Abstract
Alcohol consumption has been identified in epidemiological studies as a risk factor for cancer of the esophagus, pharynx, larynx, and oral cavity (collectively known as upper aerodigestive tract, UADT) and breast, as well as liver cancer. While inconsistent relationship between alcohol drinking and UADT cancer mortality was reported, heavy alcohol consumption could increase UADT cancer mortality, especially if combined with tobacco use. Epidemiological studies on alcohol and breast cancer are also inconsistent. The variability in outcomes in alcohol and cancer epidemiological studies stems from: (1) studies rely on self-report to determine the amount of alcoholic beverage consumed, which introduces “recall bias”; (2) cancer develops over a long period and epidemiological studies that capture current or recent alcohol consumption after clinical diagnosis cannot correlate that with carcinogenesis; (3) cancer is a multifactorial disease that culminates by complex interactions of numerous risk factors—genetic and environmental—over time. Thus, epidemiological studies cannot determine cause-and-effect relationships and are unable to give biological and clinical insights into carcinogenesis. This review focuses on hepatocellular carcinoma (HCC) and enumerates known plausible mechanisms of hepatocarcinogenesis.
Chronic heavy drinking invariably results in a spectrum of alcoholic liver disease terminating in cirrhosis in only ~10 %. Among patients with cirrhosis, about 1–2 % develops HCC. Other comorbid conditions such as HBV or HCV infection, obesity, or hemochromatosis and lifestyle factors such as smoking or ingestion of food contaminated with aflatoxin can exacerbate liver damage and amplify HCC.
This review discusses plausible mechanisms for HCC genesis due to heavy alcohol consumption including: (1) alcohol metabolism, which results in acetaldehyde generation and formation of DNA adducts, reactive oxygen species formation, changes in redox state of hepatocytes, and derangement of metabolic pathways, especially retinoic acid synthesis and transport; (2) epigenetic effects of alcohol especially on the DNA methylation, histone modification, and miRNA availability and function; (3) circadian rhythm perturbation; (4) immune system modifications; and (5) angiogenesis.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Alcohol metabolism
- Cirrhosis
- Hepatocellular carcinoma
- Metabolism
- Epigenetics
- Reactive oxygen species
- Retinoic acid
- Oxidative stress
- Immune dysfunction
- Circadian rhythm
- Angiogenesis
Introduction
While epidemiological studies point to an increased risk of various cancers associated with heavy alcohol consumption (cancers of the upper aerodigestive tract [oral cavity, pharynx, larynx, and esophagus], colorectum, liver, and female breast) and decreased risk of other cancers (renal cancer and non-Hodgkin’s lymphoma), these studies cannot determine cause and effect. While heavy drinking may be linked to certain cancers, the ultimate outcome could be due to alcohol consumption alone, to its interactions with other lifestyle factors such as smoking, or to the presence of comorbid conditions such as obesity or viral hepatitis. Various cancers are tissue specific and comprise a multistage and complex process characterized by molecular alterations that underlie their initiation, promotion, and progression over a long time. Many hypotheses have been spawned to explain plausible mechanisms by which heavy drinking may be linked to carcinogenesis. Chronic heavy ethanol (hereinafter referred to as alcohol) consumption is a risk factor for cancer of the esophagus and oral cavity and an etiological factor in liver cancer [1]. This article focuses mainly on alcohol-associated hepatocellular carcinoma (HCC) and briefly discusses other cancers attributed to heavy alcohol consumption.
Hepatocellular carcinoma comprises approximately 85 % of primary liver cancer and is the third leading cause of cancer deaths worldwide. In 2012, it resulted in approximately ¾ million deaths globally [2]. HCC usually occurs as a consequence of chronically damaged livers due to cirrhosis, chronic infection with hepatitis B (HBV) and C (HCV) viruses, chronic heavy alcohol consumption, or cirrhosis associated with nonalcoholic steatohepatitis and primary hemochromatosis [3]. Other established causes of liver cancer include contraceptives high in estrogen and progesterone, smoking, obesity, and ingestion of food contaminated with fungal aflatoxin in subtropical regions.
Hepatocellular Carcinoma
While moderate alcohol consumption by adults could be a part of a healthy lifestyle for a large segment of the population [4], chronic heavy drinking invariably results in a spectrum of alcoholic liver disease (ALD) ranging from fatty liver (steatosis) in the majority of excessive drinkers to steatohepatitis and fibrosis [5] in about 35 % and only about 10 % progress to cirrhosis [6]. Among patients with cirrhosis, about 1–2 % develops HCC [7]. In addition, heavy alcohol consumption may act synergistically with HBV or HCV infection or obesity to induce HCC. These factors have some common features/mechanisms that exacerbate liver damage when they coexist [8]. The molecular aberrations in HCC pathogenesis are elegantly reviewed elsewhere [9, 10].
Mechanisms of Alcohol-Induced Hepatocarcinogenesis
Alcohol Metabolism
Ingested alcohol is readily absorbed from the gastrointestinal tract. Over 90 % of absorbed alcohol is metabolized mainly by oxidative pathways in the liver and to a small extent by non-oxidative pathways in extrahepatic tissues. Although alcohol metabolism is often considered a predominant factor in causing alcohol-associated liver damage, other factors, such as inflammatory cytokines, immunologic and metabolic pathway derangements, effects on signal transduction, proteasome inhibition, increased gut leakiness and LPS absorption, activation of Kupffer and hepatic stellate cells, genetic and epigenetic factors, etc., contribute to ALD.
The major pathway of oxidative metabolism of alcohol in the liver involves multiple isoforms of cytosolic alcohol dehydrogenase (ADH) , which results in the production of acetaldehyde. The cytochrome P450 isozymes , including CYP2E1, 1A2, and 3A4, which are predominantly localized to the endoplasmic reticulum (ER), also contribute to alcohol’s oxidation to acetaldehyde in the liver. CYP2E1 is induced by chronic alcohol consumption and assumes an important role in metabolizing alcohol to acetaldehyde at elevated alcohol concentration. Accumulation of acetaldehyde , a highly reactive and toxic molecule, contributes to liver damage. The oxidation of alcohol is accompanied by the reduction of NAD+ to NADH and, thereby, generates a highly reduced cytosolic environment in hepatocytes. It also produces highly reactive oxygen species (ROS) , including hydroxyethyl, superoxide anion, and hydroxyl radicals. Another enzyme, catalase, located in peroxisomes, is capable of oxidizing alcohol in the presence of a hydrogen peroxide (H2O2)-generating system, such as NADPH oxidase or xanthine oxidase. Quantitatively, however, this is considered a minor pathway of alcohol oxidation.
Acetaldehyde, produced by alcohol oxidation through any of these enzymes, is rapidly metabolized mainly by mitochondrial aldehyde dehydrogenase (ALDH2), and to a small extent by cytosolic aldehyde dehydrogenase (ALDH1), to form acetate and NADH. Mitochondrial NADH is reoxidized by the electron transport chain. Most of the acetate resulting from alcohol metabolism escapes the liver into the bloodstream and is eventually metabolized to CO2 by way of the tricarboxylic acid (TCA) cycle in cells with mitochondria capable of converting acetate to the metabolically active intermediate acetyl-CoA. This occurs primarily in tissues such as heart, skeletal muscle, and brain (Fig. 13.1).
Consequences of Alcohol Metabolism by Oxidative Pathways: Cancer Implication
Alcohol metabolism in the liver results in various products/effects with implications for hepatocarcinogenesis.
Acetaldehyde Generation/Adduct Formation
Acetaldehyde produced by alcohol oxidation, if accumulated to appreciable concentrations, can form adducts with DNA and RNA and decrease DNA repair. Acetaldehyde also has the capacity to react with lysine residues on proteins including enzymes, microsomal proteins, and microtubules and affect their function. Formation of protein adducts in hepatocytes may contribute to impaired protein secretion, resulting in hepatomegaly. In addition, there is evidence that acetaldehyde and malondialdehyde (a by-product of lipid peroxidation) can combine and react with lysine residues on proteins, giving rise to stable malondialdehyde–acetaldehyde (MAA) protein adducts that can be immunogenic and, thus, can contribute to immune-mediated liver damage. Also, MAA adducts have proinflammatory and profibrogenic properties.
The most relevant acetaldehyde adducts that impact the genome function and have implications to carcinogenesis are their interaction with the exocyclic amino group of deoxyguanosine to form DNA adducts (Fig. 13.2). These adducts involve the reaction of one molecule of acetaldehyde with DNA to form N2-ethylidenedeoxyguanosine, which is relatively unstable and abundant in human liver even in the absence of exogenous acetaldehyde [11]. This adduct is reduced in vivo with glutathione or vitamin C to form the stable N2-ethyldeoxyguanosine (Et-dG). In addition, two molecules of acetaldehyde, or crotonaldehyde, form an adduct known as N2-propano-2′-deoxyguanosine [12], which is maintained at a low steady state by DNA repair. A secondary acetaldehyde-related DNA adduct, 1,N2-etheno-dG, is formed from acetaldehyde-stimulated lipid peroxidation. For a detailed discussion of these adducts and their genotoxic effects, the reader is referred to the review article by Brooks and Zakhari [13].
It should be noted that some results obtained from ADH1B polymorphisms do not concord with the acetaldehyde hypothesis. For example, a decreased UADT cancer risk was observed in drinkers who carried the ADH1B*2 allele that codes for the more active enzyme, leading to high acetaldehyde exposure [14].
Formation of Reactive Oxygen Species, Reactive Nitrogen Species, and Oxidative Stress
Hepatic mitochondria produce ROS through the activity of the electron transport chain (ETC) as a by-product of oxidative phosphorylation. Normally, a small fraction of electrons entering the ETC can prematurely escape from complexes I and III and directly react with ~1–3 % of respiratory oxygen molecules to generate the superoxide anion radical, which is then dismutated by the mitochondrial manganese superoxide dismutase (MnSOD) into hydrogen peroxide (H2O2). Mitochondrial glutathione peroxidase (GPx) then converts H2O2 into water by using reduced glutathione (GSH) as a cofactor. Thus, most of the ROS generated by the ETC in the normal state are detoxified by the mitochondrial antioxidant defenses. The non-detoxified portion of ROS diffuses out of mitochondria and affects signal transduction pathways and gene expression, triggering cytokines, hormones, and growth factors, which if excessive may lead to hepatic inflammation, necrosis, and/or apoptosis. In addition, metals (e.g., iron and copper) can further react with H2O2 to produce hydroxyl radicals via the Fenton reaction. Nitric oxide (NO) , a reactive nitrogen species critical for hepatocyte biology, can interact with peroxides to generate peroxynitrite (ONOO−), which, depending on the amount and duration, could be detrimental to the liver (Table 13.1). NO is produced from l-arginine and oxygen by iNOS, which is expressed in all liver cells (hepatocytes, stellate cells, Kupffer cells, and vascular endothelial cells), and its expression is induced by IL-1β alone or in combination with TNF-α, IFNγ, and/or LPS. Alcohol not only produces ROS and reactive nitrogen species (RNS), but also depletes antioxidants in cells resulting in “oxidative stress.” This condition regulates both genetic and epigenetic cascades underlying altered gene expression in human cancer [15].
