Abstract
To generate gene deletion mutants in Aspergillus niger, we combined the use of nonhomologous end-joining (NHEJ) mutants (ku70 mutant) and the split marker approach. The combination of both tools resulted in efficient PCR amplification because of the reduced length of the PCR fragments and efficient homologous recombination frequencies. A set of five selection markers, two dominant selection markers (hph; hygromycin B resistance and BLE; phleomycin resistance) and three auxotrophic markers (pyrG, argB, and nicB) were successfully used in a split marker approach to obtain amyR knock outs with high efficiency. AmyR encodes a transcription factor that is required for the expression of starch degrading enzymes and disruption of amyR results in the inability to grow on starch. The strategy to generate the gene deletion constructs is such that with one set of four gene-specific primers, a gene deletion mutant can be generated with either one of the five selection markers. The strategy is based on fusion PCR and omits the necessity for cloning the disruption cassettes. This accelerates the process of generating gene deletion cassettes which can now be accomplished within eight hours. The split marker approach can also be used to make gene deletions in a wild-type background instead of a Δku70 background. In this chapter, we present protocols and considerations that we used to generate gene knock out constructs by fusion PCR and to obtain and verify gene knock outs with any of the five marker genes using the split marker approach. The method is easily transferable to other filamentous fungi.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- ku70
- Gene targeting
- Homologous recombination
- Nicotinamide auxotrophy
- Uracil auxotrophy
- Arginin auxotrophy
- Phleomycin resistance
- Hygromycin resistance
1 Introduction
Targeted deletion of a Gene of Interest (GOI) is a powerful method to address gene functions and requires a double crossover homologous recombination (HR) event to exchange the GOI with a selection marker. In filamentous fungi, DNA integrates preferably via the nonhomologous end joining (NHEJ) pathway, which results in low frequencies of HR and consequently, in low efficiencies in obtaining gene deletion mutants. A successful approach to obtain gene deletion mutants with high efficiency has been the construction of mutants in the NHEJ-pathway, first described for Neurospora crassa (Ninomiya et al. 2004), and followed up by numerous other filamentous fungi including Aspergillus niger (Meyer et al. 2007; Carvalho et al. 2010; Arentshorst et al. 2012). Most often the fungal gene homologous to the gene encoding the Ku70 is used to generate a NHEJ mutant, but also Ku80 and Lig4 homologs have been disrupted to obtain NHEJ-deficient mutants (for reviews see Meyer 2008, Kuck and Hoff 2010 and references therein). The use of NHEJ mutants has greatly reduced time and effort to generate gene deletion mutants. The construction of a gene deletion cassette is also an important and time consuming factor. In principle, a gene deletion construct consists of a selection marker, flanked by upstream (5′) and downstream (3′) sequences of the GOI. Several approaches to generate gene deletion cassettes include traditional restriction enzyme and ligation-based cloning, GATEWAY cloning, fusion PCR, or in vivo assembly either in Escherichia coli or Saccharomyces cerevisiae.
An additional tool for improving gene targeting efficiencies is making use of the split marker technology. In this approach the gene deletion construct is split in two parts and each part contains the flanking region and a truncated form of the selection marker (Fairhead et al. 1996, Nielsen et al. 2006, Goswami 2012).
For the selection of transformants in A. niger (and also other filamentous fungi) the number of available markers is limited. Dominant selection markers for A. niger include markers giving resistance to hygromycin (pAN7.1) (Punt et al. 1987) or phleomycin (pAN8.1) (Punt and van den Hondel 1992), which are well established and commonly used. The uridine and arginine markers (pyrG (An12g03570) and argB (An14g03400), respectively), have been described earlier and are used in this study (Buxton et al. 1985; Van Hartingsveldt et al. 1987; Lenouvel et al. 2002). The pyrG gene encodes for the enzyme orotidine-5′-phosphate-decarboxylase and is required for uracil biosynthesis. The argB gene, encoding for an ornithine carbamoyltransferase, is essential for arginine biosynthesis. In addition, a new auxotrophic mutant which requires nicotinamide for growth based on the nicB gene (An11g10910) was made. The A. niger nicB gene encodes a nicotinate–nucleotide pyrophosphorylase. Identification and the construction of a gene deletion cassette to disrupt nicB is based on a previous work by Verdoes et al. 1994, and will be described elsewhere in detail (Niu et al. manuscript in preparation). The ΔnicB strain is auxotrophic for nicotinamide and needs supplementation of nicotinamide to be able to grow. In addition, we reconstructed an argB deletion mutant (Niu et al. manuscript in preparation) to have all auxotrophic strains in the same strain background (Table 25.1).
