Abstract
The lipid bilayer that constitutes cell membranes imposes environmental constraints on the structure, folding and function of integral membrane proteins. The cell membrane is an enormously heterogeneous and dynamic system in its chemical composition and associated physical forces. The lipid compositions of cell membranes not only vary over the tree of life but also differ by subcellular compartments within the same organism. Even in the same subcellular compartment, the membrane composition shows strong temporal and spatial dependence on the environmental or biological cues. Hence, one may expect that the membrane protein conformations and their equilibria strongly depend on the physicochemical variables of the lipid bilayer. Contrary to this expectation, the structures of homologous membrane proteins belonging to the same family but from evolutionary distant organisms exhibit a striking similarity. Furthermore, the atomic structures of the same protein in different lipid environments are also very similar. This suggests that certain stable folds optimized for a specific function have been selected by evolution. On the other hand, there is growing evidence that, despite the overall stability of the protein folds, functions of certain membrane proteins require a particular lipid composition in the bulk bilayer or binding of specific lipid species. Here I discuss the specific and nonspecific modulation of folding, misfolding and function of membrane proteins by lipids and introduce several diseases that are caused by misfolding of membrane proteins.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Membrane protein folding and misfolding
- Membrane protein misfolding diseases
- Membrane protein topology
- Lipid-protein interaction
- Bilayer curvature stress
- Bilayer lateral pressure profile
1.1 Introduction
At the border between life and environment, membrane proteins carry out numerous critical cellular processes such as energy generation, uptake and secretion of metabolites, maintenance of ion balance, signal transduction, catalysis, cell-cell communication, and more.
Protein folding occurs through a delicate balance of various driving forces, which is largely determined by the folding environment. While water-soluble proteins fold in an isotropic aqueous medium, membrane proteins fold in a heterogeneous anisotropic lipid bilayer. For water-soluble proteins, whose folding is majorly driven by the hydrophobic effect, achieving the integrity of the hydrophobic core is crucial for the successful folding (Dill 1990). Effective hydration of the protein surface is an important factor that determines the protein’s propensity toward misfolding and aggregation (Chong and Ham 2014). The exposed hydrophobic surfaces during the folding and unfolding are major targets for chaperones and degradation machinery (Wickner et al. 1999). On the other hand, membrane proteins are stabilized and function in a highly anisotropic and chemically heterogeneous lipid bilayer. Then, what are the roles of the lipid bilayer and its individual components in the structure, folding and function of membrane proteins? Are lipids just a passive solvent that mediates the assembly of membrane proteins or active modulators of the protein structure, folding and stability?
Membrane proteins can be classified as α-helical or β-barrel types depending on the secondary structural elements within the membrane (Fig. 1.1). α-helical proteins are distributed in the cytoplasmic membranes of prokaryotes and eukaryotes, and in the membranes of subcellular compartments of eukaryotes (Popot and Engelman 2000). β-barrel proteins dominate the outer membranes of Gram-negative bacteria and also exist in the outer membranes of mitochondria and chloroplasts (Tamm et al. 2004). While both types of proteins are crucial in the maintenance of the cellular function, they fold in the membranes by entirely different mechanistic and thermodynamic principles. The main focus of this chapter is on the role of the lipid environment in the folding, stability and function of α-helical membrane proteins, which are more widespread in kingdoms of life. However, valuable general folding principles have been obtained from the folding studies of β-barrel membrane proteins, which are also described.
Thermodynamic folding models of integral membrane proteins. (a) Two stage model of α-helical membrane protein folding (Engelman et al. 2003; Popot and Engelman 1990). At the first step, individually stable hydrophobic transmembrane segments are integrated into the membrane via the hydrophobic effect. At the second step, individual helices assemble into a functional 3D structure. (b) Folding of β-barrel membrane proteins can be described as a cooperative step, where the bilayer insertion and folding are coupled (Hong and Tamm 2004; Moon et al. 2013; Huysmans et al. 2010)
1.2 Folding of α-Helical Membrane Proteins
1.2.1 How Does Lipid Bilayer Constrain the Structure of Membrane Proteins?
Transmembrane (TM) segments of a polypeptide chain are largely composed of hydrophobic amino acids for their favorable partitioning into the nonpolar core of the lipid bilayer (White and Wimley 1999). The hydrophobic nature of α-helical membrane proteins allows the prediction of the TM segments from their amino acid sequences by using the hydropathy plot based on various hydrophobicity scales (Popot and Engelman 2000). In addition, the peptide groups of individual TM segments form ordered α-helical structure to fulfill the hydrogen bond-forming capability, and thereby reduce the desolvation cost in burying the polar peptide group within the nonpolar environment (Bental et al. 1997). Although the hydrophobic thickness of the cell membranes (the distance between phosphate groups in phospholipid bilayers) is maintained within 30–45 Å in various organisms and subcellular compartments (Mitra et al. 2004), the lengths of the TM helices of known structures vary widely from 14 to 36 residues (Bowie 1997). Longer helices are adapted by tilting their axis relative to the bilayer normal or by bending or kinking, while shorter helices can be stabilized by the tertiary contacts with neighboring helices or induce local thinning of the bilayer. The TM helices are packed against each other preferentially at an angle around 20°, while the packing angles for water-soluble proteins widely vary from about −40° to 30° (Bowie 1997). The tertiary contacts are predominantly made by the packing of the TM helices, but the prosthetic groups and folded or unstructured interhelical loops also contribute to the packing.
1.2.2 General Features in the Folding of α-Helical Membrane Proteins
α-helical TM segments are extremely resistant to heat and chemical denaturants (Haltia and Freire 1995). Thus, individual α-helices in the helix bundle membrane proteins can be regarded as independent folding domains packed with one another to form a native 3D structure. Based on the denaturation/refolding studies and structural information of membrane proteins known until 1980s, Popot and Engelman proposed the two-stage model, in which the folding of α-helical membrane proteins was divided into two energetically distinct steps (Fig. 1.1, left) (Engelman et al. 2003; Popot and Engelman 1990). This model greatly simplifies the folding problem and has served as a conceptual framework in studying membrane protein folding. The two proposed stages are as follows.
-
1.
Insertion of independently stable α-helical segments into the membrane. This step is largely driven by the hydrophobic effect (Hessa et al. 2005, 2007). In cells, the TM segments, which are targeted by the signal recognition particle (SRP) complex, are co-translationally inserted into the membrane through a membrane protein complex called the translocon (SecYEG in the inner membranes of E. coli, Sec61αβγ in the endocytoplasmic reticulum (ER) membranes in eukaryotes, and SecYβE in archaea) (Park and Rapoport 2012; du Plessis et al. 2011). The translocon acts as a thermodynamic machine integrating hydrophobic nascent segments into the membrane through the lateral gate or by passing the hydrophilic segments across the membrane through the vertical channel (White and von Heijne 2008).
-
2.
Association of the TM helices to form a native 3D structure. In this step, the hydrophobic effect cannot strongly drive the condensation of the TM helices because water molecules are scarce in the core of the lipid bilayer (Joh et al. 2009). Hence, other forces such as van der Waals and polar interactions must drive the folding. Although polar interactions may be strong due to the low dielectric constant in the nonpolar bilayer environment, many experimental results indicate that the strengths of interhelical hydrogen bonds are comparable to those in water-soluble proteins (Bowie 2011). Structural and statistical analyses of membrane proteins of known structure revealed that membrane proteins tend to bury more fractional area of side chains than water-soluble proteins (Oberai et al. 2009). Extensive packing around small side chains of Gly, Ala, Ser and Thr is an important feature of the interior of membrane proteins, whereas the hydrophobic core packing in water-soluble proteins prefers larger nonpolar side chains and aromatic residues (Eilers et al. 2000; Adamian and Liang 2001; Adamian et al. 2005).
Although our knowledge on the driving forces for membrane protein folding is rapidly growing, it is still at the fledgling stage compared to that of water-soluble proteins. There are relatively unexplored forces such as aromatic-aromatic interactions (Burley and Petsko 1986), π-cation interactions (Gallivan and Dougherty 1999) and the polar nature of the polypeptide backbone that may contribute to the folding of membrane proteins (Wimley and White 1992, 1996).
1.3 Chemical and Physical Properties of Lipid Bilayer
1.3.1 Chemical Properties of Cell Membranes
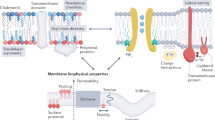
Since Singer and Nicholson proposed the first unified model of the cell membranes (Singer and Nicolson 1972), the model has been substantially updated (Engelman 2005) (Fig. 1.2). While the membrane is crowded with proteins whose amount is comparable to that of lipids (protein-to-lipid mass ratios vary in the range of 0.2–4) (Lodish et al. 2000), membrane proteins associate transiently or tightly with one another to form structural and functional complexes (Wu et al. 2003). Their diffusion within the membrane plane is significantly affected by cytoskeletons anchored on the membrane and by the “lipid rafts” or membrane domains enriched with cholesterol and sphingomyelin (Kusumi et al. 2005; Winckler et al. 1999; Simons and Sampaio 2011). The hydrophobic thicknesses of integral membrane proteins do not always match those of the lipid bilayer so that the resultant membrane thickness is reciprocally modulated by both proteins and lipids (Mitra et al. 2004).
Models of cell membranes. (a) Original “fluid-mosaic” model (Singer and Nicolson 1972). The solid bodies represent integral membrane proteins that randomly diffuse in a fluid phase of the lipid bilayer. Some integral membrane proteins form complexes. There are no membrane-associated structures and no phase separation of lipids (domains) with different lipid compositions. (b) Updated membrane model showing membrane domains of various sizes with associated cytoskeletons, glycosylated lipids and proteins, and with asymmetric lipid composition (Nicolson 2014). Peripheral and integral membrane proteins often form complexes, and some preferentially partition into membrane domains or domain interfaces (Adopted with permission from Nicolson 2014)
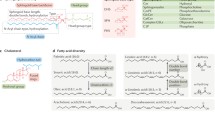
Lipid components that play structural roles in cell membranes are classified as phosphoglycerolipids, sphingolipids, cholesterol and cardiolipin, depending on the backbone structure to which fatty acyl chains and polar head groups are attached (Spector and Yorek 1985) (Table 1.1). The lipid composition of cell membranes is highly variable and dynamic depending on the species, subcellular compartment within a cell, tissue in a body, developmental stage, environmental and physiological conditions, and disease state (Dawidowicz 1987; Chan et al. 2012). Furthermore, even within the same subcellular compartment, the lipid composition is vastly different between the inner and outer leaflets of the membrane (van Meer et al. 2008). This lipid asymmetry results from the balanced action of various flippases, which cause the asymmetry, and scramblases, which equilibrate the lipid distribution between the two leaflets (Clark 2011).
1.3.2 Physical Properties of Lipid Bilayers: Lipid Polymorphism and Lateral Pressure Profile
Isolated lipid components of cell membranes form various forms of mesoscopic assemblies in water (Seddon and Templer 1995). The key concept underlying the “lipid polymorphism” is that the structure of the lipid assembly depends on the shape of the lipid molecules, e. g. the relative volumes of the head group and the hydrocarbon chains (Kumar 1991) (Fig. 1.3). For example, phosphatidylcholines (PC) lipids with relatively large head groups have a cylindrical shape so that such lipids in water spontaneously form lamellar phase, Lα, comprised of multiple stacks of relatively flat bilayers. Phosphatidylethanolamine (PE) lipids with a small head group and longer unsaturated acyl chains have the inverted cone shape and prefer the inverted hexagonal phase, HΙΙ (Shyamsunder et al. 1988; Gruner et al. 1985). The HΙΙ phase can be envisioned as stacks of cylindrical lipid arrays, with hydrocarbon chains pointing outward and polar head groups surrounding the central aqueous pore. Cardiolipin lipids (CLs) form the HII phase in the presence of divalent ions. Single-chain lipids or detergents with a cone shape (i. e. relatively large head group compared to the acyl chain cross-section) form the hexagonal phase, HΙ, with the molecular arrangement similar to that in micelles.
