Abstract
During skin development and keratinocyte differentiation, the nucleus undergoes characteristic changes in shape, size, and transcriptional output. Many of these changes are regulated by the interface between the nuclear interior and the inner nuclear membrane, a region called the nuclear lamina. The nuclear lamina is composed of a meshwork of the nuclear lamins, which interact with integral inner nuclear membrane proteins, and the associated chromatin. Studies in the last decade have revealed a view of the nuclear lamina as a hub for biochemical and mechanical inputs. In this chapter, we will discuss how the structure and organization of the nucleus allows biochemical and mechanical signals to regulate gene expression, genome integrity, cell and tissue level mechanics, and disease in the context of skin homeostasis and regeneration.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
11.1 Introduction: Skin Epithelial Keratinocytes and the Nucleus
The skin and its appendages, including hair follicles, sweat glands, and sebaceous glands, provide essential functions for animal survival, such as protection from water loss and environmental insults, as well as the capacity for tactile sense. The interfollicular epidermis (IFE), which composes the outermost layer of the skin, consists of a series of functionally and structurally distinct stratified layers. The basal layer contains proliferative progenitor keratinocytes that tightly bind to extracellular matrix (ECM) molecules in the basement membrane, which separates the epidermis from the underlying dermis. During embryonic development, basal keratinocytes divide asymmetrically, giving rise to daughter cells that move upwards to form the suprabasal layers (spinous, granular, and stratum corneum); the resulting cells of the stratified epithelium generate a cornified envelope essential for the barrier function of the skin [1]. A subset of basal cells also gives rise to hair follicles and other appendages [2,3,4].
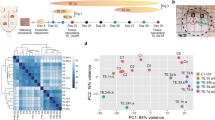
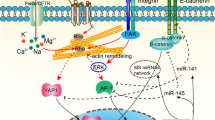
Keratinocyte differentiation during epidermal development requires coordinated changes in the cytoskeleton that support the generation and mechanical stability of cell-cell adhesions, including adherens junctions (AJs), desmosomes, and tight junctions. Keratin intermediate filament proteins, the major structural component of the IFE, interact with integrin-based hemidesmosome adhesions that connect the basal IFE to the dermis (Fig. 11.1a). E-cadherin-mediated cell-cell junctions, located throughout the epidermis and its appendages, connect to actin and microtubule networks through catenin molecules [5]. As basal cells move upward during differentiation, another class of intercellular junction, desmosomes, which contain desmosomal cadherins and are connected to keratins, becomes abundant (Fig. 11.1a). Desmosome density and desmosomal cadherin composition markedly change as cells differentiate [6]. Finally, tight junctions assemble in the granular layer [7,8,9]. These junctions provide mechanical stability to the epidermis and are essential for the barrier function of the skin.
(a) The nuclear lamina allows crosstalk between the nuclear envelope and plasma membrane. The nuclear periphery is a richly complicated cellular compartment that functions in chromatin organization, transcriptional regulation, mechanosensing, integration of the nucleus into the cytoskeleton, and DNA damage repair, among other functions. In epidermal keratinocytes, the nuclear lamina network is mechanically integrated into the cytoskeleton via nuclear envelope-spanning LINC complexes. Actin, microtubules, and intermediate filaments interact with the LINC complex through either direct or indirect interactions. Through these cytoskeletal connections, forces exerted on plasma membrane adhesions from either the extracellular matrix, mediated by Fig. 11.1 (continued) integrin-based focal adhesions and hemidesmosomes, or from adjacent cells, mediated by adherens junctions and desmosomes, can be transmitted to the nuclear interior to alter cellular function. Likewise, recent work indicates that changes at the nuclear envelope can alter the function of the cytoskeleton and plasma membrane adhesions. Thus, crosstalk exists between plasma membrane adhesions, the cytoskeleton, and components of the nuclear periphery. (b) Schematic of nuclear periphery components involved in epidermal homeostasis. The nuclear envelope is composed of both inner and outer nuclear membranes (INM and ONM), as well as conduits for macromolecular transport known as nuclear pore complexes (NPCs). Constituents of the nuclear periphery, including integral INM proteins, peripheral membrane proteins, and chromatin-binding proteins have all been shown to influence epidermal development and function. The LINC complex, which consists of SUN and KASH domain proteins, spans the INM, perinuclear space, and ONM to connect the nuclear periphery to the cytoskeleton. A number of other significant integral INM proteins function at the nuclear periphery, including the LAP2-Emerin-MAN1 (LEM) domain proteins, lamin B receptor (LBR), LAP1/Torsin, and a multitude of additional nuclear envelope transmembrane proteins (NETs) with currently unknown functions. Lamins A/C and lamin B1 and B2 form a meshwork underlying the INM. A number of chromatin-binding proteins, which also exhibit crosstalk with other nuclear periphery components, modulate chromatin organization and function, including the chromatin remodeling factor Brg1, the chromatin organizer Satb1, polycomb group proteins, heterochromatin-binding protein HP-1, histone methyltransferases (HMTs), histone deacetylases (HDACs), and a number of transcription factors including p63 and the AP-1 complex. In general, lamina-associated domains (LADs) are rich in the histone mark H3K9me, while H3K27me is commonly found at the LAD borders, known as variable LADs (vLAD). In the context of epidermal differentiation, a switch from H3K27me to H3K4me and H3K79me marks has been shown to mediate the transcriptional activation of lineage commitment genes
During the process of terminal differentiation in the IFE and during hair follicle stem cell activation and differentiation, the cell nucleus goes through remarkable changes in shape and size [10]. In the most differentiated cells in the IFE and hair follicle, a poorly understood process leads to beneficial loss of the nucleus. When intact, the nucleus is bounded by the nuclear envelope that serves to physically and functionally segregate the genome from the cytoplasmic milieu. Research over the last several decades has highlighted that, in addition to this role, the nucleus additionally serves to integrate transcriptional, biochemical, and mechanical networks within cells and tissues. In this chapter, we will discuss how the structure and organization of the nucleus impacts gene expression, genome integrity, cell and tissue level mechanics, and disease in the context of skin homeostasis and regeneration.
11.2 The Nuclear Lamina and Nuclear Organization
The nucleus is organized into distinct but dynamic subnuclear compartments, including chromosome territories and nuclear bodies (nucleoli, Cajal bodies, etc). One critical nuclear landmark is the nuclear envelope, which is contiguous with the endoplasmic reticulum and is comprised of both an inner and outer nuclear membrane (INM and ONM, respectively). The interface between the nuclear interior and the INM is characterized by the nuclear lamina, a highly complex proteinaceous structure that consists of a meshwork of the nuclear lamins, an array of lamin-interacting integral INM proteins, and the associated chromatin, which is typically heterochromatic in nature [11]. In addition, the nuclear lamina is physically connected across the nuclear envelope lumen to integral ONM proteins that mechanically attach the cytoskeleton to the chromatin in the nuclear interior [12].
In vitro studies have provided some insight into the potential structure of lamins, which are members of the type V intermediate filament protein family [13]. Much like other intermediate filament proteins, the lamins exist as parallel coiled-coil dimers in vitro, and interact in a head-to-tail fashion to generate long polymers [14]. These polymers also associate with one another in parallel to give rise to large networks [15,16,17]. In vivo studies using super-resolution microscopy suggest that the A-type and B-type lamins are organized into distinct meshworks associated with the INM [18, 19]. The lamins are classified as A-type or B-type depending on their chemical and structural qualities. The B-type lamins, lamin B1 and B2, are transcribed from the separate LMNB1 and LMNB2 genes, while A-type lamins, lamins A and C (lamin A/C), are derived from alternative splicing of the single LMNA gene. While B-type lamins are expressed in all metazoan cell types throughout development, including embryonic stem cells, A-type lamins are only found in differentiated cells [20,21,22,23,24,25]. Most A- and B-type lamins are farnesylated during biosynthesis, the exception being lamin C. While farnesylation of B-type lamins appears to be constitutive and promotes association with the INM, A-type lamins undergo proteolytic cleavage by the zinc metalloproteinase Zmpste24 to remove the farnesylated C-terminal region [26]. This processing is critical, as a mutation that disrupts cleavage of the lamin A farnesylated tail gives rise to the pathogenic “progerin” form of lamin A and is the cause of the accelerated aging disorder progeria [27].
Interestingly, lamin composition varies with both differentiation state and tissue type [22, 23]. Although the broad requirements for lamin A/C in skin biology have not yet been examined in great depth, lamins have been shown to play roles in epidermal differentiation and development. Interestingly, lamin A/C and LBR appear to have temporally distinct functions during cellular differentiation in various mammalian cell types, although at least one must be present for proper tethering of heterochromatin to the INM [28]. Using the hair follicle as a model tissue that contains both highly differentiated and undifferentiated cells, the authors of this study found that transit amplifying matrix progenitors and bulge stem cells in the hair follicle express only LBR, while the more differentiated cells of the follicular dermal papilla, epidermis, and dermis express lamin A/C [28]. Emerin and other LEM domain proteins may be particularly important in dermal papilla cells [28]. Further, mice expressing the progerin form of lamin A that is associated with premature aging exhibit aberrant LBR expression in suprabasal layers, in many cases concomitant with aberrant compaction of DNA at the nuclear periphery [29]. Epidermal differentiation appears sensitive to these defects, as these mice also display epidermal thickening and mislocalization of keratin 5 to suprabasal layers [29]. Other work has shown that simultaneous ablation of lamins A/C, lamin B1, and lamin B2 in mice produces a thickened epidermis with abnormal development of the stratum corneum and hypotrophic hair follicles, coincident with incursion of endoplasmic reticulum components into the chromatin of epidermal keratinocytes [30].
While lamin B1 and B2 are uniformly distributed in human epidermal cells, lamin A/C may be expressed at lower levels in human basal layer cells of the IFE [24]. However, a similar analysis of lamin expression in murine skin found high levels of lamin A/C in the basal layer, with decreased expression in suprabasal layers, suggesting either differences in the expression pattern of these proteins in mice and humans, or differences in antibody accessibility [25, 31].
The disassembly of lamin A/C during keratinocyte terminal differentiation is required for epidermal function [32]. Normally, lamin A/C are highly phosphorylated by the serine/threonine kinase AKT1 in the granular and cornified layers of the IFE, and loss of this activity results in parakeratosis, the retention of undegraded nuclei [32]. Deletion of AKT1 also produces increased BMP2/SMAD1 signaling and altered terminal differentiation in the IFE, including decreased keratin 1/10 and loricrin expression [32]. Interestingly, mice lacking keratins 1/10 exhibit premature nuclear degradation, with concomitant reductions in the levels of lamin A/C and the INM proteins emerin and SUN1 in suprabasal keratinocytes [33]. These results suggest that a feedback mechanism exists in the IFE between the nuclear lamina and cellular signaling, although the mechanistic basis for this signaling circuit remains unclear.
In addition to farnesylation, multiple lamin-binding integral INM proteins reinforce interactions between the lamins and the INM. The largest class of these proteins is comprised of members of the LAP2, emerin, and MAN1 (LEM) domain family [34]. These proteins share a common 40 amino acid motif known as the LEM domain that facilitates their interaction with chromatin through the DNA-binding Barrier-to-Autointegration Factor (BAF or BANF1) . Additional proteins such as LAP1 (lamina-associated polypeptide 1), LAP2 (lamina-associated polypeptide 2) and LBR (lamin B receptor) also interact with lamins, as well as carrying out distinct biochemical functions [35,36,37]. Of particular interest are the lamin-binding SUN (Sad2/UNC-84) domain proteins that make up the nuclear aspect of the LInker of Nucleoskeleton and Cytoskeleton (LINC) complex. SUN domain proteins, found in the INM, bind to the ONM-resident KASH (Klarsicht/ANC-1/SYNE homology) domain proteins (also called Nesprins) in the nuclear envelope lumen, allowing LINC complexes to bridge both membranes of the nuclear envelope [12]. KASH domain proteins facilitate either direct or indirect interactions with actin, microtubules, and intermediate filaments in the cytoplasm, thereby facilitating a continuous connection between the nuclear interior and cytoplasmic cytoskeleton [12]. Interestingly, mutations in many of the lamin-associated INM proteins drive diseases (so-called “nuclear envelopathies”) that phenocopy those associated with mutations in the lamins themselves [38], suggesting both complex and integrated functions for the nuclear lamina.
11.3 The Nuclear Lamina Is Mechanically Integrated into the Cell by the LINC Complex
As in most tissues, the cells of the epidermis and dermis contain an interconnected cytoskeletal network that can propagate forces from externally applied mechanical stimuli. Forces from both ECM adhesions and cell-cell junctions can propagate to the nuclear interior [39, 40] (Fig. 11.1a) and reorganize nuclear proteins [22, 41, 42]. One of the earliest demonstrations of this continuous connection was performed by applying pulling forces to plasma membrane-localized integrins on single cells using RGD-coated microbeads and monitoring nuclear distention in the direction of pulling [40]. Magnetic twisting cytometry induces dissociation of the nuclear-localized Cajal body components coilin and SMN within seconds in HeLa cells, indicating that mechanical stimulation is sufficient to induce changes to nuclear architecture [42]. Intranuclear movements have also been observed in response to compressive and shear forces [43].
