimentwise hit selection threshold for the entire screen. If systematic response errors are detected on the plates then the final scoring for hit selection needs to be done with the corrected activity values, otherwise the hit list will be clearly biased. The scoring expressions are based on estimated center \( {\tilde {x}} \) and scale \( {\tilde {s}} \) values and correspond to a t-statistic, but the actual null-distribution of the inactive samples in a primary screen will usually contain a very large number of values, so that we can assume \( {f}_0\sim N\left(0,1\right) \) for the R-scores of this null-effect subset after correction. We will revisit this aspect in the section on hit-selection strategies.
2.9 Well Data Normalization, Detection and Correction of Systematic Errors in Activity Data
In this section we deal with the detection, estimation and correction of and assay response artifacts. The series of different process steps in assay execution and the different types of equipment and environmental factors in a typical plate based screen often lead to assay response differences across the plate surfaces (i.e. non-random location effects), both as smooth trends (e.g. due to time drifts for measurements of different wells in a plate), bowl shapes (often due to environmental factors like evaporation, temperature gradients), or step-like discrete striping patterns (most often due to liquid handling equipment imperfections (dispensing head, needle or pin) leading to consistently lower or higher than average readings, and combinations of some or all of these types of effects. Also gradients in the step-like artifacts can sometimes be observed due to the time ordering of dispensing steps. Often, but not always, these effects are rather similar on a whole series of plates within a measurement batch which obviously will help in estimation and subsequent correction procedures. Individual well patterns can obviously only be estimated if they repeat on all or a subset (batch) of plates, otherwise they are confounded with the effect of the individual compound samples. Automatic partitioning of the sets of readouts to reflect common patterns and identification of those respective different systematic patterns is an important aspect for efficient and effective response correction steps. Temporal changes of non-random response patterns are related to batch-wise assay execution, reagent aging effects, detection sensitivity changes or changes in environmental factors and may appear gradual or sudden.
It is obvious that systematic spatial or temporal response artifacts will introduce bias and negatively affect the effectiveness of hit finding especially for samples with weak and moderate size effects and will influence the respective false decision rates in hit selection. Such effects should thus be corrected before attempting hit selection or using these data for other calculation steps. Especially when considering fragment based screens with low-molecular samples of relatively low potency, natural product (extract) screens with low amounts of particular active ingredients, or RNA interference screens where small to moderate size effects can be of interest (corresponding to full knockdown of a gene with a small effect, or partial knockdown of a gene with strong effects), or if one simply is interested in detecting active samples in the whole range of statistically significant modulating effects then these response correction methods become crucial to allow optimized and meaningful analyses. The positive influence of the response correction on the hit confirmation rate, the reproducibility of the activity in follow-up screens or secondary assays can be clearly demonstrated (Wu et al. 2008).
An important prerequisite for successful estimation of response corrections using the actual screening sample data is the assumption that the majority of these samples are inactive and that active samples are randomly placed on the various plates. For screens with high rates of non-zero response wells it is advised to place neutral control wells (showing no effect on the assay readout) spread ‘uniformly’ across the plates, and not, as is often the case, in the first or last columns of the plates, and use them to check for the occurrence, estimation and correction of systematic response errors. Sometimes the plate layout designs which can be produced by compound management and logistics groups in an automated way are limited due to restrictions of the robotic liquid handling equipment, but in order to produce reliable screening results an optimal design of control well placement is important and liquid handling procedures should be adapted to be able to produce assay plates in such a way. Such specially designed plates with a suitable spatial spread and location of control wells can be used to derive ‘smooth’ (e.g. polynomial) average response models for each plate (or for a set of plates) which do not rely on the assumption that the majority of the test samples are inactive. For example in RNAi screening many of the samples or sample pools have usually some effect in the assay, leading to a large range of responses and related problems to efficiently use correction methods which rely on estimations of the null response based on the sample wells themselves. The response level of ‘truly inactive’ samples is difficult to determine in this case and consequently an efficient plate designs with well-chosen controls in the described sense or the use of interspersed control plates for error correction in a measurement batch can become important (Zhang 2008). Cell-based screens, including RNAi screens, often need additional and prolonged incubation periods which often exacerbate assay noise, response bias and artifacts in the border regions of the plates.
In actual practice it also happens that structurally and bioactivity-wise similar compounds are placed near each other because of the way the stored compound master plates were historically constructed (often from groups of similar compounds delivered to the central compound archives in a batch) even for ‘random’ HTS libraries, or simply due to the makeup of plates in focused libraries which exhibit larger rates of activity on selected target classes (e.g. enzyme inhibitors). Modern compound management setups today allow a more flexible creation of screening plates, but the presence of spatial clusters of activity or of known subsets of very likely active samples of on particular plates need to be considered for the decision to include or exclude selected plate subsets from the standard way of background response estimation and related processing.
Well-level assay response normalization and correction of the spatial and temporal patterns is in essence just a ‘more sophisticated’ form of the sample based normalization mentioned in the previous paragraph. Because of the expected presence of at least some ‘active’ wells (i.e. outliers for the purpose of background response estimation) it is highly advisable to use robust (outlier resistant) estimation methods when relying on the actual screening sample wells to derive the response models. The robustness breakdown limits for different methods are of course quite variable and need to be considered separately for each. The breakdown bounds for the median polish procedure were elaborated by Hoaglin et al. (1983).
As mentioned in the process monitoring section graphical displays are important to visually detect and often also quickly diagnose the potential sources of the error patterns. Also the visualizations of the error patterns and of the subsequently corrected data, including suitable graphics of the corresponding (now improved) quality metrics is an important practical element of screening quality assurance (Brideau et al. 2003; Coma et al. 2009b).
It is also important to note that data correction steps should only be applied if there is evidence for actual systematic errors, otherwise their application can result in variance bias, albeit with a magnitude strongly dependent on the actual correction method used. Such variance bias can have an influence on the hit identification step because the corresponding activity threshold choices can be affected in an unfavorable fashion. Suitably constructed quality metrics which are based e.g. on Moran’s I spatial autocorrelation coefficient (see Table 5.4 item d) can be used to determine whether a systematic error is present and whether corresponding response correction should be applied or not. ‘Suitably’ means that the weight matrix needs to minimally cover the neighborhood region which is expected to exhibit strong correlations when systematic errors of a particular type are present, e.g. by using a neighborhood range \( \delta =2 \) around grid point {i 0, j 0} for setting the weights \( {w}_{ij}=1 \) for all \( \left\{i,j:\left(0\le \left|i-{i}_0\right|\le\ \delta \right)\wedge \left(0\le \left|j-{j}_0\right|\le\ \delta \right)\right\} \) with \( {w}_{i_0{j}_0}=0 \) and \( {w}_{ij}=0 \) otherwise for the situations where discrete response ‘striping’ effects in every second row or column can occur due to some liquid handling errors besides the possible smoother spatial responds trends across the plate surface. Different δ extents in row and column directions or use of different weighting functions altogether may be more optimal for other types of expected patterns.
We have separated the assay response normalization into plate-level normalization as outlined in the previous section, including the calculation of various assay quality metrics according to Tables 5.1 and 5.2, and the possible subsequent row, col and well-level effect response adjustment procedures. In essence the latter can be considered as a location-dependent sample-based data normalization step. In Table 5.6 we show several such modeling and correction methods for location dependent systematic errors of well-level data in plate based screening which have been developed and described within the past 10–15 years (Heyse 2002; Heuer et al. 2002; Brideau et al. 2003; Kevorkov and Makarenkov 2005; Gubler 2006; Malo et al. 2006; Makarenkov et al. 2007; Birmingham et al. 2009; Bushway et al. 2010; Dragiev et al. 2011; Zhang 2011a; Mangat et al. 2014)
In Fig. 5.2 we have already seen data containing typical systematic response errors. As an illustration of model performance obtained with some of the approaches listed in Table 5.6 we show the same data in Fig. 5.3a with three different error model representations in Fig. 5.3b, c and the resulting corrected data, after applying the loess error model based correction in Fig. 5.3e. The corresponding row-wise boxplot of the corrected data in Fig. 5.3e can be compared to uncorrected case in Fig. 5.2d and the resulting smaller variances as well as better centering on zero are immediately evident.
(a) Same normalized plate data as in Fig. 5.2a. (b) Median polish representation of systematic error, which is inadequate in this case because a R*C interaction term is not included. (c) loess model representation of systematic error using 2nd degree polynomial local regression. (d) Robust mixed linear effects model representation of systematic error using rlmer function with R*C interaction terms. (e) Corrected data after loess error pattern subtraction. (f) Boxplot of row summaries corresponding to corrected data of Fig. 5.3e (compare with Fig. 5.2d, without correction)
For further illustration of various diagnostics and response modeling methods we will here use a simulated screening plate data of limited size (50 384-well plates) with normalized percent inhibition data scaled between 0 (null effect) and −100 (full inhibition) exhibiting several features which are typically found in real HTS data sets: Edge effects due to evaporation, response trends due to temperature gradients, liquid handling response stripes which vanish after a series of dispensing steps in interleaved row-wise dispensing, single dispensing needle malfunction, plate batch-effects with different dominant error patterns between batches, assay response noise \( \sim N\left(0,{\sigma}^2\right) \) with σ = 5 , and a response distribution of the randomly placed hits with an overall hit rate of 5 % which is approximated as ∼Gamma(1, 0.025), leading to a median inhibition of hits around −30 %, i.e. obtaining a smaller number of hits with strong inhibition and a larger number with moderate and small values which is the usual situation screening.