Lipid polymorphism and lateral pressure profile. (a) Aqueous aggregates of lipids show different mesoscopic phase behavior depending on the relative size of the lipid head group and the acyl chains. PC, which has relatively large head group, forms lamellar phase, Lα. PE with smaller head group forms inverted hexagonal phase, HII. Lyso-PC, which has only one acyl chain, forms micellar or hexagonal phase, HI (Escriba et al. 1997). C spont represents spontaneous monolayer curvature. (b) Lateral pressure profile (p) along the bilayer normal (z). A tension-free bilayer is maintained by the balanced forces of the head group repulsion, interfacial tension and chain repulsion (The plot for lateral pressure profiles was adopted with permission from Cantor 1997)
Intrinsic curvature hypothesis (Gruner 1985). When lipids with positive or negative intrinsic spontaneous curvature are incorporated into the bilayer, a curvature stress is induced in each monolayer leaflet. The origin of this phenomenon is the change in the lateral pressure profile depending on the lipid composition (The plot for lateral pressure profiles was adopted with permission from Cantor 1997)
The lipid bilayer at its free energy minimum is maintained by several balanced forces (Cantor 1997, 1999) (Fig. 1.3b): (1) Line tension at the interface between nonpolar hydrocarbon chains and polar head groups. This attractive force originates from the hydrophobic effect of burying the nonpolar chains away from water. (2) Repulsions between hydrocarbon chains within the bilayer core and between head groups, which are caused by the dynamic collisions in each region. These balancing forces generate the characteristic lateral pressure profile along the bilayer normal (z-axis). In a tension-free bilayer, the integration over the z-axis yields zero net pressure. The lateral pressure profile has not been determined experimentally but has been calculated using the mean-field theory (Cantor 1997). Surprisingly, the calculated lateral pressure in the bilayer core amounts to several hundred atmospheres.
The lateral pressure profile is the physical origin of an important elastic property of the bilayer called “curvature stress” (Gruner et al. 1985) (Fig. 1.3). For example, in the lamellar phase of a PC bilayer, incorporation of PE reduces the head group repulsion because of the small PE head group and the hydrogen bonding between the ethanolamine group in PE and the phosphate group in an adjacent lipid (Gruner et al. 1988; Tate and Gruner 1987). However, under the constraint of the zero integrated pressure without a change in the attractive line tension, the repulsion between hydrocarbon chains will increase to compensate the reduced head group repulsion. The overall lateral pressure profile will be redistributed to induce the spontaneous negative curvature in each monolayer leaflet (Cantor 1999). Interestingly, the cell membranes, which are regarded as “lamellar phase”, are enriched with non-bilayer-forming PE and CL, suggesting an important role of the curvature stress and lateral pressure profile in the structure and function of membrane proteins (van den Brink-van der Laan et al. 2004b). Indeed, the lipid composition of E. coli cell membranes is maintained near the lamellar-to-inverted hexagonal phase transitions (Morein et al. 1996).
This suggests that integral membrane proteins are constantly subjected to mechanical stress, and the changes in the pressure profile may significantly impact their folding, stability and conformational equilibria. Indeed, modification of the lateral pressure profile was suggested to underlie the mechanisms of general anesthesia by modulating the activity of several ion channels, the sensation of mechanical stimuli by mechanosensitive channels (Milutinovic et al. 2007; Perozo et al. 2002b), the oligomerization and stability of the pH-gated potassium channel KcsA (van den Brink-van der Laan et al. 2004a) and the stabilization of fusion intermediates in the viral entry and the intracellular vesicular trafficking (Siegel 1993; Yang and Huang 2002). More evidence on this subject will be discussed in Sect. 1.5.
1.4 Lipid-Integral Membrane Protein Interactions for Folding, Structure and Function
In cell membranes, the lipid bilayer serves as a medium that supports the folding, structure and function of membrane proteins. How do constituent lipid molecules in the bilayer interact with proteins? Do they tightly bind to the protein or rapidly exchange with the bulk lipids? Are there binding sites for specific lipid molecules on the surface of membrane proteins? These questions are important for the following reasons. (1) Lipid binding selectivity and specificity could explain the heterogeneous lipid composition of cell membranes that may have evolved for an optimal function of the membrane proteome, or vice versa. (2) Lipid selectivity for a specific membrane protein may be implicated in the protein stability and function. (3) Specific protein-lipid interactions may explain the sorting of membrane proteins between different co-existing membrane domains.
1.4.1 Lipid Contact with Membrane Proteins: Annular Lipids
The first shell of lipids that surround membrane proteins, called “annular lipids” (Contreras et al. 2011), connect the proteins to bulk lipids. Electron paramagnetic resonance (EPR) spectroscopy using spin-labeled lipids revealed that lipid molecules in proteoliposomes exist in restricted (annular) and mobile (bulk) forms (East et al. 1985; Knowles et al. 1979). The two distinct lipid fractions exchange with a time scale of 10–100 ns, while the exchange between unperturbed bulk lipids occurs one to two order of magnitude faster. From the analysis of various-size proteins reconstituted mostly in PCs, approximately two lipid molecules bind to one TM helix on average (Marsh 2008). Thus, the surfaces of integral membrane proteins make favorable but transient contacts with “solvating” lipid molecules.
The selectivity for specific lipid species was estimated by measuring Trp fluorescence quenching (London and Feigenson 1981) or the changes in the immobile lipid fraction from EPR spectra (Powell et al. 1985) by using the competition between a reference lipid (typically PC) and spin-labeled or brominated lipids with various head groups and acyl-chain lengths (Krishnamachary et al. 1994). Lipid selectivity varies for the tested membrane proteins (Marsh 2008). For example, cytochrome c oxidase, which is the last enzyme in the electron transfer pathway in mitochondria and bacteria, strongly prefers CL, while PE, PS and PG bind less selectively (see Sect. 1.4.2 for more details). Mitochondrial ADP/ATP carrier strongly binds CL, PA and stearic acid. While sarcoplasmic reticulum Ca2+ ATPase does not show a noticeable preference for any lipid, PE and stearic acid are excluded from the protein surface. On the other hand, rhodopsin does not show any preference for the tested lipids (Marsh 2008).
Further experimental support for the existence of annular lipids was obtained from crystallographic studies of several membrane proteins (Lee 2011). Structural studies of membrane proteins often involve rigorous delipidation steps during purification, such as detergent solubilization and detergent exchange, as well as the addition of external lipids. Reconstitution in lipid environments is necessary to obtain 2D crystals for electron crystallography and 3D crystals for X-ray crystallography in lipid cubic or bicelle phase crystallizations. However, the electron densities of bound lipids obtained in those conditions are often either not observed or weak, hindering a straightforward modeling.
The first detailed pictures of the annular lipids were presented by the structures of bacteriorhodopsin (bR) from Halobacteria. These structures were solved by electron crystallography using 2D crystals in natural purple membranes (Grigorieff et al. 1996) and by X-ray crystallography using 3D crystals prepared by the detergent-induced fusion of 2D crystals and the lipid-cubic phase crystallization (Luecke et al. 1999; Takeda et al. 1998). A total of 18 lipid chains was identified from the structure in the lipid cubic phase at a resolution of 1.55 Å, including four diether lipids and one squalene per monomer, although the unequivocal identification of all lipids was not possible (Fig. 1.5a) (Luecke et al. 1999).
Annular lipids observed in the crystal structures of bacteriorhodopsin (bR) and aquaporin-0 (AQP0). (a) Atomic structure of bR in the cubic lipid phase determined at 1.55 Å resolution by X-ray crystallography (PDB ID: 1C3W) (Luecke et al. 1999). The annular lipids identified in the structure originate from the native purple membranes. (b) Left: AQP0 structure in DMPC (diC14:0PC) solved by 2D electron crystallography at 1.9 Å (PDB ID: 2B6O) (Gonen et al. 2005). Right: AQP0 structure in polar lipid extracts from E. coli solved by 2D electron crystallography at 2.5 Å (PDB ID: 3M9I) (Figures modified with permission from Luecke et al. 1999 and Hite et al. 2010)
More recent observation of annular lipids bound to aquaporin-0 (AQP0), which is abundant in lens fiber cells, using electron crystallography is intriguing. Walz group solved the structures of AQP0 in 2D crystals reconstituted in two different lipids, DMPC (diC14:0PC) and E. coli phospholipids whose most abundant lipid species is C16:0C16:1c9PE (Hite et al. 2010; Gonen et al. 2005) (Fig. 1.4b). Strikingly, the two structures in different lipid environments were essentially the same (0.48 Å root mean square deviation for the backbone atoms) and exhibited the same number of bound lipid molecules (seven lipids per monomer, four in the extracellular leaflet and three in the cytoplasmic leaflet) at similar lipid-protein interfaces.
Although the exact conformations of the corresponding lipids in the two structures were slightly different, the results strongly suggest that there are generic lipid binding motifs on the surface of membrane proteins that can accommodate lipids with varying head groups and acyl chain lengths. Furthermore, the bilayer thicknesses in the two environments deduced from the average distance between the annular lipids in two leaflets are significantly different (27.0 Å in E. coli lipids and 31.2 Å in diC14:0PC) (Hite et al. 2010). This difference may be due to the cis-unsaturation of E. coli lipids (~52 % of acyl chains) and the resulting thinning of the bilayer. Despite this difference, the protein structure remained unchanged, implying the adaptation of lipid conformation to the protein. Similar lipid shells around the membrane protein surface have been identified in the cytochrome bc 1 complex (Lange et al. 2001).
1.4.2 Lipid Contacts with Membrane Proteins: Non-annular Lipids
In contrast to the annular lipids that surround the protein as the first shell, another class of bound lipids, called non-annular, interact with the protein more specifically (Contreras et al. 2011; Lee 2011) (Figs. 1.6 and 1.7). These crystallographically identified lipid molecules often originate from native membranes despite rigorous protein delipidation. Therefore, these lipid molecules bind to the protein with high affinity and specificity and may help stabilize the protein. Although it is difficult to distinguish non-annular from annular lipids just by looking at the structure, several criteria can be applied (Contreras et al. 2011). (1) Non-annular lipids play structural roles within the membrane protein complexes, e. g. in the buried locations of the protein matrix, or at the protein-protein interfaces. (2) Non-annular lipids tightly bind to the protein. (3) Non-annular lipids may play a functional role as allosteric effectors of enzymatic activity. In this section, the role of specific binding of a phospholipid in the structural integrity of a membrane protein will be described. Specific binding of cholesterol will be discussed in Sects. 1.6.3, 1.6.4, and 1.6.5.
Non-annular lipids in protein structures. Left: Non-annular lipids specifically bound to cytochrome bc 1 complex (PDB ID: 1 KB9, 2.5 Å resolution) (Lange et al. 2001). Only the membrane-spanning subunits are shown (solid ribbons). Upper right: Cardiolipin binding site (CL in stick, residues in space-filling model) is formed at the subunit interface between cytochrome b (COB, green) and cytochrome c 1 (CYT, cyan). Bound CL is stabilized by the electrostatic interactions with Lys288 and Lys289 from CYT and by hydrogen bonding to Tyr28, Trp29 and Lys228 from COB, and to Tyr281 from CYT. Lower left: Acyl chain of phosphatidylinositol (PI) is wrapped around the Rieske protein (RIP1) TM domain (red). The PI head group makes contacts with the COB, CYT and RIP1 subunits (Figures modified with permission from Lange et al. 2001)
Lipids in the structure of cytochrome c oxidase. Top: All 26 lipids (shown in sticks) identified in the crystal structure of cytochrome c oxidase complex (PDB ID: 1KB9, 1.8 Å resolution) (Shinzawa-Itoh et al. 2007). Bottom left: Cardiolipin (red), PE (blue) and PG (in green) stabilize the dimer interface. Subunits from two different monomers are in grey and yellow. Bottom right: Two PGs and one PE are deeply embedded into subunit III, stabilizing the tertiary interactions (Figures modified with permission from Shinzawa-Itoh et al. 2007)
The crystal structure of yeast cytochrome bc 1 complex, a mitochondrial inner membrane protein in the electron transfer pathway of the respiratory chain, revealed interesting structural and functional role of specific bound lipids (Lange et al. 2001) (Fig. 1.6). Out of five lipid molecules (two PEs, one PC, one PI and one CL), all of which are natural membrane components, the acyl chains of PI are wrapped around the TM helix of the Rieske protein subunit (RIP1 red), and the head group forms extensive hydrogen bonds to the polar side chains in the interfacial TM regions of cytochrome b (COB, green) subunit (Fig. 1.6, lower right). The double-negatively charged head group of CL is stabilized by two positively charged “clamp” residues, Lys288 and Lys289 of cytochrome c 1 (CYT1, cyan), packing against Tyr28 of COB subunit (Fig. 1.6, upper right); this packing involves a water-mediated hydrogen bond with Lys228, and a hydrogen bond with Trp29 of COB subunit. The four acyl chains are also making extensive van der Waals contacts with the large flat surface comprised of three TM helices from COB and one TM helix from CYT. Double or triple mutations of three Lys residues led to severe growth defects and dramatically reduced the protein expression level, suggesting the role of CL binding in the structural and functional integrity of the protein (Lange et al. 2001).
