Abstract
A large variation in drug response exists between patients, with susceptible individuals being at risk of experiencing an adverse drug reaction (ADR). This susceptibility is attributable to environmental, clinical and genetic factors although the contribution of each varies with the drug, ADR and ethnicity. The variation in drug response makes personalisation of pharmacological therapy appealing to minimise ADRs whilst promoting efficacy. Pharmacogenetics seeks to contribute through genetic-guided drug and dose selection strategies. ADR pharmacogenetics was first highlighted in the 1950s, but it is only in the last decade that it has seen a rapid expansion, aided by significant advances in our knowledge of the human genome and improved genotyping technologies. ADRs can be classified according to whether the dominant mechanism is immune- or nonimmune-mediated. Several ADRs have been strongly associated with specific human leukocyte antigen (HLA) alleles. There is growing evidence for a central role of these alleles in the pathogenesis of immune-mediated delayed hypersensitivity ADRs through facilitation of ‘off-target’ interactions that lead to the presentation of ‘altered self,’ drugs and/or their metabolites to the T-cell receptor in an HLA-restricted fashion. Genetic variation can also predispose to nonimmune-mediated ADRs through perturbing drug pharmacokinetics or by altering nonimmune pharmacodynamic processes. In particular, genetic variants of phase I and phase II biotransformation enzymes and drug transporters alter the availability of a drug at the site(s) responsible for the ADR. Depending on the drug and ADR, these sites may be the therapeutic target site, the same molecular site in another tissue or distinct off-target sites. A prominent example of pharmacogenetics improving drug safety and enhancing the cost-effective use of limited healthcare resources is the reduction in the incidence of the abacavir hypersensitivity syndrome . It is apparent though that the success of ameliorating the abacavir hypersensitivity syndrome by genetic screening is proving difficult to emulate for other drug-ADR combinations. This highlights the considerable hurdles encountered in translating a pharmacogenetic association into a clinical test that benefits patient safety. The development of international consortia alongside the potential of next generation sequencing technologies and other innovations offer tantalising prospects for future advances in pharmacogenetics to reduce the burden of ADRs.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Adverse drug reaction
- Pharmacogenetics
- Predictive genotyping
- Translation
- Abacavir
- Hypersensitivity
- Malignant hyperthermia
- Codeine
- Warfarin
- Statin
1 Introduction
For many drugs, substantial evidence exists at the population level to advocate their use. However there is considerable inter-individual variability in drug response, affecting both drug efficacy and safety [1]. Over 961.5 million prescription items were dispensed in England in 2011 [2]. This high drug usage and the individuality of drug response contribute to the high frequency of adverse drug reactions (ADRs). A prospective study in England estimated that 6.5 % of hospital admissions for patients > 16 years old were related to an ADR, with a median inpatient stay of 8 days [3]. This was contextualised through extrapolation to the entire hospital bed base of England to suggest that the equivalent of up to seven 800 bed hospitals in England could be occupied at any one time with patients admitted with ADRs [3]. Studies from other countries have reported similar ADR-related hospitalisation rates [4–8]. Clearly, ADRs pose a significant international challenge to the health and safety of individual patients and to the efficient use of limited resources by healthcare services.
An ADR, as defined by the World Health Organisation (WHO), is ‘a response to a drug that is noxious and unintended and occurs at doses normally used in man for prophylaxis, diagnosis or therapy of disease or for the modification of physiologic function’ [9]. Table 1 provides definitions and examples of the related pharmacovigilance terms [9–12]. In essence, an adverse event (AE) is an umbrella term for any harm occurring to a patient temporally associated with but not necessarily directly attributable to a therapeutic intervention [10, 13]. A subdivision of AE is an adverse drug event, which describes maleficence associated with the use of a drug and includes overdoses, medication error and ADRs [11, 14, 15].
There is considerable variability between ADRs in terms of presentation and level of current aetiological understanding. This poses a challenge to their accurate categorisation and so, different classifications have been developed. The most well-known system delineates ADRs into types A and B. Type A (‘augmented’) ADRs constitute over 80 % of ADRs; they are dose-dependent and predictable from the main pharmacological action of a drug [13, 16]. This is because Type A ADRs are ‘on-target’ and manifest through excessive drug action at the therapeutic target site. Type B (‘bizarre’) ADRs are dose-independent and are not predictable from a drug’s conventional pharmacology [13, 16] as they represent idiosyncratic ‘off-target’ drug effects. This classification was first defined in 1977 [17] and has been variously extended subsequently to include additional categories as shown in Table 2 [13, 18]. However, it can prove difficult to categorise ADRs using this system. For example, some type B ADRs, including statin-induced muscle toxicity, are clearly dose related and other type B ADRs, such as hypersensitivity to abacavir, are now predictable. A second system is the DoTS classification, which categorises ADRs according to dose relatedness, timing and patient susceptibility factors [19]. This descriptive system improves the accuracy of ADR classification, but its complexity makes it more difficult to use.
In this chapter, ADRs will be classified as immune- or nonimmune-mediated. Immune-mediated ADRs result principally from a deleterious immune reaction mounted following drug exposure. Nonimmune-mediated ADRs encompass all other ADRs and as a point of clarification, include infections that result from predictable immunosuppression by biologics and disease-modifying agents. This is a simple classification, but it reflects the clinical presentation and predominant pathogenic processes of many ADRs and is helpful when considering pharmacogenetics.
The reasons for the heterogeneity in inter-individual drug response are often not known but there are a trilogy of implicated factors: environmental (e.g. drug-drug and drug-food interactions), clinical (e.g. age, co-morbidities, body mass index (BMI), pregnancy) and genetic [20]. The contribution of each postulated factor likely varies with the drug, ADR and patient ethnicity [21]. A genetic basis for specific ADRs was first suggested in the 1950s when perturbed drug metabolism was associated with abnormal drug responses, such as butyrylcholinesterase deficiency and prolonged apnoea after succinylcholine administration. Over the last decade there has been a rapid growth in our understanding of ADR pharmacogenetics. This has been facilitated by an increased knowledge of the human genome and its variation through the Human Genome Project and HapMap Projects. Further, advances in genetic technologies and a reduction in processing costs have increased the volume of pharmacogenetic research conducted and its capacity to yield associations. At present, disproportionately more is known about associations of strong individual effect size with specific ADRs. However for some ADRs it may be that the genetic contribution is polygenic with distinct loci of individual low effect size collectively contributing. Despite the advances of the last decade, elucidation of potential complex interplays within and between different biological systems is proving challenging.
The rest of this chapter explores further the pharmacogenetics of immune- and nonimmune-mediated ADRs. A discussion about all known genetic associations is beyond the scope of this chapter and so a range of genetic associations with distinct ADRs have been selected, although the tables included provide additional examples. The selected associations facilitate expansion on the following key themes:
-
the effect of specific genetic mutations on protein function,
-
the variable extent of genetic contribution to ADRs,
-
the pathogenesis of ADRs,
-
the clinical application of specific genetic-ADR associations through predictive genotyping and
-
the current variable evidence base supporting their use.
Lastly, the many challenges faced by pharmacogenetics in translating an observed genetic-ADR association from the ‘bench’ to the ‘bedside’ will be highlighted and contemporary strategies and future possibilities to overcome these obstacles and deepen our understanding of pharmacogenetics will be outlined .
2 Immune-Mediated Adverse Drug Reactions
Immune-mediated ADRs are off-target ADRs and more specifically, represent a form of hypersensitivity reaction . Hypersensitivity reactions can be classified according to the Gell and Coombs system into types I-IV representing IgE-mediated allergic reactions (type I), direct antibody-mediated (type II), immune complex-mediated (type III) and delayed-type hypersensitivity (DTH) reactions (type IV). At the time of writing, comparatively less is known about the pharmacogenetics of type I-III hypersensitivity reactions and therefore this section will concentrate on DTH reactions.
Over the last decade, the increasing use of genome-wide association studies (GWAS) in pharmacogenetic research has identified a growing number of ADRs that are strongly associated with specific human leukocyte antigen (HLA) haplotypes, genes and/or alleles. Table 3 provides an overview of HLA-ADR associations [22–59].
The HLA class I and II genes, located on chromosome 6, are the most polymorphic of the human genome and over 7000 classical alleles have been identified between them [60]. There is strong linkage disequilibrium between the alleles [61]. Classical HLA class I molecules (encoded on 3 loci: HLA-A, -B, -C) are expressed on the surface of most nucleated cells and present peptide antigen to the T-cell receptor (TCR) of CD8+ T-cells [62]. The peptides presented by HLA class I molecules are mostly derived from the degradation of intracellular proteins, although HLA class I molecules on specific dendritic cell subsets are additionally capable of presenting extracellular peptides through ‘cross-presentation’ [63]. Classical HLA class II molecule expression (encoded on 3 loci: HLA-DP, -DQ, -DR) is restricted to professional antigen-presenting cells (e.g. dendritic cells, macrophages, B-cells) and they present extracellular-derived peptides to the TCR of CD4+ T-cells [62]. HLA polymorphisms localise to the sequence motifs that encode residues of the peptide-binding groove [60, 64]. These polymorphisms alter the stereochemistry of pockets within the groove, creating individual HLA allotypes with distinct peptide-binding portfolios [62, 65]. The HLA system is integral to the development of T-cell tolerance to ‘self’ and to the development of adaptive immunity in response to ‘non-self’ peptide. HLA incompatibility is also known to be important in the pathogenesis of allogeneic transplant rejection and several HLA associations have been previously reported for autoimmune diseases including ankylosing spondylitis (with HLA-B27) and rheumatoid arthritis (e.g. with HLA-DRB1 alleles [66]).
Most of the hypersensitivity ADRs with HLA associations, including the specific reactions to abacavir, carbamazepine, allopurinol and flucloxacillin discussed below, are considered DTH reactions. In keeping with DTH reactions, they normally present ≥ 72 h after drug exposure, may resolve with drug cessation and often re-present more rapidly and with a more severe phenotype following drug re-exposure. A T-cell mediated immunopathogenesis is thought to underlie this temporal pattern. Analogous to the development of pathogen-induced adaptive immune responses, it is thought that a T-cell clone(s) can be primed by presentation of culprit antigen on an HLA molecule during primary drug exposure and effector memory T-cells are rapidly activated with secondary exposure [62, 67, 68]. The isolation of drug-specific T-cells from patients that have suffered DTH ADRs supports T-cell involvement [69, 70].
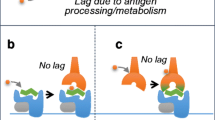
Two hypotheses have conventionally been proposed to describe potential off-target pharmacodynamic processes that may lead to the neo-antigen formation necessary for DTH drug-specific T-cell development: the hapten (or pro-hapten) model and the pharmacologic interaction with immune-receptors (p-i) model [71]. The hapten model proposes that drugs and their metabolites are too small to be independently immunogenic and so covalently bind to self-protein and the resulting de novo hapten-self peptide adduct is antigenic [71, 72]. The p-i hypothesis proposes that drugs may interact directly with HLA molecules, without specific self-peptides, to elicit a T-cell response [73]. Regardless of the mechanism of neo-antigen formation, it is widely assumed that additional ‘danger’ signals are required to overcome the immune system’s default tolerance and permit generation of an adaptive immune response. This concept is referred to as the ‘danger hypothesis’ [74] . Amongst the other key themes of this chapter, the following ADR examples illustrate how prior understanding of genetic susceptibility can facilitate elucidation of underlying mechanisms of antigen formation and presentation .
2.1 HLA-B*57:01 and Abacavir Hypersensitivity Syndrome
Abacavir represents the epitome of translational pharmacogenetics as the loop from laboratory observation to improved patient care for the genetic association between HLA-B*57:01 and the abacavir hypersensitivity syndrome (AHS) has been closed [75]. Abacavir is a nucleoside reverse transcriptase inhibitor indicated to treat HIV and is prescribed as a constituent of highly active antiretroviral treatment (HAART). AHS occurs in 2.3–9 % of patients [76] with a median time to onset of 8 days therapy [77]. The clinical diagnostic criteria require ≥ 2 of: fever, rash, nausea, vomiting, arthralgia, myalgia, headache, lethargy or gastrointestinal symptoms and importantly, onset must occur within 6 weeks of commencing therapy and remit within 72 h of abacavir cessation [76]. Unlike other drug hypersensitivity reactions, the mild to moderate rash is not a consistent feature [67] and eosinophilia is unusual [78]. Although the initial reaction is unpleasant, the significant morbidity and mortality occurs upon rechallenge [67, 78], consistent with a DTH reaction.
In 2002, two groups independently reported an association between AHS and HLA-B*57:01 [22, 23] and subsequent further observational research confirmed the association [79, 80]. The Prospective Randomised Evaluation of DNA Screening in a Clinical Trial (PREDICT-1) study was a multicentre, double-blind randomised controlled trial (RCT) that demonstrated pre-therapy HLA-B*57:01 screening significantly decreased the incidence of AHS [77]. Briefly, 1956 patients were enrolled and randomised on a 1:1 basis. The interventional group received pre-therapy HLA-B*57:01 genotyping and either HAART with abacavir for HLA-B*57:01 negative patients or HAART without abacavir for HLA-B*57:01 positive patients. The control group received HAART with abacavir and retrospective HLA-B*57:01 genotyping from blood samples taken pre-therapy. All participants with clinically diagnosed hypersensitivity reactions underwent skin patch testing for immunological corroboration to improve the specificity for the hypersensitivity phenotype. The study demonstrated that avoiding abacavir in HLA-B*57:01 positive patients in the prospective screening interventional group eliminated immunologically confirmed hypersensitivity reactions (0 vs. 2.7 % in control group, p < 0.001) with positive and negative predictive values of 47.9 % (PPV) and 100 % (NPV), respectively [77]. An estimate of the number needed to screen (NNS) to prevent one case of AHS, given an HLA-B*57:01 carriage prevalence of 6 %, was ~ 25 [77]. However 84 % of participants were Caucasian, limiting generalisation. The Study of Hypersensitivity to Abacavir and Pharmacogenetic Evaluation (SHAPE) was a retrospective case-control study that addressed this and demonstrated that HLA-B*57:01 has 100 % sensitivity for immunologically confirmed AHS in both US White and Black patients [81]. Pharmacoeconomic evaluations have demonstrated a cost effectiveness to pre-prescription HLA-B*57:01 screening [80, 82, 83]. Observational data from open-screening studies has addressed practical matters of implementation [84–86] and shown genotyping to reduce the frequency of abacavir discontinuation due to clinically suspected as well as true immunological hypersensitivity reactions [85, 87]. This is most likely because the former clinician strategy of over-diagnosing AHS to ensure high sensitivity to avoid AHS maleficence at the expense of lower specificity [78] is no longer required given the exclusivity of the association between HLA-B*57:01 and AHS. In accordance with the substantial evidence base, the drug label has been updated and clinical guidelines either mandate or strongly recommend prospective screening [76]. To summarise, abacavir represents a pioneering example of ADR translational pharmacogenetics and has charted a course that other genetic-ADR associations might follow from initial observations to a RCT to studies that address generalisation, pharmacoeconomics and applicability in widespread clinical practice.
Identifying the genetic basis for AHS has directly benefitted patient care but until recently, insight into the underlying immunopathogenesis has been limited. However 3 recent independent studies have begun to expose the pharmacodynamic off-target molecular mechanisms [88–90]. Native abacavir can bind non-covalently with exquisite specificity to HLA-B*57:01 at the base of its peptide-binding groove, extending into the deep F pocket [88, 89]. The specificity for the interaction is accounted for by the F-pocket architecture and in particular residue 116 [88]. In the absence of abacavir, the C-terminus of peptides that bind to the F-pocket of HLA-B*57:01 have large hydrophobic residues, such as tryptophan and less commonly phenylalanine [89]. In the presence of abacavir, the peptide repertoire of HLA-B*57:01 shifts, so that around 20–25 % of recoverable peptides are novel [88] and have alternative residues including isoleucine or leucine at their C-terminus [88, 90]. In essence, abacavir alters the stereochemistry of HLA-B*57:01 to create an HLA neo-allotype with a novel peptide portfolio. This model of antigen presentation is distinct from both the hapten and conventional p-i models. It is proposed that T-cells will not have been exposed to the novel range of HLA-B*57:01-restricted peptides during thymic maturation and so will lack tolerance. The formation of memory T-cells will lead to systemic AHS that is more deleterious upon abacavir re-exposure. In support of the large peptide shift and subsequent large array of ‘altered immunological self’, the observed CD8+ T-cell response is polyclonal [65]. Further, the effector T-memory cells from HLA-B*57:01 positive patients with a clinical history of AHS respond preferentially in the presence of specific peptide and abacavir together rather than to peptide or abacavir alone [89]. This indicates that the memory T-cell response to self-peptide requires abacavir for efficient presentation.
Although the exact intracellular site(s) where abacavir associates with HLA-B*57:01-peptide complexes is currently unclear, there is evidence to suggest the endoplasmic reticulum [65]. However, a minority of T-cell clones in vitro appear to react to abacavir too quickly to be explained by de novo HLA-B*57:01-novel peptide assembly [91]. This suggests that abacavir may additionally bind to HLA-B*57:01-native peptide complexes already present on the cell surface, possibly distorting their stereochemistry [65]. This mechanism is in keeping with the conventional p-i hypothesis. An inadequately resolved question is why the PPV of HLA-B*57:01 for AHS is < 50 % [77]. Postulated mechanisms to account for this include (a) the inter-individual polygenic influence on the novel peptide portfolio itself [89]; and (b) heterologous immunity as a result of pre-existing viral infections, in keeping with data which show that hypersensitivity reactions are often associated with re-activation of viruses such as Epstein-Barr virus [92].
2.2 HLA-B*15:02, HLA-A*31:01 and Carbamazepine Hypersensitivity
Carbamazepine is indicated in the treatment of epilepsy, trigeminal neuralgia and bipolar affective disorder but in up to 10 % of patients, it can provoke a cutaneous ADR [93]. Drug-induced skin injury (DISI) encompasses a spectrum of manifestations and can be caused by a diverse range of drugs including anticonvulsants, allopurinol and β-lactam antibiotics [94]; Table 3 lists drugs with known associations between DISI and genetic variants. There exists both inter- and intra-drug heterogeneity in DISI presentation but fortunately most reactions are mild [94]. Standardising phenotypic definitions for serious DISI conditions has been challenging and required an international collaborative approach. Carbamazepine itself can cause DISI ranging from mild maculopapular exanthema (MPE) of increasing severity to the hypersensitivity syndrome (HSS), also referred to as drug reaction with eosinophilia and systemic symptoms (DRESS) or drug-induced hypersensitivity syndrome (DIHS) [94], to the distinct Stevens-Johnson syndrome (SJS) and toxic epidermal necrolysis (TEN) [95].
HSS/DRESS/DIHS (herein referred to as HSS) and SJS-TEN represent severe cutaneous adverse reactions (SCARs) [96]. HSS is a multisystem disorder that carries a mortality rate of 10 % [96]. HSS can be diagnosed by the presence of at least 3 of: cutaneous involvement, internal organ involvement, fever, lymphadenopathy and eosinophilia (and/or atypical lymphocytes) with at least 1 of the first 2 criteria listed being present [94]. The skin manifestation is most commonly an exanthematous eruption and the internal organ involvement includes hepatic dysfunction and interstitial nephritis [94, 97]. SJS and TEN represent different severities along a spectrum of the same disease and are characterised by epidermal detachment involving the skin and mucous membranes with systemic manifestations including fever, intestinal and pulmonary involvement [94, 98]. SJS is diagnosed when epidermal detachment affects ≤ 10 % of the body surface area, TEN is diagnosed when > 30 % is affected and an overlap syndrome exists for 10–30 % epidermal detachment [99]. SJS and TEN have estimated mortality rates of up to 5 and 50 %, respectively [98].