This simulated data set is represented as a heat map series in Fig. 5.4a and as the corresponding ‘assay map’ (Gubler 2006) in Fig. 5.4b. The assay map is a single heat map of the complete data set where the 2D row- and col- location is transformed into a linear well-index and data displayed as a contiguous (well-index, plate measurement sequence index) map. Response errors affecting particular wells or well-series in a set of sequentially processed plates are then immediately visible, as e.g. in our case a consistently lower response of well 233 in plates 1 to 40 due to a “defective” dispensing tip. Other batch effects are visible in both types of heatmaps.
(a) Heatmaps of simulated data set of 50 plates. Different types of systematic errors are visible, see text (b) Assay-map of the same data set, in this arrangement and ordering of the well-index allowing a quick identification of row- and individual well-effects (e.g. response error for well 233 in a subset of plates). (c) Plate-by-plate correlation matrix of well values for the complete data set, allowing the identification and grouping of plates with similar error patterns either visually or using hierarchical clustering
Our principal goal in performing HT screens is the identification of interesting compounds which perturb the response of the biological test system. As per Eq. (5.1) and extending the indexes to two dimensions i,j we can model the observed response x ijp in row i, column j and plate p as \( {x}_{ijp} = {\overline{x}}_p + {\gamma}_{ijp} + {\epsilon}_{ijp} \), allowing for a simple estimation of the relative compound effect size as
The compound effect γ ijp is of course fully confounded with ϵ ijp for the n = 1 primary screening case. Separate estimation of γ ijp and ϵ ijp can only be made when measuring replicates, as is often done in RNAi screens, or when running confirmation screens with replicates. As previously discussed, in actual screening practice we almost always have systematic response errors S ijp present in the experimental data, and thus now find correspondingly disturbed readout values
In Table 5.6 we have listed various methods which are used to determine the systematic error terms S and focus now on the influence of this term on the apparent compound effect γ ′ ijp as determined in the data normalization step. For a given measured data value x ijp Eqs. (5.1) and (5.3) give
for an otherwise unchanged null effect readout level \( {\overline{x}}_p \) and observed activity value x ijp . Without a suitable estimate for S ijp we have again complete confounding of compound effect, systematic error and random error components. By using information based on spatial and temporal correlation of the observed experimental response values from a series of wells an estimate \( {\hat{S}}_{ijp} \) for the location- and plate-sequence related systematic error can be obtained. Using this estimated value the term \( \left({\overline{x}}_p+{\hat{S}}_{ijp}\right) \) is now in essence the shifted plate- and location-dependent actual null-effect reference level which needs to be used in the different normalization expression if we want to consider the influence of \( {\hat{S}}_{ijp} \) on the relative compound responses on the % scale. Equation (3) describes the behavior of assay response values around the average null-effect (neutral control) response \( {\overline{x}}_p = {\overline{x}}_{N,p} \), but when e.g. calculating normalized percent inhibition values a ‘similar’ systematic error \( {\hat{S}}_{ijp}^{'} \) is also assumed to be present at the level of the full effect controls \( {\overline{x}}_{P,p} \). This value which participates in defining the %-effect scaling at location i,j, cannot be directly estimated from the data. While it is usually possible to determine \( {\hat{S}}_{ijp} \) near the \( {\overline{x}}_{N,p} \) response level quite reliably because in small molecule compound HTS most samples are inactive, and we thus possess a lot of data for its estimation, or we are able to base the estimation on designed control well positions and their responses, we can only make assumptions about the influence of the systematic error factors on response values around the \( {\overline{x}}_{P,p} \) level, unless detailed experiments about this aspect would be made. A purely additive response shift as per Eq. (5.3) for all possible magnitudes of the assay responses is an unreasonable assumption, especially if all values derived from a particular readout technology are limited to values ≥ 0. In the case of an inhibition assay the positive control \( {\overline{x}}_P \) corresponds usually to small readout signal values \( \left({\overline{x}}_{P,p}<{\overline{x}}_{N,p}\right) \) and we can either assume that \( {S}_{ijp}^{'}=0 \) at small signal levels, or we can assume a fractional reduction of the amplitude of the systematic error which is scaled linearly with the corresponding “High/Low” ratio as
i.e. a multiplicative influence of the systematic errors on the assay signal magnitude. A typical normalization expression (e.g. for NPI) with explicit consideration of the systematic error contributions at the \( {\overline{x}}_N \) and—if relevant—at the \( {\overline{x}}_P \) level forms the basis for integrating the response value correction into a reformulated expression for corrected NPI values:
noting that this is a now a plate- and well-location dependent normalization of the assay response data x i which eliminates the systematic batch-, plate- and well-level response distortions. As mentioned previously the corresponding modification for other simple normalization expressions can be derived in an analogous way. The difference between varying assumptions for the behavior of S ′ with changing absolute response leads to slightly different values for NPI corrected , the difference between the two being larger for smaller High/Low ratios of the assay controls (dynamic range of the assay response). Using Eq. (5.5) and the NPI expression from Table 5.5 we obtain:
or for the \( {S}_{i,p}^{'}=0 \) case:
These are useful relationships because in practice the simple NPI i,p which are based on plate-level controls without consideration of location-dependent effects are already available from the preceding data analysis step which converts the ‘arbitrary’ raw data values to a common % or fold scale. The NPI values are then used for various diagnostic graphics (heat maps, scatterplots, boxplots, etc.) and provide a basis for comparing the uncorrected and the subsequently corrected response values in a quick overview fashion (and to help in the visual detection of systematic response artifacts). The described \( {\hat{S}}_p^{'} \) correction ambiguity does of course not have any influence on the scoring methods which only rely on data centering with respect to the null-effect levels \( {\overline{x}}_N \), or better, with respect to the estimated actual null-effect response levels \( \left({\overline{x}}_N+{\hat{S}}_{ij}\right) \).
It is clear that the modeling approaches have to be chosen according to the actual types of patterns occurring in the data set, hence visual inspection, possible partitioning of data sets, and choice of an optimal model have to go hand in hand. Automatic partitioning of the plate data set can be done very efficiently by clustering the p × p correlation matrix of the pairwise inter-plate well values for all p plates. Response values for the samples or for samples together with neutral controls can be included in the correlations. Larger correlation values will be found for the plates which exhibit similar spatial response distortions, while we will have \( E\left( corr\left({\boldsymbol{x}}_k,\ {\boldsymbol{x}}_{\boldsymbol{l}}\right)\right)=0 \) for independent random distributions of the responses on plates k and l. The correlation matrix for our example data set with added (hierarchical) clustering information is shown in Fig. 5.4c. The main 4 sub-clusters can clearly be associated with the 4 discernable groups of plates with distinct error patterns (edge effects: 1- 20, edge effects + striping: 21–25, striping only 26–40, no systematic pattern: 41–50), in complete agreement with the structure of the data set.
Another method for the grouping of ‘similar’ plates which can be used for the purpose of partitioning is changepoint analysis (Hinkley 1970; Horvath 1993; Barry and Hartigan 1993) in a suitable set of QC indicators which are sensitive to specific differences in the systematic error pattern in the ordered sequence of plates. The first two components \( {\boldsymbol{U}}_i{\boldsymbol{D}}_i,\ i=1..2 \) of a robust principal pattern analysis using the robpca method (Hubert et al. 2005) for the entire simulated plate data set are shown in Fig. 5.5a, b.
(a) First principal pattern, and (b) second principal pattern extracted from the example plate data set with a robust principal component analysis, reflecting the dominant error patterns in the data: evaporation edge effects, tapered striping in alternate rows and defective pipetting tip in well 233. (c) loading factors of first principal pattern, and (d) loading factors for 2nd principal pattern, both with overlaid changepoint detection segments (red lines)
The two represented principal patterns clearly correspond to the main visible systematic error features of particular subsets of the plates with evaporation edge effects, liquid handling stripes in alternate rows which taper off at higher column numbers, as well as the single-well pipetting failure at (row = 10, col = 17). The corresponding principal component loadings V ′ i are shown in Fig 5.5c, d, respectively. Now we can use these PCA loadings, or similarly, the ICA mixture weights, from such an exploratory diagnostic analysis of the complete data set for changepoint (i.e. pattern- or plate batch-boundary) detection as indicated in these figures. The red horizontal lines indicate the extent of the data series with common mean and variance properties according to an analysis using the PELT method (Killick et al. 2012) which is also implemented in the R changepoint package (Killick and Eckley 2014). The superset of the changepoint locations from these 2 principal component loadings is in complete agreement with the pattern- and plate ‘batch’ boundaries (k = 20, 25, 40) which we had identified before and correspond to the properties of the generated data set. For plates 41 to 50 the average contribution from either of these two pattern components is close to 0, as indicated by the position of the red (mean) lines. The information from such an overview analysis, jointly together with the indicators of the significance of the Moran autocorrelation coefficients I from each plate allow us to quickly judge which plate sets we should include or exclude from any pattern modeling steps.
The principal pattern methods can be used both as quick diagnostic pattern detection tool, as just described, and of course also as basis for the data correction step. In Fig. 5.6 we demonstrate the application of this 2-component robust principal component model to the whole data set, with normalized data shown in a, error pattern in b, and resulting corrected data set in c. We also illustrate the behavior of the spatial autocorrelation coefficient before and after data correction and clearly see that the autocorrelated response contributions are removed to such an extent that Moran’s I values (Fig. 5.6d, e) are significantly reduced and most of the corresponding p-values (Fig. 5.6f, g) are now below the α = 0.05 level which is indicated by the dashed line.