The 1.8 Å-resolution crystal structure of bovine cytochrome c oxidase is also a wonderful showcase of structurally and functionally important phospholipids (13 identified lipids per monomer in the dimer, including 2 CLs, 1 PC, 3 PEs, 4 PGs and 3 triglycerides) (Shinzawa-Itoh et al. 2007) (Fig. 1.7). Three lipids (two PG in green and one PE in blue) bound to subunit III are the most rigidly bound among all 13 lipid molecules (Fig. 1.7, bottom right). On average, the head group and acyl chains of each lipid are stabilized by 12 hydrogen bonds and 98.3 van der Waals contacts, and the B-factors of these lipids, which reflect molecular ordering, are comparable to those of the protein around the lipid binding site. Structurally, these lipids are buried within the protein matrix, mediating the packing of TM1 and TM2 helices of subunit III with the rest of the subunit. Functionally, the two PG lipids lie in the predicted pathway of molecular oxygen. The conformation of acyl chains was suggested to dynamically modulate the passage of O2 into the protein. Another class of lipids (one CL in red, two Pes in blue, and one PG in green) participates in the stabilization of the dimer and the subunit interface (Fig. 1.7, bottom left). Specially, CL is deeply inserted into the interface and makes favorable contacts with four subunits (subunits III and VIa from one monomer and subunits I and II from the other monomer).
Specific lipid-protein interactions of structural and functional importance have also been inferred from the crystal structures of several other membrane proteins including tetrameric K+-channel, KcsA (Zhou et al. 2001; Valiyaveetil et al. 2002), photosystem II complex (Guskov et al. 2009), nitrate reductase A (Bertero et al. 2003), photosynthetic reaction center from Rhodobacter sphaeroides (Fyfe and Jones 2005), Na+K+-ATPase (Shinoda et al. 2009), etc.
1.4.3 Identification of Stabilizing Lipids Using Mass Spectrometry
Recent advances in mass spectrometry made possible the quantitative identification of the high-affinity lipid molecules and their stabilizing role in the gas phase (Laganowsky et al. 2013; Barrera et al. 2008). The key to the success was the combination of the following implements. (1) Optimizing the number of collisions of nano-electrospray droplets (nano-ESI). By this step, detergents are released from protein-detergent-lipid aggregates while maintaining the intact protein complexed with high-affinity lipid molecules. (2) Employment of ion mobility mass spectrometry (IM-MS) for measuring the rotationally-averaged collision cross-section (CSS). CSS is sensitive to the protein shape and, therefore, to the conformational state (e. g. folded or unfolded) (Bush et al. 2010). (3) Use of non-ionic detergents that best support the structure and function of the folded protein. By measuring the changes in CCS values as a function of collision voltages, the “unfolding transition curve” can be constructed for measuring the stability of membrane proteins in the gas phase.
Using IM-MS, Laganowsky et al. (2013) determined the differential effects of various lipid species on the stability of mechanosensitive channels of large conductance (MscL), aquaporin Z (AqpZ) and the ammonia channel (Amt) from E. coli. Notably, identification of the high-affinity binding of CL to AqpZ led to the demonstration of the pivotal role of CL in water transport in vitro. Identification of the high-affinity binding of PG to Amt also enabled the elucidation of the lipid role in stabilizing the structure of Amt at the subunit interface. Therefore, this method provides a promising tool for studying the role of specific lipids in the structure, stability and function of membrane proteins. However, it is not clear whether the bound lipids identified in these studies are annular or non-annular.
1.5 Role of Physical Properties of Lipid Bilayers in the Folding and Assembly of Integral Membrane Proteins
How do the chemical and physical properties of cell membranes influence the folding, stability and conformational equilibria of membrane proteins? Researchers have approached this problem from two viewpoints (Lee 2011). First, individual lipid components influence protein conformation by modulating the physical properties of lipid bilayers such as lateral pressure profile, curvature stress, and local lipid deformation by hydrophobic mismatch. Second, lipid components take effects by selective binding to proteins. Differentiating these two distinct phenomena is not an easy task and requires careful control experiments. There is a wealth of experimental data that support either mechanism; the common conclusion is that the lipid bilayer is actively involved in the folding, structure and function of membrane proteins.
1.5.1 Role of Physical Properties in the Function of Membrane Proteins
A classic example connecting physical properties of a bilayer to the conformation and function of membrane proteins is the work by (Keller et al. 1993) who studied the ion channel activities of alamethicin as a function of the curvature stress of the lipid bilayer. Alamethicin is a 20-amino acid antimicrobial peptide from fungus Trichderma viride, which acts as a voltage-gated ion channel by forming oligomeric helical bundles (higher than pentamer) (Cafiso 1994; Tieleman et al. 2002). The curvature stress was modulated by increasing the fraction of diC18:1c9PE relative to diC18:1c9PC in the planar bilayer. diC18:1c9PE has a high tendency to form HII phase with a negative spontaneous curvature of the monolayer (lamellar-to-hexagonal transition temperature 15 °C). The spontaneous curvature was strongly correlated with the probability of the higher-conductance open states (Keller et al. 1993). This result suggests that the increased curvature stress induced the formation of higher-order oligomeric states, leading to the enhanced channel activity. The physical origin of the curvature-induced enhancement of channel activity is not well understood. Similar enhancement of membrane transport induced by PE was also observed for sarcoplasmic Ca2+-ATPase reconstituted in diC18:1c9PE-containing lipid vesicles (Navarro et al. 1984).
Another example is the dependence of the activity of mechanosensitive channel of large conductance (MscL) on lipid composition. In PC vesicles, the channel exists in a closed state. Addition into the outer leaflet of lysoPC, which by itself forms a micellar phase (HI), induces the channel opening (Perozo et al. 2002a). When incorporated into the bilayer, lysoPC induces the positive spontaneous monolayer curvature, and the resulting asymmetric distribution of the lateral pressure profile drives the opening of MscL (Perozo et al. 2002a). Further, electron-paramagnetic resonance (EPR) studies suggested that the addition of lysoPC changes the tilt angle of the TM helices. The free energy difference between the closed and open states significantly increases as the bilayer thickness increases, from 4 RT for diC16:1c9PC, to 9 RT for diC18:1c9PC, and 20 RT for diC20:1c9PC, where RT is free energy of thermal motion (Perozo et al. 2002b). This result implies that the hydrophobic thicknesses of the open and closed states of MscL are significantly different, and the lipid deformation by the hydrophobic mismatch between MscL and the bilayer modulates the conformational equilibrium of the channel. Indeed, the estimated lipid deformation energy is similar to the free energy differences between the open and closed states of the channel (Wiggins and Phillips 2004).
1.5.2 Role of Physical Properties in the Folding Kinetics of Membrane Proteins
Physical forces of the lipid bilayer also modulate the kinetics and thermodynamics of membrane proteins folding. The first connection between the two subjects was made by Booth group using bacteriorhodopsin as a model (Booth et al. 1997; Curran et al. 1999). The refolding yield of bacteriorhodopsin (bR) decreased as the fraction of PE in the lipid bilayer increased (Curran et al. 1999). PE induces the negative monolayer curvature and thus, increases the lateral pressure in the bilayer core, which was measured by the increase in the excimer fluorescence of pyrene-labeled lipids at the acyl chain region.
Inhibition of refolding into PE-containing lipid bilayers was also observed for bacterial β-barrel outer membrane proteins. In contrast to the α-helical membrane proteins that are largely composed of stretches of hydrophobic TM segments, β-barrel membrane proteins contain residues that have alternating hydrophobicity in their TM domains. As a consequence, the barrel lumen is composed of hydrophilic residues whereas the lipid-contacting surface is largely hydrophobic (Tamm et al. 2004; Wimley 2003). Because β-barrel membrane proteins are less hydrophobic, they can be solubilized in high concentrations of urea or GdnHCl in the fully unfolded state, and then refolded into lipid vesicles by dilution of denaturants. While a number of OMPs such as OmpA (Kleinschmidt and Tamm 1996, 2002), OmpLA (Moon and Fleming 2011), OmpW (Moon et al. 2013), PagP (Moon et al. 2013; Huysmans et al. 2010), OmpX (Gessmann et al. 2014), etc. can readily refold into thin PC bilayers (Burgess et al. 2008), the incorporation of PE significantly slows down the refolding kinetics (Gessmann et al. 2014; Patel and Kleinschmidt 2013). Probably the increased lateral pressure within the nonpolar core of the bilayer increases its compressibility modulus, thereby decreasing the lipid packing defects necessary for the protein insertion.
Interestingly, incorporation into lipid vesicles of the barrel assembly machinery complex (BAM), which is a conserved assembly factor of OMPs in the bacterial outer membranes, enhanced the refolding rate and yield (Gessmann et al. 2014; Patel and Kleinschmidt 2013). The crystal structures and molecular dynamics simulations of two homologues of BamA, a core component of the BAM complex, from N. gonorrhoeae and H. ducreyi indicated that the particularly small hydrophobic thickness (9 Å) near the putative lateral gate formed between β-strands 1 and 16 of the barrel domain may induce the local thinning and disorder of the lipid bilayer (Fig. 1.8) (Noinaj et al. 2013). Thus, the lipid perturbation of the outer membrane by BamA was suggested to facilitate the insertion of OMPs by reducing the elastic stiffness of the bilayer (Gessmann et al. 2014). The conformational changes involving the opening of the contacts between the barrel and the periplasmic POTRA5 domain, and the deeper insertion of the conserved loop 6 into the barrel would also contribute to the opening of the lateral gate and the insertion of nascent polypeptide chains into the outer membrane (Noinaj et al. 2014).
Crystal structures of BamA homologues. Protein structures from and H. ducreyi (Hd) (PDB code: 4K3C, left) and N. gonorrhoeae (Ng) (PDB code: 4K3B, right). The structural features that are predicted to contribute to the insertion of outer-membrane proteins are indicated. The region near the lateral gate formed between strands β1 and β16 in the C-terminal barrel domain has reduced hydrophobic thickness (9 Å), which may perturb the outer membrane. The inter-strand hydrogen bonds are significantly perturbed in the NgBamA (right). The local thinning of the bilayer, the opening of the lateral gate, the separation of the POTRA domain 5 (P5) from the barrel, and the deeper insertion of loop6 (orange arrows) may collectively induce the direct interaction of nascent polypeptide chains, with the barrel lumen facilitating the insertion of outer membrane proteins (Figure adapted with permission from Noinaj et al. 2013)
1.5.3 Role of Physical Properties in the Thermodynamic Stability of Membrane Proteins
Thermodynamic study of membrane protein stability, which requires reversibility of the folding-unfolding transition, is more difficult than the kinetic study. Achieving reversibility involves tedious screenings of folding conditions such as temperature, detergents, lipids, denaturants, pH and ionic strength. Despite this challenge, the thermodynamic stability of several β-barrel membrane proteins including OmpA, PagP, OmpLA and OmpW has been studied in lipid bilayers (Hong and Tamm 2004; Huysmans et al. 2010; Moon and Fleming 2011; Moon et al. 2013). The lipid dependence of the OmpA stability in small unilamellar vesicles (SUVs) suggests prominent role of non-specific physical bilayer forces in the membrane protein stability (Hong and Tamm 2004) (Fig. 1.9). While PEs slow down the folding kinetics of OmpA, they increase the stability of the folded protein (ΔGo unfold). As the molar fraction of C16:0C18:1c9PE increased from 0 to 40 % (the remaining lipid was C16:0C18:1c9PC), ΔGo unfold increased from 3.4 to 5.0 kcal/mol. This stability enhancement could be due to the increase in the lateral pressure caused by the negative monolayer curvature, and the favorable contacts of HII-forming PE around the hour-grass shape of the OmpA TM region. Interestingly, the effect of PE can be mimicked by high content of PCs with long unsaturated acyl chains (diC16:1c9PC to diC20:1c9PC), which also induce the negative monolayer curvature (Szule et al. 2002). Thus, the protein stability enhancement by PEs originates from the non-specific physical properties of the bilayer rather than from the specific head group interactions.
Elastic coupling of membrane protein stability to lipid bilayer forces. Cartoon depicting structures and bilayer forces acting on OmpA folding/unfolding under equilibrium conditions. Folding into most bilayers is a two-state process (left path). Large black arrows indicate lateral bilayer pressure imparted on the lipid-protein interface in the hydrophobic core (red) of bilayers composed of lipids with negative intrinsic spontaneous curvature. Increasing this pressure increases the thermodynamic stability of the protein. Small black arrows indicate lipid deformation forces caused by hydrophobic mismatch between the protein and the unstressed bilayer. These forces decrease the thermodynamic stability of the protein. Folding into thin bilayers is a multistep process (right path) with at least one equilibrium intermediate. Water molecules penetrate more easily into the hydrophobic core (blue arrows) of more flexible and more dynamic thin bilayers, stabilizing the equilibrium intermediates and decreasing the m-value of unfolding. (m-value is a linear slope in the dependence of protein thermodynamic stability on denaturant concentration; lower m-values indicate smaller change in the solvent-accessible hydrophobic surface area upon unfolding and lower unfolding cooperativity.) Ultimately, in very thin bilayers composed of saturated lipids, complete unfolding (second step) can no longer be observed under any experimental conditions tested (Hong and Tamm 2004) (Figure adapted with permission from Hong and Tamm 2004)
Lipid-dependence of the thermodynamic stability of polytopic TM α-helical membrane proteins has not been studied yet in a lipid bilayer. However, the influence of lipid composition on the stability of TM helix-helix interactions has been studied for the prototype TM helical dimer model system, glycophorin A TM (GpATM) (Hong and Bowie 2011; Hong et al. 2010). The dimer stability of GpATM had been studied mostly in detergent micelles using analytical ultracentrifugation (AUC) (Fleming et al. 1997; Fleming and Engelman 2001) and Förster resonance energy transfer (FRET) (Fisher et al. 1999, 2003; Anbazhagan and Schneider 2010). In these methods, the relative populations of the monomeric and oligomeric states are modulated by diluting the protein with detergents or lipids. However, GpATM dimer is too strong in a bilayer (Adair and Engelman 1994) to be analyzed using conventional dilution methods for measuring the dimer dissociation constant, K d, dimer. To this end, a new technique called the steric trapping was developed to measure strong protein-protein interactions in lipid bilayers (Hong et al. 2013). The method couples dissociation of biotin-tagged TM dimer to tag-binding monovalent streptavidin. By exploiting the large free energy released from biotin- streptavidin interaction (ΔGo binding ~−19 kcal/mol), the measurable low limit of K d, dimer can be extended to 10−13–10−12M, which is three to four orders lower than that measurable by AUC and FRET.