Genetic association studies have shown HLA-B*15:02 to be a susceptibility factor for carbamazepine-induced SJS-TEN in people of Han Chinese descent [32] and certain other Asian ethnicities including Thai, Malaysian and Indian [33, 100–102]; a meta-analysis of studies with Asian patients derived a pooled odds ratio (OR) of 113.4 (95 % confidence interval (CI) 51.2–251.0) [34]. A recent open-label prospective study in Taiwan has demonstrated the beneficence of pre-therapy HLA-B*15:02 screening in Han Chinese patients and reported no cases of SJS-TEN in both the carbamazepine-taking HLA-B*15:02 negative cohort and HLA-B*15:02 positive cohort administered alternative medication or advised to continue their pre-study medication [103]. Due to ethical and sample size considerations, an estimated historical annual incidence of carbamazepine SJS-TEN (0.23 %) was used as the comparator rather than a prospective non-screened control group (with appropriate blinding), unlike the PREDICT-1 study for AHS [77]. Furthermore this genetic correlation is both phenotypically restricted to SJS-TEN [34, 41] and ethnically restricted: it has not been reproduced in Europeans [104, 105] or other specific Asian populations including South Koreans [26] and Japanese [106–108]. The reason(s) for the latter are not clear but allelic frequency should be considered since HLA-B*15:02 is present in 8.6 % of the Han-Chinese population but in < 1 % of Europeans, Koreans and Japanese [26, 109], reducing the power of studies in these populations to detect statistically significant associations [109].
A more recently reported carbamazepine DISI association is with HLA-A*31:01 and importantly, it is associated with MPE, HSS and SJS-TEN and is present in multiple diverse ethnicities, including Han Chinese [41], Koreans [26], Japanese [107] and Europeans [27]. Interestingly, the allele frequency of HLA-A*31:01 for these ethnic groups is 1.8, 10.3, 9.1 and 2–5 %, respectively [26, 109]. The magnitude of the association though is smaller than with HLA-B*15:02, with a meta-analysis pooled OR for HLA-A*31:01 of 9.5 (95 % CI 6.4–13.9) [34]. However, the estimated NNS to prevent one case of carbamazepine DISI with HLA-A*31:01 is lower and ranges from 47–67 depending on ethnicity. This is in contrast to the estimated NNS of 461 Asian patients with HLA-B*15:02 to specifically prevent one case of SJS-TEN [34]. This difference in NNS is largely attributable to the higher incidence of ADRs (up to 10 % [93]) that are associated with HLA-A*31:01 compared to HLA*B15:02 (circa 0.23 % [103]), due to the relationship of HLA-A*31:01 with a broader range of phenotypes. These findings form a credible foundation for a future prospective study to assess the clinical benefit of pre-therapy HLA-A*31:01 screening.
It has been shown that carbamazepine can non-covalently associate with HLA-B*15:02 molecules and in the presence of carbamazepine, there is a shift in the peptide repertoire of HLA-B*15:02 with a novel preference for smaller residues at the 4th and 6th peptide positions and an increase in hydrophobic residues at several positions [88]. The scale of this peptide shift is smaller (approximately 15 %) than for abacavir [88]. This off-target pharmacodynamic effect does resonate with the novel model proposed for abacavir and HLA-B*57:01, but overall the mechanisms leading to carbamazepine hypersensitivity ADRs are likely to be more complex. This is because firstly, it has been shown that carbamazepine and its metabolite, carbamazepine-10,11-epoxide, can associate with other structurally related HLA-B75 family members [110]. Secondly, the availability of a restricted range of T-cell clonotypes with specific TCR rearrangements has been demonstrated to be an important determinant in the pathogenesis of HLA-B*15:02-associated carbamazepine-induced SJS-TEN [111]. This indicates that both the TCR repertoire and HLA genotype modulate the risk of carbamazepine-induced SJS-TEN and likely explains why some HLA-B*15:02 carriers tolerate carbamazepine [109]. Further research is still required to understand other complicating observations. These include how two seemingly disparate HLA alleles, HLA-B*15:02 and HLA-A*31:01, are linked to carbamazepine ADRs and furthermore, how HLA-A*31:01 can be associated with multiple phenotypes.
2.3 HLA-B*58:01 and Allopurinol Hypersensitivity
Allopurinol, an analogue of hypoxanthine, inhibits xanthine oxidase (XO) and is indicated in the management of gout and other hyperuricaemic conditions including tumour lysis syndrome. Although generally well tolerated, ~ 2–3 % of patients suffer mild hypersensitivity reactions including MPE [39, 112] and crucially, ~ 0.1–0.4 % of patients develop HSS or SJS-TEN [112]. The SCARs, HSS and SJS-TEN, normally present within weeks to months of commencing allopurinol but may take considerably longer [112]. Allopurinol is a major cause of SCARs [113, 114] and the combined mortality from allopurinol-induced SCARs approaches 25 % [112].
In 2005, a strong genetic association between HLA-B*58:01 and allopurinol-induced SCARs (HSS/SJS-TEN) was described in the Taiwan Han-Chinese population (OR 580.3, 95 % CI 34.4–9780.9); all 51 cases (100 %) carried HLA-B*58:01 in comparison to only 20 of 135 allopurinol-tolerant controls (15 %) [24]. This association has been replicated in Thai [31], Korean [25], European [35] and Japanese patients [115], although its magnitude was more modest for the latter 3 ethnic groups, possibly reflecting the lower prevalence of HLA-B*58:01 in these populations. A meta-analysis has confirmed the association between allopurinol-induced SJS-TEN and HLA-B*58:01 in both Asian and non-Asian patients compared to allopurinol-tolerant controls (combined OR 96.6, 95 % CI 24.5–381.0) [116]. Based on data from the Han Chinese and Thai populations, current estimates for the PPV and NPV for HLA-B*58:01 are ~ 1.5 and 100 %, respectively [112], although these values will be lower for other ethnic groups. An exciting development is the ongoing prospective study in Taiwan to assess the clinical benefit of pre-therapy genotyping for HLA-B*58:01 prior to commencing allopurinol [112]. Similarly to the prospective study discussed above for HLA-B*15:02 screening prior to initiating carbamazepine [103], an estimated historical ADR incidence is being used for the control. At the time of writing, no results from this study have been published.
Interestingly, unlike carbamazepine and HLA-A*31:01, it is less clear at the current time whether HLA-B*58:01 also predisposes to MPE in patients taking allopurinol. A study in Australia demonstrated no association between HLA-B*58:01 and MPE [117], but a study of Han-Chinese patients in mainland China reported HLA-B*58:01 as a risk factor for both allopurinol-induced MPE and SCARs [39]. More research into this area is required, but one hypothesis from the available literature is that the risk of MPE with HLA-B*58:01 may be ethnically-restricted. If this association is confirmed, it will increase the PPV further for the affected ethnic groups and so augment the potential utility of pre-therapy HLA-B*58:01 screening in these groups to reduce the burden of allopurinol-induced ADRs.
The exact underlying mechanism(s) by which allopurinol, or its long-circulating active metabolite oxypurinol, interact with HLA-B*58:01 for the generation of drug-specific T-cells has yet to be elucidated. However, as the PPV of HLA-B*58:01 for SCARs is low (~ 1.5 %) [112], this alludes to other contributing factors in their pathogenesis. It has long been thought that viruses play a role in drug hypersensitivity and there is increasing recognition that the reactivation of herpes viridae is important in the aetiology of HSS [92, 118]. However, any interaction(s) between allopurinol and/or oxypurinol and viruses is poorly understood. Prior to the discovery of HLA-B*58:01, several non-genetic risk factors were espoused including renal dysfunction, higher allopurinol doses, diuretic use and concomitant antibiotic therapy [112]. Although verification of these variables is difficult as SCARs are fortunately rare events, patients on allopurinol with renal insufficiency have been shown to be almost 5 times more likely to develop SCARs [24]. In addition, patients on a daily dose of ≥ 200 mg allopurinol seem to be at an increased risk of SJS-TEN compared to lower doses [113]. By assimilation of these 2 observations, it can be hypothesised that increasing the plasma concentration of allopurinol and/or oxypurinol increases the risk of drug-specific T-cell development [109]. To mitigate the risk of ADRs the dose of allopurinol could be reduced, but it is well established that the most commonly used doses of allopurinol (≤ 300 mg daily) are frequently ineffective already for the long term treatment of hyperuricaemia in gout [119].
Nevertheless, HLA-B*58:01 is the single largest predictor of allopurinol-induced SCARs and this makes genetic screening appealing to directly prevent HLA-B*58:01-associated SCARs. In addition, it is conceivable that a successful genetic screening programme may indirectly improve the overall benefit: harm ratio of allopurinol further. This is because genotyping may empower clinicians to titrate allopurinol doses up to optimise efficacy in HLA-B*58:01 negative patients with normal renal function. However, any benefit derived from genetic screening in the ongoing Taiwan study will require follow up studies to determine the extent of generalisation. This is because for other ethnic groups and especially Europeans, allopurinol-induced SCARs also occur in HLA-B*58:01 negative patients. Furthermore, the identification of other (non)-genetic risk factors may be required to improve the PPV of the test, as currently many HLA-B*58:01 patients will be unnecessarily denied allopurinol in place of other urate-lowering therapies, with unmeasured effects as yet on cost-effectiveness and treatment efficacy.
2.4 HLA-B*57:01 and Flucloxacillin-Induced Liver Injury
Flucloxacillin is a narrow-spectrum beta-lactam antibiotic indicated in Gram-positive bacterial infections and in particular, is used to treat non-methicillin resistant Staphylococcus aureus infections. In approximately 8.5 per 100,000 patients treated with flucloxacillin, cholestatic liver injury occurs [120]. Drug-induced liver injury (DILI) is associated with a structurally disparate range of drugs but notably these include non-steroidal anti-inflammatory drugs (NSAIDs) and certain antimicrobials including flucloxacillin [121]. Although rare, drug-induced liver injury (DILI) can be severe and accounts for up to 15 % of all cases of acute hepatic failure [122–124]. Analogous to DISI, the type and severity of DILI vary between causative drugs and for a given drug, presentation is variable [121]. Consequently, standardising the DILI phenotype is not straightforward but the diagnosis can be made from clinical, biochemical and histopathological parameters [125].
The aetiology of DILI can be divided into immune- and nonimmune-mediated processes [121]. Pharmacogenetic associations with DILI have now been identified for several drugs and the associated genetic variants reflect both immune and nonimmune aetiologies (see Tables 3 and 4, respectively for examples). However, DILI can be difficult to categorise by this means. This is because, although recognition of clinically suggestive features of hypersensitivity is relatively easy, the absence of such features, such as eosinophilia, does not preclude immune system involvement [126].
To date, the strongest DILI genetic association described is between flucloxacillin and HLA-B*57:01 with an OR of 80.6 (95 % CI 22.8–284.9) [47]. This is intriguing as the HLA-B*57:01 allele is also strongly associated with AHS, yet AHS rarely involves hepatitis [78].
Unlike AHS, it is improbable that this association will lead to a screening test for clinical practice because, despite an adequate estimated sensitivity and specificity (84 and 94 %, respectively) [47], the rarity of flucloxacillin-induced liver injury diminishes the PPV to 0.12 % [127]. An estimate of the NNS to prevent one flucloxacillin-induced liver injury case is 13,513 and the screening approach would unnecessarily deny almost 7 % of patients first line flucloxacillin therapy, with unmeasured adverse effects on infectious disease treatment efficacy and cost effectiveness [127]. However genetic testing may help establish the diagnosis of flucloxacillin-induced liver injury when the underlying cause of liver dysfunction is unclear [47].
The off-target pharmacodynamics that underpin flucloxacillin-induced liver injury are being unravelled. Flucloxacillin can adduct covalently to proteins to form neo-antigen drug-protein conjugates and specific flucloxacillin-modifiable lysine residues on albumin, the major circulating protein, have been identified [128]. It is predicted that several albumin-derived peptides containing flucloxacillin-modifiable lysine residues have high-affinity for HLA-B*57:01 [129] and could be presented on HLA-B*57:01 by professional antigen presenting cells through cross-presentation. Flucloxacillin-responsive CD4+ and CD8+ T-cells have been characterised in vitro from patients who have previously suffered flucloxacillin cholestatic liver injury and their activation is dependent on peptide processing. In vitro, the CD8+ T-cell activation is restricted to HLA-B*57:01 and the very similar allotype, HLA-B*58:01 [129]. The proposed model, which aligns with the hapten hypothesis, suggests that immunogenic peptide neo-antigens are derived from natural processing of flucloxacillin-protein conjugates and can be presented on HLA-B*57:01 to generate an adaptive immune response [129]. Interestingly, besides this proposed alternative mechanism of neo-antigen formation, this immunopathogenesis differs to that of abacavir in at least 2 ways. Firstly, CD4+ as well as CD8+ flucloxacillin-responsive T-cells have been cloned from patients and secondly, the T-cell clones show cross-reactivity in vitro with other commonly prescribed beta-lactam antibiotics including amoxicillin and piperacillin [129].
In summary, this section illustrates that immune-mediated DTH reactions are an emerging prominent type of off-target ADR with the potential for significant morbidity and mortality. However, pharmacogenetics has been pivotal in reducing the healthcare burden associated with abacavir, may have important future roles in the prevention of carbamazepine and allopurinol DISI and is facilitating elucidation of underlying immune-mediated aetiologies.
3 Nonimmune-mediated Adverse Drug Reactions
Nonimmune-mediated ADRs are a heterogeneous group in aetiology and presentation. However, over the last decade it has been increasingly recognised that susceptibility to many nonimmune ADRs is associated with gene variants of drug metabolising enzymes (DMEs) and less frequently, with drug transporters . It is thought that perturbed pharmacokinetics increases the availability of drug/metabolite(s) at the target site(s), increasing the likelihood of developing an ADR. The sites that mediate nonimmune ADRs include both on-target and off-target sites. On-target ADRs manifest through excessive drug/metabolite(s) action either at the therapeutic target site or at the same molecular site located in other tissues. The latter occurs for instance with NSAID-induced upper gastrointestinal ADRs.
It is important to note that, although the majority of ADRs with a genetically-influenced pharmacokinetic-mediated susceptibility found to date are nonimmune ADRs, perturbed pharmacokinetics is also relevant in the genesis of a few immune-mediated ADRs. This was described earlier for the case of allopurinol-induced SCARs and non-genetic pharmacokinetic factors. Furthermore, genetic susceptibility to ticlopidine-induced hepatotoxicity has been demonstrated to be greatest in patients with HLA-A*33:03 in combination with variants of a DME (Table 3).
In the following section, the effects of gene variants of phase I and phase II biotransformation enzymes on susceptibility to ADRs will be discussed in the context of codeine/warfarin and azathioprine, respectively. Then, the effects of gene variation for a drug transporter will be illustrated for statin-induced muscle toxicity. However as the first example of malignant hyperthermia shows, genetic susceptibility to nonimmune-mediated ADRs can occur through plausible pharmacodynamic mechanisms too. Table 4 lists examples of ADRs associated with nonimmune-related genetic variants .
3.1 RYR1 and Anaesthesia-Induced Malignant Hyperthermia
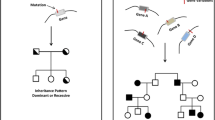
In 1962, a paper was published about a pedigree that contained 10 relatives who had unfortunately and unexpectedly died during or shortly following general anaesthesia [163]. The deaths were associated with core body temperatures, when measured, in excess of 41 °C and followed an autosomal dominant inheritance pattern [163]. Other pedigrees have since been described [164, 165] and over 500 cases of malignant hyperthermia (MH) have now been reported in the medical literature [166].
MH is precipitated by volatile anaesthetics in genetically susceptible individuals. All halogenated inhalation anaesthetics have been implicated including halothane, isoflurane, sevoflurane and desflurane [167]. The depolarising neuromuscular blocker, succinylcholine, augments the adverse response to these potent inhalation anaesthetics but its role as an independent precipitant of fulminant MH is controversial [167, 168]. Rarely, non-pharmacological stressors including environmental heat [169, 170], infections [170] and severe exercise or emotional strain [171] have been implicated in MH-like episodes.
The incidence of anaesthetic-induced MH is approximately 1 per 50,000 adults and 1 per 15,000 paediatric patients [172] and it occurs in all ethnic groups [173]. The basis of MH is hypermetabolism which can present as tachypnoea, a rise in end-tidal carbon dioxide exhalation, tachycardia, cyanosis, cardiac arrhythmias, skeletal muscle rigidity, hyperthermia [174], convulsions and eventual death [163]. Associated electrolyte complications include acidosis, hyperkalaemia, elevated creatine kinase (CK) and acute kidney injury (AKI) [174]. Timely intervention improves prognosis [175]. However, an early diagnosis of MH can be challenging as the initial clinical signs are nonspecific and variable in their time course, making them easily mistaken for other pathologies (e.g. sepsis, thyrotoxic crisis) [176]. Nevertheless, the mortality from MH has dramatically fallen from 70 % in the 1970s [169] to < 5 % today [173]. This reduction has been aided by the introduction of the muscle relaxant dantrolene for treatment of suspected MH [166] and testing for susceptible relatives (see later) [169]. A clinical grading scale has been introduced to help researchers retrospectively assess the likelihood of MH following an adverse anaesthetic event, which enables accurate phenotyping and determination of future susceptibility [177].
RYR1 is located on chromosome 19 and encodes ryanodine receptor 1 (RyR1). There are 3 RyRs isoforms (RyR1–3) and each forms a homotetrameric assembly within the endoplasmic (or sarcoplasmic) reticulum and functions as a Ca2+ channel [178]. They have evolved into the largest ion channels found to date (~ 2.2MDa) [179]. This is undoubtedly to facilitate tight channel regulation through interaction with numerous regulatory small molecules and proteins, which is important as Ca2+ is a potent intracellular mediator of several cell processes [179]. RyR1 is widely expressed in skeletal muscle and is pivotal to excitation-contraction coupling [180]. RyR1 opens in response to nerve impulses and releases Ca2+, from the sarcoplasmic reticulum where it has been sequestered, into the cytoplasm to drive muscle contraction. It is thought that a direct physical connection exists between the voltage-gated Ca2+ channel CaV1.1 (the skeletal dihydropyridine receptor) in the transverse tubule and RyR1 [181, 182], which induces conformational changes that open RyR1 when the wave of depolarisation from the neuromuscular endplate is detected by CaV1.1 [183].
Approximately 70 % of MH susceptible families carry RYR1 variants [156]. A nonsynonymous mutation of RYR1 was found to cause the porcine stress syndrome in inbred pigs, which is an animal model of MH [184]. The analogous C1843T mutation in humans was subsequently identified in an analysis of 1 of 35 MH susceptible pedigrees [185]. Currently, over 200 RYR1 mutants have been described and most are single nucleotide polymorphisms (SNPs) , but only 31 have been designated as causative of MH according to the specific criteria set out by the European Malignant Hyperthermia Group (EMHG) [155] .
Impaired Ca2+ homeostasis underlies the pathogenesis of MH [173]. Gain-of-function RYR1 mutations have been shown in vitro to lead to RyR1 hyperactivation [186, 187]. The increase in intracellular Ca2+ concentration results in sustained muscle contraction and heat generation [173]. Attempts to restore the Ca2+ balance and the contracting muscle filaments deplete the cell of adenosine triphosphate resulting in muscle rigidity, loss of integrity to the sarcolemma and leakage of intracellular contents (e.g. K+, myoglobin) out into the extracellular fluid predisposing to systemic sequelae [172]. However, the exact mechanism(s) by which volatile anaesthetics precipitate this potentially fatal cascade has not been clearly elucidated [188].
Interestingly, ~ 20 % of patients have undergone previous uneventful general anaesthesia with potent inhalation agents before experiencing MH [166]. The reasons for this incomplete penetrance are not fully understood but hypotheses include dose and/or duration dependency effects of the volatile anaesthetic agents [167], the ambient temperature and the simultaneous use of possible mitigating drugs [173].
The majority of MH occurs in asymptomatic individuals and they are considered to have a genetically-determined subclinical myopathy [176]. However, there are at least 3 rare clinical myopathies likely associated with MH susceptibility: central core disease (CCD), multiminicore disease (MmD) and King-Denborough syndrome [189, 190]. Within each syndrome there is clinical, genetic and histological variability and considerable overlap exists, in particular, between CCD and MmD [189]. Importantly, the majority of CCD cases are associated with RYR1 variants and furthermore, RYR1 mutations have also been found in cases of MmD [189] and King-Denborough syndrome [191], although these links are less certain [192]. Of the 200 RYR1 variants, ≥ 150 are associated with MH alone (subclinical myopathy), ~ 100 with CCD and ≥ 20 with both MH and CCD [192]. RYR1 alleles are also implicated in instances of exercise-induced rhabdomyolysis [193, 194]. Clearly, RYR1 is involved in a spectrum of muscle disorders, but at the present time the degree of genotype to phenotype concordance is incompletely understood.