(a) Example plate set, normalized data. (b) Systematic errors of plate data as estimated with a two-component principal pattern model. (c) Corrected plate data set after elimination of estimated systematic error values. (d) Moran autocorrelation coefficient for example plate data set before correction, and (e) after correction. (f) p-values of Moran coefficients before correction, and (g) their p-values after correction
Based on these ‘cleaned’ screening plate data sets where now all or most of the discernable systematic response errors have been eliminated using one of the described error-modeling and correction approaches the further screening data analysis steps are either (a) hit sample selection in primary screening, or (b) calculation of further quantities of interest based on data from a whole series of individual wells originating from one or several plates (e.g. concentration-response curve characterizations, compound potency determinations).
The ultimate optimality criterion for the error modeling and data correction steps is of course a maximized confirmation rate, i.e. a minimized rate of false discoveries in the list of selected hits, possibly under the constraint of an upper limit of the number of samples which can be selected and progressed into a next screening stage because of economic and resource constraints, and as small as possible false negative rate. Minimizing the false discovery rate in the primary screening hit list is expected to result in maximized confirmation rate of the putative hits in follow-up experiment, which is either a screen measuring the activity of the hits with independent replicates or a screen with concentration dependent measurements (Zhang et al. 2000; Wu and Liu 2008; Prummer 2012).
2.10 Sample Activity Ranking, Scoring and Tests of Significance
Since the hit identification from a single-concentration primary HTS with many expected actives is a large scale multiple hypothesis testing problem we need to consider practical methods for controlling the false-positive rate. The basic goal is to identify as many significant actives as possible which are not part of the null distribution, while at the same time incurring only a low proportion of false positives. The ‘false discovery rate’ statistic introduced two decades ago (Benjamini and Hochberg 1995) and related follow-up work by many authors (Efron et al. 2001; Efron 2004, 2010; Storey 2002; Storey and Tibshirani 2003; Dalmasso et al. 2005; Strimmer 2008a) has led to further development of the methodology and to its adaptation to the practical needs of large scale biological experimentation in genomics and screening. It is an ideal statistic for this purpose, because a straightforward normal p-value based threshold is not very informative in terms of presence of the interesting non-null component, because it is only related to the proportion of the samples with null-activity above this threshold. Methods like the q-value analysis and others (Storey and Tibshirani 2003; Strimmer 2008a) were shown to maintain high statistical power in many situations of biological experimentation and convey the necessary information about the proportion of the significant actives which are statistical false positives. These methods are very useful in practice because the ‘success rate’ of the subsequent follow-up verification of the detected activity from the primary screen is directly related to the q-value threshold, and the follow-up experimental confirmation and verification work (and related ‘costs’) can then be optimized in this respect.
In the screening data analysis context the mentioned false discovery rate FDR (proportion of false positives from the potentially interesting number of samples above the selection threshold, FDR = FP/(TP + FP)) and the false negative rate FNR (proportion of missed actives from the total number of real actives, FNR = FN/(TP + FN)) are the most informative and useful criteria for decision making at the primary hit selection stage.
From the explanations in the previous sections it is clear that the overall activity ranking of the probes to select the most ‘interesting’ should be done based on data which are corrected for any systematic response artifacts on a particular plate and also across the various plates measured in a screen. Activity ranking in primary screening can either be done using the normalized responses or by considering a related score value. Whereas the well–level data correction approaches have adjusted for systematic bias in the average responses, there may still be remaining differences in terms of systematic shifts of the assay variance, either plate-to-plate, batch-to-batch, or systematic differences related to the sample classes exposed to the screen (e.g. natural product fractions or extracts, single small molecular compounds, compound-mixtures, combinatorial library pools, siRNA pools etc.). Performing the analysis of the screening results based on scoring approaches can account for such differences in the assay “noise”. Its calculation can be based either on the (robust) variance estimation for the test sample area of the plates, or on the neutral control samples. In all analyses of corrected %-scale normalized or scored data we are working with scaled residuals \( {r}_{ijp} \propto {x}_{ijp}-{\overline{x}}_p \) for hit ranking and selection.
If the estimation of the scale value \( {\hat{s}}_p \), which is used for determining Z- or R-scores is based on the control samples, then the often limited number of them on a single plate can result in a large uncertainty of this estimate, especially for the lower plate densities of 96 and 384. For smaller n it may be advisable to use shrinkage estimation methods for the determination of the variance (Cui et al. 2005; Tong and Wang 2007; Murie et al. 2009) and obtain a regularized and more efficient estimate \( {\tilde {s}}_p \) by borrowing information from other (‘neighboring’) plates to prevent individual increases in the rates of false positives and/or false negatives when the corresponding platewise score values are over- or underestimated. But also for higher density plates variance shrinkage can be advantageous if the control well areas on some individual plates contain systematic errors which may not have been eliminated by the previously described response corrections if the ‘process-history’ of sample wells and control wells differ, which can be the case in some types of assays. The decomposition of the plate set into subsets belonging to particular measurement batches with similar response properties and separate estimation within these batch boundaries as well as adaptive local adjustment of the shrinkage parameters λ and w p in
on the time-ordered set of plates p will likely lead to a more efficient scale estimate \( {\tilde {s}}_p \), but the resulting score values may not be bias-free (see further below). The shrinkage parameters can be optimized by e.g. minimizing the calculated false discovery rate for a given Z- or R-score threshold, or maximizing the number of putative hits for a given preset FDR value, i.e. the fraction of identified hits which are “not interesting” based on purely statistical considerations. λ describes the mixing between the value \( {\hat{s}}_p \) of a particular individual plate and a component \( {\overline{s}}_{p,{w}_p} \) which has higher bias and lower variance, and which itself depends on a ‘smoothing’ parameter w p . The calculation of \( {\overline{s}}_{p,{w}_p} \) can e.g. be based on local averaging or on kernel smoothing of the values from ‘neighboring’ plates (Prummer 2012).
A scatterplot of (normal distribution) p-values from Z- or R-scores on the y-axis and the NPI, percent change, or fold change values (Cui and Churchill 2003) on the x-axis can be very useful to identify samples which exhibit a certain minimal %-effect change, and at the same time assess their statistical ‘significance’ (probability that the null-hypothesis of ‘no biological’ activity is true). Similar types of visualizations are also used in gene expression analysis, genome scale RNAi screens, or genome-wide association studies (GWAS).
As an illustration of the hit analysis we now return to the actual biochemical example screen which we have used in the section on assay- and screening QC metrics to show the typical behavior of selected measures for a complete screen with around 1 Mio pure compound samples in 692 1536-well plates. In this particular screen the plate data set was corrected with a ‘robust’ SVD modeling procedure which was composed of an initial median polish run, trimming of those data points which exhibit large residual values \( {r}_{ijp}>4\ ma{d}_p\left({x}_{ijp}\right) \) by replacing their values with the model values, and finally calculating a SVD model across the whole screen using this modified data set where the set of wells with large activity (NPI) values were thus effectively omitted from the final modeling step (method not listed in Table 5.6). This is the REOF procedure which is available, among several others previously listed in the standard Novartis NIBR in-house HTS data analysis software system (Gubler 2006). The scatterplot of normalized activity values (negative NPI values) and the corresponding normal distribution p-values calculated from the R-scores are shown in Fig. 5.7. This (half-) volcano plot allows a good simultaneous assessment of activity values and their statistical significance. A similar plot can of course be generated from %-activity and R-score values for situations where proper p-values cannot be obtained.
A threshold along the %-activity or along the p-value or score axis can e.g. be chosen so that the total number of hits is below some maximal number of samples which can be progressed to the next screening stage, while simultaneously considering the false discovery rate as outlined below. A %-activity threshold can also include a consideration of the minimal potency of potential hit samples, i.e. an estimate of the maximal acceptable IC 50 value which is of interest in further analysis by transforming the NPI values to such an IC 50 by assuming an ‘ideal’ concentration response relationship (see Eq. (5.11) below). For example when setting a threshold at 50 % inhibition we would expect to be able to detect hit compounds with IC 50 values which are smaller than the concentration used in the screening run (Gubler et al. 2013). For samples without known concentration in screening (e.g. siRNA, shRNA) such a direct translation can of course not be done.
A two component mixture model for the overall distribution function of the normalized activity values of all results from a particular screen can be defined as
with proportion π 0 of the null-distribution function f 0(x) for the inactive samples and proportion \( \left(1-{\pi}_0\right) \) for the alternative distribution f A (x) of the active samples. When applying this approach in practice to a large number of Z-, R- or equivalent score values, then it is reasonable to assume \( {f}_0(x)\sim N\left(0,1\right) \), provided that centering, possible error correction and the scaling values were estimated correctly for the calculation of x. Especially the unbiased estimation of \( {\tilde {s}} \) is crucial for transforming scores into normal probability p-values, as \( {f}_0(p)\sim U\left(0,1\right) \) only when this prerequisite is fulfilled. In this sense the shape of the distribution for ‘larger’ p-values, where the influence of f A becomes negligible, can be used as a practical consistency check for the appropriateness of the scale estimates \( {\tilde {s}} \) used in the score calculations.