Growth of all three auxotrophic strains (pyrG −, argB −, and nicB −) on minimal medium requires the addition of uridine, L-arginine or nicotinamide, respectively,Footnote 1 and no growth is observed in the absence of the relevant supplements (data not shown). To minimize HR of the selection markers used in the disruption cassettes, the argB and nicB homologs from Aspergillus nidulans (ANID_04409.1 and ANID_03431.1 respectively) and the pyrG homolog from Aspergillus oryzae (AO090011000868) were PCR amplified. All genes are able to complement the auxotrophy of the relevant strain. The hygromycin and phleomycin cassettes also contain only nonhomologous sequences as both resistance genes are flanked by the A. nidulans gpdA promoter (PgpdA) and trpC terminator (TtrpC) (Table 25.2).
2 General Methods
2.1 General Split Marker Approach
The split marker approach used for deleting the GOI is schematically depicted in Fig. 25.1 and consists of two overlapping DNA fragments to disrupt the GOI. The first fragment contains the 5′ flank of the GOI and a truncated version of the selection marker. The second DNA fragment contains an overlapping, but truncated version of the selection marker and the 3′ flank of the GOI. Both fragments are generated by fusion PCR as described below and transformed simultaneously to the recipient A. niger strain. The truncation of the selection marker at either site of the construct results in a nonfunctional marker and as a consequence transformation of only a single split marker fragment does not result in any transformants (data not shown).
Schematic representation of the split marker gene deletion approach. 5′ and 3′ sequences flanking the GOI (5′ and 3′) are transformed simultaneously to the recipient strain. By recombination of the selection maker and homologous integration of the cassette in the genome, a successful gene deletion mutant can be obtained
2.2 Generation of Split Marker Fragments for A. niger Transformation
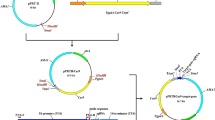
In this section the experimental design for creating the split marker fragments is discussed. The split marker DNA fragments can be obtained in three steps (Fig. 25.2). Each step is described in detail below.
2.2.1 Experimental Design for Amplification of Flanking Regions of the GOI (Step 1)
Once the GOI has been identified, primers need to be designed for making gene deletion cassettes. First, two primers are required for the amplification of the 5′ flank of the GOI. The first primer (P1) is chosen between 700 and 900 bases upstream of the start codon. The reverse primer (P2) is as close to the start codon as possible and contains a 5′-CAATTCCAGCAGCGGCTT-3′ sequence, which is overlapping with all five selection markers and included for the subsequent fusion PCR. Also, two primers are required for the amplification of the 3′ flank of the GOI (P3 and P4). Again, the aim is to generate a 700–900 base pair long flank. In this case, the forward primer (P3) needs a 5′-ACACGGCACAATTATCCATCG-3′ sequence, which is also overlapping with all five selection markers for the subsequent fusion PCR (Step 3).