The gold standard for MH diagnosis in patients and unaffected relatives is the in vitro muscle biopsy contracture test (IVCT), which assesses muscle contraction in response to caffeine and halothane [195]. However, the IVCT is invasive, costly and confined to specialist centres. Therefore, genetic testing has been increasingly used since 2001 [156] to determine MH susceptibility in family members of MH patients that have been shown to carry a causative RYR1 mutation, as classified by the EMHG [196]. A relative not carrying the familial RYR1 mutation should still undergo an IVCT though as the absence of a RYR1 mutation does not exclude MH susceptibility [176].
~ 75 % of MH events occur in patients with no reported family history [166] and therefore universal pre-anaesthetic genetic screening is appealing. However, genetic screening for MH is currently untenable, due to the heterogeneous and incompletely understood genetics underpinning MH susceptibility. The complexity of the RyR1 molecule makes structural and functional predictions of RYR1 variants challenging [197] and regardless, 30 % of MH cases are not associated with RYR1. At least 5 other genetic loci have been implicated [172] but of these to date, only nonsynonymous SNPs in CACNA1S, the gene encoding the α1 subunit of CaV1.1, have been linked to MH and in only 1 % of cases [157, 158, 172].
In summary MH is a potentially fatal disorder with a strong genetic predisposition, although the full spectrum of genetic risk variants and associated genotype-phenotype correlations are incompletely characterised. However, this strong genetic susceptibility lends itself to the future prospect of successful genetic screening to reduce the incidence of drug-induced MH.
3.2 CYP2D6 and Codeine Analgesia and Safety
Codeine is a weak opioid that is indicated for analgesia in mild to moderately severe pain and as an antitussive and anti-diarrhoeal agent. Although it has been used for many years, recent concerns are mounting over its variable efficacy and safety.
Figure 1 shows the principal pharmacokinetic pathways for codeine . Codeine is considered a prodrug whose function is derived from conversion into 2 active metabolites: morphine and morphine-6-glucuronide (M6G). Both are agonists for the widespread µ-opioid receptor, which is largely responsible for the therapeutic effects and opioidergic ADRs [198, 199]. The affinity of morphine for µ-opioid receptors is 200-fold stronger than compared to codeine [200]. The polymorphic cytochrome 2D6 enzyme (CYP2D6) catalyses the O-demethylation of codeine into morphine. Uridine diphosphate glucuronosyltransferase 2B7 (UGT2B7) catalyses morphine into M6G and, in conjunction with UGT1A isoforms, into inactive morphine-3-glucuronide (M3G). Only a minority of codeine biotransformation (≤ 15 %) [201] is via CYP2D6; the majority of codeine is converted directly into the inactive metabolites codeine-6-glucuronide and norcodeine by UGT2B7 and CYP3A4, respectively. The major codeine metabolites, including morphine and M6G, are excreted renally [201].
Codeine metabolism. C6G codeine-6-glucuronide, NC norcodeine, M6G morphine-6-glucuronide, M3G morphine-3-glucuronide, NM normorphine, CYP2D6 cytochrome P450 2D6, CYP3A4 cytochrome P450 3A4, UGT2B7 uridine diphosphate glucuronosyltransferase 2B7, UGT1A1 uridine diphosphate glucuronosyltransferase 1A1
CYP2D6 belongs to the superfamily of cytochrome P450 (CYP) genes. They encode haemoproteins that catalyse oxidative, phase I metabolism [202] and account for ~ 75 % of all drug metabolism reactions [203]. Although 57 CYP genes have been identified, ~ 95 % of these reactions are catalysed by just 5 isoenzymes including CYP2D6 [203]. Direct clinical measurement of CYP2D6 phenotypic activity is unfeasible as it is primarily expressed in the liver and indirect measurements of CYP2D6 metabolites in the plasma or urine are susceptible to other factors including renal dysfunction and drug interference. Consequently, CYP2D6 genotyping as a phenotype surrogate is appealing for clinical practice.
CYP2D6 is located on chromosome 22 and over 80 alleles have been identified [204]. They are formed by a range of genetic alterations including SNPs, insertions and deletions and can be grouped functionally into increased, normal, reduced and non-functional alleles [137]. An individual’s CYP2D6 genotype can in turn be categorised into 1 of 4 predicted phenotype classes based on the combination of CYP2D6 alleles they carry: an extensive, intermediate, poor or ultrarapid metaboliser (EM, IM, PM and UM, respectively) [205]. The EM is the wild-type CYP2D6 phenotype, IMs have reduced activity and PMs have no enzymatic activity as they carry no functional alleles. If multiple copies of functional alleles are detected this is denoted the UM phenotype as high enzymatic activity is expected [137]. There is considerable variability in the prevalence of CYP2D6 alleles and in the prevalence of the extreme phenotypes in different ethnic groups (0–10 and 0–29 % for PMs and UMs, respectively) [137].
It has been shown that following codeine administration, PMs have significantly lower plasma morphine concentrations, reduced urinary active metabolite excretion and decreased analgesia compared to EMs [206, 207]. Conversely, plasma morphine concentrations and urinary active metabolite excretion are significantly higher in UMs compared to EMs [208]. Furthermore although there is no definitive study, a growing series of case reports are documenting severe ADRs after standard codeine use associated with the UM phenotype [136, 209–213]. These case reports are from neonatal [209], paediatric [210–212] and adult populations [136, 213] and the documented on-target (opioidergic) ADRs include: severe epigastric pain, euphoria and dizziness [213], central nervous system/respiratory depression [136, 211] and death [209, 210, 212]. One especially poignant case was the death of a 13-day old neonate who was breastfed by a mother taking codeine (and paracetamol) for episiotomy pain [209]. The autopsy found an extremely high level of morphine in the neonate’s blood and a sample of stored maternal breast milk from day 10 showed an elevated morphine concentration. The mother was found to have a CYP2D6 gene duplication indicative of the UM phenotype [209]. Following this report, the US Food and Drug Administration (FDA) issued a warning on codeine use by nursing mothers [214].
Although there is increasing concern regarding the efficacy and safety of codeine , several barriers exist that hamper the translation of CYP2D6 genotyping into widespread clinical practice. Firstly, the ADR profile of PMs is incompletely understood [137]. Secondly, when compared to the prevalence of the UM phenotype (0–10 %), the documented case reports of severe ADRs are rare, suggesting that there are additional genetic and non-genetic susceptibility factors. The pharmacogenetic influence of UGT2B7 is controversial at present [201]. Other risk factors may include renal dysfunction [136, 201, 215], drug inhibitors of CYP3A4 [136, 201], ontogeny [215, 216] and repeated episodes of hypoxia [215]. The paediatric case reports are from children receiving codeine after adeno(tonsillectomy) for recurrent tonsillitis and obstructive sleep apnoea (OSA) [210–212]. OSA leads to intermittent sleep hypoxia and it has been shown that opioid analgesia sensitivity increases in children after recurrent hypoxia [217]. Another factor is potential publication bias favouring selection of case reports documenting extreme but fortunately uncommon ADRs with codeine . 10 of 11 UM participants in a pharmacokinetics study felt sedation (91 %) compared to 6 of 12 (50 %) EMs (p = 0.03) suggesting that ADRs in UMs may occur more frequently than is reported [208]. Other potential barriers include the absence of prospective studies that demonstrate clinical benefit of CYP2D6 genotyping, scarce cost-effectiveness data, lack of clinician knowledge and no clear guidelines on what constitutes a suitable substitute for codeine in CYP2D6 PMs and UMs. This is important because CYP2D6 is involved in the metabolism of other opioid drugs including oxycodone, hydrocodone and tramadol. There is evidence at least for tramadol that CYP2D6 PMs experience reduced analgesia [218] and UMs a higher risk of nausea [138] when compared to EMs. There is also a case report of respiratory depression following tramadol in a UM patient with renal dysfunction [139]. Tramadol and codeine are step 2 ‘weak’ opioid drugs on the WHO analgesia ladder [219] and are often used interchangeably in clinical practice for a patient that does not tolerate one. However if tramadol is also undesirable in CYP2D6 PMs and UMs, clinical guidance regarding suitable alternative analgesic agents is warranted.
3.3 CYP2C9 and the Risk of Haemorrhage with Warfarin
Warfarin is the most frequently prescribed oral anticoagulant worldwide [220] and is indicated in the prophylaxis and treatment of venous thromboembolism (VTE) and in the prophylaxis of systemic embolism in predisposing conditions such as atrial fibrillation and following mechanical heart valve insertion [221]. It is a coumarin-derived therapeutic that is administered as a racemic mixture; the S-warfarin enantiomer is more potent than R-warfarin [222]. They disrupt the vitamin K cycle by antagonising vitamin K epoxide reductase, resulting in a decrease in vitamin K-dependent post-translational γ-carboxylation of protein glutamate residues [223, 224]. This notably diminishes the activity of clotting cascade proteins including the procoagulant factors II, VII, IX and X and anticoagulant molecules protein C and protein S [225]. The overall anticoagulant effect is quantified by the prothrombin time-derived international normalised ratio (INR); the usual desired therapeutic INR is 2.5 [226]. However, certain high thrombotic risk conditions such as recurrent VTE(s) on warfarin and mechanical heart valves warrant higher anticoagulation levels (e.g. a desired INR range of 3.0–4.0) [226].
Epidemiological evidence has implicated warfarin as a major cause of ADRs; it is the therapeutic associated with the greatest number of preventable ADRs in Sweden [227] and the third most common cause of ADR-related hospitalisations in the UK [3]. Haemorrhage is an on-target ADR and is the predominant ADR associated with warfarin [221], especially during therapy initiation [228]. It is highly correlated to the intensity of anticoagulation [229, 230] and the risk of clinically significant bleeding increases when the desired INR range is higher [221]. The safe management of warfarin therapy is notoriously challenging because of the wide inter-individual range of optimal dose requirements (0.6–15.5 mg/day) and its narrow therapeutic index [231]. It is worth noting also that there is evidence to suggest a pharmacogenetic association between CYP2C19*17 carriage and increased bleeding risk in patients taking clopidogrel (Table 4) [130, 131], although for now, the genetic susceptibility to haemorrhage on warfarin will be outlined.
CYP2C9 , like CYP2D6 , is 1 of the 5 main human CYP DMEs [203]. CYP2C9 is the principal enzyme involved in the metabolism of the potent S-warfarin stereoisomer, while R-warfarin is cleared via CYP1A1/CYP1A2/CYP3A4 [228]. Over 30 allelic variants of CYP2C9 are known, but their relative prevalence varies with ethnicity [220]. The CYP2C9 reference genotype *1/*1 produces the normal (EM) phenotype [220] and a resultant estimated warfarin half-life of 30–37 h [232]. The 2 most frequent reduction-of-function minor alleles amongst people with European ancestry are CYP2C9*2 (rs1799853) and CYP2C*3 (rs1057910) [202]. Both are characterised by one nonsynonymous SNP, prolong the half-life of warfarin (up to 92–203 h in *3/*3 homozygotes [233, 234]) and are associated with reduced maintenance warfarin dose requirements [235].
A recent meta-analysis has reported hazard ratios for the risk of bleeding in patients on warfarin with *1/*3 or *3/*3 genotypes, compared to *1/*1 patients, to be 2.05 (95 % CI 1.36–3.10) and 4.87 (95 % CI 1.38–17.14), respectively, suggestive of a gene-dose effect [135]. Although CYP2C9*2 was also significantly associated with bleeding, albeit with a lower pooled effect size than CYP2C9*3, after stratification into *1/*2 and *2/*2 genotypes through synthesis of studies that reported these individual genotypes, neither genotype was significantly associated with bleeding. Overall, CYP2C9*3 is the main risk factor for bleeding on warfarin, which is biologically plausible as the *3 allele has a more deleterious effect than *2 on CYP2C9 enzyme function [135]. For a more comprehensive account of overall warfarin pharmacogenetics , dosing strategies to incorporate multiple environmental, clinical and genetic factors and a discussion regarding the recently published prospective warfarin pharmacogenetic RCTs, the reader at this point is referred to Chap. 11.
3.4 TPMT, Azathioprine- and 6-Mercaptopurine-Induced Myelosuppression
The immunosuppressive agent azathioprine (AZA) is a pro-drug of 6-mercaptopurine (6-MP). AZA is indicated in both the prophylaxis of transplant rejection and in the treatment of many autoimmune conditions including inflammatory bowel disease (IBD), rheumatoid arthritis and severe eczema [236]. 6-MP is conventionally used with haematological malignancies and in particular acute lymphoblastic leukaemia [237], although it also has a role in IBD [238]. AZA/6-MP can induce several ADRs including myelosuppression (predisposing to neutropaenic sepsis), DILI, pancreatitis, nausea and vomiting [239]. Although ADRs occur in 10–28 % of patients [240], the rate of fatal ADRs among AZA users is estimated at 1 in 10,000 [241].
Approximately 90 % of AZA is converted to 6-MP by ubiquitous non-enzymatic processes [242, 243]. Figure 2 depicts the 3 main competing enzyme pathways for metabolism of 6-MP: thiopurine methyltransferase (TPMT), XO and the main anabolic pathway via hypoxanthine phosphoribosyltransferase (HPRT) [244]. Both therapeutics are subject to extensive intestinal and hepatic first pass metabolism following oral dosing [244, 245]. Although there is an incomplete understanding of the modes of action of AZA/6-MP [246], the accumulation of 6-thioguanine nucleotide (6-TGN) metabolites formed in vivo via HPRT is thought to contribute to both their efficacy [247] and when in relative excess, the increased risk of myelosuppression [242, 248]. The immunosuppressive mechanisms include incorporation of 6-TGNs into DNA inhibiting leukocyte DNA synthesis [244, 249] and blockade of Rac1 protein by the 6-TGN derivative, 6-thioguanine triphosphate (6-TGTP), inducing T-cell apoptosis [246]. TPMT can methylate both 6-MP and the intermediate metabolite, 6-thioinosine monophosphate (6-TIMP), to give 6-methylmercaptopurine (6-meMP) and methyl-TIMP (meTIMP), respectively. meTIMP may be efficacious through de novo purine synthesis inhibition [240, 250] whilst high levels of TPMT methylated thiopurine metabolites (and further phosphorylated metabolites) may be associated with DILI [251–255].
Azathioprine (AZA) and 6-mercaptopurine (6-MP) metabolism (simplified). me prefix methyl, XO xanthine oxidase, TU thiouric acid, TPMT thiopurine methyltransferase, HPRT hypoxanthine phosphoribosyltransferase, TIMP thioinosine monophosphate, IMPDH inosine monophosphate dehydrogenase, TXMP thioxanthosine monophosphate, TGN thioguanine nucleotides, TGTP thioguanine triphosphate
TPMT is a phase II biotransformation enzyme, encoded by TPMT on chromosome 6 [243], and is a major pharmacokinetic determinant for active 6-TGN metabolite levels [240], which are inversely related to TPMT activity [244, 256, 257]. It is variably expressed in several tissues; the highest levels of TPMT are present in the liver and the lowest in the brain and lung [240]. Erythrocyte TPMT activity correlates with hepatic TPMT activity [258] permitting direct TPMT phenotypic assessment of patients in clinical practice, which is unusual for a DME [259]. TPMT enzymatic activity follows a trimodal distribution; ~ 90 % of individuals have high activity, ~ 10 % intermediate and 0.3 % low/undetectable enzyme activity [260, 261].
Around 30 allelic variants of TPMT have been reported [20] and despite ethnic variability, 3 account for > 90 % of the minor alleles: TPMT*2, TPMT*3A and TPMT*3C [254]. They are caused by one (TPMT*2, TPMT*3C) or two (TPMT*3A) nonsynonymous SNPs that reduce enzymatic activity through enhancing the rate that the TPMT variant is catabolised [262–264]. Analogous to CYP2D6 and CYP2C9, TPMT genotype correlates with the variable TPMT enzymatic activity levels: heterozygotes have intermediate activity (IM) and individuals carrying no normally functioning alleles have low/absent activity (PM) [254]. Like CYP2D6 and CYP2C9 , homozygous deficient individuals include both those homozygous for 1 variant allele and compound heterozygotes with 2 distinct inactivating alleles [243]. TPMT *1/*1 individuals have normal phenotypic activity (EM).
Clinically, ~ 27 % of AZA/6-MP-induced myelosuppression cases are explained by inactivating TPMT alleles [265], although little correlation exists with other specific ADRs including DILI [239, 266]. A meta-analysis of patients with chronic inflammatory diseases has reported a gene-dose effect for this on-target ADR: homozygous deficient individuals carry a higher risk of leukopaenia (OR 20.84, 95 % CI 3.42–126.89) than heterozygotes (OR 4.29, 95 % CI 2.67–6.89) when compared with *1/*1 individuals [149] and in general the myelosuppression onset is earlier [265, 267] and more severe [267]. A second systematic review, not limited to a specific class of disease, has reported that 86 % of TPMT homozygous deficient patients develop myelosuppression and the pooled OR for patients with intermediate TPMT activity or one TPMT variant allele, compared with wild-type, was 4.19 (95 % CI 3.20–5.48) [268]. For both studies, their results were primarily derived from synthesis of observational studies.
As clinical evidence has grown, consensus national clinical guidelines have been published that recommend and interpret pre-therapy TPMT testing, including the UK dermatology [269] and rheumatology guidelines [270]. In patients identified as TPMT deficient (by either genotyping of homozygous deficiency or TMPT phenotypic analysis of low/absent activity), guidance advises selection of alternative immunosuppressive therapy in non-malignant conditions and a reduction in starting dose to 10 % of normal when treating malignancy [254]. For heterozygous variant/intermediate activity patients commencing AZA/6-MP therapy, a dose reduction of 30–70 % is suggested [254]. TPMT analysis has been adopted into clinical practice and a national survey reported that 94 % of dermatologists, 60 % of gastroenterologists and 47 % of rheumatologists in England requested TPMT testing [271].
Despite the relatively high, albeit variable, clinical uptake of TPMT testing, outstanding issues remain. Firstly, there is a lack of robust prospective randomised evidence assessing the utility of pre-therapy TPMT analysis in reducing myelosuppression. An RCT (n = 333) was undertaken but the recruitment target (n = 1000) was not met due to guideline-driven pre-existing routine TPMT testing at some centres adversely impacting study recruitment [272]. The one patient in the non-genotyped arm found at study completion to be TPMT homozygous deficient developed severe, early onset neutropaenia. However overall, the study found no difference in the rates of AZA cessation due to ADRs between the TPMT genotyped arm (with recommended AZA dose reduction and avoidance in heterozygous and homozygous TPMT deficient patients, respectively) and the non-genotyped arm, and no increase in AZA cessation in TPMT heterozygous patients compared to wild-type patients [272].
Secondly, whilst the evidence and recommendations for TPMT homozygous deficient individuals are relatively clear, the optimal management strategy for heterozygous patients is less certain. Although overall they appear to be at a modest increased risk of myelosuppression [149, 268], complicating factors include the observation that only ~ 30–60 % of heterozygous patients do not tolerate full doses of AZA/6-MP [254, 257, 273] and the benefit: harm ratio attributable to different thiopurine starting doses for heterozygotes likely varies depending on the disease-specific necessity for rapid therapeutic action. A higher risk of myelosuppression with a higher starting dose in a heterozygote might be justifiable for treating malignancy, but not chronic, stable immunological disease.
Thirdly, TPMT can be analysed by phenotype or genotype and the screening test protocol remains incompletely standardised. Erythrocyte TPMT activity is predominantly offered to clinicians in the UK, but it can be affected by patient ethnicity, concurrent use of interacting drugs (e.g. mesalazine, sulfasalazine, allopurinol), allogeneic erythrocyte transfusions during the preceding 120 days, and in haematological malignancies, it can be affected by disease-related influences [274]. Whilst the overall genotype to phenotype test concordance is 98.4 % in healthy volunteers, it decreases to 86 % in the intermediate TPMT activity range, attributable to both non-genetic influences on TPMT activity, as described above, and to a lesser extent, novel mutations [275]. Therefore, neither test is 100 % sensitive to correctly identify TPMT deficiency, but research from a National Centre suggests that genotyping is more accurate and should be used as the primary test, in contrast to current UK practice [276].