In practice we may also encounter situations where f 0(x) is not normally distributed as is the case e.g. after the application of the simple median polish procedure. Given an initial normal distribution of values the residuals will in this case usually appear leptokurtic with a central narrower peak and a related density depression in its immediate neighborhood due to the various ‘median’ operations, whereas the wider tail areas follow relatively closely a normal distribution density with a slight excess at larger x. In such situations it is also very likely that the scale \( {\tilde {s}} \) will be underestimated and score values come out too large when compared to the original residual scale. Due to both possible reasons it does also no longer make sense to assign normal distribution p-values to the resulting corrected score values x. If there is interest to explicitly extract f A (x) from such an activity- or score distribution then a Bayesian mixture modeling approach with appropriate empirical priors may be used, now with f 0(x) itself already being a mixture of a narrow and a broad component. If this is not done, or cannot be done, then simple activity rankings will have to suffice for hit identification in routine screening data analysis, but the additional highly useful information on e.g. estimated false discovery rates cannot be obtained from the data. In any case, a lower limit for the hit-threshold, in terms of %-activity or score values, can be derived from \( {\tilde {s}}_N \) of the N controls, because in practice we always have \( {\tilde {s}}_N\le {\tilde {s}}_C \) due to additional experimental variability for the compound samples C as compared to the more homogeneous set of neutral controls N.
In our particular illustration of screening hit analysis the R-scores were initially calculated using the platewise \( {\tilde {s}}_N \) values based on a loess based variance shrinkage procedure as described above. The corresponding score values were then used to estimate the mixture components and perform a false discovery rate analysis using the fdrtool R package (Strimmer 2008b). In order to compensate for any remaining bias in the score estimates x the following procedure was applied: An average R-score bias correction factor was derived from comparing the null distributions \( {f}_0({x}_N) \sim N(0,{s}_N^2 \)) for \( \{{x}_N:x\in N\} \) and \( {f}_0({x}_C) \sim N(0,{s}_C^2 \)) for \( \{{x}_C:x\in C\} \), then rescaling the scores x as \( {x}^{'}=x/{s}_N \) , resulting in \( {f}_0({x}_N^{'}) \sim N(0,1 \)) and \( {f}_0({x}_C^{'}) \sim N(0,{s}_C^2/{s}_N^2 \)) where s C /s N should now be close to 1 if we assume that the same average variance estimation bias is present in the control and compound data samples. In this case this is borne out by the actual data from this screen and can be seen in Fig. 5.8a where we obtain \( {f}_0({x}_C^{'}) \sim N(0,{1.02}^2) \) after the described bias correction with the normal null-density scaling factor of \( {s}_N=0.93 \).
(a) Decomposition of bias-corrected R-score values into null- and alternative mixture components using R package fdrtool. (b) p-value distribution of the same dataset with indication of hit selection threshold of p = 0.006 (red vertical line) and estimated percentage of null samples \( \hat{\pi}_0 \) = 0.84 (red horizontal line). (c) Expected false positives as function of total number of significant tests following from tail area false discovery rate analysis using R package qvalue, with red marker lines for Fdr = 0.1
Using the same set of scaled R-score values x ′ we can also see that the corresponding normal distribution p-values in Fig. 5.8b show the ‘expected’ behavior, i.e. they are essentially flat in the large p region corresponding to a normal f 0 density with an average value of the proportion of samples following the null distribution of \( {\hat{\pi}}_0=0.84 \) and with a peak related to the alternative f A distribution of the non-null samples at small p-values. Incidentally, we obtain consistent π 0 values of 0.86 also from fdrtool and 0.85 from the qvalue R packages (Storey and Tibshirani 2003). This consistency allows us to have confidence in the related tail area ‘Fdr’ (Efron 2004) values as reported by qvalue (see Fig. 5.8c) where we can see that Fdr ≤0.1 for up to a total number of identified hits (significant tests) of \( {n}_{hit}\cong \) 49,000, also in agreement with the direct p-value histogram analysis of the Fdr fraction. This q-value (≤0.1) corresponds in this particular case to a one-sided p-value of 0.006 and a Z (R-score) threshold of around 2.5. When limiting n hit to 16,000 by selecting a R-score cutoff of around 4, then we obtain an expected Fdr close to 0. This means that we can expect a very high hit confirmation rate—close to 100 %—in a follow-up verification experiment.
With an average \( {\tilde {s}}_N \) value of 4.5 % (rescaled as per the described average bias correction procedure), this score threshold can be converted into a %-inhibition (NPI) threshold of 4⋅4.5 % = 18.0 %. Disregarding the different plate-to-plate variances which are taken into account in the R-scores, but not in the NPI values, we can then loosely say that the actually chosen threshold value of 20 % inhibition for this screen is thus quite ‘conservative’ with respect to the false discovery rate criterion. From such a mixture distribution- and p-value histogram analysis we can also get estimates for the proportion of the non-detected active samples for the threshold setting corresponding to Fdr = 0.1 example value and obtain a false negative FNR = 0.39 in this case, or FNR = 0.68 for the threshold at R-score = 4. A set of p-value distribution and FDR analyses for different screening assays done by Prummer (2012) shows quite good agreement between estimated confirmation rates and the actually observed ones for a series of screens where follow-up experiments were performed as complete concentration-response measurements.
When analyzing screening data with replicates the (compound-) sample specific data variability can be estimated, resulting in the possibility to apply more powerful for hit-selection methods than for the n = 1 case where we are essentially assuming that every sample has the same variability. Of course it is always possible to simply average the normalized responses from the replicates and analyze them in the same way as the unreplicated primary data. This is not a real problem in small molecule compound screening, but e.g. for RNAi screening where we expect differences of variability among different siRNA samples it is essential to apply statistical methods which consider these possible sample specific differences. When comparing different RNAi treatment effects to each other the use of good estimates of the variability and use of efficient statistics becomes crucial. Many different approaches for analyzing and identifying hits from replicated data have been developed for gene expression array data (Kerr and Churchill 2001; McDavid et al. 2013; Baldi and Long 2001; Smyth 2004) and for RNAi screening (Birmingham et al. 2009), including Bayesian modeling and hypothesis testing approaches (Zhang et al. 2008). SSMD-based analysis (see Table 5.2) and hit selection rules for RNAi screening data with and without replication, as well as statistical methods for comparison of gene effects between groups are discussed in depth in a book by Zhang (2011a).
Gene expression experiments are usually using both biological and technical replication, i.e. measuring readout variability of responses from different sample sources and from different measurements of the same sample in a given experiment. Also in RNAi screening we can have similar situations: replicates from different sample preparations or simply multiple measurements of the same sample. Estimation and modeling of the error components from the replicated sample, the replicated measurements from of a single sample and combining these with the estimates from the neutral control samples can be done in different ways to provide more efficient variance estimation for subsequent hypothesis testing. Regularized variance estimations for use in hit selection methods often use shrinkage and Bayesian approaches and will lead to more powerful ‘regularized’ hypothesis testing (e.g. regularized t-statistics, Baldi and Long 2001; Tusher et al. 2001; Murie et al. 2009). The advantage of Bayesian methods is to allow incorporating balanced information from sample wells and control wells in the posterior distributions and then assigning probabilities to the test samples of belonging to either of the no-effect, activation or inhibition groups. The wide area of statistical methods for gene expression analysis and identification of differentially expressed genes (equivalent to effect-size assessment and hit selection for replicated screening data) was worked on extensively in the past 10–15 years and it is impossible to cover in this short chapter and we refer the reader to the extensive base of published literature on (primarily) microarray, but also RNAi screening data analysis (Birmingham et al. 2009, and references therein).
As mentioned previously in the section on data normalization for RNAi screens the siRNA effects are often better quantified by using the SSMD-statistic than by using R-scores, especially when working with replicates. In a similar fashion than shown in the previous volcano plot example for a compound-based ‘%-inhibition screen’ we can also plot p-values based on SSMD analysis and the Fold-change values, instead of using the Z- or R-score based p-values and the %-inhibition values, to arrive at an equivalent visualization (even including the various types of controls in the same plot for additional information content). The derivation of FP, FN and related FDR rates based on the SSMD-statistic is outlined in much detail in Zhang (2011a).
In contrast to the RNAi screening and gene expression analysis areas publications on the topic of replication for small-molecule HTS data in full primary screens are practically non-existent because replicate measurements are essentially only done in the initial pilot phase where the focus is primarily on assessing the degree of agreement, and then again in the confirmation stage, i.e. after the primary screening hit selection where the analysis is again more centered around verifying the initially detected primary screening effect of the putative hits. Determining responses from replicated measurements using samples from the same or from different formulations of the same compound at a single concentration and/or the determination of a verifiable and plausible concentration dependence of the sought effect is the prime focus in this phase, but this is no longer a large-scale multiple hypothesis testing process with the associated need to control false positive (false discovery) and false negative rates.
We note again that the final hit list is often additionally filtered by considering counter screening results besides the results from the main screen, and we also need to mention that the HTS community often uses the term ‘false positives’ in the sense that a chemical test sample may well be verifiably active in a given assay through a target-independent effect, but inactive towards the actual biological target, as e.g. detected by a counter screen. This is obviously not the same use of the term ‘false positive’ as in statistics.
In compound screening additional elimination and selection steps are often made to modify the primary hit list, either by applying filters which identify unwanted chemical structure elements and compounds possessing specific chemical property values, or also augmenting the hit list based on structure-activity considerations (‘hit list enrichment’) with related similar compounds for subsequent confirmation or other follow-up experiments (Sun et al. 2010; Varin et al. 2010; Mulrooney et al. 2013).