The method revealed striking features of the lipid effects on the TM helix interactions. First, the dimer was much more stable (by ~5 kcal/mol) in a neutral fluid bilayer of C16:0C18:1c9PC than in detergent micelles (Hong et al. 2010). This is largely due to the reduced translational and rotational entropic cost when the dimer equilibrium is confined in liposomes. In addition, the packing contribution of several side chains to the dimer stability increased in the lipid bilayer, i. e. the dimer became more tightly organized. Second, the dimer stability was strongly modulated by the lipid composition and the protein environment (Hong and Bowie 2011). As seen in the case of OmpA, PE lipid, which is a major component of bacterial inner membranes, moderately stabilized the GpATM dimer by increasing the lateral pressure. Strikingly, the dimer was dramatically destabilized by 4–5 kcal/mol in PG, a natural negatively charged lipid in bacterial cell membranes. This destabilization originated from the electrostatic interactions between the cytoplasmic positively charged residues of GpATM and the negatively charged membrane, which distort the optimal dimer structure. Another surprising result was further destabilization of the dimer by the total extracts of E. coli inner membrane proteins. It has been predicted that the excluded volume effects in the crowded cellular environment enhance the protein-protein interaction and protein stability (Minton 2000). Contrary to this prediction, anonymous membrane proteins in the bilayer nonspecifically competed with the specific TM helix-helix interactions, destabilizing the GpATM dimer (Hong and Bowie 2011).
1.6 Membrane Protein Misfolding and Diseases
1.6.1 Lipid-Induced Folding, Misfolding and Topogenesis of α-Helical Membrane Proteins
The self-sealed nature of cell membranes and the directionality of polypeptide chain yield a specific pattern of “in” and “out” orientations of TM helices, termed protein “topology” (von Heijne 2006). Topogenesis of polytopic α-helical membrane proteins initially occurs during the co-translational insertion of TM helices through the interaction of the topogenic signals in the polypeptide with the translocon and the membrane, and is completed in the final folding and assembly process (Dowhan and Bogdanov 2009). Thus, the initial topology of individual TM helices is expected to critically affect the mechanism of the subsequent folding and assembly (Fig. 1.1). By this process, ion channels and transporters achieve their correct membrane orientations, and the catalytic domains and the active sites of membrane-bound enzymes are localized in a proper subcellular environment.
The strongest topogenic signal known so far is the “positive-inside” rule, which represents the biased distribution of positively charged amino acids, Arg and Lys, that are roughly five times more abundant in the cytosolic side than in the opposite side of the membrane (Krogh et al. 2001; von Heijne 1992) (see Figs. 1.1 and 1.10a). The N-terminal signal sequence or the first hydrophobic segment of membrane proteins is targeted to the inner membrane of bacteria or to the endoplasmic reticulum (ER) membrane in eukaryotes, and inserted into translocon as a hairpin during translation (Rapoport 2007). The strong electrostatic interaction of the positively charged residues along the hairpin with the negatively charged membrane and with the translocon stabilizes the orientation of the inserted TM helices (Dowhan and Bogdanov 2009). Reversal of the topology may be kinetically inhibited because it involves the translocation of hydrophilic loops and water-soluble domains across the non-polar bilayer core. However, as seen from the homo-dimeric bacterial multidrug transporter EmrE, the weak bias of the charge distribution can yield a dual topology of subunits and the formation of an antiparallel native protein assembly (Morrison et al. 2012). The topology analysis of E. coli inner membranes proteins revealed that, although not frequent, a significant fraction of TM segments does not follow the positive-inside rule (Gray et al. 2011). These findings imply that the topology of membrane proteins may be more dynamic than expected, and the reverse charge bias can be counterbalanced by more favorable tertiary or quaternary protein-protein interactions.
Influence of lipid composition on the topology and folding of polytopic α-helical membrane proteins. (a) Topology and distribution of positive charges in the N- (NT) and C- termini (CT), and the cytoplasmic (C2–C10) and periplasmic loops (P1–P11) of LacY expressed in wild type E. coli inner membranes. Positive-inside rule is well presented in the native topology of LacY. Locations of positively (red) and negatively (green) charged residues are indicated. (b) Topology of LacY expressed in PE− cells. The topology of TMI-TMVI helices was completely reversed, and TMVII in wild type E. coli turned into a cytoplasmic peripheral helix. Negatively and positively charged residues forming salt bridges between TMs are indicated. (c) Change in topology of LacY assembled in PE− cells after post-assembly synthesis of PE (Figure modified with permission from Bogdanov et al. 2014)
Critical roles of lipids in the topogenesis, assembly and function of polytopic membrane proteins have been elucidated through two methodological breakthroughs using the lactose permease (LacY) from E. coli as a model system. LacY is a galactoside/H+ symporter, which actively translocates lactose into the cytoplasm by exploiting the free energy stored in the electrochemical gradient across the membrane (the active uphill energy-dependent transport). LacY also facilitates the downhill energy-independent transport of lactose (Guan and Kaback 2006). First, Dowhan group developed E. coli mutant strains whose lipid compositions can be modified by knocking out and introducing genes involved in the lipid synthesis. Synthesis of PE, a major zwitterionic lipid in the E. coli cytoplasmic membrane, can be blocked by disrupting the chromosomal phosphatidylserine synthase gene (pssA), which produces PS, a precursor of PE (DeChavigny et al. 1991). Synthesis of PE can be restored by introducing the plasmid-encoded inducible pssA. In this system (PE− strain), negatively charged lipids PG and CL substitute PE. PE can also be replaced by another zwitterionic lipid, PC, by introducing the plasmid-encoded inducible phosphatidylcholine synthetase gene (pcsA) in the background of the PE− strain (Bogdanov et al. 2010). Second, Kaback group developed a conformation-specific monoclonal antibody (Ab4B1) that recognized the correct folding of a periplasmic P7 loop connecting helices TMVII and TMVIII (Sun et al. 1996). The binding ability of the antibody was well correlated with the correct folding and function of LacY (Fig. 1.10a).
Using those tools, Dowhan team revealed the replacement of PE lipids by PG and CL in the mutant E. coli strain led to the loss of binding of Ab4B1, that is, the misfolding of LacY. Strikingly, the topologies of TMI to TMVI were completely reversed relative to those in the wild type strain (Fig. 1.10b). LacY expressed in the membrane without PE still facilitated the downhill energy-independent transport of lactose, but could not support the uphill energy-dependent transport. The expression of PssA, which restored the PE synthesis, reversibly induced the formation of the correctly folded P7 loop and functional LacY (Bogdanov et al. 2008) (Fig. 1.10c). However, the locations of the N-terminus and P1 loop remained unchanged even after the restoration of PE synthesis, while the native-like topologies were regained by TMIII – TMVII, which apparently induced a U-shaped flexible hinge conformation of TMII within the membrane. This result indicates the dynamic nature of membrane topology and the critical role of specific lipids in the folding of polytopic α-helical membrane proteins. Replacement of PE by PC did not cause either topology reversal or loss of the uphill transport function, but the degree of the conformation-specific antibody binding was significantly reduced, further suggesting the fine-tuning role of lipids in the folding and conformational equilibrium of membrane proteins (Bogdanov et al. 2010).
Lipid-dependent control of topogenesis has been also observed for phenylalanine permease (PheP) (Zhang et al. 2003) and γ-aminobutyrate permease (GabP) (Zhang et al. 2005) from E. coli. However, the ability of lipids to support the correct topology and folding seems to depend more on the nonspecific role of lipids controlling the overall chemical and physical properties of cell membranes rather than on specific interactions between lipid head groups and certain protein residues. These intriguing findings led to the coining of the term “lipochaperone”, a specific lipid facilitating membrane protein folding, by analogy with protein molecular chaperones that assist the folding of water-soluble proteins and prevent their misfolding (Bogdanov and Dowhan 1999).
1.6.2 Membrane Protein Misfolding, Quality Control and Human Diseases
Misfolding and aggregation of membrane proteins in cells are directly linked to a number of human diseases (MacGurn et al. 2012; Houck and Cyr 2012; Reinstein and Ciechanover 2006; Sanders and Myers 2004; Kuznetsov and Nigam 1998). The abnormal folding of membrane proteins seems to be a common event that cells consistently deal with rather than a rare accident. For example, under normal physiological conditions, only <50 % of wild type cystic fibrosis transmembrane regulator (CFTR) and <20 % of wild type peripheral myelin protein 22 (PMP22) that are newly synthesized on the ER membrane reach the cytoplasmic membrane in their correctly folded form (Pareek et al. 1997; Kopito 1999). A wide array of the quality control mechanisms such as chaperones and degradation machinery constantly operates for monitoring the folding, degradation and trafficking of membrane proteomes in all subcellular compartments (MacGurn et al. 2012; Houck and Cyr 2012; Tatsuta and Langer 2008).
There are three major pathways for degrading membrane proteins in eukaryotic cells (Fig. 1.11). In the ER, where most membrane and secreted proteins are synthesized, misfolded membrane proteins are recognized and degraded by the ER-assisted degradation (ERAD) pathways (Hampton 2002). In mammalian cells, three types of the ERAD machines exist in the form of protein complexes (Hrd1/Derlin1,2,3/Sel1L/Ube2G1/Ube2k; DNAJB12/Derlin1/Hsp70/Rma1/Ube2J1; and TEB4/Ube2G/Ube2J) (Kikkert et al. 2004; Grove et al. 2011; Ravid et al. 2006). They are commonly equipped with the membrane-integrated ubiquitin ligase (E3) (Hrd1, Rma1 or TEB4) for the recognition and ubiquitination of misfolded membrane proteins, and the ubiquitin conjugation enzyme (E2) (Ube2G, Ube2K, and Ube2J1). Derlins, DNAJB12 (Hsp40), Sel1L and Hsp70 participate in the recognition process by chaperoning misfolded proteins. Ubiquitinated misfolded proteins are subsequently extracted from the ER membrane by the AAA+ATPase p97 in the cytosol, and are finally degraded in the 26S proteasome. Misfolded proteins in the Golgi and cytoplasmic membranes are recognized by other types of ubiquitination enzymes (Rsp5 in Golgi and Nedd4, Cbl, and CHIP in cytoplasmic membrane), endocytosed into endosomes, and degraded in lysosomes (MacGurn et al. 2012). Aggregated membrane proteins and damaged subcellular organelles, whose sizes are too large to be degraded by these mechanisms, are engulfed by double-membrane autophagosomes and targeted for lysosomal degradation (He and Klionsky 2009). At most, ~80 % of the membrane protein turnover is carried out by the ERAD pathways (Reinstein and Ciechanover 2006).
Quality control mechanisms of integral membrane proteins in eukaryotic cells. Correctly folded membrane proteins are ultimately targeted to their final destination (plasma membrane here) through the vesicular transport (cyan arrows). (i) ERAD pathway (purple arrow): the misfolded proteins in the ER membrane are recognized and ubiquitinated by the membrane-bound ubiquitin ligase complex (Hrd1 complex here). The proteins are extracted from the membrane by p97 AAA+ATPase and ultimately degraded in the proteasome (purple arrow). (ii) Lysosomal degradation pathway (red arrows): misfolded proteins in the Golgi and plasma membranes are tagged by ubiquitin and targeted to lysosome. (iii) Autophagy degradation pathway (brown arrow): damaged subcellular compartments and protein aggregates are engulfed by autophagosome and targeted to lysosome
The pathogenesis of membrane protein misfolding diseases is partly the consequence of the abnormal interaction between misfolded proteins and the degradation machinery (Sanders and Myers 2004): (1) Proteins with loss-of-function mutations are targeted to their final destinations bypassing the quality control mechanisms. (2) Destabilizing or misfolding mutations enhance proteins’ susceptibility to degradation and thus reduce the level of functional proteins. (3) Malfunction of degradation machinery by mutations or stresses causes a failure of clearing toxic misfolded and aggregated proteins. More than 60,000 missense mutations in human genes leading to disease phenotypes have been identified, yet mutations in residues that are directly involved in the protein function are rare (Stenson et al. 2008) (http://www.biobase-international.com/product/hgmd). Rather, a more common disease mechanism seems to involve destabilizing mutations that impair the correct trafficking of the functional proteins by enhancing degradation.