Therefore, a pharmacogenetic association exists between TPMT and myelosuppression and there is strong evidence, affirmed by clinical guidelines, for avoiding thiopurine drugs or significantly reducing their dose in TPMT homozygous deficient patients, given their near universal experience of myelosuppression at conventional doses [254]. Further research is required to clarify optimal management for heterozygous patients. However, it is already cost-effective to routinely test TPMT status to identify homozygous deficient patients alone [274]. Pre-therapy TPMT testing is not a substitute for routine on-therapy blood test monitoring, given that several thiopurine ADRs are not associated with TPMT and the majority of myelosuppression cases are still not accounted for by TPMT variants [265]. Finally, in addition to TPMT testing, there is also a growing role for thiopurine metabolite level monitoring (e.g. 6-TGNs) to individualise thiopurine doses soon after starting treatment; prospective studies to evaluate this proactive approach are ongoing [277].
3.5 SLCO1B1 and Statin-Induced Muscle Toxicity
Statins are the most commonly prescribed class of medication worldwide [278] and are highly efficacious in the primary and secondary prevention of cardiovascular disease [1]. They reduce plasma low-density lipoprotein (LDL) cholesterol through competitive inhibition of 3-hydroxy-3-methylglutaryl-coenzyme A (HMG-CoA) reductase, the rate limiting enzyme in de novo cholesterol synthesis. This in turn leads to an upregulation of hepatic LDL receptors, increasing cholesterol influx into hepatocytes and reducing the plasma burden [279].
The currently licensed statins have a good safety profile, but carry a small risk of skeletal muscle toxicity [280]. The spectrum of muscle pathology varies from the most common manifestation of asymptomatic elevations in plasma CK level, to myopathies with pain and high plasma CK levels through to rhabdomyolysis with the potential sequelae of AKI and death. Alternatively, statin therapy can cause myalgias with no detectable plasma CK rise [21]. Depending on precise definitions, myopathy and rhabdomyolysis occur at frequencies of ~ 1/1000 and ~ 1/100,000, respectively [281], although this is modulated by other risk factors including higher statin dose, female gender, older age, low BMI, untreated hypothyroidism and other drug therapies, for example concomitant use of gemfibrozil [281].
The solute carrier organic anion transporter family member 1B1 (SLCO1B1) belongs to the superfamily of solute carrier (SLC) influx transporter genes and encodes the organic anion-transporting polypeptide 1B1 (OATP1B1) [282]. OATP1B1 is one of the most highly expressed influx transporters within the human liver [283]. It facilitates hepatic uptake of a variety of xenobiotic compounds and endogenous substances [284] and so affects the level of exposure of substrate drugs to intracellular hepatic DMEs [285].
Although the effects of statins on the off-target muscle tissue are incompletely defined at present [286], there exists a significant association between gene variants of SLCO1B1 and the risk of statin-induced muscle ADRs. A seminal statin GWAS used data from the Study of the Effectiveness of Additional Reductions in Cholesterol and Homocysteine (SEARCH) RCT in the UK; 85 cases of definite or incipient myopathy were contrasted with 90 controls [148]. Both the cases and controls for the GWAS had been prescribed 80 mg simvastatin daily. Only an intronic SNP variant, rs4363657, was strongly correlated with myopathy and further regional genetic analysis showed it to be in near complete linkage disequilibrium with the nonsynonymous SNP, rs4149056, in exon 6 (SLCO1B1*5; 521T > C; V174A). Further, a gene-dose relationship was demonstrated for rs4149056: the OR for myopathy in heterozygotes and homozygotes for the minor C allele was 4.5 (95 % CI 2.6–7.7) and 16.9 (95 % CI 4.7–61.1), respectively, when compared to the ancestral TT genotype. Overall, greater than 60 % of the myopathy cases in this study were attributable to the C variant [148]. The association with rs4149056 has been replicated [148, 287, 288] but the incidence of severe myopathy and the magnitude of correlation were lower in a second UK randomised trial population [289], attributable to the smaller 40 mg daily simvastatin dose used [148]. The rs4149056 variant has been subsequently associated with more mild, statin-induced muscle ADRs [290], reduced simvastatin adherence [290] and general intolerance to simvastatin defined as a composite endpoint of prescribing +/− mild biochemical changes [291]. The weight of evidence to date for rs4149056 is with simvastatin and the evidence with other statins is less compelling [287, 290, 292], suggesting that rs4149056 may represent a simvastatin-specific effect.
Mechanistically, rs4149056 may interfere with localisation of the transporter to the hepatic plasma membrane reducing its activity [284]. It is associated with higher statin, and especially simvastatin acid, plasma concentrations [293–295] that conceivably increase skeletal muscle drug exposure. However, the relationship between plasma simvastatin acid concentration and muscle toxicity is not straightforward. Clinically, current FDA guidance recommends against the 80 mg simvastatin dose unless a patient has tolerated the higher dose for over 12 months [296].
Overall, the rs4149056 variant is a plausible candidate for a predictive test to reduce simvastatin-induced skeletal muscle ADRs. Current guidance suggests that when initiating simvastatin therapy in CT or CC genotype patients, simvastatin 20 mg daily is selected rather than the normal 40 mg daily dose, possible routine CK surveillance is utilised and alternative statin therapy is commenced rather than increasing the dose of simvastatin if lipid goals are not reached. However, the effects of these recommendations on the incidence of simvastatin ADRs and adherence are currently unknown [281].
4 Outlook and Recommendations
The aspiration of pharmacogenetics is to individualise drug treatment to minimise harm and promote efficacy. Pre-therapy predictive genetic testing seeks to tailor therapy to reduce ADRs primarily through guiding drug or dose selection and has impacted positively upon clinical practice, notably with abacavir . Genetic screening may also find a role in identifying patients for whom regular biomarker surveillance may be indicated to minimise the incidence of severe ADRs. In addition to the direct patient benefit of reducing ADRs, there are at least 3 other potentially favourable spin-offs from understanding the pharmacogenetics of ADRs. Firstly, genetic-ADR associations provide novel insights that facilitate investigation into underlying pathological processes and the extrapolation of new knowledge regarding hypersensitivity reactions may have implications for cancer , autoimmune and infectious disease management. Secondly, the safety profile of new therapeutics may be improved through screening of drug candidates for affinity to high risk HLA alleles, for example HLA-B*57:01 and HLA-B*58:01 [71]. Thirdly, the beneficial side effects of some drugs have resulted in new therapeutic indications, for example with sidenafil (Viagra) and its fortuitous alleviation of erectile dysfunction. Pharmacogenetics has the potential to increase this ‘drug repositioning’ through identifying novel off target pharmacodynamic sites.
Abacavir has provided a blueprint for translational pharmacogenetics, but it has yet to be emulated. This is partly due to certain ‘favourable’ characteristics of AHS including: the high relative prevalence of AHS [76], the exclusivity of the association between HLA-B*57:01 and immunologically-mediated AHS, the reduction of false-positive clinical diagnoses mediated by the screening programme [78], the vocal patient lobby, and a physician community who were relatively amenable to changing their prescribing and clinical behaviour. It is also because there are multiple obstacles encountered when attempting translation . It is important to first understand these hurdles, and then to have a systematic approach to both developing the ADR-genotype evidence base and to implementing it in clinical practice [297].
Many ADRs are rare and some, such as the HSS, consist of varying constellations of non-specific features. As a result, international consortia using standardised definitions for these ADRs are advisable so patient samples of sufficient size with well demarcated phenotypes that are generalisable across ethnic groups can be pooled together. The ‘International Serious Advent Consortium’ and their ‘Phenotype Standardisation Project’ are both steps in the right direction [298]. These coordinated efforts are a prerequisite to reducing the risk of type I and type II errors in genetic association studies of rare and variable ADRs.
Pharmacogenetics has traditionally harnessed the candidate gene approach, whereby genes predicted to be relevant, typically through knowledge of a drug’s pharmacology, are selectively studied. However, this approach is limited to contemporary knowledge and so has largely been superseded by GWAS, which has no stipulation for a priori hypotheses [20] and can test at least 106 SNPs concurrently. However GWAS increases sample size requirements and data capture, increasing the complexity of study data management and statistical processes and potentiates the threat of selective publication reporting. Further, the lack of a preformed hypothesis augments the importance of confirming biological causality for GWAS putative associations.
Nevertheless, GWAS is a valuable asset: it can confirm in a ‘blinded’ fashion the results of previous candidate gene studies [20] and offer a novel foothold into the idiosyncratic processes of off-target ADRs. For polygenic ADRs, GWAS may detect new loci of individual small effect size and assess genotype-phenotype associations of larger haplotype signatures. The ‘1000 Genomes Project,’ which has recently described the genomes of 1092 individuals, is in turn increasing the resolution of GWAS [299]. The 1000 Genomes Project should additionally provide a baseline reference for normal human genetic variation, enable fine mapping of existing GWAS associations and aid discovery of new genetic associations, partly through its detailed identification of indels and larger deletions as well as contemporary SNPs [299]. In the near future, next generation sequencing technologies that provide high throughput whole genome capability will offer the pinnacle of DNA resolution whilst advances in our understanding of epigenetic imprinting and microRNA regulation promise new directions for the study of ADR pharmacogenetics . As genetic variation does not usually account for all of the inter-individual variation in drug response, incorporation of data from transcriptomics, metabolomics and proteomics may further improve predictive values [127].
After identification and validation of a statistically significant genetic association(s) for an ADR, several hurdles still bar adoption into clinical practice. Large, well-conducted prospective studies represent the gold standard to confirm clinical outcome benefit, although given the rarity of some ADRs these are not always practical. For other ADRs, genetic sub-studies of clinical trials and registries will likely offer the highest attainable level of evidence [300]. Subsequent pharmacoeconomic studies should base their analyses on this high quality data rather than expert opinion and retrospective data [301].
Logistical and knowledge barriers to the implementation of ADR pharmacogenetics also exist. On-demand genotyping, where the treating physician requests a specific pharmacogenetic test for a patient when seeking to prescribe a drug with a clinically established ADR-genotype association, relies on both a physician’s knowledge of pharmacogenetics and a system for following-up and acting on the pharmacogenetic test result. Robust and validated point-of-care genotyping tests may be necessary. An alternative proposed method is pre-emptive genotyping, where multiple relevant SNPs are routinely genotyped together and this genetic data is incorporated into a patient’s electronic medical record, with subsequent access by automated clinical decision support (CDS) algorithms to provide a clinically relevant alert regarding a potential drug-genotype interaction specific to the individual patient, at the point in time when the physician is seeking to prescribe the drug of interest. This approach provides the pharmacogenetic information at the most pertinent time and secondly, the CDS approach is likely better suited to keep up with our rapidly expanding understanding of ADR pharmacogenetics. However, the associated computational challenges are considerable [302].
Finally, a genetic test should be ethically acceptable to patients, clinicians and society. The emphasis of pharmacogenetics is for the beneficial personalisation of medicine, yet paradoxically the realisation of this goal requires not only very large international research collaborations but also active engagement with society as a whole. This is not least because genetic information harbours potential adverse implications, such as individual discrimination by insurance firms based on high risk genotype carriage and neglect of ethnic minorities by pharmaceuticals opting to segregate research initiatives to benefit the majority to maximise profit margins [303]. Open dialogue between patients, healthcare services, insurance providers, pharmaceuticals and the wider public is required to address these risks. If society chooses pharmacogenetics , it must safeguard against encroachment on the rights of individuals and minority groups. Ultimately, the widespread application of pharmacogenetics throughout clinical practice to ameliorate ADRs remains far off, but the examples in this chapter and the promises inherent in the new technologies foreshadow a future potential.
References
Roden DM, Johnson JA, Kimmel SE, Krauss RM, Medina MW, Shuldiner A, Wilke RA (2011) Cardiovascular pharmacogenomics. Circ Res 109(7):807–820. doi:10.1161/circresaha.110.230995
Trevelyan Square BL (2010) Search catalogue. http://www.ic.nhs.uk/searchcatalogue?productid=7930&q=general+pharmaceutical+services&topics=1%2fPrimary+care+services%2fCommunity+pharmacy+services&sort=Most+recent&size=10&page=1#top. Accessed 17 Dec 2012
Pirmohamed M, James S, Meakin S, Green C, Scott AK, Walley TJ, Farrar K, Park BK, Breckenridge AM (2004) Adverse drug reactions as cause of admission to hospital: prospective analysis of 18 820 patients. BMJ 329(7456):15–19. doi:10.1136/bmj.329.7456.15
Ahern F, Sahm LJ, Lynch D, McCarthy S (2014) Determining the frequency and preventability of adverse drug reaction-related admissions to an Irish University Hospital: a cross-sectional study. Emerg Med J 31(1):24–29. doi:10.1136/emermed-2012-201945
Conforti A, Costantini D, Zanetti F, Moretti U, Grezzana M, Leone R (2012) Adverse drug reactions in older patients: an Italian observational prospective hospital study. Drug Healthc Patient Saf 4:75–80. doi:10.2147/dhps.s29287
Hopf Y, Watson M, Williams D (2008) Adverse-drug-reaction related admissions to a hospital in Scotland. Pharm World Sci 30(6):854–862. doi:10.1007/s11096-008-9240-5
Miranda V, Fede A, Nobuo M, Ayres V, Giglio A, Miranda M, Riechelmann RP (2011) Adverse drug reactions and drug interactions as causes of hospital admission in oncology. J Pain Symptom Manage 42(3):342–353. doi:10.1016/j.jpainsymman.2010.11.014
Ruiter R, Visser LE, Rodenburg EM, Trifirò G, Ziere G, Stricker BH (2012) Adverse drug reaction-related hospitalizations in persons aged 55 years and over: a population-based study in the Netherlands. Drugs Aging 29(3):225–232. doi:10.2165/11599430-000000000-00000
World Health Organisation, WHO Collaborating Centre for International Drug Monitoring (2002) The importance of pharmacovigilance–safety monitoring of medicinal products. http://whqlibdoc.who.int/hq/2002/a75646.pdf. Accessed 13 March 2015
European Medicines Agency (1995) Clinical safety data management: definitions and standards for expedited reporting. London CPMP/ICH/377/95. http://www.ema.europa.eu/docs/en_GB/document_library/Scientific_guideline/2009/09/WC500002749.pdf. Accessed 13 March 2015
Gurwitz JH, Field TS, Avorn J, McCormick D, Jain S, Eckler M, Benser M, Edmondson AC, Bates DW (2000) Incidence and preventability of adverse drug events in nursing homes. Am J Med 109(2):87–94
National Coordinating Council for Medication Error Reporting and Prevention. What is a Medication Error? http://www.nccmerp.org/about-medication-errors. Accessed 13 March 2015
Edwards IR, Aronson JK (2000) Adverse drug reactions: definitions, diagnosis, and management. Lancet 356(9237):1255–1259. doi:10.1016/s0140-6736(00)02799-9
Bates DW, Cullen DJ, Laird N, Petersen LA, Small SD, Servi D, Laffel G, Sweitzer BJ, Shea BF, Hallisey R (1995) Incidence of adverse drug events and potential adverse drug events. Implications for prevention. ADE prevention study group. JAMA 274(1):29–34