2.11 Calculation of Derived Quantities from Normalized Plate and Well %Activity Data
The estimated standard errors and the unbiasedness of derived summary results or of parameters of calculations of derived quantities which are themselves based on a whole series of %-activity values in different wells in one or multiple plates also depend on best possible plate data quality in a very similar way as the data used for primary hit selection. Thus, every effort should be made to apply the response error correction methods outlined above also in these types of experiments. Because most of these types of experiments rely on a high percentage of non-null data the previously described modeling approaches can often not be used directly. Useful models of the systematic errors can nonetheless be established by using control data from suitable plate designs (‘uniform’ placement of control wells across the plates), or often better, by a separate set of control plates measured in parallel to the sample plates. The advantage of the latter approach is that we are then also able to model and correct for possible striping, edge effects and more ‘complicated’ patterns than e.g. linear or 2nd degree polynomial response surface dependencies which are essentially the only practical possibilities when using a limited number of control wells as pattern model anchor points on the sample plates. An example of such control plate correction approaches (CPR, control plate regression) and assessment of their success in concentration-response screens was described by Murie et al. (2014).
The most frequent case of calculations of such derived results from High-Throughput Screening plate experiments is the determination of the concentration-response curve (CRC) characteristics of the compound samples and the estimation of the parameters of the simple phenomenological sigmoidal 4-parameter-logistic (4PL) Hill curve function
if a significant concentration dependence exists in the experimental range employed in such a screen (Ritz and Streibig 2005; Formenko et al. 2006). In Eq. (5.11) we have concentration x, response y, and the 4 function parameters A 0, A inf (lower and upper plateau values), AC 50 (inflection point) and α (Hill slope parameter), and \( {f}^{4 PL}\left(A{C}_{50}\right)=\left({A}_{inf}+{A}_0\right)/2 \). Note our use of the generic term AC 50 instead of the specific terminology IC 50 for inhibition (IC: inhibitory concentration) and EC 50 for activation experiments (EC: effective concentration).
The original Hill function (Hill 1910) has a theoretical basis in simple ligand binding theory, albeit with some unrealistic mechanistic assumptions (Goutelle et al. 2008). For all practical purposes in screening, especially for more complex cellular assays where potentially a whole cascade of biochemical reaction steps is involved, it should be viewed more as a convenient empirical model. Rankings of the ‘strength’ of the validated hit compound effects are usually based on the AC 50 parameter, but for some types of assays A inf , \( \left({A}_{inf}-{A}_0\right) \) or an ‘area under the curve’ effect measure which combines potency and efficacy into a single number are also considered (Huang and Pang 2012).
Even if the compounds for concentration dependent measurements are pre-selected to have shown activity in the previous concentration-independent screening primary runs the number of cases of ‘incomplete’ curves is usually still quite high, of course dependent on chosen hit threshold, previously used primary screening concentration and actual concentration range of the CRC experiments. In the HTS confirmation and validation screens the concentration range is held constant and not adapted on a compound by compound basis as might be done in low throughput compound profiling experiments. An accurate and independent determination of all regression function parameters is then not always possible and the analysis software needs to produce sensible results also in the initial presence of high degrees of parameter correlations, e.g. by automatic identification of ill-determined parameters and sensible application of constraints, or by switching to simpler fallback models, if needed.
HTS production-quality software for automated nonlinear regression fitting of large series (i.e. thousands or tens of thousands) of concentration-response curves needs to work in a highly robust fashion, use autostarting methods for estimation of initial parameters, and be able to handle outliers, e.g. by using resistant regression algorithms (M-estimation, IRLS iterative reweighted least squares) (Normolle 1993; Street et al. 1988). Also the likely presence of outliers needs to be signaled as diagnostic information. The analysis methods implemented in the software also need to be able to deal flexibly with other frequent ‘unusual’ situations like completely inactive compounds which lead to ∼constant activity values over the full concentration range making it impossible to derive the 4 parameters of Eq. (5.11) in a meaningful way, or with the mentioned ‘incomplete curves’ where e.g. the response plateau at higher concentrations is not reached, and generally with unusual non-sigmoidal curve shapes (e.g. bell-shaped, inversely bell-shaped, multiphasic).
If experimental curve shapes do not follow the sigmoidal shape of Eq. (5.11) then derived parameters can be strongly biased, up to the level of being completely misleading to the analyst, e.g. when trying to represent a bell-shaped curve with f 4PL. Thus, the detection of lack of fit and the selection of alternate models for the regression analysis is an important topic for large scale concentration response curve fitting in screening. A pragmatic modeling approach is to move away from a parametric representation of a given curve and choose a nonparametric model f NP as per Eq. (5.12)
if this is necessary. This model switching decision can e.g. be based on predefined lack of fit criteria together with the occurrence of large parameter dependency values. Depending on the actual model choice for f NP, a preparatory optimization of one or more model parameters Θ for use in the actual curve analysis may be needed to reflect the experimental design and the typical assay variability. This can e.g. be done by assessing penalized likelihood measures, e.g. the Akaika information criterion (Akaike 1974), or using generalized cross validation (Craven and Wahba 1979) on the complete curve data set by varying Θ within appropriate ranges. A practical choice for f NP is e.g. a smoothing spline function (Frommolt and Thomas 2008), possibly including monotonicity constraints (Kelly and Rice 1990). It is understood that the x values in these functions are the logarithmic concentrations log10(x). For simplicity of notation we are not explicitly mentioning the logarithms in the remainder of this section.
Instead of the previous 4 model parameters in Eq. (5.11) we then use surrogate values which provide an equivalent characterization of the concentration-response relationship, e.g. the set of values f(x min ), f(x max ), min(f(x)), max(f(x)), arg minx f(x), arg maxx f(x), and also the ‘absolute’ AC 50 (concentration of the intersection of the fitted curve with the 50 % level as given by the N and P controls) abs \( A{C}_{50}=\left\{x:f(x)=50\right\} \), as well as an approximate ‘equivalent Hill slope’ \( \alpha \approx 4{\left(df/dx\right)}_{x= absA{C}_{50}}/\left( \ln (10)\left(f\left({x}_{max}\right)-f\left({x}_{min}\right)\right)\right) \), together with proper consideration of non-existent or multiple intersections in the set \( \left\{x:f(x)=50\right\} \).
Such a nonparametric approach is also useful for confirmation CRC experiments where only 2–4 different concentrations may be used to verify the existence of reasonable concentration dependences and to obtain a rough estimate of abs AC 50 for the putative hits, making it essentially impossible to use parametric nonlinear regression methods. For the analysis of such experiments the choice of interpolation splines for f NP are preferred over smoothing spline functions.
For some types of assays (e.g. cell proliferation inhibition) the abs AC 50 value—which can of course also be directly calculated from f 4PL(x) when using the model represented by Eq. (5.11)—can be biologically more meaningfully interpreted than the AC 50 function parameter (Sebaugh et al. 2011). In such experiments the concentration where 50 % response inhibition with respect to the control values is reached will have an easily interpretable meaning, whereas the position of the AC 50 concentration (i.e. the inflection point of the curve) can e.g. be influenced by the fact that we can obtain cell count readings at high concentrations which lie below the 100 % inhibition baseline value (when cells are killed) or that the 100 % inhibition level is not reached (cell growth cannot be completely inhibited), i.e. we can have \( \Delta A=\left|{A}_{inf}-{A}_0\right| \) >100 % or ΔA <100 %. As a consequence the position of the inflection point is less informative than the position of the intersection of the response curve with a particular prescribed inhibition level.
When considering AC 50 or abs AC 50 potency rank orders for the final selection of hits in the confirmation or validation screens the MSR (minimum significant ratio) which was optimally already derived in the assay adaptation stage (see Table 5.3) can be used as an indicator for assessing the ‘significance’ of potency differences (Eastwood et al. 2005, 2006). Other relevant measures in this context, when comparing the potency values from the target specific screen and one or several parallel selectivity screens, are the MSSR (minimum significant selectivity ratio) and MSRSR (minimum significant ratio of selectivity ratios) values. They are calculated in a similar way as MSR (Goedken et al. 2012) and give information on the confidence limits of the selectivity ratio \( SR= \) AC 50(off-target assay)/AC 50(on-target assay) and of ratios of SR values for different compounds.
3 Open Questions and Ongoing Investigations in the HTS and HCS Data Analysis Methods Field
Many aspects of small-molecule and RNAi High-Throughput Screening data analysis were explored in the past 10–15 years and several publications describe the details of the most relevant statistical aspects for the analysis of HTS data:
-
Detection, modeling and correction of systematic error patterns of the plate-based screening readouts and to minimize selection bias, and thus allowing
-
Optimal activity scoring and ranking of active features in the hit selection process to minimize the false discovery rate, minimize the number of false negatives and maximize the success rate of follow-up screening stages.
Nonetheless, in the author’s opinion several specific areas merit to be explored and reported in more depth for the benefit of the whole HTS and HCS community:
-
Which are the most optimal plate designs for replicate single concentration and concentration-response experiments? Which are the most optimal locations for replicate data points distributed on a single plate, or on multiple ‘replicate’ plates, given certain types of systematic errors?
-
Is there an overall ‘best’ modeling and correction method for certain types of systematic error patterns? Which methods are overall most efficient and have the least amount of bias?
-
Standard testing datasets and platform for comparing different analysis methods, including test and performance results.
-
Response error modeling methods involving many median operations in their calculation (like e.g. the median polish and the simplified B-score methods) often lead to strongly non-normal distributions of the residuals. What is the best and most practical way to perform false discovery rate analyses in such cases?