Cystic fibrosis transmembrane conductance regulator (CFTR), which regulates the anion balance across the plasma membranes of epithelial cells (Miller 2010), is an intensely studied model system for membrane protein misfolding diseases. CFTR is composed of 12 TM segments, two nucleotide-binding domains (NBDs) and a regulatory domain. Malfunction of CFTR is a major cause of cystic fibrosis, a severe respiratory and gastrointestinal disorder by abnormal production of viscous mucus (Sheppard and Welsh 1999). Among 1,984 mutations deposited in the cystic fibrosis mutation database (http://www.genet.sickkids.on.ca), 69 % of patients carry the ΔF508 deletion mutation. F508 is located in the NBD on the cytoplasmic side of CFTR, and ΔF508 mutation induces the misfolding of the NBD domain (Lukacs and Verkman 2012). Although ΔF508 mutation partially retains the anion transport function, the mutant is completely degraded by the ERAD pathway and fails to correctly target to the cell surface (Pasyk and Foskett 1995). A correct targeting is also completely aborted in the ERAD pathway (Sun et al. 2006) for CFTR carrying the mutations (G85E, R347P, and R334W) in the transmembrane domain.
The enhanced susceptibility to degradation upon missense mutations is an established mechanism of other diseases. Those include retinitis pigmentosa, a common retinal degeneration disease partly caused by missense mutations in rhodopsin, and Charcot-Marie-Tooth disease, a common inherited neurological disorder, which is related to missense mutations in PMP22, a tetra-membrane spanning integral membrane proteins (Rajan and Kopito 2004; Pareek et al. 1997; Sakakura et al. 2011).
Another critical disease mechanism involves mutations that do not directly affect the function or the conformational stability of membrane proteins, but impair their interactions with the degradation machinery. One example is the Liddle syndrome, a condition of severe hypertension, hypokalemia and metabolic alkalosis. The gene responsible for the Liddle syndrome encodes the β-subunit of renal epithelial sodium channel. A disease mutation occurs at the binding interface with the E3 ubiquitin ligase Nedd4 (Shimkets 1996; Staub et al. 1997). Thus, Nedd4 can neither recognize the sodium channel nor target it for degradation. The accumulation of the channel induces the excessive reabsorption of sodium ions and water, eventually causing hypertension. Impairment of ubiquitin-mediated degradation by mutations on E3 ligases is also implicated in Angelman syndrome (E6-AP E3), Parkinson’s disease (Parkin E3), von Hippel-Lindau protein-associated syndromes (von Hippel-Lindau E3), and other diseases (Reinstein and Ciechanover 2006).
Molecular features of misfolded membrane proteins that are selectively recognized by the quality control machines remain unclear. In water-soluble proteins, the integrity of the hydrophobic core is the key determinant for protein interactions with chaperones and degradation machinery. Since the stability of membrane proteins is governed by substantially different thermodynamic principles, the interactions between misfolded membrane proteins and the quality control machines should also be different from those of water-soluble proteins. One possible scenario is that polar interactions of membrane proteins in a low-dielectric lipid bilayer may play a significant role in the folding and stability. Misfolding induces the exposure of polar residues into the bilayer, facilitating their interactions with chaperones and membrane-bound ubiquitin ligases (Houck and Cyr 2012). Indeed, Sato et al. (2009) showed that mutations of the surface polar residues in the TM domain of Hrd1p ubiquitin ligase, a key component of the ERAD machinery, significantly reduced the degradation susceptibility of 3-hydroxy-3-methylglutaryl-coenzyme A reductase.
Still, it is unclear whether the polar interactions are a dominant factor determining the thermodynamic stability of integral membrane proteins. Numerous studies measuring the hydrogen bonding strength in membrane proteins consistently report moderate contributions (0.5–1.5 kcal/mol per bond), not much different from those in water-soluble proteins (Bowie 2011). Furthermore, the analysis of disease mutations mapped on the structures of rhodopsin, G-protein coupled receptors, and potassium channels did not find a dominant feature of polar-to-nonpolar or nonpolar-to-polar mutations (Oberai et al. 2009). Further biophysical studies on membrane protein folding are necessary to explain disease mechanisms caused by misfolding.
1.6.3 Role of Cholesterol in GPCR Receptor
Lipidomic studies demonstrated that changes in the lipid composition of cell membranes can provide markers of disease; however, a direct link between the lipid-mediated misfolding of membrane proteins and specific diseases is unclear (Hu et al. 2009). Several examples show the pivotal role of lipids in the structure and function of pharmaceutically important membrane proteins. These studies may ultimately help develop the pharmacological chaperones to modulate folding and stability for the optimal function (Mendre and Mouillac 2010).
Although cholesterol has long been known as a critical modulator of membrane fluidity, curvature stress and raft formation, it also acts as a specific “ligand” stabilizing the structure of β2-ardrenergic receptor (β2-AR), a member of the G-protein coupled receptor (GPCR) family (Hanson et al. 2008). β2-AR forms a complex with its effector, l-type Ca2+ channel, and senses epinephrine and norepinephrine in muscular tissues, such as bronchial vasculature and blood vessels. The receptor is an important drug target for the treatment of asthma, glaucoma, high blood pressure and cardiac arrhythmias.
The crystal structures of β2-AR were solved for apo- and carazolol (inverse agonist)-bound forms (Cherezov et al. 2007; Rasmussen et al. 2007). Later, the structure was solved in the timolol- (inverse agonist) and cholesterol-bound form (Hanson et al. 2008). This was the first structure in which a cholesterol binding motif was elucidated in atomic detail (Fig. 1.12). Two cholesterol molecules bind to the cleft formed by the helices I, II, III and IV. Two prominent residues that bind cholesterol through CH-π and van der Waals interactions are Trp1584.50 conserved in 94 % of GPCR (Ballesteros and Weinstein 1995) and Ile1544.46 conserved in 60 % of GPCR by homology and 35 % by identity (the subscripts specify an GPCR amino acid according to Ballesteros-Weinstein numbering system allowing comparison of equivalent positions in the GPCR family). Two other bulky residues, Tyr702.41 and Arg1514.43, which are less conserved, cap cholesterol in the cytoplasmic interfacial region by van der Waals and hydrogen bonding interactions. The four-residue cholesterol binding consensus motif, [4.39–4.43(R,K)]—[4.50(W,Y)]—[4.46(I,V,L)]—[2.41(F,Y)], is widely distributed in the GPCR family, suggesting that cholesterol binding may be a crucial feature of many GPCRs. Indeed, the subsequently solved GPCR structures of A2A adenosine receptor (Jaakola et al. 2008), μ-type opioid receptor (Manglik et al. 2012) and hydroxytryptoamine (serotonin) receptor 2B (Wang et al. 2013) revealed cholesterol bound to this motif, although the precise modes of interactions were slightly different. The enhancement of protein thermostability has been observed in the micellar phase for other GPCRs such as oxytocin receptor, which is a serotonin1A receptor containing a strict cholesterol consensus motif (Hanson et al. 2008). The interaction between GPCRs and cholesterol were postulated to lead to their preferential partition into the cholesterol-rich caveolae (Pucadyil and Chattopadhyay 2004; Song et al. 2014a).
Binding of cholesterol modulates stability and function of GPCR receptors. Binding of two cholesterol molecules (CHOL1 and CHOL2) to β2-aderenergic receptor (β2-AR) (Hanson et al. 2008). Binding site of CHOL1 represents a cholesterol consensus motif. Trp158 and Ile154, which are 95 and 35 % conserved by identity, are central in cholesterol binding (Figure modified with permission from Hanson et al. 2008)
1.6.4 Role of Cholesterol in Amyloid Precursor Proteins
While cholesterol binds to the “cleft” formed by TM helices of β2-AR and is locked by several bulky residues, another type of cholesterol binding site has been identified in the amyloid precursor protein (APP) (Barrett et al. 2012). APP is cleaved by an intramembrane protease, γ-secretase, which generates the C-terminal C99 TM domain and the amyloid-β (Aβ) peptide (Song et al. 2014a). Misfolding and accumulation of aggregated Aβ in the brain is the hallmark of Alzheimer’s disease (Finder and Glockshuber 2007). The solution NMR structure of APP showed that the extracellular region of C99 is composed of an amphiphilic N-helix and a short N-loop connected to the TM domain (Barrett et al. 2012) (Fig. 1.13). The TM domain is highly bent and flexible, which may promote the cleavage of C99 by γ-secretase. The N-terminal half of the TM domain contains the GxxxGxxxG glycine zipper motif, which is frequently observed in the TM helix-helix interactions ((Kim et al. 2005); also see Chap. 4 by Morgado and Garvey and Chap. 10 by Leonova and Galzitskaya in this volume). Because of this motif, the C99 protein was thought to form dimers or higher-order oligomers. However, titration of cholesterol (up to 20 mol %) in bicelles, monitored by NMR, revealed a striking binding pattern of cholesterol to C99. Contrary to the expectation, two of the three Gly in the zipper motif, G700 and G704, are critically involved in cholesterol binding rather than oligomerization. In addition, the dynamic N-loop and the polar residues N698 and E693 also appear to contribute to the binding of cholesterol. Furthermore, the dynamic N-helix, which lies parallel to the membrane and perpendicular to the TM helix to which it is linked via the N-loop, effectively locks bound cholesterol by packing with the bulky aromatic residue, F690. Thus, the flat surface presented by the GxxxG motif, the flexible N-loop and the conformation of N-helix work together to accommodate bound cholesterol.
Cholesterol binding to the C99 TM domain of amyloid precursor protein. Cholesterol binds to the flat surface created by glycine zipper motif (G700-xxx-G704-xxx-G708) and, additionally, by N-helix (F691) and N-loop (E693 and N698) (Figure adopted with permission from Song et al. 2014a)
What is the physiological significance of cholesterol binding in the generation of Aβ? γ-secretases are known to preferentially partition into the cholesterol-rich membrane rafts (Lee et al. 1998). Cholesterol binding to APP is expected to favor partitioning into membrane rafts, and thereby co-localize APP with the γ-secretase. Cholesterol binding was also proposed to make APP a better substrate for γ-secretase (Osenkowski et al. 2008). Moreover, after cleavage, Aβ association with cholesterol may facilitate amyloid fibril formation (Yanagisawa 2005). These effects provide possible explanations for the link between the plasma levels of cholesterol and the development of Alzheimer’s disease.
1.6.5 Role of PIP2 in the Gating Function of Inward Rectifier Potassium Channel
Phosphatidylinositol 4,5 diphosphate (PIP2) is a minor lipid component in the plasma membrane that is critical in a number of signaling pathways as a secondary messenger and a targeting signal of soluble proteins on the plasma membrane (Tisi et al. 2004; Cho and Stahelin 2005). PIP2 is also of particular interest as a direct modulator of several ion channels (Suh and Hille 2008; Suh et al. 2006, 2010). The crystal structures of inward rectifier potassium channel (Kir2.2) in apo- and in PIP2 – bound forms solved by (Hansen et al. 2011) demonstrated the specific role of PIP2 in the regulation of resting cell membrane potential (Fig. 1.14). In the apo form, the TM domain and the cytoplasmic domain are disengaged. Binding of PIP2 at the interface between the two domains induces a substantial conformational change. The PIP2 head group binds to the conserved highly positively charged PIP2 – binding motif located in the flexible interdomain linker (KxxRPKK motif that binds inositol 4,5 diphosphate) and to the interfacial region of the TM domain (RWR or KWR motif that binds phosphoester). Binding of the acyl chain of PIP2 to the TM domain occurs in a non-specific manner. This PIP2 binding, which induces a coil-to-helix transition of the interdomain linker, pulls away the inner helical gating region of the TM domain in the direction of the membrane plane, docks the cytoplasmic G-loop into the channel lumen, and opens up the channel.
Kir2.2 inward rectifier K+ channel gated by PIP2 binding. Structures of Kir2.2 in apo- and PIP2-bound forms (top). PIP2 binding induces a conformational change that tightens the junction between the TM domain (TMD) and cytoplasmic domain (CYT) by 6 Å and opens the channel via a proposed mechanism shown in the bottom. PIP2 is shown in purple circles (Figures adapted with permission from Hansen et al. 2011)
1.7 Concluding Remarks
This chapter describes the active role of lipids in the folding, structure and function of integral membrane proteins. The results suggest that lipids in cell membranes not only serve as a medium for stabilizing membrane proteins but also act as critical functional and structural ligands for many of these proteins. Therefore, at the origin of life, the quasi two-dimensional film that constitutes cell membranes may have started as a simple permeability barrier, but evolved to fit to the biological needs in cellular processes by incorporating lipids with various chemical and physical properties and by utilizing them as ligands and stabilizers for membrane proteins.