How are mechanical signals communicated between the cell surface and the nuclear interior? The LINC complexes at the nuclear envelope provide a conduit for forces to be transmitted from the ECM or adjacent cells, through the cytoskeleton, to the nuclear lamina (Fig. 11.1a). Indeed, the LINC complex is essential for cytoskeletal-nuclear force transmission, as its disruption eliminates the nuclear response to mechanical stimuli [44,45,46,47] and abrogates cytoskeleton-dependent nuclear positioning in a broad range of contexts (reviewed in [48]). An emerging area of research also suggests that alterations in the composition of the nuclear lamina can likewise impact cell-ECM adhesions [49,50,51,52,53,54] (Fig. 11.1a).
While the nuclear response to ECM-derived forces is well established, cell-cell adhesion can also deliver forces on the nucleus to alter nuclear position [55, 56], suggesting a link between the nucleus and intercellular adhesions. Several components of the LINC complex are expressed in the IFE and its appendages, including multiple KASH domain-containing proteins and the widely expressed SUN domain proteins SUN1 and SUN2. Surprisingly, SUN1 is absent from the hair follicle after morphogenesis, and mice lacking SUN2 display transient alopecia during the first hair growth cycle [56]. Ultrastructural and cell culture experiments revealed alterations in the integrity of cell-cell adhesion, possibly due to a loss of LINC complex-dependent organization and/or maintenance of the cytoskeletal network and desmosome based adhesions [56]. Thus, the nucleus plays an essential role in allowing epidermal keratinocytes to generate mechanically stable cell-cell adhesions.
In addition to changes in nuclear position and the organization of intranuclear components, forces propagated to the nuclear interior can also influence the nuclear proteome and nuclear mechanics (Fig. 11.2). The expression level of lamin A/C scales with tissue stiffness in vivo and with substrate stiffness in vitro: lamin A/C levels are high in stiff tissues like bone and lower in soft tissues like brain [22]. As a result, nuclear stiffness scales with ECM stiffness, giving rise to the concept of “mechanical reciprocity” between the nucleus and the substrate that cells are grown on. Interestingly, the correlation of lamin expression level with stiffness appears to be a specific function of lamin A/C, as lamin B levels do not vary as drastically between tissue or substrate types [22]. Additionally, B-type lamins do not seem to play the same role as lamin A/C in influencing nuclear mechanics [57]. Importantly, while low substrate stiffness is known to decrease keratinocyte-ECM adhesion to induce differentiation [58], the effect of cell-ECM or intercellular adhesion-derived forces on lamin A/C levels and/or their polymerization state in keratinocytes has not yet been investigated.
The mechanosensitive function of the nuclear periphery modulates epidermal function. On softer substrates/tissues or when cells are not exposed to stretch, cytoskeletal tension is low. Signaling proteins involved in the sensing of cytoskeletal tension, like MKL and YAP/TAZ, remain inactive in the cytoplasm. Low levels of force are exerted on the LINC complex via the cytoskeleton. Emerin is localized to the inner nuclear membrane, where it associates with lamins A/C. Under these conditions, epitopes in lamins A/C are available for phosphorylation, leading to rapid turnover of the lamin network. In human epidermal stem cells, epidermal differentiation loci on chromosomes 1 and 18 associate with the lamina, most likely mediated by H3K9me chromatin marks. On stiffer substrates/tissues or upon cell stretching, higher cytoskeletal forces are exerted on the LINC complex and transmitted to the nuclear interior; cellular compression forces can also be transmitted to the nuclear interior without the LINC complex. These changes in cytoskeletal organization promote nuclear localization of MKL and YAP/TAZ, and the transcriptional activation of their target genes; importantly, the nuclear periphery plays a role in modulating the activity of these factors. Lamins A/C undergo reinforcement in response to elevated cytoskeletal tension, leading to multimerization and stabilization of the lamin network. This process may involve the burial of force-sensitive epitopes in lamins A/C, preventing their phosphorylation and turnover. When under tension, the LINC complex exhibits stronger association with lamins A/C, while emerin is phosphorylated, potentially weakening its interaction with the nuclear lamina. In human epidermal stem cells, this altered association with lamins A/C could drive the relocation of emerin from the inner nuclear membrane to the outer nuclear membrane, allowing for the emerin-dependent formation of an actomyosin cage around the nucleus. Reduced emerin localization to the inner nuclear membrane facilitates loss of H3K9me and accumulation of H3K27me marks on the promoters of epidermal differentiation loci, resulting in global chromatin rearrangements and dissociation from the periphery. Combined with PRC2 association and reduced RNAPII accumulation at promoters, these changes drive transcriptional silencing of lineage commitment genes. The reinforcement of the lamin network, the formation of a perinuclear actomyosin cage, and the release of chromatin from the nuclear periphery may protect the genome from force-induced damage under conditions of high cytoskeletal strain
While the structural integrity provided by the lamin A/C polymer network is proposed to impart nuclear stiffness directly, the associated chromatin may also influence the mechanical properties of the nucleus. Mechanical measurements made with optical tweezers on isolated nuclei derived from a fission yeast model, Schizosaccharomyces pombe , which lacks lamins, have shown that chromatin association with the nuclear periphery can impart nuclear stiffness in the absence of lamins, and can influence the viscoelastic properties of nuclei [59]. Thus, the peripherally localized heterochromatin that associates with the lamina may also play a role in defining the mechanics of nuclei. Indeed, evidence suggests that heterochromatin may also buffer forces exerted on the nucleus, acting in concert with the lamins [60,61,62]. Cell migration, which is a mechanically taxing process for the nucleus, leads to an increase in facultative heterochromatin at the leading edge of the nucleus in a mouse melanoma cell line, further suggesting that chromatin condensation and nuclear mechanical challenge can be linked [63]. The lamin network may also influence local chromatin mobility, which is limited by the association of heterochromatin with the nuclear periphery [64]. Indeed, the loss of lamin A in mammalian cells [65] or INM proteins in yeast [59] results in increased chromatin mobility, which can strongly influence nuclear mechanics, particularly nuclear viscosity, and therefore nuclear function.
In addition to changes in expression level, a central theme in the response of the nuclear lamina to mechanical input appears to be force-dependent changes in lamin A/C multimerization [31], which may serve several functions (Fig. 11.2). First, reinforcement of the lamin A/C network and subsequent nuclear stiffening may improve the efficiency of force transmission from the cytoskeleton through the LINC complex to the lamin network. This change in nuclear stiffness would be important for processes where the nucleus must be actively positioned or moved, such as during polarization prior to cell migration [66]. Second, the differential exposure of binding sites for interaction partners, including chromatin, histones, and INM proteins, may serve as a regulatory mechanism for guiding signal transduction and/or transcription. Finally, modulation of the lamin network may influence how susceptible the genome is to mechanically-induced DNA damage. Defects in the remodeling of the lamina, including chromatin, in response to force could result in increased damage; this may explain why the ablation of SUN proteins or lamin A/C can produce increases in baseline DNA damage [67] and apoptosis in response to mechanical challenge [68].
11.4 The Nuclear Lamina and Chromatin Organization
In addition to its roles in defining nuclear mechanics, the nuclear lamina, and lamin A/C and LBR in particular, sequester heterochromatic regions of the genome at the nuclear periphery [28, 69, 70]. Lamina associated domains (LADs), which account for 40% of the mammalian genome [71], are recruited to lamins through the coordinated activity of transcription factors, distinctly localized heterochromatic marks, and sequence specificity (Fig. 11.1b). Lamina-associated sequences, sometimes enriched in extended GA-rich sequence motifs, in combination with the facultative heterochromatin marks H3K27me3 and H3K9me2/3, are capable of driving peripheral association of ectopic LADs [72, 73]. These marks have been identified in other studies as important facilitators of lamina tethering [71, 74, 75]. However, GA repeats are not found in all LADs [71], and indeed non-GA sequences in LADs appear to be sufficient for lamina-targeting [72]. The transcription factor YY1, as well as histone deacetylase 3 (HDAC3), LAP2β, and lamin A/C appear to be required for peripheral recruitment [72], although additional transcription factors such as Zbtb7b may also be involved in particular cases, such as in recruiting the IgH locus to the lamina in fibroblasts [73]. Interestingly, either lamin C alone, or an optimum ratio of lamin A to lamin C, may specifically be involved in this process, suggesting distinct roles for lamin A and lamin C in LAD tethering [72].
Efforts to understand whether the tethering of individual genes to the nuclear periphery is sufficient to repress gene transcription have given mixed results, depending on the experimental system analyzed [76, 77]. Importantly, lamins appear dispensable for repression at the nuclear periphery, just as transcriptional repression is not an essential requirement for peripheral recruitment. Indeed, genomic loci with transcriptionally active histone marks are also found in this subnuclear compartment (reviewed in [78]). Further, the release of LADs from the nuclear periphery is not sufficient to induce their transcriptional activation [79]. Thus, lamin A/C does not seem to directly influence heterochromatinization, and instead merely sequesters these regions at the nuclear periphery. However, several INM proteins have an established connection with the regulation of chromatin state. For example, emerin interacts with HDAC3, an HDAC found in the NCoR complex that represses transcription, recruiting it to the nuclear lamina, and promoting its activity [80].
While techniques such as ChIP-seq and Hi-C have allowed static “images” of genome-nuclear lamina interactions to be identified, dynamic alterations in genome organization have also been observed using microscopy-based techniques [75]. Using these methods, it has been observed that LADs in mammalian cells undergo stochastic rearrangements in sub-nuclear localization, from peripheral to nucleolar regions, following mitosis [75]. These results suggest that the peripheral localization of LADs is not an essential regulator of gene expression and cell function in differentiated cells. Further, Hi-C analysis of chromosome territories in single mammalian cells suggests that chromosome organization is highly dynamic and variable between individual cells [81]. Similar analyses of LAD dynamics in differentiating epidermal cells will be an essential avenue of experimental inquiry.
While association with the lamina is not generally sufficient to predict transcriptional state, several studies suggest that the nuclear lamina helps tune gene expression in different tissue types and at different stages of cellular differentiation and development [82]. The characteristic changes in nuclear shape and size that occur during epidermal terminal differentiation occur concomitantly with changes in the 3D architecture, and therefore the “transcriptional microenvironment”, of epidermal keratinocytes [10]. Indeed, while basal progenitors exhibit markers of active transcription, increasingly suprabasal cells show global decreases in active transcriptional markers and increases in pericentromeric heterochromatin [10]. Further, a chromosomal region termed the epidermal differentiation complex (EDC), that consists of sixty consecutive genes necessary for epidermal stratification and the production of a cornified envelope (reviewed in [83]), has been observed to associate more frequently with pericentromeric heterochromatin regions in suprabasal cells, and relocates away from the nuclear periphery towards the nuclear interior [10, 84, 85]. These results suggest that functional reorganization of chromatin within the nucleus occurs during epidermal differentiation. Interestingly, relocation of the EDC locus is accompanied by the increased transcription of EDC genes, and is dependent on the chromatin remodeling factor Brg1 and the chromatin organizer Satb1; both of these chromatin interactors are regulated by p63, a transcription factor essential for epidermal specification and progenitor activity [85, 86]. In addition, the differentiation process is associated with the loss of H3K27me3 chromatin marks on differentiation-specific genes, and their replacement with H3K79me2 and H3K4me3 marks, although the full complexity of this transition is still under investigation [87] (Fig. 11.1b).
More generally, regulated histone deacetylation is important for both epidermal development and hair follicle specification. The chromatin remodeling factor Brg1, which interacts with HDAC proteins, is required for epidermal differentiation [88]. Conditional deletion of Mi-2b, a component of the HDAC1 and HDAC2-containing NURD complex, results in failure of hair follicle specification and atrophic epidermis due to loss of epidermal progenitor cells [89]. Similarly, deletion of both HDAC1 and HDAC2 in murine epidermis results in thin skin due to the upregulation of the specific target genes normally repressed by p63, as well as increases in acetylated p53 [90]. Furthermore, these mutant mice do not form hair follicles, teeth, or tongue papillae, all of which derive from basal epidermal keratinocytes. Additionally, treatment of adult skin with the HDAC inhibitor trichostatin A can induce hair follicle stem cell activity [91], while deletion of HDAC1 and HDAC2 postnatally in the epidermis results in alopecia, claw dystrophy, and hyperkeratosis [92]. Whether these functions of HDACs are related to the regulation of transcriptional repression at the nuclear periphery is unknown. Interestingly, mice lacking p63 display aberrant nuclear morphology, as well as alterations in the expression of LINC complex components, including the increased expression of SUN2 and decreased expression of SUN1, Nesprin 3, lamin A/C, lamin B1, and plectin [85, 93]. These changes are further linked to relocation of repressive heterochromatin marks H3K27me3, H3K9me3, and HP1α away from the nuclear periphery, as well as reorganization of the keratin-encoding loci KtyI and KtyII towards repressive chromocenters [93]. Thus, p63 may carry out aspects of its influence on transcriptional control of epidermal differentiation in coordination with the nuclear lamina.