2.2.2 Experimental Design for Amplification of Suitable Selection Marker (Step 2)
For the amplification of the PCR fragments containing the appropriate selection marker the following plasmids can be used (see also Table 25.2):
The plasmid pAN7.1 (Punt et al. 1987) is used as template to amplify the hygromycin resistance cassette, containing the hph gene from E. coli, coding for hygromycin B phosphotransferase. Expression of the hph gene is driven by the A. nidulans gpdA promoter, and terminated by the A. nidulans trpC terminator. The plasmid pAN8.1 (Punt and van den Hondel 1992) is used as template to amplify the phleomycin resistance cassette, containing the BLE gene from Streptoalloteichus hindustanus, coding for a phleomycin-binding protein. Expression of the BLE gene is also driven by the A. nidulans gpdA promoter and terminated by the A. nidulans trpC terminator. The plasmid pAO4-13 (De Ruiter-Jacobs et al. 1989) is used as template to amplify the A. oryzae pyrG gene (AO090011000868), including promoter and terminator region. The argB gene (ANID_04409.1) and the nicB gene (ANID_03431.1) of A. nidulans, including promoter and terminator region, were amplified using primer pairs argBnidP5f and argBnidP6r or nicBnidP5f and nicBnidP6r, and genomic DNA of A. nidulans strain FGSC A234 (yA2, pabaA1, veA1), obtained from the Fungal Genetics Stock Center, as template. The resulting PCR products were ligated into PCR-cloning vector pJet1.2 (K1231, Thermo Fisher), to give plasmids pJN2.1 and pJN4.1 respectively (Table 25.2). Plasmid pJN2.1 and pJN4.1 can be used to amplify the argB gene or the nicB gene.
We developed a generic split marker approach in such a way that with a single set of four GOI primers, all five different selection markers can be used to generate the deletion cassette. Each primer, used to amplify a specific selection marker (Fig. 25.3, Table 25.3), contains sequences which are overlapping with the GOI primer sequences (see Sect. 25.2.2.1) to create gene deletion mutants with either one of the different selection markers.
2.2.3 Experimental Design for the Generation of Split Marker Fragments (Step 3)
Once both flanks of the GOI (Fig. 25.2, step 1) and the required selection marker (Fig. 25.2, step 2 and Fig. 25.3) have been amplified, the split marker fragments can be obtained by fusion PCR (Fig. 25.2, step 3). Exact details are described in Sect. 25.3.2.3. After column purification, the resulting split marker fragments can directly be used to transform A. niger. Footnote 2
3 Detailed Procedure Description
As proof of principle, the A. niger amyR gene (An04g06910), encoding the amylase transcriptional regulator, has been used. The ΔamyR strain cannot grow on starch, allowing an easy screen for ΔamyR transformants (Petersen et al. 1999). This section contains a detailed description of the whole procedure of deleting amyR, using all five selection markers, illustrated with results of the experiments. Sequences of all primers used are listed in Tables 25.3, 25.4, and 25.5.
3.1 Materials and Reagents
For the medium composition of minimal medium, the preparation of stock solutions for the medium and for a detailed protocol of genomic DNA isolation of A. niger we refer to the Materials and Reagents section in Arentshorst et al. 2012.
-
1.
PCR enzyme (we routinely use Phire Hot start II DNA Polymerase [F-122 L, Thermo Fisher]).
-
2.
dNTPs (1.25 mM): Add 0.25 mL of all 4 dNTPs (dNTP Set 100 mM Solutions (4 × 0.25 mL, R0181, Thermo Fisher)) to 19 mL of MQ, mix well, make aliquots of 0.5 mL, and store at −20 °C.
-
3.
PCR purification Kit (we routinely use Genejet Gel Extraction Kit (K0692, Thermo Fisher), also for PCR purifications).
-
4.
Hygromycin (100 mg/mL): Dissolve 1 g of hygromycin (InvivoGen, ant-hg-10p) in 10 mL of MQ, sterilize by filtration, make aliquots of 500 μL, and store at −20 °C. The final concentration in the medium is 100 μg/mL, except for transformation plates, then use 200 μg/mL.
-
5.
Phleomycin (40 mg/mL), for 10 mL: add 400 mg of phleomycin (InvivoGen, ant-ph-10p) to 8 mL of warm MQ (~60 °C) in a 15 mL tube. When phleomycin is dissolved, add MQ up to 10 mL, and filter sterilize. Make aliquots and store at −20 °C.
-
6.
Uridine (1 M), for 100 mL: add 22.4 g of uridine (Acros, 140775000) to 50 mL of warm MQ (~60 °C) in a 100 mL cylinder. When uridine is dissolved, add MQ up to 100 mL, sterilize by filtration, and store at 4 °C. Final concentration in medium is 10 mM.
-
7.