Structural studies provided the molecular details of lipid-protein interactions for several membrane proteins. Still, it remains a far-reaching goal to quantify the energetics and dynamics of these interactions and elucidate their relationship to the conformational equilibria of membrane proteins which enable their function. In this context, quantitative studies on the energetics of the mechanosensitive channel gating (Perozo et al. 2002a, b; Wiggins and Phillips 2004) and NMR studies of the amyloid precursor protein revealing the dynamic features of this disease-causing protein, are truly intriguing (Barrett et al. 2012; Beel et al. 2010; Pester et al. 2013; Song et al. 2013, 2014b). For such efforts to be extended to other classes of membrane proteins, new experimental and theoretical frameworks will be necessary. A recent development of steric trapping method that allows the reversible control of the membrane protein folding will open new opportunities for studying the forces and mechanisms of folding of membrane proteins in their native bilayer background (Blois et al. 2009; Chang and Bowie 2014; Hong et al. 2010; Hong and Bowie 2011; Jefferson et al. 2013).
Abbreviations
- APP:
-
Amyloid precursor protein
- bR:
-
Bacteriorhodopsin
- CCS:
-
Collision-cross section
- CFTR:
-
Cystic fibrosis transmembrane conductance regulator
- CL:
-
Cardiolipin
- ER:
-
Endoplasmic reticulum
- GpATM:
-
Glycophorin A transmembrane domain
- GPCR:
-
G-protein coupled receptor
- HI :
-
Hexagonal phase
- HII :
-
Inverted hexagonal phase
- Lα :
-
Lamellar phase
- Lo :
-
Liquid ordered phase
- PA:
-
Phosphatidic acid
- PC:
-
Phosphatidylcholine
- PE:
-
Phosphatidylethanolamine
- PG:
-
Phosphatidylglycerol
- PI:
-
Phosphatidylinositol
- SM:
-
Sphingomyelin
- SRP:
-
Signal recognition particle
- TM:
-
Transmembrane
References
Adair BD, Engelman DM (1994) Glycophorin A helical transmembrane domains dimerize in phospholipid bilayers: a resonance energy transfer study. Biochemistry 33(18):5539–5544
Adamian L, Liang J (2001) Helix-helix packing and interfacial pairwise interactions of residues in membrane proteins. J Mol Biol 311(4):891–907
Adamian L, Nanda V, Degrado WF, Liang J (2005) Empirical lipid propensities of amino acid residues in multispan alpha helical membrane proteins. Proteins 59(3):496–509
Anbazhagan V, Schneider D (2010) The membrane environment modulates self-association of the human GpA TM domain - implications for membrane protein folding and transmembrane signaling. Biochim Biophys Acta 1798(10):1899–1907
Ballesteros JA, Weinstein H (1995) Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors. Methods Neurosci 25(1):366–428
Barrera NP, Di Bartolo N, Booth PJ, Robinson CV (2008) Micelles protect membrane complexes from solution to vacuum. Science 321(5886):243–246
Barrett PJ, Song Y, Van Horn WD, Hustedt EJ, Schafer JM, Hadziselimovic A, Beel AJ, Sanders CR (2012) The amyloid precursor protein has a flexible transmembrane domain and binds cholesterol. Science 336(6085):1168–1171
Beel AJ, Sakakura M, Barrett PJ, Sanders CR (2010) Direct binding of cholesterol to the amyloid precursor protein: an important interaction in lipid-Alzheimer’s disease relationships? Biochim Biophys Acta 1801(8):975–982
Bental N, Sitkoff D, Topol IA, Yang AS, Burt SK, Honig B (1997) Free energy of amide hydrogen bond formation in vacuum, in water, and in liquid alkane solution. J Phys Chem B 101(3):450–457
Bertero MG, Rothery RA, Palak M, Hou C, Lim D, Blasco F, Weiner JH, Strynadka NC (2003) Insights into the respiratory electron transfer pathway from the structure of nitrate reductase A. Nat Struct Biol 10(9):681–687
Blois TM, Hong H, Kim TH, Bowie JU (2009) Protein unfolding with a steric trap. J Am Chem Soc 131(39):13914–13915
Bogdanov M, Dowhan W (1999) Lipid-assisted protein folding. J Biol Chem 274(52):36827–36830
Bogdanov M, Xie J, Heacock P, Dowhan W (2008) To flip or not to flip: lipid-protein charge interactions are a determinant of final membrane protein topology. J Cell Biol 182(5):925–935
Bogdanov M, Heacock P, Guan Z, Dowhan W (2010) Plasticity of lipid-protein interactions in the function and topogenesis of the membrane protein lactose permease from Escherichia coli. Proc Natl Acad Sci U S A 107(34):15057–15062
Bogdanov M, Dowhan W, Vitrac H (2014) Lipids and topological rules governing membrane protein assembly. Biochim Biophys Acta Mol Cell Res 1843(8):1475–1488
Booth PJ, Riley ML, Flitsch SL, Templer RH, Farooq A, Curran AR, Chadborn N, Wright P (1997) Evidence that bilayer bending rigidity affects membrane protein folding. Biochemistry 36(1):197–203
Bowie JU (1997) Helix packing in membrane proteins. J Mol Biol 272(5):780–789
Bowie JU (2011) Membrane protein folding: how important are hydrogen bonds? Curr Opin Struct Biol 21(1):42–49
Burgess NK, Dao TP, Stanley AM, Fleming KG (2008) Beta-barrel proteins that reside in the Escherichia coli outer membrane in vivo demonstrate varied folding behavior in vitro. J Biol Chem 283(39):26748–26758
Burley SK, Petsko GA (1986) Amino-aromatic interactions in proteins. FEBS Lett 203(2):139–143
Bush MF, Hall Z, Giles K, Hoyes J, Robinson CV, Ruotolo BT (2010) Collision cross sections of proteins and their complexes: a calibration framework and database for gas-phase structural biology. Anal Chem 82(22):9557–9565
Cafiso DS (1994) Alamethicin: a peptide model for voltage gating and protein-membrane interactions. Annu Rev Biophys Biomol Struct 23:141–165
Cantor RS (1997) The lateral pressure profile in membranes: a physical mechanism of general anesthesia. Biochemistry 36(9):2339–2344
Cantor RS (1999) Lipid composition and the lateral pressure profile in bilayers. Biophys J 76(5):2625–2639
Chan RB, Oliveira TG, Cortes EP, Honig LS, Duff KE, Small SA, Wenk MR, Shui G, Di Paolo G (2012) Comparative lipidomic analysis of mouse and human brain with Alzheimer disease. J Biol Chem 287(4):2678–2688
Chang YC, Bowie JU (2014) Measuring membrane protein stability under native conditions. Proc Natl Acad Sci U S A 111(1):219–224
Cherezov V, Rosenbaum DM, Hanson MA, Rasmussen SG, Thian FS, Kobilka TS, Choi HJ, Kuhn P, Weis WI, Kobilka BK, Stevens RC (2007) High-resolution crystal structure of an engineered human beta2-adrenergic G protein-coupled receptor. Science 318(5854):1258–1265
Cho W, Stahelin RV (2005) Membrane-protein interactions in cell signaling and membrane trafficking. Annu Rev Biophys Biomol Struct 34:119–151
Chong SH, Ham S (2014) Interaction with the surrounding water plays a key role in determining the aggregation propensity of proteins. Angew Chem 53(15):3961–3964
Clark MR (2011) Flippin’ lipids. Nat Immunol 12(5):373–375
Contreras FX, Ernst AM, Wieland F, Brugger B (2011) Specificity of intramembrane protein-lipid interactions. Cold Spring Harb Perspect Biol 3(6):1–18
Curran AR, Templer RH, Booth PJ (1999) Modulation of folding and assembly of the membrane protein bacteriorhodopsin by intermolecular forces within the lipid bilayer. Biochemistry 38(29):9328–9336
Dawidowicz EA (1987) Dynamics of membrane lipid metabolism and turnover. Annu Rev Biochem 56:43–61
Dechavigny A, Heacock PN, Dowhan W (1991) Sequence and inactivation of the pss gene of Escherichia coli. Phosphatidylethanolamine may not be essential for cell viability. J Biol Chem 266(8):5323–5332
Dill KA (1990) Dominant forces in protein folding. Biochemistry 29(31):7133–7155
Dowhan W, Bogdanov M (2009) Lipid-dependent membrane protein topogenesis. Annu Rev Biochem 78:515–540
Du Plessis DJ, Nouwen N, Driessen AJ (2011) The Sec translocase. Biochim Biophys Acta 1808(3):851–865
East JM, Melville D, Lee AG (1985) Exchange rates and numbers of annular lipids for the calcium and magnesium ion dependent adenosine triphosphatase. Biochemistry 24(11):2615–2623
Eilers M, Shekar SC, Shieh T, Smith SO, Fleming PJ (2000) Internal packing of helical membrane proteins. Proc Natl Acad Sci U S A 97(11):5796–5801
Engelman DM (2005) Membranes are more mosaic than fluid. Nature 438(7068):578–580
Engelman DM, Chen Y, Chin CN, Curran AR, Dixon AM, Dupuy AD, Lee AS, Lehnert U, Matthews EE, Reshetnyak YK, Senes A, Popot JL (2003) Membrane protein folding: beyond the two stage model. FEBS Lett 555(1):122–125
Escriba PV, Ozaita A, Ribas C, Miralles A, Fodor E, Farkas T, Garciasevilla JA (1997) Role of lipid polymorphism in G protein-membrane interactions: nonlamellar-prone phospholipids and peripheral protein binding to membranes. Proc Natl Acad Sci U S A 94(21):11375–11380
Finder VH, Glockshuber R (2007) Amyloid-beta aggregation. Neurodegener Dis 4(1):13–27
Fisher LE, Engelman DM, Sturgis JN (1999) Detergents modulate dimerization, but not helicity, of the glycophorin A transmembrane domain. J Mol Biol 293(3):639–651
Fisher LE, Engelman DM, Sturgis JN (2003) Effect of detergents on the association of the glycophorin a transmembrane helix. Biophys J 85(5):3097–3105
Fleming KG, Engelman DM (2001) Specificity in transmembrane helix-helix interactions can define a hierarchy of stability for sequence variants. Proc Natl Acad Sci U S A 98(25):14340–14344
Fleming KG, Ackerman AL, Engelman DM (1997) The effect of point mutations on the free energy of transmembrane alpha-helix dimerization. J Mol Biol 72(2):266–275
Fyfe PK, Jones MR (2005) Lipids in and around photosynthetic reaction centres. Biochem Soc Trans 33(Pt 5):924–930
Gallivan JP, Dougherty DA (1999) Cation-pi interactions in structural biology. Proc Natl Acad Sci U S A 96(17):9459–9464
Gessmann D, Chung YH, Danoff EJ, Plummer AM, Sandlin CW, Zaccai NR, Fleming KG (2014) Outer membrane beta-barrel protein folding is physically controlled by periplasmic lipid head groups and BamA. Proc Natl Acad Sci U S A 111(16):5878–5883
Gonen T, Cheng Y, Sliz P, Hiroaki Y, Fujiyoshi Y, Harrison SC, Walz T (2005) Lipid-protein interactions in double-layered two-dimensional AQP0 crystals. Nature 438(7068):633–638
Gray AN, Henderson-Frost JM, Boyd D, Sharafi S, Niki H, Goldberg MB (2011) Unbalanced charge distribution as a determinant for dependence of a subset of Escherichia coli membrane proteins on the membrane insertase YidC. MBio 2(6):1–10
Grigorieff N, Ceska TA, Downing KH, Baldwin JM, Henderson R (1996) Electron-crystallographic refinement of the structure of bacteriorhodopsin. J Mol Biol 259(3):393–421
Grove DE, Fan CY, Ren HY, Cyr DM (2011) The endoplasmic reticulum-associated Hsp40 DNAJB12 and Hsc70 cooperate to facilitate RMA1 E3-dependent degradation of nascent CFTR Delta F508. Mol Biol Cell 22(3):301–314
Gruner SM (1985) Intrinsic curvature hypothesis for biomembrane lipid-composition – a role for nonbilayer lipids. Proc Natl Acad Sci U S A 82(11):3665–3669
Gruner SM, Cullis PR, Hope MJ, Tilcock CP (1985) Lipid polymorphism: the molecular basis of nonbilayer phases. Annu Rev Biophys Biophys Chem 14:211–238
Gruner SM, Tate MW, Kirk GL, So PT, Turner DC, Keane DT, Tilcock CP, Cullis PR (1988) X-ray diffraction study of the polymorphic behavior of N-methylated dioleoyl phosphatidylethanolamine. Biochemistry 27(8):2853–2866
Guan L, Kaback HR (2006) Lessons from lactose permease. Annu Rev Biophys Biomol Struct 35:67–91