The following are questions, active research topics and future directions of necessary statistical work in the area of high-dimensional multivariate screening data analysis. The ‘classical’ HTS assays, readout technologies and associated data analysis methods as outlined in this chapter will keep a lot of their present importance, and even become more pervasive in academia and smaller biological research laboratories for target identification, target validation and screening for active chemical features. But active research and development of statistical data analysis methods in the HTS/HCS field now center much more on these general questions:
-
What are the most optimal normalization, feature selection, dimension reduction, error correction, scoring and classification methods for high-dimensional multivariate data from phenotypic image based High-Content Screening and other similar sources of such data? Under which conditions are the particular methods applicable? What are their advantages and disadvantages?
-
What is the influence of imaging parameters and noise on the localization of subcellular features and what is the influence of different types of image analysis artifacts on HCS data analysis? How do these affect the derived (usually lower dimensional) final measures and classification results?
-
What are the most suitable and informative analysis methods, including normalization and possible systematic error correction questions, for single cell data and multivariate time-course data?
References
Abraham VC, Taylor DL, Haskins JRL (2004) High content screening applied to large-scale cell biology. Trends Biotechnol 22(1):15–22
Abraham Y, Zhang X, Parker CN (2014) Multiparametric Analysis of Screening Data: Growing Beyond the Single Dimension to Infinity and Beyond. J Biomol Screen 19(5):628–639
Akaike H (1974) A new look at the statistical model identification. IEEE Trans Autom Contr 19(6):716–723. doi:10.1109/TAC.1974.1100705
Baker M (2010) Academic screening goes high-throughput. Nat Methods 7:787–792. doi:10.1038/nmeth1010-787
Bakheet TM, Doig AJ (2009) Properties and identification of human protein drug targets. Bioinformatics 25(4):451–457
Baldi P, Long AD (2001) A Bayesian framework for the analysis of microarray expression data: regularized t-test and statistical inferences of gene changes. Bioinformatics 17:509–519
Barry D, Hartigan JA (1993) A Bayesian analysis of change point problems. J Am Stat Assoc 88:309–319
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B 57(1):289–300
Birmingham A, Selfors LM, Forster T, Wrobel D, Kennedy CJ, Shanks E, Santoyo-Lopez J, Dunican DJ, Long A, Kelleher D, Smith Q, Beijersbergen RL, Ghazal P, Shamu CE (2009) Statistical methods for analysis of high throughput RNA interference screens. Nat Methods 6(8):569. doi:10.1038/nmeth.1351
Bland JM, Altman DG (1986) Statistical methods for assessing agreement between two methods of clinical measurements. Lancet 1:307–310
Boutros M, Bras LP, Huber W (2006) Analysis of cell-based RNAi screens. Genome Biol 7:R66. doi:10.1186/gb-2006-7-7-r66
Box GEP, Hunter WG, Hunter JS (1978) Statistics for experimeers: an introduction to design, data analysis and model building. Wiley, New York. ISBN:0-471-09315-7
Bray MA, Carpenter AE (2012) Advanced assay development guidelines for image-based high content screening and analysis. In: Sittampalam GS (ed) Assay guidance manual. http://www.ncbi.nlm.nih.gov/books/NBK53196/. Accessed 15 Oct 2014
Brideau C, Gunter B, Pikounis B, Liaw A (2003) Improved statistical methods for hit selection in high-throughput screening. J Biomol Screen 8(6):634–647
Brummelkamp TR, Fabius AWM, Mullenders J, Madiredjo M, Velds A, Kerkhoven RM, Bernards R, Beijersbergen RL (2006) An shRNA barcode scrren provides insight into cancer cell vulnerability to MDM2 inhibitors. Nat Chem Biol 2(4):202–206
Bushway PJ, Azimi B, Heynen-Genel S, Price JH, Mercola M (2010) Hybrid median filter background estimator for correcting distortions in microtiter plate data. Assay Drug Dev Technol 8(2):238–250. doi:10.1089/adt.2009.0242
Carpenter AE (2007) Image-based chemical screening. Nat Chem Biol 3(8):461–465. doi:10.1038/nchembio.2007.15
Coma I, Clark L, Diez E, Harper G, Herranz J, Hofmann G, Lennon M, Richmond N, Valmaseda M, Macarron R (2009a) Process validation and screen reproducibility in high-throughput screening. J Biomol Screen 14:66–76
Coma I, Herranz, J, Martin J (2009) Statistics and decision making in high-throughput screening. In: William P, Janzen WP, Bernasconi P (eds) High-throughput screening. Methods in molecular biology, vol 565. Humana, Totowa. ISBN:978-1-60327-257-5
Craven P, Wahba G (1979) Smoothing noisy data with spline functions. Numer Math 31:377–403
Cui X, Churchill GA (2003) Statistical tests for differential expression in cDNA microarray experiments. Genome Biol 4(4):210
Cui X, Hwang JTG, Qiu J, Blades NJ, Churchill GA (2005) Improved statistical tests for differential gene expression by shrinking variance components estimates. Biostatistics 6(1): 59–75
Dalmasso C, Broet P, Moreau T (2005) A simple procedure for estimating the false discovery rate. Bioinformatics 21(5):660–668. doi:10.1093/bioinformatics/bti063
Davies JW, Glick M, Jenkins JL (2006) Streamlining lead discovery buy aligning in silico and high-thtoughput screening. Curr Op Chem Bio 10:343–351
Dean A, Lewis S (eds) (2006) Screening: methods for experimentation in industry, drug discovery, and genetics. Springer, New York. ISBN 978-1-4419-2098-0
R Development Core Team (2013) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. http://www.Rproject.org. ISBN:3900051070
Dragiev P, Nadon R, Makarenkov V (2011) Systematic error detection in experimental high-throughput screening. BMC Bioinformatics 12:25
Duerr O, Duval F, Nichols A, Lang P, Brodte A, Heyse S, Besson D (2007) Robust Hit identification by quality assurance and multivariate data analysis of a high content, cell based assay. J Biomol Screen 12(8):1042–1049
Eastwood BJ, Chesterfield AK, Wolff MC, Felder CC (2005) Methods for the design and analysis of replicate-experiment studies to establish assay reproducibility and the equivalence of two potency assays. In: Gad S (ed) Drug discovery handbook. Wiley, New York
Eastwood BJ, Farmen MW, Iversen PW, Craft TJ, Smallwood JK, Garbison KE, Delapp NW, Smith GF (2006) The minimum significant ratio: a statistical parameter to characterize the reproducibility of potency estimates from concentration-response assays and estimation by replicate-experiment studies. J Biomol Screen 3:253–261
Echeverri CJ, Perrimon N (2006) High-throughput RNAi screening in cultured cells – a user’s guide. Nat Rev Genet 7:373–384
Efron B (2004) Large-scale simultaneous hypothesis testing: the choice of a null hypothesis. J Am Stat Assoc 99(465):96–104
Efron B (2010) Large-scale inference. Cambridge University Press, Cambridge. ISBN 978-0-521-19249-1
Efron B, Tibshirani R, Storey J, Tusher V (2001) Empirical Bayes analysis of a microarray experiment. J Am Stat Assoc 96:1151–1160
Formenko I, Durst M, Balaban D (2006) Robust regression for high-throughput screening. Comput Methods Prog Biomed 82:31–37
Fox S, Farr-Jones S, Sopchak L, Boggs A, Nicely HW, Khoury R, Biros M (2006) High-throughput screening: update on practices and success. J Biomol Screen 11(7):864–869
Fox SJ (ed) (2002) High throughput screening 2002: new strategies and technologies. High Tech Business Decisions, Moraga
Frommolt P, Thomas RK (2008) Standardized high-throughput evaluation of cell-based compound screens. BMC Bioinformatics 9:475
Geary RC (1954) The contiguity ratio and statistical mapping. Inc Stat 5(3):15–145. doi:10.2307/2986645
Goedken ER, Devanarayan V, Harris CM, Dowding LA, Jakway JP, Voss JW, Wishart N, Jordan DC, Talanian RV (2012) Minimum significant ratio of selectivity ratios (MSRSR) and confidence in ratio of selectivity ratios (CRSR): quantitative measures for selectivity ratios obtained by screening assays. J Biomol Screen 17(7):857–867. doi:10.1177/1087057112447108
Goutelle S, Maurin M, Rougier F, Barbaut X, Bourguignon L, Ducher M, Maire P (2008) The Hill equation: a review of its capabilities in pharmacological modelling. Fundam Clin Pharmacol 22(6):633–648. doi:10.1111/j.1472-8206.2008.00633.x
Gubler H (2006) Methods for statistical analysis, quality assurance and management of primary high-throughput screening data. In: Hüser J (ed) High-throughput screening in drug discovery. Methods and principles in medicinal chemistry, vol 35. Wiley-VCH GmbH, Weinheim, pp 151–205. doi:10.1002/9783527609321.ch7