Arginine (2 %), for 100 mL: add 2 g of L-arginine monohydrochloride (Sigma, A5131) to 50 mL of warm MQ (~60 °C) in a 100 mL cylinder. When arginine is dissolved, add MQ up to 100 mL, sterilize by filtration, and store at 4 °C.
-
8.
Nicotinamide (0.5 %), for 100 mL: add 0.5 g of nicotinamide (Sigma, N0636) to 50 mL of warm MQ (~60 °C) in a 100 mL cylinder. When nicotinamide is dissolved, add MQ up to 100 mL, sterilize by filtration, and store at 4 °C.
-
9.
Transformation media + phleomycin: Prepare MMS and Top agar according to Arentshorst et al. 2012. After autoclaving, and cooling down to 50 °C, add phleomycin to a final concentration of 50 μg/mL, to both the MMS and the Top agar.
-
10.
MM + agar + L-arginine: Prepare 500 mL of MM + agar according to Arentshorst et al. 2012. Add 5 mL of 2 % L-arginine after autoclaving (100× dilution).
-
11.
MM + agar + nicotinamide: Prepare 500 mL of MM + agar according to Arentshorst et al. 2012. Add 0.25 mL of 0.5 % nicotinamide after autoclaving (2,000× dilution).
-
12.
MM + agar + starch: For 500 mL: Dissolve 5 g of starch (soluble, extra pure, Merck, 1.01253) in 450 mL of warm MQ (~60 °C). Add 10 mL of 50× ASP + N, 1 ml of 1 M MgSO4, 50 μL of trace element solution, 15 mg of yeast extract (YE)Footnote 3 (Roth, 2363.2) and 7.5 g agar bact. (Scharlau, 07-004-500), and autoclave.
3.2 Methods
3.2.1 Amplification of the AmyR 5′- and 3′ Flank
-
1.
AmyR primers were designed (Fig. 25.2, Step 1 and Table 25.4), and subsequently used in PCR reactions to amplify both the amyR 5′ flank and 3′ flank.
-
2.
The PCR mix, total volume of 50 μL, contained 1 μL genomic DNA of A. niger wt strain N402 (1 μg/μL), 8 μL dNTPs (1.25 mM), 10 μL 5× Phire buffer, 1 μL Primer F (20 pmol/μL), 1 μL Primer R (20 pmol/μL), 0.5 μL Phire Hot start II DNA Polymerase and 28.5 μL of MQ.
-
3.
PCR was performed under the following conditions: initial denaturation for 5 min at 98 °C, 30 cycles of 5 s at 98 °C, 5 s at 58 °C, and 15 s per 1 kb of template at 72 °C, followed by final extension of 5 min at 72 °C.
-
4.
PCR reactions were analyzed by loading 5 μL PCR reaction on a 1 % agarose gel.
-
5.
After column purification and elution with 30 μL of MQ, DNA concentration for both flanks was ~37 ng/μL.
3.2.2 Amplification of the Selection Markers
-
1.
Primers for all five selection markers were designed (Fig 25.3, Table 25.3) and used for PCR. In these PCR reactions 1 ng of plasmid (pAO4-13, pAN7.1, pAN8.1, pJN2.1, and pJN4.1, respectively) was used as template. For PCR mix and PCR conditions see Sect. 25.3.2.1.
-
2.
After confirmation on agarose gel, selection marker PCR products were column purified, yielding DNA concentrations of ~50 ng/μL. The markers were stored at −20 °C and used repeatedly.
3.2.3 Amplification of the Split Marker Fragments
-
1.
Fusion PCR fragments were amplified according to Fig. 25.2, step 3 (see also Tables 25.5 and 25.6). Both amyR flanks and all selection markers (Sects. 25.3.2.1 and 25.3.2.2) were diluted to 2 ng/μL.
Table 25.6 Overview of templates and primers used in Fusion PCR reactions to obtain split marker fragments -
2.
For each PCR reaction, 2 ng of amyR flank and 2 ng of selection marker PCR were used as template (Table 25.6). For PCR mix and PCR conditions see Sect. 25.3.2.1.
-
3.
Two identical fusion PCR reactions were performed, in order to increase the yield of PCR product.
-
4.