Guskov A, Kern J, Gabdulkhakov A, Broser M, Zouni A, Saenger W (2009) Cyanobacterial photosystem II at 2.9-A resolution and the role of quinones, lipids, channels and chloride. Nat Struct Mol Biol 16(3):334–342
Haltia T, Freire E (1995) Forces and factors that contribute to the structural stability of membrane-proteins. Biochim Biophys Acta Bioenerg 1228(1):1–27
Hampton RY (2002) ER-associated degradation in protein quality control and cellular regulation. Curr Opin Cell Biol 14(4):476–482
Hansen SB, Tao X, Mackinnon R (2011) Structural basis of PIP2 activation of the classical inward rectifier K+ channel Kir2.2. Nature 477(7365):495–498
Hanson MA, Cherezov V, Griffith MT, Roth CB, Jaakola VP, Chien EY, Velasquez J, Kuhn P, Stevens RC (2008) A specific cholesterol binding site is established by the 2.8 A structure of the human beta2-adrenergic receptor. Structure 16(6):897–905
He C, Klionsky DJ (2009) Regulation mechanisms and signaling pathways of autophagy. Annu Rev Genet 43:67–93
Hessa T, Kim H, Bihlmaier K, Lundin C, Boekel J, Andersson H, Nilsson I, White SH, Von Heijne G (2005) Recognition of transmembrane helices by the endoplasmic reticulum translocon. Nature 433(7024):377–381
Hessa T, Meindl-Beinker NM, Bernsel A, Kim H, Sato Y, Lerch-Bader M, Nilsson I, White SH, Von Heijne G (2007) Molecular code for transmembrane-helix recognition by the Sec61 translocon. Nature 450(7172):1026–1030
Hite RK, Li Z, Walz T (2010) Principles of membrane protein interactions with annular lipids deduced from aquaporin-0 2D crystals. EMBO J 29(10):1652–1628
Hong H, Bowie JU (2011) Dramatic destabilization of transmembrane helix interactions by features of natural membrane environments. J Am Chem Soc 133(29):11389–11398
Hong H, Tamm LK (2004) Elastic coupling of integral membrane protein stability to lipid bilayer forces. Proc Natl Acad Sci U S A 101(12):4065–4070
Hong H, Blois TM, Cao Z, Bowie JU (2010) Method to measure strong protein-protein interactions in lipid bilayers using a steric trap. Proc Natl Acad Sci U S A 107(46):19802–19807
Hong H, Chang YC, Bowie JU (2013) Measuring transmembrane helix interaction strengths in lipid bilayers using steric trapping. Methods Mol Biol 1063:37–56
Houck SA, Cyr DM (2012) Mechanisms for quality control of misfolded transmembrane proteins. Biochim Biophys Acta 1818(4):1108–1114
Hu CX, Van Der Heijden R, Wang M, Van Der Greef J, Hankemeier T, Xua GW (2009) Analytical strategies in lipidomics and applications in disease biomarker discovery. J Chromatogr B Anal Tech Biomed Life Sci 877(26):2836–2846
Huysmans GH, Baldwin SA, Brockwell DJ, Radford SE (2010) The transition state for folding of an outer membrane protein. Proc Natl Acad Sci U S A 107(9):4099–4104
Jaakola VP, Griffith MT, Hanson MA, Cherezov V, Chien EY, Lane JR, Ijzerman AP, Stevens RC (2008) The 2.6 angstrom crystal structure of a human A2A adenosine receptor bound to an antagonist. Science 322(5905):1211–1217
Jefferson RE, Blois TM, Bowie JU (2013) Membrane proteins can have high kinetic stability. J Am Chem Soc 135(40):15183–15190
Joh NH, Oberai A, Yang D, Whitelegge JP, Bowie JU (2009) Similar energetic contributions of packing in the core of membrane and water-soluble proteins. J Am Chem Soc 131(31):10846–10847
Keller SL, Bezrukov SM, Gruner SM, Tate MW, Vodyanoy I, Parsegian VA (1993) Probability of alamethicin conductance states varies with nonlamellar tendency of bilayer phospholipids. Biophys J 65(1):23–27
Kikkert M, Doolman R, Dai M, Avner R, Hassink G, Van Voorden S, Thanedar S, Roitelman J, Chau V, Wiertz E (2004) Human HRD1 is an E3 ubiquitin ligase involved in degradation of proteins from the endoplasmic reticulum. J Biol Chem 279(5):3525–3534
Kim S, Jeon TJ, Oberai A, Yang D, Schmidt JJ, Bowie JU (2005) Transmembrane glycine zippers: physiological and pathological roles in membrane proteins. Proc Natl Acad Sci U S A 102(40):14278–14283
Kleinschmidt JH, Tamm LK (1996) Folding intermediates of a beta-barrel membrane protein. Kinetic evidence for a multi-step membrane insertion mechanism. Biochemistry 35(40):12993–13000
Kleinschmidt JH, Tamm LK (2002) Secondary and tertiary structure formation of the beta-barrel membrane protein OmpA is synchronized and depends on membrane thickness. J Mol Biol 324(2):319–330
Knowles PF, Watts A, Marsh D (1979) Spin-label studies of lipid immobilization in dimyristoyl phosphatidylcholine-substituted cytochrome oxidase. Biochemistry 18(21):4480–4487
Kopito RR (1999) Biosynthesis and degradation of CFTR. Physiol Rev 79(1 Suppl):S167–S173
Krishnamachary N, Stephenson FA, Steggles AW, Holloway PW (1994) Stopped-flow fluorescence studies of the interaction of a mutant form of cytochrome b5 with lipid vesicles. J Fluoresc 4(3):227–233
Krogh A, Larsson B, Von Heijne G, Sonnhammer ELL (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305(3):567–580
Kumar VV (1991) Complementary molecular shapes and additivity of the packing parameter of lipids. Proc Natl Acad Sci U S A 88(2):444–448
Kusumi A, Ike H, Nakada C, Murase K, Fujiwara T (2005) Single-molecule tracking of membrane molecules: plasma membrane compartmentalization and dynamic assembly of raft-philic signaling molecules. Semin Immunol 17(1):3–21
Kuznetsov G, Nigam SK (1998) Folding of secretory and membrane proteins. N Engl J Med 339(23):1688–1695
Laganowsky A, Reading E, Hopper JT, Robinson CV (2013) Mass spectrometry of intact membrane protein complexes. Nat Protoc 8(4):639–651
Lange C, Nett JH, Trumpower BL, Hunte C (2001) Specific roles of protein-phospholipid interactions in the yeast cytochrome bc1 complex structure. EMBO J 20(23):6591–6600
Lee AG (2011) Biological membranes: the importance of molecular detail. Trends Biochem Sci 36(9):493–500
Lee SJ, Liyanage U, Bickel PE, Xia W, Lansbury PT Jr, Kosik KS (1998) A detergent-insoluble membrane compartment contains A beta in vivo. Nat Med 4(6):730–734
Lehninger AL, Nelson DL, Cox MM (1993) Principles of biochemistry. Worth Publishers, New York
Lodish H, Berk A, Kaiser CA, Krieger T, Scott M, Pretscher A, Ploegh H, Matsudaira P (2000) Biomembrane structure (chapter 10). In: Molecular cell biology. W. H. Freeman, New York
London E, Feigenson GW (1981) Fluorescence quenching in model membranes. 1. Characterization of quenching caused by a spin-labeled phospholipid. Biochemistry 20(7):1932–1938
Luecke H, Schobert B, Richter HT, Cartailler JP, Lanyi JK (1999) Structure of bacteriorhodopsin at 1.55 A resolution. J Mol Biol 291(4):899–911
Lukacs GL, Verkman AS (2012) CFTR: folding, misfolding and correcting the DeltaF508 conformational defect. Trends Mol Med 18(2):81–91
Macgurn JA, Hsu PC, Emr SD (2012) Ubiquitin and membrane protein turnover: from cradle to grave. Annu Rev Biochem 81:231–259
Manglik A, Kruse AC, Kobilka TS, Thian FS, Mathiesen JM, Sunahara RK, Pardo L, Weis WI, Kobilka BK, Granier S (2012) Crystal structure of the micro-opioid receptor bound to a morphinan antagonist. Nature 485(7398):321–326
Marsh D (2008) Protein modulation of lipids, and vice-versa, in membranes. Biochim Biophys Acta 1778(7–8):1545–1575
Mendre C, Mouillac B (2010) Pharmacological chaperones: a potential therapeutic treatment for conformational diseases. M S Med Sci 26(6–7):627–635
Miller C (2010) CFTR: break a pump, make a channel. Proc Natl Acad Sci U S A 107(3):959–960
Milutinovic PS, Yang L, Cantor RS, Eger EI 2nd, Sonner JM (2007) Anesthetic-like modulation of a gamma-aminobutyric acid type A, strychnine-sensitive glycine, and N-methyl-d-aspartate receptors by coreleased neurotransmitters. Anesth Analg 105(2):386–392
Minton AP (2000) Implications of macromolecular crowding for protein assembly. Curr Opin Struct Biol 10(1):34–39
Mitra K, Ubarretxena-Belandia I, Taguchi T, Warren G, Engelman DM (2004) Modulation of the bilayer thickness of exocytic pathway membranes by membrane proteins rather than cholesterol. Proc Natl Acad Sci U S A 101(12):4083–4088
Moon CP, Fleming KG (2011) Side-chain hydrophobicity scale derived from transmembrane protein folding into lipid bilayers. Proc Natl Acad Sci U S A 108(25):10174–10177
Moon CP, Zaccai NR, Fleming PJ, Gessmann D, Fleming KG (2013) Membrane protein thermodynamic stability may serve as the energy sink for sorting in the periplasm. Proc Natl Acad Sci U S A 110(11):4285–4290
Morein S, Andersson A, Rilfors L, Lindblom G (1996) Wild-type Escherichia coli cells regulate the membrane lipid composition in a “window” between gel and non-lamellar structures. J Biol Chem 271(12):6801–6809
Morrison EA, Dekoster GT, Dutta S, Vafabakhsh R, Clarkson MW, Bahl A, Kern D, Ha T, Henzler-Wildman KA (2012) Antiparallel EmrE exports drugs by exchanging between asymmetric structures. Nature 481(7379):45–50
Navarro J, Toivio-Kinnucan M, Racker E (1984) Effect of lipid composition on the calcium/adenosine 5′-triphosphate coupling ratio of the Ca2+-ATPase of sarcoplasmic reticulum. Biochemistry 23(1):130–135
Nicolson GL (2014) Thefluid-mosaic model of membrane structure: still relevant to understanding the structure, function and dynamics of biological membranes after more than 40 years. Biochem Biophys Acta Biomembr 1838(6):1451–1466
Noinaj N, Kuszak AJ, Gumbart JC, Lukacik P, Chang HS, Easley NC, Lithgow T, Buchanan SK (2013) Structural insight into the biogenesis of beta-barrel membrane proteins. Nature 501(7467):385–390
Noinaj N, Kuszak AJ, Balusek C, Gumbart JC, Buchanan SK (2014) Lateral opening and exit pore formation are required for BamA function. Structure 22(7):1055–1062
Oberai A, Joh NH, Pettit FK, Bowie JU (2009) Structural imperatives impose diverse evolutionary constraints on helical membrane proteins. Proc Natl Acad Sci U S A 106(42):17747–17750
Osenkowski P, Ye W, Wang R, Wolfe MS, Selkoe DJ (2008) Direct and potent regulation of gamma-secretase by its lipid microenvironment. J Biol Chem 283(33):22529–2240
Pareek S, Notterpek L, Snipes GJ, Naef R, Sossin W, Laliberte J, Iacampo S, Suter U, Shooter EM, Murphy RA (1997) Neurons promote the translocation of peripheral myelin protein 22 into myelin. J Neurosci 17(20):7754–7762
Park E, Rapoport TA (2012) Mechanisms of Sec61/SecY-mediated protein translocation across membranes. Annu Rev Biophys 41:21–40
Pasyk EA, Foskett JK (1995) Mutant (Delta-F508) Cystic-fibrosis transmembrane conductance regulator Cl- channel is functional when retained in endoplasmic-reticulum of mammalian cells. J Biol Chem 270(21):12347–12350
Patel GJ, Kleinschmidt JH (2013) The lipid bilayer-inserted membrane protein BamA of escherichia coli facilitates insertion and folding of outer membrane protein A from its complex with Skp. Biochemistry 52(23):3974–3986
Perozo E, Cortes DM, Sompornpisut P, Kloda A, Martinac B (2002a) Open channel structure of MscL and the gating mechanism of mechanosensitive channels. Nature 418(6901):942–948
Perozo E, Kloda A, Cortes DM, Martinac B (2002b) Physical principles underlying the transduction of bilayer deformation forces during mechanosensitive channel gating. Nat Struct Biol 9(9):696–703