Nebeker JR, Barach P, Samore MH (2004) Clarifying adverse drug events: a clinician’s guide to terminology, documentation, and reporting. Ann Intern Med 140(10):795–801
Davies EC, Green CF, Mottram DR, Pirmohamed M (2007) Adverse drug reactions in hospitals: a narrative review. Curr Drug Saf 2(1):79–87
Rawlins MD, Thompson JW (1977) Pathogenesis of adverse drug reactions. In: Davies DM (ed) Textbook of adverse drug reactions. Oxford University Press, Oxford, p. 10
Medicines, Healthcare products Regulatory Agency wmgu Adverse drug reactions. http://www.mhra.gov.uk/Safetyinformation/Reportingsafetyproblems/Reportingsuspectedadversedrugreactions/Healthcareprofessionalreporting/Adversedrugreactions/index.htm#1. Accessed 18 Dec 2012
Aronson JK, Ferner RE (2003) Joining the DoTS: new approach to classifying adverse drug reactions. BMJ 327(7425):1222–1225. doi:10.1136/bmj.327.7425.1222
Evrard A, Mbatchi L (2012) Genetic polymorphisms of drug metabolizing enzymes and transporters: the long way from bench to bedside. Curr Top Med Chem 12(15):1720–1729
Pirmohamed M (2010) Pharmacogenetics of idiosyncratic adverse drug reactions. Handb Exp Pharmacol 196:477–491. doi:10.1007/978-3-642-00663-0_17
Mallal S, Nolan D, Witt C, Masel G, Martin AM, Moore C, Sayer D, Castley A, Mamotte C, Maxwell D, James I, Christiansen FT (2002) Association between presence of HLA-B*5701, HLA-DR7, and HLA-DQ3 and hypersensitivity to HIV-1 reverse-transcriptase inhibitor abacavir. Lancet 359(9308):727–732
Hetherington S, Hughes AR, Mosteller M, Shortino D, Baker KL, Spreen W, Lai E, Davies K, Handley A, Dow DJ, Fling ME, Stocum M, Bowman C, Thurmond LM, Roses AD (2002) Genetic variations in HLA-B region and hypersensitivity reactions to abacavir. Lancet 359(9312):1121–1122. doi:10.1016/s0140-6736(02)08158-8
Hung S-I, Chung W-H, Liou L-B, Chu C-C, Lin M, Huang H-P, Lin Y-L, Lan J-L, Yang L-C, Hong H-S, Chen M-J, Lai P-C, Wu M-S, Chu C-Y, Wang K-H, Chen C-H, Fann CSJ, Wu J-Y, Chen Y-T (2005) HLA-B*5801 allele as a genetic marker for severe cutaneous adverse reactions caused by allopurinol. Proc Natl Acad Sci U S A 102(11):4134–4139. doi:10.1073/pnas.0409500102
Kang H-R, Jee YK, Kim Y-S, Lee CH, Jung J-W, Kim SH, Park H-W, Chang Y-S, Jang I-J, Cho S-H, Min K-U, Kim S-H, Lee KW (2011) Positive and negative associations of HLA class I alleles with allopurinol-induced SCARs in Koreans. Pharmacogenet Genomics 21(5):303–307. doi:10.1097/FPC.0b013e32834282b8
Kim S-H, Lee KW, Song W-J, Kim S-H, Jee Y-K, Lee S-M, Kang H-R, Park H-W, Cho S-H, Park S-H, Min K-U, Chang Y-S (2011) Carbamazepine-induced severe cutaneous adverse reactions and HLA genotypes in Koreans. Epilepsy Res 97(1–2):190–197. doi:10.1016/j.eplepsyres.2011.08.010
McCormack M, Alfirevic A, Bourgeois S, Farrell JJ, Kasperavičiūtė D, Carrington M, Sills GJ, Marson T, Jia X, de Bakker PIW, Chinthapalli K, Molokhia M, Johnson MR, O'Connor GD, Chaila E, Alhusaini S, Shianna KV, Radtke RA, Heinzen EL, Walley N, Pandolfo M, Pichler W, Park BK, Depondt C, Sisodiya SM, Goldstein DB, Deloukas P, Delanty N, Cavalleri GL, Pirmohamed M (2011) HLA-A*3101 and carbamazepine-induced hypersensitivity reactions in Europeans. N Engl J Med 364(12):1134–1143. doi:10.1056/NEJMoa1013297
Littera R, Carcassi C, Masala A, Piano P, Serra P, Ortu F, Corso N, Casula B, Nasa G L, Contu L, Manconi PE (2006) HLA-dependent hypersensitivity to nevirapine in Sardinian HIV patients. AIDS 20(12):1621–1626. doi:10.1097/01.aids.0000238408.82947.09
Gatanaga H, Yazaki H, Tanuma J, Honda M, Genka I, Teruya K, Tachikawa N, Kikuchi Y, Oka S (2007) HLA-Cw8 primarily associated with hypersensitivity to nevirapine. AIDS 21(2):264–265. doi:10.1097/QAD.0b013e32801199d9
Chantarangsu S, Mushiroda T, Mahasirimongkol S, Kiertiburanakul S, Sungkanuparph S, Manosuthi W, Tantisiriwat W, Charoenyingwattana A, Sura T, Chantratita W, Nakamura Y (2009) HLA-B*3505 allele is a strong predictor for nevirapine-induced skin adverse drug reactions in HIV-infected Thai patients. Pharmacogenet Genomics 19(2):139–146. doi:10.1097/FPC.0b013e32831d0faf
Tassaneeyakul W, Jantararoungtong T, Chen P, Lin P-Y, Tiamkao S, Khunarkornsiri U, Chucherd P, Konyoung P, Vannaprasaht S, Choonhakarn C, Pisuttimarn P, Sangviroon A, Tassaneeyakul W (2009) Strong association between HLA-B*5801 and allopurinol-induced Stevens-Johnson syndrome and toxic epidermal necrolysis in a Thai population. Pharmacogenet Genomics 19(9):704–709. doi:10.1097/FPC.0b013e328330a3b8
Chung W-H, Hung S-I, Hong H-S, Hsih M-S, Yang L-C, Ho H-C, Wu J-Y, Chen Y-T (2004) Medical genetics: a marker for Stevens-Johnson syndrome. Nature 428(6982):486. doi:10.1038/428486a
Locharernkul C, Loplumlert J, Limotai C, Korkij W, Desudchit T, Tongkobpetch S, Kangwanshiratada O, Hirankarn N, Suphapeetiporn K, Shotelersuk V (2008) Carbamazepine and phenytoin induced Stevens-Johnson syndrome is associated with HLA-B*1502 allele in Thai population. Epilepsia 49(12):2087–2091. doi:10.1111/j.1528-1167.2008.01719.x
Yip VL, Marson AG, Jorgensen AL, Pirmohamed M, Alfirevic A (2012) HLA genotype and carbamazepine-induced cutaneous adverse drug reactions: a systematic review. Clin Pharmacol Ther 92(6):757–765. doi:10.1038/clpt.2012.189
Lonjou C, Borot N, Sekula P, Ledger N, Thomas L, Halevy S, Naldi L, Bouwes-Bavinck J-N, Sidoroff A, de Toma C, Schumacher M, Roujeau J-C, Hovnanian A, Mockenhaupt M (2008) A European study of HLA-B in Stevens-Johnson syndrome and toxic epidermal necrolysis related to five high-risk drugs. Pharmacogenet Genomics 18(2):99–107. doi:10.1097/FPC.0b013e3282f3ef9c
Kim S-H, Kim M, Lee KW, Kim S-H, Kang H-R, Park H-W, Jee Y-K (2010) HLA-B*5901 is strongly associated with methazolamide-induced Stevens-Johnson syndrome/toxic epidermal necrolysis. Pharmacogenomics 11(6):879–884. doi:10.2217/pgs.10.54
Carr DF, Chaponda M, Jorgensen AL, Castro EC, van Oosterhout JJ, Khoo SH, Lalloo DG, Heyderman RS, Alfirevic A, Pirmohamed M (2013) Association of human leukocyte antigen alleles and nevirapine hypersensitivity in a Malawian HIV-infected population. Clin Infect Dis 56(9):1330–1339. doi:10.1093/cid/cit021
Hung S-I, Chung W-H, Liu Z-S, Chen C-H, Hsih M-S, Hui RC-y, Chu C-Y, Chen Y-T (2010) Common risk allele in aromatic antiepileptic-drug induced Stevens-Johnson syndrome and toxic epidermal necrolysis in Han Chinese. Pharmacogenomics 11(3):349–356. doi:10.2217/pgs.09.162
Cao Z-h, Wei Z-y, Zhu Q-y, Zhang J-y, Yang L, Qin S-y, Shao L-y, Zhang Y-t, Xuan J-k, Q-l L, Xu J-H, Xu F, Ma L, Huang H-y, Xing Q-h, Luo X-q (2012) HLA-B*58:01 allele is associated with augmented risk for both mild and severe cutaneous adverse reactions induced by allopurinol in Han Chinese. Pharmacogenomics 13(10):1193–1201. doi:10.2217/pgs.12.89
Romano A, De Santis A, Romito A, Fonso M D, Venuti A, Gasbarrini GB, Manna R (1998) Delayed hypersensitivity to aminopenicillins is related to major histocompatibility complex genes. Ann Allergy Asthma Immunol 80(5):433–437
Hung S-I, Chung W-H, Jee S-H, Chen W-C, Chang Y-T, Lee W-R, Hu S-L, Wu M-T, Chen G-S, Wong T-W, Hsiao P-F, Chen W-H, Shih H-Y, Fang W-H, Wei C-Y, Lou Y-H, Huang Y-L, Lin J-J, Chen Y-T (2006) Genetic susceptibility to carbamazepine-induced cutaneous adverse drug reactions. Pharmacogenet Genomics 16(4):297–306. doi:10.1097/01.fpc.0000199500.46842.4a
Vitezica ZG, Milpied B, Lonjou C, Borot N, Ledger TN, Lefebvre A, Hovnanian A (2008) HLA-DRB1*01 associated with cutaneous hypersensitivity induced by nevirapine and efavirenz. AIDS 22(4):540–541. doi:10.1097/QAD.0b013e3282f37812
Likanonsakul S, Rattanatham T, Feangvad S, Uttayamakul S, Prasithsirikul W, Tunthanathip P, Nakayama EE, Shioda T (2009) HLA-Cw*04 allele associated with nevirapine-induced rash in HIV-infected Thai patients. AIDS Res Ther 6:22. doi:10.1186/1742-6405-6-22
Sharma SK, Balamurugan A, Saha PK, Pandey RM, Mehra NK (2002) Evaluation of clinical and immunogenetic risk factors for the development of hepatotoxicity during antituberculosis treatment. Am J Respir Crit Care Med 166(7):916–919
Lucena MI, Molokhia M, Shen Y, Urban TJ, Aithal GP, Andrade RJ, Day CP, Ruiz-Cabello F, Donaldson PT, Stephens C, Pirmohamed M, Romero-Gomez M, Navarro JM, Fontana RJ, Miller M, Groome M, Bondon-Guitton E, Conforti A, Stricker BHC, Carvajal A, Ibanez L, Yue Q-Y, Eichelbaum M, Floratos A, Pe'er I, Daly MJ, Goldstein DB, Dillon JF, Nelson MR, Watkins PB, Daly AK (2011) Susceptibility to amoxicillin-clavulanate-induced liver injury is influenced by multiple HLA class I and II alleles. Gastroenterology 141(1):338–347. doi:10.1053/j.gastro.2011.04.001
O’Donohue J, Oien KA, Donaldson P, Underhill J, Clare M, MacSween RN, Mills PR (2000) Co-amoxiclav jaundice: clinical and histological features and HLA class II association. Gut 47(5):717–720
Daly AK, Donaldson PT, Bhatnagar P, Shen Y, Pe’er I, Floratos A, Daly MJ, Goldstein DB, John S, Nelson MR, Graham J, Park BK, Dillon JF, Bernal W, Cordell HJ, Pirmohamed M, Aithal GP, Day CP (2009) HLA-B*5701 genotype is a major determinant of drug-induced liver injury due to flucloxacillin. Nat Genet 41(7):816–819. doi:10.1038/ng.379
Spraggs CF, Budde LR, Briley LP, Bing N, Cox CJ, King KS, Whittaker JC, Mooser VE, Preston AJ, Stein SH, Cardon LR (2011) HLA-DQA1*02:01 is a major risk factor for lapatinib-induced hepatotoxicity in women with advanced breast cancer. J Clin Oncol 29(6):667–673. doi:10.1200/jco.2010.31.3197
Singer JB, Lewitzky S, Leroy E, Yang F, Zhao X, Klickstein L, Wright TM, Meyer J, Paulding CA (2010) A genome-wide study identifies HLA alleles associated with lumiracoxib-related liver injury. Nat Genet 42(8):711–714. doi:10.1038/ng.632
Yuan J, Guo S, Hall D, Cammett AM, Jayadev S, Distel M, Storfer S, Huang Z, Mootsikapun P, Ruxrungtham K, Podzamczer D, Haas DW (2011) Toxicogenomics of nevirapine-associated cutaneous and hepatic adverse events among populations of African, Asian, and European descent. AIDS 25(10):1271–1280. doi:10.1097/QAD.0b013e32834779df
Hirata K, Takagi H, Yamamoto M, Matsumoto T, Nishiya T, Mori K, Shimizu S, Masumoto H, Okutani Y (2008) Ticlopidine-induced hepatotoxicity is associated with specific human leukocyte antigen genomic subtypes in Japanese patients: a preliminary case-control study. Pharmacogenomics J 8(1):29–33. doi:10.1038/sj.tpj.6500442
Ariyoshi N, Iga Y, Hirata K, Sato Y, Miura G, Ishii I, Nagamori S, Kitada M (2010) Enhanced susceptibility of HLA-mediated ticlopidine-induced idiosyncratic hepatotoxicity by CYP2B6 polymorphism in Japanese. Drug Metab Pharmacokinet 25(3):298–306
Kindmark A, Jawaid A, Harbron CG, Barratt BJ, Bengtsson OF, Andersson TB, Carlsson S, Cederbrant KE, Gibson NJ, Armstrong M, Lagerström-Fermér ME, Dellsén A, Brown EM, Thornton M, Dukes C, Jenkins SC, Firth MA, Harrod GO, Pinel TH, Billing-Clason SME, Cardon LR, March RE (2008) Genome-wide pharmacogenetic investigation of a hepatic adverse event without clinical signs of immunopathology suggests an underlying immune pathogenesis. Pharmacogenomics J 8(3):186–195. doi:10.1038/sj.tpj.6500458
Athanasiou MC, Dettling M, Cascorbi I, Mosyagin I, Salisbury BA, Pierz KA, Zou W, Whalen H, Malhotra AK, Lencz T, Gerson SL, Kane JM, Reed CR (2011) Candidate gene analysis identifies a polymorphism in HLA-DQB1 associated with clozapine-induced agranulocytosis. J Clin Psychiatry 72(4):458–463. doi:10.4088/JCP.09m05527yel
Diez RA (1990) HLA-B27 and agranulocytosis by levamisole. Immunol Today 11(8):270
Lee HY, Lee JW, Lee KW, Park MH, Park HS (2009) The HLA allele marker for differentiating ASA hypersensitivity phenotypes. Allergy 64(9):1385–1387. doi:10.1111/j.1398-9995.2009.02048.x
Kim S-H, Hur G-Y, Choi J-H, Park H-S (2008) Pharmacogenetics of aspirin-intolerant asthma. Pharmacogenomics 9(1):85–91. doi:10.2217/14622416.9.1.85
Hakala M, van Assendelft AH, Ilonen J, Jalava S, Tiilikainen A (1986) Association of different HLA antigens with various toxic effects of gold salts in rheumatoid arthritis. Ann Rheum Dis 45(3):177–182
Shah P, Griffith SM, Shadforth MF, Fisher J, Dawes PT, Poulton KV, Thomson W, Ollier WER, Mattey DL (2004) Can gold therapy be used more safely in rheumatoid arthritis? Adverse drug reactions are more likely in patients with nodular disease, independent of HLA-DR3 status. J Rheumatol 31(10):1903–1905
Erlich H (2012) HLA DNA typing: past, present, and future. Tissue Antigens 80(1):1–11. doi:10.1111/j.1399-0039.2012.01881.x
Horton R, Wilming L, Rand V, Lovering RC, Bruford EA, Khodiyar VK, Lush MJ, Povey S, Talbot CC Jr, Wright MW, Wain HM, Trowsdale J, Ziegler A, Beck S (2004) Gene map of the extended human MHC. Nat Rev Genet 5(12):889–899. doi:10.1038/nrg1489
Bharadwaj M, Illing P, Theodossis A, Purcell AW, Rossjohn J, McCluskey J (2012) Drug hypersensitivity and human leukocyte antigens of the major histocompatibility complex. Annu Rev Pharmacol Toxicol 52:401–431. doi:10.1146/annurev-pharmtox-010611-134701
Joffre OP, Segura E, Savina A, Amigorena S (2012) Cross-presentation by dendritic cells. Nat Rev Immunol 12(8):557–569. doi:10.1038/nri3254
Reche PA, Reinherz EL (2003) Sequence variability analysis of human class I and class II MHC molecules: functional and structural correlates of amino acid polymorphisms. J Mol Biol 331(3):623–641
Illing PT, Vivian JP, Purcell AW, Rossjohn J, McCluskey J (2013) Human leukocyte antigen-associated drug hypersensitivity. Curr Opin Immunol 25(1):81–89. doi:10.1016/j.coi.2012.10.002
Kurkó J, Besenyei T, Laki J, Glant TT, Mikecz K, Szekanecz Z (2013) Genetics of rheumatoid arthritis—a comprehensive review. Clin Rev Allergy Immunol 45(2):170-179. doi:10.1007/s12016-012-8346-7
Hetherington S, McGuirk S, Powell G, Cutrell A, Naderer O, Spreen B, Lafon S, Pearce G, Steel H (2001) Hypersensitivity reactions during therapy with the nucleoside reverse transcriptase inhibitor abacavir. Clin Ther 23(10):1603–1614
Pichler W, Yawalkar N, Schmid S, Helbling A (2002) Pathogenesis of drug-induced exanthems. Allergy 57(10):884–893
Nassif A, Bensussan A, Boumsell L, Deniaud A, Moslehi H, Wolkenstein P, Bagot M, Roujeau J-C (2004) Toxic epidermal necrolysis: effector cells are drug-specific cytotoxic T cells. J Allergy Clin Immunol 114(5):1209–1215. doi:10.1016/j.jaci.2004.07.047
Phillips EJ, Sullivan JR, Knowles SR, Shear NH (2002) Utility of patch testing in patients with hypersensitivity syndromes associated with abacavir. AIDS 16(16):2223–2225
Alfirevic A, Park BK, Pirmohamed M, Naisbitt DJ (2012) Research highlights: explanation for HLA-B*57:01-linked immune-mediated abacavir-induced hypersensitivity. Pharmacogenomics 13(14):1567–1569. doi:10.2217/pgs.12.146
Park BK, Naisbitt DJ, Gordon SF, Kitteringham NR, Pirmohamed M (2001) Metabolic activation in drug allergies. Toxicology 158(1–2):11–23
Pichler WJ, Beeler A, Keller M, Lerch M, Posadas S, Schmid D, Spanou Z, Zawodniak A, Gerber B (2006) Pharmacological interaction of drugs with immune receptors: the p-i concept. Allergol Int 55(1):17–25. doi:10.2332/allergolint.55.17
Pirmohamed M, Naisbitt DJ, Gordon F, Park BK (2002) The danger hypothesis-potential role in idiosyncratic drug reactions. Toxicology 181-182:55–63
Alfirevic A, Pirmohamed M (2010) Drug-induced hypersensitivity reactions and pharmacogenomics: past, present and future. Pharmacogenomics 11(4):497–499. doi:10.2217/pgs.10.12
Chaponda M, Pirmohamed M (2011) Hypersensitivity reactions to HIV therapy. Br J Clin Pharmacol 71(5):659–671. doi:10.1111/j.1365-2125.2010.03784.x
Mallal S, Phillips E, Carosi G, Molina J-M, Workman C, Tomazic J, Jägel-Guedes E, Rugina S, Kozyrev O, Cid JF, Hay P, Nolan D, Hughes S, Hughes A, Ryan S, Fitch N, Thorborn D, Benbow A (2008) HLA-B*5701 screening for hypersensitivity to abacavir. N Engl J Med 358(6):568–579. doi:10.1056/NEJMoa0706135
Phillips EJ, Mallal SA (2010) Pharmacogenetics of drug hypersensitivity. Pharmacogenomics 11(7):973–987. doi:10.2217/pgs.10.77
Hughes AR, Mosteller M, Bansal AT, Davies K, Haneline SA, Lai EH, Nangle K, Scott T, Spreen WR, Warren LL, Roses AD (2004) Association of genetic variations in HLA-B region with hypersensitivity to abacavir in some, but not all, populations. Pharmacogenomics 5(2):203–211. doi:10.1517/phgs.5.2.203.27481
Hughes DA, Vilar FJ, Ward CC, Alfirevic A, Park BK, Pirmohamed M (2004) Cost-effectiveness analysis of HLA B*5701 genotyping in preventing abacavir hypersensitivity. Pharmacogenetics 14(6):335–342
Saag M, Balu R, Phillips E, Brachman P, Martorell C, Burman W, Stancil B, Mosteller M, Brothers C, Wannamaker P, Hughes A, Sutherland-Phillips D, Mallal S, Shaefer M (2008) High sensitivity of human leukocyte antigen-b*5701 as a marker for immunologically confirmed abacavir hypersensitivity in white and black patients. Clin Infect Dis 46(7):1111–1118. doi:10.1086/529382
Dervieux T, Bala MV (2006) Overview of the pharmacoeconomics of pharmacogenetics. Pharmacogenomics 7(8):1175–1184. doi:10.2217/14622416.7.8.1175
Schackman BR, Scott CA, Walensky RP, Losina E, Freedberg KA, Sax PE (2008) The cost-effectiveness of HLA-B*5701 genetic screening to guide initial antiretroviral therapy for HIV. AIDS 22(15):2025–2033. doi:10.1097/QAD.0b013e3283103ce6