In addition to histone deacetylation and chromatin remodelers, the nuclear lamina can also serve as a platform for the concentration of transcription factors and components of signaling pathways, thereby influencing gene expression. For example, the lamin A/C scaffold is postulated to regulate the transcription factor c-Fos, which heterodimerizes with c-Jun to form the AP-1 complex. The sequestration of c-Fos at the nuclear periphery by lamin A/C prevents c-Fos/c-Jun heterodimerization, attenuating AP-1 DNA binding and transcription [94]. The interaction of c-Fos with lamin A/C appears to be regulated by ERK1/2-dependent c-Fos phosphorylation, which allows it to be released from the periphery and activate transcription [95]. Interestingly, AP-1 regulates EDC gene expression in both proliferating and differentiating keratinocytes in vitro [96] and coordinates with Ezh2 in the polycomb complex to regulate epidermal differentiation [97]. Thus, the nuclear lamina may support epidermal differentiation by regulating AP-1 activity.
Several additional nuclear envelope components can influence specific signal transduction cascades, including signaling through the Wnt pathway, itself an important regulator of stem cell renewal and differentiation in the epidermis [98]. Emerin interacts with β-catenin via its adenomatous polyposis coli (APC) homology domain and antagonizes Wnt signaling, potentially by promoting β-catenin export from the nucleus [99,100,101]. Conversely, the LINC complex component Nesprin-2 may positively regulate the nuclear localization of β-catenin [102]. Thus, because the level of β-catenin signaling influences stem cell lineage choice (reviewed in [98, 103]), the nuclear lamina-dependent tuning of nuclear β-catenin levels may impact lineage selection.
TGF-β is another important modulator of epidermal stem cell activity and wound healing that is regulated by nuclear lamina components. MAN1 antagonizes TGF-β/BMP signaling by binding Smad2 and Smad3, upstream regulators of TGF-β signaling, although the mechanism of inhibition is unclear [104, 105]. Disruption of this activity by a loss-of-function mutation in MAN1 is linked to Buschke-Ollendorff syndrome, characterized by skeletal dysplasia and skin abnormalities [106]. Interestingly, increased TGF-β signaling is often seen in fibrotic skin disorders, such as those that characterize the early aging disorder progeria , associated with expression of the progerin form of lamin A [107, 108]. Once again, Nesprin-2 may act conversely to promote Smad activity, as mice expressing a mutant actin-binding domain-null Nesprin-2G exhibit delayed nuclear accumulation of Smad2/3, as well as delayed wound healing in vivo [109]. These changes have also been linked to aberrant fibroblast differentiation and keratinocyte proliferation in response to wounding [109]. Thus, in addition to the lamins, other nuclear envelope components likely act to modulate signaling cascades leading to proper stimulation and repression of signaling circuits in the epidermis.
11.5 Maintenance of Genome Integrity
In addition to regulating chromatin dynamics, activation state, and organization, increasing data suggest that the lamina also plays a role in maintaining genome integrity. Although this topic has been less well studied to date, there are likely two primary mechanisms that underlie the functions of the lamina in genome integrity: one in the prevention of DNA double-strand breaks (DSBs), and the other in modulating the efficiency and fidelity of repairing the DSBs that do occur. Mechanical stresses have been shown to induce DNA damage, leading to apoptosis, in vascular smooth muscle cells [110]. A role for the nuclear lamina in supporting mechanical stability when high force is exerted on the nucleus may protect the genome from damage. For example, mechanical stresses in the form of nuclear deformation during cellular migration through pores have been shown to induce apoptosis in lung carcinoma A549 cells with partial knockdown of lamin A [68]. While this study did not directly examine whether DNA damage was the driver of the apoptotic response, increases in genome instability, including DNA lesions and telomere dysfunction, have been observed in response to mutations in nuclear lamina components . For example, mouse embryonic fibroblasts (MEFs) from Sun1/Sun2−/− mice exhibit elevated basal DNA damage and increased sensitivity to DNA damaging agents [67], as do C. elegans deficient for the SUN domain-containing protein UNC-84 [111].
In addition, the nuclear lamina likely contributes to DSB repair pathway choice. Components of the nuclear lamina have been found to directly interact with DNA damage signaling or repair-associated proteins. Nesprin-1 physically interacts with MSH2 and MSH6, components of the mismatch repair pathway [112]. A Nesprin-2 isoform lacking a KASH domain may also influence the DNA damage response (DDR), specifically influencing ATM localization to sites of damage through its interaction with ERK1/2 [113]. Further, SUN1 and SUN2 have been implicated in modulating DDR in MEFs [67], and the budding yeast SUN protein Mps3p recruits DSBs to the nuclear periphery [114, 115]. Interestingly, the bridging of persistent DSBs through the LINC complex to dynamic cytoplasmic microtubules promotes homology-directed repair in fission yeast [116], while a similar LINC complex-mediated process may inhibit non-homologous end joining in favor of homology-directed repair in the C. elegans germline [111]. Lamin A/C may also influence DDR by directly interacting with DDR components, such as Ku70 and γ-H2AX [117], and influencing the formation of repair foci (reviewed in [118,119,120,121,122]). Indeed, MEFs null for Zmpste24 or expressing unprocessed prelamin A exhibit delayed 53BP1 and Rad51 recruitment to sites of damage, leading to defective repair and irreparable DSBs [121]. Further, defects in DNA damage repair were identified in a restrictive dermopathy-like disease – characterized by severe epidermal defects – which arose through mutation of lamin A and subsequent postnatal loss of mature lamin A expression [123]. This lamin A mutation resulted in the accumulation of DSBs in vivo, as well as decreased 53BP1 localization at breaks and impaired DNA repair in human fibroblasts [123].
While it is not yet mechanistically clear if or how defects in DNA repair contribute to progeria and other diseases of the lamina, fibroblasts from progeria patients, Zmpste24−/− mice (defective in lamin A processing), or MEFs overexpressing unprocessed prelamin A all show increased susceptibility to DNA damage [121]. Interestingly, SUN1 has been shown to accumulate in progeroid-expressing and lamin A-null mouse fibroblasts [124]. Given the potential role of SUN proteins in the DDR, Lei et al. proposed a mechanism for progerin-toxicity where excessive accumulation of SUN1 at the nuclear envelope could lead to hyperactive DDR signaling [67]. Indeed, the loss of SUN1 is capable of rescuing many of the defects in a mouse model of progeria [124]. The loss of SUN1 in this context would theoretically reduce DDR signaling in lamin A mutant mice, and minimize the associated defects. However, it remains possible that the amelioration of the progeria phenotype may instead be due to a concomitant decrease in the number of LINC complexes, and therefore the force exerted on the nucleus. The emerging connection between lamins A/C, lamin B1, and autophagy [125,126,127,128,129,130] may additionally play a role in modulating cellular senescence and aging in progeria patients. For example, epidermal keratinocytes deficient for Atg7 exhibit decreased lamin B1 expression coupled with increased DNA damage foci, altered lipid metabolism, and cellular senescence [129]. As an alternative model, progerin itself may interfere with the activity of repair factors, thereby promoting irreparable DSBs [119, 131].
As genome integrity is essential for skin homeostasis and regeneration (reviewed in [132]), a potential role for the nuclear lamina in protecting the genome and even repairing DNA has significant consequences for epidermal function. The homeostasis of different components of the epidermis, including the stratified epithelium, hair follicles, and the sebaceous glands, is maintained by resident stem cell niches; in the case of the hair follicle, stem cells located in the bulge region promote hair follicle regeneration throughout adulthood [133]. Adult bulge stem cells may utilize distinct DNA repair strategies at different developmental stages, as these cells rely on non-homologous end joining and expression of the anti-apoptotic protein Bcl2 during the resting phase of the hair cycle [134]. Interestingly, progerin has been implicated in suppressing 53BP1-mediated non-homologous end joining in human keratinocytes following UVA radiation [135]. Further, ablation of BRCA1, an important mediator of homologous recombination, in the murine epidermis results in DNA damage accumulation, apoptosis, and loss of transit amplifying cells and bulge stem cells, preventing adult regeneration of hair follicles [136]. Additional work must be carried out to determine if the nuclear lamina’s regulation of DNA repair has implications for skin homeostasis and tumorigenesis.
11.6 Mechanosensing: The Lamina and the LINC Complex
In addition to biochemical signaling, mechanical signals are also known to influence chromatin structure, gene expression, and differentiation (reviewed in [137, 138] (Fig. 11.2). Cell geometry can influence gene expression downstream of MKL1/SRF signaling and HDAC3 localization [139], while modulation of the cytoskeleton and/or LINC complexes can modify nuclear morphology, chromatin organization, and gene expression [140,141,142]. Similarly, cell geometry, substrate stiffness, and intracellular tension can influence stem cell lineage selection [58, 143, 144].
Interaction of the nucleus with the cytoskeleton through LINC complexes provides a mechanism for external forces to be propagated from the ECM or adjacent cells to the nuclear interior directly. Thus, in addition to the modulation of nuclear mechanics in response to extracellular biochemical cues, mechanosensing at the nuclear lamina is also likely critical for the cell to integrate and respond to mechanical cues from adhesions and the cytoskeleton. An attractive model is the possibility that the nuclear lamina responds to mechanical inputs in a fashion similar to that described for E-cadherin-based AJs and integrin-based focal adhesions, which increase in size in response to exogenous force [145,146,147]. The acute application of force on Nesprin-1 molecules on isolated nuclei using magnetic tweezers has been shown to produce nuclear stiffening within seconds, an effect dependent on the LINC complex, emerin, and lamin A/C [46]. Further, force application to nuclei in this context increases the association of lamin A/C with LINC complexes [46]. Nuclear stiffening in response to stress has been observed in other contexts as well: endothelial cell nuclei stiffen in response to shear stress [148], while HeLa cells exposed to shear exhibit increased peripheral recruitment of lamin A/C [149]. Interestingly, acute force application to isolated nuclei also results in emerin phosphorylation [46]. This process is required for force-induced nuclear stiffening, as the expression of a phosphomutant emerin prevents nuclear stiffening and abrogates an increased association of LINC complexes with lamin A/C [46]. As both SUN2 and emerin bind to the same region of lamin A/C, the force-induced phosphorylation of emerin in this context may decrease lamin A/C-emerin affinity and promote a lamin A/C-SUN2 interaction. In support of this notion, emerin-null nuclei exhibit increased nuclear stiffness, suggesting that the association of lamin A/C with SUN proteins is even greater than in control nuclei [46] (Fig. 11.2).
Interestingly, uniaxial stretch applied to human epidermal progenitor cells drives the redistribution of emerin from the INM to the ONM, leading to the formation of a perinuclear non-muscle myosin II-actin scaffold, nuclear actin depletion and inactivation of RNAPII, and the subsequent H3K27me3- and PRC2-dependent transcriptional repression of genes associated with epidermal differentiation [150] (Fig. 11.2). While a role for emerin phosphorylation in this process has not yet been identified, reduced association between lamin A/C and emerin could facilitate the redistribution of emerin to the ONM in this context. Additional nuclear lamina components may have the capacity to swap between the INM and ONM, such as short nesprin isoforms [151,152,153,154] and LUMA [155].
Like emerin, lamin A/C has also been shown to undergo force-induced phosphorylation, perhaps in response to unfolding of its immunoglobulin-like domain [22, 41]. During mitosis, lamin A is phosphorylated on Ser22, prompting disassembly of the lamin network in preparation for nuclear envelope breakdown [156]. This same modification appears to be sensitive to mechanical input: increased intracellular cytoskeletal tension, induced by growing mesenchymal stem cells on stiff substrates, decreases phosphorylation of Ser22, while reduced tension, either through plating on soft substrates or inhibition of myosin II, increases phosphorylation [22, 41] (Fig. 11.2). These changes in phosphorylation level are seen within tens of minutes, and correspond to changes in nuclear stiffness as measured by micropipette aspiration [41].
While a definitive role for forced unfolding has not yet been identified in lamin A/C function, the discovery of two mechanically-regulated epitopes of lamin A/C hints at such a mechanism [31]. The first epitope is located within the first 50 residues of the N-terminus, while the second is a conformational epitope consisting of sequences in the Ig domain and the Ig-proximal unstructured linker. These regions become inaccessible to antibodies at the basal side of nuclei in response to compressive forces, including those applied by an apical actin cap [31], an actin-rich meshwork found in close association with the nucleus in various cell types [157]. The observed differential lamin A epitope accessibility is regulated by cytoskeletal tension, and requires intact LINC complexes [31]. Further, steered molecular dynamics simulations show that the loss of the epitope can be linked to lamin A/C multimerization, and therefore reinforcement of the lamin network [31]. Several previously identified stress-sensitive phosphorylation sites [22], including Ser22, are present in the two epitopes. These results suggest that cytoskeletal tension could regulate the phosphorylation state of lamin A/C by influencing the exposure of phosphorylation sites to kinases and/or phosphatases, potentially through partial unfolding of the Ig domain and concomitant multimerization of lamins (Fig. 11.2).