Gubler H, Schopfer U, Jacoby E (2013) Theoretical and experimental relationships between percent inhibition and IC50 data observed in high-throughput screening. J Biomol Screen 18(1):1–13. doi:10.1177/1087057112455219
Gunter B, Brideau C, Pikounis B, Liaw A (2003) Statistical and graphical methods for quality control determination of high-throughput screening data. J Biomol Screen 8(6):624–633
Haney SA (2014) Rapid assessment and visualization of normality in high-content and other cell-level data and its impact on the interpretation of experimental results. J Biomol Screen 19(5):672–684
Heuer C, Haenel T, Prause B (2002) A novel approach for quality control and correction of HTS based on artificial intelligence. Pharmaceutical Discovery and Development 2002/03, PharmaVentures Ltd., Oxford
Heyse S (2002) Comprehensive analysis of high-throughput screening data. In: Bornhop DJ et al (eds) Proceedings of the SPIE, Biomedical Nanotechnology Architectures and Applications 4626, pp 535–547
Hill AV (1910) The possible effects of the aggregation of the molecules of haemoglobin on its dissociation curves. J Physiol 40:iv–vii
Hill AA, LaPan P, Li Y, Haney SA (2007) Analysis of multiparametric HCS data. In: Haney SA (ed) High content screening: science, techniques and applications. Wiley, New York. doi:10.1002/9780470229866.ch15
Hinkley DV (1970) Inference about the change-point in a sequence of random variables. Biometrika 57(1):1–17
Hoaglin DC, Mosteller F, Tukey JW (1983) Understanding robust and exploratory data analysis. Wiley, New York. ISBN 0-471-09777-2
Horvath L (1993) The maximum likelihood method for testing changes in the paramaters of normal observations. Ann Stat 21(2):671–680
Huang S, Pang L (2012) Comparing statistical methods for quantifying drug sensitivity based on in vitro dose–response assays. Assay Drug Dev Technol 10(1):88–96. doi:10.1089/adt.2011.0388
Hubert M, Rousseuw PJ, Vanden Branden K (2005) ROBPCA: a new approach to robust principal component analysis. Technometrics 47:64–79
Hughes JP, Rees S, Kalindjian SB, Philpott KL (2011) Principles of early drug discovery. Br J Pharmacol 162:1239–1249
Hughes M, Inglese J, Kurtz A, Andalibi A, Patton L, Austin C, Baltezor M, Beckloff M, Sittampalam S, Weingarten M, Weir S (2012) Early drug discovery and development guidelines: for academic researchers, collaborators, and start-up companies. In: Sittampalam S et al (eds) Assay guidance manual. Eli Lilly & Company and the National Center for Advancing Translational Sciences, Bethesda. http://www.ncbi.nlm.nih.gov/books/NBK92015/
Hyvarinen A, Karhunen J, Oja E (2001) Independent component analysis. Wiley, New York. ISBN 978-0471405405
Ilouga PE, Hesterkamp T (2012) On the prediction of statistical parameters in high-throughput screening using resampling techniques. J Biomol Screen 17(6):705–712. doi:10.1177/1087057112441623
Iversen PW, Eastwood BJ, Sittampalam GS (2006) A comparison of assay performance measures in screening assays: signal window. Z′-factor and assay variability ratio. J Biomol Screen 11(3):247–252
Kaiser J (2008) Industrial-style screening meets academic biology. Science 321(5890):764–766. doi:10.1126/science.321.5890.764
Kelly C, Rice J (1990) Monotone smoothing with application to dose-response curves and the assessment of synergism. Biometrics 46(4):1071–1085
Kerr MK, Churchill GA (2001) Experimental design for gene expression microarrays. Biostatistics 2(2):183–201
Kevorkov D, Makarenkov V (2005) Statistical analysis of systematic errors in high-throughput screening. J Biomol Screen 10(6):557–567
Killick R, Eckley IA (2014) Changepoint: an R package for changepoint analysis. J Stat Software 58(3):1--19
Killick R, Fearnhead P, Eckley IA (2012) Optimal detection of changepoints with a linear computational cost. J Am Stat Assoc 107(500):1590–1598
König R, Chiang CY, Tu BP, Yan SF, DeJesus PD, Romero A, Bergauer T, Orth A, Krueger U, Zhou Y, Chanda SK (2007) A probability-based approach for the analysis large scale RNAi screens. Nat Methods 4(10):847–849
Kramer R, Cohen D (2004) Functional genomics to new drug targets. Nat Rev Drug Discov 3(11):965–972
Kümmel A, Selzer P, Siebert D, Schmidt I, Reinhardt J, Götte M, Ibig-Rehm Y, Parker CN, Gabriel D (2012) Differentiation and visualization of diverse cellular phenotypic responses in primary high-content screening. J Biomol Screen 17(6):843–849. doi:10.1177/1087057112439324
Loo LH, Wu LF, Altschuler SJ (2007) Image-based multivariate profiling of drug responses from single cells. Nat Methods 4(5):445. doi:10.1038/NMETH1032
Macarron R, Hertzberg RP (2011) Design and implementation of high-throughput screening assays. Mol Biotechnol 47(3):270–285. doi:10.1007/s12033-010-9335-9
Macarron R, Banks MN, Bojanic D, Burns DJ, Cirovic DA, Garyantes T, Green DVS, Hertzberg RP, Janzen WP, Paslay JW, Schopfer U, Sittampalam GS (2011) Impact of high-throughput screening in biomedical research. Nat Rev Drug Discov 10:188–195. doi:10.1038/nrd3368
Majumdar A, Stock D (2011) Large sample inference for an assay quality measure used in high-throughput screening. Pharm Stat 1:227–231
Makarenkov V, Zentilli P, Kevorkov D, Gagarin A, Malo N, Nadon R (2007) An efficient method for the detection and elimination of systematic error in high-throughput screening. Bioinformatics 23:1648–1657
Malo N, Hanley JA, Cerquozzi S, Pelletier J, Nadon R (2006) Statistical practice in high-throughput screening data analysis. Nat Biotechnol 24:167–175
Mangat CS, Bharat A, Gehrke SS, Brown ED (2014) Rank ordering plate data facilitates data visualization and normalization in high-throughput screening. Biomol Screen 19(9): 1314–1320. doi:10.1177/1087057114534298
Matson RS (2004) Applying genomic and proteomic microarray technology in drug discovery. CRC, Boca Raton. ISBN 978-0849314698
Mayr LM, Bojanic D (2009) Novel trends in high-throughput screening. Curr Opin Pharmacol 9(5):580–588
Mayr LM, Fuerst P (2008) The future of high-throughput screening. J Biomol Screen 13:443–448
McDavid A, Finak G, Chattopadyay PK, Dominguez M, Lamoreaux L, Ma SS, Roederer M, Gottardo R (2013) Data exploration, quality control and testing in single-cell qPCR-based gene expression experiments. Bioinformatics 29(4):461–467. doi:10.1093/bioinformatics/bts714
Millard BL, Niepel M, Menden MP, Muhlich JL, Sorger PK (2011) Adaptive informatics for multifactorial and high-content biological data. Nat Methods 8(6):487
Moran PAP (1950) Notes on continuous stochastic phenomena. Biometrika 37(1):17–23. doi:10.2307/2332142
Mosteller F, Tukey J (1977) Data analysis and regression. Addison-Wesley, Reading. ISBN 0-201-04854-X
Mulrooney CA, Lahr DL, Quintin MJ, Youngsaye W, Moccia D, Asiedu JK, Mulligan EL, Akella LB, Marcaurelle LA, Montgomery P, Bittker JA, Clemons PA, Brudz S, Dandapani S, Duvall JR, Tolliday NJ, De Souza A (2013) An informatic pipeline for managing high-throughput screening experiments and analyzing data from stereochemically diverse libraries. J Comput Aided Mol Des 27(5):455–468. doi:10.1007/s10822-013-9641-y
Murie C, Woody O, Lee AY, Nadon R (2009) Comparison of small n statistical tests of differential expression applied to microarrays. BMC Bioinformatics 10:45. doi:10.1186/1471-2105-10-45
Murie C, Barette C, Lafanechere L, Nadon R (2013) Single assay-wide variance experimental (SAVE) design for high-throughput screening. Bioinformatics 29(23):3067–3072. doi:10.1093/bioinformatics/btt538
Murie C, Barette C, Lafanechere L, Nadon R (2014) Control-plate regression (CPR) normalization for high-throughput screens with many active features. J Biomol Screen 19(5):661–671. doi:10.1177/1087057113516003
Murray CW, Rees DC (2008) The rise of fragment-based drug discovery. In: Edward Zartler E, Shapiro M (eds) Fragment-based drug discovery: a practical approach. Wiley, Hoboken. ISBN 978-0-470-05813-8
Ngo VN, Davis RE, Lamy L, Yu X, Zhao H, Lenz G, Lam LT, Dave S, Yang L, Powell J, Staudt LM (2006) A loss-of-function RNA interference screen for molecular targets in cancer. Nature 441:106–110
Nichols A (2007) High content screening as a screening tool in drug discovery. Methods Mol Biol 356:379–387
Normolle DP (1993) An algorithm for robust non-linear analysis of radioimmunoassay and other bioassays. Stat Med 12:2025–2042
Oakland J (2002) Statistical process control. Routledge, Milton Park. ISBN 0-7506-5766-9
Pereira DA, Williams JA (2007) Origin and evolution of high throughput screening. Br J Pharmacol 152:53–61
Perlman ZE, Slack MD, Feng Y, Mitchison TJ, Wu LF, Altschuler SJ (2004) Multidimensional drug profiling by automated microscopy. Science 306(5699):1194–1198