Lalonde RG, Thomas R, Rachlis A, Gill MJ, Roger M, Angel JB, Smith G, Higgins N, Trottier B (2010) Successful implementation of a national HLA-B*5701 genetic testing service in Canada. Tissue Antigens 75(1):12–18. doi:10.1111/j.1399-0039.2009.01383.x
Rauch A, Nolan D, Martin A, McKinnon E, Almeida C, Mallal S (2006) Prospective genetic screening decreases the incidence of abacavir hypersensitivity reactions in the Western Australian HIV cohort study. Clin Infect Dis 43(1):99–102. doi:10.1086/504874
Waters LJ, Mandalia S, Gazzard B, Nelson M (2007) Prospective HLA-B*5701 screening and abacavir hypersensitivity: a single centre experience. AIDS 21(18):2533–2534. doi:10.1097/QAD.0b013e328273bc07
Zucman D, Truchis Pd, Majerholc C, Stegman S, Caillat-Zucman S (2007) Prospective screening for human leukocyte antigen-B*5701 avoids abacavir hypersensitivity reaction in the ethnically mixed French HIV population. J Acquir Immune Defic Syndr 45(1):1–3. doi:10.1097/QAI.0b013e318046ea31
Illing PT, Vivian JP, Dudek NL, Kostenko L, Chen Z, Bharadwaj M, Miles JJ, Kjer-Nielsen L, Gras S, Williamson NA, Burrows SR, Purcell AW, Rossjohn J, McCluskey J (2012) Immune self-reactivity triggered by drug-modified HLA-peptide repertoire. Nature 486(7404):554–558. doi:10.1038/nature11147
Ostrov DA, Grant BJ, Pompeu YA, Sidney J, Harndahl M, Southwood S, Oseroff C, Lu S, Jakoncic J, de Oliveira CAF, Yang L, Mei H, Shi L, Shabanowitz J, English AM, Wriston A, Lucas A, Phillips E, Mallal S, Grey HM, Sette A, Hunt DF, Buus S, Peters B (2012) Drug hypersensitivity caused by alteration of the MHC-presented self-peptide repertoire. Proc Natl Acad Sci U S A 109(25):9959–9964. doi:10.1073/pnas.1207934109
Norcross MA, Luo S, Lu L, Boyne MT, Gomarteli M, Rennels AD, Woodcock J, Margulies DH, McMurtrey C, Vernon S, Hildebrand WH, Buchli R (2012) Abacavir induces loading of novel self-peptides into HLA-B*57: 01: an autoimmune model for HLA-associated drug hypersensitivity. AIDS 26(11):F21–29. doi:10.1097/QAD.0b013e328355fe8f
Adam J, Eriksson KK, Schnyder B, Fontana S, Pichler WJ, Yerly D (2012) Avidity determines T-cell reactivity in abacavir hypersensitivity. Eur J Immunol 42(7):1706–1716. doi:10.1002/eji.201142159
Picard D, Janela B, Descamps V, D'Incan M, Courville P, Jacquot S, Rogez S, Mardivirin L, Moins-Teisserenc H, Toubert A, Benichou J, Joly P, Musette P (2010) Drug reaction with eosinophilia and systemic symptoms (DRESS): a multiorgan antiviral T cell response. Sci Transl Med 2(46):46ra62. doi:10.1126/scitranslmed.3001116
Marson AG, Al-Kharusi AM, Alwaidh M, Appleton R, Baker GA, Chadwick DW, Cramp C, Cockerell OC, Cooper PN, Doughty J, Eaton B, Gamble C, Goulding PJ, Howell SJL, Hughes A, Jackson M, Jacoby A, Kellett M, Lawson GR, Leach JP, Nicolaides P, Roberts R, Shackley P, Shen J, Smith DF, Smith PEM, Smith CT, Vanoli A, Williamson PR (2007) The SANAD study of effectiveness of carbamazepine, gabapentin, lamotrigine, oxcarbazepine, or topiramate for treatment of partial epilepsy: an unblinded randomised controlled trial. Lancet 369(9566):1000–1015. doi:10.1016/s0140-6736(07)60460-7
Pirmohamed M, Friedmann PS, Molokhia M, Loke YK, Smith C, Phillips E, Grenade L L, Carleton B, Papaluca-Amati M, Demoly P, Shear NH (2011) Phenotype standardization for immune-mediated drug-induced skin injury. Clin Pharmacol Ther 89(6):896–901. doi:10.1038/clpt.2011.79
Roujeau JC, Stern RS (1994) Severe adverse cutaneous reactions to drugs. N Engl J Med 331(19):1272–1285. doi:10.1056/nejm199411103311906
Profaizer T, Eckels D (2012) HLA alleles and drug hypersensitivity reactions. Int J Immunogenet 39(2):99–105. doi:10.1111/j.1744-313X.2011.01061.x
Criado PR, Avancini J, Santi CG, Medrado ATA, Rodrigues CE, de Carvalho JF (2012) Drug reaction with eosinophilia and systemic symptoms (DRESS): a complex interaction of drugs, viruses and the immune system. Isr Med Assoc J 14(9):577–582
Borchers AT, Lee JL, Naguwa SM, Cheema GS, Gershwin ME (2008) Stevens-Johnson syndrome and toxic epidermal necrolysis. Autoimmun Rev 7(8):598–605. doi:10.1016/j.autrev.2008.06.004
Bastuji-Garin S, Rzany B, Stern RS, Shear NH, Naldi L, Roujeau JC (1993) Clinical classification of cases of toxic epidermal necrolysis, Stevens-Johnson syndrome, and erythema multiforme. Arch Dermatol 129(1):92–96
Ding WY, Lee CK, Choon SE (2010) Cutaneous adverse drug reactions seen in a tertiary hospital in Johor, Malaysia. Int J Dermatol 49(7):834–841. doi:10.1111/j.1365-4632.2010.04481.x
Kulkantrakorn K, Tassaneeyakul W, Tiamkao S, Jantararoungtong T, Prabmechai N, Vannaprasaht S, Chumworathayi P, Chen P, Sritipsukho P (2012) HLA-B*1502 strongly predicts carbamazepine-induced Stevens-Johnson syndrome and toxic epidermal necrolysis in Thai patients with neuropathic pain. Pain Pract 12(3):202–208. doi:10.1111/j.1533-2500.2011.00479.x
Mehta TY, Prajapati LM, Mittal B, Joshi CG, Sheth JJ, Patel DB, Dave DM, Goyal RK (2009) Association of HLA-B*1502 allele and carbamazepine-induced Stevens-Johnson syndrome among Indians. Indian J Dermatol Venereol Leprol 75(6):579–582. doi:10.4103/0378-6323.57718
Chen P, Lin J-J, Lu C-S, Ong C-T, Hsieh PF, Yang C-C, Tai C-T, Wu S-L, Lu C-H, Hsu Y-C, Yu H-Y, Ro L-S, Lu C-T, Chu C-C, Tsai J-J, Su Y-H, Lan S-H, Sung S-F, Lin S-Y, Chuang H-P, Huang L-C, Chen Y-J, Tsai P-J, Liao H-T, Lin Y-H, Chen C-H, Chung W-H, Hung S-I, Wu J-Y, Chang C-F, Chen L, Chen Y-T, Shen C-Y (2011) Carbamazepine-induced toxic effects and HLA-B*1502 screening in Taiwan. N Engl J Med 364(12):1126–1133. doi:10.1056/NEJMoa1009717
Alfirevic A, Jorgensen AL, Williamson PR, Chadwick DW, Park BK, Pirmohamed M (2006) HLA-B locus in Caucasian patients with carbamazepine hypersensitivity. Pharmacogenomics 7(6):813–818. doi:10.2217/14622416.7.6.813
Lonjou C, Thomas L, Borot N, Ledger N, de Toma C, LeLouet H, Graf E, Schumacher M, Hovnanian A, Mockenhaupt M, Roujeau JC (2006) A marker for Stevens-Johnson syndrome…: ethnicity matters. Pharmacogenomics J 6(4):265–268. doi:10.1038/sj.tpj.6500356
Ikeda H, Takahashi Y, Yamazaki E, Fujiwara T, Kaniwa N, Saito Y, Aihara M, Kashiwagi M, Muramatsu M (2010) HLA class I markers in Japanese patients with carbamazepine-induced cutaneous adverse reactions. Epilepsia 51(2):297–300. doi:10.1111/j.1528-1167.2009.02269.x
Ozeki T, Mushiroda T, Yowang A, Takahashi A, Kubo M, Shirakata Y, Ikezawa Z, Iijima M, Shiohara T, Hashimoto K, Kamatani N, Nakamura Y (2011) Genome-wide association study identifies HLA-A*3101 allele as a genetic risk factor for carbamazepine-induced cutaneous adverse drug reactions in Japanese population. Hum Mol Genet 20(5):1034–1041. doi:10.1093/hmg/ddq537
Kaniwa N, Saito Y, Aihara M, Matsunaga K, Tohkin M, Kurose K, Furuya H, Takahashi Y, Muramatsu M, Kinoshita S, Abe M, Ikeda H, Kashiwagi M, Song Y, Ueta M, Sotozono C, Ikezawa Z, Hasegawa R (2010) HLA-B*1511 is a risk factor for carbamazepine-induced Stevens-Johnson syndrome and toxic epidermal necrolysis in Japanese patients. Epilepsia 51(12):2461–2465. doi:10.1111/j.1528-1167.2010.02766.x
Yun J, Adam J, Yerly D, Pichler WJ (2012) Human leukocyte antigens (HLA) associated drug hypersensitivity: consequences of drug binding to HLA. Allergy 67(11):1338–1346. doi:10.1111/all.12008
Wei C-Y, Chung W-H, Huang H-W, Chen Y-T, Hung S-I (2012) Direct interaction between HLA-B and carbamazepine activates T cells in patients with Stevens-Johnson syndrome. J Allergy Clin Immunol 129(6):1562–1569.e1565. doi:10.1016/j.jaci.2011.12.990
Ko T-M, Chung W-H, Wei C-Y, Shih H-Y, Chen J-K, Lin C-H, Chen Y-T, Hung S-I (2011) Shared and restricted T-cell receptor use is crucial for carbamazepine-induced Stevens-Johnson syndrome. J Allergy Clin Immunol 128(6):1266–1276.e11. doi:10.1016/j.jaci.2011.08.013
Hershfield MS, Callaghan JT, Tassaneeyakul W, Mushiroda T, Thorn CF, Klein TE, Lee MTM (2013) Clinical Pharmacogenetics Implementation Consortium guidelines for human leukocyte antigen-B genotype and allopurinol dosing. Clin Pharmacol Ther 93(2):153–158. doi:10.1038/clpt.2012.209
Halevy S, Ghislain P-D, Mockenhaupt M, Fagot J-P, Bouwes Bavinck JN, Sidoroff A, Naldi L, Dunant A, Viboud C, Roujeau J-C (2008) Allopurinol is the most common cause of Stevens-Johnson syndrome and toxic epidermal necrolysis in Europe and Israel. J Am Acad Dermatol 58(1):25–32. doi:10.1016/j.jaad.2007.08.036
Cacoub P, Musette P, Descamps V, Meyer O, Speirs C, Finzi L, Roujeau JC (2011) The DRESS syndrome: a literature review. Am J Med 124(7):588–597. doi:10.1016/j.amjmed.2011.01.017
Kaniwa N, Saito Y, Aihara M, Matsunaga K, Tohkin M, Kurose K, Sawada J-i, Furuya H, Takahashi Y, Muramatsu M, Kinoshita S, Abe M, Ikeda H, Kashiwagi M, Song Y, Ueta M, Sotozono C, Ikezawa Z, Hasegawa R (2008) HLA-B locus in Japanese patients with anti-epileptics and allopurinol-related Stevens-Johnson syndrome and toxic epidermal necrolysis. Pharmacogenomics 9(11):1617–1622. doi:10.2217/14622416.9.11.1617
Somkrua R, Eickman EE, Saokaew S, Lohitnavy M, Chaiyakunapruk N (2011) Association of HLA-B*5801 allele and allopurinol-induced Stevens Johnson syndrome and toxic epidermal necrolysis: a systematic review and meta-analysis. BMC Med Genet 12:118. doi:10.1186/1471-2350-12-118
Lee MH, Stocker SL, Anderson J, Phillips EJ, Nolan D, Williams KM, Graham GG, Sullivan JR, Day RO (2012) Initiating allopurinol therapy: do we need to know the patient’s human leucocyte antigen status? Intern Med J 42(4):411–416. doi:10.1111/j.1445-5994.2011.02567.x
Shiohara T, Inaoka M, Kano Y (2006) Drug-induced hypersensitivity syndrome (DIHS): a reaction induced by a complex interplay among herpesviruses and antiviral and antidrug immune responses. Allergol Int 55(1):1–8. doi:10.2332/allergolint.55.1
Chao J, Terkeltaub R (2009) A critical reappraisal of allopurinol dosing, safety, and efficacy for hyperuricemia in gout. Curr Rheumatol Rep 11(2):135–140
Russmann S, Kaye JA, Jick SS, Jick H (2005) Risk of cholestatic liver disease associated with flucloxacillin and flucloxacillin prescribing habits in the UK: cohort study using data from the UK general practice research database. Br J Clin Pharmacol 60(1):76–82. doi:10.1111/j.1365-2125.2005.02370.x
Daly AK, Day CP (2012) Genetic association studies in drug-induced liver injury. Drug Metab Rev 44(1):116–126. doi:10.3109/03602532.2011.605790
Ostapowicz G, Fontana RJ, Schiødt FV, Larson A, Davern TJ, Han SHB, McCashland TM, Shakil AO, Hay JE, Hynan L, Crippin JS, Blei AT, Samuel G, Reisch J, Lee WM (2002) Results of a prospective study of acute liver failure at 17 tertiary care centers in the United States. Ann Intern Med 137(12):947–954
Russo MW, Galanko JA, Shrestha R, Fried MW, Watkins P (2004) Liver transplantation for acute liver failure from drug induced liver injury in the United States. Liver Transpl 10(8):1018–1023. doi:10.1002/lt.20204
Sgro C, Clinard F, Ouazir K, Chanay H, Allard C, Guilleminet C, Lenoir C, Lemoine A, Hillon P (2002) Incidence of drug-induced hepatic injuries: a French population-based study. Hepatology 36(2):451–455. doi:10.1053/jhep.2002.34857
Aithal GP, Watkins PB, Andrade RJ, Larrey D, Molokhia M, Takikawa H, Hunt CM, Wilke RA, Avigan M, Kaplowitz N, Bjornsson E, Daly AK (2011) Case definition and phenotype standardization in drug-induced liver injury. Clin Pharmacol Ther 89(6):806–815. doi:10.1038/clpt.2011.58
Uetrecht J (2009) Immunoallergic drug-induced liver injury in humans. Semin Liver Dis 29(4):383–392. doi:10.1055/s-0029-1240007
Alfirevic A, Pirmohamed M (2012) Predictive genetic testing for drug-induced liver injury: considerations of clinical utility. Clin Pharmacol Ther 92(3):376–380. doi:10.1038/clpt.2012.107
Jenkins RE, Meng X, Elliott VL, Kitteringham NR, Pirmohamed M, Park BK (2009) Characterisation of flucloxacillin and 5-hydroxymethyl flucloxacillin haptenated HSA in vitro and in vivo. Proteomics Clin Appl 3(6):720–729. doi:10.1002/prca.200800222
Monshi M, Faulkner L, Gibson A, Jenkins RE, Farrell J, Earnshaw CJ, Alfirevic A, Cederbrant K, Daly AK, French N, Pirmohamed M, Park BK, Naisbitt DJ (2013) HLA-B*57:01-restricted activation of drug-specific T-cells provides the immunological basis for flucloxacillin-induced liver injury. Hepatology 57(2):727–739. doi:10.1002/hep.26077
Li Y, Tang HL, Hu YF, Xie HG (2012) The gain-of-function variant allele CYP2C19*17: a double-edged sword between thrombosis and bleeding in clopidogrel-treated patients. J Thromb Haemost 10(2):199–206. doi:10.1111/j.1538-7836.2011.04570.x
Zabalza M, Subirana I, Sala J, Lluis-Ganella C, Lucas G, Tomás M, Masiá R, Marrugat J, Brugada R, Elosua R (2012) Meta-analyses of the association between cytochrome CYP2C19 loss- and gain-of-function polymorphisms and cardiovascular outcomes in patients with coronary artery disease treated with clopidogrel. Heart 98(2):100–108. doi:10.1136/hrt.2011.227652
Higashi MK, Veenstra DL, Kondo LM, Wittkowsky AK, Srinouanprachanh SL, Farin FM, Rettie AE (2002) Association between CYP2C9 genetic variants and anticoagulation-related outcomes during warfarin therapy. JAMA 287(13):1690–1698
Aithal GP, Day CP, Kesteven PJ, Daly AK (1999) Association of polymorphisms in the cytochrome P450 CYP2C9 with warfarin dose requirement and risk of bleeding complications. Lancet 353(9154):717–719. doi:10.1016/s0140-6736(98)04474-2
Limdi NA, McGwin G, Goldstein JA, Beasley TM, Arnett DK, Adler BK, Baird MF, Acton RT (2008) Influence of CYP2C9 and VKORC1 1173C/T genotype on the risk of hemorrhagic complications in African-American and European-American patients on warfarin. Clin Pharmacol Ther 83(2):312–321. doi:10.1038/sj.clpt.6100290
Yang J, Chen Y, Li X, Wei X, Chen X, Zhang L, Zhang Y, Xu Q, Wang H, Li Y, Lu C, Chen W, Zeng C, Yin T (2013) Influence of CYP2C9 and VKORC1 genotypes on the risk of hemorrhagic complications in warfarin-treated patients: a systematic review and meta-analysis. Int J Cardiol 168(4):4234–4243. doi:10.1016/j.ijcard.2013.07.151
Gasche Y, Daali Y, Fathi M, Chiappe A, Cottini S, Dayer P, Desmeules J (2004) Codeine intoxication associated with ultrarapid CYP2D6 metabolism. N Engl J Med 351(27):2827–2831. doi:10.1056/NEJMoa041888
Crews KR, Gaedigk A, Dunnenberger HM, Klein TE, Shen DD, Callaghan JT, Kharasch ED, Skaar TC (2012) Clinical Pharmacogenetics Implementation Consortium (CPIC) guidelines for codeine therapy in the context of cytochrome P450 2D6 (CYP2D6) genotype. Clin Pharmacol Ther 91(2):321–326. doi:10.1038/clpt.2011.287
Kirchheiner J, Keulen J-THA, Bauer S, Roots I, Brockmöller J (2008) Effects of the CYP2D6 gene duplication on the pharmacokinetics and pharmacodynamics of tramadol. J Clin Psychopharmacol 28(1):78–83. doi:10.1097/JCP.0b013e318160f827
Stamer UM, Stüber F, Muders T, Musshoff F (2008) Respiratory depression with tramadol in a patient with renal impairment and CYP2D6 gene duplication. Anesth Analg 107(3):926–929. doi:10.1213/ane.0b013e31817b796e
Cai Y, Yi J, Zhou C, Shen X (2012) Pharmacogenetic study of drug-metabolising enzyme polymorphisms on the risk of anti-tuberculosis drug-induced liver injury: a meta-analysis. PLoS ONE 7(10):e47769. doi:10.1371/journal.pone.0047769
Huang Y-S, Chern H-D, Su W-J, Wu J-C, Chang S-C, Chiang C-H, Chang F-Y, Lee S-D (2003) Cytochrome P450 2E1 genotype and the susceptibility to antituberculosis drug-induced hepatitis. Hepatology 37(4):924–930. doi:10.1053/jhep.2003.50144
Roy B, Chowdhury A, Kundu S, Santra A, Dey B, Chakraborty M, Majumder PP (2001) Increased risk of antituberculosis drug-induced hepatotoxicity in individuals with glutathione S-transferase M1 ‘null’ mutation. J Gastroenterol Hepatol 16(9):1033–1037
Daly AK, Aithal GP, Leathart JBS, Swainsbury RA, Dang TS, Day CP (2007) Genetic susceptibility to diclofenac-induced hepatotoxicity: contribution of UGT2B7, CYP2C8, and ABCC2 genotypes. Gastroenterology 132(1):272–281. doi:10.1053/j.gastro.2006.11.023
Simon T, Becquemont L, Mary-Krause M, de Waziers I, Beaune P, Funck-Brentano C, Jaillon P (2000) Combined glutathione-S-transferase M1 and T1 genetic polymorphism and tacrine hepatotoxicity. Clin Pharmacol Ther 67(4):432–437. doi:10.1067/mcp.2000.104944
Watanabe I, Tomita A, Shimizu M, Sugawara M, Yasumo H, Koishi R, Takahashi T, Miyoshi K, Nakamura K, Izumi T, Matsushita Y, Furukawa H, Haruyama H, Koga T (2003) A study to survey susceptible genetic factors responsible for troglitazone-associated hepatotoxicity in Japanese patients with type 2 diabetes mellitus. Clin Pharmacol Ther 73(5):435–455
Iyer L, King CD, Whitington PF, Green MD, Roy SK, Tephly TR, Coffman BL, Ratain MJ (1998) Genetic predisposition to the metabolism of irinotecan (CPT-11). Role of uridine diphosphate glucuronosyltransferase isoform 1A1 in the glucuronidation of its active metabolite (SN-38) in human liver microsomes. J Clin Invest 101(4):847–854. doi:10.1172/jci915
de Leon J, Susce MT, Pan R-M, Fairchild M, Koch WH, Wedlund PJ (2005) The CYP2D6 poor metabolizer phenotype may be associated with risperidone adverse drug reactions and discontinuation. J Clin Psychiatry 66(1):15–27
Link E, Parish S, Armitage J, Bowman L, Heath S, Matsuda F, Gut I, Lathrop M, Collins R (2008) SLCO1B1 variants and statin-induced myopathy-a genomewide study. N Engl J Med 359(8):789–799. doi:10.1056/NEJMoa0801936