This force-dependent exposure of lamin A/C epitopes may play an important role in stem cell lineage selection. Fascinatingly, human mesenchymal stem cells that are shunted down osteogenic or adipogenic lineages exhibit different levels of basal epitope exposure, with the lamin epitope more polarized in osteoblasts than in adipocytes [31]. Previous work has also identified actin cap and LINC complex-dependent polarization of lamin A/C to the apical side of nuclei, as well as the apical polarization of histone marks H3K12ac and H4K5ac, which are associated with active transcription [158]. Interestingly, the conformational epitope in lamin A/C overlaps with binding sites for DNA, histones, emerin, and SUN proteins [31]. Thus, the force-dependent differential exposure of these binding sites may influence the interaction of lamin A/C with the genome, either directly or through modulation of the activity of its binding partners. As described earlier, emerin is capable of binding to DNA, and can also interact with and activate HDAC3 [80]. Again, these responses harken back to the force-dependent modulation of adhesion morphology and function at the cell surface.
Interestingly, Ihalainen et al. showed that integrin-based connectivity to the nucleus might play a crucial role in determining the polarity of lamin A/C epitope exposure, suggesting this activity of lamins may be important for cell types that are exposed to both cell-cell and cell-ECM adhesions. Thus, the polarized, force-dependent exposure of lamin A/C epitopes may be particularly relevant for the skin, where polarized basal layer cells – which contact both the basal lamina and other keratinocytes – must differentiate in a highly regulated manner to correctly form the layers of the IFE. Further, other work has shown that the geometry of exogenous force application to cells may influence nuclear lamina-dependent regulation of gene expression by modulating the extent of chromatin stretch [159].
While mechanosensing at the nuclear interior by lamin A requires the LINC complex, likely for force transduction across the nuclear envelope, it is also possible that the LINC complex is itself mechanoresponsive. The majority of the extranuclear domains of Nesprin proteins are composed of spectrin repeats, known to undergo force-induced unfolding, which could serve multiple functions [160, 161]. As with other mechanosensitive proteins, such as talin [162], this unfolding could either abrogate or reveal cryptic interacting sites, potentially influencing signaling pathways or LINC complex oligomerization [153, 163]. Increased affinity between Nesprins under tension could allow increased force to be transmitted to the nuclear lamina and could therefore mediate changes in lamina-regulated signaling, chromatin interactions, and transcription. Alternatively, unfolding of the spectrin repeats could instead release tension on LINC complexes, preventing excessive and potentially dangerous levels of force from being exerted on the nuclear interior. Indeed, using an optical tweezers assay to apply force to LINC complexes on isolated nuclei, Balikov et al. showed that nesprin spectrin repeats may undergo unfolding in response to physiological levels of force [164]. Interestingly, the LINC complex is not only capable of adapting to the sustained application of high forces, such as during the formation of TAN lines for nuclear positioning, but also to transient (10s of milliseconds) [42, 45] or low magnitude [165] mechanical signals. This suggests that the LINC complex may be able to coordinate differential responses tuned to the magnitude and geometries of forces exerted on the nucleus, which could in turn differentially influence lamin A/C activity.
Thus far, studies of nuclear lamina mechanosensitivity have focused on isolated cells in which mechanical cues are driven exclusively by cell-ECM adhesions. Indeed, little is known about the potential role intercellular adhesions, and the balance between forces generated at cell-cell and cell-ECM adhesions, may have on the lamina, particularly in tissues. In the current model, mechanical signals originating at intercellular adhesions would be relayed to the nucleus in the same way as signals from cell-ECM adhesions, resulting in changes in lamin A/C phosphorylation and structure. However, the types of mechanical inputs studied to date represent unnaturally high levels of force or asymmetrical compressive forces (due to apical actin cap formation). In an epithelial sheet, the force distribution at intercellular adhesions reaches a homeostatic equilibrium, suggesting that these cells may respond differently to changes in tension in vivo [166].
Many of the mechanosensing functions of cells are dependent upon YAP/TAZ and/or MKL/SRF signaling [167]. YAP1 regulates epidermal stem cell proliferation and tissue expansion [168, 169] and is downstream of α-catenin, which links YAP signaling to cadherin junctions [170]. MKL/SRF signaling is an essential regulator of epidermal differentiation [171,172,173] and hair follicle morphogenesis [174]. SRF further regulates epidermal differentiation by modulating actomyosin-guided mitotic spindle orientation during asymmetric divisions of basal layer cells to promote epidermal stratification [171], as well as controlling the expression and localization of AP-1 family members, and the C/EBPα transcription factor [174]. Postnatal deletion of SRF in mice yields alterations in cell-cell and cell-ECM adhesion as well as altered differentiation and actin organization, leading to an inflammatory hyperproliferative phenotype similar to psoriasis [172].
Interestingly, YAP/TAZ and MKL/SRF are regulated by the nuclear lamina (Fig. 11.2). In MEFs, the transcription of the mechanically sensitive immediate early genes egr-1 and iex-1 in response to cyclic strain requires lamin A/C and emerin [175, 176]. Further, YAP/TAZ and MKL/SRF signaling are perturbed upon loss of lamin A/C [22, 177, 178] and a nonphosphorylatable mutant lamin A/C has been shown to increase gene expression compared to a phosphomimetic, suggesting that the structure of the lamin network can influence mechanosensitive transcription [41]. Further, emerin, through its mechanosensing capacity, is specifically required for MKL/SRF-dependent gene expression on stiff versus soft substrates, implicating it in the regulation of fibrosis and other pathologies involving tissue stiffening [179].
Thus, a model is emerging in which a mechanical stimulus can be rapidly transmitted from the cytoskeleton to the nuclear interior through LINC complexes, leading to force-induced changes in posttranslational modifications of nuclear lamina components, subsequent lamina reinforcement, and transcriptional regulation. This model is consistent with recent work showing that YAP/TAZ works with AP-1 to activate target genes via chromatin looping [180]. Phosphorylation of emerin may link nuclear lamina changes to these signaling pathways by promoting MKL/SRF signaling, potentially by promoting actin polymerization and/or organization in the cytoplasm and nucleoplasm [177, 181]. However, mutant emerin protein that cannot be phosphorylated does not influence YAP/TAZ transcription [46], suggesting that modulation of these two pathways may occur through distinct mechanisms. These changes may promote increased stiffness of the nuclear lamina through reinforcement of the lamin network [22, 41, 46]. How the nuclear lamina functions in mechanosensing and through these signaling pathways during epidermal differentiation and homeostasis will be an interesting area of future investigation.
11.7 The Nuclear Lamina and Human Disease
Disruption of the nuclear lamina leads to a host of rare diseases, known as laminopathies and nuclear envelopathies, which include muscle dystrophies, premature aging, lipodystrophies, peripheral nerve disorders, and bone diseases (reviewed in [182]). These disorders are often associated with hallmark defects in epidermal structure and function, which hints at the special role the nuclear lamina plays in epidermal homeostasis. While we are just beginning to dissect the etiology of human diseases associated with the nuclear lamina, the burgeoning field of epidermal nuclear biology highlight the importance of this domain in regulating epidermal development and integrity.
In particular, mutations in lamin A, associated with either Hutchinson-Gilford Progeria Syndrome (HGPS) or other laminopathies, have been implicated in altering interactions between LADs and the nuclear lamina. Numerous studies have shown that loss of lamin A/C, or expression of lamin A/C mutants, can result in global changes in 3D genome organization and gene expression. Further, global changes in H3K27me3 marks, dissociation of chromatin from lamin A/C and the lamina, and gene expression changes have been identified in epidermal fibroblasts from HGPS patients [183]. However, in line with the fact that lamin association does not necessarily influence transcriptional state, these changes do not always induce altered gene expression. Other work suggests that progerin expression is not sufficient to induce such vast perturbation of chromatin organization and transcription [79]. While progerin preferentially interacts with a subset of genes distinct from lamin A/C, these altered associations are not associated with global expression changes [79]. A complex array of interactions between nuclear lamina proteins and chromatin-interactors or –modifiers has been identified. These interactions may be disrupted in diseases where the nuclear lamina is altered, such as in HGPS and restrictive dermopathy [184]. Further work will be required to understand the exact changes that occur upon lamin A/C disruption, and the mechanisms by which these occur. The use of sophisticated tools to map the dynamic interactions of chromatin with the nuclear periphery over time [75], including in embryonic development, will prove an essential component in understanding the etiology of laminopathies and nuclear envelopathies.
11.8 Summary
Skin is subject to constant mechanical challenge through external insults and as a result of the cellular changes that occur during development and homeostasis. Recent data suggest that tissue homeostasis requires crosstalk between the cell surface (in the form of cell-ECM and cell-cell adhesions) and the nucleus to coordinate biochemical and mechanical cues. The LINC complex, which spans the nuclear envelope, plays a critical role in communicating mechanical information from the cytoskeleton to the nuclear interior. The nuclear lamina responds to forces transduced by the LINC complex by coordinating changes in chromatin organization and transcriptional output; the lamina also helps to maintain genome integrity, although the mechanisms at play will require further investigation. Understanding how the lamina coordinates these integrated functions will be critical to defining how lamin dysfunction contributes to defects in epidermal structure and function.
Abbreviations
- AJs:
-
Adherens junctions
- APC:
-
Adenomatous polyposis coli
- BAF:
-
Barrier-to-autointegration factor
- ChIP-seq:
-
Chromatin immunoprecipitation sequencing
- DDR:
-
DNA damage response
- DSB:
-
Double-strand break
- ECM:
-
Extracellular Matrix
- EDC:
-
Epidermal differentiation complex
- GA repeat:
-
Guanine-adenine repeat
- H2K9me2/3:
-
Di- or trimethylation of lysine 9 on histone H2
- H3K27me3:
-
Trimethylation of lysine 27 on histone H3
- HDAC:
-
Histone deacetylase
- HGPS:
-
Hutchinson-Gilford Progeria Syndrome
- Hi-C:
-
High-throughput chromosome capture
- IFE:
-
Interfollicular epidermis
- INM:
-
Inner nuclear membrane
- LEM:
-
LAP2, emerin and MAN1
- ONM:
-
Outer nuclear membrane
- LAD:
-
Lamina associated domain
- LAP1:
-
Lamina-associated polypeptide 1
- LAP2:
-
Lamina-associated polypeptide 2
- LBR:
-
Lamin B receptor
- LINC:
-
Linker of Nucleoskeleton and Cytoskeleton
- MEF:
-
Mouse embryonic fibroblast
- RGD:
-
Arginylglycylaspartic acid
- TAN lines:
-
Transmembrane actin-associated nuclear lines
References
Candi E, Schmidt R, Melino G. The cornified envelope: a model of cell death in the skin. Nat Rev Mol Cell Biol. 2005;6:328–40. https://doi.org/10.1038/nrm1619.
Blanpain C, Fuchs E. Epidermal homeostasis: a balancing act of stem cells in the skin. Nat Rev Mol Cell Biol. 2009;10:207–17. https://doi.org/10.1038/nrm2636.
Lechler T, Fuchs E. Asymmetric cell divisions promote stratification and differentiation of mammalian skin. Nature. 2005;437:275–80. https://doi.org/10.1038/nature03922.
Koster MI, Roop DR. Asymmetric cell division in skin development: a new look at an old observation. Dev Cell. 2005;9:444–6. https://doi.org/10.1016/j.devcel.2005.09.009.
Simpson CL, Patel DM, Green KJ. Deconstructing the skin: cytoarchitectural determinants of epidermal morphogenesis. Nat Rev Mol Cell Biol. 2011;12:565–80. https://doi.org/10.1038/nrm3175.
Delva E, Tucker DK, Kowalczyk AP. The desmosome. Cold Spring Harb Perspect Biol. 2009;1:a002543. https://doi.org/10.1101/cshperspect.a002543.
Furuse M, Hata M, Furuse K, Yoshida Y, Haratake A, Sugitani Y, Noda T, Kubo A, Tsukita S. Claudin-based tight junctions are crucial for the mammalian epidermal barrier: a lesson from claudin-1-deficient mice. J Cell Biol. 2002;156:1099–111. https://doi.org/10.1083/jcb.200110122.
Morita K, Itoh M, Saitou M, Ando-Akatsuka Y, Furuse M, Yoneda K, Imamura S, Fujimoto K, Tsukita S. Subcellular distribution of tight junction-associated proteins (occludin, ZO-1, ZO-2) in rodent skin. J Invest Dermatol. 1998;110:862–6. https://doi.org/10.1046/j.1523-1747.1998.00209.x.
Schlüter H, Wepf R, Moll I, Franke WW. Sealing the live part of the skin: the integrated meshwork of desmosomes, tight junctions and curvilinear ridge structures in the cells of the uppermost granular layer of the human epidermis. Eur J Cell Biol. 2004;83:655–65. https://doi.org/10.1078/0171-9335-00434.