It has been suggested that ROS participate in tumor progression by promoting DNA damage and/or altering cellular signaling pathways [16]. A reliable biomarker of oxidative stress and ROS-induced carcinogenesis is 8-oxo-7,8-dihydroguanine (8-oxoGua), which is strongly implicated in all stages of carcinogenesis [17]. Formation of 8-oxoGua lesions has been shown to induce DNA base mutations in the TP53 tumor suppressor gene in liver cancer cells [18]. Elevated levels of 8-OHdG in transgenic mice infected with HBV can lead to development of hepatocellular carcinoma [19]. In addition, oxidative stress often renders repair mechanisms ineffective. ROS have been reported to promote hypermethylation of the promoter region of the tumor suppressor E-cadherin, a regulator of the epithelial-to-mesenchymal transition, in HCC cells [20]. ROS induce hypermethylation of the E-cadherin promoter by increasing Snail expression. Alcohol promotes breast and colon cancer progression through stimulating the EMT program via a Snail-mediated pathway [21] and may have a similar effect in HCC. Exacerbating oxidative stress in livers infected with HCV by ROS induction and by hampering the antioxidant system facilitates hepatocarcinogenesis [22]. Ironically, increases in Nrf2 protein, a transcription factor that regulates important antioxidant and phase II detoxification genes, were observed in hepatocytes of alcohol-fed mice, suggesting that Nrf2 plays a key role in the adaptive response against increased oxidative stress caused by CYP2E1 [23].
Increase in NADH/NAD+ Ratio
Alcohol metabolism produces a significant increase in the hepatic NADH/NAD+ ratio in both the cytosol and the mitochondria, as evidenced by an increase in the lactate/pyruvate and β-hydroxybutyrate/acetoacetate ratios, respectively [24] Consequently, alcohol oxidation vastly increases the availability of oxidizable NADH to the electron transport chain in the mitochondria. The liver responds to alcohol exposure in part by increasing the rate of oxygen uptake, which may lead to periods of hypoxia, particularly in the downstream (pericentral) parts of the liver lobule. Increased NADH/NAD+ ratio provides reducing equivalents and thus enhances the activity of the respiratory chain, including heightened oxygen use and ROS formation [25]. Furthermore, the increase in the NADH/NAD+ ratio results in derangement of carbohydrate metabolism and modulation of gene expression of, among others, SIRT1, a NAD+-dependent deacetylase [26] whose substrates include histones and the transcription factor p53 [27]. Increased NADH in hepatocytes due to alcohol metabolism may promote tumor growth by favoring the generation of lactate through a NADH-dependent enzyme, lactate dehydrogenase A (LDH-A), which catalyzes the conversion of pyruvate to lactate during glycolysis. Tumor cells utilize more glucose than normal tissue, favor aerobic glycolysis, and rely on lactate production for their survival. A molecular mechanism underlying the enhanced lactate production in cancer cells involves tyrosine phosphorylation which enhances LDH-A enzyme activity to promote tumor growth by regulating the NADH/NAD redox homeostasis [28].
Derangement of Metabolic Pathways
Increased NADH/NAD+ ratios in both the cytosol and mitochondria of hepatocytes influence the direction of several reversible reactions leading to alterations in hepatic lipid, carbohydrate, protein, lactate, and uric acid metabolism. These changes include (1) alcoholic hypoglycemia , the increase in NADH prevents pyruvate conversion to glucose by lowering the concentration of pyruvate, which in turn decreases the pyruvate carboxylase reaction, one of the rate-limiting steps of gluconeogenesis; (2) hampering of the tricarboxylic acid (TCA) cycle function , the increase in mitochondrial NADH in hepatocytes contributes to the saturation of NADH dehydrogenase; and (3) alcoholic acidosis, ketoacidosis is common in chronically malnourished alcoholics and is due to the formation of ketone bodies, primarily β-hydroxybutyrate [29]. In addition, the increase in NADH favors the conversion of pyruvate to lactate, resulting in lactic acidosis. The increase in NADH/NAD+ ratio diminishes pyruvate dehydrogenase (PDH) activity in the mitochondria, resulting in diminished conversion of pyruvate to acetyl-CoA. PDH activity is further diminished in chronic alcoholics due to hypomagnesemia and thiamine deficiency, resulting in the inhibition of pyruvate utilization in the TCA cycle; (4) hypoxia, alcohol metabolism by hepatocytes tends to increase oxygen uptake, resulting in significant hypoxia in the perivenous hepatocytes, the site of early liver damage due to chronic alcohol consumption.
Perhaps the derangement most relevant to HCC is alcohol impairment of retinoic acid (RA) synthesis and transport. Alcohol dramatically changes vitamin A and RA availability [30] and thus can impact carcinogenesis [31]. RA deficiency in alcoholics results from poor dietary intake and decreased absorption of retinoids and the significant overlap in metabolic pathways of alcohol and retinol, the alcohol form of vitamin A. Alcohol and retinol can be oxidized by similar, and sometimes identical, enzymes. In addition to the competitive inhibition of RA biosynthesis, prolonged alcohol consumption decreases tissue RA concentrations by enhancing its catabolism through the induction of cytochrome P450 enzymes and increasing mobilization of retinoids from the liver to extrahepatic tissues [32]. As a result, alcohol profoundly depletes hepatic retinoids and alters their distribution in other tissues [33, 34]. In addition to directly affecting RA metabolism and transport, alcohol may also affect plasma retinol concentration and its organ distribution indirectly through LPS-induced inflammation that reduces the level of RBP mRNA in the liver resulting in the impairment of the transport of retinol from the liver to plasma [35].
Changes in RA availability due to dysregulation of retinoid transport by alcohol are also becoming more evident [1]. It is strongly linked to alterations in differentiation/proliferation status of hepatocytes.
RA acts as a signaling molecule and regulates gene expression by binding to two subclasses of nuclear receptors, retinoic acid receptors (RARα, β, and γ isotypes) and retinoid X receptors (RXRα, β, and γ isotypes), encoded by distinct genes [36]. Alcohol affects the expression and activation of RA receptors, which in turn can impair the signaling events and induce harmful effects on cell survival and differentiation [37]. Recent developments indicate that alcohol can contribute to the aberrancy of retinoid nuclear receptor function and increased risk of cancer development through epigenetic alterations [38]. Alterations in the level of expression or functional activity of retinoid nuclear receptors are associated with a variety of cancers despite normal vitamin A levels. Hepatic retinoid level reduction by alcohol can lead to enhanced fibrogenesis that, in turn, may eventually constitute an irreversible process with regenerative diffuse parenchymal nodular transformation, cirrhosis, and HCC. Alcohol-related HCCs are associated with cirrhosis in a majority of cases, indicating that the pathological events leading to cirrhosis precede those causing cancer or that the structural alterations of cirrhosis favor hepatocyte dedifferentiation [39]. Since dedifferentiation is ultimately associated with increased proliferation rate, the dedifferentiation hypothesis fits perfectly with findings that low hepatic RA concentration due to alcohol leads to an upregulation of AP-1 (c-jun and c-fos) and beta-catenin-dependent gene expression that may promote proliferation and malignant transformation of hepatocytes by alcohol [34, 40]. Interestingly, alcohol-induced RA-dependent hepatocyte hyperproliferation may not only lead to the neoplastic transformation of preexisting hepatocytes but may also compromise organ regeneration by liver stem cells. The activation of liver stem cells requires a uniform inhibition of parenchymal proliferation [41] and their differentiation depends on RA [42, 43]. It is interesting to note that liver stem cells are highly responsive to vitamin A deprivation [44] and always appear in close proximity to activated hepatic stellate cells (HSC) suggesting a possible involvement of RA in control of their behavior. Deficiency of RA in alcoholic livers blocks differentiation and apoptosis in the progeny of liver stem cells, while promoting their proliferation. This may explain the development of anaplastic poorly differentiated HCCs without preexisting cirrhosis, which are also observed in alcoholics, albeit rarely [45, 46].
Variations in Metabolic Enzymes
Class I ADH and ALDH2 play a central role in alcohol metabolism. Allelic variations in the genes encoding ADH and ALDH produce alcohol- and acetaldehyde-metabolizing enzymes that vary in activity. These genotypes modify the susceptibility to tissue damage. The ADH gene family encodes for enzymes that metabolize various substrates, including retinol, and are differentially expressed in different organs. This highlights the important issue of substrate competition in alcohol-induced tissue damage. Genetic polymorphism occurs at the ADH1B and ADH1C loci [47] with different catalytic activities for alcohol. The ADH1B alleles occur at different frequencies in different populations. For example, the ADH1B*1 form is found predominantly in Caucasian and Black populations, while ADH1B*2 frequency is higher in Chinese and Japanese populations and in 25 % of people with Jewish ancestry. A significant interaction exists between ADH1B polymorphism and heavy alcohol consumption especially for those with ADH1B*1/*1 genotype and esophageal [48] and UADT [49] cancer. Several isozymes of ALDH have been identified, but only the cytosolic ALDH1 and the mitochondrial ALDH2 metabolize acetaldehyde. There is one significant genetic polymorphism of the ALDH2 gene, resulting in allelic variants ALDH2*1 and ALDH2*2 (glutamine to lysine substitution at position 487, resulting in 100-fold increase in the Km for NAD+, making the gene product virtually inactive). The low activity ALDH2*2 is a deficient phenotype, which is present in about 50 % of the Taiwanese, Han Chinese, and Japanese populations [50] and shows virtually no acetaldehyde-metabolizing activity in vitro. The activity of ADH and ALDH isozymes contributes to alcohol-induced tissue damage. Alcoholic cirrhosis is reduced over 70 % in populations carrying the ALDH2*2 allele [51]. The activities of class I ADH are much higher in cancerous than in healthy tissues [52], and individuals with the ADH1C*1 allele have an increased risk to develop breast cancer from alcohol [53]. Furthermore, ALDH2-deficient individuals are at much higher risk of esophageal cancer (specifically squamous cell carcinoma) from alcohol consumption than individuals with fully active ALDH2 [54]. The correlation between genetic polymorphism of ADH and ALDH and esophageal, head, and neck cancers was reviewed by Yokoyama and Omori [55] and summarized in Table 13.2.