Booth RA, Ansari MT, Loit E, Tricco AC, Weeks L, Doucette S, Skidmore B, Sears M, Sy R, Karsh J (2011) Assessment of thiopurine S-methyltransferase activity in patients prescribed thiopurines: a systematic review. Ann Intern Med 154(12):814–823. doi:10.1059/0003-4819-154-12-201106210-00009W-295-298
Musumba CO, Jorgensen A, Sutton L, Van Eker D, Zhang E, O'Hara N, Carr DF, Pritchard DM, Pirmohamed M (2013) CYP2C19*17 gain-of-function polymorphism is associated with peptic ulcer disease. Clin Pharmacol Ther 93(2):195–203. doi:10.1038/clpt.2012.215
Yen T, Nightingale BN, Burns JC, Sullivan DR, Stewart PM (2003) Butyrylcholinesterase (BCHE) genotyping for post-succinylcholine apnea in an Australian population. Clin Chem 49(8):1297–1308
Mega JL, Simon T, Collet J-P, Anderson JL, Antman EM, Bliden K, Cannon CP, Danchin N, Giusti B, Gurbel P, Horne BD, Hulot J-S, Kastrati A, Montalescot G, Neumann F-J, Shen L, Sibbing D, Steg PG, Trenk D, Wiviott SD, Sabatine MS (2010) Reduced-function CYP2C19 genotype and risk of adverse clinical outcomes among patients treated with clopidogrel predominantly for PCI: a meta-analysis. JAMA 304(16):1821–1830. doi:10.1001/jama.2010.1543
Amstutz U, Froehlich TK, Largiadèr CR (2011) Dihydropyrimidine dehydrogenase gene as a major predictor of severe 5-fluorouracil toxicity. Pharmacogenomics 12(9):1321–1336. doi:10.2217/pgs.11.72
Dávila-Fajardo CL, Swen JJ, Cabeza Barrera J, Guchelaar H-J (2013) Genetic risk factors for drug-induced liver injury in rheumatoid arthritis patients using low-dose methotrexate. Pharmacogenomics 14(1):63–73. doi:10.2217/pgs.12.183
European Malignant Hyperthermia Group: Home. http://www.emhg.org/. Accessed 8 Feb 2013
Robinson R, Carpenter D, Shaw M-A, Halsall J, Hopkins P (2006) Mutations in RYR1 in malignant hyperthermia and central core disease. Hum Mutat 27(10):977–989. doi:10.1002/humu.20356
Monnier N, Procaccio V, Stieglitz P, Lunardi J (1997) Malignant-hyperthermia susceptibility is associated with a mutation of the alpha 1-subunit of the human dihydropyridine-sensitive L-type voltage-dependent calcium-channel receptor in skeletal muscle. Am J Hum Genet 60(6):1316–1325
Toppin PJ, Chandy TT, Ghanekar A, Kraeva N, Beattie WS, Riazi S (2010) A report of fulminant malignant hyperthermia in a patient with a novel mutation of the CACNA1S gene. Can J Anaesth 57(7):689–693. doi:10.1007/s12630-010-9314-4
Mulder H, Cohen D, Scheffer H, Gispen-de Wied C, Arends J, Wilmink FW, Franke B, Egberts ACG (2009) HTR2C gene polymorphisms and the metabolic syndrome in patients with schizophrenia: a replication study. J Clin Psychopharmacol 29(1):16–20. doi:10.1097/JCP.0b013e3181934462
Minucci A, Moradkhani K, Hwang MJ, Zuppi C, Giardina B, Capoluongo E (2012) Glucose-6-phosphate dehydrogenase (G6PD) mutations database: review of the "old" and update of the new mutations. Blood Cells Mol Dis 48(3):154–165. doi:10.1016/j.bcmd.2012.01.001
Youngster I, Arcavi L, Schechmaster R, Akayzen Y, Popliski H, Shimonov J, Beig S, Berkovitch M (2010) Medications and glucose-6-phosphate dehydrogenase deficiency: an evidence-based review. Drug Saf 33(9):713–726. doi:10.2165/11536520-000000000-00000
Jennings BA, Kwok CS, Willis G, Matthews V, Wawruch P, Loke YK (2012) Functional polymorphisms of folate metabolism and response to chemotherapy for colorectal cancer, a systematic review and meta-analysis. Pharmacogenet Genomics 22(4):290–304. doi:10.1097/FPC.0b013e328351875d
Denborough MA, Forster JF, Lovell RR, Maplestone PA, Villiers JD (1962) Anaesthetic deaths in a family. Br J Anaesth 34:395–396
Larard DG, Rice CP, Robinson R, Spencer RW, Westhead RA (1972) Malignant hyperthermia: a study of an affected family. Br J Anaesth 44(1):93–96
Matos AR, Sambuughin N, Rumjanek FD, Amoedo ND, Cunha LBP, Zapata-Sudo G, Sudo RT (2009) Multigenerational Brazilian family with malignant hyperthermia and a novel mutation in the RYR1 gene. Braz J Med Biol Res 42(12):1218–1224
Strazis KP, Fox AW (1993) Malignant hyperthermia: a review of published cases. Anesth Analg 77(2):297–304
Hopkins PM (2011) Malignant hyperthermia: pharmacology of triggering. Br J Anaesth 107(1):48–56. doi:10.1093/bja/aer132
Dexter F, Epstein RH, Wachtel RE, Rosenberg H (2013) Estimate of the relative risk of succinylcholine for triggering malignant hyperthermia. Anesth Analg 116(1):118–122. doi:10.1213/ANE.0b013e31826f5e3b
Denborough M (1998) Malignant hyperthermia. Lancet 352(9134):1131–1136. doi:10.1016/s0140-6736(98)03078-5
Groom L, Muldoon SM, Tang ZZ, Brandom BW, Bayarsaikhan M, Bina S, Lee H-S, Qiu X, Sambuughin N, Dirksen RT (2011) Identical de novo mutation in the type 1 ryanodine receptor gene associated with fatal, stress-induced malignant hyperthermia in two unrelated families. Anesthesiology 115(5):938–945. doi:10.1097/ALN.0b013e3182320068
Gronert GA, Thompson RL, Onofrio BM (1980) Human malignant hyperthermia: awake episodes and correction by dantrolene. Anesth Analg 59(5):377–378
Correia ACdC, Silva PCB, da Silva BA (2012) Malignant hyperthermia: clinical and molecular aspects. Rev Bras Anestesiol 62(6):820–837. doi:10.1016/s0034-7094(12)70182-4
Rosenberg H, Davis M, James D, Pollock N, Stowell K (2007) Malignant hyperthermia. Orphanet J Rare Dis 2:21. doi:10.1186/1750-1172-2-21
Sinkovich DD, Mitch-Resignalo AE (1991) Malignant hyperthermia. Orthop Nurs 10(1):39–43
Larach MG, Gronert GA, Allen GC, Brandom BW, Lehman EB (2010) Clinical presentation, treatment, and complications of malignant hyperthermia in North America from 1987-2006. Anesth Analg 110(2):498–507. doi:10.1213/ANE.0b013e3181c6b9b2
Bandschapp O, Girard T (2012) Malignant hyperthermia. Swiss Med Wkly 142:w13652. doi:10.4414/smw.2012.13652
Larach MG, Localio AR, Allen GC, Denborough MA, Ellis FR, Gronert GA, Kaplan RF, Muldoon SM, Nelson TE, Ording H (1994) A clinical grading scale to predict malignant hyperthermia susceptibility. Anesthesiology 80(4):771–779
Lanner JT, Georgiou DK, Joshi AD, Hamilton SL (2010) Ryanodine receptors: structure, expression, molecular details, and function in calcium release. Cold Spring Harb Perspect Biol 2(11):a003996. doi:10.1101/cshperspect.a003996
Van Petegem F (2012) Ryanodine receptors: structure and function. J Biol Chem 287(38):31624–31632. doi:10.1074/jbc.R112.349068
Takeshima H, Nishimura S, Matsumoto T, Ishida H, Kangawa K, Minamino N, Matsuo H, Ueda M, Hanaoka M, Hirose T (1989) Primary structure and expression from complementary DNA of skeletal muscle ryanodine receptor. Nature 339(6224):439–445. doi:10.1038/339439a0
Block BA, Imagawa T, Campbell KP, Franzini-Armstrong C (1988) Structural evidence for direct interaction between the molecular components of the transverse tubule/sarcoplasmic reticulum junction in skeletal muscle. J Cell Biol 107(6 Pt 2):2587–2600
Inui M, Saito A, Fleischer S (1987) Purification of the ryanodine receptor and identity with feet structures of junctional terminal cisternae of sarcoplasmic reticulum from fast skeletal muscle. J Biol Chem 262(4):1740–1747
Meissner G, Lu X (1995) Dihydropyridine receptor-ryanodine receptor interactions in skeletal muscle excitation-contraction coupling. Biosci Rep 15(5):399–408
Fujii J, Otsu K, Zorzato F, de Leon S, Khanna VK, Weiler JE, O'Brien PJ, MacLennan DH (1991) Identification of a mutation in porcine ryanodine receptor associated with malignant hyperthermia. Science 253(5018):448–451
Gillard EF, Otsu K, Fujii J, Khanna VK, de Leon S, Derdemezi J, Britt BA, Duff CL, Worton RG, MacLennan DH (1991) A substitution of cysteine for arginine 614 in the ryanodine receptor is potentially causative of human malignant hyperthermia. Genomics 11(3):751–755
Fill M, Stefani E, Nelson TE (1991) Abnormal human sarcoplasmic reticulum Ca2 + release channels in malignant hyperthermic skeletal muscle. Biophys J 59(5):1085–1090. doi:10.1016/s0006-3495(91)82323-2
Ohta T, Endo M, Nakano T, Morohoshi Y, Wanikawa K, Ohga A (1989) Ca-induced Ca release in malignant hyperthermia-susceptible pig skeletal muscle. Am J Physiol Cell Physiol 256(2):C358–C367
Kim D-C (2012) Malignant hyperthermia. Korean J Anesthesiol 63(5):391–401. doi:10.4097/kjae.2012.63.5.391
Klingler W, Rueffert H, Lehmann-Horn F, Girard T, Hopkins PM (2009) Core myopathies and risk of malignant hyperthermia. Anesth Analg 109(4):1167–1173. doi:10.1213/ANE.0b013e3181b5ae2d
Isaacs H, Badenhorst ME (1992) Dominantly inherited malignant hyperthermia (MH) in the King-Denborough syndrome. Muscle Nerve 15(6):740–742. doi:10.1002/mus.880150619
D’Arcy CE, Bjorksten A, Yiu EM, Bankier A, Gillies R, McLean CA, Shield LK, Ryan MM (2008) King-denborough syndrome caused by a novel mutation in the ryanodine receptor gene. Neurology 71(10):776–777. doi:10.1212/01.wnl.0000324929.33780.2f
Hirshey Dirksen SJ, Larach MG, Rosenberg H, Brandom BW, Parness J, Lang RS, Gangadharan M, Pezalski T (2011) Special article: future directions in malignant hyperthermia research and patient care. Anesth Analg 113(5):1108–1119. doi:10.1213/ANE.0b013e318222af2e
Capacchione JF, Sambuughin N, Bina S, Mulligan LP, Lawson TD, Muldoon SM (2010) Exertional rhabdomyolysis and malignant hyperthermia in a patient with ryanodine receptor type 1 gene, L-type calcium channel alpha-1 subunit gene, and calsequestrin-1 gene polymorphisms. Anesthesiology 112(1):239–244. doi:10.1097/ALN.0b013e3181c29504
Wappler F, Fiege M, Steinfath M, Agarwal K, Scholz J, Singh S, Matschke J, Schulte Am Esch J (2001) Evidence for susceptibility to malignant hyperthermia in patients with exercise-induced rhabdomyolysis. Anesthesiology 94(1):95–100
(1984) A protocol for the investigation of malignant hyperpyrexia (MH) susceptibility. The European Malignant Hyperpyrexia Group. Br J Anaesth 56 (11):1267–1269
Urwyler A, Deufel T, McCarthy T, West S (2001) Guidelines for molecular genetic detection of susceptibility to malignant hyperthermia. Br J Anaesth 86(2):283–287
Ramachandran S, Chakraborty A, Xu L, Mei Y, Samso M, Dokholyan NV, Meissner G (2013) Structural determinants of skeletal muscle ryanodine receptor gating. J Biol Chem 288(9):6154–6165. doi:10.1074/jbc.M112.433789
Dewire SM, Yamashita DS, Rominger DH, Liu G, Cowan CL, Graczyk TM, Chen X-T, Pitis PM, Gotchev D, Yuan C, Koblish M, Lark MW, Violin JD (2013) A G protein-biased ligand at the mu-opioid receptor is potently analgesic with reduced gastrointestinal and respiratory dysfunction compared to morphine. J Pharmacol Exp Ther 344(3):708–717. doi:10.1124/jpet.112.201616
Kieffer BL (1999) Opioids: first lessons from knockout mice. Trends Pharmacol Sci 20(1):19–26
Mignat C, Wille U, Ziegler A (1995) Affinity profiles of morphine, codeine, dihydrocodeine and their glucuronides at opioid receptor subtypes. Life Sci 56(10):793–799
Eissing T, Lippert J, Willmann S (2012) Pharmacogenomics of codeine, morphine, and morphine-6-glucuronide: model-based analysis of the influence of CYP2D6 activity, UGT2B7 activity, renal impairment, and CYP3A4 inhibition. Mol Diagn Ther 16(1):43–53. doi:10.2165/11597930-000000000-00000
Lee CR, Goldstein JA, Pieper JA (2002) Cytochrome P450 2C9 polymorphisms: a comprehensive review of the in-vitro and human data. Pharmacogenetics 12(3):251–263
Guengerich FP (2008) Cytochrome p450 and chemical toxicology. Chem Res Toxicol 21(1):70–83. doi:10.1021/tx700079z
Sim SC, Ingelman-Sundberg M (2010) The Human Cytochrome P450 (CYP) Allele Nomenclature website: a peer-reviewed database of CYP variants and their associated effects. Hum Genomics 4(4):278–281. doi:10.1186/1479-7364-4-4-278
Gaedigk A, Simon SD, Pearce RE, Bradford LD, Kennedy MJ, Leeder JS (2008) The CYP2D6 activity score: translating genotype information into a qualitative measure of phenotype. Clin Pharmacol Ther 83(2):234–242. doi:10.1038/sj.clpt.6100406
Eckhardt K, Li S, Ammon S, Schänzle G, Mikus G, Eichelbaum M (1998) Same incidence of adverse drug events after codeine administration irrespective of the genetically determined differences in morphine formation. Pain 76(1–2):27–33
Sindrup SH, Brøsen K, Bjerring P, Arendt-Nielsen L, Larsen U, Angelo HR, Gram LF (1990) Codeine increases pain thresholds to copper vapor laser stimuli in extensive but not poor metabolizers of sparteine. Clin Pharmacol Ther 48(6):686–693
Kirchheiner J, Schmidt H, Tzvetkov M, Keulen JTHA, Lötsch J, Roots I, Brockmöller J (2007) Pharmacokinetics of codeine and its metabolite morphine in ultra-rapid metabolizers due to CYP2D6 duplication. Pharmacogenomics J 7(4):257–265. doi:10.1038/sj.tpj.6500406
Koren G, Cairns J, Chitayat D, Gaedigk A, Leeder SJ (2006) Pharmacogenetics of morphine poisoning in a breastfed neonate of a codeine-prescribed mother. Lancet 368 (9536):704. doi:10.1016/s0140-6736(06)69255-6
Kelly LE, Rieder M, van den Anker J, Malkin B, Ross C, Neely MN, Carleton B, Hayden MR, Madadi P, Koren G (2012) More codeine fatalities after tonsillectomy in North American children. Pediatrics 129(5):e1343–e1347. doi:10.1542/peds.2011-2538
Voronov P, Przybylo HJ, Jagannathan N (2007) Apnea in a child after oral codeine: a genetic variant—an ultra-rapid metabolizer. Paediatr Anaesth 17(7):684–687. doi:10.1111/j.1460-9592.2006.02182.x
Ciszkowski C, Madadi P, Phillips MS, Lauwers AE, Koren G (2009) Codeine, ultrarapid-metabolism genotype, and postoperative death. N Engl J Med 361(8):827–828. doi:10.1056/NEJMc0904266
Dalén P, Frengell C, Dahl ML, Sjöqvist F (1997) Quick onset of severe abdominal pain after codeine in an ultrarapid metabolizer of debrisoquine. Ther Drug Monit 19(5):543–544
Food and Drug Administration (2007) FDA Warning on Codeine Use by Nursing Mothers. http://www.fda.gov/NewsEvents/Newsroom/PressAnnouncements/2007/ucm108968.htm. Accessed 26 Jan 2013
Niesters M, Overdyk F, Smith T, Aarts L, Dahan A (2013) Opioid-induced respiratory depression in paediatrics: a review of case reports. Br J Anaesth 110(2):175–182. doi:10.1093/bja/aes447
Tong TF, Ng KK (2001) Codeine ingestion and apparent life-threatening event in a neonate. Pediatr Int 43(5):517–518
Brown KA, Laferrière A, Lakheeram I, Moss IR (2006) Recurrent hypoxemia in children is associated with increased analgesic sensitivity to opiates. Anesthesiology 105(4):665–669
Stamer UM, Lehnen K, Höthker F, Bayerer B, Wolf S, Hoeft A, Stuber F (2003) Impact of CYP2D6 genotype on postoperative tramadol analgesia. Pain 105(1–2):231–238
Prommer E, Ficek B (2012) Management of pain in the elderly at the end of life. Drugs Aging 29(4):285–305. doi:10.2165/11599210-000000000-00000
Johnson JA, Gong L, Whirl-Carrillo M, Gage BF, Scott SA, Stein CM, Anderson JL, Kimmel SE, Lee MTM, Pirmohamed M, Wadelius M, Klein TE, Altman RB (2011) Clinical pharmacogenetics implementation consortium guidelines for CYP2C9 and VKORC1 genotypes and warfarin dosing. Clin Pharmacol Ther 90(4):625–629. doi:10.1038/clpt.2011.185
Hirsh J, Fuster V, Ansell J, Halperin JL (2003) American heart association/American college of cardiology foundation guide to warfarin therapy. Circulation 107(12):1692–1711. doi:10.1161/01.cir.0000063575.17904.4e
Choonara IA, Haynes BP, Cholerton S, Breckenridge AM, Park BK (1986) Enantiomers of warfarin and vitamin K1 metabolism. Br J Clin Pharmacol 22(6):729–732
Stenflo J, Fernlund P, Egan W, Roepstorff P (1974) Vitamin K dependent modifications of glutamic acid residues in prothrombin. Proc Natl Acad Sci U S A 71(7):2730–2733
Wallin R, Hutson SM (2004) Warfarin and the vitamin K-dependent gamma-carboxylation system. Trends Mol Med 10(7):299–302. doi:10.1016/j.molmed.2004.05.003
Ansell J, Hirsh J, Hylek E, Jacobson A, Crowther M, Palareti G (2008) Pharmacology and management of the vitamin K antagonists: American college of chest physicians evidence-based clinical practice guidelines (8th Ed). Chest 133(6 Suppl):160S–198S. doi:10.1378/chest.08-0670
Baglin TP, Keeling DM, Watson HG (2006) Guidelines on oral anticoagulation (warfarin): third edition-2005 update. Br J Haematol 132(3):277–285. doi:10.1111/j.1365-2141.2005.05856.x
Lövborg H, Eriksson LR, Jönsson AK, Bradley T, Hägg S (2012) A prospective analysis of the preventability of adverse drug reactions reported in Sweden. Eur J Clin Pharmacol 68(8):1183–1189. doi:10.1007/s00228-012-1237-2
Gage BF, Lesko LJ (2008) Pharmacogenetics of warfarin: regulatory, scientific, and clinical issues. J Thromb Thrombolysis 25(1):45–51. doi:10.1007/s11239-007-0104-y
Connolly SJ, Pogue J, Eikelboom J, Flaker G, Commerford P, Franzosi MG, Healey JS, Yusuf S (2008) Benefit of oral anticoagulant over antiplatelet therapy in atrial fibrillation depends on the quality of international normalized ratio control achieved by centers and countries as measured by time in therapeutic range. Circulation 118(20):2029–2037. doi:10.1161/circulationaha.107.750000
Reynolds MW, Fahrbach K, Hauch O, Wygant G, Estok R, Cella C, Nalysnyk L (2004) Warfarin anticoagulation and outcomes in patients with atrial fibrillation: a systematic review and metaanalysis. Chest 126(6):1938–1945. doi:10.1378/chest.126.6.1938
Owen RP, Gong L, Sagreiya H, Klein TE, Altman RB (2010) VKORC1 pharmacogenomics summary. Pharmacogenet Genomics 20(10):642–644. doi:10.1097/FPC.0b013e32833433b6
Eriksson N, Wadelius M (2012) Prediction of warfarin dose: why, when and how? Pharmacogenomics 13(4):429–440. doi:10.2217/pgs.11.184