Gdula MR, Poterlowicz K, Mardaryev AN, Sharov AA, Peng Y, Fessing MY, Botchkarev VA. Remodeling of three-dimensional organization of the nucleus during terminal keratinocyte differentiation in the epidermis. J Invest Dermatol. 2013;133:2191–201. https://doi.org/10.1038/jid.2013.66.
Burke B, Stewart CL. The nuclear lamins: flexibility in function. Nat Rev Mol Cell Biol. 2013;14:13–24. https://doi.org/10.1038/nrm3488.
Chang W, Worman HJ, Gundersen GG. Accessorizing and anchoring the LINC complex for multifunctionality. J Cell Biol. 2015;208:11–22. https://doi.org/10.1083/jcb.201409047.
Peter A, Stick R. Evolutionary aspects in intermediate filament proteins. Curr Opin Cell Biol. 2015;32:48–55. https://doi.org/10.1016/j.ceb.2014.12.009.
Heitlinger E, Peter M, Lustig A, Villiger W, Nigg EA, Aebi U. The role of the head and tail domain in lamin structure and assembly: analysis of bacterially expressed chicken lamin A and truncated B2 lamins. J Struct Biol. 1992;108:74–89.
Herrmann H, Aebi U. Intermediate filaments: molecular structure, assembly mechanism, and integration into functionally distinct intracellular Scaffolds. Annu Rev Biochem. 2004;73:749–89. https://doi.org/10.1146/annurev.biochem.73.011303.073823.
Gruenbaum Y, Medalia O. Lamins: the structure and protein complexes. Curr Opin Cell Biol. 2015;32:7–12. https://doi.org/10.1016/j.ceb.2014.09.009.
Turgay Y, Eibauer M, Goldman AE, Shimi T, Khayat M, Ben-Harush K, Dubrovsky-Gaupp A, Sapra KT, Goldman RD, Medalia O. The molecular architecture of lamins in somatic cells. Nature. 2017;543:261–4. https://doi.org/10.1038/nature21382.
Shimi T, Kittisopikul M, Tran J, Goldman AE, Adam SA, Zheng Y, Jaqaman K, Goldman RD. Structural organization of nuclear lamins A, C, B1 and B2 revealed by super-resolution microscopy. Mol Biol Cell. 2015;26(22):4075–86. https://doi.org/10.1091/mbc.E15-07-0461.
Xie W, Chojnowski A, Boudier T, Lim JSY, Ahmed S, Ser Z, Stewart C, Burke B. A-type Lamins form distinct filamentous networks with differential nuclear pore complex associations. Curr Biol. 2016;26:2651–8. https://doi.org/10.1016/j.cub.2016.07.049.
Worman HJ, Lazaridis I, Georgatos SD. Nuclear lamina heterogeneity in mammalian cells. Differential expression of the major lamins and variations in lamin B phosphorylation. J Biol Chem. 1988;263:12135–41.
Constantinescu D, Gray HL, Sammak PJ, Schatten GP, Csoka AB. Lamin A/C expression is a marker of mouse and human embryonic stem cell differentiation. Stem Cells. 2006;24:177–85. https://doi.org/10.1634/stemcells.2004-0159.
Swift J, Ivanovska IL, Buxboim A, Harada T, Dingal PCDP, Pinter J, Pajerowski JD, Spinler KR, Shin J-W, Tewari M, Rehfeldt F, Speicher DW, Discher DE. Nuclear lamin-A scales with tissue stiffness and enhances matrix-directed differentiation. Science. 2013;341:1240104. https://doi.org/10.1126/science.1240104.
Shin J-W, Spinler KR, Swift J, Chasis JA, Mohandas N, Discher DE. Lamins regulate cell trafficking and lineage maturation of adult human hematopoietic cells. Proc Natl Acad Sci USA. 2013;110:18892–7. https://doi.org/10.1073/pnas.1304996110.
Venables RS, McLean S, Luny D, Moteleb E, Morley S, Quinlan RA, Lane EB, Hutchison CJ. Expression of individual lamins in basal cell carcinomas of the skin. Br J Cancer. 2001;84:512–9. https://doi.org/10.1054/bjoc.2000.1632.
Hanif M, Rosengardten Y, Sagelius H, Rozell B, Eriksson M. Differential expression of A-type and B-type lamins during hair cycling. PLoS One. 2009;4:e4114. https://doi.org/10.1371/journal.pone.0004114.
Rusiñol AE, Sinensky MS. Farnesylated lamins, progeroid syndromes and farnesyl transferase inhibitors. J Cell Sci. 2006;119:3265–72. https://doi.org/10.1242/jcs.03156.
De Sandre-Giovannoli A, Bernard R, Cau P, Navarro C, Amiel J, Boccaccio I, Lyonnet S, Stewart CL, Munnich A, Le Merrer M, Lévy N. Lamin a truncation in Hutchinson-Gilford progeria. Science. 2003;300:2055. https://doi.org/10.1126/science.1084125.
Solovei II, Wang ASA, Thanisch KK, Schmidt CSC, Krebs SS, Zwerger MM, Cohen TVT, Devys DD, Foisner RR, Peichl LL, Herrmann HH, Blum HH, Engelkamp DD, Stewart CLC, Leonhardt HH, Joffe BB. LBR and lamin A/C sequentially tether peripheral heterochromatin and inversely regulate differentiation. Cell. 2013;152:584–98.
McKenna T, Rosengardten Y, Viceconte N, Baek J-H, Grochová D, Eriksson M. Embryonic expression of the common progeroid lamin A splice mutation arrests postnatal skin development. Aging Cell. 2014;13:292–302. https://doi.org/10.1111/acel.12173.
Jung H-J, Tatar A, Tu Y, Nobumori C, Yang SH, Goulbourne CN, Herrmann H, Fong LG, Young SG. An absence of nuclear lamins in keratinocytes leads to ichthyosis, defective epidermal barrier function, and intrusion of nuclear membranes and endoplasmic reticulum into the nuclear chromatin. Mol Cell Biol. 2014;34:4534–44. https://doi.org/10.1128/MCB.00997-14.
Ihalainen TO, Aires L, Herzog FA, Schwartlander R, Moeller J, Vogel V. Differential basal-to-apical accessibility of lamin A/C epitopes in the nuclear lamina regulated by changes in cytoskeletal tension. Nat Mater. 2015. https://doi.org/10.1038/nmat4389.
Naeem AS, Zhu Y, Di WL, Marmiroli S, O'Shaughnessy RFL. AKT1-mediated Lamin A/C degradation is required for nuclear degradation and normal epidermal terminal differentiation. Cell Death Differ. 2015. https://doi.org/10.1038/cdd.2015.62.
Wallace L, Roberts-Thompson L, Reichelt J. Deletion of K1/K10 does not impair epidermal stratification but affects desmosomal structure and nuclear integrity. J Cell Sci. 2012;125:1750–8. https://doi.org/10.1242/jcs.097139.
Barton LJ, Soshnev AA, Geyer PK. Networking in the nucleus: a spotlight on LEM-domain proteins. Curr Opin Cell Biol. 2015;34:1–8. https://doi.org/10.1016/j.ceb.2015.03.005.
Shin J-Y, Dauer WT, Worman HJ. Lamina-associated polypeptide 1: protein interactions and tissue-selective functions. Semin Cell Dev Biol. 2014;29:164–8. https://doi.org/10.1016/j.semcdb.2014.01.010.
Gesson K, Vidak S, Foisner R. Lamina-associated polypeptide (LAP)2α and nucleoplasmic lamins in adult stem cell regulation and disease. Semin Cell Dev Biol. 2014;29:116–24. https://doi.org/10.1016/j.semcdb.2013.12.009.
Olins AL, Rhodes G, Welch DBM, Zwerger M, Olins DE. Lamin B receptor: multi-tasking at the nuclear envelope. Nucleus. 2010;1:53–70. https://doi.org/10.4161/nucl.1.1.10515.
Dauer WT, Worman HJ. The nuclear envelope as a signaling node in development and disease. Dev Cell. 2009;17:626–38. https://doi.org/10.1016/j.devcel.2009.10.016.
Wang N, Tytell JD, Ingber DE. Mechanotransduction at a distance: mechanically coupling the extracellular matrix with the nucleus. Nat Rev Mol Cell Biol. 2009;10:75–82. https://doi.org/10.1038/nrm2594.
Maniotis AJ, Ingber DE, Chen CS. Demonstration of mechanical connections between integrins, cytoskeletal filaments, and nucleoplasm that stabilize nuclear structure. Proc Natl Acad Sci USA. 1997;94:849–54.
Buxboim A, Swift J, Irianto J, Spinler KR, Dingal PCDP, Athirasala A, Kao Y-RC, Cho S, Harada T, Shin J-W, Discher DE. Matrix elasticity regulates lamin-A,C phosphorylation and turnover with feedback to actomyosin. Curr Biol. 2014;24:1909–17. https://doi.org/10.1016/j.cub.2014.07.001.
Poh Y-C, Shevtsov SP, Chowdhury F, Wu DC, Na S, Dundr M, Wang N. Dynamic force-induced direct dissociation of protein complexes in a nuclear body in living cells. Nat Commun. 2012;3:866. https://doi.org/10.1038/ncomms1873.
Booth-Gauthier EA, Alcoser TA, Yang G, Dahl KN. Force-induced changes in subnuclear movement and rheology. Biophys J. 2012;103:2423–31. https://doi.org/10.1016/j.bpj.2012.10.039.
Lombardi ML, Jaalouk DE, Shanahan CM, Burke B, Roux KJ, Lammerding J. The interaction between nesprins and sun proteins at the nuclear envelope is critical for force transmission between the nucleus and cytoskeleton. J Biol Chem. 2011;286:26743–53. https://doi.org/10.1074/jbc.M111.233700.
Chambliss AB, Khatau SB, Erdenberger N, Robinson DK, Hodzic D, Longmore GD, Wirtz D. The LINC-anchored actin cap connects the extracellular milieu to the nucleus for ultrafast mechanotransduction. Sci Rep. 2013;3:1087. https://doi.org/10.1038/srep01087.
Guilluy C, Osborne LD, Van Landeghem L, Sharek L, Superfine R, Garcia-Mata R, Burridge K. Isolated nuclei adapt to force and reveal a mechanotransduction pathway in the nucleus. Nat Cell Biol. 2014;16:376–81. https://doi.org/10.1038/ncb2927.
Morgan JT, Pfeiffer ER, Thirkill TL, Kumar P, Peng G, Fridolfsson HN, Douglas GC, Starr DA, Barakat AI. Nesprin-3 regulates endothelial cell morphology, perinuclear cytoskeletal architecture, and flow-induced polarization. Mol Biol Cell. 2011;22:4324–34. https://doi.org/10.1091/mbc.E11-04-0287.
Gundersen GG, Worman HJ. Nuclear positioning. Cell. 2013;152:1376–89. https://doi.org/10.1016/j.cell.2013.02.031.
Thakar K, May CK, Rogers A, Carroll CW. Opposing roles for distinct LINC complexes in regulation of the small GTPase RhoA. Mol Biol Cell. 2017;28:182–91. https://doi.org/10.1091/mbc.E16-06-0467.
Mounkes LC, Kozlov SV, Rottman JN, Stewart CL. Expression of an LMNA-N195K variant of A-type lamins results in cardiac conduction defects and death in mice. Hum Mol Genet. 2005;14:2167–80. https://doi.org/10.1093/hmg/ddi221.
Frock RL, Chen SC, Da D-F, Frett E, Lau C, Brown C, Pak DN, Wang Y, Muchir A, Worman HJ, Santana LF, Ladiges WC, Rabinovitch PS, Kennedy BK. Cardiomyocyte-specific expression of lamin a improves cardiac function in Lmna-/- mice. PLoS One. 2012;7:e42918. https://doi.org/10.1371/journal.pone.0042918.
Hale CM, Shrestha AL, Khatau SB, Stewart-Hutchinson PJ, Hernandez L, Stewart CL, Hodzic D, Wirtz D. Dysfunctional connections between the nucleus and the actin and microtubule networks in laminopathic models. Biophys J. 2008;95:5462–75. https://doi.org/10.1529/biophysj.108.139428.
Lee JSH, Hale CM, Panorchan P, Khatau SB, George JP, Tseng Y, Stewart CL, Hodzic D, Wirtz D. Nuclear lamin A/C deficiency induces defects in cell mechanics, polarization, and migration. Biophys J. 2007;93:2542–52. https://doi.org/10.1529/biophysj.106.102426.
Kim D-H, Khatau SB, Feng Y, Walcott S, Sun SX, Longmore GD, Wirtz D. Actin cap associated focal adhesions and their distinct role in cellular mechanosensing. Sci Rep. 2012;2:555. https://doi.org/10.1038/srep00555.
Dupin I, Camand E, Etienne-Manneville S. Classical cadherins control nucleus and centrosome position and cell polarity. J Cell Biol. 2009;185:779–86. https://doi.org/10.1083/jcb.200812034.