Prummer M (2012) Hypothesis testing in high-throughput screening for drug discovery. J Biomol Screen 17(4):519–529. doi:10.1177/1087057111431278
Qiu P, Simonds EF, Bendall SC, Gibbs KD Jr, Bruggner RV, Linderman MD, Sachs S, Nolan GP, Plevritis SK (2011) Extracting a cellular hierarchy from high-dimensional cytometry data with SPADE. Nat Biotechnol 29(10):886–891. doi:10.1038/nbt.1991
Ravkin I (2004) Quality measures for imaging-based cellular assays. Poster #P12024, Society for Biomolecular Screening. Annual Meeting Abstracts. http://www.ravkin.net/posters/P12024-Quality%20Measures%20for%20Imaging-based%20Cellular%20Assays.pdf. Accessed 15 Oct 2014
Reisen F, Zhang X, Gabriel D, Selzer P (2013) Benchmarking of multivariate similarity measures for high-content screening fingerprints in phenotypic drug discovery. J Biomol Screen 18(10):1284–1297. doi:10.1177/1087057113501390
Ritz C, Streibig JC (2005) Bioassay analysis using R. J Stat Softw 12(5):1–22. http://www.jstatsoft.org/
Root DE, Hacohen N, Hahn WC, Lander ES, Sabatini DM (2006) Genome-scale loss-of-function screening with a lentiviral RNAi library. Nat Methods 3(9):71. doi:10.1038/NMETH92
Rousseeuw PJ, Croux C (1993) Alternatives to the median absolute deviation. J Am Stat Assoc 88(424):1273–1283. doi:10.2307/2291267
Sakharkar MK, Sakharkar KR, Pervaiz S (2007) Druggability of human disease genes. Int J Biochem Cell Biol 39:1156–1164
Sebaugh JL (2011) Guidelines for accurate EC50/IC50 estimation. Pharm Stat 10:128–134
Sharma S, Rao A (2009) RNAi screening – tips and techniques. Nat Immunol 10(8):799–804
Shewhart WA (1931) Economic control of quality of manufactured product. Van Nostrand, New York. ISBN 0-87389-076-0
Shun TY, Lazo JS, Sharlow ER, Johnston PA (2011) Identifying actives from HTS data sets: practical approaches for the selection of an appropriate HTS data-processing method and quality control review. J Biomol Screen 16(1):1–14. doi:10.1177/1087057110389039
Sims D, Mendes-Pereira AM, Frankum J, Burgess D, Cerone MA, Lombardelli C, Mitsopoulos C, Hakas J, Murugaesu N, Isacke CM, Fenwick K, Assiotis I, Kozarewa I, Zvelebil M, Ashworth A, Lord CJ (2011) High-throughput RNA interference screening using pooled shRNA libraries and next generation sequencing. Genome Biol 12(R104):1–13
Singh S, Carpenter AE, Genovesio A (2014) Increasing the content of high-content screening: an overview. J Biomol Screen 19(5):640–650. doi:10.1177/1087057114528537
Sittampalam GS, Iversen PW, Boadt JA, Kahl SD, Bright S, Zock JM, Janzen WP, Lister MD (1997) Design of signal windows in high-throughput screening assays for drug discovery. J Biomol Screen 2:159
Sittampalam GS, Gal-Edd N, Arkin M, Auld D, Austin C, Bejcek B, Glicksman M, Inglese J, Lemmon V, Li Z, McGee J, McManus O, Minor L, Napper A, Riss T, Trask OJ, Weidner J (eds) (2004) Assay guidance manual. Eli Lilly & Company and the National Center for Advancing Translational Sciences, Bethesda. http://www.ncbi.nlm.nih.gov/books/NBK53196/
Smith K, Horvath P (2014) Active learning strategies for phenotypic profiling of high-content screens. J Biomol Screen 19(5):685–695
Smyth GK (2004) Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3(1):1–26
Storey JD (2002) A direct approach to false discovery rates. J R Stat Soc Ser B 64:479–498
Storey JD, Tibshirani R (2003) Statistical significance for genome-wide experiments. Proc Natl Acad Sci 100:9440–9445
Street JO, Carrol RJ, Ruppert D (1988) A note on computing robust regression estimates via iteratively reweighted least squares. Am Stat 42:152–154
Strimmer K (2008a) A unified approach to false discovery rate estimation. BMC Bioinformatics 9:303. doi:10.1186/1471-2105-9-303
Strimmer K (2008b) fdrtool: a versatile R package for estimating local and tail area-based false discovery rates. Bioinformatics 24(12):1461–1462
Sui Y, Wu Z (2007) Alternative statistical parameter for high-throughput screening assay quality assessment. J Biomol Screen 12(2):229–234
Sun D, Whitty A, Papadatos J, Newman M, Donnelly J, Bowes S, Josiah S (2005) Adopting a practical statistical approach for evaluating assay agreement in drug discovery. J Biomol Screen 10(5):508–516
Sun D, Jung J, Rush TS, Xu Z, Weber MJ, Bobkova E, Northrup A, Kariv I (2010) Efficient identification of novel leads by dynamic focused screening: PDK1 case study. Comb Chem High Throughput Screen 13(1):16–26
Taylor PB, Stewart FP, Dunnington DJ, Quinn ST, Schulz CK, Vaidya KS, Kurali E, Lane TR, Xiong WC, Sherrill TP, Snider JS, Terpstra ND, Hertzberg RP (2000) Automated assay optimization with integrated statistics and smart robotics. J Biomol Screen 5(4):213–226
Thorne N, Auld DS, Inglese J (2010) Apparent activity in high-throughput screening: origins of compound-dependent assay interference. Curr Opin Chem Biol 14(3):315–324. doi:10.1016/j.cbpa.2010.03.020
Tong T, Wang Y (2007) Optimal shrinkage estimation of variances with applications to microarray data analysis. J Am Stat Assoc 102(477):113–122. doi:10.1198/01621450600000126
Tusher VG, Tibshirani R, Chu G (2001) Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci 98(9):5116–5121
van Oostrum J, Calonder C, Rechsteiner D, Ehrat M, Mestan J, Fabbro D, Voshol H (2009) Tracing pathway activities with kinase inhibitors and reverse phase protein arrays. Proteomics Clin Appl 3(4):412–422
Varin T, Gubler H, Parker CN, Zhang JH, Raman P, Ertl P, Schuffenhauer A (2010) Compound set enrichment: a novel approach to analysis of primary HTS data. J Chem Inf Model 50(12): 2067–2078. doi:10.1021/ci100203e
Wu Z, Liu D, Sui Y (2008) Quantitative assessment of hit detection and confirmation in single and duplicate high-throughput screenings. J Biomol Screen 13(2):159–167. doi:10.1177/1087057107312628
Wunderlich ML, Dodge ME, Dhawan RK, Shek WR (2011) Multiplexed fluorometric immunoassay testing methodology and troubleshooting. J Vis Exp 12(58):pii:3715
Yin Z, Zhou X, Bakal C, Li F, Sun Y, Perrimon N, Wong ST (2008) Using iterative cluster merging with improved gap statistics to perform online phenotype discovery in the context of high-throughput RNAi screens. BMC Bioinformatics 9:264. doi:10.1186/1471-2105-9-264
Zhang XD (2008) Novel analytic criteria and effective plate designs for quality control in genome-scale RNAi screens. J Biomol Screen 13:363–377
Zhang XD (2011a) Optimal high-throughput screening: practical experimental design and data analysis for genome-scale RNAi research. Cambridge University Press, Cambridge. ISBN 978-0-521-73444-8
Zhang XD (2011b) Illustration of SSMD, Z Score, SSMD*, Z* score and t statistic for hit selection in RNAi high-throughput screening. J Biomol Sreen 16(7):775–785
Zhang XD, Zhang Z (2013) displayHTS: a R package for displaying data and results from high-throughput screening experiments. Bioinformatics 29(6):794–796. doi:10.1093/bioinformatics/btt060
Zhang JH, Chung TD, Oldenburg KR (1999) A simple statistical parameter for use in evaluation and validation of high throughput screening assays. J Biomol Screen 4:67–73
Zhang JH, Chung TD, Oldenburg KR (2000) Confirmation of primary active substances from high throughput screening of chemical and biological populations: a statistical approach and practical considerations. J Comb Chem 2(3):258–265
Zhang JH, Wu X, Sills MA (2005) Probing the primary screening efficiency by multiple replicate testing: a quantitative analysis of hit confirmation and false screening results of a biochemical assay. J Biomol Screen 10:695. doi:10.1177/1087057105279149
Zhang XD, Ferrer M, Espeseth AS, Marine SD, Stec EM, Crackower MA, Holder DJ, Heyse JF, Strulovici B (2007) The use of strictly standardized mean difference for hit selection in primary RNA interference high-throughput screening experiments. J Biomol Screen 12(4):497–509. doi:10.1177/1087057107300646
Zhang XD, Kuan PF, Ferrer M, Shu X, Liu YC, Gates AT, Kunapuli P, Stec EM, Xu M, Marine SD, Holder DJ, Strulovici B, Heyse JF, Espeseth AS (2008) Hit selection with false discovery rate control in genome-scale RNAi screens. Nucleic Acids Res 36(14):4667–4679. doi:10.1093/nar/gkn435
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Gubler, H. (2016). High-Throughput Screening Data Analysis. In: Zhang, L. (eds) Nonclinical Statistics for Pharmaceutical and Biotechnology Industries. Statistics for Biology and Health. Springer, Cham. https://doi.org/10.1007/978-3-319-23558-5_5
Download citation
DOI: https://doi.org/10.1007/978-3-319-23558-5_5
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-23557-8
Online ISBN: 978-3-319-23558-5
eBook Packages: Mathematics and StatisticsMathematics and Statistics (R0)