Although several CYP2E1 polymorphisms have been identified, only a few studies were undertaken to determine the effect on alcohol metabolism and tissue damage. In one study, the presence of the rare c2 allele was associated with higher alcohol metabolism in Japanese alcoholics but only at high blood alcohol concentrations of 0.25 g/dL [56]. In addition, induction of CYP2E1 also contributes to carcinogenesis through activation of pro-carcinogens, such as nitrosamines present in diets and in tobacco smoke, to their carcinogenic metabolites [57]. The correlations between genetic polymorphisms and risk of alcohol-related cancers are reviewed elsewhere [58].
Epigenetic Modifications
Functional genomic studies (GWAS, whole-genome sequencing, global DNA copy numbers and methylation, and gene or noncoding RNA expression profiling) revealed that genetic polymorphisms of immune-related genes, such as IL28B and MHC class I and II molecules (e.g., MICA), and somatic mutations of TP53 and ARID2 (a novel liver cancer-related gene that encodes a component of the SWI/SNF chromatin-remodeling complex) as well as activated β-catenin mutations are associated with HCC initiation and progression [59]. In addition, epigenetic mechanisms play an important role during the development and progression of HCC, and numerous studies have identified a large number of genes and pathways that are subject to epigenetic dysregulation [60]. Global DNA hypomethylation, histone modifications, promoter methylation, aberrant expression of noncoding RNAs, and dysregulated expression of epigenetic regulatory genes such as EZH2 are the best-known epigenetic abnormalities [61]. As mentioned above, alcohol metabolism alters the ratio of NAD+ to NADH and promotes the formation of ROS and acetate, all of which impact epigenetic regulatory mechanisms. Furthermore, the activities of enzymes involved in epigenetic modifications, such as DNA and histone methylation and histone acetylation, are influenced by the levels of metabolites such as NAD+, adenosine triphosphate (ATP), and S-adenosylmethionine (SAM). Chronic alcohol consumption leads to significant reductions in SAM levels, thereby contributing to DNA hypomethylation. These epigenetic changes are discussed in detail elsewhere [62] and are briefly mentioned below.
Epigenetic Effects
Since only about 10 % of heavy drinkers develop cirrhosis, the complex biological processes underlying states of health and disease could be determined by interactions between many genes and the external environment, and are likely to be driven by both genetic defects and by modifications that affect the transcriptional capacity of these genes (epigenetics). Epigenetic changes (e.g., DNA methylation or histone modification) affect gene expression directly or through the way DNA is packaged into chromatin, thus altering accessibility to transcription factors. In general, methylation of DNA represses gene expression by changing the chromatin structure or by interfering with the binding of some transcription factors to the promoter. DNA methylation has been shown to play a critical role in many cellular and biological processes, including cancer, aging, development, and the maintenance and differentiation of stem cells [63]. The successful establishment and maintenance of transcriptional profiles depend on the interplay between epigenetic modifications, interacting proteins, noncoding RNAs, and inter- and intrachromosomal interactions. Perturbation of any one of these regulatory elements may have profound consequences on the liver.
Chronic alcohol consumption has been shown to affect epigenetic regulation of gene expression involving DNA methylation, histone modification, and RNA-mediated gene silencing, thus modulating the expression of many genes. Alcohol-induced changes in gene expression may affect various biochemical and signaling pathways influencing the function of cells and organs, leading to liver disease and even cancer. Understanding alcohol-induced epigenetic regulation will provide mechanistic insights, diagnostic biomarkers, and therapeutic targets for alcohol-related liver injury.
Alcohol and DNA Methylation
DNA methylation tags cytosine, one of the four chemical bases that make up the genetic code, with a methyl group by transferring a methyl group from SAM onto the cytosine residue, which protrudes into the major groove of the DNA. Although acetaldehyde, the first metabolite of alcohol, can form DNA adducts and may cause sequence alteration of DNA, most of the short or long-lasting effects of alcohol may be independent of DNA sequence changes. Recent studies have shown that epigenetic regulation of gene expression is an important mechanism for alcohol’s action in the cell and alcohol-induced liver damage [38].
Alcohol-induced epigenetic changes may also affect stem cell differentiation and liver repair and regeneration. These processes are characterized by rapid, well-synchronized patterns of gene expression in which the status of DNA methylation shifts dramatically, involving both loss of methylation and de novo methylation. Many epigenetic changes during stem cell differentiation involve genes known to function in cell cycle, growth, apoptosis, and oxidative stress, all of which play a critical role in alcohol-induced liver damage and cancer. Evidently, purely sequence-based genetic or genomic approaches to study gene regulation are not sufficient to explain alcohol-related HCC.
Effects of Alcohol on the Availability and Transfer of Methyl Groups
Chronic alcohol consumption is associated with abnormal methionine metabolism, increased plasma homocysteine level, decreased level of SAM, and folate deficiency [5, 64]. SAM, the major methyl donor for DNA methylation, is primarily generated in the liver from l-methionine and ATP by methionine adenosyltransferase (MAT), which is encoded by two genes, MAT1A and MAT2A. MAT1A encodes the isoenzymes MATI and MATIII, whereas MAT2A gene encodes the isoenzyme MATII. MATI and MATIII are primarily responsible for maintaining high intracellular SAM levels in adult liver, while MATII is predominantly active in fetal and regenerating liver tissues.
Alcohol impairs the transfer of methyl groups to the cytosine residues of DNA by reducing the levels and activity of DNA methyltransferases (DNMT) , resulting in DNA hypomethylation. The alcohol metabolite acetaldehyde can also inhibit DNMT activity. Studies have shown that livers of MAT1A knockout mice had SAM deficiency and increased expression of genes involved in proliferation and consequently developed hepatomegaly, fatty liver, and eventually HCC. They also regenerated abnormally after partial hepatectomy and were more sensitive to developing steatosis in response to a methionine- and choline-deficient diet [65]. It has been reported that during hepatocarcinogenesis or hepatectomy, MAT1A itself is severely downregulated due to the hypermethylation of its promoter [5]. These observations suggest that alcohol may contribute to HCC development via its inhibition of MATI and MATIII activity as well as reducing SAM levels [66].
In addition to its effects on MAT and SAM synthesis, alcohol also inhibits a number of methyl group transfer-related enzymes such as methionine synthase and cystathionine-β-synthase. The latter removes homocysteine through trans-sulfuration to cystathionine, which is used to generate the antioxidant glutathione (GSH). The net result is a decreased level of GSH, leading to increased oxidative stress, also contributing to liver damage. Excessive alcohol intake can decrease the GSH level by inhibiting hepatic GSH synthesis and the enzymatic activities involved in GSH-related peroxide detoxification such as GSH peroxidase and glutathione S-transferase (GST), thus increasing the susceptibility of the liver to oxidative injury [67].
Another major site of alcohol’s actions on methyl group transfer is the folate metabolism cycle. Chronic alcohol consumption causes malabsorption of folates and increases their renal excretion resulting in a significant decrease in hepatic folate content [68]. In addition, it inhibits methionine synthase which transfers a methyl group from 5-methyl tetrahydrofolate to homocysteine to form methionine.
Although it is clear that chronic alcohol ingestion alters availability and transfer of methyl groups, its impact on DNA methylation appears to be complex. While DNA methylation in the promoter regions of class I ADH genes is elevated in an alcohol-treated human hepatoma cell line [69], the expression of DNA methyl transferase (DNMT-3b) is decreased in alcoholic patients [70]. The effect of alcohol on DNA methylation may depend on genomic context, cell type, and target organ.
Histone Modification
Chromatin structure is dynamically regulated to selectively facilitate the expression of some genes while maintaining others in a quiescent state and to allow for DNA repair and replication. Genes within highly condensed “heterochromatin” regions are generally silenced, whereas uncondensed “euchromatin” is permissive for gene expression. The various states of chromatin are largely attributable to the posttranslational modification of histones. These occur primarily on the N-terminal histone “tails” that protrude from the nucleosome structure and include mono-, di-, or tri-lysine methylation, lysine acetylation, lysine ubiquitination, arginine methylation (mono- or di-), and serine or threonine phosphorylation, among others.
Histone acetylation is associated with loosely packed chromatin and actively transcribed genes. As depicted in Fig. 13.3, histone acetylation is determined by the opposing activities of histone acetyltransferases (HATs) and deacetylases (HDACs). Histone methylation results from the action of histone methyltransferases (HMTs) and is reversed by histone demethylases. Enzymes involved in histone acetylation are relatively few in number and promiscuous in terms of which lysines they modify. In contrast, HMTs and histone demethylases are typically specific for a single H3 or H4 residue and, consequently, are more numerous [71].
Alcohol and Histone Modification
Emerging evidence points to the potential of alcohol to exert its health effects by altering the state of chromatin. Acute alcohol administration to rats was shown to increase H3K9 acetylation in selected tissues, including liver, but not in others, indicating that epigenetic effects of alcohol will likely vary by tissue [72]. Consistent with these in vivo findings, in vitro studies found that alcohol also promotes the acetylation of H3K9 in primary hepatocyte cultures without affecting the acetylation status of other H3 lysines, including K14, K18, K23, or K27 [73]. In addition, this specific effect on H3K9 was associated with increased HAT activity by the acetate derived from alcohol metabolism. Accompanying this increase in H3K9 acetylation, there was an overall reduction of methylated H3K9 and a concomitant increase in methylated H3K4 in alcohol-treated hepatocyte cultures [74]. At the individual gene level, genes whose promoters exhibited this predominant pattern were found to be transcriptionally active, while those exhibiting the inverse pattern (i.e., increased H3K9me, decreased H3K4me) were silenced. Together, these findings highlight the potential for alcohol to alter patterns of histone modification and the expression of associated genes, raising the possibility that oncogenes and/or tumor suppressors might be among the affected genes and represent a mechanism by which alcohol contributes to HCC.
The decrease in NAD+/NADH ratio due to alcohol metabolism has the potential to diminish the activity of NAD+-dependent enzymes, including the SIRT family of histone deacetylases (HDACs). Indeed, inhibition of hepatic SIRT1 activity by alcohol was associated with an increase in the acetylated active nuclear form of SREBP-1c in the livers of alcohol-fed mice leading to impairment of lipid metabolism [75].
MicroRNAs, Cancer, and Alcohol
MicroRNAs (miRNAs) often regulate the expression of cancer pathway components, including oncogenes and tumor suppressors. While miRNAs are often globally downregulated in human tumors [76], some miRNAs are frequently dysregulated in many types of cancer. The degree to which the expression of some miRNAs is altered is correlated with clinical or pathologic indicators of malignancy, offering opportunities for early detection and insights into early pathogenesis [77].