Avery PJ, Jorgensen A, Hamberg AK, Wadelius M, Pirmohamed M, Kamali F (2011) A proposal for an individualized pharmacogenetics-based warfarin initiation dose regimen for patients commencing anticoagulation therapy. Clin Pharmacol Ther 90(5):701–706. doi:10.1038/clpt.2011.186
Linder MW, Bon Homme M, Reynolds KK, Gage BF, Eby C, Silvestrov N, Valdes R Jr (2009) Interactive modeling for ongoing utility of pharmacogenetic diagnostic testing: application for warfarin therapy. Clin Chem 55(10):1861–1868. doi:10.1373/clinchem.2009.125898
Jorgensen AL, FitzGerald RJ, Oyee J, Pirmohamed M, Williamson PR (2012) Influence of CYP2C9 and VKORC1 on patient response to warfarin: a systematic review and meta-analysis. PLoS ONE 7(8):e44064. doi:10.1371/journal.pone.0044064
Patel AA, Swerlick RA, McCall CO (2006) Azathioprine in dermatology: the past, the present, and the future. J Am Acad Dermatol 55(3):369–389. doi:10.1016/j.jaad.2005.07.059
Gervasini G, Vagace JM (2012) Impact of genetic polymorphisms on chemotherapy toxicity in childhood acute lymphoblastic leukemia. Front Genet 3:249. doi:10.3389/fgene.2012.00249
Manz M, Vavricka SR, Wanner R, Lakatos PL, Rogler G, Frei P, Safroneeva E, Schoepfer AM (2012) Therapy of steroid-resistant inflammatory bowel disease. Digestion 86(Suppl 1):11–15. doi:10.1159/000341952
Roberts RL, Barclay ML (2012) Current relevance of pharmacogenetics in immunomodulation treatment for Crohn’s disease. J Gastroenterol Hepatol 27(10):1546–1554. doi:10.1111/j.1440-1746.2012.07220.x
Ford LT, Berg JD (2010) Thiopurine S-methyltransferase (TPMT) assessment prior to starting thiopurine drug treatment; a pharmacogenomic test whose time has come. J Clin Pathol 63(4):288–295. doi:10.1136/jcp.2009.069252
Gurwitz D, Rodríguez-Antona C, Payne K, Newman W, Gisbert JP, de Mesa EG, Ibarreta D (2009) Improving pharmacovigilance in Europe: TPMT genotyping and phenotyping in the UK and Spain. Eur J Hum Genet 17(8):991–998. doi:10.1038/ejhg.2009.10
Bradford K, Shih DQ (2011) Optimizing 6-mercaptopurine and azathioprine therapy in the management of inflammatory bowel disease. World J Gastroenterol 17(37):4166–4173. doi:10.3748/wjg.v17.i37.4166
Sahasranaman S, Howard D, Roy S (2008) Clinical pharmacology and pharmacogenetics of thiopurines. Eur J Clin Pharmacol 64(8):753–767. doi:10.1007/s00228-008-0478-6
Lennard L (1992) The clinical pharmacology of 6-mercaptopurine. Eur J Clin Pharmacol 43(4):329–339
Kurowski V, Iven H (1991) Plasma concentrations and organ distribution of thiopurines after oral application of azathioprine in mice. Cancer Chemother Pharmacol 28(1):7–14
Tiede I, Fritz G, Strand S, Poppe D, Dvorsky R, Strand D, Lehr HA, Wirtz S, Becker C, Atreya R, Mudter J, Hildner K, Bartsch B, Holtmann M, Blumberg R, Walczak H, Iven H, Galle PR, Ahmadian MR, Neurath MF (2003) CD28-dependent Rac1 activation is the molecular target of azathioprine in primary human CD4+ T lymphocytes. J Clin Invest 111(8):1133–1145. doi:10.1172/jci16432
Osterman MT, Kundu R, Lichtenstein GR, Lewis JD (2006) Association of 6-thioguanine nucleotide levels and inflammatory bowel disease activity: a meta-analysis. Gastroenterology 130(4):1047–1053. doi:10.1053/j.gastro.2006.01.046
Dubinsky MC (2004) Azathioprine, 6-mercaptopurine in inflammatory bowel disease: pharmacology, efficacy, and safety. Clin Gastroenterol Hepatol 2(9):731–743
Cuffari C, Hunt S, Bayless T (2001) Utilisation of erythrocyte 6-thioguanine metabolite levels to optimise azathioprine therapy in patients with inflammatory bowel disease. Gut 48(5):642–646. doi:10.1136/gut.48.5.642
Bökkerink JP, Stet EH, De Abreu RA, Damen FJ, Hulscher TW, Bakker MA, van Baal JA (1993) 6-Mercaptopurine: cytotoxicity and biochemical pharmacology in human malignant T-lymphoblasts. Biochem Pharmacol 45(7):1455–1463
Adam de Beaumais T, Fakhoury M, Medard Y, Azougagh S, Zhang D, Yakouben K, Jacqz-Aigrain E (2011) Determinants of mercaptopurine toxicity in paediatric acute lymphoblastic leukemia maintenance therapy. Br J Clin Pharmacol 71(4):575–584. doi:10.1111/j.1365-2125.2010.03867.x
Dubinsky MC, Lamothe S, Yang HY, Targan SR, Sinnett D, Théorêt Y, Seidman EG (2000) Pharmacogenomics and metabolite measurement for 6-mercaptopurine therapy in inflammatory bowel disease. Gastroenterology 118(4):705–713
Nygaard U, Toft N, Schmiegelow K (2004) Methylated metabolites of 6-mercaptopurine are associated with hepatotoxicity. Clin Pharmacol Ther 75(4):274–281. doi:10.1016/j.clpt.2003.12.001
Relling MV, Gardner EE, Sandborn WJ, Schmiegelow K, Pui CH, Yee SW, Stein CM, Carrillo M, Evans WE, Klein TE (2011) Clinical Pharmacogenetics Implementation Consortium guidelines for thiopurine methyltransferase genotype and thiopurine dosing. Clin Pharmacol Ther 89(3):387–391. doi:10.1038/clpt.2010.320
van Asseldonk DP, Seinen ML, de Boer NKH, van Bodegraven AA, Mulder CJ (2012) Hepatotoxicity associated with 6-methyl mercaptopurine formation during azathioprine and 6-mercaptopurine therapy does not occur on the short-term during 6-thioguanine therapy in IBD treatment. J Crohns Colitis 6(1):95–101. doi:10.1016/j.crohns.2011.07.009
Lennard L, Van Loon JA, Lilleyman JS, Weinshilboum RM (1987) Thiopurine pharmacogenetics in leukemia: correlation of erythrocyte thiopurine methyltransferase activity and 6-thioguanine nucleotide concentrations. Clin Pharmacol Ther 41(1):18–25
Relling MV, Hancock ML, Rivera GK, Sandlund JT, Ribeiro RC, Krynetski EY, Pui CH, Evans WE (1999) Mercaptopurine therapy intolerance and heterozygosity at the thiopurine S-methyltransferase gene locus. J Natl Cancer Inst 91(23):2001–2008
Szumlanski CL, Honchel R, Scott MC, Weinshilboum RM (1992) Human liver thiopurine methyltransferase pharmacogenetics: biochemical properties, liver-erythrocyte correlation and presence of isozymes. Pharmacogenetics 2(4):148–159
Wu AH (2011) Drug metabolizing enzyme activities versus genetic variances for drug of clinical pharmacogenomic relevance. Clin Proteomics 8(1):12. doi:10.1186/1559-0275-8-12
McLeod HL, Lin JS, Scott EP, Pui CH, Evans WE (1994) Thiopurine methyltransferase activity in American white subjects and black subjects. Clin Pharmacol Ther 55(1):15–20
Weinshilboum RM, Sladek SL (1980) Mercaptopurine pharmacogenetics: monogenic inheritance of erythrocyte thiopurine methyltransferase activity. Am J Hum Genet 32(5):651–662
Krynetski EY, Schuetz JD, Galpin AJ, Pui CH, Relling MV, Evans WE (1995) A single point mutation leading to loss of catalytic activity in human thiopurine S-methyltransferase. Proc Natl Acad Sci U S A 92(4):949–953
Tai HL, Fessing MY, Bonten EJ, Yanishevsky Y, d’Azzo A, Krynetski EY, Evans WE (1999) Enhanced proteasomal degradation of mutant human thiopurine S-methyltransferase (TPMT) in mammalian cells: mechanism for TPMT protein deficiency inherited by TPMT*2, TPMT*3A, TPMT*3B or TPMT*3C. Pharmacogenetics 9(5):641–650
Tai HL, Krynetski EY, Schuetz EG, Yanishevski Y, Evans WE (1997) Enhanced proteolysis of thiopurine S-methyltransferase (TPMT) encoded by mutant alleles in humans (TPMT*3A, TPMT*2): mechanisms for the genetic polymorphism of TPMT activity. Proc Natl Acad Sci U S A 94(12):6444–6449
Colombel JF, Ferrari N, Debuysere H, Marteau P, Gendre JP, Bonaz B, Soulé JC, Modigliani R, Touze Y, Catala P, Libersa C, Broly F (2000) Genotypic analysis of thiopurine S-methyltransferase in patients with Crohn’s disease and severe myelosuppression during azathioprine therapy. Gastroenterology 118(6):1025–1030
Dong X-W, Zheng Q, Zhu M-M, Tong J-L, Ran Z-H (2010) Thiopurine S-methyltransferase polymorphisms and thiopurine toxicity in treatment of inflammatory bowel disease. World J Gastroenterol 16(25):3187–3195. doi:10.3748/wjg.v16.i25.3187
Gisbert JP, Gomollón F (2008) Thiopurine-induced myelotoxicity in patients with inflammatory bowel disease: a review. Am J Gastroenterol 103(7):1783–1800. doi:10.1111/j.1572-0241.2008.01848.x
Higgs JE, Payne K, Roberts C, Newman WG (2010) Are patients with intermediate TPMT activity at increased risk of myelosuppression when taking thiopurine medications? Pharmacogenomics 11(2):177–188. doi:10.2217/pgs.09.155
Meggitt SJ, Anstey AV, Mohd Mustapa MF, Reynolds NJ, Wakelin S (2011) British Association of Dermatologists’ guidelines for the safe and effective prescribing of azathioprine 2011. Br J Dermatol 165(4):711–734. doi:10.1111/j.1365-2133.2011.10575.x
Chakravarty K, McDonald H, Pullar T, Taggart A, Chalmers R, Oliver S, Mooney J, Somerville M, Bosworth A, Kennedy T, British Society for Rheumatology BHPiRSG, Audit Working G, British Association of D (2008) BSR/BHPR guideline for disease-modifying anti-rheumatic drug (DMARD) therapy in consultation with the British Association of Dermatologists. Rheumatology (Oxford) 47(6):924–925. doi:10.1093/rheumatology/kel216a
Fargher EA, Tricker K, Newman W, Elliott R, Roberts SA, Shaffer JL, Bruce I, Payne K (2007) Current use of pharmacogenetic testing: a national survey of thiopurine methyltransferase testing prior to azathioprine prescription. J Clin Pharm Ther 32(2):187–195. doi:10.1111/j.1365-2710.2007.00805.x
Newman WG, Payne K, Tricker K, Roberts SA, Fargher E, Pushpakom S, Alder JE, Sidgwick GP, Payne D, Elliott RA, Heise M, Elles R, Ramsden SC, Andrews J, Houston JB, Qasim F, Shaffer J, Griffiths CEM, Ray DW, Bruce I, Ollier WER, team Tsr (2011) A pragmatic randomized controlled trial of thiopurine methyltransferase genotyping prior to azathioprine treatment: the TARGET study. Pharmacogenomics 12(6):815–826. doi:10.2217/pgs.11.32
Stocco G, Cheok MH, Crews KR, Dervieux T, French D, Pei D, Yang W, Cheng C, Pui CH, Relling MV, Evans WE (2009) Genetic polymorphism of inosine triphosphate pyrophosphatase is a determinant of mercaptopurine metabolism and toxicity during treatment for acute lymphoblastic leukemia. Clin Pharmacol Ther 85(2):164–172. doi:10.1038/clpt.2008.154
Lennard L (2014) Implementation of TPMT testing. Br J Clin Pharmacol 77(4):704–714. doi:10.1111/bcp.12226
Schaeffeler E, Fischer C, Brockmeier D, Wernet D, Moerike K, Eichelbaum M, Zanger UM, Schwab M (2004) Comprehensive analysis of thiopurine S-methyltransferase phenotype-genotype correlation in a large population of German-Caucasians and identification of novel TPMT variants. Pharmacogenetics 14(7):407–417
Hindorf U, Appell ML (2012) Genotyping should be considered the primary choice for pre-treatment evaluation of thiopurine methyltransferase function. J Crohns Colitis 6(6):655–659. doi:10.1016/j.crohns.2011.11.014
Beswick L, Friedman AB, Sparrow MP (2014) The role of thiopurine metabolite monitoring in inflammatory bowel disease. Expert Rev Gastroenterol Hepatol 8(4):383–392. doi:10.1586/17474124.2014.894878
Postmus I, Verschuren JJW, de Craen AJM, Slagboom PE, Westendorp RGJ, Jukema JW, Trompet S (2012) Pharmacogenetics of statins: achievements, whole-genome analyses and future perspectives. Pharmacogenomics 13(7):831–840. doi:10.2217/pgs.12.25
Goldstein JL, Brown MS (2009) The LDL receptor. Arterioscler Thromb Vasc Biol 29(4):431–438. doi:10.1161/atvbaha.108.179564
Baigent C, Keech A, Kearney PM, Blackwell L, Buck G, Pollicino C, Kirby A, Sourjina T, Peto R, Collins R, Simes R (2005) Efficacy and safety of cholesterol-lowering treatment: prospective meta-analysis of data from 90,056 participants in 14 randomised trials of statins. The Lancet 366(9493):1267–1278. doi:10.1016/s0140-6736(05)67394-1
Wilke RA, Ramsey LB, Johnson SG, Maxwell WD, McLeod HL, Voora D, Krauss RM, Roden DM, Feng Q, Cooper-Dehoff RM, Gong L, Klein TE, Wadelius M, Niemi M (2012) The clinical pharmacogenomics implementation consortium: CPIC guideline for SLCO1B1 and simvastatin-induced myopathy. Clin Pharmacol Ther 92(1):112–117. doi:10.1038/clpt.2012.57
Wei C-Y, Lee M-TM, Chen Y-T (2012) Pharmacogenomics of adverse drug reactions: implementing personalized medicine. Hum Mol Genet 21(R1):R58–65. doi:10.1093/hmg/dds341
Hilgendorf C, Ahlin G, Seithel A, Artursson P, Ungell A-L, Karlsson J (2007) Expression of thirty-six drug transporter genes in human intestine, liver, kidney, and organotypic cell lines. Drug Metab Dispos 35(8):1333–1340. doi:10.1124/dmd.107.014902
Tirona RG, Leake BF, Merino G, Kim RB (2001) Polymorphisms in OATP-C: identification of multiple allelic variants associated with altered transport activity among European- and African-Americans. J Biol Chem 276(38):35669–35675. doi:10.1074/jbc.M103792200
Karlgren M, Vildhede A, Norinder U, Wisniewski JR, Kimoto E, Lai Y, Haglund U, Artursson P (2012) Classification of inhibitors of hepatic organic anion transporting polypeptides (OATPs): influence of protein expression on drug-drug interactions. J Med Chem 55(10):4740–4763. doi:10.1021/jm300212s
Laaksonen R (2006) On the mechanisms of statin-induced myopathy. Clin Pharmacol Ther 79(6):529–531. doi:10.1016/j.clpt.2006.02.013
Brunham LR, Lansberg PJ, Zhang L, Miao F, Carter C, Hovingh GK, Visscher H, Jukema JW, Stalenhoef AF, Ross CJD, Carleton BC, Kastelein JJP, Hayden MR (2012) Differential effect of the rs4149056 variant in SLCO1B1 on myopathy associated with simvastatin and atorvastatin. Pharmacogenomics J 12(3):233–237. doi:10.1038/tpj.2010.92
Carr DF, O'Meara H, Jorgensen AL, Campbell J, Hobbs M, McCann G, van Staa T, Pirmohamed M (2013) SLCO1B1 genetic variant associated with statin-induced myopathy: a proof-of-concept study using the clinical practice research datalink. Clin Pharmacol Ther 94(6):695–701. doi:10.1038/clpt.2013.161
MRC/BHF Heart Protection Study of cholesterol lowering with simvastatin in 20,536 high-risk individuals: a randomised placebo-controlled trial (2002). Lancet 360(9326):7–22. doi:10.1016/s0140-6736(02)09327-3
Voora D, Shah SH, Spasojevic I, Ali S, Reed CR, Salisbury BA, Ginsburg GS (2009) The SLCO1B1*5 genetic variant is associated with statin-induced side effects. J Am Coll Cardiol 54(17):1609–1616. doi:10.1016/j.jacc.2009.04.053
Donnelly LA, Doney ASF, Tavendale R, Lang CC, Pearson ER, Colhoun HM, McCarthy MI, Hattersley AT, Morris AD, Palmer CNA (2011) Common non-synonymous substitutions in SLCO1B1 predispose to statin intolerance in routinely treated individuals with type 2 diabetes: a Go-DARTS study. Clin Pharmacol Ther 89(2):210–216. doi:10.1038/clpt.2010.255
Puccetti L, Ciani F, Auteri A (2010) Genetic involvement in statins induced myopathy. Preliminary data from an observational case-control study. Atherosclerosis 211(1):28–29. doi:10.1016/j.atherosclerosis.2010.02.026
Niemi M (2010) Transporter pharmacogenetics and statin toxicity. Clin Pharmacol Ther 87(1):130–133. doi:10.1038/clpt.2009.197
Pasanen MK, Fredrikson H, Neuvonen PJ, Niemi M (2007) Different effects of SLCO1B1 polymorphism on the pharmacokinetics of atorvastatin and rosuvastatin. Clin Pharmacol Ther 82(6):726–733. doi:10.1038/sj.clpt.6100220
Pasanen MK, Neuvonen M, Neuvonen PJ, Niemi M (2006) SLCO1B1 polymorphism markedly affects the pharmacokinetics of simvastatin acid. Pharmacogenet Genomics 16(12):873–879. doi:10.1097/01.fpc.0000230416.82349.90
Research C for DE and Drug Safety and Availability—FDA Drug Safety Communication: new restrictions, contraindications, and dose limitations for Zocor (simvastatin) to reduce the risk of muscle injury. http://www.fda.gov/Drugs/DrugSafety/ucm256581.htm. Accessed 30 Dec 2012
Pirmohamed M (2010) Acceptance of biomarker-based tests for application in clinical practice: criteria and obstacles. Clin Pharmacol Ther 88(6):862–866. doi:10.1038/clpt.2010.245
Pirmohamed M, Aithal GP, Behr E, Daly A, Roden D (2011) The phenotype standardization project: improving pharmacogenetic studies of serious adverse drug reactions. Clin Pharmacol Ther 89(6):784–785. doi:10.1038/clpt.2011.30
Abecasis GR, Auton A, Brooks LD, DePristo MA, Durbin RM, Handsaker RE, Kang HM, Marth GT, McVean GA (2012) An integrated map of genetic variation from 1,092 human genomes. Nature 491(7422):56–65. doi:10.1038/nature11632
Voora D, Ginsburg GS (2012) Clinical application of cardiovascular pharmacogenetics. J Am Coll Cardiol 60(1):9–20. doi:10.1016/j.jacc.2012.01.067
Payne K, Newman W, Fargher E, Tricker K, Bruce IN, Ollier WER (2007) TPMT testing in rheumatology: any better than routine monitoring? Rheumatology (Oxford) 46(5):727–729. doi:10.1093/rheumatology/kel427
Manolio TA, Chisholm RL, Ozenberger B, Roden DM, Williams MS, Wilson R, Bick D, Bottinger EP, Brilliant MH, Eng C, Frazer KA, Korf B, Ledbetter DH, Lupski JR, Marsh C, Mrazek D, Murray MF, O'Donnell PH, Rader DJ, Relling MV, Shuldiner AR, Valle D, Weinshilboum R, Green ED, Ginsburg GS (2013) Implementing genomic medicine in the clinic: the future is here. Genet Med 15(4):258–267. doi:10.1038/gim.2012.157
Hudson J (2012) Pharmacogenomics: where does Britain stand? Pharmacogenomics 13(1):1–3. doi:10.2217/pgs.11.159
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Turner, R., Pirmohamed, M. (2015). Pharmacogenetics of Adverse Drug Reactions. In: Grech, G., Grossman, I. (eds) Preventive and Predictive Genetics: Towards Personalised Medicine. Advances in Predictive, Preventive and Personalised Medicine, vol 9. Springer, Cham. https://doi.org/10.1007/978-3-319-15344-5_6
Download citation
DOI: https://doi.org/10.1007/978-3-319-15344-5_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-15343-8
Online ISBN: 978-3-319-15344-5
eBook Packages: MedicineMedicine (R0)