Stewart RM, Zubek AE, Rosowski KA, Schreiner SM, Horsley V, King MC. Nuclear-cytoskeletal linkages facilitate cross talk between the nucleus and intercellular adhesions. J Cell Biol. 2015;209:403–18. https://doi.org/10.1083/jcb.201502024.
Lammerding J, Fong LG, Ji JY, Reue K, Stewart CL, Young SG, Lee RT. Lamins A and C but not lamin B1 regulate nuclear mechanics. J Biol Chem. 2006;281:25768–80. https://doi.org/10.1074/jbc.M513511200.
Trappmann B, Gautrot JE, Connelly JT, Strange DGT, Li Y, Oyen ML, Cohen Stuart MA, Boehm H, Li B, Vogel V, Spatz JP, Watt FM, Huck WTS. Extracellular-matrix tethering regulates stem-cell fate. Nat Mater. 2012;11:642–9. https://doi.org/10.1038/nmat3339.
Schreiner SM, Koo PK, Zhao Y, Mochrie SGJ, King MC. The tethering of chromatin to the nuclear envelope supports nuclear mechanics. Nat Commun. 2015;6:7159. https://doi.org/10.1038/ncomms8159.
Dahl KN, Kahn SM, Wilson KL, Discher DE. The nuclear envelope lamina network has elasticity and a compressibility limit suggestive of a molecular shock absorber. J Cell Sci. 2004;117:4779–86. https://doi.org/10.1242/jcs.01357.
Stephens AD, Banigan EJ, Adam SA, Goldman RD, Marko JF. Chromatin and lamin A determine two different mechanical response regimes of the cell nucleus. Mol Biol Cell. 2017;28:1984–96. https://doi.org/10.1091/mbc.E16-09-0653.
Furusawa T, Rochman M, Taher L, Dimitriadis EK, Nagashima K, Anderson S, Bustin M. Chromatin decompaction by the nucleosomal binding protein HMGN5 impairs nuclear sturdiness. Nat Commun. 2015;6:6138. https://doi.org/10.1038/ncomms7138.
Gerlitz G, Bustin M. Efficient cell migration requires global chromatin condensation. J Cell Sci. 2010;123:2207–17. https://doi.org/10.1242/jcs.058271.
Chubb JR, Boyle S, Perry P, Bickmore WA. Chromatin motion is constrained by association with nuclear compartments in human cells. Curr Biol. 2002;12:439–45.
Bronshtein I, Kepten E, Kanter I, Berezin S, Lindner M, Redwood AB, Mai S, Gonzalo S, Foisner R, Shav-Tal Y, Garini Y. Loss of lamin A function increases chromatin dynamics in the nuclear interior. Nat Commun. 2015;6:8044. https://doi.org/10.1038/ncomms9044.
Luxton GWG, Gomes ER, Folker ES, Vintinner E, Gundersen GG. Linear arrays of nuclear envelope proteins harness retrograde actin flow for nuclear movement. Science. 2010;329:956–9. https://doi.org/10.1126/science.1189072.
Lei K, Zhu X, Xu R, Shao C, Xu T, Zhuang Y, Han M. Inner nuclear envelope proteins SUN1 and SUN2 play a prominent role in the DNA damage response. Curr Biol. 2012;22:1609–15. https://doi.org/10.1016/j.cub.2012.06.043.
Harada T, Swift J, Irianto J, Shin J-W, Spinler KR, Athirasala A, Diegmiller R, Dingal PCDP, Ivanovska IL, Discher DE. Nuclear lamin stiffness is a barrier to 3D migration, but softness can limit survival. J Cell Biol. 2014;204:669–82. https://doi.org/10.1083/jcb.201308029.
Mattout A, Pike BL, Towbin BD, Bank EM, Gonzalez-Sandoval A, Stadler MB, Meister P, Gruenbaum Y, Gasser SM. An EDMD mutation in C. elegans lamin blocks muscle-specific gene relocation and compromises muscle integrity. Curr Biol. 2011;21:1603–14. https://doi.org/10.1016/j.cub.2011.08.030.
Towbin BD, Meister P, Pike BL, Gasser SM. Repetitive transgenes in C. elegans accumulate heterochromatic marks and are sequestered at the nuclear envelope in a copy-number- and lamin-dependent manner. Cold Spring Harb Symp Quant Biol. 2010;75:555–65. https://doi.org/10.1101/sqb.2010.75.041.
Guelen L, Pagie L, Brasset E, Meuleman W, Faza MB, Talhout W, Eussen BH, de Klein A, Wessels L, de Laat W, van Steensel B. Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature. 2008;453:948–51. https://doi.org/10.1038/nature06947.
Harr JC, Luperchio TR, Wong X, Cohen E, Wheelan SJ, Reddy KL. Directed targeting of chromatin to the nuclear lamina is mediated by chromatin state and A-type lamins. J Cell Biol. 2015;208:33–52. https://doi.org/10.1083/jcb.201405110.
Zullo JM, Demarco IA, Piqué-Regi R, Gaffney DJ, Epstein CB, Spooner CJ, Luperchio TR, Bernstein BE, Pritchard JK, Reddy KL, Singh H. DNA sequence-dependent compartmentalization and silencing of chromatin at the nuclear lamina. Cell. 2012;149:1474–87. https://doi.org/10.1016/j.cell.2012.04.035.
Towbin BD, González-Aguilera C, Sack R, Gaidatzis D, Kalck V, Meister P, Askjaer P, Gasser SM. Step-wise methylation of histone H3K9 positions heterochromatin at the nuclear periphery. Cell. 2012;150:934–47. https://doi.org/10.1016/j.cell.2012.06.051.
Kind J, Pagie L, Ortabozkoyun H, Boyle S, de Vries SS, Janssen H, Amendola M, Nolen LD, Bickmore WA, van Steensel B. Single-cell dynamics of genome-nuclear lamina interactions. Cell. 2013;153:178–92. https://doi.org/10.1016/j.cell.2013.02.028.
Kumaran RI, Spector DL. A genetic locus targeted to the nuclear periphery in living cells maintains its transcriptional competence. J Cell Biol. 2008;180:51–65. https://doi.org/10.1083/jcb.200706060.
Reddy KL, Zullo JM, Bertolino E, Singh H. Transcriptional repression mediated by repositioning of genes to the nuclear lamina. Nature. 2008;452:243–7. https://doi.org/10.1038/nature06727.
Van de Vosse DW, Wan Y, Wozniak RW, Aitchison JD. Role of the nuclear envelope in genome organization and gene expression. Wiley Interdiscip Rev Syst Biol Med. 2011;3:147–66. https://doi.org/10.1002/wsbm.101.
Kubben N, Adriaens M, Meuleman W, Voncken JW, van Steensel B, Misteli T. Mapping of lamin A- and progerin-interacting genome regions. Chromosoma. 2012;121:447–64. https://doi.org/10.1007/s00412-012-0376-7.
Demmerle J, Koch AJ, Holaska JM. The nuclear envelope protein emerin binds directly to histone deacetylase 3 (HDAC3) and activates HDAC3 activity. J Biol Chem. 2012;287:22080–8. https://doi.org/10.1074/jbc.M111.325308.
Nagano T, Lubling Y, Stevens TJ, Schoenfelder S, Yaffe E, Dean W, Laue ED, Tanay A, Fraser P. Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature. 2013;502:59–64. https://doi.org/10.1038/nature12593.
Andrés V, González JM. Role of A-type lamins in signaling, transcription, and chromatin organization. J Cell Biol. 2009;187:945–57. https://doi.org/10.1083/jcb.200904124.
Kypriotou M, Huber M, Hohl D. The human epidermal differentiation complex: cornified envelope precursors, S100 proteins and the “fused genes” family. Exp Dermatol. 2012;21:643–9. https://doi.org/10.1111/j.1600-0625.2012.01472.x.
Williams RRE, Broad S, Sheer D, Ragoussis J. Subchromosomal positioning of the epidermal differentiation complex (EDC) in keratinocyte and lymphoblast interphase nuclei. Exp Cell Res. 2002;272:163–75. https://doi.org/10.1006/excr.2001.5400.
Mardaryev AN, Gdula MR, Yarker JL, Emelianov VU, Emelianov VN, Poterlowicz K, Sharov AA, Sharova TY, Scarpa JA, Joffe B, Solovei I, Chambon P, Botchkarev VA, Fessing MY. p63 and Brg1 control developmentally regulated higher-order chromatin remodelling at the epidermal differentiation complex locus in epidermal progenitor cells. Development. 2014;141:101–11. https://doi.org/10.1242/dev.103200.
Fessing MY, Mardaryev AN, Gdula MR, Sharov AA, Sharova TY, Rapisarda V, Gordon KB, Smorodchenko AD, Poterlowicz K, Ferone G, Kohwi Y, Missero C, Kohwi-Shigematsu T, Botchkarev VA. p63 regulates Satb1 to control tissue-specific chromatin remodeling during development of the epidermis. J Cell Biol. 2011;194:825–39. https://doi.org/10.1083/jcb.201101148.
Lien W-H, Guo X, Polak L, Lawton LN, Young RA, Zheng D, Fuchs E. Genome-wide maps of histone modifications unwind in vivo chromatin states of the hair follicle lineage. Cell Stem Cell. 2011;9:219–32. https://doi.org/10.1016/j.stem.2011.07.015.
Indra AK, Dupé V, Bornert J-M, Messaddeq N, Yaniv M, Mark M, Chambon P, Metzger D. Temporally controlled targeted somatic mutagenesis in embryonic surface ectoderm and fetal epidermal keratinocytes unveils two distinct developmental functions of BRG1 in limb morphogenesis and skin barrier formation. Development. 2005;132:4533–44. https://doi.org/10.1242/dev.02019.
Kashiwagi M, Morgan BA, Georgopoulos K. The chromatin remodeler Mi-2beta is required for establishment of the basal epidermis and normal differentiation of its progeny. Development. 2007;134:1571–82. https://doi.org/10.1242/dev.001750.
LeBoeuf M, Terrell A, Trivedi S, Sinha S, Epstein JA, Olson EN, Morrisey EE, Millar SE. Hdac1 and Hdac2 act redundantly to control p63 and p53 functions in epidermal progenitor cells. Dev Cell. 2010;19:807–18. https://doi.org/10.1016/j.devcel.2010.10.015.
Frye M, Fisher AG, Watt FM. Epidermal stem cells are defined by global histone modifications that are altered by Myc-induced differentiation. PLoS One. 2007;2:e763. https://doi.org/10.1371/journal.pone.0000763.
Hughes MW, Jiang T-X, Lin S-J, Leung Y, Kobielak K, Widelitz RB, Chuong CM. Disrupted ectodermal organ morphogenesis in mice with a conditional histone deacetylase 1, 2 deletion in the epidermis. J Invest Dermatol. 2014;134:24–32. https://doi.org/10.1038/jid.2013.283.
Rapisarda V, Malashchuk I, Asamaowei IE, Poterlowicz K, Fessing MY, Sharov AA, Karakesisoglou I, Botchkarev VA, Mardaryev A. p63 transcription factor regulates nuclear shape and expression of nuclear envelope-associated genes in epidermal keratinocytes. J Invest Dermatol. 2017. https://doi.org/10.1016/j.jid.2017.05.013.
Ivorra C, Kubicek M, González JM, Sanz-González SM, Alvarez-Barrientos A, O'Connor J-E, Burke B, Andrés V. A mechanism of AP-1 suppression through interaction of c-Fos with lamin A/C. Genes Dev. 2006;20:307–20. https://doi.org/10.1101/gad.349506.
González JM, Navarro-Puche A, Casar B, Crespo P, Andrés V. Fast regulation of AP-1 activity through interaction of lamin A/C, ERK1/2, and c-Fos at the nuclear envelope. J Cell Biol. 2008;183:653–66. https://doi.org/10.1083/jcb.200805049.
Oh IY, Albea DM, Goodwin ZA, Quiggle AM, Baker BP, Guggisberg AM, Geahlen JH, Kroner GM, de Guzman Strong C. Regulation of the dynamic chromatin architecture of the epidermal differentiation complex is mediated by a c-Jun/AP-1-modulated enhancer. J Invest Dermatol. 2014;134:2371–80. https://doi.org/10.1038/jid.2014.44.
Ezhkova E, Pasolli HA, Parker JS, Stokes N, Su I-H, Hannon G, Tarakhovsky A, Fuchs E. Ezh2 orchestrates gene expression for the stepwise differentiation of tissue-specific stem cells. Cell. 2009;136:1122–35. https://doi.org/10.1016/j.cell.2008.12.043.
Blanpain C, Fuchs E. Epidermal stem cells of the skin. Annu Rev Cell Dev Biol. 2006;22:339–73. https://doi.org/10.1146/annurev.cellbio.22.010305.104357.
Markiewicz E, Tilgner K, Barker N, van de Wetering M, Clevers H, Dorobek M, Hausmanowa-Petrusewicz I, Ramaekers FCS, Broers JLV, Blankesteijn WM, Salpingidou G, Wilson RG, Ellis JA, Hutchison CJ. The inner nuclear membrane protein emerin regulates beta-catenin activity by restricting its accumulation in the nucleus. EMBO J. 2006;25:3275–85. https://doi.org/10.1038/sj.emboj.7601230.