HCC is of particular relevance to alcohol and has been the subject of several miRNA profiling studies. Murakami et al. [78] identified three miRNAs that were overexpressed in HCC (miR-224, miR-18, and pre-miR-18) and five others that were under-expressed (miR-199a, miR-199a*, miR-200a, miR-125a, miR-195). Furthermore, they demonstrated the effectiveness of a signature, based on these changes, in distinguishing HCC and non-HCC cases and identified three miRNAs (miR-92, miR-20, and miR-18) whose expression was inversely correlated with the degree of HCC differentiation. Another study defined miRNA changes that allowed differentiation of HCC (increased miR-21, miR-10b, and miR-222) from benign hepatocellular adenomas (decreased miR-200c and miR-203) [79]. Importantly, this study also revealed specific miRNA markers of alcohol-related HCC (decreased miR-126) and HCC associated with hepatitis B viral exposure (increased miR-96). The molecular pathways regulated by miRNA in HCC are detailed in a review by Milazzo and colleagues [80].
Effect of Alcohol on miRNA Expression and Function
Given the breadth of processes that miRNAs are known to regulate, it is reasonable to expect that they will play significant roles in mediating the effects of alcohol, including cancer. Several recent reports have identified miRNAs whose levels are altered by alcohol and that mediate alcohol’s ability to promote gut leakiness [81]. Chronic alcohol consumption increased miR-21 expression during liver regeneration in ethanol-fed rats [82] and enhanced miR-155 in macrophages via NF-6B, which contributed to the elevation in TNF-α production [83]. In addition, chronic alcohol feeding resulted in significant alteration in several miRNAs that regulate hepatic metabolism (miR-34a, miR-103, miR-107, and miR-122) [84], as well as downregulation of miR-199, which may contribute to HIF-1α augmentation [85]. In addition, large-scale miRNA screens indicate that alcohol alters the expression of 2–3 % of miRNAs (miR-320, miR-486, miR-705, and miR-1224 and a decreased expression for miR-27b, miR-214, miR-199a-3p, miR-182, miR-183, miR-200a, and miR-322) in murine models of alcohol-induced steatohepatitis [86]. These changes in miRNAs and their significance were reviewed elsewhere [87].
Circadian Rhythm Perturbation
The heterodimer of transcription factors CLOCK and BMAL1, which activates transcription of the period (Per) and cryptochrome (Cry) genes, comprises the centerpiece of the mammalian circadian network [88]. Their products PER and CRY interact to form the PER/CRY complex which translocates into the nucleus to inhibit CLOCK/BMAL1 transactivation, which in turn results in the repression of the Per and Cry genes. Release of PER/CRY complex through proteasome degradation can relieve repression and start the negative feedback loop again.
Despite the fact that there is no direct evidence connecting circadian rhythm disturbance to alcohol-induced HCC, dysfunctions of circadian rhythms are involved in many diseases that are known to be modulated by alcohol. The hypothesis that disturbance in circadian rhythms by alcohol may be involved in cancer is supported by the following research:
-
Disturbance in circadian rhythm genes expression is a common feature in certain types of cancer, including HCC [89]. Many studies have suggested indirectly that circadian rhythms may play an important role in hepatocarcinogenesis. Key clock genes have been found to be disrupted in HCC patients [90], demonstrating that HCC impacts the orchestrated circadian rhythm of liver cells. In addition, a long noncoding RNA (lncRNA), highly upregulated in liver cancer (HULC), has been reported to contribute to the perturbations in circadian rhythm of hepatoma cells [91]. Furthermore, a single functional polymorphism of one SNP rs2640908 in PER3 gene was significantly associated with overall survival of HCC patients [92].
-
Liver metabolism can be greatly affected by circadian rhythms and changes in feeding status. A large number of metabolic enzymes, such as CYP2E1, CYP3A4, CYP3A11, ADH, and ALDH—many of which are involved in alcohol metabolism—are regulated by the circadian clock [93]. Many other important metabolic pathways such as glycolysis, fatty-acid metabolism, cholesterol biosynthesis, and xenobiotic and intermediate metabolism are also under circadian regulation [94]. The rate-limiting steps of these metabolic pathways are often the target sites of circadian control. Because of their important roles in metabolism, mutations in the clock genes often cause metabolic disorders. For example, a mutation of the Clock gene caused mice to be hyperphagic and obese, exhibiting hyperlipidemia, hepatic steatosis, hyperglycemia, and hypoinsulinemia [95]. Conversely, circadian clocks can be entrained by various metabolism-related external cues, such as food intake and alcohol consumption. A high-fat diet in mice can change the expression of clock genes and clock-controlled genes [96], delay the circadian expression of adiponectin signaling components, and inhibit AMPK expression in mouse liver [97]. Furthermore, some nuclear receptors, such as PPARα, PPARγ, glucocorticoid receptor, RARα, and RXRα, have been directly connected to the key components of the circadian system [98] and have been also implicated in alcohol-induced tissue injury.
-
Some biological pathways closely related to alcohol’s actions are involved in, or affected by, circadian rhythms. The redox state of cells plays an important role in the function of the circadian rhythm. The presence of NADH or NADPH promotes the binding of the heterodimeric clock transcription factor complexes to DNA [99]. In addition, studies suggest that the histone deacetylase SIRT1 may be involved in the integration of circadian and metabolic transcription networks. It has been shown that SIRT1 interacts directly with CLOCK and deacetylates BMAL1 and PER2. Malondialdehyde, a marker of oxidative stress affected by alcohol, is found to exhibit circadian patterns of expression in mice liver [100]. The retinoic acid receptors RXR and RAR can interact with CLOCK and NPAS2 (a homolog of CLOCK), and these interactions can be increased 15-fold by retinoic acid [101]. Other consideration is that circadian rhythm disturbances lead to immunodeficiency [102], suppression of natural killer cell activity, and alteration in the T-helper 1/T-helper 2 cytokine balance, resulting in decreases in cellular immunity and tumor immune surveillance [103], all of which are affected by alcohol.
-
Lipopolysaccharide (LPS) , which is increased in blood after alcohol consumption, suppresses clock genes, suggesting that circadian rhythms play an important role in response to systemic inflammatory stimulation [104].
The relative contributions of disrupted circadian rhythm and circadian genes to cancer risk may be informative as to the pathogenesis of various cancers and new treatment interventions.
Immune Modification
The first line of host defense against HCC is innate immunity, including natural killer (NK) cells and T lymphocytes. The involvement of the immune system in HCC carcinogenesis has been previously proposed in clinical studies, where the percentage and absolute number of NK cells were decreased significantly during the development and progression of HCC [105]. Effective adaptive immune response depends on the antigen-specific activation of T and B cells. Increased activity of helper T cells, which promote inflammation, is associated with HCC [106], and chronic inflammation has been implicated in the development of liver cancer in humans [107]. While the immunoglobulin protein family (CD28 and cytotoxic T-lymphocyte antigen-4) plays important roles in the control of T-cell responses against infection and cancer, tumors seem to have exploited these pathways to evade immune surveillance. Activation and proliferation of cytotoxic T lymphocytes is suppressed in individuals with HCC [108].
Recent studies suggested a role of the immune system in constitutional susceptibility to HCC. A genome-wide association study (GWAS) , focusing on HCC, revealed that constitutional genetic variations are risk factors for HCC [109]. Three susceptibility loci have been strongly associated with HCC including the class II MHC complex, whose protein products present antigen to T-cell receptors and mediate immune surveillance (rs9267673, rs2647073, and rs3997872) resulting in an ineffective T-cell response. MHC class II molecules present antigen to CD4. Thus, genes involved in the immune response play a critical role in the development of HCC. Since only a subset of liver cirrhosis patients develop HCC, the transition from cirrhosis to HCC has been attributed to two SNPs whose allele frequencies differ significantly between HCC and cirrhosis (one lies in the PTEN homolog TPTE2, and the second variant lies within an intron of TPTE2, which encodes a homolog of the PTEN tumor suppressor protein [110]). Multiple SNP analysis showed that “antigen processing and presentation” emerged as the pathway with the strongest association with HCC. Thus, the T-cell repertoire of each individual plays a critical role in HCC susceptibility and that biological processes affecting T-cell maturation or immune surveillance may represent important etiologic mechanisms for the development of HCC in humans.
The NK (natural killer cells) are able to recognize and kill invading pathogens and cancer cells. This capability depends on the balance between activating (CD16, NKG2D, NKG2C, CD226, CD244, and the natural cytotoxicity receptors) and inhibitory (killer cell immunoglobulin-like receptors [KIRs], CD94/NKG2A, and leukocyte immunoglobulin-like receptor 1 [CD85], most of which recognize MHC class I molecules) receptor signaling, a complex process requiring several NK cell surface receptors acting synergistically. By inducing the upregulation of inhibitory receptors and downregulation of activating receptors on NK cells, cancer cells can become “invisible” to immune surveillance. Furthermore, NK cell dysfunction may promote the escape of tumor cells. Numerous studies have found a reduction in the proportion of NK cells in peripheral blood of HCC patients [111] and a decrease in the expression of NK cell-activating receptors during the development and progression of HCC [112].
Chronic alcohol consumption could have profound effect on both innate and adaptive immunity [113]. Immune function inhibition by alcohol could allow tumor evasion from immune surveillance and ultimately establishing tumor growth. NK cell activity was decreased in abstinent alcoholics compared to nondrinkers [114] and alcoholics show reduced numbers of T cells (including CD4+ T cells, CD8+ T cells) with alterations in their cytokine expression [115].
As mentioned earlier, antitumor defense requires both effective antigen presentation and the capacity of the T cells to respond to the antigen priming. Chronic alcohol impairs antigen presentation by dendritic cells and monocytes and damages the primary response of CD8+ cells, but not CD4+ response to specific antigen priming in mice [116]. In addition, CD8+ cells from the ethanol-fed mice exhibited poorer proliferation and poorer IFNγ production after priming. Thus, chronic alcohol consumption impairs the interactions between the antigen-presenting cells and the T cells [117] and causes intrinsic defects in the T cells themselves [99].
Neoangiogenesis
The hallmarks of hepatic circulation in liver cirrhosis are vasoconstriction, sinusoidal remodeling, angiogenesis, and venous thrombosis, which all contribute to increase hepatic vascular resistance and portal hypertension [118]. Angiogenesis is the result of two opposing processes regulated by proangiogenic factors (e.g., vascular endothelial growth factor (VEGF), fibroblast growth factor (FGF), angiopoietin, EGF, and PDGF, which induce angiogenic signaling via RAS/RAF/MEK/ERK, mTOR, and Wnt signal transduction pathways) and inhibitory factors (thrombospondin [TSP] and angiostatin) [119]. Increased expression and secretion of VEGFA due to hypoxia (mediated by hypoxia-inducible factor 2-α) of cancer cells [120] induces endothelial cells’ proliferation, migration, survival, and angiogenesis which promote tumor growth [121]. Normally, HCC displays active angiogenesis, which not only contributes to increased vascular resistance and portal hypertension, but also allows cancer cells to invade vessels and metastasize [122].