Fusion PCR products were analyzed on agarose gel, followed by column purification. The DNA concentration for all fragments varied between 120 and 160 ng/μL in a total volume of 20 μL (Table 25.6, column DNA Yield).Footnote 4
3.2.4 Transformation of Split Marker Fragments to A. niger Δku70 Strains
-
1.
Split marker fragments were combined and transformed to different A. niger strains (Table 25.6, column Transformed strain), according to Arentshorst et al. 2012. Results of these transformations are shown in Fig. 25.4.
Fig. 25.4 Phenotypic analysis of putative amyR disruptant strains using five different selection markers (hph, hygromycin resistance; BLE, phleomycin resistance; pyrG, uridine requiring; argB, arginine requiring; nicB, nicotinamide requiring). (a) Transformation plates after transforming split marker fragment combinations for each of the five amyR deletion cassettes to the relevant recipient strain (Table 25.6). (b, c) Purified transformants were analyzed for their ability to grow on starch. The inability to grow on starch is indicative for the deletion of the amyR gene
-
2.
As a control, also separate split marker fragments were transformed. None of the separately transformed split marker fragments yielded any transformants (data not shown).
-
3.
Four transformants were purified for each selection marker tested.Footnote 5 For purification protocol, see Arentshorst et al. 2012.
-
4.
After the second purification, all purified transformants were tested for growth on MM + starch (Fig. 25.4). All transformants analyzed showed a ΔamyR phenotype.
-
5.
Purified transformants can be further analyzed by isolating genomic DNA, followed by both Southern blot analysis and diagnostic PCR (Arentshorst et al. 2012).
3.2.5 Transformation of Split Marker Fragments to A. niger wt Strains
For some experimental set-ups, it is preferred to analyze gene deletions in a ku70 wild-type strain. In order to show that the split marker approach also can be applied to a wild-type (ku70 plus) strain, both A. niger strains AB4.1 (Van Hartingsveldt et al. 1987) (pyrG −) and MA169.4 (Δku70, pyrG −) were transformed with ΔamyR::pyrG split marker fragments. After purification and screening on MM + starch, 25 out of 60 AB4.1-transformants (41 %) showed a ΔamyR phenotype.Footnote 6 For MA169.4, 39 out of 40 transformants (98 %) showed a ΔamyR phenotype. This result clearly shows that the split marker approach can also be used to make gene deletions in a wt background instead of a Δku70 background.
Notes
- 1.
The argB and nicB auxotrophic mutants are also pyrG − and therefore the growth medium for these strains needs to be supplemented with uridine.
- 2.
A small sample of PCR fragments is routinely analyzed for purity and size. Optional is to confirm PCR product integrity by restriction analysis or sequencing.
- 3.
YE is added to a final concentration of 0.003 % to stimulate germination of A. niger. On MM + starch without YE, the wt strain also does not germinate very well.
- 4.
The split marker fragments are not purified from gel and template DNA (pyrG, hygB, Ble, argB, and nicB genes, respectively) used for amplification of the split marker might remain present in the next steps. We therefore include control transformations with both split markers separately. As no transformants are obtained in the transformation with only one flank (data not shown), the purification of the split marker fragment is not required, but is optional.
- 5.
Only the sporulating transformants on the phleomycin transformation plate (see Fig. 25.4) can grow on MM + phleomycin. The non-sporulating transformants do not grow, and are probably transient transformants, in which the split marker fragments have not integrated into the genome.
- 6.
The percentages of HR for the amyR gene are very high (41 % for wt, 98 % for Δku70). Usually we find 5–10 % HR for wt, and 80–100 % for Δku70 (Meyer et al. 2007).