Tilgner K, Wojciechowicz K, Jahoda C, Hutchison C, Markiewicz E. Dynamic complexes of A-type lamins and emerin influence adipogenic capacity of the cell via nucleocytoplasmic distribution of beta-catenin. J Cell Sci. 2009;122:401–13. https://doi.org/10.1242/jcs.026179.
Stubenvoll A, Rice M, Wietelmann A, Wheeler M, Braun T. Attenuation of Wnt/β-catenin activity reverses enhanced generation of cardiomyocytes and cardiac defects caused by the loss of emerin. Hum Mol Genet. 2015;24:802–13. https://doi.org/10.1093/hmg/ddu498.
Neumann S, Schneider M, Daugherty RL, Gottardi CJ, Eming SA, Beijer A, Noegel AA, Karakesisoglou I. Nesprin-2 interacts with {alpha}-catenin and regulates Wnt signaling at the nuclear envelope. J Biol Chem. 2010;285:34932–8. https://doi.org/10.1074/jbc.M110.119651.
Lim X, Nusse R. Wnt signaling in skin development, homeostasis, and disease. Cold Spring Harb Perspect Biol. 2013;5:a008029. https://doi.org/10.1101/cshperspect.a008029.
Lin F, Morrison JM, Wu W, Worman HJ. MAN1, an integral protein of the inner nuclear membrane, binds Smad2 and Smad3 and antagonizes transforming growth factor-beta signaling. Hum Mol Genet. 2005;14:437–45. https://doi.org/10.1093/hmg/ddi040.
Bengtsson L. What MAN1 does to the Smads. TGFbeta/BMP signaling and the nuclear envelope. FEBS J. 2007;274:1374–82.
Hellemans J, Preobrazhenska O, Willaert A, Debeer P, Verdonk PCM, Costa T, Janssens K, Menten B, Van Roy N, Vermeulen SJT, Savarirayan R, Van Hul W, Vanhoenacker F, Huylebroeck D, De Paepe A, Naeyaert J-M, Vandesompele J, Speleman F, Verschueren K, Coucke PJ, Mortier GR. Loss-of-function mutations in LEMD3 result in osteopoikilosis, Buschke-Ollendorff syndrome and melorheostosis. Nat Genet. 2004. Published online: 10 December 2001; doi:101038/ng789 36:1213–18. doi: https://doi.org/10.1038/ng1453.
Asano Y, Ihn H, Yamane K, Kubo M, Tamaki K. Impaired Smad7-Smurf-mediated negative regulation of TGF-beta signaling in scleroderma fibroblasts. J Clin Invest. 2004;113:253–64. https://doi.org/10.1172/JCI16269.
Mori Y, Chen S-J, Varga J. Expression and regulation of intracellular SMAD signaling in scleroderma skin fibroblasts. Arthritis Rheum. 2003;48:1964–78. https://doi.org/10.1002/art.11157.
Rashmi RN, Eckes B, Glöckner G, Groth M, Neumann S, Gloy J, Sellin L, Walz G, Schneider M, Karakesisoglou I, Eichinger L, Noegel AA. The nuclear envelope protein Nesprin-2 has roles in cell proliferation and differentiation during wound healing. Nucleus. 2012;3:172–86. https://doi.org/10.4161/nucl.19090.
Mayr M, Hu Y, Hainaut H, Xu Q. Mechanical stress-induced DNA damage and rac-p38MAPK signal pathways mediate p53-dependent apoptosis in vascular smooth muscle cells. FASEB J. 2002;16:1423–5. https://doi.org/10.1096/fj.02-0042fje.
Lawrence KS, Tapley EC, Cruz VE, Li Q, Aung K, Hart KC, Schwartz TU, Starr DA, Engebrecht J. LINC complexes promote homologous recombination in part through inhibition of nonhomologous end joining. J Cell Biol. 2016;215:801–21. https://doi.org/10.1083/jcb.201604112.
Sur I, Neumann S, Noegel AA. Nesprin-1 role in DNA damage response. Nucleus. 2014;5:173–91. https://doi.org/10.4161/nucl.29023.
Warren DT, Tajsic T, Porter LJ, Minaisah RM, Cobb A, Jacob A, Rajgor D, Zhang QP, Shanahan CM. Nesprin-2-dependent ERK1/2 compartmentalisation regulates the DNA damage response in vascular smooth muscle cell ageing. Cell Death Differ. 2015;22:1540–50. https://doi.org/10.1038/cdd.2015.12.
Oza P, Jaspersen SL, Miele A, Dekker J, Peterson CL. Mechanisms that regulate localization of a DNA double-strand break to the nuclear periphery. Genes Dev. 2009;23:912–27. https://doi.org/10.1101/gad.1782209.
Kalocsay M, Hiller NJ, Jentsch S. Chromosome-wide Rad51 spreading and SUMO-H2A.Z-dependent chromosome fixation in response to a persistent DNA double-strand break. Mol Cell. 2009;33:335–43. https://doi.org/10.1016/j.molcel.2009.01.016.
Swartz RK, Rodriguez EC, King MC. A role for nuclear envelope-bridging complexes in homology-directed repair. Mol Biol Cell. 2014;25:2461–71. https://doi.org/10.1091/mbc.E13-10-0569.
Kubben N, Voncken JW, Demmers J, Calis C, van Almen G, Pinto Y, Misteli T. Identification of differential protein interactors of lamin A and progerin. Nucleus. 2010;1:513–25. https://doi.org/10.4161/nucl.1.6.13512.
Gonzalez-Suarez I, Gonzalo S. Nurturing the genome: A-type lamins preserve genomic stability. Nucleus. 2010;1:129–35. https://doi.org/10.4161/nucl.1.2.10797.
Gonzalo S. DNA damage and lamins. Adv Exp Med Biol. 2014;773:377–99. https://doi.org/10.1007/978-1-4899-8032-8_17.
Manju K, Muralikrishna B, Parnaik VK. Expression of disease-causing lamin A mutants impairs the formation of DNA repair foci. J Cell Sci. 2006;119:2704–14. https://doi.org/10.1242/jcs.03009.
Liu B, Wang J, Chan KM, Tjia WM, Deng W, Guan X, Huang J-D, Li KM, Chau PY, Chen DJ, Pei D, Pendas AM, Cadiñanos J, López-Otín C, Tse HF, Hutchison C, Chen J, Cao Y, Cheah KSE, Tryggvason K, Zhou Z. Genomic instability in laminopathy-based premature aging. Nat Med. 2005;11:780–5. https://doi.org/10.1038/nm1266.
Mahen R, Hattori H, Lee M, Sharma P, Jeyasekharan AD, Venkitaraman AR. A-type lamins maintain the positional stability of DNA damage repair foci in mammalian nuclei. PLoS One. 2013;8:e61893. https://doi.org/10.1371/journal.pone.0061893.
Starke S, Meinke P, Camozzi D, Mattioli E, Pfaeffle R, Siekmeyer M, Hirsch W, Horn LC, Paasch U, Mitter D, Lattanzi G, Wehnert M, Kiess W. Progeroid laminopathy with restrictive dermopathy-like features caused by an isodisomic LMNA mutation p.R435C. Aging (Albany NY). 2013;5:445–59.
Chen C-Y, Chi Y-H, Mutalif RA, Starost MF, Myers TG, Anderson SA, Stewart CL, Jeang K-T. Accumulation of the inner nuclear envelope protein Sun1 is pathogenic in progeric and dystrophic laminopathies. Cell. 2012;149:565–77. https://doi.org/10.1016/j.cell.2012.01.059.
Choi JC, Worman HJ. Reactivation of autophagy ameliorates LMNA cardiomyopathy. Autophagy. 2013;9:110–1. https://doi.org/10.4161/auto.22403.
Choi JC, Muchir A, Wu W, Iwata S, Homma S, Morrow JP, Worman HJ. Temsirolimus activates autophagy and ameliorates cardiomyopathy caused by lamin A/C gene mutation. Sci Transl Med. 2012;4:144ra102. https://doi.org/10.1126/scitranslmed.3003875.
Lenain C, Gusyatiner O, Douma S, van den Broek B, Peeper DS. Autophagy-mediated degradation of nuclear envelope proteins during oncogene-induced senescence. Carcinogenesis. 2015;36:1263–74. https://doi.org/10.1093/carcin/bgv124.
Dou Z, Xu C, Donahue G, Shimi T, Pan J-A, Zhu J, Ivanov A, Capell BC, Drake AM, Shah PP, Catanzaro JM, Ricketts MD, Lamark T, Adam SA, Marmorstein R, Zong W-X, Johansen T, Goldman RD, Adams PD, Berger SL. Autophagy mediates degradation of nuclear lamina. Nature. 2015;527:105–9. https://doi.org/10.1038/nature15548.
Song X, Narzt MS, Nagelreiter IM, Hohensinner P, Terlecki-Zaniewicz L, Tschachler E, Grillari J, Gruber F. Autophagy deficient keratinocytes display increased DNA damage, senescence and aberrant lipid composition after oxidative stress in vitro and in vivo. Redox Biol. 2017;11:219–30. https://doi.org/10.1016/j.redox.2016.12.015.
Akinduro O, Sully K, Patel A, Robinson DJ, Chikh A, McPhail G, Braun KM, Philpott MP, Harwood CA, Byrne C, O'Shaughnessy RFL, Bergamaschi D. Constitutive autophagy and Nucleophagy during epidermal differentiation. J Invest Dermatol. 2016;136:1460–70. https://doi.org/10.1016/j.jid.2016.03.016.
Musich PR, Zou Y. DNA-damage accumulation and replicative arrest in Hutchinson-Gilford progeria syndrome. Biochem Soc Trans. 2011;39:1764–9. https://doi.org/10.1042/BST20110687.
Sotiropoulou PA, Blanpain C. Development and homeostasis of the skin epidermis. Cold Spring Harb Perspect Biol. 2012;4:a008383. https://doi.org/10.1101/cshperspect.a008383.
Goldstein J, Horsley V. Home sweet home: skin stem cell niches. CMLS, Cell Mol Life Sci. 2012;69:2573–82. https://doi.org/10.1007/s00018-012-0943-3.
Sotiropoulou PA, Candi A, Mascré G, De Clercq S, Youssef KK, Lapouge G, Dahl E, Semeraro C, Denecker G, Marine J-C, Blanpain C. Bcl-2 and accelerated DNA repair mediates resistance of hair follicle bulge stem cells to DNA-damage-induced cell death. Nat Cell Biol. 2010;12:572–82. https://doi.org/10.1038/ncb2059.
Huang X, Pan Y, Cao D, Fang S, Huang K, Chen J, Chen A. UVA-induced upregulation of progerin suppresses 53BP1-mediated NHEJ DSB repair in human keratinocytes via progerin-lamin A complex formation. Oncol Rep. 2017;37:3617–24. https://doi.org/10.3892/or.2017.5603.
Sotiropoulou PA, Karambelas AE, Debaugnies M, Candi A, Bouwman P, Moers V, Revenco T, Rocha AS, Sekiguchi K, Jonkers J, Blanpain C. BRCA1 deficiency in skin epidermis leads to selective loss of hair follicle stem cells and their progeny. Genes Dev. 2013;27:39–51. https://doi.org/10.1101/gad.206573.112.
Vogel V. Mechanotransduction involving multimodular proteins: converting force into biochemical signals. Annu Rev Biophys Biomol Struct. 2006;35:459–88. https://doi.org/10.1146/annurev.biophys.35.040405.102013.
Ivanovska IL, Shin J-W, Swift J, Discher DE. Stem cell mechanobiology: diverse lessons from bone marrow. Trends Cell Biol. 2015;25:523–32. https://doi.org/10.1016/j.tcb.2015.04.003.
Jain N, Iyer KV, Kumar A, Shivashankar GV. Cell geometric constraints induce modular gene-expression patterns via redistribution of HDAC3 regulated by actomyosin contractility. Proc Natl Acad Sci USA. 2013;110:11349–54. https://doi.org/10.1073/pnas.1300801110.
Ramdas NM, Shivashankar GV. Cytoskeletal control of nuclear morphology and chromatin organization. J Mol Biol. 2015;427:695–706. https://doi.org/10.1016/j.jmb.2014.09.008.
Versaevel M, Braquenier J-B, Riaz M, Grevesse T, Lantoine J, Gabriele S. Super-resolution microscopy reveals LINC complex recruitment at nuclear indentation sites. Sci Rep. 2014;4:7362. https://doi.org/10.1038/srep07362.
Alam SG, Zhang Q, Prasad N, Li Y, Chamala S, Kuchibhotla R, Kc B, Aggarwal V, Shrestha S, Jones AL, Levy SE, Roux KJ, Nickerson JA, Lele TP. The mammalian LINC complex regulates genome transcriptional responses to substrate rigidity. Sci Rep. 2016;6:38063. https://doi.org/10.1038/srep38063.