Other signaling pathways involved in hepatocarcinogenesis include phosphatidylinositol-3 kinase (PI3K)/AKT/mTOR), Wnt/β-catenin, insulin-like growth factor, and hepatocyte growth factor/c-MET [123]. Wnt/β-catenin pathway contributes to HCC formation by influencing cell adhesion and transcriptional activation of target genes such as c-myc and cyclin D [124]. In fact, β-catenin accumulation, a hallmark of the activated Wnt/FZ signaling, has been observed in 33–67 % of HCC tumors [125]. In addition, miR-610 was downregulated in human HCC, thus promoting HCC cell proliferation and tumorigenicity by activating Wnt/β-catenin signaling [126]. On the other hand, overexpression of miR-153 was able to promote β-catenin transcriptional activity, leading to cell-cycle progression, proliferation, and colony formation of HCC cells [127].
Although no studies have examined the relationship between alcohol consumption and HCC angiogenesis, other studies showed that this is a plausible hypothesis. In experimental animals, alcohol consumption (equivalent to ~2 drinks/day in humans) by immune-competent mice implanted with mouse melanoma cells resulted in an increase in VEGF transcript and protein levels, doubling of tumor volume, and enhanced microvascular density [128]. Further studies showed that alcohol intake enhances angiogenesis in a rat model of choroidal neovascularization [129]. Chronic alcohol consumption increased HCC angiogenesis, progression, and metastasis through NFκB-dependent VEGF and MCP-1 upregulation [130].
In addition, alcohol-induced oxidative stress and inflammation further amplify vasoconstriction and portal hypertension and the ensuing angiogenesis, a hallmark in tumor maintenance [131].
Breast Cancer
Breast cancer is a heterogeneous disease that encompasses more than 20 different subtypes and has a wide range of known risk factors involved in its development. Many of the primary risk factors for breast cancer are beyond women’s control, such as aging, inherited changes in certain genes and family history of breast cancer, prenatal history (e.g., daughters born to mothers who used diethylstilbestrol (DES) during pregnancy), and reproductive parameters such as first full-term pregnancy, miscarriage, and abortion. However, modifiable lifestyle risk factors under women’s control include dietary habits (consumption of polyunsaturated fats and excessive alcohol), smoking, exposure to radiation or synthetic estrogens, viral infection, physical inactivity, use of HRT, obesity, diabetes, breast implants, and even changes in circadian rhythm homeostasis, such as night shift work. The interactions between genetic susceptibility for breast cancer and the environmental factors add another layer of complexity to this picture.
Epidemiological studies on alcohol and breast cancer are inconsistent. For example, in one study [132], consumption of one or two drinks/day increased breast cancer risk by 40 %, whereas consumption of two or more drinks/day resulted in no increase in risk. For moderate drinking, one study [133] stated that drinking <1.5 drink/day was associated with a 42 % decrease in breast cancer risk, whereas another [134] reported that drinking 3–6 drinks/week was associated with a 15 % increase in risk. A meta-analysis [135] stated “the modest size of the association and variation in results across studies leave the causal role of alcohol in question.” The variability in outcomes could be ascribed to: (1) the majority of the studies rely on self-report to determine the amount of alcoholic beverage consumed, which introduces “recall bias” into the studies that makes the alcohol consumption variable notoriously inaccurate; (2) since breast cancer develops over a period of more than 20 years [136], the correlations between current or recent alcohol consumption and breast cancer cannot be determined in epidemiological studies that captures intake after clinical diagnosis; (3) epidemiological study rarely differentiates between specific subtypes of breast cancer—which are associated with unique risk factors and might have been preprogrammed at an early stage of the disease—and risk of alcohol consumption. Better standardization of study designs in assessing alcohol intake and timing of exposure may improve our understanding of the heterogeneity of results across studies [137].
Despite the challenge in understanding the epidemiological findings, several molecular mechanisms have been postulated [138] for alcohol-associated breast cancer, including formation of acetaldehyde and ROS, epigenetic effect through the folate cycle, and estrogen formation. For a comprehensive review of alcohol and breast cancer, the reader is referred to the article by Zakhari and Hoek [139].
Colon Cancer
Colorectal cancer (CRC) is the third most commonly diagnosed cancer in the world with an estimated 1.24 million new cases each year [140]. In the United States, the annual incidence of CRC is about 148,300, with 56,600 deaths per year, and the lifetime risk in the general population is about 5–6 % [141]. It appears that chronic heavy consumption of alcohol may increase the relative risk for colon cancer.
In the colon, acetaldehyde is primarily produced from ethanol by resident bacteria and, to a lesser extent, by mucosal ADHs. Human colon mucosal cells harbor ADH1, ADH3, and ADH5, with the ADH1 and ADH3 isozymes being most active [52]. In an in vitro experiment, human colon contents were able to generate 60–250 μM acetaldehyde when incubated with 10–100 mg % of ethanol [142]. The high levels of acetaldehyde attained in the colon likely underlie the correlation between chronic, heavy ethanol consumption and CRC in humans. In alcohol-treated rats, a high concentration of acetaldehyde (50–350 μM) in the colon mucosa has been shown to correlate positively with hyperproliferation of the colon crypt cells [143]. Another evidence for a role of acetaldehyde in CRC initiation emanates from studies showing 3.4 times increase in colon cancer risk among Asians who possess a polymorphism in their ALDH2 enzyme known as ALDH2*2 [144].
As mentioned above, acetaldehyde is metabolized to acetate by ALDH2, ALDH1B1, and ALDH1A1 [145]. The ability of these ALDHs to detoxify acetaldehyde levels is consistent with a role for ALDHs in colon cancer. This hypothesis is supported by the association of ALDH2 deficiency with high incidence of CRC in heavy drinkers [144]. In addition to detoxifying acetaldehyde, ALDH1A enzymes are involved in the formation of RA from retinaldehyde, which plays an important role in cellular proliferation and differentiation [146]. Therefore, RA-generating ALDHs play a critical role in modulating carcinogenesis. In addition, ALDH activity is used as a molecular tool to isolate normal and cancer stem cells of various lineages [147]. Furthermore, the high ALDH expression in cancer stem cells is associated with poor prognosis in CRC [148]. These ALDH bright cells (cells with very high ALDH expression) are more tumorigenic, as reflected by colony-forming capability in vitro and in xenograft-induced tumor formation in vivo [149]. The high expression of ALDH1B1 in both human colon cancers and in animal model of colon polyps has been identified, specifically adenomatous polyposis coli multiple intestinal neoplasia (Apc (Min)/+) in mice [150]. These mice have the tumor suppressor Apc gene mutated, which upregulates oncogenes like c-Myc via a dysregulated Wnt signaling [151].
In vivo studies in rats have revealed that retinoids added to the diet reduced colon cancer cell proliferation and prevented azoxymethane-induced aberrant crypt foci (putative precancerous lesions in colon) and colon tumor formation [152, 153].
Alcohol and Upper Aerodigestive Tract Cancer
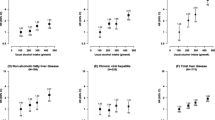
UADT cancer is among the most frequent cancers in the world. Cancers of the larynx account for approximately 12,000 new cancer cases per year in the United States. Worldwide, approximately 260,000 new cases of oral cancer occur, and more than 125,000 mortalities are attributed to oral cancers each year. Epidemiological studies report an inconsistent relationship between alcohol drinking and UADT cancer mortality; heavy alcohol consumption could increase UADT cancer mortality, especially if combined with tobacco use [154].
For detailed information about alcohol and cancer, the reader is referred to the monographs “Alcohol and Cancer” [155] and “Biological Basis of Alcohol-Induced Cancer” [156].
References
Seitz HK, Stickel F. Risk factors and mechanisms of hepatocarcinogenesis with special emphasis on alcohol and oxidative stress. Biol Chem. 2006;387:349–60.
Ferlay J, Soerjomataram I, Ervik M, et al. GLOBOCAN 2012 v1.2, cancer incidence and mortality worldwide: IARC Cancer Base No. 11. 2015. Available from http://globocan.iarc.fr
El-Serag HB, Rudolph KL. Hepatocellular carcinoma: epidemiology and molecular carcinogenesis. Gastroenterology. 2007;132:2557–76.
Poli A, Marangoni F, Avogaro A, Barba G, et al. Moderate alcohol use and health: A consensus document. Nutr Metab Cardiovasc Dis. 2013;23:487–504.
Friedman SL. Mechanisms of hepatic fibrogenesis. Gastroenterology. 2008;134:1655–69.
Bellantani S, Saccoccio G, Costa G, et al. Drinking habits as cofactors of risk for alcohol-induced liver damage. Gut. 1997;41:845–50.
Seitz HK, Stickel F. Molecular mechanisms of alcohol-mediated carcinogenesis. Nat Rev Cancer. 2007;7:599–612.
Zakhari S. Bermuda Triangle for the liver: alcohol, obesity, and viral hepatitis. J Gastroenterol Hepatol. 2013;28 Suppl 1:18–25.
Cornellà H, Alsinet C, Villanueva A. Molecular pathogenesis of hepatocellular carcinoma. Alcohol Clin Exp Res. 2011;35:821–5.
Zender L, Villanueva A, Tovar V, et al. Cancer gene discovery in hepatocellular carcinoma. J Hepatol. 2010;52:921–9.
Balbo S, Hashibe M, Gundy S, Brennan P, Canova C, Simonato L, Merletti F, Richiardi L, Agudo A, Castellsagué X, et al. N2-ethyldeoxyguanosine as a potential biomarker for assessing effects of alcohol consumption on DNA. Cancer Epidemiol Biomarkers Prev. 2008;17:3026–32.
Garcia CC, Angeli JP, Freitas FP, Gomes OF, de Oliveira TF, Loureiro AP, Di Mascio P, Medeiros MH. [13C2]-Acetaldehyde promotes unequivocal formation of 1, N2-propano-2′-deoxyguanosine in human cells. J Am Chem Soc. 2011;133:9140–3.
Brooks PJ, Zakhari S. Acetaldehyde and the genome: beyond nuclear DNA adducts and carcinogenesis. Environ Mol Mutagen. 2014;55:77–91.
Yokoyama A, Omori T. Genetic polymorphisms of alcohol and aldehyde dehydrogenases and risk for esophageal and head and neck cancers. Alcohol. 2005;35:175–85.