References
Arentshorst M, Ram AFJ, Meyer V (2012) Using non-homologous end-joining-deficient strains for functional gene analyses in filamentous fungi. Methods Mol Biol 835:133–150
Bos CJ, Debets AJ, Swart K, Huybers A, Kobus G (1988) Genetic analysis and the construction of master strains for assignment of genes to six linkage groups in Aspergillus niger. Curr Genet 14:437–443
Buxton FP, Gwynne DI, Davies RW (1985) Transformation of Aspergillus niger using the argB gene of Aspergillus nidulans. Gene 37:207–214
Carvalho ND, Arentshorst M, Kwon MJ, Meyer V, Ram AFJ (2010) Expanding the ku70 toolbox for filamentous fungi: establishment of complementation vectors and recipient strains for advanced gene analyses. Appl Microbiol Biotechnol 87:1463–1473
De Ruiter-Jacobs YM, Broekhuijsen M, Unkles SE, Campbell EI, Kinghorn JR, Contreras R, Pouwels PH, van den Hondel CAMJJ (1989) A gene transfer system based on the homologous pyrG gene and efficient expression of bacterial genes in Aspergillus oryzae. Curr Genet 16:159–163
Fairhead C, Llorente B, Denis F, Soler M, Dujon B (1996) New vectors for combinatorial deletions in yeast chromosomes and for gap-repair cloning using ‘split-marker’ recombination. Yeast 12:1439–1457
Goswami RS (2012) Targeted gene replacement in fungi using a split-marker approach. Methods Mol Biol 835:255–269
Kuck U, Hoff B (2010) New tools for the genetic manipulation of filamentous fungi. Appl Microbiol Biotechnol 86:51–62
Lenouvel F, van de Vondervoort PJ, Visser J (2002) Disruption of the Aspergillus niger argB gene: a tool for transformation. Curr Genet 41:425–432
Meyer V (2008) Genetic engineering of filamentous fungi—progress, obstacles and future trends. Biotechnol Adv 26:177–185
Meyer V, Arentshorst M, El-Ghezal A, Drews AC, Kooistra R, van den Hondel CAMJJ, Ram AFJ (2007) Highly efficient gene targeting in the Aspergillus niger kusA mutant. J Biotechnol 128:770–775
Nielsen ML, Albertsen L, Lettier G, Nielsen JB, Mortensen UH (2006) Efficient PCR-based gene targeting with a recyclable marker for Aspergillus nidulans. Fungal Genet Biol 43:54–64
Ninomiya Y, Suzuki K, Ishii C, Inoue H (2004) Highly efficient gene replacements in Neurospora strains deficient for nonhomologous end-joining. Proc Natl Acad Sci U S A 101:12248–12253
Petersen KL, Lehmbeck J, Christensen T (1999) A new transcriptional activator for amylase genes in Aspergillus. Mol Gen Genet 262:668–676
Punt PJ, van den Hondel CAMJJ (1992) Transformation of filamentous fungi based on hygromycin B and phleomycin resistance markers. Methods Enzymol 216:447–457
Punt PJ, Oliver RP, Dingemanse MA, Pouwels PH, van den Hondel CAMJJ (1987) Transformation of Aspergillus based on the hygromycin B resistance marker from Escherichia coli. Gene 56:117–124
Van Hartingsveldt W, Mattern IE, van Zeijl CM, Pouwels PH, van den Hondel CAMJJ (1987) Development of a homologous transformation system for Aspergillus niger based on the pyrG gene. Mol Gen Genet 206:71–75
Verdoes JC, Punt PJ, van der Berg P, Debets F, Stouthamer AH, van den Hondel CAMJJ (1994) Characterization of an efficient gene cloning strategy for Aspergillus niger based on an autonomously replicating plasmid: cloning of the nicB gene of A. niger. Gene 146:159–165
Acknowledgments
Jing Niu was supported by a grant from the China Scholarship Council. The research group of A.F.J. Ram is part of the Kluyver Centre for Genomics of Industrial Fermentation which is supported by the Netherlands Genomics Initiative.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Arentshorst, M., Niu, J., Ram, A.F.J. (2015). Efficient Generation of Aspergillus niger Knock Out Strains by Combining NHEJ Mutants and a Split Marker Approach. In: van den Berg, M., Maruthachalam, K. (eds) Genetic Transformation Systems in Fungi, Volume 1. Fungal Biology. Springer, Cham. https://doi.org/10.1007/978-3-319-10142-2_25
Download citation
DOI: https://doi.org/10.1007/978-3-319-10142-2_25
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-10141-5
Online ISBN: 978-3-319-10142-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)