McBeath R, Pirone DM, Nelson CM, Bhadriraju K, Chen CS. Cell shape, cytoskeletal tension, and RhoA regulate stem cell lineage commitment. Dev Cell. 2004;6:483–95. https://doi.org/10.1038/nrm1413.
Engler AJ, Sen S, Sweeney HL, Discher DE. Matrix elasticity directs stem cell lineage specification. Cell. 2006;126:677–89. https://doi.org/10.1016/j.cell.2006.06.044.
le Duc Q, Shi Q, Blonk I, Sonnenberg A, Wang N, Leckband D, de Rooij J. Vinculin potentiates E-cadherin mechanosensing and is recruited to actin-anchored sites within adherens junctions in a myosin II-dependent manner. J Cell Biol. 2010;189:1107–15. https://doi.org/10.1083/jcb.201001149.
Yonemura S, Wada Y, Watanabe T, Nagafuchi A, Shibata M. Alpha-catenin as a tension transducer that induces adherens junction development. Nat Cell Biol. 2010;12:533–42. https://doi.org/10.1038/ncb2055.
Galbraith CG, Yamada KM, Sheetz MP. The relationship between force and focal complex development. J Cell Biol. 2002;159:695–705. https://doi.org/10.1083/jcb.200204153.
Deguchi S, Maeda K, Ohashi T, Sato M. Flow-induced hardening of endothelial nucleus as an intracellular stress-bearing organelle. J Biomech. 2005;38:1751–9. https://doi.org/10.1016/j.jbiomech.2005.06.003.
Philip JT, Dahl KN. Nuclear mechanotransduction: response of the lamina to extracellular stress with implications in aging. J Biomech. 2008;41:3164–70. https://doi.org/10.1016/j.jbiomech.2008.08.024.
Le HQ, Ghatak S, Yeung C-YC, Tellkamp F, Günschmann C, Dieterich C, Yeroslaviz A, Habermann B, Pombo A, Niessen CM, Wickström SA. Mechanical regulation of transcription controls Polycomb-mediated gene silencing during lineage commitment. Nat Cell Biol. 2016;18:864–75. https://doi.org/10.1038/ncb3387.
Morris GE, Randles KN. Nesprin isoforms: are they inside or outside the nucleus? Biochem Soc Trans. 2010;38:278–80. https://doi.org/10.1042/BST0380278.
Wheeler MA, Davies JD, Zhang Q, Emerson LJ, Hunt J, Shanahan CM, Ellis JA. Distinct functional domains in nesprin-1alpha and nesprin-2beta bind directly to emerin and both interactions are disrupted in X-linked Emery-Dreifuss muscular dystrophy. Exp Cell Res. 2007;313:2845–57. https://doi.org/10.1016/j.yexcr.2007.03.025.
Mislow JMK, Holaska JM, Kim MS, Lee KK, Segura-Totten M, Wilson KL, McNally EM. Nesprin-1alpha self-associates and binds directly to emerin and lamin A in vitro. FEBS Lett. 2002;525:135–40.
Haque F, Lloyd DJ, Smallwood DT, Dent CL, Shanahan CM, Fry AM, Trembath RC, Shackleton S. SUN1 interacts with nuclear lamin A and cytoplasmic nesprins to provide a physical connection between the nuclear lamina and the cytoskeleton. Mol Cell Biol. 2006;26:3738–51. https://doi.org/10.1128/MCB.26.10.3738-3751.2006.
Bengtsson L, Otto H. LUMA interacts with emerin and influences its distribution at the inner nuclear membrane. J Cell Sci. 2008;121:536–48. https://doi.org/10.1242/jcs.019281.
Heald R, McKeon F. Mutations of phosphorylation sites in lamin A that prevent nuclear lamina disassembly in mitosis. Cell. 1990;61:579–89.
Khatau SB, Kim D-H, Hale CM, Bloom RJ, Wirtz D. The perinuclear actin cap in health and disease. Nucleus. 2010;1:337–42. https://doi.org/10.4161/nucl.1.4.12331.
Kim D-H, Wirtz D. Cytoskeletal tension induces the polarized architecture of the nucleus. Biomaterials. 2015;48:161–72. https://doi.org/10.1016/j.biomaterials.2015.01.023.
Tajik A, Zhang Y, Wei F, Sun J, Jia Q, Zhou W, Singh R, Khanna N, Belmont AS, Wang N. Transcription upregulation via force-induced direct stretching of chromatin. Nat Mater. 2016;15:1287–96. https://doi.org/10.1038/nmat4729.
Autore F, Pfuhl M, Quan X, Williams A, Roberts RG, Shanahan CM, Fraternali F. Large-scale modelling of the divergent spectrin repeats in nesprins: giant modular proteins. PLoS One. 2013;8:e63633. https://doi.org/10.1371/journal.pone.0063633.
Johnson CP, Tang H-Y, Carag C, Speicher DW, Discher DE. Forced unfolding of proteins within cells. Science. 2007;317:663–6. https://doi.org/10.1126/science.1139857.
Fillingham I, Gingras AR, Papagrigoriou E, Patel B, Emsley J, Critchley DR, Roberts GCK, Barsukov IL. A vinculin binding domain from the talin rod unfolds to form a complex with the vinculin head. Structure. 2005;13:65–74. https://doi.org/10.1016/j.str.2004.11.006.
Lu W, Schneider M, Neumann S, Jaeger V-M, Taranum S, Munck M, Cartwright S, Richardson C, Carthew J, Noh K, Goldberg M, Noegel AA, Karakesisoglou I. Nesprin interchain associations control nuclear size. CMLS, Cell Mol Life Sci. 2012;69:3493–509. https://doi.org/10.1007/s00018-012-1034-1.
Balikov DA, Brady SK, Ko UH, Shin JH, de Pereda JM, Sonnenberg A, Sung H-J, Lang MJ. The nesprin-cytoskeleton interface probed directly on single nuclei is a mechanically rich system. Nucleus. 2017;5:1–14. https://doi.org/10.1080/19491034.2017.1322237.
Uzer G, Thompson WR, Sen B, Xie Z, Yen SS, Miller S, Bas G, Styner M, Rubin CT, Judex S, Burridge K, Rubin J. Cell Mechanosensitivity to extremely low-magnitude signals is enabled by a LINCed nucleus. Stem Cells. 2015;33:2063–76. https://doi.org/10.1002/stem.2004.
Tambe DT, Hardin CC, Angelini TE, Rajendran K, Park CY, Serra-Picamal X, Zhou EH, Zaman MH, Butler JP, Weitz DA, Fredberg JJ, Trepat X. Collective cell guidance by cooperative intercellular forces. Nat Mater. 2011;10:469–75. https://doi.org/10.1038/nmat3025.
Dupont S, Morsut L, Aragona M, Enzo E, Giulitti S, Cordenonsi M, Zanconato F, Le Digabel J, Forcato M, Bicciato S, Elvassore N, Piccolo S. Role of YAP/TAZ in mechanotransduction. Nature. 2011;474:179–83. https://doi.org/10.1038/nature10137.
Zhang H, Pasolli HA, Fuchs E. Yes-associated protein (YAP) transcriptional coactivator functions in balancing growth and differentiation in skin. Proc Natl Acad Sci USA. 2011;108:2270–5. https://doi.org/10.1073/pnas.1019603108.
Schlegelmilch K, Mohseni M, Kirak O, Pruszak J, Rodriguez JR, Zhou D, Kreger BT, Vasioukhin V, Avruch J, Brummelkamp TR, Camargo FD. Yap1 acts downstream of α-catenin to control epidermal proliferation. Cell. 2011;144:782–95. https://doi.org/10.1016/j.cell.2011.02.031.
Silvis MR, Kreger BT, Lien W-H, Klezovitch O, Rudakova GM, Camargo FD, Lantz DM, Seykora JT, Vasioukhin V. α-catenin is a tumor suppressor that controls cell accumulation by regulating the localization and activity of the transcriptional coactivator Yap1. Sci Signal. 2011;4:ra33. https://doi.org/10.1126/scisignal.2001823.
Luxenburg C, Pasolli HA, Williams SE, Fuchs E. Developmental roles for Srf, cortical cytoskeleton and cell shape in epidermal spindle orientation. Nat Cell Biol. 2011;13:203–14. https://doi.org/10.1038/ncb2163.
Koegel H, Tobel von L, Schäfer M, Alberti S, Kremmer E, Mauch C, Hohl D, Wang X-J, Beer H-D, Bloch W, Nordheim A, Werner S. Loss of serum response factor in keratinocytes results in hyperproliferative skin disease in mice. J Clin Invest. 2009;119:899–910. https://doi.org/10.1172/JCI37771.
Connelly JT, Gautrot JE, Trappmann B, Tan DW-M, Donati G, Huck WTS, Watt FM. Actin and serum response factor transduce physical cues from the microenvironment to regulate epidermal stem cell fate decisions. Nat Cell Biol. 2010;12:711–8. https://doi.org/10.1038/ncb2074.
Lin C, Hindes A, Burns CJ, Koppel AC, Kiss A, Yin Y, Ma L, Blumenberg M, Khnykin D, Jahnsen FL, Crosby SD, Ramanan N, Efimova T. Serum response factor controls transcriptional network regulating epidermal function and hair follicle morphogenesis. J Invest Dermatol. 2013;133:608–17. https://doi.org/10.1038/jid.2012.378.
Lammerding J, Schulze PC, Takahashi T, Kozlov S, Sullivan T, Kamm RD, Stewart CL, Lee RT. Lamin A/C deficiency causes defective nuclear mechanics and mechanotransduction. J Clin Invest. 2004;113:370–8. https://doi.org/10.1172/JCI19670.
Lammerding J, Hsiao J, Schulze PC, Kozlov S, Stewart CL, Lee RT. Abnormal nuclear shape and impaired mechanotransduction in emerin-deficient cells. J Cell Biol. 2005;170:781–91. https://doi.org/10.1083/jcb.200502148.
Ho CY, Jaalouk DE, Vartiainen MK, Lammerding J. Lamin A/C and emerin regulate MKL1-SRF activity by modulating actin dynamics. Nature. 2013;497:507–11. https://doi.org/10.1038/nature12105.
Bertrand AT, Ziaei S, Ehret C, Duchemin H, Mamchaoui K, Bigot A, Mayer M, Quijano-Roy S, Desguerre I, Lainé J, Ben Yaou R, Bonne G, Coirault C. Cellular microenvironments reveal defective mechanosensing responses and elevated YAP signaling in LMNA-mutated muscle precursors. J Cell Sci. 2014;127:2873–84. https://doi.org/10.1242/jcs.144907.
Willer MK, Carroll CW. Substrate stiffness-dependent regulation of the SRF-Mkl1 co-activator complex requires the inner nuclear membrane protein Emerin. J Cell Sci. 2017;130:2111–8. https://doi.org/10.1242/jcs.197517.
Zanconato F, Forcato M, Battilana G, Azzolin L, Quaranta E, Bodega B, Rosato A, Bicciato S, Cordenonsi M, Piccolo S. Genome-wide association between YAP/TAZ/TEAD and AP-1 at enhancers drives oncogenic growth. Nat Cell Biol. 2015;17:1218–27. https://doi.org/10.1038/ncb3216.
Holaska JM, Kowalski AK, Wilson KL. Emerin caps the pointed end of actin filaments: evidence for an actin cortical network at the nuclear inner membrane. PLoS Biol. 2004;2:E231. https://doi.org/10.1371/journal.pbio.0020231.
Schreiber KH, Kennedy BK. When lamins go bad: nuclear structure and disease. Cell. 2013;152:1365–75. https://doi.org/10.1016/j.cell.2013.02.015.
McCord RP, Nazario-Toole A, Zhang H, Chines PS, Zhan Y, Erdos MR, Collins FS, Dekker J, Cao K. Correlated alterations in genome organization, histone methylation, and DNA-lamin A/C interactions in Hutchinson-Gilford progeria syndrome. Genome Res. 2013;23:260–9. https://doi.org/10.1101/gr.138032.112.
Gonzalo S, Kreienkamp R. DNA repair defects and genome instability in Hutchinson-Gilford progeria syndrome. Curr Opin Cell Biol. 2015;34:75–83. https://doi.org/10.1016/j.ceb.2015.05.007.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG, part of Springer Nature
About this chapter
Cite this chapter
Stewart, R.M., King, M.C., Horsley, V. (2018). Integration of Biochemical and Mechanical Signals at the Nuclear Periphery: Impacts on Skin Development and Disease. In: Botchkarev, V., Millar, S. (eds) Epigenetic Regulation of Skin Development and Regeneration. Stem Cell Biology and Regenerative Medicine. Humana Press, Cham. https://doi.org/10.1007/978-3-319-16769-5_11
Download citation
DOI: https://doi.org/10.1007/978-3-319-16769-5_11
Published:
Publisher Name: Humana Press, Cham
Print ISBN: 978-3-319-16768-8
Online ISBN: 978-3-319-16769-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)