Ziecha D, Franco R, Pappa A, Panayiotidis MI. Reactive oxygen species (ROS)-induced genetic and epigenetic alterations in human carcinogenesis. Mutat Res. 2011;711:167–73.
Hussain SP, Hofseth LJ, Harris CC. Radical causes of cancer. Nat Rev Cancer. 2003;3:276–85.
Trachootham D, Alexander J, Huang P. Targeting cancer cells by ROS-mediated mechanism: a radical therapeutic approach? Nat Rev. 2009;8:579–91.
Tudek B, Winczura A, Janik J, et al. Involvement of oxidatively damaged DNA and repair in cancer development and aging. Am J Transl Res. 2010;2:254–84.
Hagen TM, Huang S, Curnutte J, et al. Extensive oxidative DNA damage in hepatocytes of transgenic mice with chronic active hepatitis destined to develop hepatocellular carcinoma. Proc Natl Acad Sci U S A. 1994;91:12808–12.
Lim SO, Gu JM, Kim MS, Kim HS, et al. Epigenetic changes induced by reactive oxygen species in hepatocellular carcinoma: methylation of the E-cadherin promoter. Gastroenterology. 2008;135:2128–40.
Forsyth CB, Tang Y, Shaikh M, Zhang L, Keshavarzian A. Alcohol stimulates activation of Snail, epidermal growth factor receptor signaling, and biomarkers of epithelial-mesenchymal transition in colon and breast cancer cells. Alcohol Clin Exp Res. 2010;34:19–31.
Fujinaga H, Tsutsumi T, Yotsuyanagi H, et al. Hepatocarcinogenesis in hepatitis C: HCV shrewdly exacerbates oxidative stress by modulating both production and scavenging of reactive oxygen species. Oncology. 2011;81 Suppl 1:11–7.
Gong P, Cerderbaum A. Nrf2 is increased by CYP2E1 in rodent liver and HepG2 cells and protects against oxidative stress caused by CYP2E1. Hepatology. 2006;43(1):144–53.
Cederbaum AI. Alcohol metabolism. Clin Liver Dis. 2012;16:667–85.
Cederbaum A. Nrf2 and antioxidant defense against CYP2E1 toxicity. Expert Opin Drug Metab Toxicol. 2009;5:1223–44.
Imai SI, Armstrong CM, Kaeberlein M, Guarente L. Transcriptional silencing and longevity protein Sir2 is an NAD-dependent histone deacetylase. Nature. 2000;403:795–800.
Vaziri H, et al. hSIR2 SIRT1 functions as an NAD-dependent p53 deacetylase. Cell. 2001;107:149–59.
Fan J, Hitosugi T, Chung TW, et al. Tyrosine phosphorylation of lactate dehydrogenase A is important for NADH/NAD redox homeostasis in cancer cells. Mol Cell Biol. 2011;31:4938–50.
Gauthier PM, Szerlip HM. Metabolic acidosis in the intensive care unit. Crit Care Clin. 2002;18:289–308.
Clugston RD, Blaner WS. The adverse effects of alcohol on vitamin A metabolism. Nutrients. 2012;4(5):356–71.
Shiota G, Kanki K. Retinoids and their target genes in liver functions and diseases. J Gastroenterol Hepatol. 2013;Suppl 1:33–7.
Wang XD. Alcohol, vitamin A, and cancer. Alcohol. 2005;35:251–8.
Mobarhan S, Seitz HK, Russell RM, Mehta R, Hupert J, Friedman H, Layden TJ, Meydani M, Langenberg P. Age-related effects of chronic ethanol intake on vitamin A status in Fisher 344 rats. J Nutr. 1991;121:510–7.
Mercer KE, Hennings L, Sharma N, Lai K, Cleves MA, Wynne RA, Badger TM, Ronis MJ. Alcohol consumption promotes diethylnitrosamine-induced hepatocarcinogenesis in male mice through activation of the Wnt/β-catenin signaling pathway. Cancer Prev Res (Phila). 2014;7(7):675–85.
Rosales FJ, Ritter SJ, Zolfaghari R, Smith JE, Ross AC. Effects of acute inflammation on plasma retinol, retinol-binding protein, and its mRNA in the liver and kidneys of vitamin A-sufficient rats. J Lipid Res. 1996;37:962–71.
di Masi A, Leboffe L, De Marinis E, Pagano F, Cicconi L, Rochette-Egly C, Lo-Coco F, Ascenzi P, Nervi C. Retinoic acid receptors: From molecular mechanisms to cancer therapy. Mol Aspects Med. 2015;41C:1–115.
Kumar A, Singh CK, Dipette DD, Singh US. Ethanol impairs activation of retinoic acid receptors in cerebellar granule cells in a rodent model of fetal alcohol spectrum disorders. Alcohol Clin Exp Res. 2010;34(5):928–37.
Shukla SD, Velazquez J, French SW, Lu SC, Ticku MK, Zakhari S. Emerging role of epigenetics in the actions of alcohol. Alcohol Clin Exp Res. 2008;32:1525–34.
Stickel F, Schuppan D, Hahn EG, Seitz HK. Cocarcinogenic effects of alcohol in hepatocarcinogenesis. Gut. 2002;51:132–9.
Wang XD, Liu C, Chung J, Stickel F, Seitz HK, Russell RM. Chronic alcohol intake reduces retinoic acid concentration and enhances AP-1 (c-Jun and c-Fos) expression in rat liver. Hepatology. 1998;28(3):744–50.
Fausto N, Campbell JS. The role of hepatocytes and oval cells in liver regeneration and repopulation. Mech Dev. 2003;120:117–30.
Huang J, Bi Y, Zhu GH, He Y, Su Y, He BC, Wang Y, Kang Q, Chen L, Zuo GW, Luo Q, Shi Q, Zhang BQ, Huang A, Zhou L, Feng T, Luu HH, Haydon RC, He TC, Tang N. Retinoic acid signalling induces the differentiation of mouse fetal liver-derived hepatic progenitor cells. Liver Int. 2009;29(10):1569–81.
Zhang Y, Guan DX, Shi J, Gao H, Li JJ, Zhao JS, Qiu L, Liu J, Li N, Guo WX, Xue J, Zhou FG, Wu MC, Wang HY, Xie D, Cheng SQ. All-trans retinoic acid potentiates the chemotherapeutic effect of cisplatin by inducing differentiation of tumor initiating cells in liver cancer. J Hepatol. 2013;59(6):1255–63.
Hu Z, Fujio K, Marsden ER, Thorgeirsson SS, Evarts RP. Hepatic regeneration in vitamin A-deficient rats: changes in the expression of transforming growth factor alpha/epidermal growth factor receptor and retinoic acid receptors alpha and beta. Cell Growth Differ. 1994;5:503–8.
Kuper H, Ye W, Broomé U, Romelsjö A, Mucci LA, Ekbom A, Adami HO, Trichopoulos D, Nyrén O. The risk of liver and bile duct cancer in patients with chronic viral hepatitis, alcoholism, or cirrhosis. Hepatology. 2001;34:714–8.
Fattovich G, Stroffolini T, Zagni I, Donato F. Hepatocellular carcinoma in cirrhosis: incidence and risk factors. Gastroenterology. 2004;127:S35–50.
Agarwal DP. Genetic polymorphisms of alcohol metabolizing enzymes. Pathol Biol (Paris). 2001;49:703–9.
Boonyaphiphat P, Thongsuksai P, Sriplung H, Puttawibul P. Lifestyle habits and genetic susceptibility and the risk of esophageal cancer in the Thai population. Cancer Lett. 2002;186(2):193–9.
Hiraki A, Matsuo K, Wakai K, Suzuki T, Hasegawa Y, Tajima K. Gene–gene and gene–environment interactions between alcohol drinking habit and polymorphisms in alcohol-metabolizing enzyme genes and the risk of head and neck cancer in Japan. Cancer Sci. 2007;98:1087–91.
Shen YC, et al. Polymorphism of ADH and ALDH genes among four ethnic groups in China and effects upon the risk for alcoholism. Alcohol Clin Exp Res. 1997;21:1272–7.
Nagata N, et al. Assessment of a difference in ALDH2 heterozygotes and alcoholic liver injury. Alcohol Clin Exp Res. 2002;26:11S–4.
Jelski W, Zalewski B, Chrostek L, Szmitkowski M. The activity of class I, II, III, and IV alcohol dehydrogenase isoenzymes and aldehyde dehydrogenase in colorectal cancer. Dig Dis Sci. 2004;49:977–81.
Coutelle C, Höhn B, Benesova M, Oneta CM, et al. Risk factors in alcohol associated breast cancer: alcohol dehydrogenase polymorphism and estrogens. Int J Oncol. 2004;25:1127–32.
Brooks PJ, Enoch MA, Goldman D, et al. The alcohol flushing response: an unrecognized risk factor for esophageal cancer from alcohol consumption. PLoS Med. 2009;6(3):e1000050.
Yokoyama A, Omori T. Genetic polymorphisms of alcohol and aldehyde dehydrogenases and risk for esophageal and head and neck cancers. Jpn J Clin Oncol. 2003;33:111–21.
Ueno Y, et al. Effect of the cytochrome P-450IIE1 genotype on ethanol elimination rate in alcoholics and control subjects. Alcohol Clin Exp Res. 1996;20(Suppl):17A–21.
Seitz HK, Wang XD. The role of cytochrome P450 2E1 in ethanol-mediated carcinogenesis. Subcell Biochem. 2013;67:131–43.
Druesne-Pecollo N, Tehard B, Mallet Y, Gerber M, Norat T, et al. Alcohol and genetic polymorphisms: effect on risk of alcohol-related cancer. Lancet Oncol. 2009;10:173–80.
Han Z-G. Functional genomic studies: insights into the pathogenesis of liver cancer. Annu Rev Genomics Hum Genet. 2012;13:171–205.
Herceg Z, Paliwal A. Epigenetic mechanisms in hepatocellular carcinoma: How environmental factors influence the epigenome. Mutat Res. 2011;727:55–61.
Ozen C, Yildiz G, Dagcan AT, et al. Genetics and epigenetics of liver cancer. N Biotechnol. 2013;30:381–4.
Zakhari S. Alcohol metabolism and epigenetics changes. Alcohol Res. 2013;35:6–16.
Zhang T-Y, Meaney MJ. Epigenetics and the environmental regulation of the genome and its function. Annu Rev Psychol. 2010;61:439–66.
Bleich S, Hillemacher T. Homocysteine, alcoholism and its molecular networks. Pharmacopsychiatry. 2009;42:S102–9.
Mato JM, Martínez-Chantar ML, Lu SC. Methionine metabolism and liver disease. Annu Rev Nutr. 2008;28:273–93.