Pester O, Barrett PJ, Hornburg D, Hornburg P, Probstle R, Widmaier S, Kutzner C, Durrbaum M, Kapurniotu A, Sanders CR, Scharnagl C, Langosch D (2013) The backbone dynamics of the amyloid precursor protein transmembrane helix provides a rationale for the sequential cleavage mechanism of gamma-secretase. J Am Chem Soc 135(4):1317–1329
Popot JL, Engelman DM (1990) Membrane protein folding and oligomerization: the two-stage model. Biochemistry 29(17):4031–4037
Popot JL, Engelman DM (2000) Helical membrane protein folding, stability, and evolution. Annu Rev Biochem 69:881–922
Powell GL, Knowles PF, Marsh D (1985) Association of spin-labelled cardiolipin with dimyristoyl phosphatidylcholine-substituted bovine heart cytochrome c oxidase. A generalized specificity increase rather than highly specific binding sites. Biochim Biophys Acta 816(1):191–194
Pucadyil TJ, Chattopadhyay A (2004) Cholesterol modulates ligand binding and G-protein coupling to serotonin (1A) receptors from bovine hippocampus. Biochim Biophys Acta 1663(1–2):188–200
Rajan RS, Kopito RR (2004) Role of protein misfolding in retinal degeneration caused by a mutation in rhodopsin. Abstr Am Chem Soc 227:U220
Rapoport TA (2007) Protein translocation across the eukaryotic endoplasmic reticulum and bacterial plasma membranes. Nature 450(7170):663–669
Rasmussen SG, Choi HJ, Rosenbaum DM, Kobilka TS, Thian FS, Edwards PC, Burghammer M, Ratnala VR, Sanishvili R, Fischetti RF, Schertler GF, Weis WI, Kobilka BK (2007) Crystal structure of the human beta2 adrenergic G-protein-coupled receptor. Nature 450(7168):383–387
Ravid T, Kreft SG, Hochstrasser M (2006) Membrane and soluble substrates of the Doa10 ubiquitin ligase are degraded by distinct pathways. EMBO J 25(3):533–543
Reinstein E, Ciechanover A (2006) Narrative review: protein degradation and human diseases: the ubiquitin connection. Ann Intern Med 145(9):676–684
Sakakura M, Hadziselimovic A, Wang Z, Schey KL, Sanders CR (2011) Structural basis for the trembler-J phenotype of Charcot-Marie-Tooth disease. Structure 19(8):1160–1169
Sanders CR, Myers JK (2004) Disease-related misassembly of membrane proteins. Annu Rev Biophys Biomol Struct 33:25–51
Sato BK, Schulz D, Do PH, Hampton RY (2009) Misfolded membrane proteins are specifically recognized by the transmembrane domain of the Hrd1p ubiquitin ligase. Mol Cell 34(2):212–222
Seddon JM, Templer RH (1995) Polymorphism of lipid-water systems (chapter 3). In: Lipowsky R, Sackmann E (eds) Handbook of biological physics, volume 1. Elsevier, Amsterdam
Sheppard DN, Welsh MJ (1999) Structure and function of the CFTR chloride channel. Physiol Rev 79(1 Suppl):S23–S45
Shimkets RA (1996) Hot papers – hypertension genetics – Liddle’s syndrome: heritable human hypertension caused by mutations of the beta subunit of the epithelial sodium channel by R.A. Shimkets DG, Warnock CM, Bositis C, Nelson-Williams JH, Hansson M, Schambelan JR, Gill S, Ulick RV, Milora JW, Findling CM, Canessa BC, Rossier RP. Lifton – Comments. Scientist 10(22):14
Shinoda T, Ogawa H, Cornelius F, Toyoshima C (2009) Crystal structure of the sodium-potassium pump at 2.4 A resolution. Nature 459(7245):446–450
Shinzawa-Itoh K, Aoyama H, Muramoto K, Terada H, Kurauchi T, Tadehara Y, Yamasaki A, Sugimura T, Kurono S, Tsujimoto K, Mizushima T, Yamashita E, Tsukihara T, Yoshikawa S (2007) Structures and physiological roles of 13 integral lipids of bovine heart cytochrome c oxidase. EMBO J 26(6):1713–1725
Shyamsunder E, Gruner SM, Tate MW, Turner DC, So PT, Tilcock CP (1988) Observation of inverted cubic phase in hydrated dioleoylphosphatidylethanolamine membranes. Biochemistry 27(7):2332–2336
Siegel DP (1993) Energetics of intermediates in membrane fusion: comparison of stalk and inverted micellar intermediate mechanisms. Biophys J 65(5):2124–2140
Simons K, Sampaio JL (2011) Membrane organization and lipid rafts. Cold Spring Harb Perspect Biol 3(10):a004697
Singer SJ, Nicolson GL (1972) The fluid mosaic model of the structure of cell membranes. Science 175(4023):720–731
Song Y, Hustedt EJ, Brandon S, Sanders CR (2013) Competition between homodimerization and cholesterol binding to the C99 domain of the amyloid precursor protein. Biochemistry 52(30):5051–5064
Song Y, Kenworthy AK, Sanders CR (2014a) Cholesterol as a co-solvent and a ligand for membrane proteins. Protein Sci 23(1):1–22
Song Y, Mittendorf KF, Lu Z, Sanders CR (2014b) Impact of bilayer lipid composition on the structure and topology of the transmembrane amyloid precursor C99 protein. J Am Chem Soc 136(11):4093–4096
Spector AA, Yorek MA (1985) Membrane lipid composition and cellular function. J Lipid Res 26(9):1015–1035
Staub O, Gautschi I, Ishikawa T, Breitschopf K, Ciechanover A, Schild L, Rotin D (1997) Regulation of stability and function of the epithelial Na+channel (ENaC) by ubiquitination. EMBO J 16(21):6325–6336
Stenson PD, Ball E, Howells K, Phillips A, Mort M, Cooper DN (2008) Human gene mutation database: towards a comprehensive central mutation database. J Med Genet 45(2):124–126
Suh BC, Hille B (2008) PIP2 is a necessary cofactor for ion channel function: how and why? Annu Rev Biophys 37:175–195
Suh BC, Inoue T, Meyer T, Hille B (2006) Rapid chemically induced changes of PtdIns(4,5)P2 gate KCNQ ion channels. Science 314(5804):1454–1457
Suh BC, Leal K, Hille B (2010) Modulation of high-voltage activated Ca(2+) channels by membrane phosphatidylinositol 4,5-bisphosphate. Neuron 67(2):224–238
Sun J, Wu J, Carrasco N, Kaback HR (1996) Identification of the epitope for monoclonal antibody 4B1 which uncouples lactose and proton translocation in the lactose permease of Escherichia coli. Biochemistry 35(3):990–998
Sun F, Zhang RL, Gong XY, Geng XH, Drain PF, Frizzell RA (2006) Derlin-1 promotes the efficient degradation of the cystic fibrosis transmembrane conductance regulator (CFTR) and CFTR folding mutants. J Biol Chem 281(48):36856–36863
Szule JA, Fuller NL, Rand RP (2002) The effects of acyl chain length and saturation of diacylglycerols and phosphatidylcholines on membrane monolayer curvature. Biophys J 83(2):977–984
Takeda K, Sato H, Hino T, Kono M, Fukuda K, Sakurai I, Okada T, Kouyama T (1998) A novel three-dimensional crystal of bacteriorhodopsin obtained by successive fusion of the vesicular assemblies. J Mol Biol 283(2):463–474
Tamm LK, Hong H, Liang BY (2004) Folding and assembly of beta-barrel membrane proteins. Biochem Biophys Acta Biomembr 1666(1–2):250–263
Tate MW, Gruner SM (1987) Lipid polymorphism of mixtures of dioleoylphosphatidylethanolamine and saturated and monounsaturated phosphatidylcholines of various chain lengths. Biochemistry 26(1):231–236
Tatsuta T, Langer T (2008) Quality control of mitochondria: protection against neurodegeneration and ageing. EMBO J 27(2):306–314
Tieleman DP, Hess B, Sansom MS (2002) Analysis and evaluation of channel models: simulations of alamethicin. Biophys J 83(5):2393–2407
Tisi R, Belotti F, Wera S, Winderickx J, Thevelein JM, Martegani E (2004) Evidence for inositol triphosphate as a second messenger for glucose-induced calcium signalling in budding yeast. Curr Genet 45(2):83–89
Valiyaveetil FI, Zhou Y, Mackinnon R (2002) Lipids in the structure, folding, and function of the KcsA K+ channel. Biochemistry 41(35):10771–10777
Van Den Brink-Van Der Laan E, Chupin V, Killian JA, De Kruijff B (2004a) Stability of KcsA tetramer depends on membrane lateral pressure. Biochemistry 43(14):4240–4250
Van Den Brink-Van Der Laan E, Killian JA, De Kruijff B (2004b) Nonbilayer lipids affect peripheral and integral membrane proteins via changes in the lateral pressure profile. Biochim Biophys Acta 1666(1–2):275–288
Van Meer G, Voelker DR, Feigenson GW (2008) Membrane lipids: where they are and how they behave. Nat Rev Mol Cell Biol 9(2):112–124
Von Heijne G (1992) Membrane protein structure prediction. Hydrophobicity analysis and the positive-inside rule. J Mol Biol 225(2):487–494
Von Heijne G (2006) Membrane-protein topology. Nat Rev Mol Cell Biol 7(12):909–918
Wang C, Jiang Y, Ma J, Wu H, Wacker D, Katritch V, Han GW, Liu W, Huang XP, Vardy E, Mccorvy JD, Gao X, Zhou XE, Melcher K, Zhang C, Bai F, Yang H, Yang L, Jiang H, Roth BL, Cherezov V, Stevens RC, Xu HE (2013) Structural basis for molecular recognition at serotonin receptors. Science 340(6132):610–614
White SH, Von Heijne G (2008) How translocons select transmembrane helices. Annu Rev Biophys 37:23–42
White SH, Wimley WC (1999) Membrane protein folding and stability: physical principles. Annu Rev Biophys Biomol Struct 28:319–365
Wickner S, Maurizi MR, Gottesman S (1999) Posttranslational quality control: folding, refolding, and degrading proteins. Science 286(5446):1888–18893
Wiggins P, Phillips R (2004) Analytic models for mechanotransduction: gating a mechanosensitive channel. Proc Natl Acad Sci U S A 101(12):4071–4076
Wimley WC (2003) The versatile beta-barrel membrane protein. Curr Opin Struct Biol 13(4):404–411
Wimley WC, White SH (1992) Partitioning of tryptophan side-chain analogs between water and cyclohexane. Biochemistry 31(51):12813–12818
Wimley WC, White SH (1996) Experimentally determined hydrophobicity scale for proteins at membrane interfaces. Nat Struct Biol 3(10):842–848
Winckler B, Forscher P, Mellman I (1999) A diffusion barrier maintains distribution of membrane proteins in polarized neurons. Nature 397(6721):698–701
Wu CC, Maccoss MJ, Howell KE, Yates JR 3rd (2003) A method for the comprehensive proteomic analysis of membrane proteins. Nat Biotechnol 21(5):532–538
Yanagisawa K (2005) Cholesterol and amyloid beta fibrillogenesis. Subcell Biochem 38:179–202
Yang L, Huang HW (2002) Observation of a membrane fusion intermediate structure. Science 297(5588):1877–1879
Zhang W, Bogdanov M, Pi J, Pittard AJ, Dowhan W (2003) Reversible topological organization within a polytopic membrane protein is governed by a change in membrane phospholipid composition. J Biol Chem 278(50):50128–50135
Zhang W, Campbell HA, King SC, Dowhan W (2005) Phospholipids as determinants of membrane protein topology – phosphatidylethanolamine is required for the proper topological organization of the gamma-aminobutyric acid permease (GabP) of Escherichia coli. J Biol Chem 280(28):26032–26038
Zhou Y, Morais-Cabral JH, Kaufman A, Mackinnon R (2001) Chemistry of ion coordination and hydration revealed by a K+ channel-Fab complex at 2.0 A resolution. Nature 414(6859):43–48
Acknowledgements
This works was supported by start-up fund from Michigan State University and Hunt for Cure foundation cystic fibrosis research grant to H.H. The author is thankful for critical comments and suggestions by Hong lab members, anonymous reviewers and Dr. Olga Gursky, the volume editor.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Hong, H. (2015). Role of Lipids in Folding, Misfolding and Function of Integral Membrane Proteins. In: Gursky, O. (eds) Lipids in Protein Misfolding. Advances in Experimental Medicine and Biology, vol 855. Springer, Cham. https://doi.org/10.1007/978-3-319-17344-3_1
Download citation
DOI: https://doi.org/10.1007/978-3-319-17344-3_1
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-17343-6
Online ISBN: 978-3-319-17344-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)