Varela-Rey M, Woodhoo A, Martinez-Chantar M-L, Mato JM, Lu SC. Alcohol, DNA methylation, and cancer. Alcohol Res. 2013;35:25–35.
Lieber CS. ALCOHOL: its metabolism and interaction with nutrients. Annu Rev Nutr. 2000;20:395–430.
Hamid A, Wani NA, Kaur J. New perspectives on folate transport in relation to alcoholism-induced folate malabsorption—association with epigenome stability and cancer development. FEBS J. 2009;276:2175–91.
Dannenberg LO, Chen HJ, Tian H, Edenberg HJ. Differential regulation of the alcohol dehydrogenase 1B (ADH1B) and ADH1C genes by DNA methylation and histone deacetylation. Alcohol Clin Exp Res. 2006;30:928–37.
Bonsch D, Lenz B, Fiszer R, Frieling H, Kornhuber J, Bleich S. Lowered DNA methyltransferase (DNMT-3b) mRNA expression is associated with genomic DNA hypermethylation in patients with chronic alcoholism. J Neural Transm. 2006;113:1299–304.
Kouzarides T. Chromatin modifications and their function. Cell. 2007;128:693–705.
Kim JS, Shukla SD. Acute in vivo effect of ethanol (binge drinking) on histone H3 modifications in rat tissues. Alcohol Alcohol. 2006;41:126–32.
Choudhury M, Shukla SD. Surrogate alcohols and their metabolites modify histone H3 acetylation: involvement of histone acetyl transferase and histone deacetylase. Alcohol Clin Exp Res. 2008;32:829–39.
Pal-Bhadra M, Bhadra U, Jackson DE, Mamatha L, Park PH, Shukla SD. Distinct methylation patterns in histone H3 at Lys-4 and Lys-9 correlate with up- & down-regulation of genes by ethanol in hepatocytes. Life Sci. 2007;81:979–87.
You M, Liang X, Ajmo JM, Ness GC. Involvement of mammalian sirtuin 1 in the action of ethanol in the liver. Am J Physiol Gastrointest Liver Physiol. 2008;294:G892–8.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, Sweet-Cordero A, Ebert BL, Mak RH, Ferrando AA, Downing JR, Jacks T, Horvitz HR, Golub TR. MicroRNA expression profiles classify human cancers. Nature. 2005;435:834–8.
Calin GA, Croce CM. MicroRNA-cancer connection: the beginning of a new tale. Cancer Res. 2006;66:7390–4.
Murakami Y, Yasuda T, Saigo K, Urashima T, Toyoda H, Okanoue T, Shimotohno K. Comprehensive analysis of microRNA expression patterns in hepatocellular carcinoma and non-tumorous tissues. Oncogene. 2006;25:2537–45.
Ladeiro Y, Couchy G, Balabaud C, Bioulac-Sage P, Pelletier L, Rebouissou S, Zucman-Rossi J. MicroRNA profiling in hepatocellular tumors is associated with clinical features and oncogene/tumor suppressor gene mutations. Hepatology. 2008;47:1955–63.
Milazzo M, Fornari F, Gramanteri L. MicroRNA and hepatocellular carcinoma: biology and prognostic significance. Minerva Gastroenterol Dietol. 2011;57:257–71.
Tang Y, Banan A, Forsyth CB, Fields JZ, Lau CK, Zhang LJ, Keshavarzian A. Effect of alcohol on miR-212 expression in intestinal epithelial cells and its potential role in alcoholic liver disease. Alcohol Clin Exp Res. 2008;32:355–64.
Dippold RP, Vadigepalli R, Gonye GE, et al. Chronic ethanol feeding enhances miR-21 induction during liver regeneration while inhibiting proliferation in rats. Am J Physiol Gastrointest Liver Physiol. 2012;303:G733–43.
Bala S, Marcos M, Kodys K, et al. Upregulation of microRNA-155 in macrophages contributes to increased tumor necrosis factor alpha (TNF{alpha}) production via increased mRNA half-life in alcoholic liver disease. J Biol Chem. 2011;286:1436–44.
Dippold RP, Vadigepalli R, Gonye GE, et al. Chronic ethanol feeding alters miRNA expression dynamics during liver regeneration. Alcohol Clin Exp Res. 2013;37 Suppl 1:E59–69.
Yeligar S, Tsukamoto H, Kalra VK. Ethanol-induced expression of ET-1 and ET-BR in liver sinusoidal endothelial cells and human endothelial cells involves hypoxia-inducible factor-1alpha and microrNA-199. J Immunol. 2009;183:5232–43.
Dolganiuc A, Petrasek J, Kodys K, Catalano D, Mandrekar P, Velayudham A, Szabo G. MicroRNA expression profile in Lieber-DeCarli diet-induced alcoholic and methionine choline deficient diet-induced nonalcoholic steatohepatitis models in mice. Alcohol Clin Exp Res. 2009;33:1704–10.
Bala S, Szabo G. MicroRNA signature in alcoholic liver disease. Int J Hepatol. 2012;2012:498232.
Reick M, Garcia JA, Dudley C, McKnight SL. NPAS2: an analog of clock operative in the mammalian forebrain. Science. 2001;293:506–9.
Filipski E, Subramanian P, Carrière J, Guettier C, Barbason H, Lévi F. Circadian disruption accelerates liver carcinogenesis in mice. Mutat Res. 2009;680:95–105.
Lin YM, Chang JH, Yeh KT, Yang MY, Liu TC, Lin SF, Su WW, Chang JG. Disturbance of circadian gene expression in hepatocellular carcinoma. Mol Carcinog. 2008;47:925–33.
Cui M, Zheng M, Sun B, Wang Y, et al. A long noncoding RNA perturbs the circadian rhythm of hepatoma cells to facilitate hepatocarcinogenesis. Neoplasia. 2015;17:79–88.
Zhao B, Lu J, Yin J, Liu H, et al. A functional polymorphism in PER3 gene is associated with prognosis in hepatocellular carcinoma. Liver Int. 2012;32:1451–9.
Zhang YK, Yeager RL, Klaassen CD. Circadian expression profiles of drug-processing genes and transcription factors in mouse liver. Drug Metab Dispos. 2009;37:106–15.
Green CB, Takahashi JS, Bass J. The meter of metabolism. Cell. 2008;134:728–42.
Turek FW, Joshu C, Kohsaka A, Lin E, Ivanova G, McDearmon E, et al. Obesity and metabolic syndrome in circadian Clock mutant mice. Science. 2005;308:1043–5.
Kohsaka A, Laposky AD, Ramsey KM, Estrada C, Joshu C, Kobayashi Y, et al. High-fat diet disrupts behavioral and molecular circadian rhythms in mice. Cell Metab. 2007;6:414–21.
Barnea M, Madar Z, Froy O. High-fat diet delays and fasting advances the circadian expression of adiponectin signaling components in mouse liver. Endocrinology. 2009;150:161–8.
Yang X, Downes M, Yu RT, Bookout AL, He W, Straume M, et al. Nuclear receptor expression links the circadian clock to metabolism. Cell. 2006;126:801–10.
Rutter J, Reick M, McKnight SL. Metabolism and the control of circadian rhythms. Annu Rev Biochem. 2002;71:307–31.
Sani M, Ghanem-Boughanmi N, Gadacha W, Sebai H, Boughattas NA, Reinberg A, Ben-Attia M. Malondialdehyde content and circadian variations in brain, kidney, liver, and plasma of mice. Chronobiol Int. 2007;24:671–85.
McNamara P, Seo SB, Rudic RD, Sehgal A, Chakravarti D, FitzGerald GA. Regulation of CLOCK and MOP4 by nuclear hormone receptors in the vasculature: a humoral mechanism to reset a peripheral clock. Cell. 2001;105:877–89.
Nelson RJ. Seasonal immune function and sickness responses. Trends Immunol. 2004;25:187–92.
Dimitrov S, Lange T, Tieken S, Fehm HL, Born J. Sleep associated regulation of T helper 1/T helper 2 cytokine balance in humans. Brain Behav Immun. 2004;18:341–8.
Okada K, Yano M, Doki Y, Azama T, Iwanaga H, Miki H, et al. Injection of LPS causes transient suppression of biological clock genes in rats. J Surg Res. 2008;145:5–12.
Chew V, Chen J, Lee D, Loh E, Lee J, Lim KH, et al. Chemokine-driven lymphocyte infiltration: an early intratumoural event determining long-term survival in resectable hepatocellular carcinoma. Gut. 2012;61:427–38.
Pentcheva-Hoang T, Corse E, Allison JP. Negative regulators of T-cell activation: potential targets for therapeutic intervention in cancer, autoimmune disease, and persistent infections. Immunol Rev. 2009;229:67–87.
Mantovani A, Allavena P, Sica A, Balkwill F. Cancer-related inflammation. Nature. 2008;454:436–44.
Ormandy LA, Hillemann T, Wedemeyer H, Manns MP, Greten TF, Korangy F. Increased populations of regulatory T cells in peripheral blood of patients with hepatocellular carcinoma. Cancer Res. 2005;65:2457–64.
Clifford RJ, Zhang J, Meerzaman DM, et al. Genetic variations at loci involved in the immune response are risk factors for hepatocellular carcinoma. Hepatology. 2010;52:2034–43.
Walker SM, Downes CP, Leslie NR. TPIP: a novel phosphoinositide 3-phosphatase. Biochem J. 2001;360:277–83.
Cai L, Zhang Z, Zhou L, Wang H, Fu J, Zhang S, et al. Functional impairment in circulating and intrahepatic NK cells and relative mechanism in hepatocellular carcinoma patients. Clin Immunol. 2008;129:428–37.
Chinnery F, King CA, Elliott T, Bateman AR, James E. Viral antigen mediated NKp46 activation of NK cells results in tumor rejection via NK–DC crosstalk. Oncoimmunology. 2012;1:874–83.
Szabo G, Mandrekar P. A recent perspective on alcohol, immunity and host defense. Alcohol Clin Exp Res. 2009;33:1–13.
Motivala SJ, Dang J, Obradovic T, Meadows GG, Butch AW, Irwin MR. Leptin and cellular and innate immunity in abstinent alcoholics and controls. Alcohol Clin Exp Res. 2003;27:1819–24.
Zhang H, Meadows GG. Chronic alcohol consumption in mice increases the proportion of peripheral memory T cells by homeostatic proliferation. J Leukoc Biol. 2005;78:1070–80.
Gurung P, Young BM, Coleman RA, Wiechert S, Turner LE, Ray NB, Waldschmidt TJ, Legge KL, Cook RT. Chronic ethanol induces inhibition of antigen-specific CD8+ but not CD4+ immunodominant T cell responses following Listeria monocytogenes inoculation. J Leukoc Biol. 2009;85:34–43.
Cook RT, Zhu X, Coleman RA, Ballas ZK, Waldschmidt TJ, Ray NB, LaBrecque DR, Cook BL. T-cell activation after chronic ethanol ingestion in mice. Alcohol. 2004;33:175–81.
Iwakiri Y, Groszmann RJ. The hyperdynamic circulation of chronic liver diseases: from the patient to the molecule. Hepatology. 2006;43:S121–31.
Carmeliet P, Jain RK. Angiogenesis in cancer and other diseases. Nature. 2000;407:249–57.
Harris AL. Hypoxia—a key regulatory factor in tumour growth. Nat Rev Cancer. 2002;2:38–47.
Kim KJ, Li B, Winer J, Armanini M, Gillett N, Phillips HS, Ferrara N. Inhibition of vascular endothelial growth factor-induced angiogenesis suppresses tumour growth in vivo. Nature. 1993;362:841–4.
Fernández M, Semela D, Bruix J, Colle I, Pinzani M, Bosch J. Angiogenesis in liver disease. J Hepatol. 2009;50:604–20.
Whittaker S, Marais R, Zhu AX. The role of signaling pathways in the development and treatment of hepatocellular carcinoma. Oncogene. 2010;29:4989–5005.
Villanueva A, Newell P, Chiang DY, Friedman SL, Llovet JM. Genomics and signaling pathways in hepatocellular carcinoma. Semin Liver Dis. 2007;27:55–76.
Giles RH, van Es JH, Clevers H. Caught up in a Wnt storm: Wnt signaling in cancer. Biochim Biophys Acta. 2003;1653:1–24.
Zeng XC, Liu FQ, Yan R, et al. Downregulation of miR-610 promotes proliferation and tumorigenicity and activates Wnt/β-catenin signaling in human hepatocellular carcinoma. Mol Cancer. 2014;13:261–76.
Hua HW, Jiang F, Huang Q, Liao Z, Ding G. MicroRNA-153 promotes Wnt/β-catenin activation in hepatocellular carcinoma through suppression of WWOX. Oncotarget. 2015;6:3840–7.
Tan W, Bailey AP, Shparago M, Busby B, Covington J, Johnson JW, Young E, Gu JW. Chronic alcohol consumption stimulates VEGF expression, tumor angiogenesis and progression of melanoma in mice. Cancer Biol Ther. 2007;6:1211–7.
Kaliappan S, Jha P, Lyzogubov VV, Tytarenko RG, Bora NS, Bora PS. Alcohol and nicotine consumption exacerbates choroidal neovascularization by modulating the regulation of complement system. FEBS Lett. 2008;582:3451–8.
Wang F, Yang J-L, Yu KK, et al. Activation of the NF-κB pathway as a mechanism of alcohol enhanced progression and metastasis of human hepatocellular carcinoma. Mol Cancer. 2015;14:10–24.
Gitto S, Micco L, Conti F, Andreone P, Bernardi M. Alcohol and viral hepatitis: a mini-review. Dig Liver Dis. 2009;41:67–70.
Kinney AY, Millikan RC, Lin YH, Moorman PG, Newman B. Alcohol consumption and breast cancer among black and white women in North Carolina (United States). Cancer Causes Control. 2000;11:345–57.
Bessaoud F, Daurès JP. Patterns of alcohol (especially wine) consumption and breast cancer risk: a case-control study among a population in Southern France. Ann Epidemiol. 2008;18:467–75.
Chen WY, Rosner B, Hankinson SE, Colditz GA, Willett WC. Moderate alcohol consumption during adult life, drinking patterns, and breast cancer risk. JAMA. 2011;306:1884–90.
Longnecker MP. Alcoholic beverage consumption in relation to risk of breast cancer: meta-analysis and review. Cancer Causes Control. 1994;5:73–82.
Brooks PJ, Zakhari S. Moderate alcohol consumption and breast cancer in women: from epidemiology to mechanisms and interventions. Alcohol Clin Exp Res. 2013;37:23–30.
Scoccianti C, Lauby-Secretan B, Bellp P-Y, et al. Female breast cancer and alcohol consumption: a review of the literature. Am J Prev Med. 2014;46:S16–25.
Castro GD, Castro JA. Alcohol drinking and mammary cancer: pathogenesis and potential dietary preventive alternatives. World J Clin Oncol. 2014;5:713–29.
Zakhari S, Hoek JB. Alcohol and breast cancer: reconciling epidemiological and molecular data. Adv Exp Med Biol. 2015;815:7–39.
Ferlay J, Shin HR, Bray F, Forman D, Mathers C, Parkin DM. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer. 2010;127:2893–917.
Jemal A, Thomas A, Murray T, Thun M. Cancer statistics, 2002. CA Cancer J Clin. 2002;52:23–47.
Salaspuro M. Microbial metabolism of ethanol and acetaldehyde and clinical consequences. Addict Biol. 1997;2:35–46.
Seitz HK, Simanowski UA, Garzon FT, Rideout JM, Peters TJ, Koch A, Berger MR, et al. Possible role of acetaldehyde in ethanol-related rectal cocarcinogenesis in the rat. Gastroenterology. 1990;98:406–13.
Yokoyama A, Muramatsu T, Ohmori T, Yokoyama T, Okuyama K, Takahashi H, Hasegawa Y, et al. Alcohol-related cancers and aldehyde dehydrogenase-2 in Japanese alcoholics. Carcinogenesis. 1998;19:1383–7.
Stagos D, Chen Y, Brocker C, Donald E, Jackson BC, Orlicky DJ, Thompson DC, et al. Aldehyde dehydrogenase 1B1: molecular cloning and characterization of a novel mitochondrial acetaldehyde-metabolizing enzyme. Drug Metab Dispos. 2010;38:1679–87.
Singh S, Brocker C, Koppaka V, Chen Y, Jackson BC, Matsumoto A, Thompson DC, et al. Aldehyde dehydrogenases in cellular responses to oxidative/electrophilic stress. Free Radic Biol Med. 2013;56:89–101.
Hess DA, Meyerrose TE, Wirthlin L, Craft TP, Herrbrich PE, Creer MH, Nolta JA. Functional characterization of highly purified human hematopoietic repopulating cells isolated according to aldehyde dehydrogenase activity. Blood. 2004;104:1648–55.
Langan RC, Mullinax JE, Ray S, Raiji MT, Schaub N, Xin HW, Koizumi T, et al. A pilot study assessing the potential role of non-CD133 colorectal cancer stem cells as biomarkers. J Cancer. 2012;3:231–40.
Carpentino JE, Hynes MJ, Appelman HD, Zheng T, Steindler DA, Scott EW, Huang EH. Aldehyde dehydrogenase-expressing colon stem cells contribute to tumorigenesis in the transition from colitis to cancer. Cancer Res. 2009;69:8208–15.
Chen Y, Orlicky DJ, Matsumoto A, Singh S, Thompson DC, Vasiliou V. Aldehyde dehydrogenase 1B1 (ALDH1B1) is a potential biomarker for human colon cancer. Biochem Biophys Res Commun. 2011;405:173–9.
McCart AE, Vickaryous NK, Silver A. Apc mice: models, modifiers and mutants. Pathol Res Pract. 2008;204:479–90.
Zheng Y, Kramer PM, Lubet RA, Steele VE, Kelloff GJ, Pereira MA. Effect of retinoids on AOM-induced colon cancer in rats: modulation of cell proliferation, apoptosis and aberrant crypt foci. Carcinogenesis. 1999;20:255–60.
Wargovich MJ, Jimenez A, McKee K, Steele VE, Velasco M, Woods J, Price R, et al. Efficacy of potential chemopreventive agents on rat colon aberrant crypt formation and progression. Carcinogenesis. 2000;21:1149–55.
Li Y, Mao Y, Zhang Y, Cai S, Chen G, Ding Y, Guo J, et al. Alcohol drinking and upper aerodigestive tract cancer mortality: a systematic review and meta-analysis. Oral Oncol. 2014;50:269–75.
Zakhari S, Vasiliou V, Guo QM, editors. Alcohol and cancer. New York: Springer; 2011.
Vasiliou V, Zakhari S, Seitz HK, Hoek JB, editors. Biological basis of alcohol-induced cancer. Cham: Springer; 2015.
Asakage T, Yokoyama A, Haneda T, Yamakazi M, Muto M, Yokoyama T, et al. Genetic polymorphisms of alcohol and aldehyde dehydrogenases, and drinking, smoking and diet in Japanese men with oral and pharyngeal squamous cell carcinoma. Carcinogenesis. 2007;28:865–74.
Yokoyama A, Muramatsu T, Ohmori T, Makuuchi H, Higuchi A, et al. Multiple primary esophageal and concurrent upper aerodigestive tract. Cancer Epidemiol Biomarkers Prev. 2006;15:696–703.
Visapaa JP, Gotte K, Benesova M, Li J, Homann N, Conradt C, et al. Increased cancer risk in heavy drinkers with the alcohol dehydrogenase 1C*1 allele, possibly due to salivary acetaldehyde. Gut. 2004;53:871–6.
Homann N, Stickel F, Konig IR, Jacobs A, Junghanns K, et al. Alcohol dehydrogenase 1C*1 allele is a genetic marker for alcohol-associated cancer in heavy drinkers. Int J Cancer. 2006;118:1998–2002.
Schwartz SM, Doody DR, Fitzgibbons ED, Ricks S, Porter PL, Chen C. Oral squamous cell cancer risk in relation to alcohol consumption and alcohol dehydrogenase-3 genotypes. Cancer Epidemiol Biomarkers Prev. 2001;10:1137–44.
Peters ES, McLean MD, Liu M, Eisen EA, Mueller N, Kelsey KT. The ADH1C polymorphism modifies the risk of squamous cell carcinoma of the head and neck associated with alcohol and tobacco use. Cancer Epidemiol Biomarkers Prev. 2005;14:476–82.
Chen YJ, Chen C, Wu DC, Lee CH, Lee JM, Goan YG, et al. Interactive effect of lifetime alcohol consumption and alcohol and aldehyde dehydrogenase polymorphisms on esophageal cancer risks. Int J Cancer. 2006;119:2827–31.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Zakhari, S., Radaeva, S., Vasiliou, V. (2016). Hepatic and Extrahepatic Malignancies in Alcoholic Liver Disease. In: Chalasani, N., Szabo, G. (eds) Alcoholic and Non-Alcoholic Fatty Liver Disease. Springer, Cham. https://doi.org/10.1007/978-3-319-20538-0_13
Download citation
DOI: https://doi.org/10.1007/978-3-319-20538-0_13
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-20537-3
Online ISBN: 978-3-319-20538-0
eBook Packages: MedicineMedicine (R0)